Probing the Intracellular Bio-Nano Interface in Different Cell Lines with Gold Nanostars
Abstract
1. Introduction
2. Materials and Methods
2.1. Gold Nanostar Synthesis
2.2. Cell Cultures and Incubation with Nanostars
2.3. SERS Experiments
2.4. Cryo Soft X-ray Nanotomography
3. Results and Discussion
3.1. Optical and Morphological Properties of Gold Nanostars
3.2. Nanostar–Biomolecule Interactions Differ in Different Cell Lines
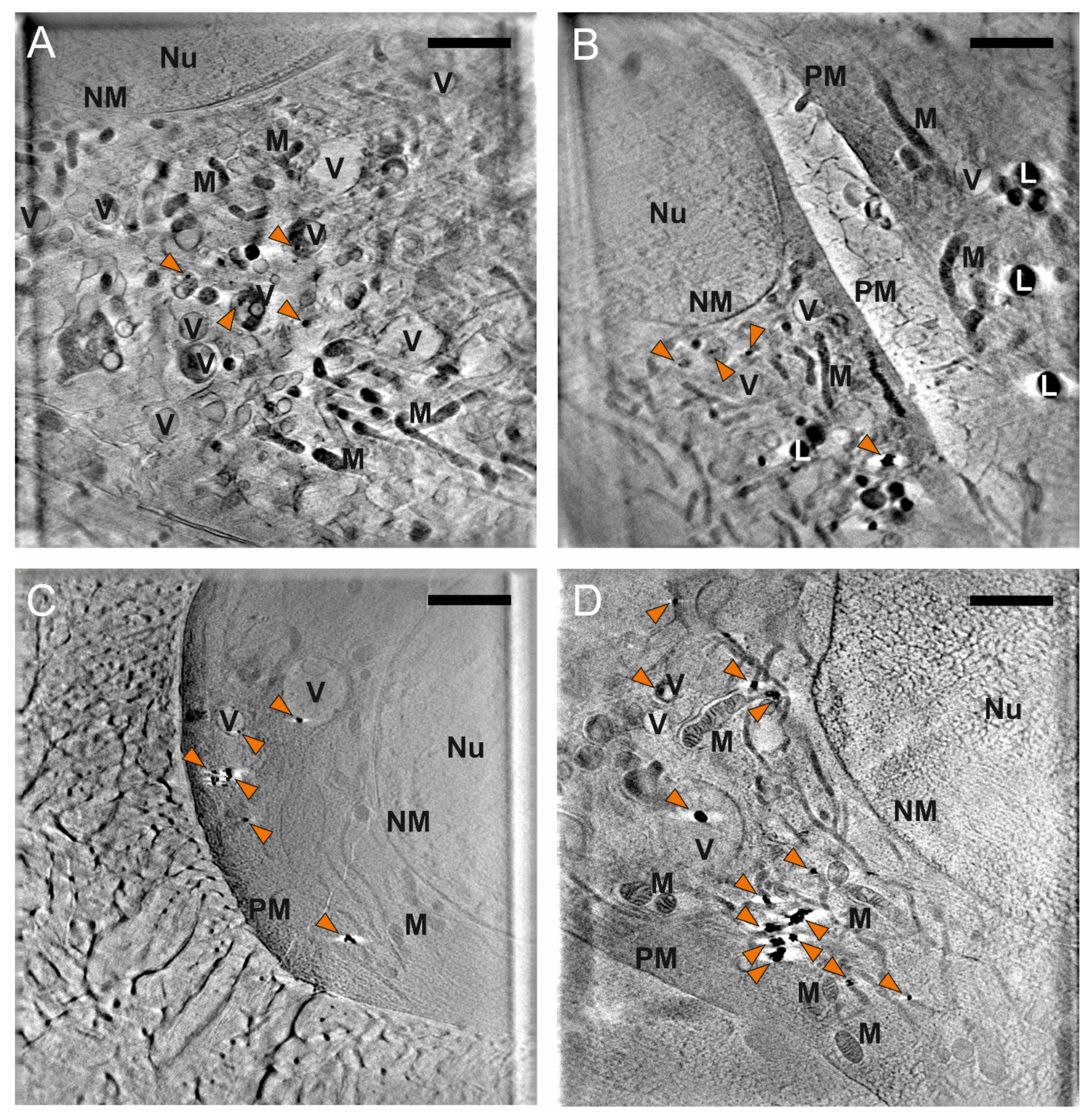
3.3. Nanostar Processing and Distribution in Cellular Compartments
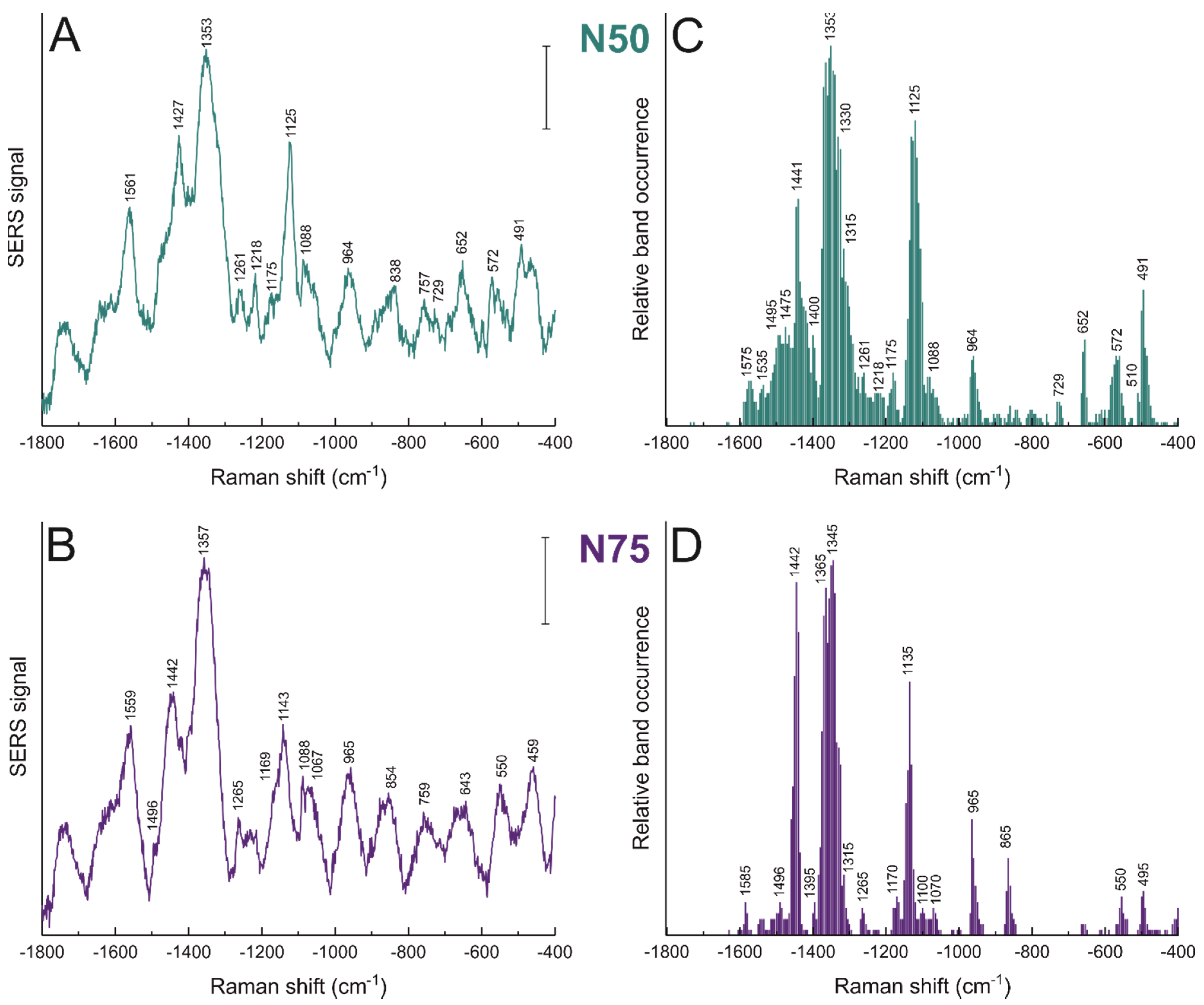
3.4. Morphology Dictates Nanostar–Biomolecule Interaction
3.5. Nanostar–Biomolecule Interaction Variability with Time
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Khoury, C.G.; Vo-Dinh, T. Gold nanostars for surface-enhanced Raman scattering: Synthesis, characterization and optimization. J. Phys. Chem. C 2008, 112, 18849–18859. [Google Scholar] [CrossRef]
- Senthil Kumar, P.; Pastoriza-Santos, I.; Rodriguez-Gonzalez, B.; Javier Garcia de Abajo, F.; Liz-Marzan, L.M. High-yield synthesis and optical response of gold nanostars. Nanotechnology 2008, 19, 015606. [Google Scholar] [CrossRef] [PubMed]
- Perevedentseva, E.; Ali, N.; Lin, Y.C.; Karmenyan, A.; Chang, C.C.; Bibikova, O.; Skovorodkin, I.; Prunskaite-Hyyrylainen, R.; Vainio, S.J.; Kinnunen, M.; et al. Au nanostar nanoparticle as a bio-imaging agent and its detection and visualization in biosystems. Biomed. Opt. Express 2020, 11, 5872–5885. [Google Scholar] [CrossRef]
- Ou, Y.C.; Wen, X.; Johnson, C.A.; Shae, D.; Ayala, O.D.; Webb, J.A.; Lin, E.C.; DeLapp, R.C.; Boyd, K.L.; Richmond, A.; et al. Multimodal multiplexed immunoimaging with nanostars to detect multiple immunomarkers and monitor response to immunotherapies. ACS Nano 2020, 14, 651–663. [Google Scholar] [CrossRef]
- Phiri, M.M.; Mulder, D.W.; Vorster, B.C. Seedless gold nanostars with seed-like advantages for biosensing applications. Roy. Soc. Open Sci. 2019, 6, 181971. [Google Scholar] [CrossRef] [PubMed]
- Jana, D.; Matti, C.; He, J.; Sagle, L. Capping agent-free gold nanostars show greatly increased versatility and sensitivity for biosensing. Anal. Chem. 2015, 87, 3964–3972. [Google Scholar] [CrossRef]
- Yuan, H.; Gomez, J.A.; Chien, J.S.; Zhang, L.; Wilson, C.M.; Li, S.; Fales, A.M.; Liu, Y.; Grant, G.A.; Mirotsou, M.; et al. Tracking mesenchymal stromal cells using an ultra-bright TAT-functionalized plasmonic-active nanoplatform. J. Biophotonics 2016, 9, 406–413. [Google Scholar] [CrossRef] [PubMed]
- Ahijado-Guzmán, R.; Sánchez-Arribas, N.; Martínez-Negro, M.; González-Rubio, G.; Santiago-Varela, M.; Pardo, M.; Piñeiro, A.; López-Montero, I.; Junquera, E.; Guerrero-Martínez, A. Intercellular trafficking of gold nanostars in uveal melanoma cells for plasmonic photothermal therapy. Nanomaterials 2020, 10, 590. [Google Scholar] [CrossRef]
- Li, Y.; Wang, X.; Yang, D.; Hu, P.; Gao, L.; Chen, D.; Qiao, Y.; Wu, Y.; Jiang, X.; Li, G. Polydopamine-coated gold nanostars for near-infrared cancer photothermal therapy by multiple pathways. J. Mater. Sci. 2019, 54, 12036–12048. [Google Scholar] [CrossRef]
- Miao, D.; Yu, Y.; Chen, Y.; Liu, Y.; Su, G. Facile construction of i-motif DNA-conjugated gold nanostars as near-infrared and pH dual-responsive targeted drug delivery systems for combined cancer therapy. Mol. Pharm. 2020, 17, 1127–1138. [Google Scholar] [CrossRef] [PubMed]
- Hao, F.; Nehl, C.L.; Hafner, J.H.; Nordlander, P. Plasmon resonances of a gold nanostar. Nano Lett. 2007, 7, 729–732. [Google Scholar] [CrossRef]
- Tsoulos, T.V.; Fabris, L. Interface and bulk standing waves drive the coupling of plasmonic nanostar antennas. J. Phys. Chem. C 2018, 122, 28949–28957. [Google Scholar] [CrossRef]
- Theodorou, I.G.; Jiang, Q.; Malms, L.; Xie, X.; Coombes, R.C.; Aboagye, E.O.; Porter, A.E.; Ryan, M.P.; Xie, F. Fluorescence enhancement from single gold nanostars: Towards ultra-bright emission in the first and second near-infrared biological windows. Nanoscale 2018, 10, 15854–15864. [Google Scholar] [CrossRef] [PubMed]
- Hrelescu, C.; Sau, T.K.; Rogach, A.L.; Jäckel, F.; Feldmann, J. Single gold nanostars enhance Raman scattering. Appl. Phys. Lett. 2009, 94, 153113. [Google Scholar] [CrossRef]
- De Albuquerque, C.D.L.; Schultz, Z.D. Super-resolution surface-enhanced Raman scattering imaging of single particles in cells. Anal. Chem. 2020, 92, 9389–9398. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.; Sheng, Z.; Li, P.; Wu, M.; Zhang, N.; Yu, X.F.; Wang, Y.; Hu, D.; Zheng, H.; Wang, G.P. Indocyanine green-loaded gold nanostars for sensitive SERS imaging and subcellular monitoring of photothermal therapy. Nanoscale 2017, 9, 11888–11901. [Google Scholar] [CrossRef] [PubMed]
- Oliveira, M.J.; de Almeida, M.P.; Nunes, D.; Fortunato, E.; Martins, R.; Pereira, E.; Byrne, H.J.; Aguas, H.; Franco, R. Design and simple assembly of gold nanostar bioconjugates for surface-enhanced Raman spectroscopy immunoassays. Nanomaterials 2019, 9, 1561. [Google Scholar] [CrossRef]
- Pei, Y.; Wang, Z.; Zong, S.; Cui, Y. Highly sensitive SERS-based immunoassay with simultaneous utilization of self-assembled substrates of gold nanostars and aggregates of gold nanostars. J. Mater. Chem. B 2013, 1, 3992–3998. [Google Scholar] [CrossRef] [PubMed]
- Dardir, K.; Wang, H.; Martin, B.E.; Atzampou, M.; Brooke, C.B.; Fabris, L. SERS nanoprobe for intracellular monitoring of viral mutations. J. Phys. Chem. C 2020, 124, 3211–3217. [Google Scholar] [CrossRef]
- Nguyen, T.D.; Song, M.S.; Ly, N.H.; Lee, S.Y.; Joo, S.W. Nanostars on nanopipette tips: A Raman probe for quantifying oxygen levels in hypoxic single cells and tumours. Angew. Chem. Int. Ed. 2019, 58, 2710–2714. [Google Scholar] [CrossRef]
- He, S.; Kyaw, Y.M.E.; Tan, E.K.M.; Bekale, L.; Kang, M.W.C.; Kim, S.S.; Tan, I.; Lam, K.P.; Kah, J.C.Y. Quantitative and label-free detection of protein kinase a activity based on surface-enhanced Raman spectroscopy with gold nanostars. Anal. Chem. 2018, 90, 6071–6080. [Google Scholar] [CrossRef]
- Childs, A.; Vinogradova, E.; Ruiz-Zepeda, F.; Velazquez-Salazar, J.J.; Jose-Yacaman, M. Biocompatible gold/silver nanostars for surface-enhanced Raman scattering. J. Raman Spectrosc. 2016, 47, 651–655. [Google Scholar] [CrossRef]
- Spedalieri, C.; Szekeres, G.P.; Werner, S.; Guttmann, P.; Kneipp, J. Intracellular optical probing with gold nanostars. Nanoscale 2021, 13, 968–979. [Google Scholar] [CrossRef] [PubMed]
- Chithrani, B.D.; Ghazani, A.A.; Chan, W.C. Determining the size and shape dependence of gold nanoparticle uptake into mammalian cells. Nano Lett. 2006, 6, 662–668. [Google Scholar] [CrossRef]
- Carnovale, C.; Bryant, G.; Shukla, R.; Bansal, V. Identifying trends in gold nanoparticle toxicity and uptake: Size, shape, capping ligand, and biological corona. ACS Omega 2019, 4, 242–256. [Google Scholar] [CrossRef]
- Arnida; Janat-Amsbury, M.M.; Ray, A.; Peterson, C.M.; Ghandehari, H. Geometry and surface characteristics of gold nanoparticles influence their biodistribution and uptake by macrophages. Eur. J. Pharm. Biopharm. 2011, 77, 417–423. [Google Scholar] [CrossRef]
- Xie, X.; Liao, J.; Shao, X.; Li, Q.; Lin, Y. The effect of shape on cellular uptake of gold nanoparticles in the forms of stars, rods, and triangles. Sci. Rep. 2017, 7, 3827. [Google Scholar] [CrossRef]
- Nativo, P.; Prior, I.A.; Brust, M. Uptake and intracellular fate of surface-modified gold nanoparticles. ACS Nano 2008, 2, 1639–1644. [Google Scholar] [CrossRef]
- Ding, L.; Yao, C.; Yin, X.; Li, C.; Huang, Y.; Wu, M.; Wang, B.; Guo, X.; Wang, Y.; Wu, M. Size, shape, and protein corona determine cellular uptake and removal mechanisms of gold nanoparticles. Small 2018, 14, e1801451. [Google Scholar] [CrossRef]
- Walkey, C.D.; Olsen, J.B.; Guo, H.; Emili, A.; Chan, W.C. Nanoparticle size and surface chemistry determine serum protein adsorption and macrophage uptake. J. Am. Chem. Soc. 2012, 134, 2139–2147. [Google Scholar] [CrossRef] [PubMed]
- Szekeres, G.; Werner, S.; Guttmann, P.; Spedalieri, C.; Drescher, D.; Živanović, V.; Montes-Bayón, M.; Bettmer, J.; Kneipp, J. Relating the composition and interface interactions in the hard corona of gold nanoparticles to the induced response mechanisms in living cells. Nanoscale 2020, 12, 17450–17461. [Google Scholar] [CrossRef]
- Szekeres, G.P.; Montes-Bayón, M.; Bettmer, J.; Kneipp, J. Fragmentation of proteins in the corona of gold nanoparticles as observed in live cell surface-enhanced Raman scattering. Anal. Chem. 2020, 92, 8553–8560. [Google Scholar] [CrossRef] [PubMed]
- Drescher, D.; Büchner, T.; Guttmann, P.; Werner, S.; Schneider, G.; Kneipp, J. X-ray tomography shows the varying three-dimensional morphology of gold nanoaggregates in the cellular ultrastructure. Nanoscale Adv. 2019, 1, 2937–2945. [Google Scholar] [CrossRef]
- Huefner, A.; Kuan, W.L.; Muller, K.H.; Skepper, J.N.; Barker, R.A.; Mahajan, S. Characterization and visualization of vesicles in the endo-lysosomal pathway with surface-enhanced Raman spectroscopy and chemometrics. ACS Nano 2016, 10, 307–316. [Google Scholar] [CrossRef] [PubMed]
- Kneipp, J.; Kneipp, H.; McLaughlin, M.; Brown, D.; Kneipp, K. In vivo molecular probing of cellular compartments with gold nanoparticles and nanoaggregates. Nano Lett. 2006, 6, 2225–2231. [Google Scholar] [CrossRef] [PubMed]
- Ando, J.; Fujita, K.; Smith, N.I.; Kawata, S. Dynamic SERS imaging of cellular transport pathways with endocytosed gold nanoparticles. Nano Lett. 2011, 11, 5344–5348. [Google Scholar] [CrossRef] [PubMed]
- Xia, Q.; Huang, J.; Feng, Q.; Chen, X.; Liu, X.; Li, X.; Zhang, T.; Xiao, S.; Li, H.; Zhong, Z.; et al. Size- and cell type-dependent cellular uptake, cytotoxicity and in vivo distribution of gold nanoparticles. Int. J. Nanomed. 2019, 14, 6957–6970. [Google Scholar] [CrossRef]
- Kuhn, D.A.; Vanhecke, D.; Michen, B.; Blank, F.; Gehr, P.; Petri-Fink, A.; Rothen-Rutishauser, B. Different endocytotic uptake mechanisms for nanoparticles in epithelial cells and macrophages. Beilstein J. Nanotechnol. 2014, 5, 1625–1636. [Google Scholar] [CrossRef]
- Sharma, G.; Valenta, D.T.; Altman, Y.; Harvey, S.; Xie, H.; Mitragotri, S.; Smith, J.W. Polymer particle shape independently influences binding and internalization by macrophages. J. Control Release 2010, 147, 408–412. [Google Scholar] [CrossRef]
- Ba Fakih, F.; Shanti, A.; Stefanini, C.; Lee, S. Optimization of gold nanoparticles for efficient delivery of catalase to macrophages for alleviating inflammation. ACS Appl. Nano Mater. 2020, 3, 9510–9519. [Google Scholar] [CrossRef]
- Foroozandeh, P.; Aziz, A.A. Insight into cellular uptake and intracellular trafficking of nanoparticles. Nanoscale Res. Lett. 2018, 13, 339. [Google Scholar] [CrossRef]
- Saverot, S.; Geng, X.; Leng, W.; Vikesland, P.J.; Grove, T.Z.; Bickford, L.R. Facile, tunable, and SERS-enhanced HEPES gold nanostars. RSC Adv. 2016, 6, 29669–29673. [Google Scholar] [CrossRef]
- Schneider, C.A.; Rasband, W.S.; Eliceiri, K.W. NIH image to ImageJ: 25 years of image analysis. Nat. Methods 2012, 9, 671–675. [Google Scholar] [CrossRef]
- Schneider, G.; Guttmann, P.; Heim, S.; Rehbein, S.; Mueller, F.; Nagashima, K.; Heymann, J.B.; Muller, W.G.; McNally, J.G. Three-dimensional cellular ultrastructure resolved by X-ray microscopy. Nat. Methods 2010, 7, 985–987. [Google Scholar] [CrossRef] [PubMed]
- Guttmann, P.; Werner, S.; Siewert, F.; Sokolov, A.; Schmidt, J.-S.; Mast, M.; Brzhezinskaya, M.; Jung, C.; Follath, R.; Schneider, G. The new HZB X-ray microscopy beamline U41-PGM1-XM at BESSY II. Microsc. Microanal. 2018, 24, 206–207. [Google Scholar] [CrossRef]
- Plascencia-Villa, G.; Bahena, D.; Rodriguez, A.R.; Ponce, A.; Jose-Yacaman, M. Advanced microscopy of star-shaped gold nanoparticles and their adsorption-uptake by macrophages. Metallomics 2013, 5, 242–250. [Google Scholar] [CrossRef] [PubMed]
- Favi, P.M.; Gao, M.; Johana Sepulveda Arango, L.; Ospina, S.P.; Morales, M.; Pavon, J.J.; Webster, T.J. Shape and surface effects on the cytotoxicity of nanoparticles: Gold nanospheres versus gold nanostars. J. Biomed. Mater. Res. Part A 2015, 103, 3449–3462. [Google Scholar] [CrossRef] [PubMed]
- Habib, A.; Tabata, M.; Wu, Y.G. Formation of gold nanoparticles by Good’s buffers. Bull. Chem. Soc. Jpn. 2005, 78, 262–269. [Google Scholar] [CrossRef]
- Webb, J.A.; Erwin, W.R.; Zarick, H.F.; Aufrecht, J.; Manning, H.W.; Lang, M.J.; Pint, C.L.; Bardhan, R. Geometry-dependent plasmonic tunability and photothermal characteristics of multibranched gold nanoantennas. J. Phys. Chem. C 2014, 118, 3696–3707. [Google Scholar] [CrossRef]
- Xie, J.; Lee, J.Y.; Wang, D.I.C. Seedless, surfactantless, high-yield synthesis of branched gold nanocrystals in HEPES buffer solution. Chem. Mater. 2007, 19, 2823–2830. [Google Scholar] [CrossRef]
- Chandra, K.; Culver, K.S.B.; Werner, S.E.; Lee, R.C.; Odom, T.W. Manipulating the anisotropic structure of gold nanostars using Good’s buffers. Chem. Mater. 2016, 28, 6763–6769. [Google Scholar] [CrossRef]
- Buchner, T.; Drescher, D.; Traub, H.; Schrade, P.; Bachmann, S.; Jakubowski, N.; Kneipp, J. Relating surface-enhanced Raman scattering signals of cells to gold nanoparticle aggregation as determined by LA-ICP-MS micromapping. Anal. Bioanal. Chem. 2014, 406, 7003–7014. [Google Scholar] [CrossRef] [PubMed]
- Talari, A.C.S.; Movasaghi, Z.; Rehman, S.; Rehman, I.U. Raman spectroscopy of biological tissues. Appl. Spectrosc. Rev. 2014, 50, 46–111. [Google Scholar] [CrossRef]
- Zivanovic, V.; Kochovski, Z.; Arenz, C.; Lu, Y.; Kneipp, J. SERS and cryo-EM directly reveal different liposome structures during interaction with gold nanoparticles. J. Phys. Chem. Lett. 2018, 9, 6767–6772. [Google Scholar] [CrossRef] [PubMed]
- Zivanovic, V.; Seifert, S.; Drescher, D.; Schrade, P.; Werner, S.; Guttmann, P.; Szekeres, G.P.; Bachmann, S.; Schneider, G.; Arenz, C.; et al. Optical nanosensing of lipid accumulation due to enzyme inhibition in live cells. ACS Nano 2019, 13, 9363–9375. [Google Scholar] [CrossRef] [PubMed]
- Rygula, A.; Majzner, K.; Marzec, K.M.; Kaczor, A.; Pilarczyk, M.; Baranska, M. Raman spectroscopy of proteins: A review. J. Raman Spectrosc. 2013, 44, 1061–1076. [Google Scholar] [CrossRef]
- Czamara, K.; Majzner, K.; Pacia, M.Z.; Kochan, K.; Kaczor, A.; Baranska, M. Raman spectroscopy of lipids: A review. J. Raman Spectrosc. 2015, 46, 4–20. [Google Scholar] [CrossRef]
- Szekeres, G.P.; Kneipp, J. Different binding sites of serum albumins in the protein corona of gold nanoparticles. Analyst 2018, 143, 6061–6068. [Google Scholar] [CrossRef]
- Pauwels, A.M.; Trost, M.; Beyaert, R.; Hoffmann, E. Patterns, receptors, and signals: Regulation of phagosome maturation. Trends Immunol. 2017, 38, 407–422. [Google Scholar] [CrossRef]
- Szekeres, G.P.; Fernández-Iglesias, N.; Kneipp, J.; Montes-Bayón, M.; Bettmer, J. Mass spectrometric approach for the analysis of the hard protein corona of nanoparticles in living cells. J. Proteom. 2020, 212, 103582. [Google Scholar] [CrossRef]
- Kapishnikov, S.; Weiner, A.; Shimoni, E.; Guttmann, P.; Schneider, G.; Dahan-Pasternak, N.; Dzikowski, R.; Leiserowitz, L.; Elbaum, M. Oriented nucleation of hemozoin at the digestive vacuole membrane in plasmodium falciparum. Proc. Natl. Acad. Sci. USA 2012, 109, 11188–11193. [Google Scholar] [CrossRef] [PubMed]
- Xu, H.; Aizpurua, J.; Kall, M.; Apell, P. Electromagnetic contributions to single-molecule sensitivity in surface-enhanced Raman scattering. Phys. Rev. E Stat. Phys. Plasmas Fluids Relat. Interdiscip. Top. 2000, 62, 4318–4324. [Google Scholar] [CrossRef] [PubMed]
- Lopes Rodrigues, R.; Xie, F.; Porter, A.E.; Ryan, M.P. Geometry-induced protein reorientation on the spikes of plasmonic gold nanostars. Nanoscale Adv. 2020, 2, 1144–1151. [Google Scholar] [CrossRef]
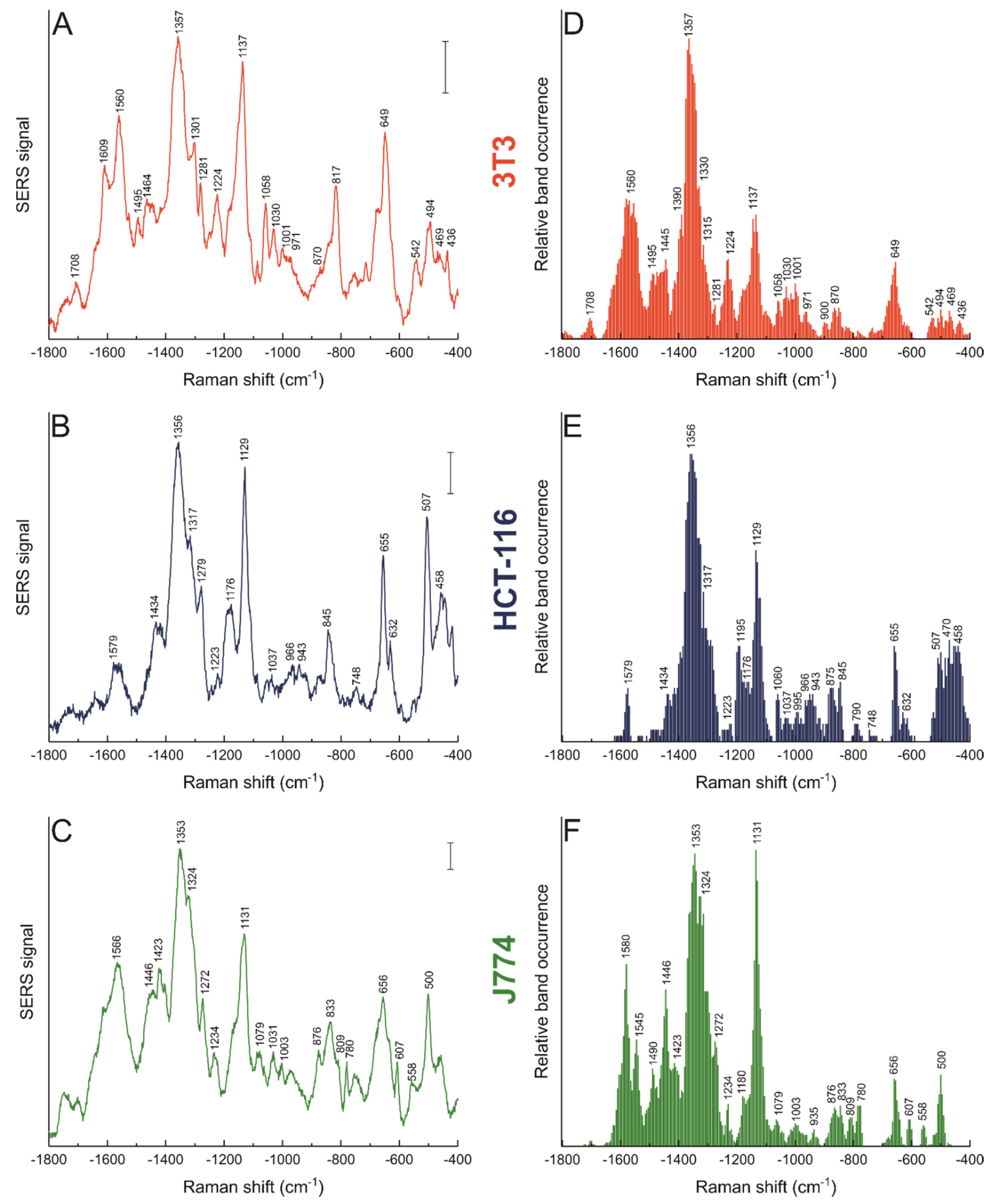
| Raman Shift (cm−1) | Tentative Band Assignment | ||
|---|---|---|---|
| 3T3 | HCT-116 | J774 | |
| 436 | Cholesterol | ||
| 458 | 457 | Protein S-S str, C-S str, Trp ring def | |
| 469 | 470 | C-S str | |
| 494 | 507 | 500 | Protein S-S str |
| 542 | S-S str, Cholesterol | ||
| 558 | S-S str | ||
| 607 | COO- def, ring def, C-H def | ||
| 632 | C-S str | ||
| 649 | 655 | 656 | Cys C-S str, Tyr C-C twist, Phe |
| 752 | 748 | Trp br; Pro; C-S, C-C str; T | |
| 790 | 780 | O-P-O str; C, U, T ring br | |
| 809 | Pro, Tyr, C-C str, lipid O–P–O str | ||
| 817 | O-P-O str; COO- def | ||
| 845 | 833 | Tyr ring br, Phe, Cα-N and C-C str, O-P-O str | |
| 870 | 875 | 876 | Pro, Val C-C str; C-N str |
| 900 | Trp, C-C, C-N str | ||
| 935 | Pro, C-C str | ||
| 943 | C-C str | ||
| 971 | 966 | Lipids C-C str, Pro, Val | |
| 1001 | 995 | 1003 | Phe ring br |
| 1030 | 1037 | 1031 | Phe, C-C str, Tyr ring def |
| 1058 | 1060 | Protein C-C and C-N str | |
| 1079 | Lipid O-P-O str, C-C str; C-N str | ||
| 1137 | 1129 | 1131 | Protein backbone C-C str; C-N str |
| 1176 | 1180 | Tyr C-H bend; Phe | |
| 1195 | Trp, Tyr, Phe; Aromatic C-O and C-N | ||
| 1224 | 1223 | Amide III; Trp ring | |
| 1234 | Amide III, CH2 wag, O-P-O str | ||
| 1281 | 1279 | 1272 | Amide III; CH/CH2/CH3 def |
| 1301 | Amide III, A and C, CH/CH2/CH3 def | ||
| 1315 | 1317 | Amide III, G, lipids CH2/CH3 def | |
| 1324 | Amide III; G; protein CH2/CH3 twist | ||
| 1330 | Amide III; CH2/CH3 def | ||
| 1357 | 1356 | 1353 | Protein CH/CH2/CH3 def; Trp |
| 1390 | CH rock | ||
| 1423 | A, G, CH3CH2 twist | ||
| 1434 | CH2 def | ||
| 1445 | 1446 | CH2/CH3 def | |
| 1464 | CH2 def | ||
| 1495 | 1490 | Amide II, NH3+ | |
| 1545 | Amide II, lipid CH2 sciss | ||
| 1560 | 1566 | Amide II; Trp, Tyr, COO– str | |
| 1575 | 1579 | 1580 | Amide II, C-C str,COO– str |
| 1609 | Trp, Tyr, Phe | ||
| 1708 | Asp, Glu C=O str | ||
| Raman Shift (cm−1) | Tentative Band Assignment | |
|---|---|---|
| NS 50 | NS 75 | |
| 459 | Protein S-S str, C-S str, Trp ring def | |
| 491 | 495 | Protein S-S str |
| 550 | S-S str | |
| 572 | Trp | |
| 652 | Cys C-S str, Tyr C-C twist, Phe | |
| 729 | Lipid C-N str; C-S str | |
| 757 | Trp br; C-N str | |
| 838 | Tyr ring br, Phe, Cα-N and C-C str, O-P-O str | |
| 854 | Tyr ring br; Phe, Cα-N and C-C str | |
| 865 | Lipid C-N str | |
| 964 | 965 | Lipids C-C str, Pro, Val |
| 1067 | C-C and C-N str; DNA/RNA O-P-O str; Pro | |
| 1088 | Lipid C-C str | |
| 1100 | Phe; C-N str | |
| 1125 | Lipid C-C str | |
| 1135 | Protein backbone C-C str; C-N str | |
| 1143 | Lipid C-C str; C-N str | |
| 1169 | Tyr and lipids C-H bend | |
| 1175 | Tyr C-H bend; Phe | |
| 1218 | Amide III; T, A; O-P-O str; C-N str | |
| 1261 | 1265 | Amide III |
| 1315 | 1315 | Amide III, G, lipids CH2/CH3 def |
| 1330 | Amide III; CH2/CH3 def | |
| 1353 | 1357 | Protein CH/CH2/CH3 def; Trp |
| 1365 | Lipids CH2 def | |
| 1400 | 1395 | CH rock; C=O str |
| 1427 | A, G, CH3CH2 twist | |
| 1441 | 1442 | CH2/CH3 def |
| 1475 | Lipid CH2/CH3 def | |
| 1495 | 1496 | Amide II, NH3+ |
| 1535 | Amide II; N-H def | |
| 1561 | 1559 | Amide II; Trp, Tyr, COO– str |
| 1575 | Amide II, C-C str,COO– str | |
| 1585 | C=C str, COO– str; Phe | |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Spedalieri, C.; Szekeres, G.P.; Werner, S.; Guttmann, P.; Kneipp, J. Probing the Intracellular Bio-Nano Interface in Different Cell Lines with Gold Nanostars. Nanomaterials 2021, 11, 1183. https://doi.org/10.3390/nano11051183
Spedalieri C, Szekeres GP, Werner S, Guttmann P, Kneipp J. Probing the Intracellular Bio-Nano Interface in Different Cell Lines with Gold Nanostars. Nanomaterials. 2021; 11(5):1183. https://doi.org/10.3390/nano11051183
Chicago/Turabian StyleSpedalieri, Cecilia, Gergo Péter Szekeres, Stephan Werner, Peter Guttmann, and Janina Kneipp. 2021. "Probing the Intracellular Bio-Nano Interface in Different Cell Lines with Gold Nanostars" Nanomaterials 11, no. 5: 1183. https://doi.org/10.3390/nano11051183
APA StyleSpedalieri, C., Szekeres, G. P., Werner, S., Guttmann, P., & Kneipp, J. (2021). Probing the Intracellular Bio-Nano Interface in Different Cell Lines with Gold Nanostars. Nanomaterials, 11(5), 1183. https://doi.org/10.3390/nano11051183