Endothelial Cell Targeting by cRGD-Functionalized Polymeric Nanoparticles under Static and Flow Conditions
Abstract
1. Introduction
2. Materials and Methods
2.1. Chemicals
2.2. Synthesis of Cy5 Labeled Poly(D,L-lactic-co-glycolic-co-hydroxymethyl glycolic acid) (Cy5-PLGHMGA)
2.3. Preparation and Characterization of the Polymeric Nanoparticles (NPs)
2.4. Conjugation of cRGD to the NPs
2.5. Quantification of cRGD by HPLC to Determine the Conjugation Efficiency to the NPs
2.6. Cell Culture Conditions
2.7. Association of Cys-NPs and cRGD-NPs with HUVECs under Static Conditions
2.8. Uptake of Cys-NPs and cRGD-NPs by HUVECs under Static Incubation Conditions
2.9. Association of Cys-NPs and RGD-NPs with HUVECs under Flow
2.9.1. Experiments under Flow: 1 h Incubation in EBM-2 Medium
2.9.2. Experiments under Flow: 16 h Incubation in EBM-2 Medium
2.9.3. Experiments under Flow, 16 h Incubation in Washed Red Blood Cells (RBCs)
2.10. Statistical Analysis
3. Results
3.1. Preparation and Characterization of the Polymeric Nanoparticles (NPs)
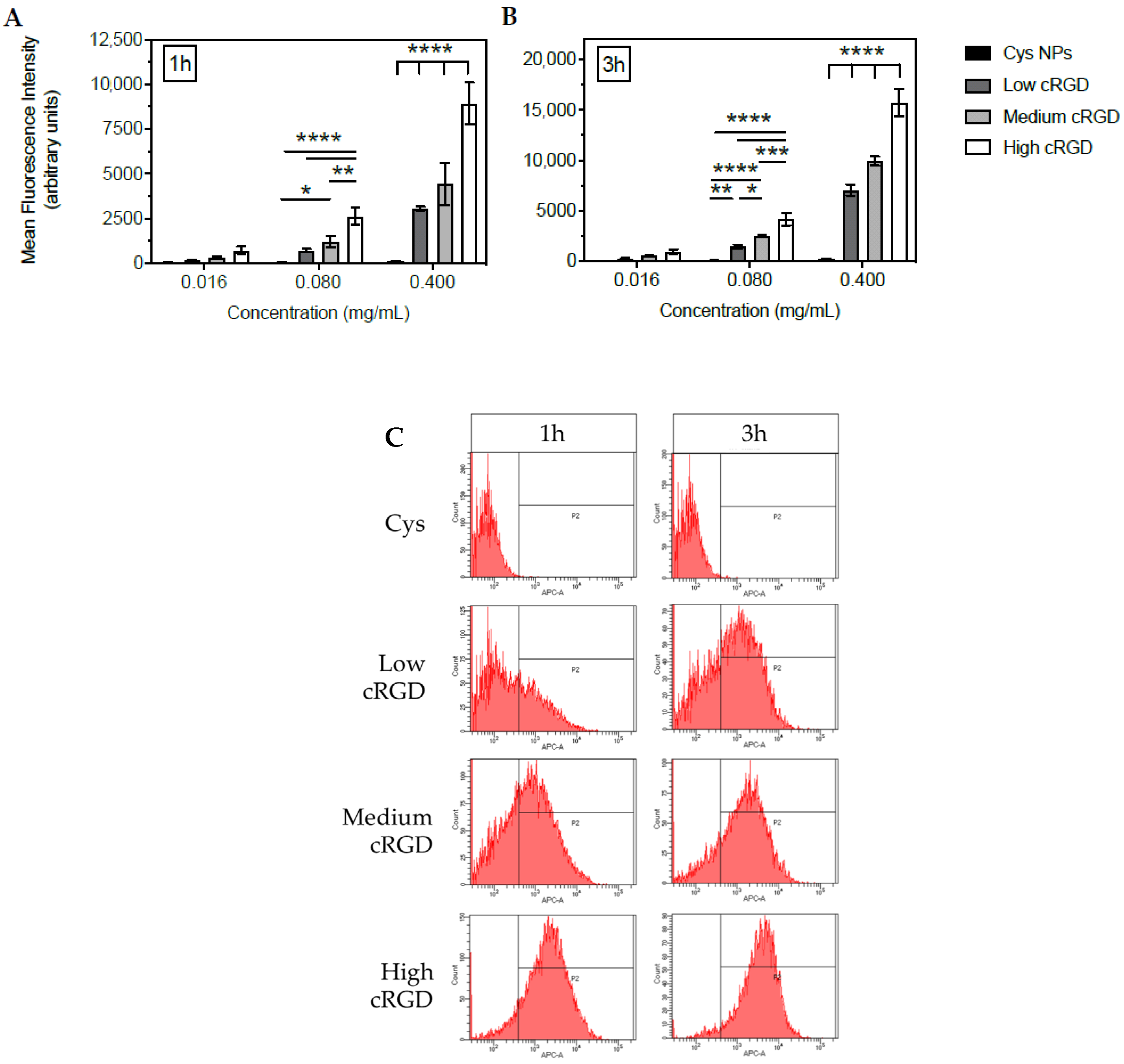
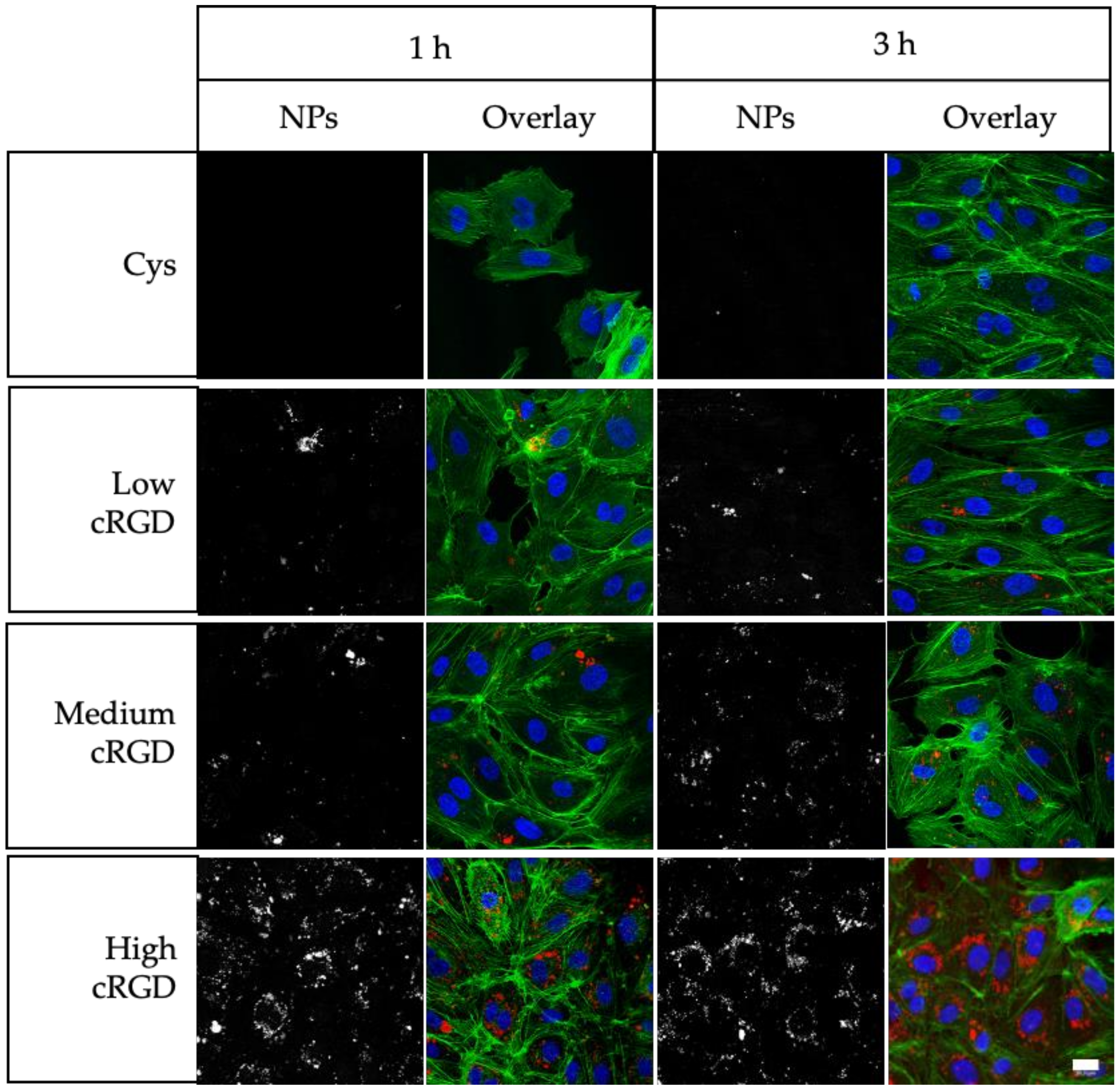
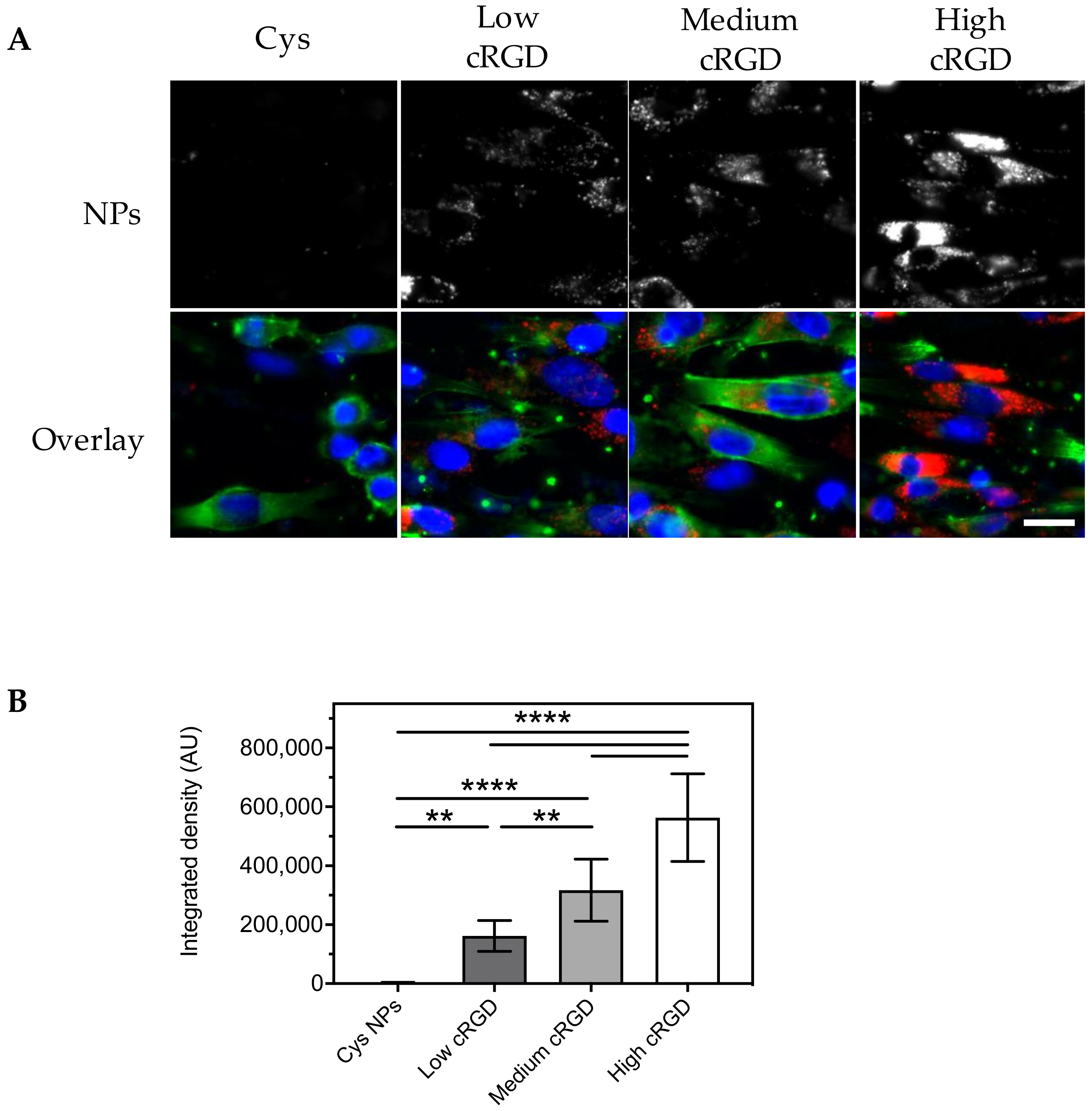
3.2. Association of Cys-NPs and RGD-NPs with HUVECs under Static Incubation Conditions
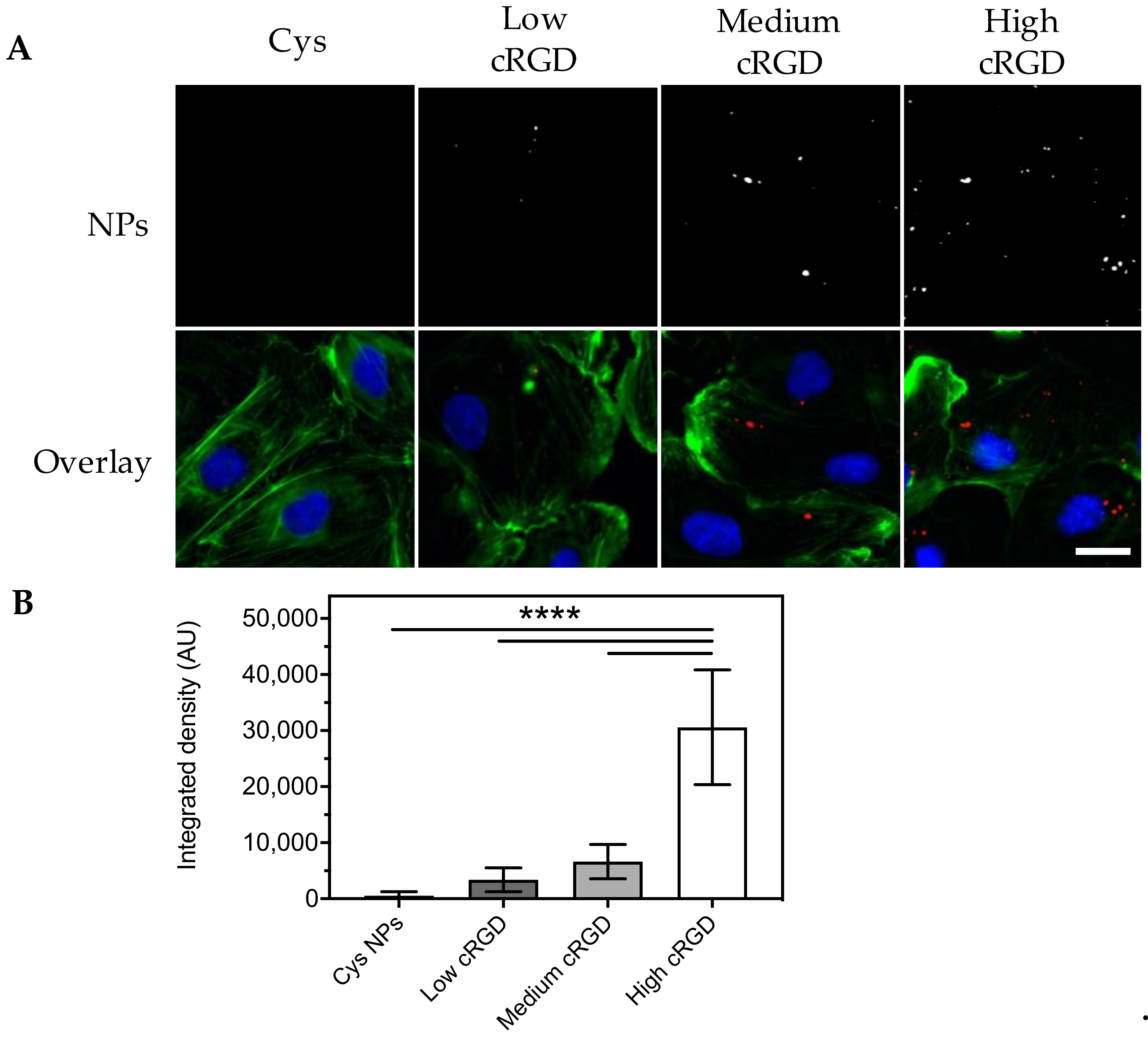
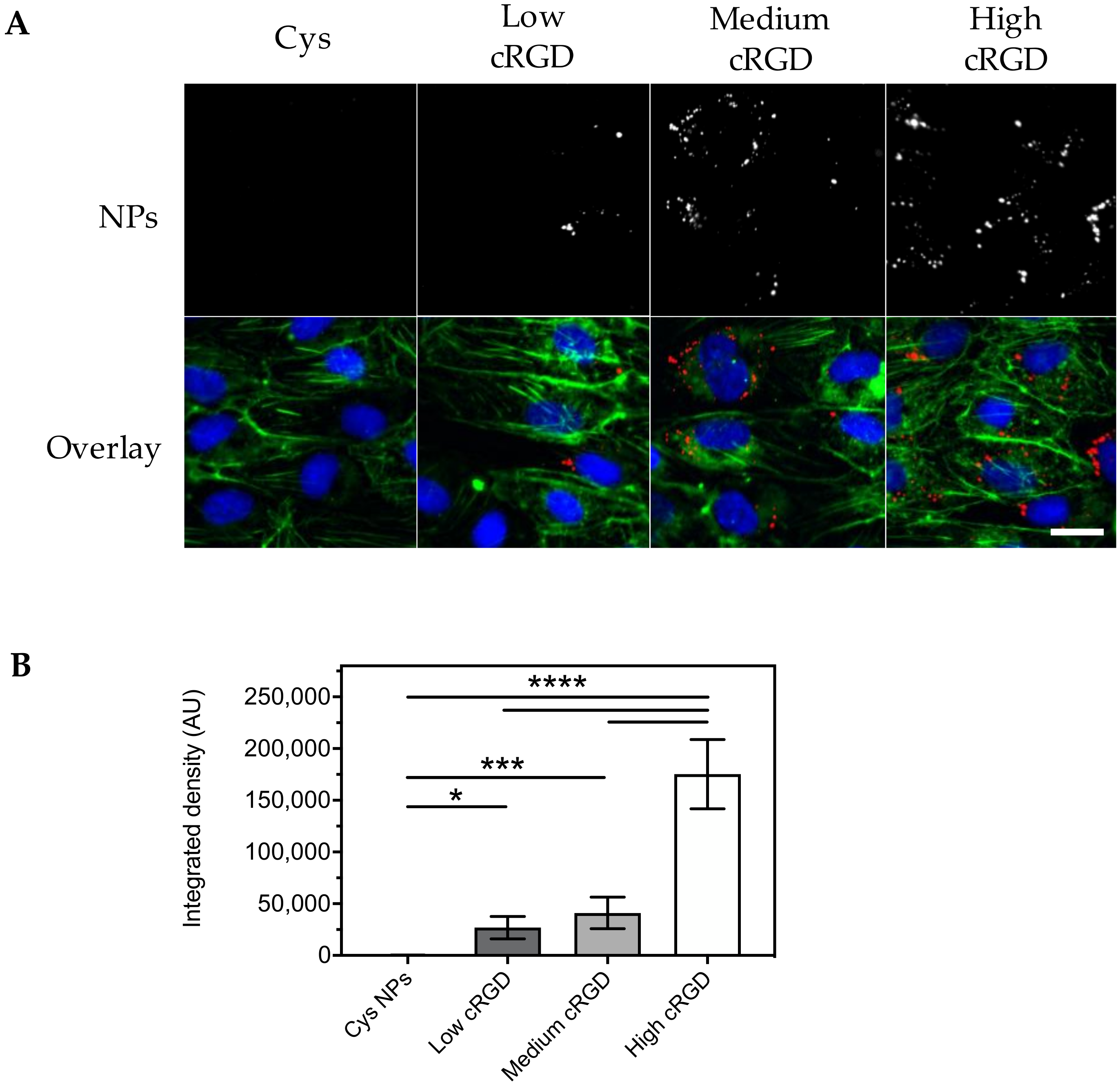
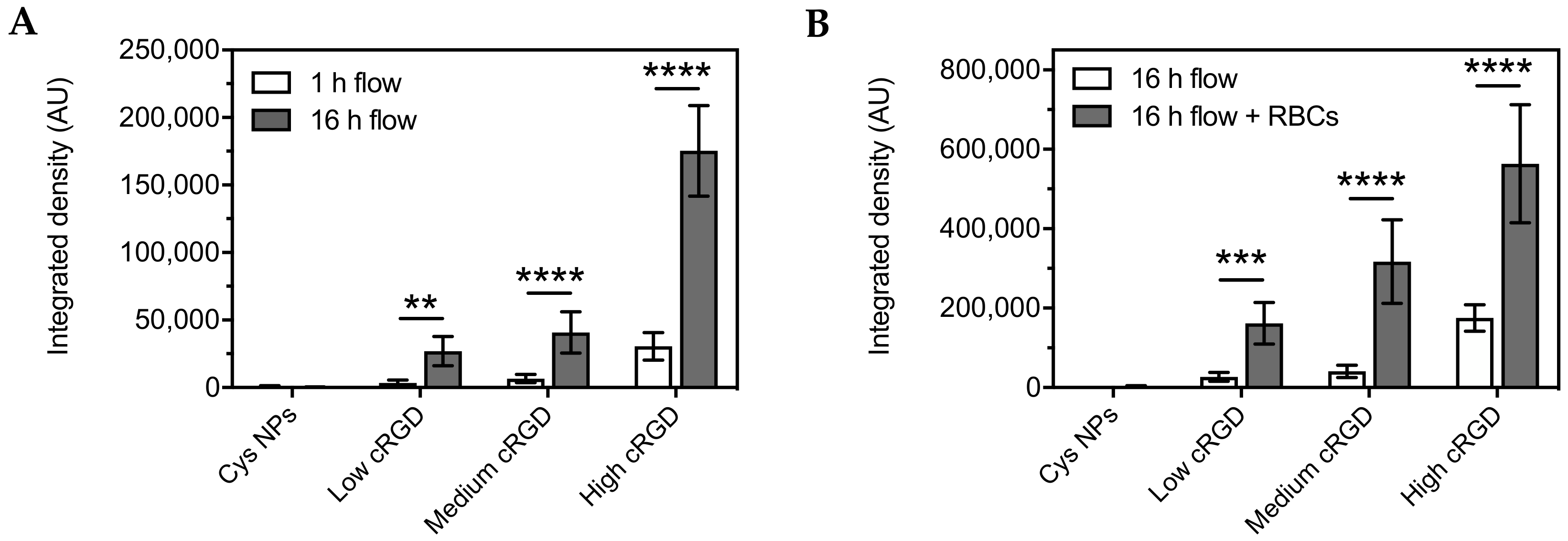
3.3. Association of Cys-NPs and RGD-NPs with HUVECs under Flow
4. Discussion
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Appendix A
References
- Carmeliet, P.; Jain, R.K. Molecular mechanisms and clinical applications of angiogenesis. Nature 2011, 473, 298–307. [Google Scholar] [CrossRef] [PubMed]
- Otrock, Z.K.; Mahfouz, R.A.; Makarem, J.A.; Shamseddine, A.I. Understanding the biology of angiogenesis: Review of the most important molecular mechanisms. Blood Cells Mol. Dis. 2007, 39, 212–220. [Google Scholar] [CrossRef] [PubMed]
- Stromblad, S.; Cheresh, D.A. Cell adhesion and angiogenesis. Trends Cell Biol. 1996, 6, 462–468. [Google Scholar] [CrossRef]
- Hynes, R.O. Integrins: Bidirectional, allosteric signaling machines. Cell 2002, 110, 673–687. [Google Scholar] [CrossRef]
- Avraamides, C.J.; Garmy-Susini, B.; Varner, J.A. Integrins in angiogenesis and lymphangiogenesis. Nat. Rev. Cancer 2008, 8, 604–617. [Google Scholar] [CrossRef]
- Brooks, P.C.; Clark, R.A.; Cheresh, D.A. Requirement of vascular integrin alpha v beta 3 for angiogenesis. Science 1994, 264, 569–571. [Google Scholar] [CrossRef]
- Kapp, T.G.; Rechenmacher, F.; Neubauer, S.; Maltsev, O.; Cavalcanti-Adam, E.; Zarka, R.; Reuning, U.; Notni, J.; Wester, H.-J.; Mas-Moruno, C.; et al. A Comprehensive Evaluation of the Activity and Selectivity Profile of Ligands for RGD-Binding Integrins. Sci Rep. 2017, 7, 39805. [Google Scholar] [CrossRef] [PubMed]
- Ruoslahti, E. RGD and other recognition sequences for integrins. Annu Rev. Cell Dev. Biol. 1996, 12, 697–715. [Google Scholar] [CrossRef]
- Desgrosellier, J.S.; Cheresh, D.A. Integrins in cancer: Biological implications and therapeutic opportunities. Nat. Rev. Cancer 2010, 10, 9–22. [Google Scholar] [CrossRef]
- Mizejewski, G.J. Role of integrins in cancer: Survey of expression patterns. Proc. Soc. Exp. Biol. Med. 1999, 222, 124–138. [Google Scholar] [CrossRef] [PubMed]
- Nieberler, M.; Reuning, U.; Reichart, F.; Notni, J.; Wester, H.J.; Schwaiger, M.; Weinmuller, M.; Rader, A.; Steiger, K.; Kessler, H. Exploring the Role of RGD-Recognizing Integrins in Cancer. Cancers 2017, 9, 116. [Google Scholar] [CrossRef]
- Eble, J.A.; Haier, J. Integrins in cancer treatment. Curr. Cancer Drug Targets 2006, 6, 89–105. [Google Scholar] [CrossRef] [PubMed]
- Kumar, C.C. Integrin alpha v beta 3 as a therapeutic target for blocking tumor-induced angiogenesis. Curr. Drug Targets 2003, 4, 123–131. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; Wang, F.; Chen, X. Integrin alpha(v)beta(3)-Targeted Cancer Therapy. Drug Dev. Res. 2008, 69, 329–339. [Google Scholar] [CrossRef] [PubMed]
- Raab-Westphal, S.; Marshall, J.F.; Goodman, S.L. Integrins as Therapeutic Targets: Successes and Cancers. Cancers 2017, 9, 110. [Google Scholar] [CrossRef] [PubMed]
- Gajbhiye, K.R.; Gajbhiye, V.; Siddiqui, I.A.; Gajbhiye, J.M. cRGD functionalised nanocarriers for targeted delivery of bioactives. J. Drug Target 2019, 27, 111–124. [Google Scholar] [CrossRef]
- Kunjachan, S.; Pola, R.; Gremse, F.; Theek, B.; Ehling, J.; Moeckel, D.; Hermanns-Sachweh, B.; Pechar, M.; Ulbrich, K.; Hennink, W.E.; et al. Passive versus active tumor targeting using RGD- and NGR-modified polymeric nanomedicines. Nano Lett. 2014, 14, 972–981. [Google Scholar] [CrossRef]
- Park, J.; Singha, K.; Son, S.; Kim, J.; Namgung, R.; Yun, C.O.; Kim, W.J. A review of RGD-functionalized nonviral gene delivery vectors for cancer therapy. Cancer Gene Ther. 2012, 19, 741–748. [Google Scholar] [CrossRef]
- Schiffelers, R.M.; Ansari, A.; Xu, J.; Zhou, Q.; Tang, Q.; Storm, G.; Molema, G.; Lu, P.Y.; Scaria, P.V.; Woodle, M.C. Cancer siRNA therapy by tumor selective delivery with ligand-targeted sterically stabilized nanoparticle. Nucleic Acids Res. 2004, 32, e149. [Google Scholar] [CrossRef]
- Wang, F.; Li, Y.; Shen, Y.; Wang, A.; Wang, S.; Xie, T. The functions and applications of RGD in tumor therapy and tissue engineering. Int. J. Mol. Sci. 2013, 14, 13447–13462. [Google Scholar] [CrossRef]
- Zhu, Y.; Zhang, J.; Meng, F.; Deng, C.; Cheng, R.; Feijen, J.; Zhong, Z. cRGD-functionalized reduction-sensitive shell-sheddable biodegradable micelles mediate enhanced doxorubicin delivery to human glioma xenografts in vivo. J. Control. Release 2016, 233, 29–38. [Google Scholar] [CrossRef] [PubMed]
- Danhier, F.; Vroman, B.; Lecouturier, N.; Crokart, N.; Pourcelle, V.; Freichels, H.; Jerome, C.; Marchand-Brynaert, J.; Feron, O.; Preat, V. Targeting of tumor endothelium by RGD-grafted PLGA-nanoparticles loaded with paclitaxel. J. Control. Release 2009, 140, 166–173. [Google Scholar] [CrossRef] [PubMed]
- Graf, N.; Bielenberg, D.R.; Kolishetti, N.; Muus, C.; Banyard, J.; Farokhzad, O.C.; Lippard, S.J. alpha(V)beta(3) integrin-targeted PLGA-PEG nanoparticles for enhanced anti-tumor efficacy of a Pt(IV) prodrug. ACS Nano 2012, 6, 4530–4539. [Google Scholar] [CrossRef] [PubMed]
- Liu, P.; Qin, L.; Wang, Q.; Sun, Y.; Zhu, M.; Shen, M.; Duan, Y. cRGD-functionalized mPEG-PLGA-PLL nanoparticles for imaging and therapy of breast cancer. Biomaterials 2012, 33, 6739–6747. [Google Scholar] [CrossRef] [PubMed]
- Yin, P.; Wang, Y.; Qiu, Y.; Hou, L.; Liu, X.; Qin, J.; Duan, Y.; Liu, P.; Qiu, M.; Li, Q. Bufalin-loaded mPEG-PLGA-PLL-cRGD nanoparticles: Preparation, cellular uptake, tissue distribution, and anticancer activity. Int. J. Nanomed. 2012, 7, 3961–3969. [Google Scholar] [CrossRef]
- Garanger, E.; Boturyn, D.; Coll, J.L.; Favrot, M.C.; Dumy, P. Multivalent RGD synthetic peptides as potent alphaVbeta3 integrin ligands. Org. Biomol. Chem. 2006, 4, 1958–1965. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.B.; Cai, W.; Cao, Q.; Chen, K.; Wu, Z.; He, L.; Chen, X. (64)Cu-labeled tetrameric and octameric RGD peptides for small-animal PET of tumor alpha(v)beta(3) integrin expression. J. Nucl. Med. 2007, 48, 1162–1171. [Google Scholar] [CrossRef]
- Liu, S. Radiolabeled Cyclic RGD Peptide Bioconjugates as Radiotracers Targeting Multiple Integrins. Bioconjugate Chem. 2015, 26, 1413–1438. [Google Scholar] [CrossRef]
- Ye, Y.; Bloch, S.; Xu, B.; Achilefu, S. Design, synthesis, and evaluation of near infrared fluorescent multimeric RGD peptides for targeting tumors. J. Med. Chem. 2006, 49, 2268–2275. [Google Scholar] [CrossRef]
- Goodwin, A.M. In vitro assays of angiogenesis for assessment of angiogenic and anti-angiogenic agents. Microvasc. Res. 2007, 74, 172–183. [Google Scholar] [CrossRef]
- Zimrin, A.B.; Villeponteau, B.; Maciag, T. Models of in vitro angiogenesis: Endothelial cell differentiation on fibrin but not matrigel is transcriptionally dependent. Biochem. Biophys. Res. Commun. 1995, 213, 630–638. [Google Scholar] [CrossRef] [PubMed]
- Valencia, P.M.; Farokhzad, O.C.; Karnik, R.; Langer, R. Microfluidic technologies for accelerating the clinical translation of nanoparticles. Nat. Nanotechnol. 2012, 7, 623–629. [Google Scholar] [CrossRef] [PubMed]
- Van der Meer, A.D.; Poot, A.A.; Duits, M.H.; Feijen, J.; Vermes, I. Microfluidic technology in vascular research. J. Biomed. Biotechnol. 2009, 2009, 823148. [Google Scholar] [CrossRef] [PubMed]
- Cicha, I.; Beronov, K.; Ramirez, E.L.; Osterode, K.; Goppelt-Struebe, M.; Raaz, D.; Yilmaz, A.; Daniel, W.G.; Garlichs, C.D. Shear stress preconditioning modulates endothelial susceptibility to circulating TNF-alpha and monocytic cell recruitment in a simplified model of arterial bifurcations. Atherosclerosis 2009, 207, 93–102. [Google Scholar] [CrossRef] [PubMed]
- Klingberg, H.; Loft, S.; Oddershede, L.B.; Moller, P. The influence of flow, shear stress and adhesion molecule targeting on gold nanoparticle uptake in human endothelial cells. Nanoscale 2015, 7, 11409–11419. [Google Scholar] [CrossRef]
- Samuel, S.P.; Jain, N.; O’Dowd, F.; Paul, T.; Kashanin, D.; Gerard, V.A.; Gun’ko, Y.K.; Prina-Mello, A.; Volkov, Y. Multifactorial determinants that govern nanoparticle uptake by human endothelial cells under flow. Int. J. Nanomed. 2012, 7, 2943–2956. [Google Scholar] [CrossRef]
- Chen, Y.Y.; Syed, A.M.; MacMillan, P.; Rocheleau, J.V.; Chan, W.C.W. Flow Rate Affects Nanoparticle Uptake into Endothelial Cells. Adv. Mater. 2020, 32, e1906274. [Google Scholar] [CrossRef]
- Jurney, P.; Agarwal, R.; Singh, V.; Choi, D.; Roy, K.; Sreenivasan, S.V.; Shi, L. Unique size and shape-dependent uptake behaviors of non-spherical nanoparticles by endothelial cells due to a shearing flow. J. Control. Release 2017, 245, 170–176. [Google Scholar] [CrossRef]
- Fede, C.; Fortunati, I.; Weber, V.; Rossetto, N.; Bertasi, F.; Petrelli, L.; Guidolin, D.; Signorini, R.; De Caro, R.; Albertin, G.; et al. Evaluation of gold nanoparticles toxicity towards human endothelial cells under static and flow conditions. Microvasc. Res. 2015, 97, 147–155. [Google Scholar] [CrossRef]
- Matuszak, J.; Baumgartner, J.; Zaloga, J.; Juenet, M.; da Silva, A.E.; Franke, D.; Almer, G.; Texier, I.; Faivre, D.; Metselaar, J.M.; et al. Nanoparticles for intravascular applications: Physicochemical characterization and cytotoxicity testing. Nanomedicine 2016, 11, 597–616. [Google Scholar] [CrossRef]
- Martinez-Jothar, L.; Beztsinna, N.; van Nostrum, C.F.; Hennink, W.E.; Oliveira, S. Selective Cytotoxicity to HER2 Positive Breast Cancer Cells by Saporin-Loaded Nanobody-Targeted Polymeric Nanoparticles in Combination with Photochemical Internalization. Mol. Pharm. 2019, 16, 1633–1647. [Google Scholar] [CrossRef]
- Leemhuis, M.; Van Nostrum, C.F.; Kruijtzer, J.A.W.; Zhong, Z.Y.; ten Breteler, M.R.; Dijkstra, P.J.; Feijen, J.; Hennink, W.E. Functionalized poly(alpha-hydroxy acid)s via ring-opening polymerization: Toward hydrophilic polyesters with pendant hydroxyl groups. Macromolecules 2006, 39, 3500–3508. [Google Scholar] [CrossRef]
- Samadi, N.; van Steenbergen, M.J.; van den Dikkenberg, J.B.; Vermonden, T.; van Nostrum, C.F.; Amidi, M.; Hennink, W.E. Nanoparticles based on a hydrophilic polyester with a sheddable PEG coating for protein delivery. Pharm. Res. 2014, 31, 2593–2604. [Google Scholar] [CrossRef] [PubMed]
- Zambaux, M.F.; Bonneaux, F.; Gref, R.; Maincent, P.; Dellacherie, E.; Alonso, M.J.; Labrude, P.; Vigneron, C. Influence of experimental parameters on the characteristics of poly(lactic acid) nanoparticles prepared by a double emulsion method. J. Control. Release 1998, 50, 31–40. [Google Scholar] [CrossRef]
- Martinez-Jothar, L.; Doulkeridou, S.; Schiffelers, R.M.; Torano, J.S.; Oliveira, S.; van Nostrum, C.F.; Hennink, W.E. Insights into maleimide-thiol conjugation chemistry: Conditions for efficient surface functionalization of nanoparticles for receptor targeting. J. Control. Release 2018, 282, 101–109. [Google Scholar] [CrossRef]
- Yameen, B.; Choi, W.I.; Vilos, C.; Swami, A.; Shi, J.; Farokhzad, O.C. Insight into nanoparticle cellular uptake and intracellular targeting. J. Control. Release 2014, 190, 485–499. [Google Scholar] [CrossRef] [PubMed]
- Boeneman, K.; Delehanty, J.B.; Blanco-Canosa, J.B.; Susumu, K.; Stewart, M.H.; Oh, E.; Huston, A.L.; Dawson, G.; Ingale, S.; Walters, R.; et al. Selecting improved peptidyl motifs for cytosolic delivery of disparate protein and nanoparticle materials. ACS Nano 2013, 7, 3778–3796. [Google Scholar] [CrossRef] [PubMed]
- Salomone, F.; Cardarelli, F.; Di Luca, M.; Boccardi, C.; Nifosi, R.; Bardi, G.; Di Bari, L.; Serresi, M.; Beltram, F. A novel chimeric cell-penetrating peptide with membrane-disruptive properties for efficient endosomal escape. J. Control. Release 2012, 163, 293–303. [Google Scholar] [CrossRef] [PubMed]
- Javiya, C.; Jonnalagadda, S. Physicochemical characterization of spray-dried PLGA/PEG microspheres, and preliminary assessment of biological response. Drug Dev. Ind. Pharm. 2016, 42, 1504–1514. [Google Scholar] [CrossRef]
- Malzert, A.; Boury, F.; Saulnier, P.; Benoit, J.P.; Proust, J.E. Interfacial properties of mixed polyethylene glycol/poly (d,l-lactide-co-glycolide) films spread at the air/water interface. Langmuir 2000, 16, 1861–1867. [Google Scholar] [CrossRef]
- Chen, Y.; van Steenbergen, M.J.; Li, D.; van de Dikkenberg, J.B.; Lammers, T.; van Nostrum, C.F.; Metselaar, J.M.; Hennink, W.E. Polymeric Nanogels with Tailorable Degradation Behavior. Macromol. Biosci. 2016, 16, 1122–1137. [Google Scholar] [CrossRef]
- Zhang, X.P.; Sun, J.G.; Yao, J.; Shan, K.; Liu, B.H.; Yao, M.D.; Ge, H.M.; Jiang, Q.; Zhao, C.; Yan, B. Effect of nanoencapsulation using poly (lactide-co-glycolide) (PLGA) on anti-angiogenic activity of bevacizumab for ocular angiogenesis therapy. Biomed. Pharmacother. 2018, 107, 1056–1063. [Google Scholar] [CrossRef] [PubMed]
- Trif, M.; Florian, P.E.; Roseanu, A.; Moisei, M.; Craciunescu, O.; Astete, C.E.; Sabliov, C.M. Cytotoxicity and intracellular fate of PLGA and chitosan-coated PLGA nanoparticles in Madin-Darby bovine kidney (MDBK) and human colorectal adenocarcinoma (Colo 205) cells. J. Biomed. Mater. Res. A 2015, 103, 3599–3611. [Google Scholar] [CrossRef]
- Alqahtani, S.; Simon, L.; Astete, C.E.; Alayoubi, A.; Sylvester, P.W.; Nazzal, S.; Shen, Y.; Xu, Z.; Kaddoumi, A.; Sabliov, C.M. Cellular uptake, antioxidant and antiproliferative activity of entrapped alpha-tocopherol and gamma-tocotrienol in poly (lactic-co-glycolic) acid (PLGA) and chitosan covered PLGA nanoparticles (PLGA-Chi). J. Colloid Interface Sci. 2015, 445, 243–251. [Google Scholar] [CrossRef]
- Zhang, N.; Chittasupho, C.; Duangrat, C.; Siahaan, T.J.; Berkland, C. PLGA nanoparticle--peptide conjugate effectively targets intercellular cell-adhesion molecule-1. Bioconjugate Chem. 2008, 19, 145–152. [Google Scholar] [CrossRef]
- Canton, I.; Battaglia, G. Endocytosis at the nanoscale. Chem. Soc. Rev. 2012, 41, 2718–2739. [Google Scholar] [CrossRef]
- Petros, R.A.; DeSimone, J.M. Strategies in the design of nanoparticles for therapeutic applications. Nat. Rev. Drug Discov. 2010, 9, 615–627. [Google Scholar] [CrossRef] [PubMed]
- Verma, A.; Stellacci, F. Effect of surface properties on nanoparticle-cell interactions. Small 2010, 6, 12–21. [Google Scholar] [CrossRef] [PubMed]
- Mailander, V.; Landfester, K. Interaction of nanoparticles with cells. Biomacromolecules 2009, 10, 2379–2400. [Google Scholar] [CrossRef] [PubMed]
- Danhier, F.; Le Breton, A.; Preat, V. RGD-based strategies to target alpha(v) beta(3) integrin in cancer therapy and diagnosis. Mol. Pharm. 2012, 9, 2961–2973. [Google Scholar] [CrossRef]
- Fens, M.H.; Mastrobattista, E.; de Graaff, A.M.; Flesch, F.M.; Ultee, A.; Rasmussen, J.T.; Molema, G.; Storm, G.; Schiffelers, R.M. Angiogenic endothelium shows lactadherin-dependent phagocytosis of aged erythrocytes and apoptotic cells. Blood 2008, 111, 4542–4550. [Google Scholar] [CrossRef] [PubMed]
- Amin, M.; Mansourian, M.; Koning, G.A.; Badiee, A.; Jaafari, M.R.; Ten Hagen, T.L.M. Development of a novel cyclic RGD peptide for multiple targeting approaches of liposomes to tumor region. J. Control. Release 2015, 220, 308–315. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Chui, W.K.; Ho, P.C. Design of a multifunctional PLGA nanoparticulate drug delivery system: Evaluation of its physicochemical properties and anticancer activity to malignant cancer cells. Pharm. Res. 2009, 26, 1162–1171. [Google Scholar] [CrossRef] [PubMed]
- Hoekman, J.D.; Srivastava, P.; Ho, R.J. Aerosol-stable peptide-coated liposome nanoparticles: A proof-of-concept study with opioid fentanyl in enhancing analgesic effects and reducing plasma drug exposure. J. Pharm. Sci. 2014, 103, 2231–2239. [Google Scholar] [CrossRef]
- Shan, D.; Li, J.; Cai, P.; Prasad, P.; Liu, F.; Rauth, A.M.; Wu, X.Y. RGD-conjugated solid lipid nanoparticles inhibit adhesion and invasion of alphavbeta3 integrin-overexpressing breast cancer cells. Drug Deliv. Transl. Res. 2015, 5, 15–26. [Google Scholar] [CrossRef]
- Valencia, P.M.; Hanewich-Hollatz, M.H.; Gao, W.; Karim, F.; Langer, R.; Karnik, R.; Farokhzad, O.C. Effects of ligands with different water solubilities on self-assembly and properties of targeted nanoparticles. Biomaterials 2011, 32, 6226–6233. [Google Scholar] [CrossRef] [PubMed]
- Kok, R.J.; Schraa, A.J.; Bos, E.J.; Moorlag, H.E.; Asgeirsdottir, S.A.; Everts, M.; Meijer, D.K.; Molema, G. Preparation and functional evaluation of RGD-modified proteins as alpha(v)beta(3) integrin directed therapeutics. Bioconjugate Chem. 2002, 13, 128–135. [Google Scholar] [CrossRef]
- Sancey, L.; Garanger, E.; Foillard, S.; Schoehn, G.; Hurbin, A.; Albiges-Rizo, C.; Boturyn, D.; Souchier, C.; Grichine, A.; Dumy, P.; et al. Clustering and internalization of integrin alphavbeta3 with a tetrameric RGD-synthetic peptide. Mol. Ther. 2009, 17, 837–843. [Google Scholar] [CrossRef]
- Shukla, R.; Thomas, T.P.; Peters, J.; Kotlyar, A.; Myc, A.; Baker, J.R., Jr. Tumor angiogenic vasculature targeting with PAMAM dendrimer-RGD conjugates. Chem. Commun. (Camb.) 2005, 5739–5741. [Google Scholar] [CrossRef]
- Temming, K.; Schiffelers, R.M.; Molema, G.; Kok, R.J. RGD-based strategies for selective delivery of therapeutics and imaging agents to the tumour vasculature. Drug Resist. Updat. 2005, 8, 381–402. [Google Scholar] [CrossRef]
- Wu, Y.; Zhang, X.; Xiong, Z.; Cheng, Z.; Fisher, D.R.; Liu, S.; Gambhir, S.S.; Chen, X. microPET imaging of glioma integrin {alpha}v{beta}3 expression using (64)Cu-labeled tetrameric RGD peptide. J. Nucl. Med. 2005, 46, 1707–1718. [Google Scholar] [PubMed]
- Reneman, R.S.; Arts, T.; Hoeks, A.P. Wall shear stress--an important determinant of endothelial cell function and structure--in the arterial system in vivo. Discrepancies with theory. J. Vasc. Res. 2006, 43, 251–269. [Google Scholar] [CrossRef] [PubMed]
- Stroev, P.V.; Hoskins, P.R.; Easson, W.J. Distribution of wall shear rate throughout the arterial tree: A case study. Atherosclerosis 2007, 191, 276–280. [Google Scholar] [CrossRef] [PubMed]
- Lin, A.; Sabnis, A.; Kona, S.; Nattama, S.; Patel, H.; Dong, J.F.; Nguyen, K.T. Shear-regulated uptake of nanoparticles by endothelial cells and development of endothelial-targeting nanoparticles. J. Biomed. Mater. Res. A 2010, 93, 833–842. [Google Scholar] [CrossRef]
- Rinkenauer, A.C.; Press, A.T.; Raasch, M.; Pietsch, C.; Schweizer, S.; Schworer, S.; Rudolph, K.L.; Mosig, A.; Bauer, M.; Traeger, A.; et al. Comparison of the uptake of methacrylate-based nanoparticles in static and dynamic in vitro systems as well as in vivo. J. Control. Release 2015, 216, 158–168. [Google Scholar] [CrossRef]
- Howard, M.; Zern, B.J.; Anselmo, A.C.; Shuvaev, V.V.; Mitragotri, S.; Muzykantov, V. Vascular targeting of nanocarriers: Perplexing aspects of the seemingly straightforward paradigm. ACS Nano 2014, 8, 4100–4132. [Google Scholar] [CrossRef]
- Zhang, J.; Johnson, P.C.; Popel, A.S. Effects of erythrocyte deformability and aggregation on the cell free layer and apparent viscosity of microscopic blood flows. Microvasc. Res. 2009, 77, 265–272. [Google Scholar] [CrossRef]
- Tan, J.; Thomas, A.; Liu, Y. Influence of Red Blood Cells on Nanoparticle Targeted Delivery in Microcirculation. Soft Matter 2011, 8, 1934–1946. [Google Scholar] [CrossRef]
- Tan, J.; Keller, W.; Sohrabi, S.; Yang, J.; Liu, Y. Characterization of Nanoparticle Dispersion in Red Blood Cell Suspension by the Lattice Boltzmann-Immersed Boundary Method. Nanomaterials 2016, 6, 30. [Google Scholar] [CrossRef]
- Charoenphol, P.; Huang, R.B.; Eniola-Adefeso, O. Potential role of size and hemodynamics in the efficacy of vascular-targeted spherical drug carriers. Biomaterials 2010, 31, 1392–1402. [Google Scholar] [CrossRef]
- Kiani, M.F.; Yuan, H.; Chen, X.; Smith, L.; Gaber, M.W.; Goetz, D.J. Targeting microparticles to select tissue via radiation-induced upregulation of endothelial cell adhesion molecules. Pharm. Res. 2002, 19, 1317–1322. [Google Scholar] [CrossRef] [PubMed]
- Qi, R.; Guo, R.; Shen, M.; Cao, X.; Zhang, L.; Xu, J.; Yu, J.; Shi, X. Electrospun poly(lactic-co-glycolic acid)/halloysite nanotube composite nanofibers for drug encapsulation and sustained release. J. Mater. Chem. 2010, 20, 10622–10629. [Google Scholar] [CrossRef]
| Formulation | Diameter (nm) | PDI | Zeta Potential (mV) | Conjugation Efficiency (%) a | cRGD Peptides Coupled per NP b | Density (cRGD/ µm2) b |
|---|---|---|---|---|---|---|
| NPs | 303 ± 34 | 0.25 ± 0.10 | −10.1 ± 0.7 | NA c | NA c | NA c |
| Cys-NPs | 316 ± 19 | 0.22 ± 0.02 | −12.2 ± 0.6 | 39 ± 12 | NA c | NA c |
| Low cRGD-NPs | 313 ± 11 | 0.21 ± 0.00 | −12.0 ± 0.4 | 38 ± 5 | ~3400 | ~11,000 |
| Medium cRGD-NPs | 312 ± 14 | 0.19 ± 0.03 | −11.4 ± 0.2 | 50 ± 8 | ~9700 | ~31,700 |
| High cRGD-NPs | 332 ± 22 | 0.23 ± 0.04 | −10.1 ± 0.6 | 71 ± 8 | ~40,700 | ~117,600 |
| Concentration of Fluorescent NPs (mg/mL) | Percentage of Fluorescence Positive HUVECs | |||||||
|---|---|---|---|---|---|---|---|---|
| Cys-NPs | Low cRGD-NPs | Med cRGD-NPs | High cRGD-NPs | |||||
| 1 h | 3 h | 1 h | 3 h | 1 h | 3 h | 1 h | 3 h | |
| 0.016 | 0.3 ± 0.2 | 0.2 ± 0.1 | 8 ± 1 | 15 ± 5 | 15 ± 4 | 36 ± 3 | 43 ± 8 | 60 ± 9 |
| 0.080 | 0.5 ± 0.2 | 0.9 ± 0.4 | 30 ± 4 | 58 ± 8 | 51 ± 11 | 83 ± 2 | 83 ± 5 | 93 ± 3 |
| 0.400 | 3 ± 1 | 12 ± 7 | 71 ± 1 | 91 ± 4 | 85 ± 5 | 98 ± 1 | 97 ± 1 | 99 ± 0 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Martínez-Jothar, L.; Barendrecht, A.D.; de Graaff, A.M.; Oliveira, S.; van Nostrum, C.F.; Schiffelers, R.M.; Hennink, W.E.; Fens, M.H.A.M. Endothelial Cell Targeting by cRGD-Functionalized Polymeric Nanoparticles under Static and Flow Conditions. Nanomaterials 2020, 10, 1353. https://doi.org/10.3390/nano10071353
Martínez-Jothar L, Barendrecht AD, de Graaff AM, Oliveira S, van Nostrum CF, Schiffelers RM, Hennink WE, Fens MHAM. Endothelial Cell Targeting by cRGD-Functionalized Polymeric Nanoparticles under Static and Flow Conditions. Nanomaterials. 2020; 10(7):1353. https://doi.org/10.3390/nano10071353
Chicago/Turabian StyleMartínez-Jothar, Lucía, Arjan D. Barendrecht, Anko M. de Graaff, Sabrina Oliveira, Cornelus F. van Nostrum, Raymond M. Schiffelers, Wim E. Hennink, and Marcel H. A. M. Fens. 2020. "Endothelial Cell Targeting by cRGD-Functionalized Polymeric Nanoparticles under Static and Flow Conditions" Nanomaterials 10, no. 7: 1353. https://doi.org/10.3390/nano10071353
APA StyleMartínez-Jothar, L., Barendrecht, A. D., de Graaff, A. M., Oliveira, S., van Nostrum, C. F., Schiffelers, R. M., Hennink, W. E., & Fens, M. H. A. M. (2020). Endothelial Cell Targeting by cRGD-Functionalized Polymeric Nanoparticles under Static and Flow Conditions. Nanomaterials, 10(7), 1353. https://doi.org/10.3390/nano10071353





