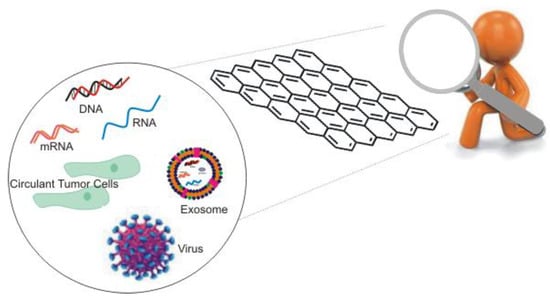
Graphene-Based Strategies in Liquid Biopsy and in Viral Diseases Diagnosis
Abstract
1. Introduction
2. Graphene Nanomaterials
3. Tumor Biomarkers in Liquid Biopsy
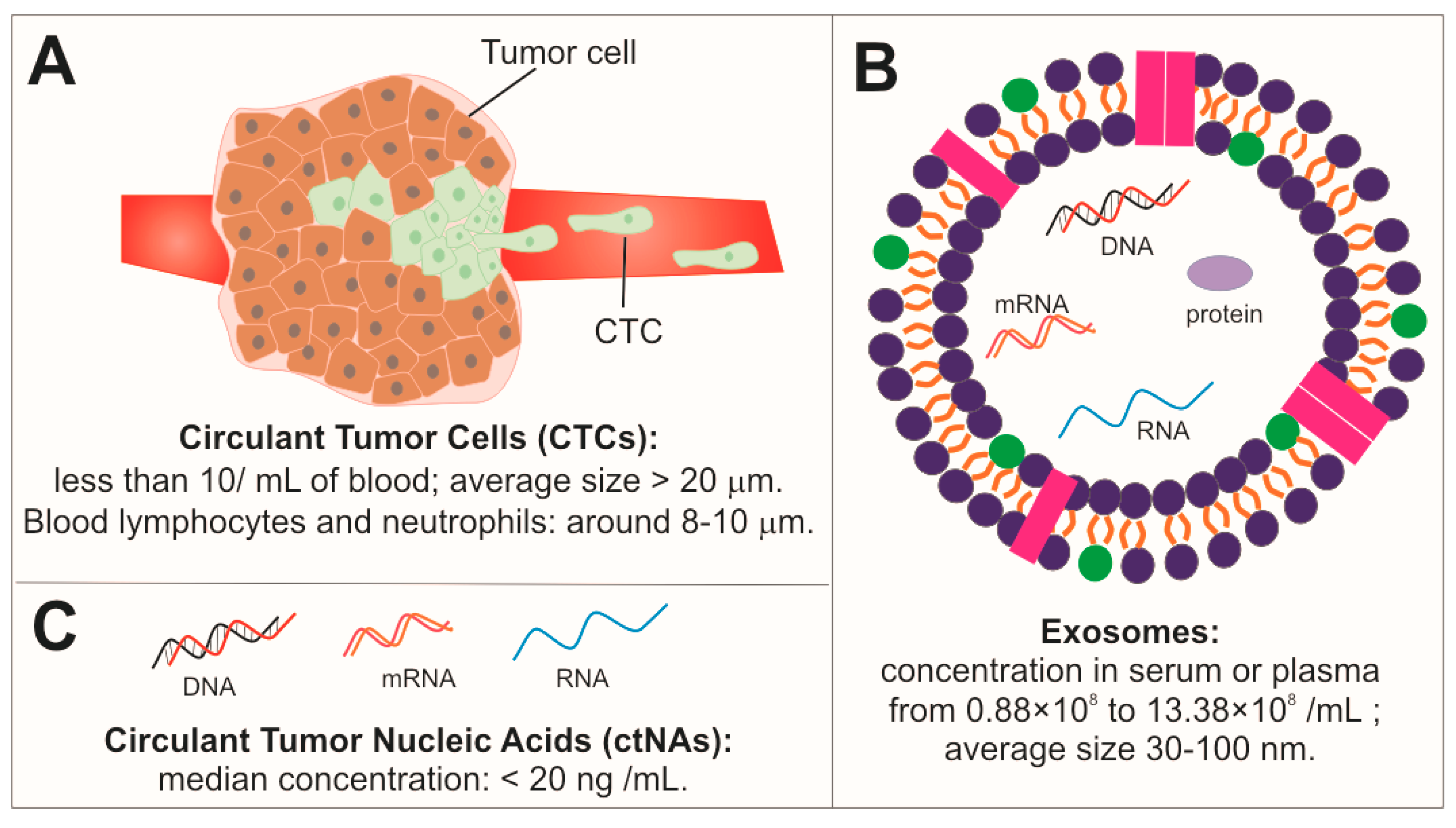
4. Circulant Tumor Cells (CTCs)
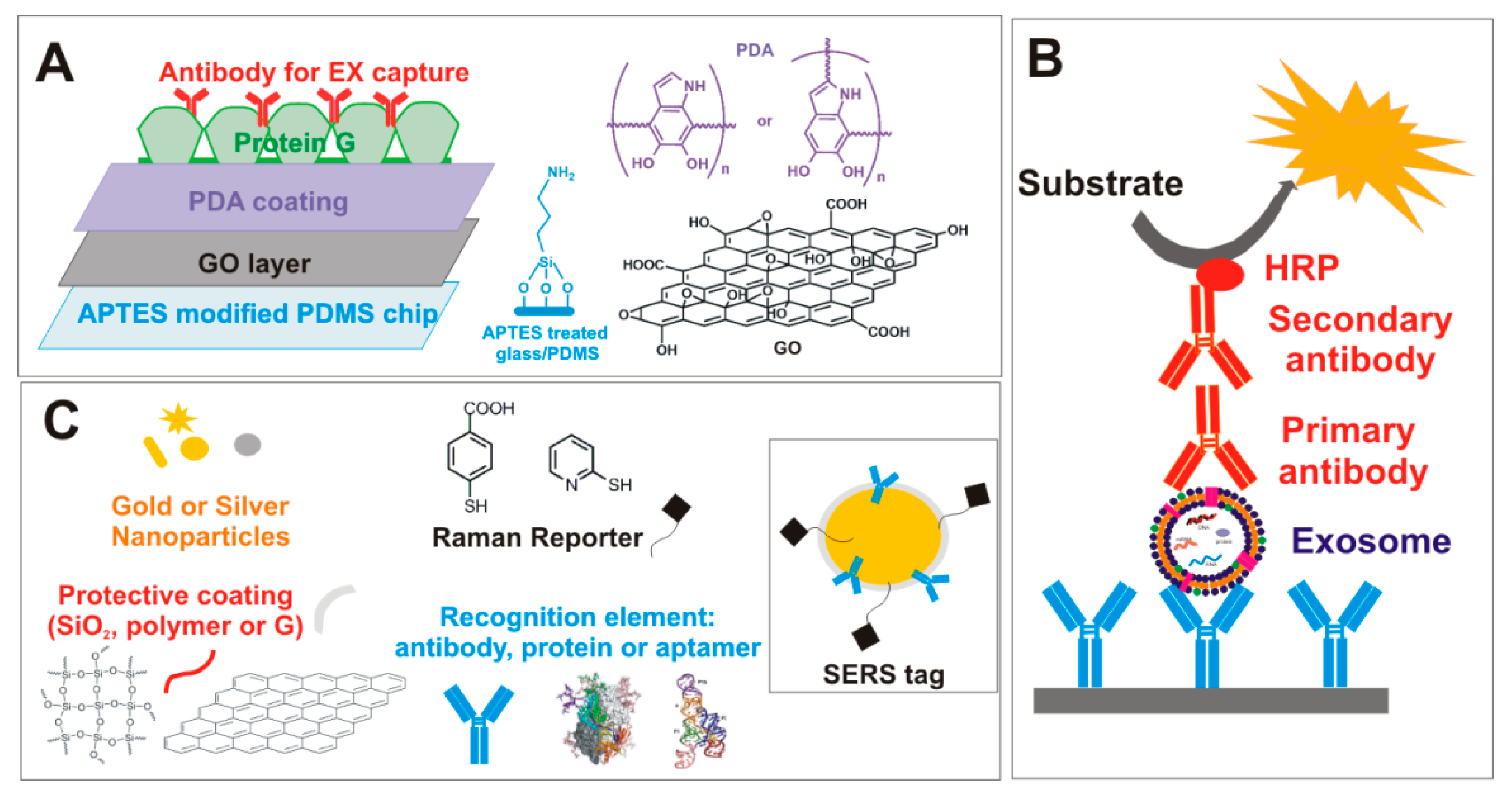
5. Exosomes (EXs)
6. Circulant Tumor Nucleic Acids (ctNAs)
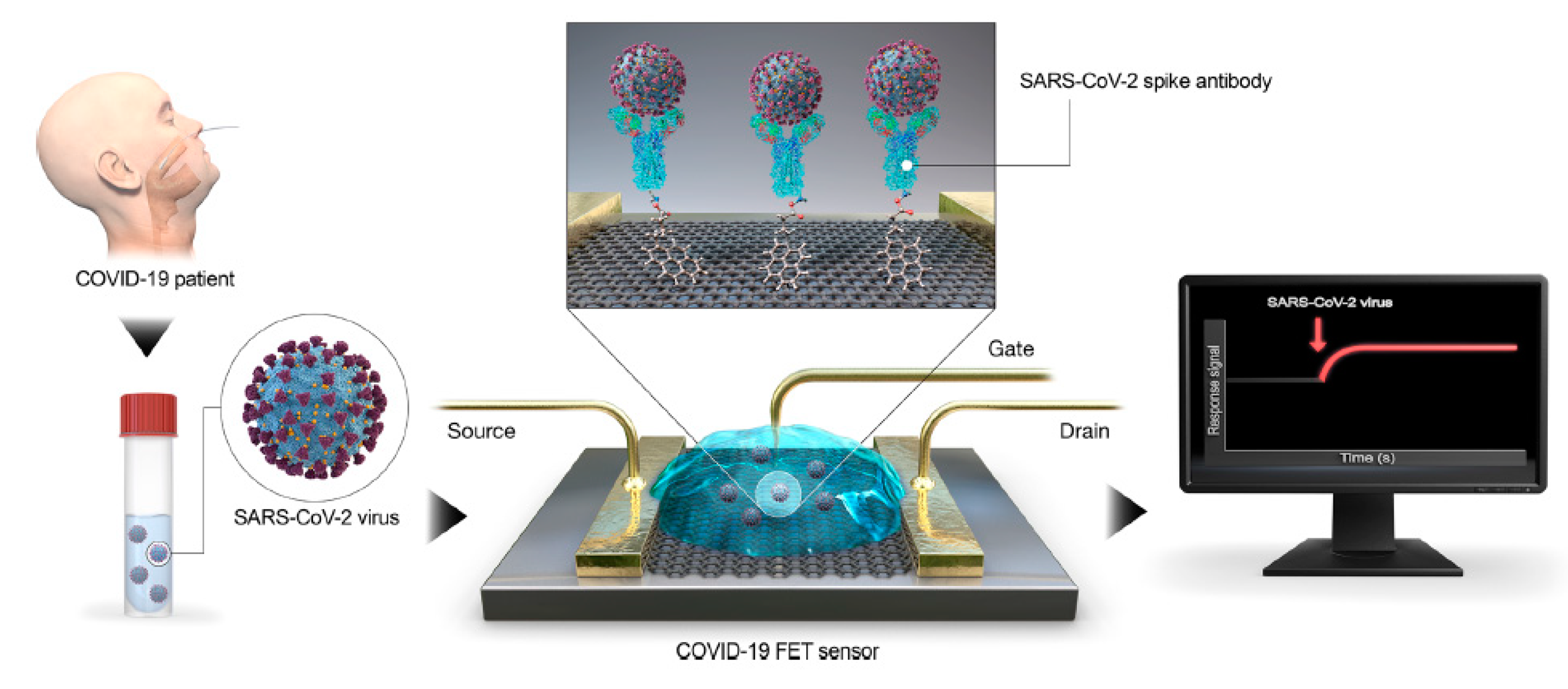
7. Graphene-Based Strategies in the Diagnosis of Viral Diseases
8. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Schmidt, B.; Fleischhacker, M. Is liquid biopsy ready for the litmus test and what has been achieved so far to deal with pre-analytical issues? Transl. Cancer Res. 2018, 7, S130–S139. [Google Scholar] [CrossRef]
- Vaidyanathan, R.; Soon, R.H.; Zhang, P.; Jiang, K.; Lim, C.T. Cancer diagnosis: From tumor to liquid biopsy and beyond. Lab Chip 2019, 19, 11–34. [Google Scholar] [CrossRef]
- La Marca, V.; Fierabracci, A. Insights into the diagnostic potential of extracellular vesicles and their mirna signature from liquid biopsy as early biomarkers of diabetic micro/macrovascular complications. Int. J. Mol. Sci. 2017, 18, 1974. [Google Scholar] [CrossRef]
- Stefancu, A.; Badarinza, M.; Moisoiu, V.; Iancu, S.D.; Serban, O.; Leopold, N.; Fodor, D. Sers-based liquid biopsy of saliva and serum from patients with Sjögren’s syndrome. Anal. Bioanal. Chem. 2019, 411, 5877–5883. [Google Scholar] [CrossRef]
- Suades, R.; Padró, T.; Crespo, J.; Sionis, A.; Alonso, R.; Mata, P.; Badimon, L. Liquid biopsy of extracellular microvesicles predicts future major ischemic events in genetically characterized familial hypercholesterolemia patients. Arterioscler. Thromb. Vasc. Biol. 2019, 39, 1172–1181. [Google Scholar] [CrossRef]
- Macías, M.; Alegre, E.; Díaz-Lagares, A.; Patiño, A.; Pérez-Gracia, J.L.; Sanmamed, M.; López-López, R.; Varo, N.; González, A. Chapter three-liquid biopsy: From basic research to clinical practice. In Advances in Clinical Chemistry; Makowski, G.S., Ed.; Elsevier: Amsterdam, The Netherlands, 2018; Volume 83, pp. 73–119. [Google Scholar]
- Li, W.; Ye, L.; Li, S.; Yao, H.; Ade, H.; Hou, J. A high-efficiency organic solar cell enabled by the strong intramolecular electron push–pull effect of the nonfullerene acceptor. Adv. Mater. 2018, 30, 1707170. [Google Scholar] [CrossRef]
- Gribko, A.; Künzel, J.; Wünsch, D.; Lu, Q.; Nagel, S.M.; Knauer, S.K.; Stauber, R.H.; Ding, G.-B. Is small smarter? Nanomaterial-based detection and elimination of circulating tumor cells: Current knowledge and perspectives. Int. J. Nanomed. 2019, 14, 4187–4209. [Google Scholar] [CrossRef]
- Zhu, Y.; Murali, S.; Cai, W.; Li, X.; Suk, J.W.; Potts, J.R.; Ruoff, R.S. Graphene and graphene oxide: Synthesis, properties, and applications. Adv. Mater. 2010, 22, 3906–3924. [Google Scholar] [CrossRef]
- Gu, H.; Tang, H.; Xiong, P.; Zhou, Z. Biomarkers-based biosensing and bioimaging with graphene for cancer diagnosis. Nanomaterials 2019, 9, 130. [Google Scholar] [CrossRef]
- Criado, A.; Melchionna, M.; Marchesan, S.; Prato, M. The covalent functionalization of graphene on substrates. Angew. Chem. Int. Ed. 2015, 54, 10734–10750. [Google Scholar] [CrossRef]
- Lee, J.-H.; Park, S.-J.; Choi, J.-W. Electrical property of graphene and its application to electrochemical biosensing. Nanomaterials 2019, 9, 297. [Google Scholar] [CrossRef] [PubMed]
- Neri, G.; Scala, A.; Barreca, F.; Fazio, E.; Mineo, P.G.; Mazzaglia, A.; Grassi, G.; Piperno, A. Engineering of carbon based nanomaterials by ring-opening reactions of a reactive azlactone graphene platform. Chem. Commun. 2015, 51, 4846–4849. [Google Scholar] [CrossRef] [PubMed]
- Piperno, A.; Scala, A.; Mazzaglia, A.; Neri, G.; Pennisi, R.; Sciortino, M.T.; Grassi, G. Cellular signaling pathways activated by functional graphene nanomaterials. Int. J. Mol. Sci. 2018, 19, 3365. [Google Scholar] [CrossRef] [PubMed]
- Tour, J.M. Top-down versus bottom-up fabrication of graphene-based electronics. Chem. Mater. 2014, 26, 163–171. [Google Scholar] [CrossRef]
- Neri, G.; Fazio, E.; Mineo, P.G.; Scala, A.; Piperno, A. SERS sensing properties of new graphene/gold nanocomposite. Nanomaterials 2019, 9, 1236. [Google Scholar] [CrossRef] [PubMed]
- Neri, G.; Scala, A.; Fazio, E.; Mineo, P.G.; Rescifina, A.; Piperno, A.; Grassi, G. Repurposing of oxazolone chemistry: Gaining access to functionalized graphene nanosheets in a top-down approach from graphite. Chem. Sci. 2015, 6, 6961–6970. [Google Scholar] [CrossRef]
- Ryzhii, V.; Otsuji, T.; Shur, M. Graphene based plasma-wave devices for terahertz applications. Appl. Phys. Lett. 2020, 116, 140501. [Google Scholar] [CrossRef]
- Zhang, L.; Yang, Z.; Gong, T.; Pan, R.; Wang, H.; Guo, Z.; Zhang, H.; Fu, X. Recent advances in emerging janus two-dimensional materials: From fundamental physics to device applications. J. Mater. Chem. A 2020, 8, 8813–8830. [Google Scholar] [CrossRef]
- Balandin, A.A. Phononics of graphene and related materials. ACS Nano 2020. [Google Scholar] [CrossRef]
- Morales-Narváez, E.; Merkoçi, A. Graphene oxide as an optical biosensing platform: A progress report. Adv. Mater. 2019, 31, 1805043. [Google Scholar] [CrossRef]
- Mousavi, S.M.; Hashemi, S.A.; Ghasemi, Y.; Amani, A.M.; Babapoor, A.; Arjmand, O. Applications of graphene oxide in case of nanomedicines and nanocarriers for biomolecules: Review study. Drug Metab. Rev. 2019, 51, 12–41. [Google Scholar] [CrossRef] [PubMed]
- Singh, R.K.; Kumar, R.; Singh, D.P. Graphene oxide: Strategies for synthesis, reduction and frontier applications. RSC Adv. 2016, 6, 64993–65011. [Google Scholar] [CrossRef]
- Chen, D.; Feng, H.; Li, J. Graphene oxide: Preparation, functionalization, and electrochemical applications. Chem. Rev. 2012, 112, 6027–6053. [Google Scholar] [CrossRef] [PubMed]
- Smith, A.T.; LaChance, A.M.; Zeng, S.; Liu, B.; Sun, L. Synthesis, properties, and applications of graphene oxide/reduced graphene oxide and their nanocomposites. Nano Mater. Sci. 2019, 1, 31–47. [Google Scholar] [CrossRef]
- Neri, G.; Micale, N.; Scala, A.; Fazio, E.; Mazzaglia, A.; Mineo, P.G.; Montesi, M.; Panseri, S.; Tampieri, A.; Grassi, G.; et al. Silibinin-conjugated graphene nanoplatform: Synthesis, characterization and biological evaluation. FlatChem 2017, 1, 34–41. [Google Scholar] [CrossRef]
- Narayanan, K.B.; Kim, H.D.; Han, S.S. Biocompatibility and hemocompatibility of hydrothermally derived reduced graphene oxide using soluble starch as a reducing agent. Colloids Surf. B Biointerfaces 2020, 185, 110579. [Google Scholar] [CrossRef]
- Li, L.; Wu, G.; Yang, G.; Peng, J.; Zhao, J.; Zhu, J.-J. Focusing on luminescent graphene quantum dots: Current status and future perspectives. Nanoscale 2013, 5, 4015–4039. [Google Scholar] [CrossRef]
- Kumawat, M.K.; Thakur, M.; Gurung, R.B.; Srivastava, R. Graphene quantum dots from mangifera indica: Application in near-infrared bioimaging and intracellular nanothermometry. Sustain. Chem. Eng. 2017, 5, 1382–1391. [Google Scholar] [CrossRef]
- Li, M.; Chen, T.; Gooding, J.J.; Liu, J. Review of carbon and graphene quantum dots for sensing. ACS Sens. 2019, 4, 1732–1748. [Google Scholar] [CrossRef]
- Wu, Y.; Zhu, J.; Huang, L. A review of three-dimensional graphene-based materials: Synthesis and applications to energy conversion/storage and environment. Carbon 2019, 143, 610–640. [Google Scholar] [CrossRef]
- Novoselov, K.S.; Geim, A.K.; Morozov, S.V.; Jiang, D.; Zhang, Y.; Dubonos, S.V.; Grigorieva, I.V.; Firsov, A.A. Electric field effect in atomically thin carbon films. Science 2004, 306, 666–669. [Google Scholar] [CrossRef] [PubMed]
- Li, H.; Gan, S.; Wang, H.; Han, D.; Niu, L. Intercorrelated superhybrid of AgBr supported on graphitic-C3N4-decorated nitrogen-doped graphene: High engineering photocatalytic activities for water purification and CO2 reduction. Adv. Mater. 2015, 27, 6906–6913. [Google Scholar] [CrossRef] [PubMed]
- Masoumi, S.; Hajghassem, H.; Erfanian, A.; Molaei Rad, A. Design and manufacture of TNT explosives detector sensors based onGFET. Sens. Rev. 2018, 38, 181–193. [Google Scholar] [CrossRef]
- Zhang, W.; Xia, W.; Lv, Z.; Ni, C.; Xin, Y.; Yang, L. Liquid biopsy for cancer: Circulating tumor cells, circulating free DNA or exosomes? Cell. Physiol. Biochem. 2017, 41, 755–768. [Google Scholar] [CrossRef] [PubMed]
- Lohr, J.G.; Adalsteinsson, V.A.; Cibulskis, K.; Choudhury, A.D.; Rosenberg, M.; Cruz-Gordillo, P.; Francis, J.M.; Zhang, C.-Z.; Shalek, A.K.; Satija, R.; et al. Whole-exome sequencing of circulating tumor cells provides a window into metastatic prostate cancer. Nat. Biotechnol. 2014, 32, 479–484. [Google Scholar] [CrossRef] [PubMed]
- Miller, M.A.; Oudin, M.J.; Sullivan, R.J.; Wang, S.J.; Meyer, A.S.; Im, H.; Frederick, D.T.; Tadros, J.; Griffith, L.G.; Lee, H.; et al. Reduced proteolytic shedding of receptor tyrosine kinases is a post-translational mechanism of kinase inhibitor resistance. Cancer Discov. 2016, 6, 382–399. [Google Scholar] [CrossRef]
- Im, H.; Shao, H.; Park, Y.I.; Peterson, V.M.; Castro, C.M.; Weissleder, R.; Lee, H. Label-free detection and molecular profiling of exosomes with a nano-plasmonic sensor. Nat. Biotechnol. 2014, 32, 490–495. [Google Scholar] [CrossRef]
- Spellman, P.T.; Gray, J.W. Detecting cancer by monitoring circulating tumor DNA. Nat. Med. 2014, 20, 474–475. [Google Scholar] [CrossRef]
- Schwarzenbach, H.; Stoehlmacher, J.; Pantel, K.; Goekkurt, E. Detection and monitoring of cell-free DNA in blood of patients with colorectal cancer. Ann. N. Y. Acad. Sci. 2008, 1137, 190–196. [Google Scholar] [CrossRef]
- Sozzi, G.; Conte, D.; Mariani, L.; Lo Vullo, S.; Roz, L.; Lombardo, C.; Pierotti, M.A.; Tavecchio, L. Analysis of circulating tumor DNA in plasma at diagnosis and during follow-up of lung cancer patients. Cancer Res. 2001, 61, 4675–4678. [Google Scholar]
- Diehl, F.; Schmidt, K.; Choti, M.A.; Romans, K.; Goodman, S.; Li, M.; Thornton, K.; Agrawal, N.; Sokoll, L.; Szabo, S.A.; et al. Circulating mutant DNA to assess tumor dynamics. Nat. Med. 2008, 14, 985–990. [Google Scholar] [CrossRef] [PubMed]
- Fan, T.W.M.; Zhang, X.; Wang, C.; Yang, Y.; Kang, W.-Y.; Arnold, S.; Higashi, R.M.; Liu, J.; Lane, A.N. Exosomal lipids for classifying early and late stage non-small cell lung cancer. Anal. Chim. Acta 2018, 1037, 256–264. [Google Scholar] [CrossRef] [PubMed]
- Sinha, A.; Ignatchenko, V.; Ignatchenko, A.; Mejia-Guerrero, S.; Kislinger, T. In-depth proteomic analyses of ovarian cancer cell line exosomes reveals differential enrichment of functional categories compared to the NCI60 proteome. Biochem. Biophys. Res. Commun. 2014, 445, 694–701. [Google Scholar] [CrossRef] [PubMed]
- Thierry, A.R.; Mouliere, F.; El Messaoudi, S.; Mollevi, C.; Lopez-Crapez, E.; Rolet, F.; Gillet, B.; Gongora, C.; Dechelotte, P.; Robert, B.; et al. Clinical validation of the detection of KRAS and FRAF mutations from circulating tumor DNA. Nat. Med. 2014, 20, 430–435. [Google Scholar] [CrossRef] [PubMed]
- Marrugo-Ramírez, J.; Mir, M.; Samitier, J. Blood-based cancer biomarkers in liquid biopsy: A promising non-invasive alternative to tissue biopsy. Int. J. Mol. Sci. 2018, 19, 2877. [Google Scholar] [CrossRef]
- Song, Y.; Tian, T.; Shi, Y.; Liu, W.; Zou, Y.; Khajvand, T.; Wang, S.; Zhu, Z.; Yang, C. Enrichment and single-cell analysis of circulating tumor cells. Chem. Sci. 2017, 8, 1736–1751. [Google Scholar] [CrossRef]
- Shen, Z.; Wu, A.; Chen, X. Current detection technologies for circulating tumor cells. Chem. Soc. Rev. 2017, 46, 2038–2056. [Google Scholar] [CrossRef]
- Xue, T.; Wang, S.; Ou, G.; Li, Y.; Ruan, H.; Li, Z.; Ma, Y.; Zou, R.; Qiu, J.; Shen, Z.; et al. Detection of circulating tumor cells based on improved SERS-active magnetic nanoparticles. Anal. Methods 2019, 11, 2918–2928. [Google Scholar] [CrossRef]
- Desitter, I.; Guerrouahen, B.S.; Benali-Furet, N.; Wechsler, J.; Janne, P.A.; Kuang, Y.; Yanagita, M.; Wang, L.; Berkowitz, J.A.; Distel, R.J.; et al. A new device for rapid isolation by size and characterization of rare circulating tumor cells. Anticancer Res. 2011, 31, 427–441. [Google Scholar]
- Khoo, B.L.; Grenci, G.; Lim, Y.B.; Lee, S.C.; Han, J.; Lim, C.T. Expansion of patient-derived circulating tumor cells from liquid biopsies using a ctc microfluidic culture device. Nat. Protoc. 2018, 13, 34–58. [Google Scholar] [CrossRef]
- Cho, H.; Kim, J.; Song, H.; Sohn, K.Y.; Jeon, M.; Han, K.-H. Microfluidic technologies for circulating tumor cell isolation. Analyst 2018, 143, 2936–2970. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Balasubramanian, P.; Chen, A.P.; Kummar, S.; Evrard, Y.A.; Kinders, R.J. Promise and limits of the cell search platform for evaluating pharmacodynamics in circulating tumor cells. Semin. Oncol. 2016, 43, 464–475. [Google Scholar] [CrossRef] [PubMed]
- Dobrovolskaia, M.A.; McNeil, S.E. Immunological properties of engineered nanomaterials. Nat. Nanotechnol. 2007, 2, 469–478. [Google Scholar] [CrossRef] [PubMed]
- Huang, Q.; Wang, Y.; Chen, X.; Wang, Y.; Li, Z.; Du, S.; Wang, L.; Chen, S. Nanotechnology-based strategies for early cancer diagnosis using circulating tumor cells as a liquid biopsy. Nanotheranostics 2018, 2, 21–41. [Google Scholar] [CrossRef] [PubMed]
- PramaniK, A.; Jones, S.; Gao, Y.; Sweet, C.; Vangara, A.; Begum, S.; Ray, P.C. Multifunctional hybrid graphene oxide for circulating tumor cell isolation and analysis. Adv. Drug Deliv. Rev. 2018, 125, 21–35. [Google Scholar] [CrossRef]
- Shi, Y.; Pramanik, A.; Tchounwou, C.; Pedraza, F.; Crouch, R.A.; Chavva, S.R.; Vangara, A.; Sinha, S.S.; Jones, S.; Sardar, D.; et al. Multifunctional biocompatible graphene oxide quantum dots decorated magnetic nanoplatform for efficient capture and two-photon imaging of rare tumor cells. ACS Appl. Mater. Interfaces 2015, 7, 10935–10943. [Google Scholar] [CrossRef]
- Dou, B.; Xu, L.; Jiang, B.; Yuan, R.; Xiang, Y. Aptamer-functionalized and gold nanoparticle array-decorated magnetic graphene nanosheets enable multiplexed and sensitive electrochemical detection of rare circulating tumor cells in whole blood. Anal. Chem. 2019, 91, 10792–10799. [Google Scholar] [CrossRef]
- Kim, T.H.; Yoon, H.J.; Fouladdel, S.; Wang, Y.; Kozminsky, M.; Burness, M.L.; Paoletti, C.; Zhao, L.; Azizi, E.; Wicha, M.S.; et al. Characterizing circulating tumor cells isolated from metastatic breast cancer patients using graphene oxide based microfluidic assay. Adv. Biosyst. 2019, 3, 1800278. [Google Scholar] [CrossRef]
- Yoon, H.J.; Shanker, A.; Wang, Y.; Kozminsky, M.; Jin, Q.; Palanisamy, N.; Burness, M.L.; Azizi, E.; Simeone, D.M.; Wicha, M.S.; et al. Tunable thermal-sensitive polymer–graphene oxide composite for efficient capture and release of viable circulating tumor cells. Adv. Mater. 2016, 28, 4891–4897. [Google Scholar] [CrossRef] [PubMed]
- Wu, L.; Ji, H.; Guan, Y.; Ran, X.; Ren, J.; Qu, X. A graphene-based chemical nose/tongue approach for the identification of normal, cancerous and circulating tumor cells. NPG Asia Mater. 2017, 9, e356. [Google Scholar] [CrossRef]
- Li, F.; Hu, S.; Zhang, R.; Gu, Y.; Li, Y.; Jia, Y. Porous graphene oxide enhanced aptamer specific circulating-tumor-cell sensing interface on light addressable potentiometric sensor: Clinical application and simulation. ACS Appl. Mater. Interfaces 2019, 11, 8704–8709. [Google Scholar] [CrossRef] [PubMed]
- Hu, S.; Wang, Z.; Gu, Y.; Li, Y.; Jia, Y. Clinical available circulating tumor cell assay based on tetra(4-aminophenyl) porphyrin mediated reduced graphene oxide field effect transistor. Electrochim. Acta 2019, 313, 415–422. [Google Scholar] [CrossRef]
- Li, N.; Xiao, T.; Zhang, Z.; He, R.; Wen, D.; Cao, Y.; Zhang, W.; Chen, Y. A 3D graphene oxide microchip and a Au-enwrapped silica nanocomposite-based supersandwich cytosensor toward capture and analysis of circulating tumor cells. Nanoscale 2015, 7, 16354–16360. [Google Scholar] [CrossRef] [PubMed]
- Bu, J.; Kim, Y.J.; Kang, Y.-T.; Lee, T.H.; Kim, J.; Cho, Y.-H.; Han, S.-W. Polyester fabric sheet layers functionalized with graphene oxide for sensitive isolation of circulating tumor cells. Biomaterials 2017, 125, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Yin, S.; Wu, Y.-L.; Hu, B.; Wang, Y.; Cai, P.; Tan, C.K.; Qi, D.; Zheng, L.; Leow, W.R.; Tan, N.S.; et al. Three-dimensional graphene composite macroscopic structures for capture of cancer cells. Adv. Mater. Interfaces 2014, 1, 1300043. [Google Scholar] [CrossRef]
- Wang, B.; Song, Y.; Ge, L.; Zhang, S.; Ma, L. Antibody-modified reduced graphene oxide film for circulating tumor cell detection in early-stage prostate cancer patients. RSC Adv. 2019, 9, 9379–9385. [Google Scholar] [CrossRef]
- Nahand, J.S.; Mahjoubin-Tehran, M.; Moghoofei, M.; Pourhanifeh, M.H.; Mirzaei, H.R.; Asemi, Z.; Khatami, A.; Bokharaei-Salim, F.; Mirzaei, H.; Hamblin, M.R. Exosomal miRNAs: Novel players in viral infection. Epigenomics 2020, 12, 353–370. [Google Scholar] [CrossRef]
- Hill, A.F. Extracellular vesicles and neurodegenerative diseases. J. Neurosci. 2019, 39, 9269–9273. [Google Scholar] [CrossRef]
- Cui, F.; Zhou, Z.; Zhou, H.S. Review—measurement and analysis of cancer biomarkers based on electrochemical biosensors. J. Electrochem. Soc. 2020, 167, 037525. [Google Scholar] [CrossRef]
- Fang, X.; Duan, Y.; Adkins, G.B.; Pan, S.; Wang, H.; Liu, Y.; Zhong, W. Highly efficient exosome isolation and protein analysis by an integrated nanomaterial-based platform. Anal. Chem. 2018, 90, 2787–2795. [Google Scholar] [CrossRef]
- Rider, M.A.; Hurwitz, S.N.; Meckes, D.G. Extrapeg: A polyethylene glycol-based method for enrichment of extracellular vesicles. Sci. Rep. 2016, 6, 23978. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.; Wang, C.; Li, T.; Liu, Z.; Li, L. Comparison of ultracentrifugation and density gradient separation methods for isolating Tca8113 human tongue cancer cell line-derived exosomes. Oncol. Lett. 2014, 8, 1701–1706. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Wu, H.-j.; Fine, D.; Schmulen, J.; Hu, Y.; Godin, B.; Zhang, J.X.J.; Liu, X. Ciliated micropillars for the microfluidic-based isolation of nanoscale lipid vesicles. Lab Chip 2013, 13, 2879–2882. [Google Scholar] [CrossRef] [PubMed]
- Zhang, P.; He, M.; Zeng, Y. Ultrasensitive microfluidic analysis of circulating exosomes using a nanostructured graphene oxide/polydopamine coating. Lab Chip 2016, 16, 3033–3042. [Google Scholar] [CrossRef] [PubMed]
- Lenzi, E.; Jimenez de Aberasturi, D.; Liz-Marzán, L.M. Surface-enhanced raman scattering tags for three-dimensional bioimaging and biomarker detection. ACS Sens. 2019, 4, 1126–1137. [Google Scholar] [CrossRef]
- Zou, Y.; Huang, S.; Liao, Y.; Zhu, X.; Chen, Y.; Chen, L.; Liu, F.; Hu, X.; Tu, H.; Zhang, L.; et al. Isotopic graphene–isolated-Au-nanocrystals with cellular raman-silent signals for cancer cell pattern recognition. Chem. Sci. 2018, 9, 2842–2849. [Google Scholar] [CrossRef]
- Gorgannezhad, L.; Umer, M.; Islam, M.N.; Nguyen, N.-T.; Shiddiky, M.J.A. Circulating tumor DNA and liquid biopsy: Opportunities, challenges, and recent advances in detection technologies. Lab Chip 2018, 18, 1174–1196. [Google Scholar] [CrossRef]
- Otandault, A.; Anker, P.; Al Amir Dache, Z.; Guillaumon, V.; Meddeb, R.; Pastor, B.; Pisareva, E.; Sanchez, C.; Tanos, R.; Tousch, G.; et al. Recent advances in circulating nucleic acids in oncology. Ann. Oncol. 2019, 30, 374–384. [Google Scholar] [CrossRef]
- Bellassai, N.; Spoto, G. Biosensors for liquid biopsy: Circulating nucleic acids to diagnose and treat cancer. Anal. Bioanal. Chem. 2016, 408, 7255–7264. [Google Scholar] [CrossRef]
- Peterlinz, K.A.; Georgiadis, R.M.; Herne, T.M.; Tarlov, M.J. Observation of hybridization and dehybridization of thiol-tethered DNA using two-color surface plasmon resonance spectroscopy. J. Am. Chem. Soc. 1997, 119, 3401–3402. [Google Scholar] [CrossRef]
- Das, J.; Kelley, S.O. High-performance nucleic acid sensors for liquid biopsy applications. J. Am. Chem. Soc. 2020, 59, 2554–2564. [Google Scholar]
- Compton, J. Nucleic acid sequence-based amplification. Nature 1991, 350, 91–92. [Google Scholar] [CrossRef] [PubMed]
- Demidov, V.V. Rolling Circle Amplification (RCA), 1st ed.; Springer International Publishing: Berlin/Heidelberg, Germany, 2016; p. 176. [Google Scholar]
- Saiki, R.; Gelfand, D.; Stoffel, S.; Scharf, S.; Higuchi, R.; Horn, G.; Mullis, K.; Erlich, H. Primer-directed enzymatic amplification of DNA with a thermostable DNA polymerase. Science 1988, 239, 487–491. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.; Park, S.-J.; Min, D.-H. Emerging approaches for graphene oxide biosensor. Anal. Chem. 2017, 89, 232–248. [Google Scholar] [CrossRef] [PubMed]
- Liu, B.; Salgado, S.; Maheshwari, V.; Liu, J. DNA adsorbed on graphene and graphene oxide: Fundamental interactions, desorption and applications. Curr. Opin. Colloid Interface Sci. 2016, 26, 41–49. [Google Scholar] [CrossRef]
- Yan, H.; Xu, Y.; Lu, Y.; Xing, W. Reduced graphene oxide-based solid-phase extraction for the enrichment and detection of microrna. Anal. Chem. 2017, 89, 10137–10140. [Google Scholar] [CrossRef]
- Hizir, M.S.; Balcioglu, M.; Rana, M.; Robertson, N.M.; Yigit, M.V. Simultaneous detection of circulating oncomirs from body fluids for prostate cancer staging using nanographene oxide. ACS Appl. Mater. Interfaces 2014, 6, 14772–14778. [Google Scholar] [CrossRef]
- Robertson, N.M.; Salih Hizir, M.; Balcioglu, M.; Wang, R.; Yavuz, M.S.; Yumak, H.; Ozturk, B.; Sheng, J.; Yigit, M.V. Discriminating a single nucleotide difference for enhanced miRNA detection using tunable graphene and oligonucleotide nanodevices. Langmuir 2015, 31, 9943–9952. [Google Scholar] [CrossRef]
- Cui, L.; Lin, X.; Lin, N.; Song, Y.; Zhu, Z.; Chen, X.; Yang, C.J. Graphene oxide-protected DNA probes for multiplex microRNA analysis in complex biological samples based on a cyclic enzymatic amplification method. Chem. Commun. 2012, 48, 194–196. [Google Scholar] [CrossRef]
- Wu, M.; Kempaiah, R.; Huang, P.-J.J.; Maheshwari, V.; Liu, J. Adsorption and desorption of DNA on graphene oxide studied by fluorescently labeled oligonucleotides. Langmuir 2011, 27, 2731–2738. [Google Scholar] [CrossRef]
- Huang, P.-J.J.; Liu, J. Separation of short single- and double-stranded DNA based on their adsorption kinetics difference on graphene oxide. Nanomaterials 2013, 3, 221–228. [Google Scholar] [CrossRef]
- Ruiyi, L.; Ling, L.; Hongxia, B.; Zaijun, L. Nitrogen-doped multiple graphene aerogel/gold nanostar as the electrochemical sensing platform for ultrasensitive detection of circulating free DNA in human serum. Biosens. Bioelectron. 2016, 79, 457–466. [Google Scholar] [CrossRef]
- Huang, H.; Bai, W.; Dong, C.; Guo, R.; Liu, Z. An ultrasensitive electrochemical DNA biosensor based on graphene/Au nanorod/polythionine for human papillomavirus DNA detection. Biosens. Bioelectron. 2015, 68, 442–446. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.W.; Kim, S.; Jang, Y.-h.; Lim, K.-i.; Lee, W.H. Attomolar detection of virus by liquid coplanar-gate graphene transistor on plastic. Nanotechnology 2019, 30, 345502. [Google Scholar] [CrossRef] [PubMed]
- Omar, N.A.S.; Fen, Y.W.; Abdullah, J.; Sadrolhosseini, A.R.; Mustapha Kamil, Y.; Fauzi, N.I.M.; Hashim, H.S.; Mahdi, M.A. Quantitative and selective surface plasmon resonance response based on a reduced graphene oxide–polyamidoamine nanocomposite for detection of dengue virus e-proteins. Nanomaterials 2020, 10, 569. [Google Scholar] [CrossRef] [PubMed]
- Mahase, E. Coronavirus: Covid-19 has killed more people than sars and mers combined, despite lower case fatality rate. BMJ 2020, 368, m641. [Google Scholar] [CrossRef]
- Gentile, D.; Patamia, V.; Scala, A.; Sciortino, M.T.; Piperno, A.; Rescifina, A. Putative inhibitors of SARS-CoV-2 main protease from a library of marine natural products: A virtual screening and molecular modeling study. Mar. Drugs 2020, 18, 225. [Google Scholar] [CrossRef]
- Hogan, A.C.; Caya, C.; Papenburg, J. Rapid and simple molecular tests for the detection of respiratory syncytial virus: A review. Expert Rev. Mol. Diagn. 2018, 18, 617–629. [Google Scholar] [CrossRef]
- Udugama, B.; Kadhiresan, P.; Kozlowski, H.N.; Malekjahani, A.; Osborne, M.; Li, V.Y.C.; Chen, H.; Mubareka, S.; Gubbay, J.B.; Chan, W.C.W. Diagnosing covid-19: The disease and tools for detection. ACS Nano 2020, 14, 3822–3835. [Google Scholar] [CrossRef]
- Cheng, M.P.; Papenburg, J.; Desjardins, M.; Kanjilal, S.; Quach, C.; Libman, M.; Dittrich, S.; Yansouni, C.P. Diagnostic testing for severe acute respiratory syndrome–related coronavirus-2: A narrative review. Ann. Intern. Med. 2020. [Google Scholar] [CrossRef]
- Qiu, G.; Gai, Z.; Tao, Y.; Schmitt, J.; Kullak-Ublick, G.A.; Wang, J. Dual-functional plasmonic photothermal biosensors for highly accurate severe acute respiratory syndrome coronavirus 2 detection. ACS Nano 2020. [Google Scholar] [CrossRef] [PubMed]
- Seo, G.; Lee, G.; Kim, M.J.; Baek, S.-H.; Choi, M.; Ku, K.B.; Lee, C.-S.; Jun, S.; Park, D.; Kim, H.G.; et al. Rapid detection of covid-19 causative virus (SARS -COV-2) in human nasopharyngeal swab specimens using field-effect transistor-based biosensor. ACS Nano 2020, 14, 5135–5142. [Google Scholar] [CrossRef] [PubMed]
- Barreca, D.; Neri, G.; Scala, A.; Fazio, E.; Gentile, D.; Rescifina, A.; Piperno, A. Covalently immobilized catalase on functionalized graphene: Effect on the activity, immobilization efficiency, and tetramer stability. Biomater. Sci. 2018, 6, 3231–3240. [Google Scholar] [CrossRef] [PubMed]
- Piperno, A.; Mazzaglia, A.; Scala, A.; Pennisi, R.; Zagami, R.; Neri, G.; Torcasio, S.M.; Rosmini, C.; Mineo, P.G.; Potara, M.; et al. Casting light on intracellular tracking of a new functional graphene-based microrna delivery system by flim and raman imaging. ACS Appl. Mater. Interfaces 2019, 11, 46101–46111. [Google Scholar] [CrossRef]
- Morales-Narváez, E.; Dincer, C. The impact of biosensing in a pandemic outbreak: Covid-19. Biosens. Bioelectron. 2020, 163, 112274. [Google Scholar] [CrossRef]
- Palmieri, V.; Papi, M. Can graphene take part in the fight against covid-19? Nano Today 2020, 100883. [Google Scholar] [CrossRef]
- Bruch, R.; Urban, G.A.; Dincer, C. Unamplified gene sensing via cas9 on graphene. Nat. Biomed. Eng. 2019, 3, 419–420. [Google Scholar] [CrossRef]
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Cordaro, A.; Neri, G.; Sciortino, M.T.; Scala, A.; Piperno, A. Graphene-Based Strategies in Liquid Biopsy and in Viral Diseases Diagnosis. Nanomaterials 2020, 10, 1014. https://doi.org/10.3390/nano10061014
Cordaro A, Neri G, Sciortino MT, Scala A, Piperno A. Graphene-Based Strategies in Liquid Biopsy and in Viral Diseases Diagnosis. Nanomaterials. 2020; 10(6):1014. https://doi.org/10.3390/nano10061014
Chicago/Turabian StyleCordaro, Annalaura, Giulia Neri, Maria Teresa Sciortino, Angela Scala, and Anna Piperno. 2020. "Graphene-Based Strategies in Liquid Biopsy and in Viral Diseases Diagnosis" Nanomaterials 10, no. 6: 1014. https://doi.org/10.3390/nano10061014
APA StyleCordaro, A., Neri, G., Sciortino, M. T., Scala, A., & Piperno, A. (2020). Graphene-Based Strategies in Liquid Biopsy and in Viral Diseases Diagnosis. Nanomaterials, 10(6), 1014. https://doi.org/10.3390/nano10061014