Internalization and Viability Studies of Suspended Nanowire Silicon Chips in HeLa Cells
Abstract
1. Introduction
2. Materials and Methods
2.1. Technology for the Fabrication of Microchips Decorated with Silicon Nanowires and Isolated Silicon Nanowire Entangled Meshes
2.2. Nanowire Growth Method Assisted by Catalyst Gold Deposition
2.3. Cell Culture and Silicon Chips Internalization Methodologies
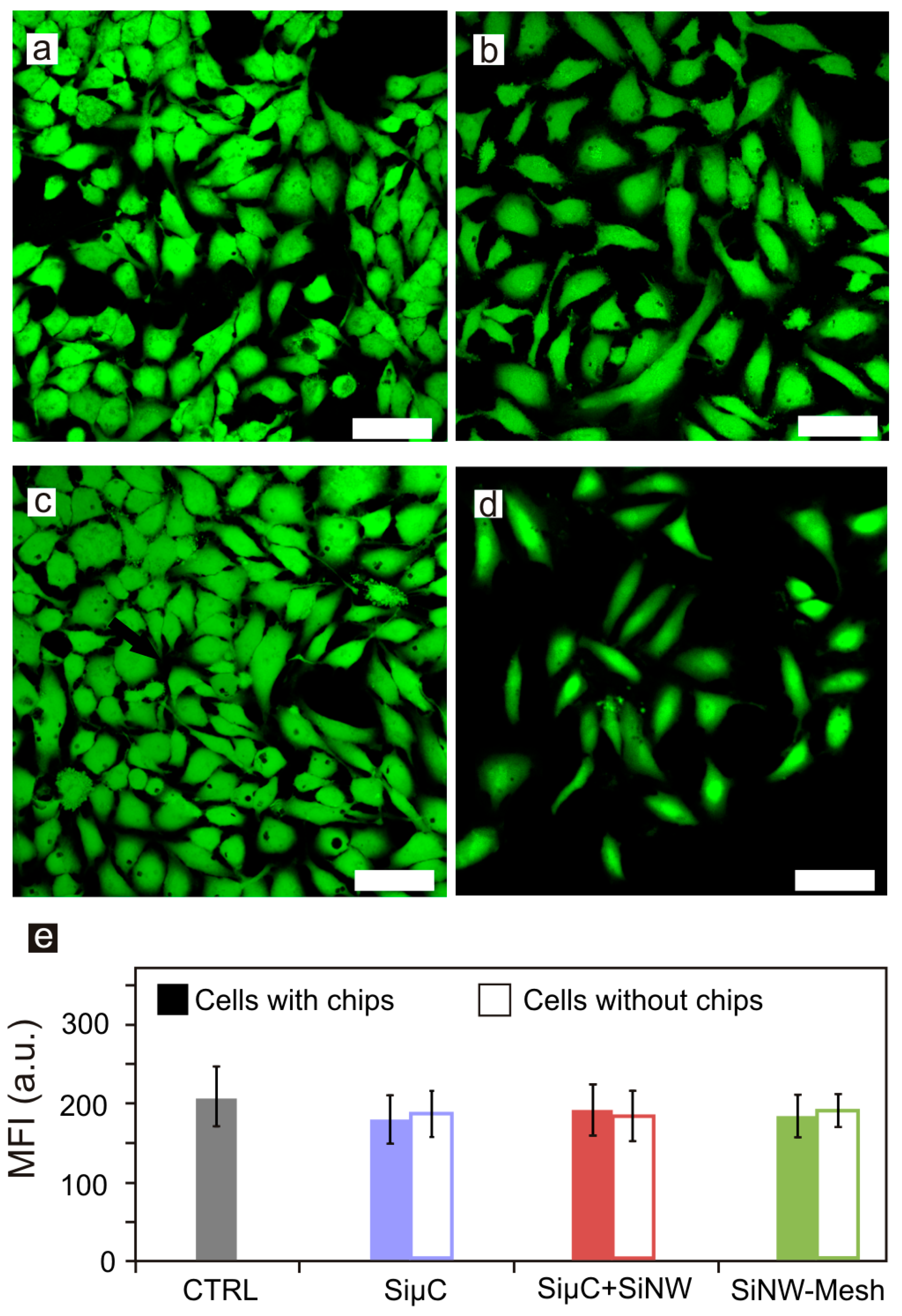
2.4. Living Cells Confocal Microscopy and Viability Assay
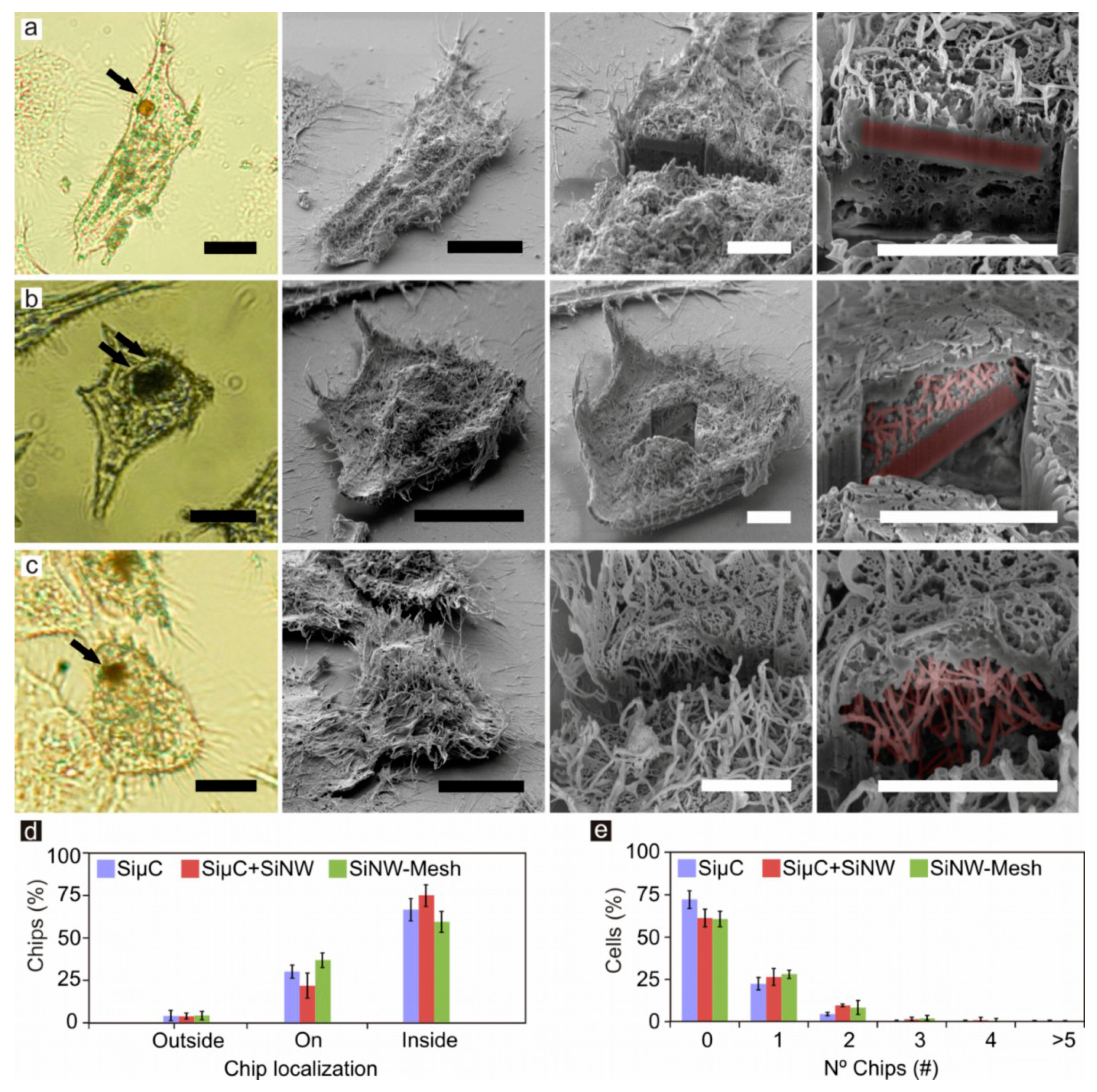
2.5. Correlative Light and Electron Microscopy of Silicon Chips in HeLa Cells
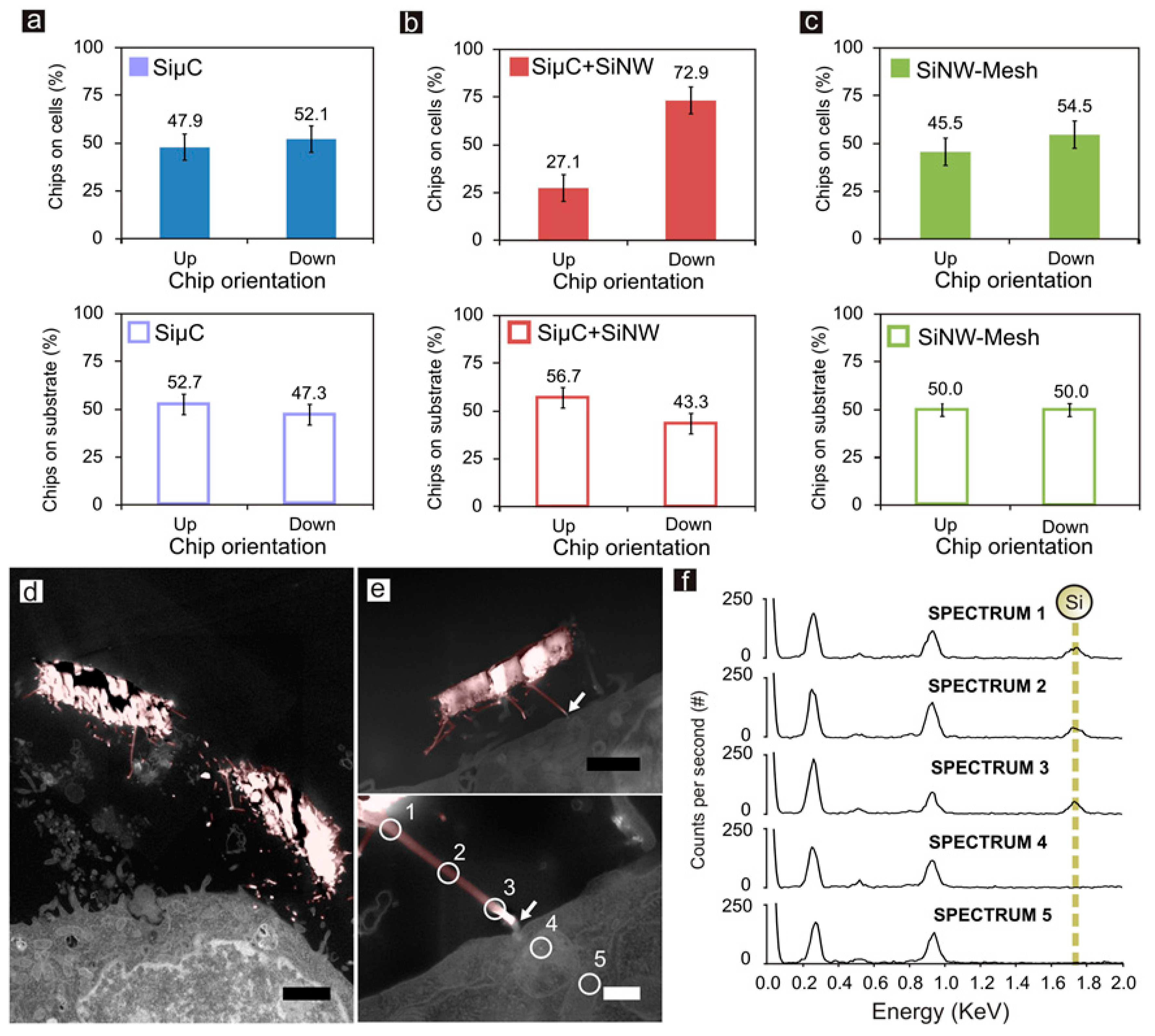
2.6. EDX Analysis of Chips and HeLa Cells
2.7. Sedimentation Tests of the Silicon Chips
3. Results
3.1. Device Fabrication and Morphological Studies
3.2. Cell Viability Assays
3.3. Chip Internalization Studies in HeLa Cells
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Whitesides, G.M. The “right” size in nanobiotechnology. Nat. Biotechnol. 2003, 21, 1161–1165. [Google Scholar] [CrossRef] [PubMed]
- Singhal, R.; Orynbayeva, Z.; Sundaram, R.V.K.; Niu, J.J.; Bhattacharyya, S.; Vitol, E.A.; Schrlau, M.G.; Papazoglou, E.S.; Friedman, G.; Gogotsi, Y. Multifunctional carbon-nanotube cellular endoscopes. Nat. Nanotechnol. 2011, 6, 57–64. [Google Scholar] [CrossRef] [PubMed]
- Tian, B.; Lieber, C.M. Synthetic nanoelectronic probes for biological cells and tissues. Annu. Rev. Anal. Chem. 2013, 6, 31–51. [Google Scholar] [CrossRef] [PubMed]
- El-Ali, J.; Sorger, P.K.; Jensen, K.F. Cells on chips. Nature 2006, 442, 403–411. [Google Scholar] [CrossRef]
- Cross, S.E.; Jin, Y.-S.; Rao, J.; Gimzewski, K.J. Nanomechanical analysis of cells from cancer patients. Nat. Nanotechnol. 2007, 2, 780–783. [Google Scholar] [CrossRef]
- Shalek, A.K.; Robinson, J.T.; Karp, E.S.; Lee, J.S.; Ahn, D.-R.M.; Yoon, H.; Sutton, A.; Jorgolli, M.; Gertner, R.S.; Gujral, T.S.; et al. Vertical silicon nanowires as a universal platform for delivering biomolecules into living cells. Proc. Natl. Acad. Sci. USA 2010, 107, 1870–1875. [Google Scholar] [CrossRef]
- Kim, W.; Ng, J.K.; Kunitake, M.E.; Conklin, B.R.; Yang, P. Interfacing silicon nanowires with mammalian cells. J. Am. Chem. Soc. 2007, 129, 7228–7229. [Google Scholar] [CrossRef]
- Fan, J.; Chu, P.K. Group IV Nanoparticles: Synthesis, properties, and biological applications. Small 2010, 6, 2080–2098. [Google Scholar] [CrossRef]
- Fernández-Rosas, E.; Gómez-Martínez, R.; Ibáñez, E.; Barrios, L.; Duch, M.; Esteve, J.; Nogués, C.; Plaza, J.A. Intracellular Polysilicon Barcodes for Cell Tracking. Small 2009, 5, 2433–2439. [Google Scholar] [CrossRef]
- Penon, O.; Novo, S.; Durán, S.; Ibáñez, E.; Nogués, C.; Samitier, J.; Duch, M.; Plaza, J.A.; Pérez-García, L. Efficient Biofunctionalization of Polysilicon Barcodes for Adhesion to the Zona Pellucida of Mouse Embryos. Bioconjugate Chem. 2012, 23, 2392–2402. [Google Scholar] [CrossRef]
- Gómez-Martínez, R.; Vázquez, P.; Duch, M.; Muriano, A.; Pinacho, D.; Sanvicens, N.; Sánchez-Baeza, F.; Boya, P.; de la Rosa, E.J.; Esteve, J.; et al. Intracellular Silicon Chips in Living Cells. Small 2010, 6, 499–502. [Google Scholar]
- Gómez-Martínez, R.; Hernández-Pinto, A.M.; Duch, M.; Vázquez, P.; Zinoviev, K.; de la Rosa, E.J.; Esteve, J.; Suárez, T.; Plaza, J.A. Silicon chips detect intracellular pressure changes in living cells. Nat. Nanotechnol. 2013, 8, 517–521. [Google Scholar] [CrossRef] [PubMed]
- Fan, H.J.; Werner, P.; Zacharias, M. Semiconductor nanowires: From self-organization to patterned growth. Small 2006, 2, 700–717. [Google Scholar] [CrossRef] [PubMed]
- Cui, Y.; Wei, Q.; Park, H.; Lieber, C.M. Nanowire nanosensors for highly sensitive and selective detection of biological and chemical species. Science 2001, 293, 1289–1292. [Google Scholar] [CrossRef]
- Schmidt, V.; Riel, H.; Senz, S.; Karg, S.; Riess, W.; Gösele, U. Realization of a silicon nanowire vertical surround-gate field-effect transistor. Small 2006, 2, 85–88. [Google Scholar] [CrossRef]
- Tian, B.; Cohen-Karni, T.; Qing, Q.; Duan, X.; Xie, P.; Lieber, C.M. Three-dimensional, flexible nanoscale field-effect transistors as localized bioprobes. Science 2010, 329, 830–834. [Google Scholar] [CrossRef]
- Gil-Santos, E.; Ramos, D.; Martínez, J.; Fernández-Regúlez, M.; García, R.; San Paulo, A.; Calleja, M.; Tamayo, J. Nanomechanical mass sensing and stiffness spectrometry based on two-dimensional vibrations of resonant nanowires. Nat. Nanotechnol. 2010, 5, 641–645. [Google Scholar] [CrossRef]
- San Paulo, A.; Arellano, N.; Plaza, J.A.; He, R.; Carraro, C.; Maboudian, R.; Howe, R.T.; Bokor, J.; Yang, P. Suspended mechanical structures based on elastic silicon nanowire arrays. Nano Lett. 2007, 7, 1100–1104. [Google Scholar] [CrossRef]
- Zheng, G.; Patolsky, F.; Cui, Y.; Wang, W.U.; Lieber, C.M. Multiplexed electrical detection of cancer markers with nanowire sensor arrays. Nat. Biotechnol. 2005, 23, 1294–1301. [Google Scholar] [CrossRef]
- Ramgir, N.S.; Yang, Y.; Zacharias, M. Nanowire-based sensors. Small 2010, 6, 1705–1722. [Google Scholar] [CrossRef]
- Lieber, C.M.; Wang, Z.L. Functional Nanowires. MRS Bull. 2007, 32, 99–108. [Google Scholar] [CrossRef]
- Zimmerman, J.F.; Parameswaran, R.; Murray, G.; Wang, Y.; Burke, M.; Tian, B. Cellular uptake and dynamics of unlabeled freestanding silicon nanowires. Sci. Adv. 2016, 2, e1601039. [Google Scholar] [CrossRef] [PubMed]
- Mumm, F.; Beckwith, K.M.; Bonde, S.; Karen, L.; Martinez, P. A transparent nanowire-based cell impalement device suitable for detailed cell nanowire interaction studies. Small 2013, 9, 263–272. [Google Scholar] [CrossRef] [PubMed]
- Hou, S.; Zhao, H.; Zhao, L.; Shen, Q.; Wei, K.S.; Suh, D.Y.; Nakao, A.; Garcia, M.A.; Song, M.; Lee, T.; et al. Capture and stimulated release of circulating tumor cells on polymer-grafted silicon nanostructures. Adv. Mater. 2012, 25, 1547–1551. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.; Wang, H.; Jiao, J.; Chen, K.-J.; Owens, G.E.; Kamei, K.; Sun, J.; Sherman, D.J.; Behrenbruch, C.P.; Wu, H.; et al. Three-dimensional nanostructured substrates toward efficient capture of circulating tumor cells. Angew. Chem. Int. Ed. Engl. 2009, 48, 8970. [Google Scholar] [CrossRef] [PubMed]
- Jokilaakso, N.; Salm, E.; Chen, A.; Millet, L.; Guevara, C.D.; Dorvel, B.; Reddy, B., Jr.; Karlstrom, A.E.; Chen, Y.; Ji, H.; et al. Ultralocalized single cell electroporation using silicon nanowires. Lab Chip 2013, 13, 336–339. [Google Scholar] [CrossRef]
- Wierzbicki, R.; Købler, C.; Jensen, M.R.B.; Łopacinska, J.; Schmidt, M.S.; Skolimowski, M.; Abeille, F.; Qvortrup, K.; Mølhave, K. Mapping the complex morphology of cell interactions with nanowire substrates using FIB-SEM. PLoS ONE 2013, 8, e53307. [Google Scholar] [CrossRef]
- Zhang, W.; Tong, L.; Yang, C. Cellular binding and internalization of functionalized silicon nanowires. Nano Lett. 2012, 12, 1002–1006. [Google Scholar] [CrossRef]
- Cui, Y.; Zhong, Z.; Wang, D.; Wang, W.U.; Lieber, C.M. High performance silicon nanowire field effect transistors. Nano Lett. 2003, 3, 149–152. [Google Scholar] [CrossRef]
- Magagnin, L.; Bertani, V.; Cavallotti, L.; Maboudian, R.; Carraro, C. Selective deposition of gold nanoclusters on silicon by a galvanic displacement process. Microelectron. Eng. 2002, 64, 479–485. [Google Scholar] [CrossRef]
- Kamins, T. Polycrystalline Silicon for Integrated Circuit Applications; Springer: Norwell, MA, USA, 1988; Volume 45. [Google Scholar]
- Gratton, S.E.A.; Ropp, P.A.; Pohlhaus, P.D.; Luft, J.C.; Madden, V.J.; Napier, M.E.; DeSimone, J.M. The effect of particle design on cellular internalization. Proc. Natl. Acad. Sci. USA 2008, 105, 11613–11618. [Google Scholar] [CrossRef] [PubMed]
- Fernández-Rosas, E.; Gómez-Martínez, R.; Ibañez, E.; Barrios, L.; Duch, M.; Esteve, J.; Plaza, J.A.; Nogués, C. Internalization and cytotoxicity analysis of silicon-based microparticles in macrophages and embryos. Biomed. Microdev. 2010, 12, 371–379. [Google Scholar] [CrossRef] [PubMed]
- Xie, X.; Xu, A.M.; Angle, M.R.; Tayebi, N.; Verma, P.; Melosh, N.A. Mechanical model of vertical nanowire cell penetration. Nano Lett. 2003, 13, 6002–6008. [Google Scholar] [CrossRef]
- Berthing, T.; Bonde, S.; Sørensen, C.B.; Utko, P.; Nygård, J.; Martinez, K.L. Intact mammalian cell function on semiconductor nanowire arrays: New perspectives for cell-based biosensing. Small 2011, 7, 640–647. [Google Scholar] [CrossRef] [PubMed]
- McNear, K.; Huang, Y.; Yang, C. Understanding cellular internalization pathways of silicon nanowires. J. Nanobiotechnol. 2017, 15, 17. [Google Scholar] [CrossRef] [PubMed]
- Robinson, J.T.; Jorgolli, M.; Shalek, A.K.; Yoon, M.-H.; Gertner, R.S.; Park, H. Vertical nanowire electrode arrays as a scalable platform for intracellular interfacing to neuronal circuits. Nat. Nanotechnol. 2012, 7, 180–184. [Google Scholar] [CrossRef]
- Chithrani, B.D.; Ghazani, A.A.; Chan, W.C.W. Determining the Size and Shape Dependence of Gold Nanoparticle Uptake into Mammalian Cells. Nano Lett. 2006, 6, 662–668. [Google Scholar] [CrossRef]
- Fischer, H.W.; Cooper, T.W. Electron microscope studies of the microvilli of HeLa cells. J. Cell Biol. 1967, 34, 569–576. [Google Scholar] [CrossRef]
- Fischer, K.E.; Jayagopal, A.; Nagaraj, G.; Daniels, R.H.; Li, E.M.; Silvestrini, M.T.; Desai, T.A. Nanoengineered surfaces enhance drug loading and adhesion. Nano Lett. 2011, 11, 1076–1081. [Google Scholar] [CrossRef]
- Durán, S. Suspended Micro-and Nanotools for Cell Biology. Ph.D. Thesis, Universidad Autónoma de Barcelona, Barcelona, Spain, December 2014. [Google Scholar]
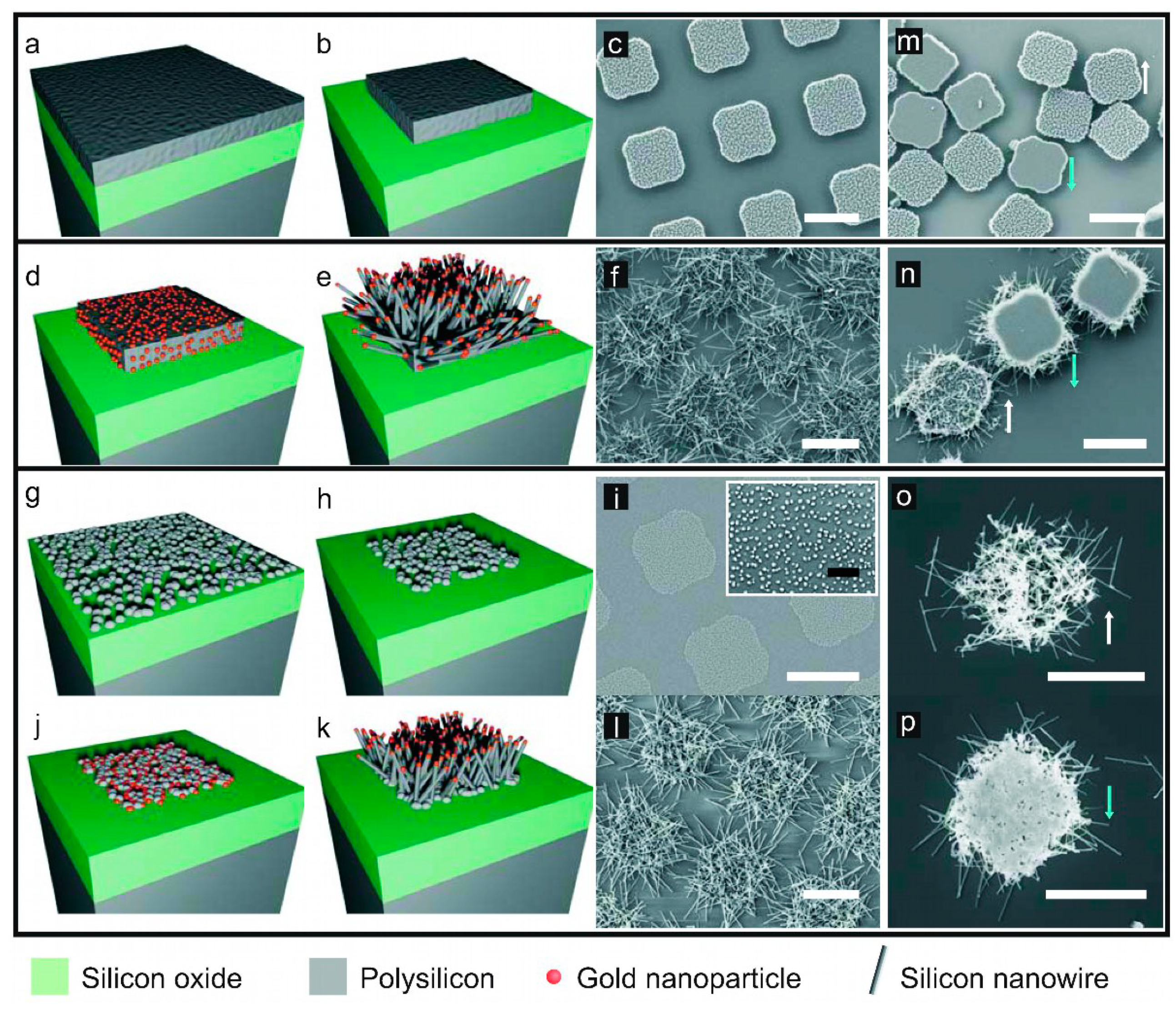


| Ø (nm) | Density (clusters/µm2) | ||
|---|---|---|---|
| Polysilicon nanoclusters | 65 ± 12 | 128 ± 8 | |
| Length (nm) | Width (nm) | Density (NWs/µm2) | |
| SiµC+SiNWs | 885 ± 231 | 60 ± 19 | 28 ± 2 |
| SiNW-Mesh | 1124 ± 405 | 71 ± 16 | 47 ± 2 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Durán, S.; Duch, M.; Gómez-Martínez, R.; Fernández-Regúlez, M.; Agusil, J.P.; Reina, M.; Müller, C.; San Paulo, Á.; Esteve, J.; Castel, S.; et al. Internalization and Viability Studies of Suspended Nanowire Silicon Chips in HeLa Cells. Nanomaterials 2020, 10, 893. https://doi.org/10.3390/nano10050893
Durán S, Duch M, Gómez-Martínez R, Fernández-Regúlez M, Agusil JP, Reina M, Müller C, San Paulo Á, Esteve J, Castel S, et al. Internalization and Viability Studies of Suspended Nanowire Silicon Chips in HeLa Cells. Nanomaterials. 2020; 10(5):893. https://doi.org/10.3390/nano10050893
Chicago/Turabian StyleDurán, Sara, Marta Duch, Rodrigo Gómez-Martínez, Marta Fernández-Regúlez, Juan Pablo Agusil, Manuel Reina, Claudia Müller, Álvaro San Paulo, Jaume Esteve, Susana Castel, and et al. 2020. "Internalization and Viability Studies of Suspended Nanowire Silicon Chips in HeLa Cells" Nanomaterials 10, no. 5: 893. https://doi.org/10.3390/nano10050893
APA StyleDurán, S., Duch, M., Gómez-Martínez, R., Fernández-Regúlez, M., Agusil, J. P., Reina, M., Müller, C., San Paulo, Á., Esteve, J., Castel, S., & Plaza, J. A. (2020). Internalization and Viability Studies of Suspended Nanowire Silicon Chips in HeLa Cells. Nanomaterials, 10(5), 893. https://doi.org/10.3390/nano10050893





