1. Introduction
In recent days, many applications involve the role of extracting useful information for data collection. The extracted information is known as feature, and it is useful in describing the target concept [
1]. However, an increment in the number of features will cause the “curse of dimensionality” in which the performance of the system is degraded and becomes complex. This is mainly due to the existence of irrelevant and redundant information, which badly affects the performance of the classification model [
2]. To resolve the issue above, a proper selection of extracted features is critically important. Hence, the feature selection problem has become one of the major concerns in most of the research areas [
3].
Feature selection is the pre-processing step to determine a subset of significant features that can strongly improve the performance of the system. It not only eliminates the redundant information, but also reduces the temporal and spatial complexity of the classification model [
4]. Generally, feature selection can be classified into two approaches: filter and wrapper. The former identifies the relevant features by using the proxy measure, mutual information, and data characteristics, while the later utilizes a predictive model to train the feature set for evaluating the nearly optimal feature subset [
5,
6]. As compared to the wrapper, filter feature selection is independent of the learning algorithm, and it is computationally less expensive. However, wrapper feature selection can usually offer better performance [
7].
As for the wrapper approach, the feature selection is considered as a combinatorial optimization problem, which can be solved by using metaheuristic algorithms [
8,
9]. The most common wrapper feature selection methods are the genetic algorithm (GA) and binary particle swarm optimization (BPSO). The GA is an evolutionary algorithm that generates the population of solutions called chromosomes. For each generation, the solutions are evolved based on the selection, crossover, and mutation operations [
10]. Several studies have shown that GA is good for the high dimensional feature selection problem [
10,
11]. However, the GA suffers from time consumption and parameter setting. Thus, Ghareb et al. [
12] performed the hybridization between an enhanced genetic algorithm (EGA) and a filter approach for text categorization. The authors first employed the filter approach to identify the potential initial solutions, and the solutions are then evaluated by the EGA. Ma and Xia [
13] introduced a novel tribe competition-based genetic algorithm (TCbGA) to tackle the feature selection problem in pattern classification. Another study proposed the adaptive multi-parent crossover GA for epileptic seizure identification [
14].
BPSO is a binary variant of particle swarm optimization (PSO). Unlike GA, BPSO is a swarm-based algorithm that generates the population of solutions called particles. The particles adjust their positions by changing their velocities according to their own experience, as well as the experience of their neighbors [
15]. BPSO is a useful tool and it has been widely applied for feature selection. However, BPSO has the disadvantages of premature convergence and stagnation, thus leading to ineffective solutions [
15,
16]. Therefore, Chuang et al. [
17] proposed a chaotic binary particle swarm optimization (CBPSO) for feature selection in which the chaotic maps were implemented for identifying the inertia weight in each iteration. Jain et al. [
18] developed an improved binary particle swarm optimization (iBPSO) for gene selection and cancer classification. The authors first applied the correlation-based feature selection (CFS) to reduce the dimensions, and then evaluated the relevant features using iBPSO. Another study introduced the BPSO with the personal best (
pbest) guide strategy to tackle the feature selection problem in electromyography signals classification [
19].
Competitive swarm optimizer (CSO) is a newly introduced variant of PSO [
20]. In comparison with other metaheuristic algorithms, CSO has shown superior performance in several benchmark tests. Generally, CSO employs the competition strategy that partitioned the solutions into winners and losers in which the winners are directly moved to the next iteration. In this way, CSO is computationally less expensive since only half of the population is used in the evaluations. In this paper, we propose the binary version of CSO to solve the feature selection problem in classification tasks. The binary version introduced here is performed by implementing the transfer functions. In this approach, the transfer functions from S-shaped and V-shaped families are used to allow the search agents to move around the binary search space. The proposed approaches are validated with 15 benchmark datasets, and the results are compared with other conventional methods.
The organization of this paper as the following:
Section 2 details the background of the competitive swarm optimizer (CSO).
Section 3 presents the proposed binary version of the competitive swarm optimizer (BCSO) and
Section 4 describes the application of BCSO in feature selection. The experimental results are discussed in
Section 5. Finally, conclusions are offered in
Section 6.
2. The Competitive Swarm Optimizer
The competitive swarm optimizer (CSO) is a recent metaheuristic optimization algorithm proposed by Cheng and Jin in 2015 [
20]. The CSO is a new variant of particle swarm optimization (PSO), and it has been proven to work more effectively on large-scale optimization. In addition, the CSO is able to find the global optimum in a very short period, which leads to fast computational speed. In the CSO, the population of particles is randomly partitioned into two groups with equal size. The competition is then made between the particles from each group. From the competition, the particle that scores a better fitness value is known as the winner and it is directly moved to the new iteration. On the contrary, the loser updates its velocity and position by learning from the winner. Mathematically, the velocity and position of loser is updated as follows:
where
vl is the velocity of loser particle,
xw is the position of the winner particle,
xl is the position of the loser particle,
is the mean position of the current swarm,
r1,
r2, and
r3 are three independent random vectors in [0, 1],
is the social factor,
d is the dimension of search space, and
t is the iteration number. The pseudocode of the CSO is presented in Algorithm 1.
| Algorithm 1. Competitive swarm optimizer |
| Input parameter:N, Tmax and |
| (1) Initialize a population of particles, x |
| (2) Calculate the fitness of particles, F(x) |
| (3) Define the best particle as gbest |
| (4) for t = 1 to maximum number of iterations, Tmax |
| // Competition Strategy // |
| (5) for i = 1 to half of population, N/2 |
| (6) Random select two particles, xk and xm |
| (7) if F(xk) better than F(xm) |
| (8) xw = xk, xl = xm |
| (9) else |
| (10) xw = xm, xl = xk |
| (11) end if |
| (12) Add xw into new population |
| (13) Remove xk and xm from the population |
| (14) next i |
| //Velocity and Position Update // |
| (15) for i = 1 to half of population, N/2 |
| (16) for d = 1 to the dimension of search space, D |
| (17) Update velocity of loser using Equation (1) |
| (18) Update position of loser as shown in Equation (2) |
| (19) next d |
| (20) Calculate the fitness of new loser, F(xl) |
| (21) Move new loser into new population |
| (22) Update gbest if there is better solution |
| (23) next i |
| (24) Pass new population to next iteration |
| (25) next t |
| Output: Global best solution |
3. Binary Version of the Competitive Swarm Optimizer
The CSO is a swarm intelligent method that mimics the concept of competition between particles in the population. As mentioned in [
20], the CSO has been tested on several benchmark functions, and it showed superior performance against other conventional optimization algorithms. The CSO algorithm utilizes the competition strategy and new velocity updating rule, which is beneficial in improving the exploration and convergence rate [
20]. This motivates us to model the CSO so that it can be useful for wrapper-based feature selection.
Generally speaking, wrapper feature selection is considered as a binary optimization problem. In wrapper feature selection, the solution is represented as either 0 or 1 [
8]. In the traditional CSO, the particles are moved around the search space by updating their positions within the continuous real domain. However, the real continuous value is not suitable for binary optimization since the solution should be represented in binary form. For such a reason, the CSO is modeled into the binary version.
One of the effective ways to convert the continuous optimization into a binary version is the utilization of a transfer function. In binary optimization, a transfer function is a mathematical function that determines the probability of changing a position vector’s dimension from 0 to 1, and vice versa [
21]. More importantly, a transfer function is an extremely cheap operator, and it can improve the exploitation and exploration of the CSO in feature selection [
22]. Hence, the transfer function has become our main focus in this work. In this paper, we propose eight versions of binary competitive swarm optimizers for feature selection.
3.1. S-Shaped Family
In general, the transfer function can be categorized into S-shaped and V-shaped families. In this sub-section, the implementation of the S-shaped transfer function is described. The S-shaped transfer function forces the search agents to move around the binary search space [
8]. Previously, S-shaped transfer functions have been successfully applied in binary particle swarm optimization (BPSO), binary antlion optimizer (BALO) and binary salp swarm algorithm (BSSA) [
8,
21,
23]. The four commonly used S-shaped transfer functions (
S1–
S4) are expressed as follows:
where
vl is the velocity of loser particle,
d is the dimension, and
t is the iteration number. The illustrations of the S-shaped transfer functions are presented in
Figure 1. In these approaches, the velocity of the loser is first calculated as shown in Equation (1). The transfer function is then used to convert the velocity into a probability value between [0, 1]. After that, the position of the loser is updated as:
where
S can be
S1,
S2,
S3, or
S4 and
r4 is a random vector distributed in [0, 1].
3.2. V-Shaped Family
In this sub-section, the implementation of the V-shaped transfer function is presented. The V-shaped transfer function allows the search agents to perform the search within the binary search space. Many studies employ the V-shaped transfer function to convert the metaheuristic algorithms into a binary version [
23,
24,
25]. The four frequently used V-shaped transfer functions (
V1–
V4) are defined as follows:
where
vl is the velocity of loser particle,
d is the dimension, and
t is the iteration number. The illustrations of the V-shaped transfer functions are shown in
Figure 2. Unlike the S-shaped transfer function, the V-shaped transfer function does not force the search agents to move on the binary search space. In this approach, the position of loser particle is updated as:
where
V can be
V1,
V2,
V3, or
V4 and
r5 is a random vector distributed in [0, 1].
The pseudocode of the binary competitive swarm optimizer (BCSO) is shown in Algorithm 2. N and Tmax are the number of particles and the maximum number of iterations. In the first step, a population of N particles is randomly initialized, and the velocity of each particle is initialized as zero. Then, the fitness of each particle is evaluated. The best particle is defined as the global best, gbest. For each iteration, the particles are randomly divided into two groups, and the competition is made between two coupled particles. From the competition, the winners are directly passed into the new population. On the other hand, the losers update their velocity using Equation (1). Then, the velocity is converted into a probability value by employing S-shaped or V-shaped transfer functions. Afterward, the position of the loser particle is updated using Equation (7) or Equation (12). Next, the fitness of each new loser is evaluated, and the new losers are moved into the new population for the next iteration. At the end of each iteration, the global best solution gbest is updated. The procedure is repeated iteratively until the maximum number of iterations is reached. Finally, the global best solution is achieved.
| Algorithm 2. Binary competitive swarm optimizer. |
| Input parameter:N, Tmax and |
| (1) Initialize a population of particles, x |
| (2) Calculate the fitness of particles, F(x) |
| (3) Define the best particle as gbest |
| (4) for t = 1 to maximum number of iterations, Tmax |
| // Competition Strategy // |
| (5) for i = 1 to half of population, N/2 |
| (6) Random select two particles, xk and xm |
| (7) if F(xk) better than F(xm) |
| (8) xw = xk, xl = xm |
| (9) else |
| (10) xw = xm, xl = xk |
| (11) end if |
| (12) Add xw into new population |
| (13) Remove xk and xm from the population |
| (14) next i |
| //Velocity and Position Update // |
| (15) for i = 1 to half of population, N/2 |
| (16) for d = 1 to the dimension of search space, D |
| (17) Update velocity of loser using Equation (1) |
| (18) Convert velocity into probability using S-shaped or V-shaped transfer function |
| (19) Update position of loser as shown in Equation (7) or Equation (12) |
| (20) next d |
| (21) Calculate the fitness of new loser, F(xl) |
| (22) Move new loser into new population |
| (23) Update gbest if there is better solution |
| (24) next i |
| (25) Pass new population to next iteration |
| (26) next t |
| Output: Global best solution |
4. Application of the Binary Competitive Swarm Optimizer for Feature Selection
In this section, the proposed binary competitive swarm optimization approaches are applied to solve the feature selection problem in classification tasks. In wrapper feature selection, the solution is represented in binary form. Bit 1 indicates that the feature is selected, while bit 0 denotes the unselected feature [
23]. For example, let solution
X = {0, 1, 0, 0, 1, 0, 0, 0, 1, 1}. As can be seen, solution
X consists of 10 dimensions (features). Among them, only four features (2nd, 5th, 9th, and 10th) are selected.
Feature selection is an
NP-hard combinatorial problem. For a dataset with feature size
D, the possible combination of feature subsets will be 2
D – 1, which is impractical for searching exhaustively. Therefore, the proposed approaches are used to evaluate the best feature subset. In this paper, the fitness function that considers both classification error rate and number of features is applied. Mathematically, the fitness function can be expressed as:
where
ER(
K) is the classification error rate computed by a classifier relative to selection decision
K of the features, |
C| is the total number of features in the dataset, |
S| is the length of selected feature subset, and
α is the parameter in [0, 1] that controls the influence of the classification error rate. According to [
8,
23,
26], the
a is set at 0.99 since classification performance is the most important measure in this framework.
6. Conclusions
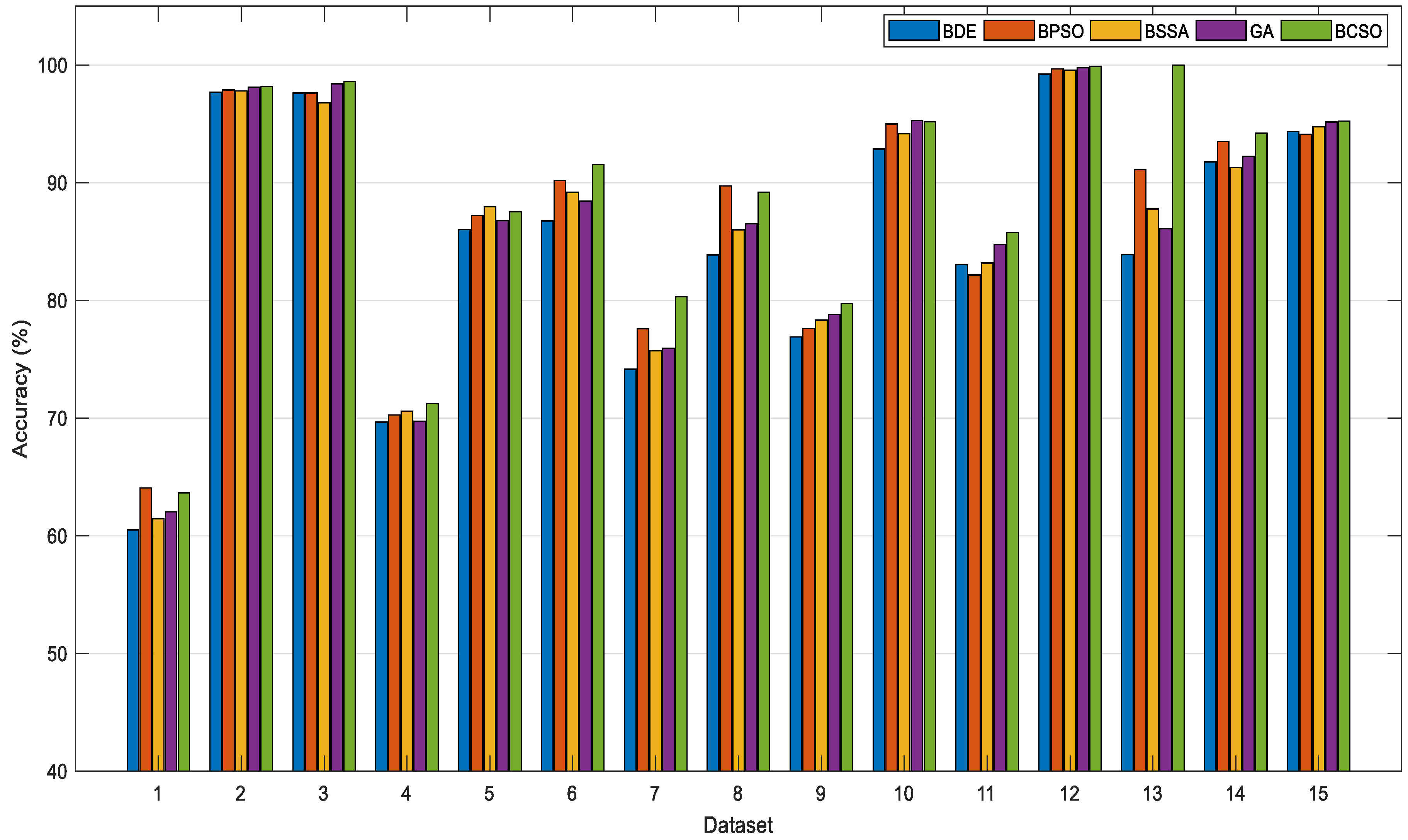
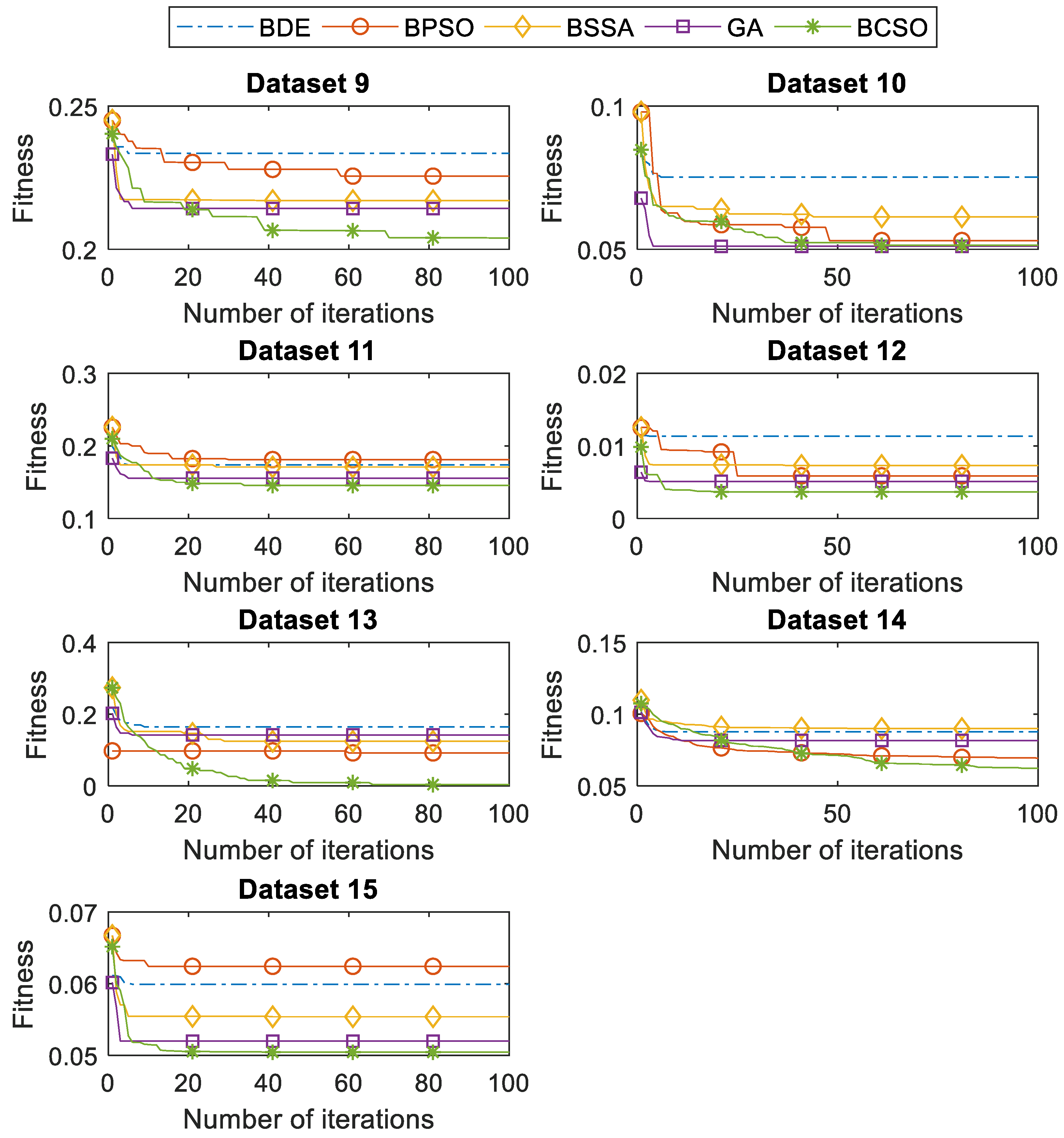
In this paper, binary variants of the CSO are proposed and applied for feature selection tasks. The continuous CSO is converted into the binary version by using transfer functions. Eight different transfer functions from S-shaped and V-shaped families are implemented in the BCSO. The S-shaped transfer functions force the search agents to move on the binary search space. On one hand, the V-shaped transfer functions allow the search agents to perform the search around the binary search space. The proposed BCSO is validated using 15 benchmark datasets. Firstly, the BCSO with the optimal transfer function is investigated. In comparison with other transfer functions, we found that the BCSO with the V2 transfer function was the most suitable, which perceived the optimal performance in current work. Secondly, the performance of the BCSO is verified with four other conventional feature selection methods. Based on the results obtained, the BCSO outperformed other methods (BDE, BSSA, BPSO, and GA) when finding the significant features, in which a high searching capability can be ensured. In addition, BCSO can often select a smaller number of significant features that contributed a high accuracy. Moreover, the processing speed of the BCSO is extremely fast, which is more appropriate in real-world applications. All in all, it can be inferred that the BCSO is a valuable feature selection tool. In the future, the BCSO can be applied to other binary optimization tasks, such as unit commitment issues, optimized neural networks, and the knapsack problem.