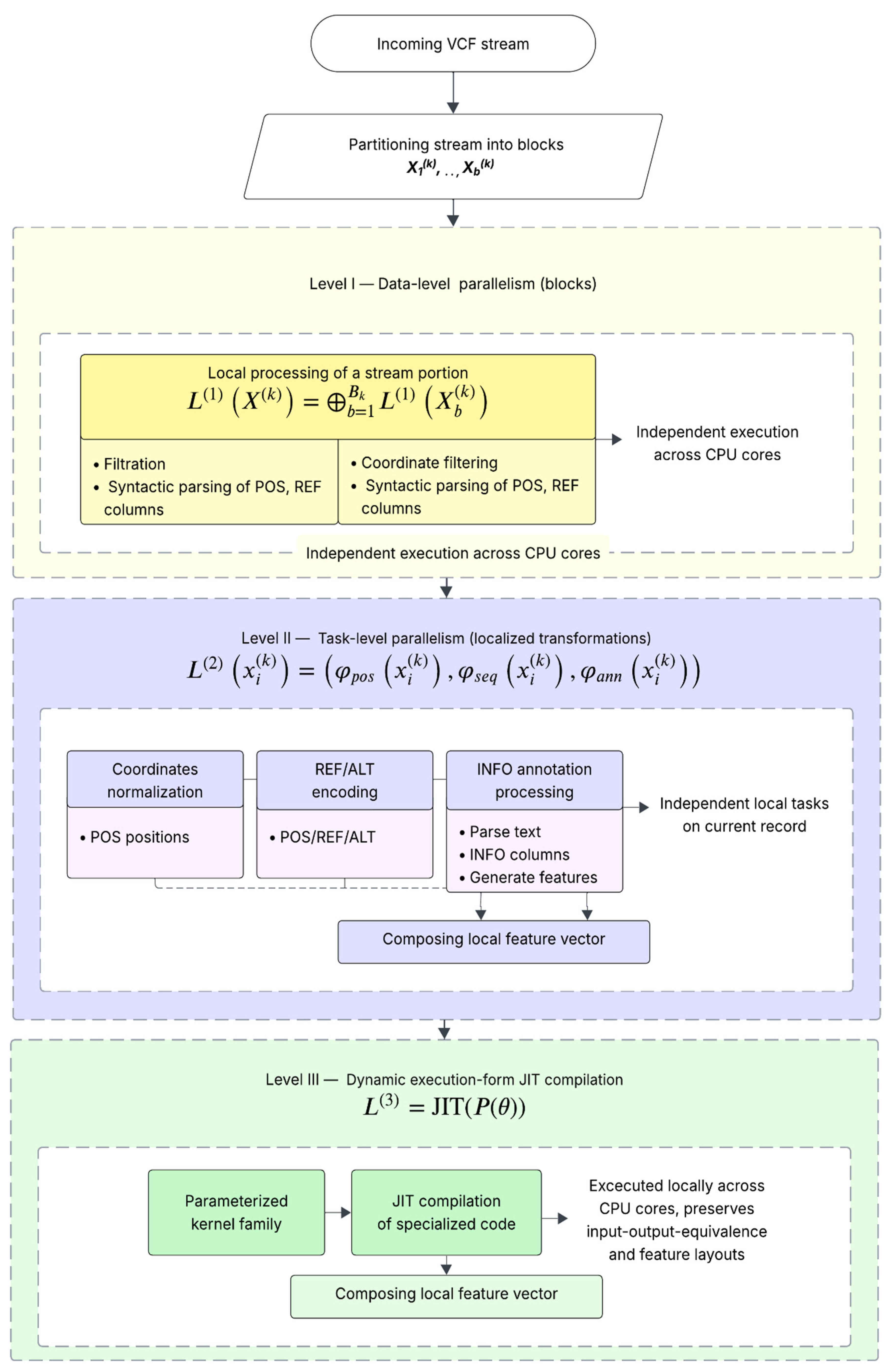
This section reports the dataset used, the evaluation protocol, and the experimental results that quantify classification quality and end-to-end execution efficiency of the proposed multi-level CPU execution method, including scalability and stage-wise ablation.
4.1. Dataset Description (BRCA1 Region of Chromosome 17)
To address the task of automated classification of mutations in the BRCA1 gene, an open human DNA variant dataset
Homo sapiens variation data (Release 110) in Variant Call Format (VCF) was used, containing information on single nucleotide variants (SNPs), multiple nucleotide variants (MNPs), insertions, and deletions (InDels) relative to the reference genome [
20]. The variants were obtained from the Ensembl resource [
21], which provides genome assemblies, gene annotations, and curated variant tracks.
For this study, the file
homo_sapiens-chr17.vcf.gz [
22] was downloaded, containing variants on chromosome 17. The downloaded VCF contained N_total = 19,772,917 variant records (excluding header lines). Variants were filtered to include only those located within the BRCA1 genomic region (GRCh38). In our experiments, the BRCA1 region was defined by the genomic interval [43,044,295; 43,125,482] in GRCh38 (chromosome 17), and all variants outside this interval were discarded. After coordinate-based filtering to the BRCA1 interval, N_BRCA1_coord = 26,548 variants were retained, which corresponds to 13% of the downloaded chromosome-17 records. Clinical labels were derived from ClinVar-derived clinical-significance attributes available in the VCF INFO field. Variants with unambiguous significance (Pathogenic/Likely_pathogenic) were assigned to class 1, and variants with unambiguous significance (Benign/Likely_benign) were assigned to class 0. Among the BRCA1-interval variants, N_excl = 21,475 records (80.89% of the BRCA1-interval subset) had undefined, ambiguous, or conflicting clinical-significance annotations and were removed. After coordinate-based filtering to the BRCA1 interval and excluding variants with undefined or conflicting clinical significance, the dataset comprised
N = 5073 variants (
N1 = 2642 pathogenic/likely pathogenic;
N0 = 2431 benign/likely benign). Overall, the final labeled dataset corresponds to 19.11% of the BRCA1-interval variants and 0.03% of the original downloaded chromosome-17 file.
The resulting dataset defines a binary classification task: clinically significant variants (pathogenic, class 1) versus clinically insignificant variants (benign, class 0). Each record includes the variant position, reference and alternative alleles, and aggregated annotations, which, after preprocessing and feature construction, are used as inputs for the classification model.
4.3. Results and Efficiency of Implementations
Experiments were executed on a system with an Intel Core i7-13700H CPU (Intel Corporation, Santa Clara, CA, USA) (14 cores, 20 threads), 16 GB RAM, and Python 3.10. The implementation was developed in Python using the standard library multiprocessing module and Numba JIT (Numba version 0.62.1) compilation. End-to-end execution time was measured with time.perf_counter() and validated across five independent runs. The measured runtime includes VCF portion reading, parsing and filtering, feature construction, model selection and training, and evaluation to ensure a consistent end-to-end comparison.
Although the dataset is stored as a finite file, the pipeline processes the VCF in a file-backed, portion-based (chunked) manner: the file is read sequentially in fixed-size portions, and the Level I–III transformations are applied to each portion before global aggregation. We report the effective throughput as (variants/s), where is the total number of processed variants and is the end-to-end runtime, and we report alongside runtime to characterize processing capacity rather than hard real-time latency guarantees.
To ensure reproducibility, we fixed random_state = 42 and used identical hyperparameter-search settings in all runs. Peak memory consumption was measured as resident set size (RSS) using psutil by sampling the main Python process and all worker processes at 0.25 s intervals during the end-to-end run. We report (i) peak total RSS (sum over processes) and (ii) peak per-process RSS (maximum over processes) to characterize memory footprint under different core counts. The dataset was split into training and test subsets using an 80/20 ratio with random_state = 42. Hyperparameters were optimized on the training subset only via RandomizedSearchCV (50 sampled configurations) with 5-fold cross-validation, and the final reported classification metrics were computed on the held-out test subset.
The reported results correspond to the execution levels defined in
Section 3: Level I (data-parallel block processing) accelerates parsing and filtering; Level II (task-parallel feature decomposition) accelerates independent feature transformations; Level III (JIT specialization) reduces numeric preprocessing overhead. To avoid nested oversubscription (
Section 3.4), preprocessing parallelism and internally threaded learning routines are executed as coordinated phases rather than unconstrained concurrent nesting.
After training and evaluating the Random Forest model, we performed a comparative analysis of sequential versus multi-level parallel execution. The analysis includes classification quality and end-to-end runtime across different CPU configurations. All experiments were conducted under identical data splits, random seeds, and hyperparameter-search settings. The results are summarized in
Table 3,
Table 4,
Table 5,
Table 6,
Table 7 and
Table 8. All memory values in
Table 5 are reported as the maximum observed peak across five independent end-to-end runs for each configuration.
As shown in
Table 3, classification quality is preserved after parallelization. Precision and F1-score slightly improve, while end-to-end runtime decreases from 291.25 s to 51.18 s. For a deeper analysis, recall was computed separately for both classes, indicating balanced sensitivity: recall for class 1 is approximately 0.83 and for class 0 approximately 0.87.
The scaling dynamics are summarized in
Table 4. With an increasing number of cores, speedup increases; however, efficiency decreases due to parallel overheads and synchronization costs [
24].
Table 5 provides the corresponding execution times. The monotonic reduction in runtime confirms the effectiveness of the parallelization strategy.
As expected, the memory footprint increases with the number of worker processes due to per-process parsing buffers and intermediate feature arrays. In our setup, the peak total RSS for 14 cores remained within the available 16 GB RAM (
Table 5), indicating feasibility for CPU-only deployment under moderate memory constraints. For memory-constrained systems, the configuration can be adapted by reducing the number of worker processes or the portion size, trading peak RSS for longer runtime. In practice, we recommend selecting the highest core count that keeps peak total RSS comfortably below available RAM to avoid paging and performance collapse.
To strengthen the portion-based processing interpretation, we additionally report the effective throughput computed from the end-to-end runtime.
Random Forest hyperparameters were automatically optimized using RandomizedSearchCV for both implementations; the selected configuration was: max_depth = None, max_features = ‘sqrt’, min_samples_split = 18, min_samples_leaf = 1, n_estimators = 236.
To address the computational overhead of model selection, we explicitly measured the runtime of the hyperparameter search phase separately from preprocessing and model training. The hyperparameter optimization stage includes all RandomizedSearchCV operations (sampling of configurations, 5-fold cross-validation, and model fitting inside each fold) and is reported as a distinct component of the end-to-end pipeline. The measured runtime for hyperparameter search (mean ± SD) was 141.87 ± 4.635 s for sequential execution (1 core) and decreased with parallelization to 100.68 ± 5.35 s (2 cores), 46.62 ± 2.92 s (4 cores), 29.66 ± 4.04 s (8 cores), and 23.64 ± 3.76 s (14 cores). This explicit separation confirms that reported end-to-end speedups remain valid even when the hyperparameter search cost is included.
Hyperparameter tuning can constitute a substantial fraction of the learning-stage cost. Therefore, the runtime of the hyperparameter-search phase (RandomizedSearchCV with cross-validation) has been measured as a standalone component and is treated separately from preprocessing and the final model fitting.
To quantify the incremental contribution of each execution component, we progressively enabled optimizations from the sequential baseline to the full workflow.
Table 7 reports the end-to-end runtime for cumulatively enabled steps, separating (i) local VCF processing and feature construction (Levels I–III) from (ii) learning-stage acceleration, decomposed into parallel hyperparameter search and parallel Random Forest fitting under resource-coordinated execution.
The ablation results demonstrate a cumulative reduction in end-to-end runtime as successive execution components are enabled. JIT compilation reduces interpreter overhead in numeric kernels; block-level data parallelism accelerates VCF parsing and filtering; and task-parallel feature processing accelerates independent feature transformations. The first four stages correspond to the local multi-level CPU execution (Levels I–III), achieving a 3.95× speedup (291.25 s → 73.82 s).
The learning stage is decomposed into two separately measured components. First, parallel hyperparameter search (RandomizedSearchCV) reduces the model-selection overhead and yields an additional reduction before the final estimator is fitted. In the parallel hyperparameter search ablation step, the acceleration is attributed to the search procedure (RandomizedSearchCV) as a standalone component, while the final model fitting is accounted for in the subsequent resource-coordinated parallel RF training step. Second, enabling resource-coordinated parallelism for the final Random Forest fitting further decreases the end-to-end runtime to 51.18 s (5.69×). This separation makes the contributions of preprocessing, hyperparameter search, and final training directly interpretable, while the resource-coordination scheme prevents nested oversubscription when externally parallel preprocessing is combined with internally threaded learning routines. Overall, the workflow achieves a 5.69× end-to-end speedup while maintaining classification quality (F1 ≈ 0.85) for BRCA1-region variant classification.
To ensure statistical robustness of the reported speedup values, each CPU configuration (1, 2, 4, 8, and 14 cores) was evaluated over five independent runs, reporting mean runtime, standard deviation, coefficient of variation, and 95% confidence intervals; paired t-tests were used to assess the significance of runtime differences relative to the sequential baseline.
The observed coefficient of variation remains below 10% across all configurations, indicating low runtime variability and stable performance behavior despite inherent multicore execution noise. The monotonic increase in speedup with the number of threads exceeds the reported confidence intervals, confirming that the observed performance gains are statistically significant and not attributable to random system fluctuations. A paired t-test between adjacent configurations confirmed statistically significant runtime reductions (p < 0.01).