Embedding-Based Alignments Capture Structural and Sequence Domains of Distantly Related Multifunctional Human Proteins
Abstract
1. Introduction
2. Materials and Methods
2.1. Workflow
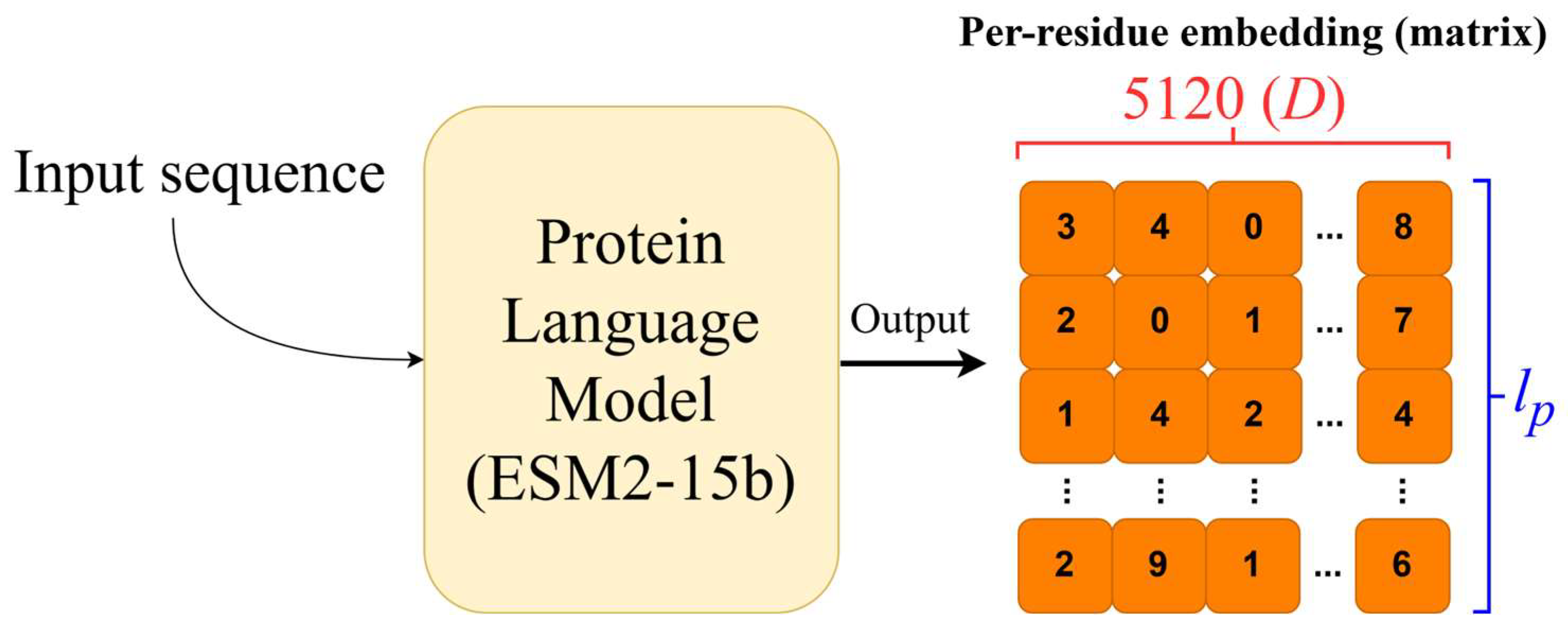
2.2. Protein Embeddings
2.3. Embedding-Based Alignment (EBA)
2.4. Clustering
2.5. Computational Time
2.6. Validation
3. Results
3.1. Validation of Unique Protein Pairs
- Pair #1 (Figure 4A). The embedded sequences have an EBA score (15.48) much higher than the reliability threshold (3.5) (see Section 2.3 for details), and the two sequences have similar length. In this case, the two structures are almost perfectly superimposed (TM-score = 0.86), supporting the notion that they are indeed remote homologs, notwithstanding the computed low sequence identity. The two proteins are multifunctional, with the same two Enzyme Commission (EC) numbers (4 and 5), indicating that they can act as lyases (4) and isomerases (5).
- Pairs #2 (Figure 4B) and #3. The embedded sequences of both pairs have an EBA score higher than the threshold (11.27 and 4.22, respectively), and the two sequences in the pairs have very different lengths. In both cases, the PDB of the longer sequence covers only a fragment of the protein, which superimposes well (TM-score > 0.6) with the other member of the pair. Despite the two proteins being very different in terms of their number of residues, the shortest protein assumes the same conformation as a domain of the longer protein. Validation indicates that the proteins in pair #2 belong to the DNA polymerase type-X family, sharing 11 InterPro signatures, and that they act as transferases (2) and lyases (4). Proteins in pair #3 belong to the protein kinase superfamily, sharing an InterPro signature, and they act as transferases (2) and hydrolases (3).
- Pairs #4 (Figure 4C), #5, and #6. The embedded sequences of all the pairs have an EBA score above the threshold (4.41, 5.96, and 5.63, respectively), and the three sequences are different in length (the difference ranges from 63 to 132 residues). In all cases, the structures in the pairs superimpose only partially (TM-score < 0.6). However, we observe a significant overlap of specific segments, suggesting that they have common structural features. Validation indicates that the proteins in pair #4 belong to the class-II aminoacyl-tRNA synthetase family, sharing two InterPro signatures (Figure 4C), and that they both act as transferases (2) and ligases (6). Proteins in pair #5 belong to the HhH-GPD superfamily, sharing three InterPro signatures, and they act as hydrolases (3) and lyases (4). Proteins in pair #6 belong to the DNA polymerase families, share two InterPro signatures, and act as transferases (2) and hydrolases (3), and one of the two also acts as a lyase (4).
| Entries | Validation | |||
|---|---|---|---|---|
| # | Protein 1 | Protein 2 | Comparison | Shared Annotation |
| 1 (Figure 4A) | Q96GA7 Len: 329 PDB: 2RKB Cov: 318 | Q9GZT4 Len: 340 PDB: 3L6B Cov: 322 | EBA: 15.48 Seq. Id.: 29% Len diff: 11 TM-score: 0.86 Aln res: 286 | InterPro: IPR000634, IPR001926, IPR036052 Family: serine/threonine dehydratase family E.C.: (4, 5) |
| 2 (Figure 4B) | P06746 Len: 335 PDB: 8VFG Cov: 327 | Q9UGP5 Len: 575 PDB: 7M09 Cov: 331 | EBA: 11.27 Seq. Id.: 21% Len diff: 240 TM-score: 0.80 Aln res: 262 | InterPro: IPR002054, IPR019843, IPR010996, IPR028207, IPR018944, IPR027421, IPR037160, IPR022312, IPR002008, IPR043519, IPR029398 Family: DNA polymerase type-X family E.C.: (2, 4) |
| 3 | Q96S44 Len: 253 PDB: 7SZC Cov: 229 | Q9BRS2 Len: 568 PDB: 4OTP Cov: 236 | EBA: 4.22 Seq. Id.: 12% Len diff: 315 TM-score: 0.67 Aln res: 153 | InterPro: IPR011009 Superfamily: protein kinase superfamily E.C.: (2, 3) |
| 4 (Figure 4C) | P41250 Len: 703 PDB: 2ZT5 Cov: 530 | Q15046 Len: 597 PDB: 6ILD Cov: 501 | EBA: 4.41 Seq. Id.: 20% Len diff: 106 TM-score: 0.36 Aln res: 176 | InterPro: IPR006195, IPR045864 Family: class-II aminoacyl-tRNA synthetase family E.C.: (2, 6) |
| 5 | O15527 Len: 345 PDB: 2XHI Cov: 316 | P78549 Len: 282 PDB: 7RDS Cov: 232 | EBA: 5.96 Seq. Id.: 21% Len diff: 63 TM-score: 0.37 Aln res: 120 | InterPro: IPR011257, IPR003265, IPR023170 Superfamily: HhH-GPD superfamily E.C.: (3, 4) |
| 6 | P54098 Len: 1239 PDB: 4ZTU Cov: 1222 | P28340 Len: 1107 PDB: 9EKB Cov: 1107 | EBA: 5.63 Seq. Id.: 18.33 Len diff: 132 TM-score: 0.25 Aln res: 87 | InterPro: IPR043502, IPR012337 Superfamily: DNA polymerase families E.C.: (2, 3); (2, 3, 4) |
| 7 | Q9NST1 Len: 481 PDB: NO | Q9UP65 Len: 541 PDB: NO | EBA: 3.95 Seq. Id.: 16% Len diff: 60 | InterPro: IPR016035 Superfamily: FabD/lysophospholipase-like superfamily E.C.: (2, 3) |
| 8 | Q14032 Len: 418 PDB: NO | Q99487 Len: 392 PDB: NO | EBA: 7.05 Seq. Id.: 20% Len diff: 26 | InterPro: IPR029058 Superfamily: α/β hydrolase superfamily E.C.: (2, 3) |
3.2. Remote Homology Validation in Protein Groups Detected with Complete-Linkage Clustering
| Groups | Validation | ||
|---|---|---|---|
| # | Group Members | Comparison | Shared Annotation |
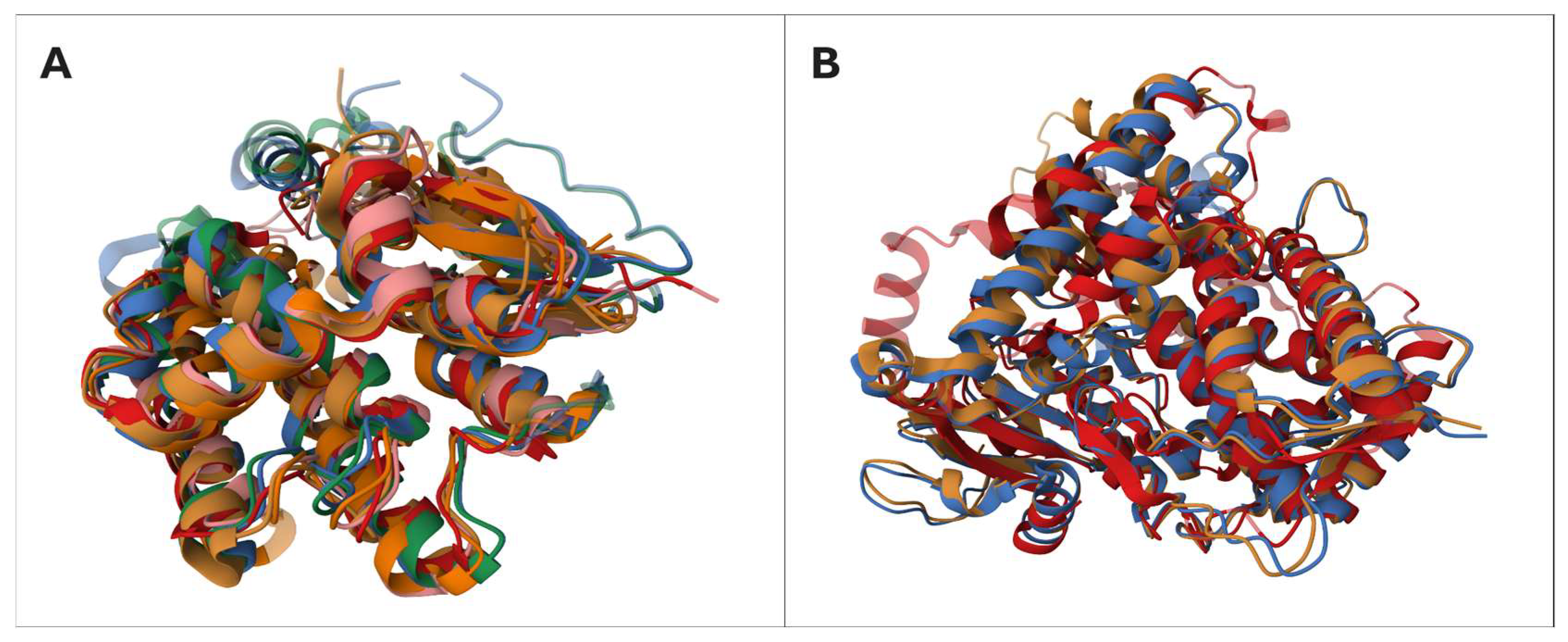
| 1 (Figure 5A) | P08263 * (222); P78417 * (241); Q9H4Y5 * (243); Q03013 * (218); O60760 * (199); O43708 * (216); | Mean EBA: 11.29 Mean TM-score: 0.75 | InterPro: IPR036249; IPR010987; IPR004045; IPR036282 Superfamily: GST superfamily E.C.: (1, 2, 5); (1, 2); (1, 2); (2, 4); (2, 5); (2, 5) |
| 2 | P11586 * (935); P13995 * (315); Q9H903 * (347) | Mean EBA: 10.21 Mean TM-score: 0.96 | InterPro: IPR020867; IPR000672; IPR020630; IPR046346; IPR020631; IPR036291 Family: tetrahydrofolate dehydrogenase/cyclohydrolase family E.C.: (1, 3, 6); (1, 3); (1, 3) |
| 3 (Figure 5B) | Q96SQ9 (504); P04798 * (512); P05177 * (516); Q16678 * (543); P51589 (502); P24557 (533); Q16647 * (500) | Mean EBA: 16.89 Mean TM-score: 0.84 | InterPro: IPR036396; IPR001128 Family: cytochrome P450 family E.C.: (1, 4, 5); (1, 4); (1, 4); (1, 4); (1, 5); (4, 5); (4, 5) |
| 4 | O14880 (152); Q99735 * (147); O14684 * (152); Q16873 * (150) | Mean EBA: 9.71 Mean TM-score: 0.79 | InterPro: IPR001129; IPR023352 Family: MAPEG family E.C.: (1, 2, 4); (1, 2, 4); (1, 2, 5); (2, 4) |
| 5 | P16118 * (471); Q16875 * (520); Q16877 (469); Q96T60 (521); O60825 * (505) | Mean EBA: 13.50 Mean TM-score: 0.92 | InterPro: IPR027417 Superfamily: P-loop containing nucleoside triphosphate hydrolases E.C.: (2, 3); (2, 3); (2, 3); (2, 3); (2, 3) |
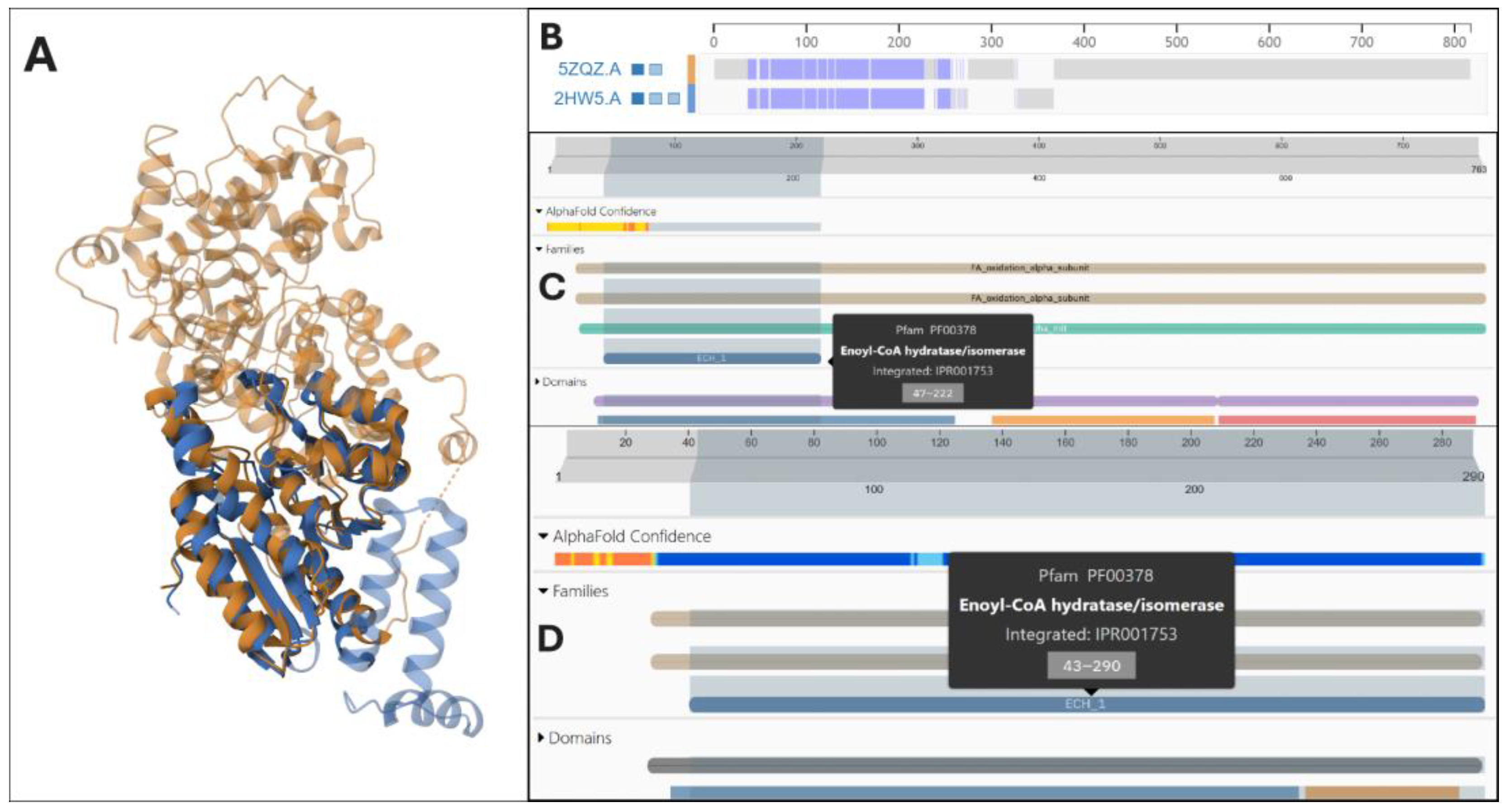
| 6 (Figure 6) | P40939 * (727); Q08426 (723); P30084 * (263) | Mean EBA: 11.85 Mean TM-score: 0.27 | InterPro: IPR029045; IPR018376; IPR001753 Family: enoyl-CoA hydratase/isomerase family E.C.: (1, 2, 4); (1, 4, 5); (4, 5) |
| 7 | P53816 * (162); Q9HDD0 (168); Q9NWW9 * (162); Q96KN8 (279); Q9UL19 * (164) | Mean EBA: 10,16 Mean TM-score: 0.88 | InterPro: IPR007053; IPR051496 Family: H-rev107 family E.C.: (2, 3); (2, 3); (2, 3); (2, 3) |
| Groups | Validation | ||
|---|---|---|---|
| # | Group Members | Comparison | Shared Annotation |
| 8 | Q14191 * (1432); Q9H8H2 (1009); Q14527 (1009) | Mean EBA: 5.92 | InterPro: IPR014001; IPR001650; IPR027417 Superfamily: helicase superfamily E.C.: (3, 5); (3, 5); (2, 3) |
| 9 | Q8TAT5 (605); Q96FI4 * (390); Q969S2 (332) | Mean EBA: 6.80 | InterPro: IPR015886; IPR012319; IPR010979 Family: FPG family E.C.: (3, 4); (3, 4); (3, 4) |
| 10 | A6NGU5 (568); P19440 * (569); P36269 (586); Q6P531 (493); Q9UJ14 (662) | Mean EBA: 17.26 | InterPro: IPR029055; IPR043137 Family: gamma-glutamyltransferase family E.C.: (2, 3); (2, 3); (2, 3); (2, 3); (2, 3) |
| 11 | P30043 * (206); P14060 (373); P26439 (372) | Mean EBA: 9.48 | InterPro: IPR036291 Superfamily: NAD(P)-binding domain superfamily E.C.: (1, 2); (1, 5); (1, 5) |
4. Conclusions and Perspectives
Supplementary Materials
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Lesk, A.M. Introduction to Protein Science: Architecture, Function, and Genomics, 3rd ed.; Oxford University Press: Oxford, UK, 2016. [Google Scholar]
- Doolittle, R.F. Of Urfs and Orfs: A Primer on How to Analyze Derived Amino Acid Sequences; University Science Books: Mill Valley, CA, USA, 1987. [Google Scholar]
- Zhang, Y.; Skolnick, J. TM-Align: A Protein Structure Alignment Algorithm Based on the TM-Score. Nucleic Acids Res. 2005, 33, 2302–2309. [Google Scholar] [CrossRef]
- Finn, R.D.; Mistry, J.; Tate, J.; Coggill, P.; Heger, A.; Pollington, J.E.; Gavin, O.L.; Gunasekaran, P.; Ceric, G.; Forslund, K.; et al. The Pfam Protein Families Database. Nucleic Acids Res. 2010, 38, D211–D222. [Google Scholar] [CrossRef]
- Paysan-Lafosse, T.; Andreeva, A.; Blum, M.; Chuguransky, S.R.; Grego, T.; Pinto, B.L.; Salazar, G.A.; Bileschi, M.L.; Llinares-López, F.; Meng-Papaxanthos, L.; et al. The Pfam Protein Families Database: Embracing AI/ML. Nucleic Acids Res. 2025, 53, D523–D534. [Google Scholar] [CrossRef]
- Blum, M.; Andreeva, A.; Florentino, L.C.; Chuguransky, S.R.; Grego, T.; Hobbs, E.; Pinto, B.L.; Orr, A.; Paysan-Lafosse, T.; Ponamareva, I.; et al. InterPro: The Protein Sequence Classification Resource in 2025. Nucleic Acids Res. 2025, 53, D444–D456. [Google Scholar] [CrossRef]
- The UniProt Consortium; Bateman, A.; Martin, M.-J.; Orchard, S.; Magrane, M.; Ahmad, S.; Alpi, E.; Bowler-Barnett, E.H.; Britto, R.; Bye-A-Jee, H.; et al. UniProt: The Universal Protein Knowledgebase in 2023. Nucleic Acids Res. 2023, 51, D523–D531. [Google Scholar] [CrossRef]
- Berman, H.M. The Protein Data Bank. Nucleic Acids Res. 2000, 28, 235–242. [Google Scholar] [CrossRef] [PubMed]
- Holm, L.; Laiho, A.; Törönen, P.; Salgado, M. DALI Shines a Light on Remote Homologs: One Hundred Discoveries. Protein Sci. 2023, 32, e4519. [Google Scholar] [CrossRef] [PubMed]
- Zhu, J.; Weng, Z. FAST: A Novel Protein Structure Alignment Algorithm. Proteins 2005, 58, 618–627. [Google Scholar] [CrossRef]
- Ortiz, A.R.; Strauss, C.E.M.; Olmea, O. MAMMOTH (Matching Molecular Models Obtained from Theory): An Automated Method for Model Comparison. Protein Sci. 2002, 11, 2606–2621. [Google Scholar] [CrossRef]
- Van Kempen, M.; Kim, S.S.; Tumescheit, C.; Mirdita, M.; Lee, J.; Gilchrist, C.L.M.; Söding, J.; Steinegger, M. Fast and Accurate Protein Structure Search with Foldseek. Nat. Biotechnol. 2024, 42, 243–246. [Google Scholar] [CrossRef]
- Bileschi, M.L.; Belanger, D.; Bryant, D.H.; Sanderson, T.; Carter, B.; Sculley, D.; Bateman, A.; DePristo, M.A.; Colwell, L.J. Using Deep Learning to Annotate the Protein Universe. Nat. Biotechnol. 2022, 40, 932–937. [Google Scholar] [CrossRef]
- Hamamsy, T.; Morton, J.T.; Blackwell, R.; Berenberg, D.; Carriero, N.; Gligorijevic, V.; Strauss, C.E.M.; Leman, J.K.; Cho, K.; Bonneau, R. Protein Remote Homology Detection and Structural Alignment Using Deep Learning. Nat. Biotechnol. 2024, 42, 975–985. [Google Scholar] [CrossRef]
- Radivojac, P. Advancing Remote Homology Detection: A Step toward Understanding and Accurately Predicting Protein Function. Cell Syst. 2022, 13, 435–437. [Google Scholar] [CrossRef]
- Jumper, J.; Evans, R.; Pritzel, A.; Green, T.; Figurnov, M.; Ronneberger, O.; Tunyasuvunakool, K.; Bates, R.; Žídek, A.; Potapenko, A.; et al. Highly Accurate Protein Structure Prediction with AlphaFold. Nature 2021, 596, 583–589. [Google Scholar] [CrossRef]
- Lin, Z.; Akin, H.; Rao, R.; Hie, B.; Zhu, Z.; Lu, W.; Smetanin, N.; Verkuil, R.; Kabeli, O.; Shmueli, Y.; et al. Evolutionary-Scale Prediction of Atomic-Level Protein Structure with a Language Model. Science 2023, 379, 1123–1130. [Google Scholar] [CrossRef] [PubMed]
- Rives, A.; Meier, J.; Sercu, T.; Goyal, S.; Lin, Z.; Liu, J.; Guo, D.; Ott, M.; Zitnick, C.L.; Ma, J.; et al. Biological Structure and Function Emerge from Scaling Unsupervised Learning to 250 Million Protein Sequences. Proc. Natl. Acad. Sci. USA 2021, 118, e2016239118. [Google Scholar] [CrossRef]
- Vaswani, A.; Shazeer, N.; Parmar, N.; Uszkoreit, J.; Jones, L.; Gomez, A.N.; Kaiser, L.; Polosukhin, I. Attention Is All You Need. arXiv 2017, arXiv:1706.03762. [Google Scholar] [CrossRef]
- Iovino, B.G.; Ye, Y. Protein embedding based alignment. BMC Bioinform. 2024, 25, 85. [Google Scholar] [CrossRef] [PubMed]
- Pantolini, L.; Studer, G.; Pereira, J.; Durairaj, J.; Tauriello, G.; Schwede, T. Embedding-Based Alignment: Combining Protein Language Models with Dynamic Programming Alignment to Detect Structural Similarities in the Twilight-Zone. Bioinformatics 2024, 40, btad786. [Google Scholar] [CrossRef]
- Spicer, R.; Raychawdhary, N.; Danwada, S.; Udomprasert, P.; Seals, C.; Bhattacharya, S. Evaluating the significance of embedding-based protein sequence alignment with clustering and double dynamic programming for remote homology. Sci. Rep. 2025, 15, 39601. [Google Scholar] [CrossRef]
- Kilinc, M.; Jia, K.; Jernigan, R.L. Major advances in protein function assignment by remote homolog detection with protein language models—A review. Curr. Opin. Struct. Biol. 2025, 90, 102984. [Google Scholar] [CrossRef] [PubMed]
- Kilinc, M.; Jia, K.; Jernigan, R.L. Improved global protein homolog detection with major gains in function identification. Proc. Natl. Acad. Sci. USA 2023, 120, e2211823120. [Google Scholar] [CrossRef] [PubMed]
- Vazzana, G.; Savojardo, C.; Martelli, P.L.; Casadio, R. Testing the Capability of Embedding-Based Alignments on the GST Superfamily Classification: The Role of Protein Length. Molecules 2024, 29, 4616. [Google Scholar] [CrossRef]
- Bertolini, E.; Babbi, G.; Savojardo, C.; Martelli, P.L.; Casadio, R. MultifacetedProtDB: A Database of Human Proteins with Multiple Functions. Nucleic Acids Res. 2024, 52, D494–D501. [Google Scholar] [CrossRef] [PubMed]
- Schmirler, R.; Heinzinger, M.; Rost, B. Fine-tuning protein language models boosts predictions across diverse tasks. Nat. Commun. 2024, 15, 7407. [Google Scholar] [CrossRef]
- Saadat, A.; Fellay, J. Fine-tuning protein language models to understand the functional impact of missense variants. Comput. Struct. Biotechnol. J. 2025, 27, 2199–2207. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Everitt, B.S.; Landau, S.; Leese, M.; Stahl, D. Cluster Analysis, 5th ed.; Wiley Series in Probability and Statistics; Wiley: Chichester, UK, 2011. [Google Scholar]
- Zhang, Y.; Skolnick, J. Scoring Function for Automated Assessment of Protein Structure Template Quality. Proteins 2004, 57, 702–710. [Google Scholar] [CrossRef]
- Sievers, F.; Higgins, D.G. Clustal Omega for Making Accurate Alignments of Many Protein Sequences. Protein Sci. 2018, 27, 135–145. [Google Scholar] [CrossRef]



Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Vazzana, G.; Manfredi, M.; Savojardo, C.; Martelli, P.L.; Casadio, R. Embedding-Based Alignments Capture Structural and Sequence Domains of Distantly Related Multifunctional Human Proteins. Computation 2026, 14, 25. https://doi.org/10.3390/computation14010025
Vazzana G, Manfredi M, Savojardo C, Martelli PL, Casadio R. Embedding-Based Alignments Capture Structural and Sequence Domains of Distantly Related Multifunctional Human Proteins. Computation. 2026; 14(1):25. https://doi.org/10.3390/computation14010025
Chicago/Turabian StyleVazzana, Gabriele, Matteo Manfredi, Castrense Savojardo, Pier Luigi Martelli, and Rita Casadio. 2026. "Embedding-Based Alignments Capture Structural and Sequence Domains of Distantly Related Multifunctional Human Proteins" Computation 14, no. 1: 25. https://doi.org/10.3390/computation14010025
APA StyleVazzana, G., Manfredi, M., Savojardo, C., Martelli, P. L., & Casadio, R. (2026). Embedding-Based Alignments Capture Structural and Sequence Domains of Distantly Related Multifunctional Human Proteins. Computation, 14(1), 25. https://doi.org/10.3390/computation14010025





