DEGAIN: Generative-Adversarial-Network-Based Missing Data Imputation
Abstract
:1. Introduction
- Removing duplicate or irrelevant data: When datasets from multiple sources, clients, etc., are combined, the chance of duplicate data creation increases. Additionally, in some analyses, the irrelevant data could also be removed. Any information that does not pertain to the issue that we are attempting to solve is considered irrelevant.
- Fixing structural errors: Structural errors occur when conventions, typos, or incorrect capitalization is observed due to the measurement or data transfer. For instance, if “N/A” and “Not Applicable” both appear, they should be analyzed as the same category.
- Filtering unwanted outliers: If an outlier proves to be irrelevant for analysis or is a mistake, it needs to be removed. Some outliers represent natural variations in the population, and they should be left as is.
- Handling missing data: Many data analytic algorithms cannot accept missing values. There are a few methods to deal with missing data. In general, missing data rows are removed or the missing values are estimated according to the existing data in the dataset. These methods are also known as data imputation.
- Data validation and quality assurance: After completing the previous steps, it is needed to validate the data and to make sure that the data have sufficient quality for the considered analytics.
- Missing completely at random (MCAR): It means that the probability of missing data does not depend on any value of attributes.
- Missing at random (MAR): Meaning that the probability of missing data does not depend on its own particular value, but on the values of other attributes.
- Not missing at random (NMAR): Meaning that the missing data depend on the missed values.
2. Related Works
2.1. Incomplete Information
- We are sure that the value exists but it is unknown for us;
- We are sure that this value does not exist;
- We do not know anything.
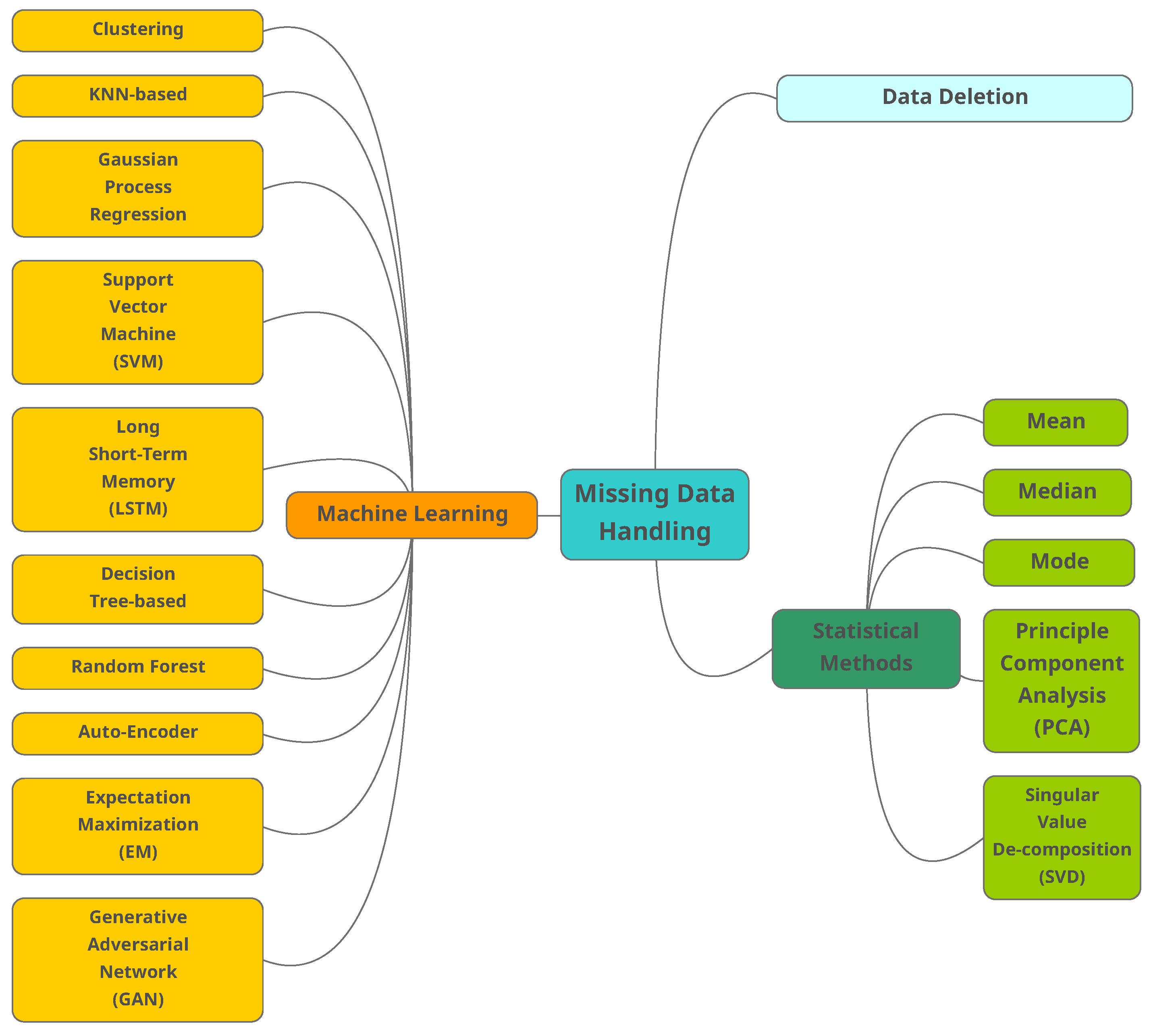
2.2. Traditional Methods
- Case deletion (CD): In CD, missing data instances are omitted. The method has two main disadvantages [25]:
- Decreasing the dataset size;
- Since the data are not always MCAR, bias occurs on data distribution and corresponding statistical analysis.
- Mean, median and mode: In these methods, the missing data are replaced with the mean (numeric attribute) of all observed cases. Median is also used to reduce the influence of exceptional data. The characteristic of the original dataset will be changed by using constants to replace missing data, ignoring the relationship among attributes. As an alternative similar solution, we may use the mode of all known values of that attributes to replace the missing data [25]. Mode is usually preferred for categorical data.
- Principal component analysis (PCA): This method is well-known in statistical data analysis and can be used to estimate the data structure level. Traditional PCA cannot deal with the missing data. Upon the measure of variance within the dataset, data will be scored by how well they fit into a principal component (PC). Since the data points will have a PC (the one that best fits), PCA can be considered a clustering analysis. Missing scores are estimated by projecting known scores back into the principal space. More details can be found in [26].
- Singular value decomposition (SVD): In this method, data are projected into another space where the attributes have different values. In the projected space, it is possible to re-construct the missing data.
2.3. Machine Learning Methods
- Clustering: These unsupervised learning algorithms group the samples with similar characters. To replace the missing value, the distance between the centroid of clusters with the sample is calculated, and the missing value of chosen cluster is replaced with the obtained value [28]. The minimum distance could be calculated by a variety of distance functions. One common function is -norm [37]. The -norm calculates the distance of the vector coordinate from the origin of the vector space.
- k-nearest neighbors (KNN): This supervised algorithm replaces the missing value by the mean (or weighted mean) of the k nearest samples. These neighbor samples are identified by calculating the distance of the missing value with the available samples [28]. Currently, many variations of the original KNN have been proposed in the literature, including the SKNN, IKNN, CKNN, and ICKNN. KNN-based algorithms require heavy calculations to find the nearest neighbors [29]. Some of the common distance functions used in KNN are the Euclidean-overlap metric [38], Value Difference Metric [38] and mean Euclidean distance [38].
- Gaussian process regression (GPR): GPR algorithms predict the output’s variance based on non-linear probabilistic techniques. GPR-based algorithms estimate a probabilistic region for the missing values instead of point estimation. The performance of GRP-based algorithms is dependent on the used kernel function. This kernel is chosen according to the data type and the effective algorithms might use a combination of different kernels. Similar to KNN, the GRP-based algorithms also need heavy calculations, which is not the ideal case for large-scale datasets [30].
- Support vector machine (SVM): SVM has applications in both classification and regression. SVM-based algorithms are non-linear and map the input to a high-dimensional feature space. SVM-based algorithms also use different kernels such as GPR [30].
- Long short-term memory (LSTM): LSTM is a deep learning (DL) algorithm. As a subcategory of recurrent neural networks, DL shows improvement for time series compared with conventional ML algorithms. The training phase of the LSTM might be complex due to the vanishing gradient problem [32].
- Decision tree (DT): DT-based algorithms partition the dataset into groups of samples, forming a tree. The missing data are estimated by the samples associated with the same leaves of the tree. Different variations of DT algorithms have been proposed by researchers, including the ID3, C4.5, CRAT, CHAILD and QUEST [39]. DT algorithms do not require prior information on the data distribution [33].
- Random forest (RF): RF consists of multiple DTs in which the average value of the DT estimation is considered the missing value [34].
- Auto-encoder (AE): As a class of unsupervised DL algorithms, AE learns a coded vector from the input space. The AE generally consists of three layers, including the input, hidden, and output. The objective in AE is to map the input layer to the hidden layer and then reconstruct the input samples through the hidden vector. The elements of input attributes can be missed randomly in the training phase. Therefore, it is expected that the vectors in the input space consist of randomly missing vectors, and the output layer has the complete set of vectors. Performing this task, the AE will learn how to complete the missing data. Different versions of AEs have been introduced, such as VAE, DAE, SAE, and SDAE [35].
- Expectation maximization (EM): EM algorithms are capable of obtaining the local maximum likelihood of a statistical model. These models entail latent variables besides the observed data and unknown parameters. These latent variables can be missing values among the data. The EM-based algorithms guarantee that the likelihood will increase. However, the price is slow convergence rate [36].
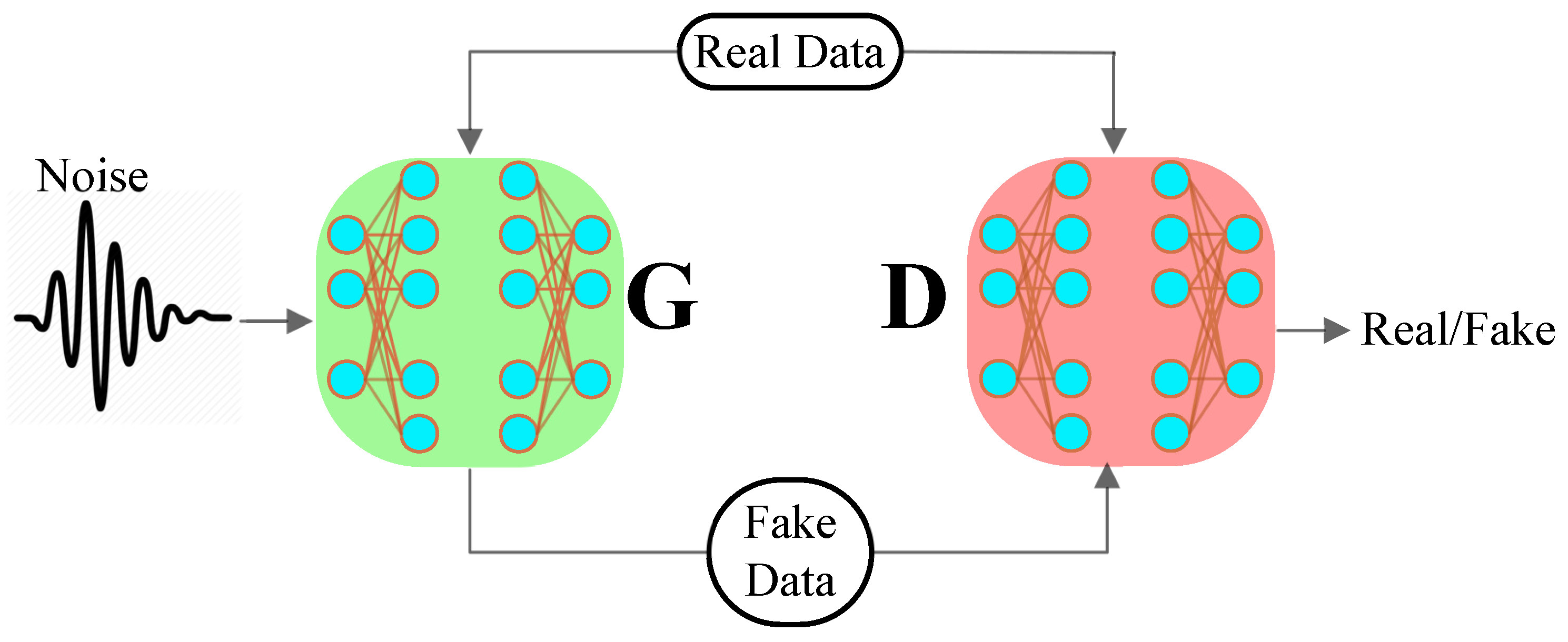
Generative Adversarial Networks
3. Proposed Method
3.1. System Model
- Linear layer: The noise vector is fed into a fully connected layer, and its output is reshaped into a tensor.
- Batch normalization layer: Stabilizes learning by normalizing inputs to zero mean and unit variance, avoiding training issues, such as vanishing or exploding gradients, and allowing the gradient to flow through the network.
- Up sample layer: Instead of using a convolutional transpose layer to up sample, it mentions using upsampling and then applying a simple convolutional layer on top of it. Convolutional transpose is sometimes used instead.
- Convolutional layer: To learn from up-sampled data, the matrix is passed through a convolutional layer with a stride of 1 and the same padding as it is up sampled.
- ReLU layer: For the generator because it allows the model to quickly saturate and cover the training distribution space.
- TanH Activation: TanH enables the model to converge more quickly.
3.2. Problem Formulation
3.3. Proposed Algorithm: DEGAIN
| Algorithm 1 (The DEGAIN algorithm: deconvolution and then training). |
|
4. Performance Evaluation
4.1. Evaluation Metrics
4.2. Dataset
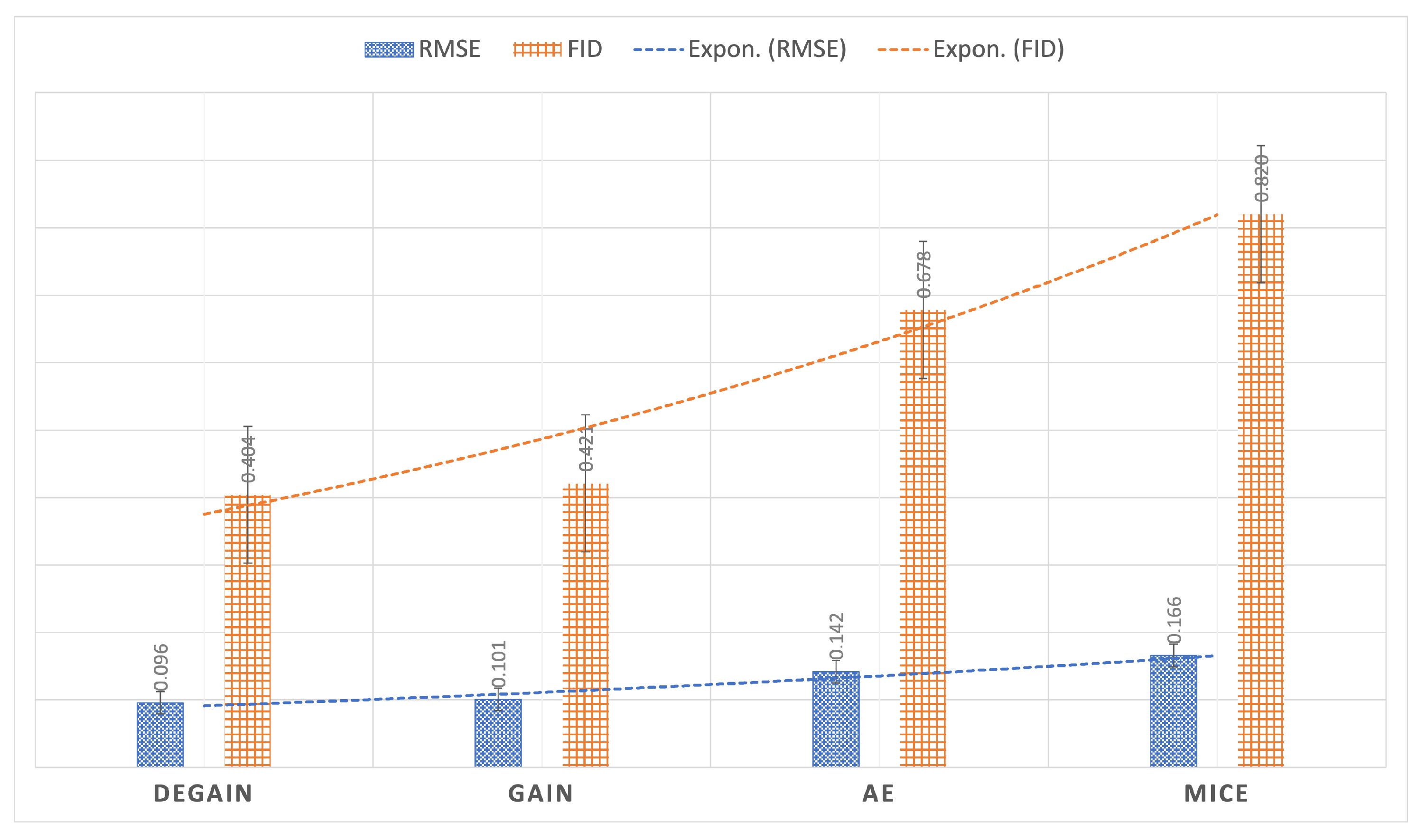
4.3. Results
5. Conclusions
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
Abbreviations
| AE | Auto-Encoder |
| CD | Case Deletion |
| DT | Decision Tree |
| DL | Deep Learning |
| EM | Expectation Maximization |
| FID | Frechet Inception Distance |
| GRP | Gaussian Process Regression |
| GAN | Generative Adversarial Network |
| KNN | k-Nearest Neighbors |
| LSTM | Long Short-Term Memory |
| ML | Machine Learning |
| MAE | Mean Absolute Error |
| MAR | Missing At Random |
| MCAR | Missing Completely At Random |
| MLP | Multi-Layer Perceptron |
| NMAR | Not Missing At Random |
| PCA | Principal Component Analysis |
| RF | Random Forest |
| RMSE | Root Mean Square Error |
| SVD | Singular Value Decomposition |
| SVM | Support Vector Machine |
References
- Ilyas, I.F.; Chu, X. Data Cleaning; Morgan & Claypool: San Rafael, CA, USA, 2019. [Google Scholar]
- O’Brien, A.D.; Stone, D.N. Yes, you can import, analyze, and create dashboards and storyboards in Tableau! The GBI case. J. Emerg. Technol. Account. 2020, 17, 21–31. [Google Scholar] [CrossRef]
- Luo, Y. Evaluating the state of the art in missing data imputation for clinical data. Briefings Bioinform. 2022, 23, bbab489. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Bao, T.; Chen, H.; Zhang, K.; Shu, X.; Chen, Z.; Hu, Y. A large-scale sensor missing data imputation framework for dams using deep learning and transfer learning strategy. Measurement 2021, 178, 109377. [Google Scholar] [CrossRef]
- Platias, C.; Petasis, G. A Comparison of Machine Learning Methods for Data Imputation. In Proceedings of the 11th Hellenic Conference on Artificial Intelligence, Athens, Greece, 2–4 September 2020; pp. 150–159. [Google Scholar]
- Austin, P.C.; White, I.R.; Lee, D.S.; van Buuren, S. Missing data in clinical research: A tutorial on multiple imputation. Can. J. Cardiol. 2021, 37, 1322–1331. [Google Scholar] [CrossRef] [PubMed]
- Yoon, J.; Jordon, J.; Schaar, M. Gain: Missing data imputation using generative adversarial nets. In Proceedings of the International Conference on Machine Learning, Stockholm, Sweden, 10–15 July 2018; pp. 5689–5698. [Google Scholar]
- Ye, C.; Evanusa, M.; He, H.; Mitrokhin, A.; Goldstein, T.; Yorke, J.A.; Fermüller, C.; Aloimonos, Y. Network deconvolution. arXiv 2019, arXiv:1905.11926. [Google Scholar]
- Gondara, L.; Wang, K. Multiple imputation using deep denoising autoencoders. arXiv 2017, arXiv:1705.02737. [Google Scholar]
- Van Buuren, S.; Groothuis-Oudshoorn, K. mice: Multivariate imputation by chained equations in R. J. Stat. Softw. 2011, 45, 1–67. [Google Scholar] [CrossRef] [Green Version]
- Greco, S.; Molinaro, C.; Trubitsyna, I. Approximation algorithms for querying incomplete databases. Inf. Syst. 2019, 86, 28–45. [Google Scholar] [CrossRef]
- Calautti, M.; Console, M.; Pieris, A. Benchmarking approximate consistent query answering. In Proceedings of the 40th ACM SIGMOD-SIGACT-SIGAI Symposium on Principles of Database Systems, Virtual Event, China, 20–25 June 2021; pp. 233–246. [Google Scholar]
- Calautti, M.; Caroprese, L.; Greco, S.; Molinaro, C.; Trubitsyna, I.; Zumpano, E. Existential active integrity constraints. Expert Syst. Appl. 2021, 168, 114297. [Google Scholar] [CrossRef]
- Calautti, M.; Greco, S.; Molinaro, C.; Trubitsyna, I. Query answering over inconsistent knowledge bases: A probabilistic approach. Theor. Comput. Sci. 2022, 935, 144–173. [Google Scholar] [CrossRef]
- Calautti, M.; Greco, S.; Molinaro, C.; Trubitsyna, I. Preference-based Inconsistency-Tolerant Query Answering under Existential Rules. Artif. Intell. 2022, 312, 103772. [Google Scholar] [CrossRef]
- Calautti, M.; Greco, S.; Molinaro, C.; Trubitsyna, I. Querying Data Exchange Settings Beyond Positive Queries. In Proceedings of the 4th International Workshop on the Resurgence of Datalog in Academia and Industry (Datalog-2.0), Genova, Italy, 5 September 2022; Volume 3203, pp. 27–41. [Google Scholar]
- Toussaint, E.; Guagliardo, P.; Libkin, L.; Sequeda, J. Troubles with nulls, views from the users. Proc. VIDB Endow. 2022, 15, 2613–2625. [Google Scholar] [CrossRef]
- Guagliardo, P.; Libkin, L. Making SQL queries correct on incomplete databases: A feasibility study. In Proceedings of the 35th ACM SIGMOD-SIGACT-SIGAI Symposium on Principles of Database Systems, San Francisco, CA, USA, 26 June–1 July 2016; pp. 211–223. [Google Scholar]
- Abiteboul, S.; Kanellakis, P.C.; Grahne, G. On the Representation and Querying of Sets of Possible Worlds. Theor. Comput. Sci. 1991, 78, 158–187. [Google Scholar] [CrossRef] [Green Version]
- Libkin, L. SQL’s three-valued logic and certain answers. ACM Trans. Database Syst. (TODS) 2016, 41, 1–28. [Google Scholar] [CrossRef] [Green Version]
- Fiorentino, N.; Greco, S.; Molinaro, C.; Trubitsyna, I. ACID: A system for computing approximate certain query answers over incomplete databases. In Proceedings of the International Conference on Management of Data (SIGMOD), Houston, TX, USA, 10–15 June 2018; pp. 1685–1688. [Google Scholar]
- Fiorentino, N.; Molinaro, C.; Trubitsyna, I. Approximate Query Answering over Incomplete Data. In Complex Pattern Mining; Springer: Berlin, Germany, 2020; pp. 213–227. [Google Scholar]
- Hu, J.; Zhou, Z.; Yang, X. Characterizing Physical-Layer Transmission Errors in Cable Broadband Networks. In Proceedings of the 19th USENIX Symposium on Networked Systems Design and Implementation (NSDI 22), Renton, WA, USA, 4–6 April 2022; USENIX Association: Renton, WA, USA, 2022; pp. 845–859. [Google Scholar]
- Yu, K.; Yang, Y.; Ding, W. Causal Feature Selection with Missing Data. ACM Trans. Knowl. Discov. Data 2022, 16, 1–24. [Google Scholar] [CrossRef]
- Peng, L.; Lei, L. A review of missing data treatment methods. Intell. Inf. Manag. Syst. Technol 2005, 1, 412–419. [Google Scholar]
- Folch-Fortuny, A.; Arteaga, F.; Ferrer, A. PCA model building with missing data: New proposals and a comparative study. Chemom. Intell. Lab. Syst. 2015, 146, 77–88. [Google Scholar] [CrossRef]
- Mirtaheri, S.L.; Shahbazian, R. Machine Learning: Theory to Applications; CRC Press: Boca Raton, FL, USA, 2022. [Google Scholar]
- Nagarajan, G.; Babu, L.D. Missing data imputation on biomedical data using deeply learned clustering and L2 regularized regression based on symmetric uncertainty. Artif. Intell. Med. 2022, 123, 102214. [Google Scholar] [CrossRef]
- Emmanuel, T.; Maupong, T.; Mpoeleng, D.; Semong, T.; Mphago, B.; Tabona, O. A survey on missing data in machine learning. J. Big Data 2021, 8, 1–37. [Google Scholar]
- Ma, Y.; He, Y.; Wang, L.; Zhang, J. Probabilistic reconstruction for spatiotemporal sensor data integrated with Gaussian process regression. Probabilistic Eng. Mech. 2022, 69, 103264. [Google Scholar] [CrossRef]
- Camastra, F.; Capone, V.; Ciaramella, A.; Riccio, A.; Staiano, A. Prediction of environmental missing data time series by Support Vector Machine Regression and Correlation Dimension estimation. Environ. Model. Softw. 2022, 150, 105343. [Google Scholar] [CrossRef]
- Saroj, A.J.; Guin, A.; Hunter, M. Deep LSTM recurrent neural networks for arterial traffic volume data imputation. J. Big Data Anal. Transp. 2021, 3, 95–108. [Google Scholar] [CrossRef]
- Cenitta, D.; Arjunan, R.V.; Prema, K. Missing data imputation using machine learning algorithm for supervised learning. In Proceedings of the 2021 International Conference on Computer Communication and Informatics (ICCCI), Coimbatore, India, 27–29 January 2021; pp. 1–5. [Google Scholar]
- Tang, F.; Ishwaran, H. Random forest missing data algorithms. Stat. Anal. Data Mining: Asa Data Sci. J. 2017, 10, 363–377. [Google Scholar] [CrossRef] [PubMed]
- Ryu, S.; Kim, M.; Kim, H. Denoising autoencoder-based missing value imputation for smart meters. IEEE Access 2020, 8, 40656–40666. [Google Scholar] [CrossRef]
- Nelwamondo, F.V.; Mohamed, S.; Marwala, T. Missing data: A comparison of neural network and expectation maximization techniques. Curr. Sci. 2007, 93, 1514–1521. [Google Scholar]
- Eirola, E.; Doquire, G.; Verleysen, M.; Lendasse, A. Distance estimation in numerical data sets with missing values. Inf. Sci. 2013, 240, 115–128. [Google Scholar] [CrossRef] [Green Version]
- Santos, M.S.; Abreu, P.H.; Wilk, S.; Santos, J. How distance metrics influence missing data imputation with k-nearest neighbours. Pattern Recognit. Lett. 2020, 136, 111–119. [Google Scholar] [CrossRef]
- Rokach, L.; Maimon, O. Decision trees. In Data Mining and Knowledge Discovery Handbook; Springer: New York, NY, USA, 2005; pp. 165–192. [Google Scholar]
- Benjdira, B.; Ammar, A.; Koubaa, A.; Ouni, K. Data-efficient domain adaptation for semantic segmentation of aerial imagery using generative adversarial networks. Appl. Sci. 2020, 10, 1092. [Google Scholar] [CrossRef] [Green Version]
- Revesz, P.Z. On the semantics of arbitration. Int. J. Algebra Comput. 1997, 7, 133–160. [Google Scholar] [CrossRef]

| Metric | Formula |
|---|---|
| RMSE | |
| FID |
| Proposed DEGAIN | GAIN [7] | AE [9] | MICE [10] | |
|---|---|---|---|---|
| RMSE (Letter dataset) | 0.096 | 0.101 | 0.142 | 0.166 |
| Normalized FID (Letter dataset) | 0.492 | 0.513 | 0.826 | 1 |
| RMSE (SPAM dataset) | 0.047 | 0.050 | 0.064 | 0.068 |
| Normalized FID (SPAM dataset) | 0.898 | 0.946 | 0.973 | 1 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Shahbazian, R.; Trubitsyna, I. DEGAIN: Generative-Adversarial-Network-Based Missing Data Imputation. Information 2022, 13, 575. https://doi.org/10.3390/info13120575
Shahbazian R, Trubitsyna I. DEGAIN: Generative-Adversarial-Network-Based Missing Data Imputation. Information. 2022; 13(12):575. https://doi.org/10.3390/info13120575
Chicago/Turabian StyleShahbazian, Reza, and Irina Trubitsyna. 2022. "DEGAIN: Generative-Adversarial-Network-Based Missing Data Imputation" Information 13, no. 12: 575. https://doi.org/10.3390/info13120575
APA StyleShahbazian, R., & Trubitsyna, I. (2022). DEGAIN: Generative-Adversarial-Network-Based Missing Data Imputation. Information, 13(12), 575. https://doi.org/10.3390/info13120575







