A Review of Environmental Challenges Facing Martian Colonisation and the Potential for Terrestrial Microbes to Transform a Toxic Extraterrestrial Environment
Abstract
1. Introduction
2. A Brief History of Anthropogenic Uses of Bacteria
2.1. Bacterial Uses in Ancient Civilisation
2.2. Modern Uses of Bacteria
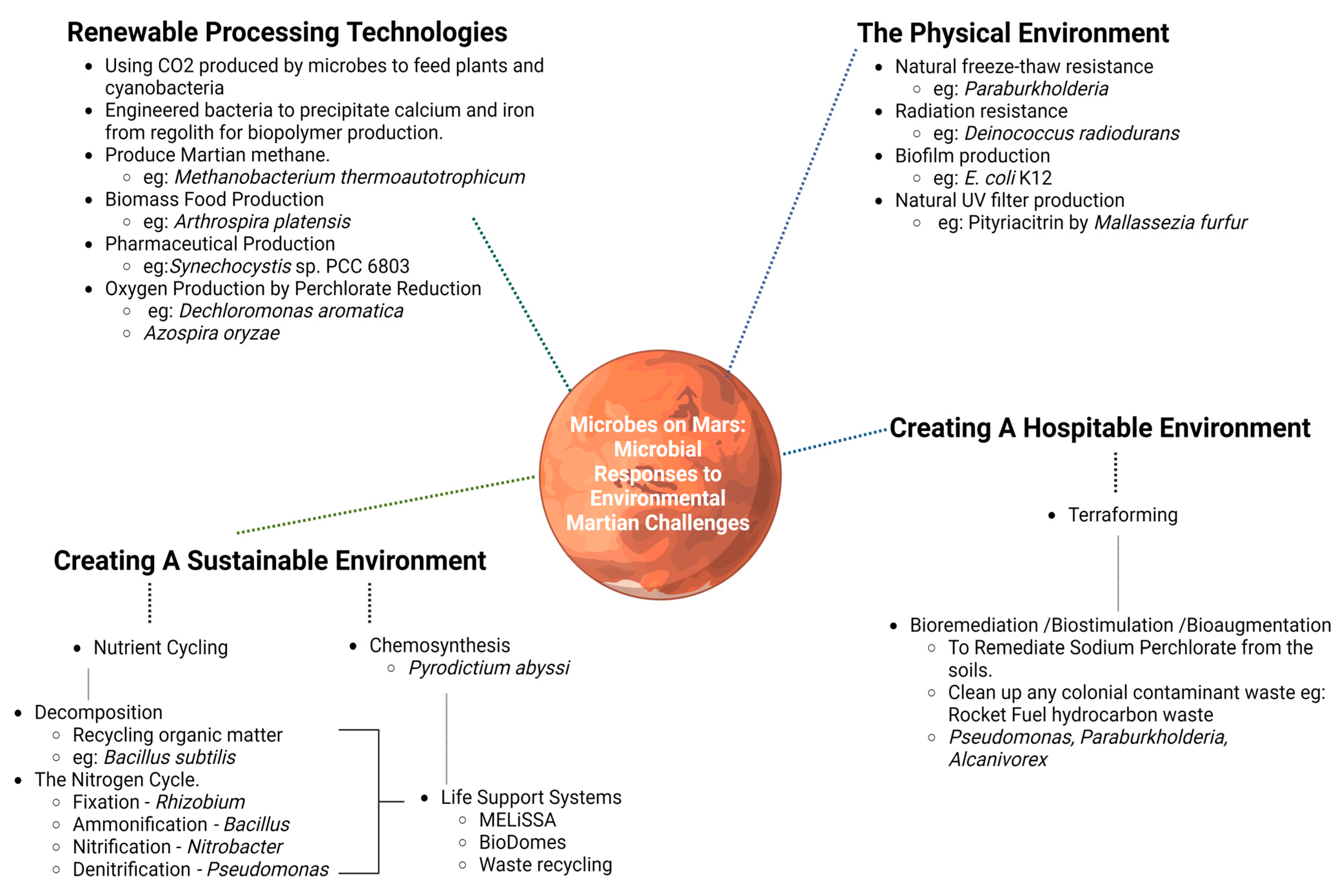
3. The Role of Bacteria in Combatting New Space Age Challenges
3.1. The Physical Environment
3.2. Terraforming
3.2.1. Microbes & Bioremediation
Understanding the Origins of Bioremediation
Transferable Environmental Uses of Microbes in the Modern World
3.3. Environmental Sustainability via Microbial Action
3.3.1. A Brief Overview of the Roles of Microbes in Nature
3.3.2. Microbial-Based Decomposition
The Nitrogen Cycle: Fixation, Ammonification, Nitrification and Denitrification
| Species Name | Role | Associated Gene(s) | Expression | References |
|---|---|---|---|---|
| Proteus mirabilis | Decomposition | ureR, fis, hybG, zapB, and PROSTU_03490 | The swarming of P. mirabilis on decaying organic material upregulates genes that produce a strong odour that specifically attracts Bot Flies (Lucilia sericata) which aid in the decomposition process. | [129] |
| Alphaproteobacteria | Decomposition | phoX | Alkaline phosphatase activity | [130] |
| Rhizobium meliloti | Nitrogen Fixation | nifHDK operon | nifHDK encodes for nitrogenase, an enzyme commonly used by bacteria in legume plant nodules which reduces free nitrogen (N2) into ammonia (NH3), which is used by the plant. | [131] |
| Bradyrhizobium japonicum | Nitrogen Fixation | FixL | A haemoprotein sensor histidine kinase to aid in fixation | [123] |
| Geobacillus kaustophilus | Chemosynthetic reduction of Chromium | ChrA | ChrA helps in the reduction of chromium from deep sea thermal vents, into less toxic forms of the metal | [78] |
3.4. Renewable Processing Technologies
4. Conclusions
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- Trevors, J.T. Review: From Chemosphere to Biosphere. World J. Microbiol. Biotechnol. 2001, 17, 651–655. [Google Scholar] [CrossRef]
- Vallet, M.; Kaftan, F.; Grabe, V.; Ghaderiardakani, F.; Fenizia, S.; Svatoš, A.; Pohnert, G.; Wichard, T. A New Glance at the Chemosphere of Macroalgal-Bacterial Interactions: In Situ Profiling of Metabolites in Symbiosis by Mass Spectrometry. Beilstein J. Org. Chem. 2021, 17, 1313–1322. [Google Scholar] [CrossRef] [PubMed]
- Oliveira, V.M.; Andreote, F.D.; Cortelo, P.C.; Castro-Gamboa, I.; Costa-Lotufo, L.V.; Polizeli, M.d.L.T.M.; Thiemann, O.H.; Setubal, J.C. Microorganisms: The Secret Agents of the Biosphere, and Their Key Roles in Biotechnology. Biota Neotrop. 2022, 22, e20221343. [Google Scholar] [CrossRef]
- Cavicchioli, R.; Ripple, W.J.; Timmis, K.N.; Azam, F.; Bakken, L.R.; Baylis, M.; Behrenfeld, M.J.; Boetius, A.; Boyd, P.W.; Classen, A.T.; et al. Scientists’ Warning to Humanity: Microorganisms and Climate Change. Nat. Rev. Microbiol. 2019, 17, 569–586. [Google Scholar] [CrossRef] [PubMed]
- Mannaa, M.; Han, G.; Seo, Y.-S.; Park, I. Evolution of Food Fermentation Processes and the Use of Multi-Omics in Deciphering the Roles of the Microbiota. Foods 2021, 10, 2861. [Google Scholar] [CrossRef]
- Hutchinson, C.R. Drug Synthesis by Genetically Engineered Microorganisms. Biotechnology 1994, 12, 375–380. [Google Scholar] [CrossRef]
- Abel, A.J.; Berliner, A.J.; Mirkovic, M.; Collins, W.D.; Arkin, A.P.; Clark, D.S. Photovoltaics-Driven Power Production Can Support Human Exploration on Mars. Front. Astron. Space Sci. 2022, 9, 868519. [Google Scholar] [CrossRef]
- Breyer, C. Low-Cost Solar Power Enables a Sustainable Energy Industry System. Proc. Natl. Acad. Sci. USA 2021, 118, e2116940118. [Google Scholar] [CrossRef]
- Kumar, V.; Paraschivoiu, M.; Paraschivoiu, I. Low Reynolds Number Vertical Axis Wind Turbine for Mars. Wind. Eng. 2010, 34, 461–476. [Google Scholar] [CrossRef]
- Morgan, P. Geothermal Energy on Mars. In Mars; Springer: Berlin/Heidelberg, Germany, 2009; pp. 331–349. ISBN 978-3-642-03629-3. [Google Scholar]
- Santomartino, R.; Zea, L.; Cockell, C.S. The Smallest Space Miners: Principles of Space Biomining. Extremophiles 2022, 26, 7. [Google Scholar] [CrossRef]
- Vincendon, M.; Mustard, J.; Forget, F.; Kreslavsky, M.; Spiga, A.; Murchie, S.; Bibring, J. Near-tropical Subsurface Ice on Mars. Geophys. Res. Lett. 2010, 37, L01202. [Google Scholar] [CrossRef]
- Mellerowicz, B.; Zacny, K.; Palmowski, J.; Bradley, B.; Stolov, L.; Vogel, B.; Ware, L.; Yen, B.; Sabahi, D.; Ridilla, A.; et al. RedWater: Water Mining System for Mars. New Space 2022, 10, 166–186. [Google Scholar] [CrossRef]
- Soureshjani, O.K.; Massumi, A.; Nouri, G. Sustainable Colonization of Mars Using Shape Optimized Structures and in Situ Concrete. Sci. Rep. 2023, 13, 15747. [Google Scholar] [CrossRef] [PubMed]
- Matarazzo, L.; Bettencourt, P.J.G. MRNA Vaccines: A New Opportunity for Malaria, Tuberculosis and HIV. Front. Immunol. 2023, 14, 1172691. [Google Scholar] [CrossRef]
- Lawler, A. Collaborative Research. Plans for Mars Unite Cancer, Space Agencies. Science 2000, 288, 415–416. [Google Scholar] [CrossRef] [PubMed]
- Dunbar, B.; Graf, A. NASA Funds Eight Studies to Protect Astronaut Health on Long Missions-NASA. Available online: https://www.nasa.gov/humans-in-space/nasa-funds-eight-studies-to-protect-astronaut-health-on-long-missions/ (accessed on 6 December 2023).
- Lovett, L. Mission to Mars: The Healthcare Challenges Facing NASA|MobiHealthNews. 2019. Available online: https://www.mobihealthnews.com/news/mission-mars-healthcare-challenges-facing-nasa (accessed on 17 September 2023).
- Manzey, D. Human Missions to Mars: New Psychological Challenges and Research Issues. Acta Astronaut. 2004, 55, 781–790. [Google Scholar] [CrossRef] [PubMed]
- Oluwafemi, F.A.; Abdelbaki, R.; Lai, J.C.-Y.; Mora-Almanza, J.G.; Afolayan, E.M. A Review of Astronaut Mental Health in Manned Missions: Potential Interventions for Cognitive and Mental Health Challenges. Life Sci. Space Res. 2021, 28, 26–31. [Google Scholar] [CrossRef]
- Horneck, G.; Facius, R.; Reitz, G.; Rettberg, P.; Baumstark-Khan, C.; Gerzer, R. Critical Issues in Connection with Human Missions to Mars: Protection of and from the Martian Environment. Adv. Space Res. 2003, 31, 87–95. [Google Scholar] [CrossRef]
- Jäkel, O. Radiation Hazard during a Manned Mission to Mars. Z. Med. Phys. 2004, 14, 267–272. [Google Scholar] [CrossRef]
- Shahab, M.; Shahab, N. Coevolution of the Human Host and Gut Microbiome: Metagenomics of Microbiota. Cureus 2022, 14, e26310. [Google Scholar] [CrossRef]
- Harper, K.N.; Armelagos, G.J. Genomics, the Origins of Agriculture, and Our Changing Microbe-Scape: Time to Revisit Some Old Tales and Tell Some New Ones. Am. J. Phys. Anthr. 2013, 152 (Suppl. S57), 135–152. [Google Scholar] [CrossRef]
- Rodríguez-Frías, F.; Quer, J.; Tabernero, D.; Cortese, M.F.; Garcia-Garcia, S.; Rando-Segura, A.; Pumarola, T. Microorganisms as Shapers of Human Civilization, from Pandemics to Even Our Genomes: Villains or Friends? A Historical Approach. Microorganisms 2021, 9, 2518. [Google Scholar] [CrossRef] [PubMed]
- Cavalieri, D.; McGovern, P.E.; Hartl, D.L.; Mortimer, R.; Polsinelli, M. Evidence for S. Cerevisiae Fermentation in Ancient Wine. J. Mol. Evol. 2003, 57, S226–S232. [Google Scholar] [CrossRef]
- Hagedorn, S.; Kaphammer, B. Microbial Biocatalysis in the Generation of Flavor and Fragrance Chemicals. Annu. Rev. Microbiol. 1994, 48, 773–800. [Google Scholar] [CrossRef] [PubMed]
- Marco, M.L.; Heeney, D.; Binda, S.; Cifelli, C.J.; Cotter, P.D.; Foligné, B.; Gä Nzle, M.; Kort, R.; Pasin, G.; Pihlanto, A.; et al. Health Benefits of Fermented Foods: Microbiota and Beyond. Curr. Opin. Biotechnol. 2017, 44, 94–102. [Google Scholar] [CrossRef]
- Lee, Y.-K.; Puong, K.-Y. Competition for Adhesion between Probiotics and Human Gastrointestinal Pathogens in the Presence of Carbohydrate. Br. J. Nutr. 2002, 88, S101. [Google Scholar] [CrossRef] [PubMed]
- Woo, J.; Ahn, J. Probiotic-Mediated Competition, Exclusion and Displacement in Biofilm Formation by Food-Borne Pathogens. Lett. Appl. Microbiol. 2013, 56, 307–313. [Google Scholar] [CrossRef] [PubMed]
- Ewaschuk, J.B.; Diaz, H.; Meddings, L.; Diederichs, B.; Dmytrash, A.; Backer, J.; Looijer-van Langen, M.; Madsen, K.L. Secreted Bioactive Factors from Bifidobacterium Infantis Enhance Epithelial Cell Barrier Function. Am. J. Physiol.-Gastrointest. Liver Physiol. 2008, 295, G1025–G1034. [Google Scholar] [CrossRef]
- Dobson, A.; Cotter, P.D.; Ross, R.P.; Hill, C. Bacteriocin Production: A Probiotic Trait? Appl. Environ. Microbiol. 2012, 78, 1–6. [Google Scholar] [CrossRef]
- Yang, S.-C.; Lin, C.-H.; Sung, C.T.; Fang, J.-Y. Antibacterial Activities of Bacteriocins: Application in Foods and Pharmaceuticals. Front. Microbiol. 2014, 5, 241. [Google Scholar] [CrossRef]
- Bernbom, N.; Licht, T.R.; Brogren, C.-H.; Jelle, B.; Johansen, A.H.; Badiola, I.; Vogensen, F.K.; Norrung, B. Effects of Lactococcus Lactis on Composition of Intestinal Microbiota: Role of Nisin. Appl. Environ. Microbiol. 2006, 72, 239–244. [Google Scholar] [CrossRef] [PubMed]
- Cleveland, J.; Montville, T.J.; Nes, I.F.; Chikindas, M.L. Bacteriocins: Safe, Natural Antimicrobials for Food Preservation. Int. J. Food Microbiol. 2001, 71, 1–20. [Google Scholar] [CrossRef] [PubMed]
- Rogers, L.A. The Inhibiting Effect of Streptococcus Lactis on Lactobacillus Bulgaricus. J. Bacteriol. 1928, 16, 321–325. [Google Scholar] [CrossRef] [PubMed]
- Zheng, Y.; Du, Y.; Qiu, Z.; Liu, Z.; Qiao, J.; Li, Y.; Caiyin, Q. Nisin Variants Generated by Protein Engineering and Their Properties. Bioengineering 2022, 9, 251. [Google Scholar] [CrossRef] [PubMed]
- Goudriaan, M.; Morales, V.H.; van der Meer, M.T.J.; Mets, A.; Ndhlovu, R.T.; van Heerwaarden, J.; Simon, S.; Heuer, V.B.; Hinrichs, K.-U.; Niemann, H. A Stable Isotope Assay with 13C-Labeled Polyethylene to Investigate Plastic Mineralization Mediated by Rhodococcus Ruber. Mar. Pollut. Bull. 2023, 186, 114369. [Google Scholar] [CrossRef] [PubMed]
- Saravanan, A.; Kumar, P.S.; Ramesh, B.; Srinivasan, S. Removal of Toxic Heavy Metals Using Genetically Engineered Microbes: Molecular Tools, Risk Assessment and Management Strategies. Chemosphere 2022, 298, 134341. [Google Scholar] [CrossRef] [PubMed]
- Singh, D.K. Biodegradation and Bioremediation of Pesticide in Soil: Concept, Method and Recent Developments. Indian. J. Microbiol. 2008, 48, 35–40. [Google Scholar] [CrossRef]
- French, K.E.; Zhou, Z.; Terry, N. Horizontal ‘Gene Drives’ Harness Indigenous Bacteria for Bioremediation. Sci. Rep. 2020, 10, 15091. [Google Scholar] [CrossRef]
- IQVIA Institute Global Market for Medicines to Rise to $1.9 Trillion by 2027, Says Report from IQVIA Institute-IQVIA. Available online: https://www.iqvia.com/newsroom/2023/01/global-market-for-medicines-to-rise-to-19-trillion-by-2027-says-report-from-iqvia-institute (accessed on 19 April 2023).
- Quianzon, C.C.; Cheikh, I. History of Insulin. J. Community Hosp. Intern. Med. Perspect. 2012, 2, 18701. [Google Scholar] [CrossRef]
- Fleming, A. Classics in Infectious Diseases: On the Antibacterial Action of Cultures of a Penicillium, with Special Reference to Their Use in the Isolation of B. Influenzae by Alexander Fleming, Reprinted from the British Journal of Experimental Pathology 10:226–236. Rev. Infect. Dis. 1929, 2, 129–139. [Google Scholar] [CrossRef]
- CDC Antibiotic/Antimicrobial Resistance|CDC. Available online: https://www.cdc.gov/drugresistance/index.html (accessed on 17 October 2018).
- Peplow, M. How Mars Got Its Rust. Nature 2004. [Google Scholar] [CrossRef]
- Hecht, M.; Kounaves, S.; Quinn, R.; West, S.; Young, S.; Ming, D.; Catling, D.; Clark, B.; Boynton, W.; Hoffman, J.; et al. Detection of Perchlorate and the Soluble Chemistry of Martian Soil at the Phoenix Lander Site. Science 2009, 325, 64–67. [Google Scholar] [CrossRef] [PubMed]
- Davila, A.F.; Willson, D.; Coates, J.D.; McKay, C.P.; Davila, A.F.; Willson, D.; Coates, J.D.; McKay, C.P. Perchlorate on Mars: A Chemical Hazard and a Resource for Humans. Int. J. Astrobiol. 2013, 12, 321–325. [Google Scholar] [CrossRef]
- Srinivasan, A.; Viraraghavan, T. Perchlorate: Health Effects and Technologies for Its Removal from Water Resources. Int. J. Environ. Res. Public Health 2009, 6, 1418–1442. [Google Scholar] [CrossRef] [PubMed]
- Sinton, W.M. Taking the Temperatures of the Moon and Planets. Astron. Soc. Pac. Leafl. 1958, 7, 361. [Google Scholar]
- Moissl-Eichinger, C.; Cockell, C.; Rettberg, P. Venturing into New Realms? Microorganisms in Space. FEMS Microbiol. Rev. 2016, 40, 722–737. [Google Scholar] [CrossRef]
- NASA Mars Facts|All about Mars–NASA’s Mars Exploration Program. Available online: https://mars.nasa.gov/all-about-mars/facts/ (accessed on 13 April 2020).
- Catling, D.C.; Cockell, C.S.; Mckay, C.P. Ultraviolet Radiation on the Surface of MARS. In Proceedings of the Fifth International Conference on Mars, Pasadena, CA, USA, 18–23 June 1999. [Google Scholar]
- Cockell, C.S.; Catling, D.C.; Davis, W.L.; Snook, K.; Kepner, R.L.; Lee, P.; McKay, C.P. The Ultraviolet Environment of Mars: Biological Implications Past, Present, and Future. Icarus 2000, 146, 343–359. [Google Scholar] [CrossRef]
- Pattison, D.I.; Davies, M.J. Actions of Ultraviolet Light on Cellular Structures. In Cancer: Cell Structures, Carcinogens and Genomic Instability; Birkhäuser-Verlag: Basel, Switzerland, 2006; pp. 131–157. [Google Scholar]
- Keaney, D.; Lucey, B.; Quinn, N.; Finn, K. The Effects of Freeze-Thaw and UVC Radiation on Microbial Survivability in a Selected Mars-like Environment. Microorganisms 2022, 10, 576. [Google Scholar] [CrossRef]
- Mathewson, S. Algae “Bioreactor” on Space Station Could Make Oxygen, Food for Astronauts|Space. Available online: https://www.space.com/space-station-algae-experiment-fresh-air.html (accessed on 11 June 2019).
- Machowinski, A.; Kramer, H.-J.; Hort, W.; Mayser, P. Pityriacitrin ? A Potent UV Filter Produced by Malassezia Furfur and Its Effect on Human Skin Microflora. Mycoses 2006, 49, 388–392. [Google Scholar] [CrossRef]
- Besemer, K. Biodiversity, Community Structure and Function of Biofilms in Stream Ecosystems. Res. Microbiol. 2015, 166, 774–781. [Google Scholar] [CrossRef]
- Baqué, M.; de Vera, J.-P.; Rettberg, P.; Billi, D. The BOSS and BIOMEX Space Experiments on the EXPOSE-R2 Mission: Endurance of the Desert Cyanobacterium Chroococcidiopsis under Simulated Space Vacuum, Martian Atmosphere, UVC Radiation and Temperature Extremes. Acta Astronaut. 2013, 91, 180–186. [Google Scholar] [CrossRef]
- DePas, W.H.; Syed, A.K.; Sifuentes, M.; Lee, J.S.; Warshaw, D.; Saggar, V.; Csankovszki, G.; Boles, B.R.; Chapman, M.R. Biofilm Formation Protects Escherichia Coli against Killing by Caenorhabditis Elegans and Myxococcus Xanthus. Appl. Environ. Microbiol. 2014, 80, 7079–7087. [Google Scholar] [CrossRef] [PubMed]
- Liu, W.; Røder, H.L.; Madsen, J.S.; Bjarnsholt, T.; Sørensen, S.J.; Burmølle, M. Interspecific Bacterial Interactions Are Reflected in Multispecies Biofilm Spatial Organization. Front. Microbiol. 2016, 7, 1366. [Google Scholar] [CrossRef] [PubMed]
- Makarova, K.S.; Aravind, L.; Wolf, Y.I.; Tatusov, R.L.; Minton, K.W.; Koonin, E.V.; Daly, M.J. Genome of the Extremely Radiation-Resistant Bacterium Deinococcus Radiodurans Viewed from the Perspective of Comparative Genomics. Microbiol. Mol. Biol. Rev. 2001, 65, 44–79. [Google Scholar] [CrossRef] [PubMed]
- Sleator, R.D.; Smith, N. Directed Panspermia: A 21st Century Perspective. Sci. Prog. 2017, 100, 187–193. [Google Scholar] [CrossRef] [PubMed]
- De Rosa, M.; Verdino, A.; Soriente, A.; Marabotti, A. The Odd Couple(s): An Overview of Beta-Lactam Antibiotics Bearing More Than One Pharmacophoric Group. Int. J. Mol. Sci. 2021, 22, 617. [Google Scholar] [CrossRef]
- Schiwon, K.; Arends, K.; Rogowski, K.M.; Fürch, S.; Prescha, K.; Sakinc, T.; Van Houdt, R.; Werner, G.; Grohmann, E. Comparison of Antibiotic Resistance, Biofilm Formation and Conjugative Transfer of Staphylococcus and Enterococcus Isolates from International Space Station and Antarctic Research Station Concordia. Microb. Ecol. 2013, 65, 638–651. [Google Scholar] [CrossRef]
- Nickerson, C.A.; Ott, C.M.; Wilson, J.W.; Ramamurthy, R.; Pierson, D.L. Microbial Responses to Microgravity and Other Low-Shear Environments. Microbiol. Mol. Biol. Rev. 2004, 68, 345–361. [Google Scholar] [CrossRef]
- Castro, S.L.; Nelman-Gonzalez, M.; Nickerson, C.A.; Ott, C.M. Induction of Attachment-Independent Biofilm Formation and Repression of Hfq Expression by Low-Fluid-Shear Culture of Staphylococcus Aureus. Appl. Environ. Microbiol. 2011, 77, 6368–6378. [Google Scholar] [CrossRef]
- Love, S. Bacteria Get Dangerously Weird in Space|Indy100. Available online: https://www.indy100.com/article/bacteria-get-dangerously-weird-in-space-7380481 (accessed on 29 October 2018).
- Haynes, R.H.; McKay, C.P. The Implantation of Life on Mars: Feasibility and Motivation. Adv. Space Res. 1992, 12, 133–140. [Google Scholar] [CrossRef]
- Zubrin, R.; McKay, C. Technological Requirements for Terraforming Mars. Available online: http://www.users.globalnet.co.uk/~mfogg/zubrin.htm (accessed on 10 November 2018).
- Hamer, G. Bioremediation: A Response to Gross Environmental Abuse. Trends Biotechnol. 1993, 11, 317–319. [Google Scholar] [CrossRef]
- Gribaldo, S.; Brochier-Armanet, C. The Origin and Evolution of Archaea: A State of the Art. Philos. Trans. R. Soc. Lond. B Biol. Sci. 2006, 361, 1007–1022. [Google Scholar] [CrossRef] [PubMed]
- Dodd, M.S.; Papineau, D.; Grenne, T.; Slack, J.F.; Rittner, M.; Pirajno, F.; O’Neil, J.; Little, C.T.S. Evidence for Early Life in Earth’s Oldest Hydrothermal Vent Precipitates. Nature 2017, 543, 60. [Google Scholar] [CrossRef]
- Schopf, J.W.; Logan, G.A.; Buick, R.; Summons, R.E. Microfossils of the Early Archean Apex Chert: New Evidence of the Antiquity of Life. Science 1993, 260, 640–646. [Google Scholar] [CrossRef] [PubMed]
- Sohlenkamp, C. Crossing the Lipid Divide. J. Biol. Chem. 2021, 297, 100859. [Google Scholar] [CrossRef] [PubMed]
- Hoagland, P.; Beaulieu, S.; Tivey, M.A.; Eggert, R.G.; German, C.; Glowka, L.; Lin, J. Deep-Sea Mining of Seafloor Massive Sulfides. Mar. Policy 2010, 34, 728–732. [Google Scholar] [CrossRef]
- Sar, P.; Kazy, S.K.; Paul, D.; Sarkar, A. Metal Bioremediation by Thermophilic Microorganisms. In Thermophilic Microbes in Environmental and Industrial Biotechnology; Springer: Dordrecht, The Netherlands, 2013; pp. 171–201. [Google Scholar]
- Al-Daghistani, H. Internet Scientific Publications. Available online: http://ispub.com/IJMB/10/1/13756# (accessed on 25 April 2019).
- van der Meer, J.R.; de Vos, W.M.; Harayama, S.; Zehnder, A.J. Molecular Mechanisms of Genetic Adaptation to Xenobiotic Compounds. Microbiol. Rev. 1992, 56, 677. [Google Scholar] [CrossRef]
- Springael, D.; Top, E.M. Horizontal Gene Transfer and Microbial Adaptation to Xenobiotics: New Types of Mobile Genetic Elements and Lessons from Ecological Studies. Trends Microbiol. 2004, 12, 53–58. [Google Scholar] [CrossRef]
- Sharma, S. Bioremediation: Features, Strategies and Applications. Asian J. Pharm. Life Sci. 2012, 2, 2. [Google Scholar]
- Pacific Northwest National Laboratory In Situ Bioremediation. Available online: https://bioprocess.pnnl.gov/resour/rt3d.in.situ.bioremediation.htm (accessed on 26 April 2019).
- Tomei, M.C.; Daugulis, A.J. Ex Situ Bioremediation of Contaminated Soils: An Overview of Conventional and Innovative Technologies. Crit. Rev. Environ. Sci. Technol. 2013, 43, 2107–2139. [Google Scholar] [CrossRef]
- US EPA. How to Evaluate Alternative Cleanup Technologies for Underground Storage Tank Sites—A Guide for Corrective Action Plan Reviewers; Biosparging. 1994; Chapter 8. Available online: https://www.epa.gov/ust/how-evaluate-alternative-cleanup-technologies-underground-storage-tank-sites-guide-corrective (accessed on 17 September 2023).
- Chapelle, F.H.; O’Neill, K.; Bradley, P.M.; Methé, B.A.; Ciufo, S.A.; Knobel, L.L.; Lovley, D.R. A Hydrogen-Based Subsurface Microbial Community Dominated by Methanogens. Nature 2002, 415, 312–315. [Google Scholar] [CrossRef]
- Aslam, M.; Charfi, A.; Lesage, G.; Heran, M.; Kim, J. Membrane Bioreactors for Wastewater Treatment: A Review of Mechanical Cleaning by Scouring Agents to Control Membrane Fouling. Chem. Eng. J. 2017, 307, 897–913. [Google Scholar] [CrossRef]
- European Commission. In-Depth Report Soil Contamination: Impacts on Human Health. Science for Environment Policy. September 2013. Available online: http://ec.europa.eu/science-environment-policy (accessed on 17 September 2023).
- Gouda, M.K.; Omar, S.H.; Nour Eldin, H.M.; Chekroud, Z.A. Bioremediation of Kerosene II: A Case Study in Contaminated Clay (Laboratory and Field: Scale Microcosms). World J. Microbiol. Biotechnol. 2008, 24, 1451–1460. [Google Scholar] [CrossRef]
- Ezzariai, A.; Hafidi, M.; Khadra, A.; Aemig, Q.; El Fels, L.; Barret, M.; Merlina, G.; Patureau, D.; Pinelli, E. Human and Veterinary Antibiotics during Composting of Sludge or Manure: Global Perspectives on Persistence, Degradation, and Resistance Genes. J. Hazard. Mater. 2018, 359, 465–481. [Google Scholar] [CrossRef] [PubMed]
- Koch, N.; Islam, N.F.; Sonowal, S.; Prasad, R.; Sarma, H. Environmental Antibiotics and Resistance Genes as Emerging Contaminants: Methods of Detection and Bioremediation. Curr. Res. Microb. Sci. 2021, 2, 100027. [Google Scholar] [CrossRef] [PubMed]
- Agwuh, K.N.; MacGowan, A. Pharmacokinetics and Pharmacodynamics of the Tetracyclines Including Glycylcyclines. J. Antimicrob. Chemother. 2006, 58, 256–265. [Google Scholar] [CrossRef] [PubMed]
- Al-Gheethi, A.A.S.; Lalung, J.; Noman, E.A.; Bala, J.D.; Norli, I. Removal of Heavy Metals and Antibiotics from Treated Sewage Effluent by Bacteria. Clean. Technol. Environ. Policy 2015, 17, 2101–2123. [Google Scholar] [CrossRef]
- Al-Gheethi, A.A.S.; Norli, I.; Lalung, J.; Megat Azlan, A.; Nur Farehah, Z.A.; Kadir, M.O.A. Biosorption of Heavy Metals and Cephalexin from Secondary Effluents by Tolerant Bacteria. Clean. Technol. Environ. Policy 2014, 16, 137–148. [Google Scholar] [CrossRef]
- Tyagi, M.; da Fonseca, M.M.R.; de Carvalho, C.C.C.R. Bioaugmentation and Biostimulation Strategies to Improve the Effectiveness of Bioremediation Processes. Biodegradation 2011, 22, 231–241. [Google Scholar] [CrossRef]
- Kanissery, R.G.; Sims, G.K. Biostimulation for the Enhanced Degradation of Herbicides in Soil. Appl. Environ. Soil. Sci. 2011, 2011, 843450. [Google Scholar] [CrossRef]
- Joshi, M.N.; Dhebar, S.V.; Dhebar, S.V.; Bhargava, P.; Pandit, A.; Patel, R.P.; Saxena, A.; Bagatharia, S.B. Metagenomics of Petroleum Muck: Revealing Microbial Diversity and Depicting Microbial Syntrophy. Arch. Microbiol. 2014, 196, 531–544. [Google Scholar] [CrossRef] [PubMed]
- Samanta, S.K.; Singh, O.V.; Jain, R.K. Polycyclic Aromatic Hydrocarbons: Environmental Pollution and Bioremediation. Trends Biotechnol. 2002, 20, 243–248. [Google Scholar] [CrossRef]
- Salinero, K.; Keller, K.; Feil, W.S.; Feil, H.; Trong, S.; Di Bartolo, G.; Lapidus, A. Metabolic Analysis of the Soil Microbe Dechloromonas Aromatica Str. RCB: Indications of a Surprisingly Complex Life-Style and Cryptic Anaerobic Pathways for Aromatic Degradation. BMC Genom. 2009, 10, 351. [Google Scholar] [CrossRef] [PubMed]
- Brim, H.; McFarlan, S.C.; Fredrickson, J.K.; Minton, K.W.; Zhai, M.; Wackett, L.P.; Daly, M.J. Engineering Deinococcus Radiodurans for Metal Remediation in Radioactive Mixed Waste Environments. Nat. Biotechnol. 2000, 18, 85–90. [Google Scholar] [CrossRef]
- Genovese, M.; Crisafi, F.; Denaro, R.; Cappello, S.; Russo, D.; Calogero, R.; Santisi, S.; Catalfamo, M.; Modica, A.; Smedile, F.; et al. Effective Bioremediation Strategy for Rapid in Situ Cleanup of Anoxic Marine Sediments in Mesocosm Oil Spill Simulation. Front. Microbiol. 2014, 5, 162. [Google Scholar] [CrossRef] [PubMed]
- Elliott, G.N.; Chen, W.-M.; Chou, J.-H.; Wang, H.-C.; Sheu, S.-Y.; Perin, L.; Reis, V.M.; Moulin, L.; Simon, M.F.; Bontemps, C.; et al. Burkholderia Phymatum Is a Highly Effective Nitrogen-Fixing Symbiont of Mimosa spp. and Fixes Nitrogen Ex. Planta. New Phytol. 2007, 173, 168–180. [Google Scholar] [CrossRef] [PubMed]
- Andreolli, M.; Lampis, S.; Zenaro, E.; Salkinoja-Salonen, M.; Vallini, G. Burkholderia Fungorum DBT1: A Promising Bacterial Strain for Bioremediation of PAHs-Contaminated Soils. FEMS Microbiol. Lett. 2011, 319, 11–18. [Google Scholar] [CrossRef]
- Mrozik, A.; Piotrowska-Seget, Z. Bioaugmentation as a Strategy for Cleaning up of Soils Contaminated with Aromatic Compounds. Microbiol. Res. 2010, 165, 363–375. [Google Scholar] [CrossRef]
- Hong, Q.; Zhang, Z.; Hong, Y.; Li, S. A Microcosm Study on Bioremediation of Fenitrothion-Contaminated Soil Using Burkholderia sp. FDS-1. Int. Biodeterior. Biodegrad. 2007, 59, 55–61. [Google Scholar] [CrossRef]
- US EPA. Deepwater Horizon–BP Gulf of Mexico Oil Spill. 2017. Available online: https://www.epa.gov/enforcement/deepwater-horizon-bp-gulf-mexico-oil-spill (accessed on 17 September 2023).
- Barron, M.G. Ecological Impacts of the Deepwater Horizon Oil Spill: Implications for Immunotoxicity. Toxicol. Pathol. 2012, 40, 315–320. [Google Scholar] [CrossRef]
- Hazen, T.C.; Dubinsky, E.A.; DeSantis, T.Z.; Andersen, G.L.; Piceno, Y.M.; Singh, N.; Jansson, J.K.; Probst, A.; Borglin, S.E.; Fortney, J.L.; et al. Deep-Sea Oil Plume Enriches Indigenous Oil-Degrading Bacteria. Science 2010, 330, 204–208. [Google Scholar] [CrossRef] [PubMed]
- Atlas, R.M.; Hazen, T.C. Oil Biodegradation and Bioremediation: A Tale of the Two Worst Spills in U.S. History. Env. Sci. Technol. 2011, 45, 6709. [Google Scholar] [CrossRef] [PubMed]
- Kessler, J.D.; Valentine, D.L.; Redmond, M.C.; Du, M.; Chan, E.W.; Mendes, S.D.; Quiroz, E.W.; Villanueva, C.J.; Shusta, S.S.; Werra, L.M.; et al. A Persistent Oxygen Anomaly Reveals the Fate of Spilled Methane in the Deep Gulf of Mexico. Science 2011, 331, 312–315. [Google Scholar] [CrossRef]
- Kalenitchenko, D.; Le Bris, N.; Dadaglio, L.; Peru, E.; Besserer, A.; Galand, P.E. Bacteria Alone Establish the Chemical Basis of the Wood-Fall Chemosynthetic Ecosystem in the Deep-Sea. ISME J. 2018, 12, 367–379. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.; Xiao, L.; Li, F.; Xiao, M.; Lin, D.; Long, X.; Wu, Z. Microbial Degradation of Pesticide Residues and an Emphasis on the Degradation of Cypermethrin and 3-Phenoxy Benzoic Acid: A Review. Molecules 2018, 23, 2313. [Google Scholar] [CrossRef]
- Metcalf, J.L.; Carter, D.O.; Knight, R. Microbiology of Death. Curr. Biol. 2016, 13, 561–563. [Google Scholar]
- Scharlemann, J.P.; Tanner, E.V.; Hiederer, R.; Kapos, V. Global Soil Carbon: Understanding and Managing the Largest Terrestrial Carbon Pool. Carbon. Manag. 2014, 5, 81–91. [Google Scholar] [CrossRef]
- Shaw, M.R.; Zavaleta, E.S.; Chiariello, N.R.; Cleland, E.E.; Mooney, H.A.; Field, C.B. Grassland Responses to Global Environmental Changes Suppressed by Elevated CO2. Science 2002, 298, 1987–1990. [Google Scholar] [CrossRef]
- Cheng, L.; Zhang, N.; Yuan, M.; Xiao, J.; Qin, Y.; Deng, Y.; Tu, Q.; Xue, K.; Van Nostrand, J.D.; Wu, L.; et al. Warming Enhances Old Organic Carbon Decomposition through Altering Functional Microbial Communities. ISME J. 2017, 11, 1825–1835. [Google Scholar] [CrossRef]
- Tsang, J. Changing CO2 Levels Require Microbial Coping Strategies. Available online: https://www.asm.org/Articles/2019/April/Changing-CO2-Levels-Means-Different-Coping-Strateg (accessed on 24 July 2019).
- Carroll, K.C.; Pfaller, M.A.; Landry, M.L.; McAdam, A.J.; Patel, R.; Richter, S.S.; Warnock, D.W. (Eds.) Manual of Clinical Microbiology, 12th ed.; American Society of Microbiology: Washington, DC, USA, 2019; ISBN 9781555819835. [Google Scholar]
- DeVries, T.; Holzer, M.; Primeau, F. Recent Increase in Oceanic Carbon Uptake Driven by Weaker Upper-Ocean Overturning. Nature 2017, 542, 215–218. [Google Scholar] [CrossRef]
- Centralblatt Für Bakteriologie, Parasitenkunde Und Infektionskrankheiten; G. Fischer: Jena, Germany, 1902. Available online: https://www.biodiversitylibrary.org/bibliography/77475 (accessed on 23 April 2019).
- Lesser, M.P.; Mazel, C.H.; Gorbunov, M.Y.; Falkowski, P.G. Discovery of Symbiotic Nitrogen-Fixing Cyanobacteria in Corals. Science 2004, 305, 997–1000. [Google Scholar] [CrossRef] [PubMed]
- Postgate, J.R. Biology Nitrogen Fixation: Fundamentals. Philos. Trans. R. Soc. Lond B Biol. Sci. 1982, 296, 375–385. [Google Scholar]
- Dixon, R.; Kahn, D. Genetic Regulation of Biological Nitrogen Fixation. Nat. Rev. Microbiol. 2004, 2, 621–631. [Google Scholar] [CrossRef]
- Jeong, H.S.; Jouanneau, Y. Enhanced Nitrogenase Activity in Strains of Rhodobacter Capsulatus That Overexpress the Rnf Genes. J. Bacteriol. 2000, 182, 1208–1214. [Google Scholar] [CrossRef] [PubMed]
- Nag, P.; Pal, S. Fe Protein Over-Expression Can Enhance the Nitrogenase Activity of Azotobacter Vinelandii. J. Basic. Microbiol. 2013, 53, 156–162. [Google Scholar] [CrossRef] [PubMed]
- Alley, R.; Wratt, D. Climate Change 2007: The Physical Science. Available online: https://web.archive.org/web/20110716022730/http://www.uclm.es/area/amf/Antoine/Fusion/Ipcc_anotado.pdf (accessed on 23 April 2019).
- Rabalais, N.N. Nitrogen in Aquatic Ecosystems. AMBIO A J. Hum. Environ. 2002, 31, 102–112. [Google Scholar] [CrossRef]
- Bhagowati, B.; Ahamad, K.U. A Review on Lake Eutrophication Dynamics and Recent Developments in Lake Modeling. Ecohydrol. Hydrobiol. 2019, 19, 155–166. [Google Scholar] [CrossRef]
- Ma, Q.; Fonseca, A.; Liu, W.; Fields, A.T.; Pimsler, M.L.; Spindola, A.F.; Tarone, A.M.; Crippen, T.L.; Tomberlin, J.K.; Wood, T.K. Proteus Mirabilis Interkingdom Swarming Signals Attract Blow Flies. ISME J. 2012, 6, 1356–1366. [Google Scholar] [CrossRef]
- Dai, J.; Chen, D.; Wu, S.; Wu, X.; Gao, G.; Tang, X.; Shao, K.; Lv, X.; Xue, W.; Yang, Q.; et al. Dynamics of Phosphorus and Bacterial PhoX Genes during the Decomposition of Microcystis Blooms in a Mesocosm. PLoS ONE 2018, 13, e0195205. [Google Scholar] [CrossRef]
- Szeto, W.W.; Lynn Zimmerman, J.; Sundaresan, V.; Ausubel, F.M. A Rhizobium Meliloti Symbiotic Regulatory Gene. Cell 1984, 36, 1035–1043. [Google Scholar] [CrossRef]
- Covich, A.P. Energy Flow and Ecosystems. Encycl. Biodivers. 2013, 237–249. [Google Scholar] [CrossRef]
- Hooper, A.B.; DiSpirito, A.A. Chemolithotrophy. Encycl. Biol. Chem. 2013, 486–492. [Google Scholar] [CrossRef]
- Boschker, H.T.S.; Vasquez-Cardenas, D.; Bolhuis, H.; Moerdijk-Poortvliet, T.W.C.; Moodley, L. Chemoautotrophic Carbon Fixation Rates and Active Bacterial Communities in Intertidal Marine Sediments. PLoS ONE 2014, 9, e101443. [Google Scholar] [CrossRef] [PubMed]
- Gazda, L.S.O.T. The Microbes That Keep Hydrothermal Vents Pumping|Smithsonian Ocean. Available online: https://ocean.si.edu/ecosystems/deep-sea/microbes-keep-hydrothermal-vents-pumping (accessed on 23 April 2019).
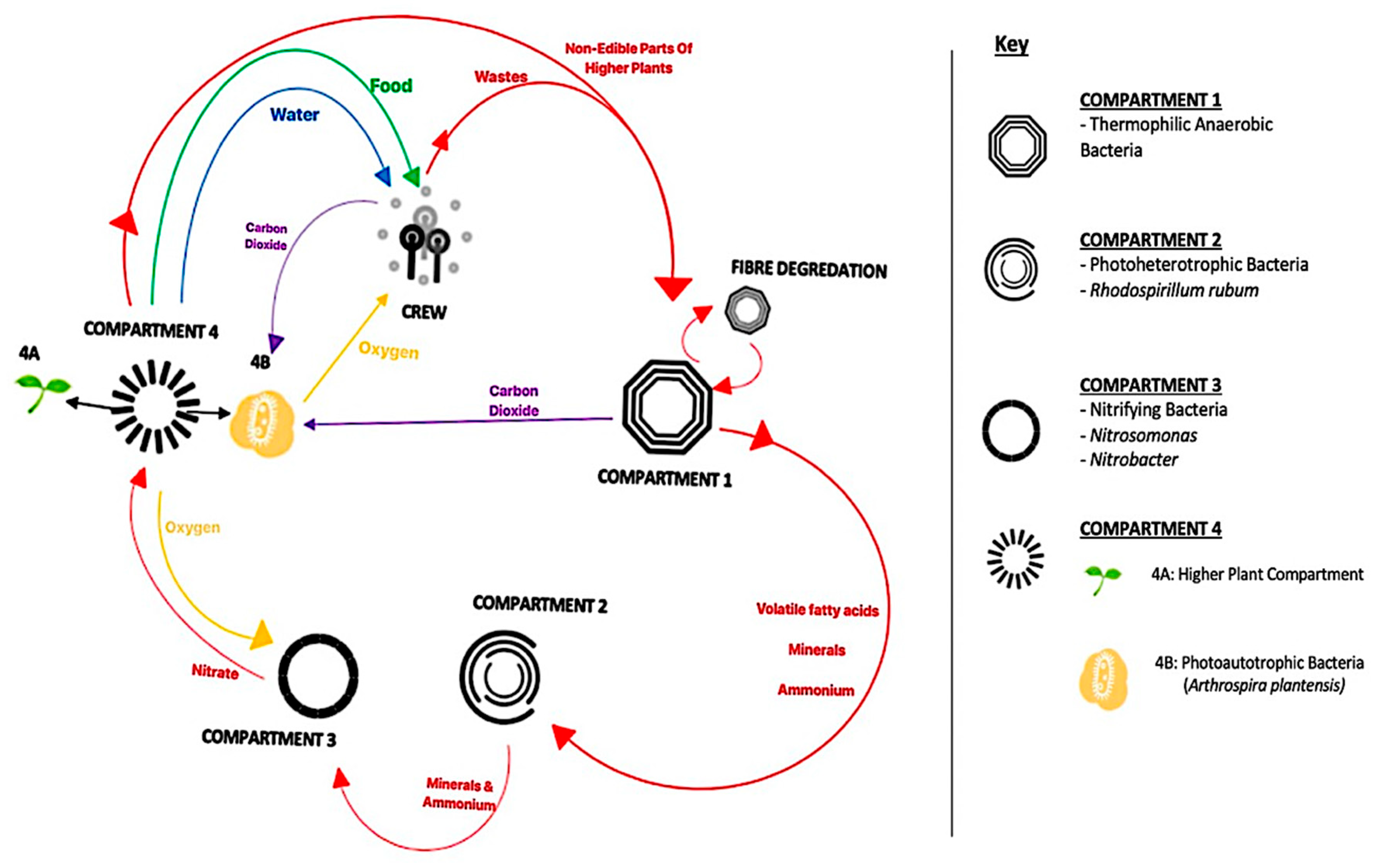
- Lasseur, C.; Brunet, J.; De Weever, H.; Dixon, M.; Dussap, G.; Godia, F.; Leys, N.; Mergeay, M.; Van Der Straeten, D. Melissa: The European Project of Closed Life Support System. Am. Soc. Gravit. Sp. Biol. 2010, 23, 3. [Google Scholar]
- Wetterer, J.K.; Miller, S.E.; Wheeler, D.E.; Olson, C.A.; Polhemus, D.A.; Pitts, M.; Ashton, I.W.; Himler, A.G.; Yospin, M.M.; Helms, K.R.; et al. Ecological Dominance by Paratrechina Longicornis (Hymenoptera: Formicidae), an Invasive Tramp Ant, in Biosphere 2. Fla. Entomol. 1999, 82, 381. [Google Scholar] [CrossRef]
- Nelson, M.; Dempster, W.F.; Allen, J.P. The Water Cycle in Closed Ecological Systems: Perspectives from the Biosphere 2 and Laboratory Biosphere Systems. Adv. Space Res. 2009, 44, 1404–1412. [Google Scholar] [CrossRef]
- Scherson, Y.D.; Wells, G.F.; Woo, S.-G.; Lee, J.; Park, J.; Cantwell, B.J.; Criddle, C.S. Nitrogen Removal with Energy Recovery through N2O Decomposition. Energy Environ. Sci. 2013, 6, 241–248. [Google Scholar] [CrossRef]
- De Muynck, W.; De Belie, N.; Verstraete, W. Microbial Carbonate Precipitation in Construction Materials: A Review. Ecol. Eng. 2010, 36, 118–136. [Google Scholar] [CrossRef]
- Menezes, A.A.; Cumbers, J.; Hogan, J.A.; Arkin, A.P. Towards Synthetic Biological Approaches to Resource Utilization on Space Missions. J. R. Soc. Interface 2015, 12, 20140715. [Google Scholar] [CrossRef]
- Stern, J.C.; Sutter, B.; Freissinet, C.; Navarro-González, R.; McKay, C.P.; Archer, P.D.; Buch, A.; Brunner, A.E.; Coll, P.; Eigenbrode, J.L.; et al. Evidence for Indigenous Nitrogen in Sedimentary and Aeolian Deposits from the Curiosity Rover Investigations at Gale Crater, Mars. Proc. Natl. Acad. Sci. USA 2015, 112, 4245–4250. [Google Scholar] [CrossRef]
- Williams, D. Mars Fact Sheet. Available online: https://nssdc.gsfc.nasa.gov/planetary/factsheet/marsfact.html (accessed on 23 April 2019).
- Motzer, W. Perchlorate: Problems, Detection, and Solutions. Environ. Forensics 2001, 2, 301–311. [Google Scholar] [CrossRef]
- Renner, R. Food Safety: Perchlorate Exposure: Tip of the Iceberg? Environ. Health Perspect. 2005, 113, A232. [Google Scholar] [CrossRef][Green Version]
- Glavin, D.P.; Freissinet, C.; Miller, K.E.; Eigenbrode, J.L.; Brunner, A.E.; Buch, A.; Sutter, B.; Archer, P.D.; Atreya, S.K.; Brinckerhoff, W.B.; et al. Evidence for Perchlorates and the Origin of Chlorinated Hydrocarbons Detected by SAM at the Rocknest Aeolian Deposit in Gale Crater. J. Geophys. Res. Planets 2013, 118, 1955–1973. [Google Scholar] [CrossRef]
- Becker, C. Prophylaxis and Treatment of Side Effects Due to Iodinated Contrast Media Relevant to Radiological Practice. Radiologe 2007, 47, 768–773. [Google Scholar] [CrossRef]
- Seiler, M.A.; Jensen, D.; Neist, U.; Deister, U.K.; Schmitz, F. Validation Data for the Determination of Perchlorate in Water Using Ion Chromatography with Suppressed Conductivity Detection. Environ. Sci. Eur. 2016, 28, 18. [Google Scholar] [CrossRef]
- Seiler, M.A.; Jensen, D.; Neist, U.; Deister, U.K.; Schmitz, F. Determination of Trace Perchlorate in Water: A Simplified Method for the Identification of Potential Interferences. Environ. Sci. Eur. 2017, 29, 30. [Google Scholar] [CrossRef]
- Luo, Y.; Naidu, R.; Fang, C. Raman Imaging towards In-Situ Visualisation of Perchlorate Adsorption. Water Res. 2023, 229, 119510. [Google Scholar] [CrossRef]
| Initiative | Description | Application | References |
|---|---|---|---|
| In-Situ Bioremediation | The use of bioremediation “on site” where the pollution has occurred. | Treatment of wastewater, groundwater, soil/sludge, gas. | [83] |
| Ex-Situ Bioremediation | The bioremediation of foreign substances by indigenous bacteria. | Composting. Land farming. | [84] |
| Biosparging | The injection of air into a contaminated area in order to stimulate the bioremediation properties of inhabiting bacteria. | Treatment of polluted soils and groundwater. | [85,86] |
| Bioaugmentation | The introduction of non-indigenous bacteria to a site of pollution in order to stimulate bioremediation. | Treatment of polluted soils and xenobiotics. | [86] |
| Bioreactors | The introduction of liquid-state polluted material into an enclosed controlled environment, home to bioremediation microbes. | Wastewater treatment. | [87] |
| Bacterium | Substrate | Uses | References |
|---|---|---|---|
| Pseudomonas stutzeri | Xylene, toluene, phenol and polycyclic aromatic hydrocarbons. | Universally in soils. | [97] |
| Pseudomonas putida | Toluene, phenanthrene and naphthalene. | Universally in soils. | [98] |
| Dechloromonas aromatica | Benzoate, perchlorate, chlorobenzoate, and toluene. | In situ bioremediation, especially in ground/surface water environments. | [99] |
| Deinococcus radiodurans | Ionic mercury and toluene. | Radioactive mixed waste environments. | [100] |
| Alcanivorex borkumensis | Fuel hydrocarbons. | Marine environments; used during the Deepwater Horizon oil spill in the Gulf of Mexico. | [101] |
| Paraburkholderia phymatum | Xylene, toluene, phenol and polycyclic aromatic hydrocarbons. | Universally in soils. | [102] |
| Paraburkholderia fungorum | Dibenzothiophene, fluorene, naphthalene and phenanthrene, along with polycyclic aromatic hydrocarbons other than condensed thiophenes. | Universally in soils. | [103] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2024 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Keaney, D.; Lucey, B.; Finn, K. A Review of Environmental Challenges Facing Martian Colonisation and the Potential for Terrestrial Microbes to Transform a Toxic Extraterrestrial Environment. Challenges 2024, 15, 5. https://doi.org/10.3390/challe15010005
Keaney D, Lucey B, Finn K. A Review of Environmental Challenges Facing Martian Colonisation and the Potential for Terrestrial Microbes to Transform a Toxic Extraterrestrial Environment. Challenges. 2024; 15(1):5. https://doi.org/10.3390/challe15010005
Chicago/Turabian StyleKeaney, Daniel, Brigid Lucey, and Karen Finn. 2024. "A Review of Environmental Challenges Facing Martian Colonisation and the Potential for Terrestrial Microbes to Transform a Toxic Extraterrestrial Environment" Challenges 15, no. 1: 5. https://doi.org/10.3390/challe15010005
APA StyleKeaney, D., Lucey, B., & Finn, K. (2024). A Review of Environmental Challenges Facing Martian Colonisation and the Potential for Terrestrial Microbes to Transform a Toxic Extraterrestrial Environment. Challenges, 15(1), 5. https://doi.org/10.3390/challe15010005





