Microbial Colonization in Marine Environments: Overview of Current Knowledge and Emerging Research Topics
Abstract
1. Introduction
- whether and to what extent the community structure of microbial biofilms changes in relation to environmental variables or substrate-related properties;
- whether there are differences in the colonization depending on the chemical nature of the artificial substrates;
- whether there is a selection of biofilm components, with preferential growth of populations that are rare members within the common microflora inhabiting the marine environment.
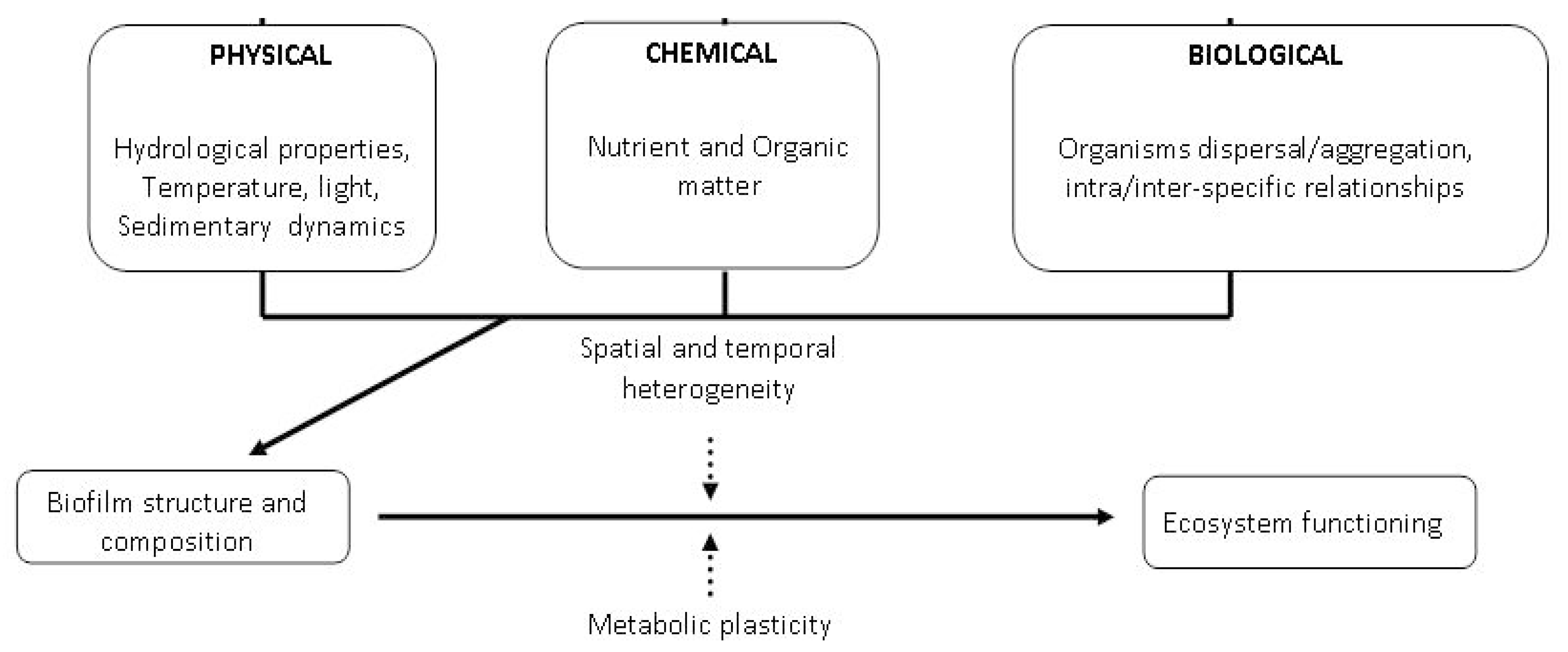
2. Drivers Potentially Affecting Microbial Colonization
3. Microbial Colonization of Several Artificial Substrates
3.1. Stainless
3.2. Glass
3.3. Ceramics
3.4. Plastic Polymers
3.4.1. Polyvinyl Chloride
3.4.2. Polyethylene and Polyethylene Terephthalate
3.4.3. Polystyrene
3.4.4. Polyurethane
3.4.5. Acrylic
3.5. Mixed Substrates
4. A Case Study: Microbial Colonization in Polar Regions
5. Significance of Microbial Biofilms in Larval Recruitment and Settlement
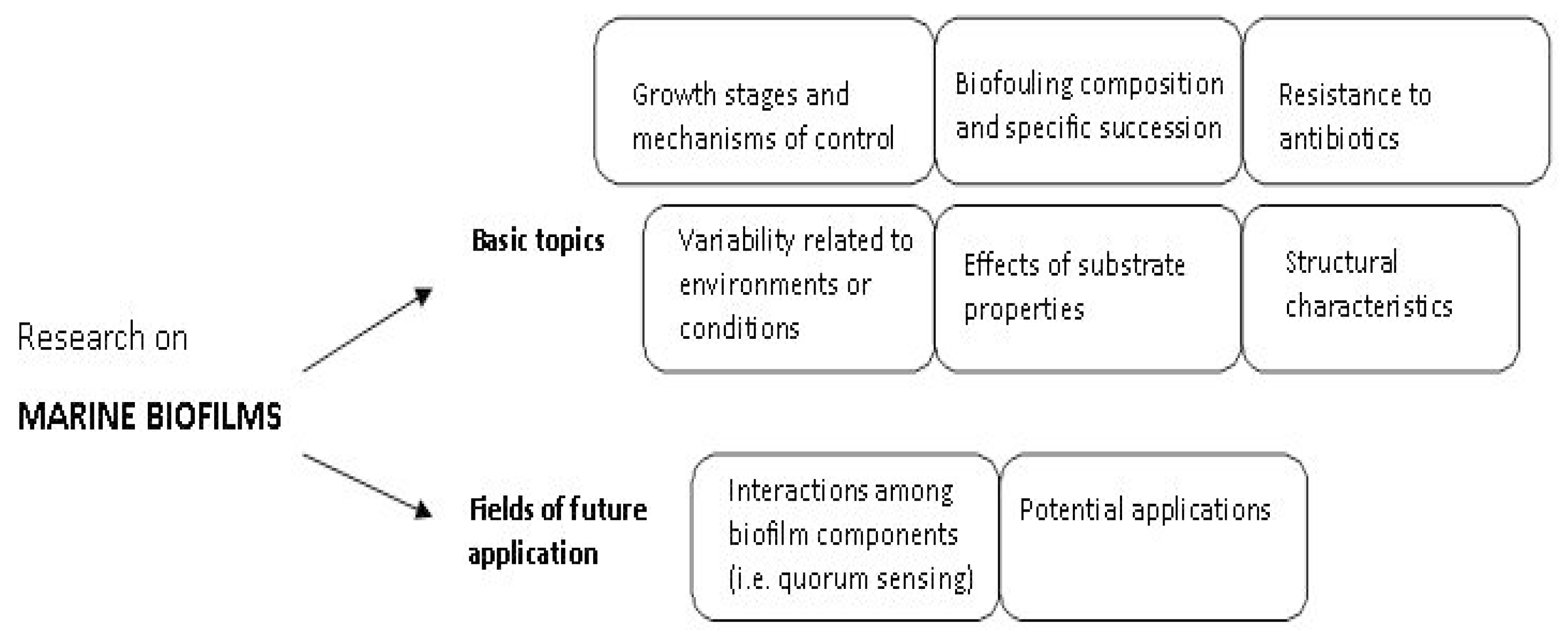
6. Future Research Directions
7. Conclusions
Funding
Acknowledgments
Conflicts of Interest
References
- Costerton, J.W.; Lewandowski, Z.; Caldwell, D.E.; Korber, D.R.; Lappin-Scott, H.M. Microbial biofilms. Annu. Rev. Microbiol. 1995, 49, 711–745. [Google Scholar] [CrossRef]
- Dang, H.; Lovell, C.R. Microbial surface colonization and biofilm development in marine environments. Microbiol. Mol. Biol. Rev. 2016, 80, 91–138. [Google Scholar] [CrossRef]
- Qian, P.-Y.; Lau, S.C.K.; Dahms, H.-U.; Dobretsov, S.; Harder, T. Marine biofilms as mediators of colonization by marine macroorganisms: Implications for antifouling and aquaculture. Mar. Biotechnol. 2007, 9, 399–410. [Google Scholar] [CrossRef]
- Dobretsov, S.; Abed, R.M.; Teplitski, M. Mini-review: Inhibition of biofouling by marine microorganisms. Biofouling 2013, 29, 423–441. [Google Scholar] [CrossRef]
- Flemming, H.C.; Wuertz, S. Bacteria and archaea on Earth and their abundance in biofilms. Nat. Rev. Microbiol. 2019, 17, 247–260. [Google Scholar] [CrossRef]
- Salta, M.; Wharton, J.A.; Blache, Y.; Stokes, K.R.; Briand, J.F. Marine biofilms on artificial surfaces: Structure and dynamics. Environ. Microbiol. 2013, 15, 2879–2893. [Google Scholar] [CrossRef] [PubMed]
- Anderson, O.R. Marine and estuarine natural microbial biofilms: Ecological and biogeochemical dimensions. AIMS Microbiol. 2018, 2, 304–331. [Google Scholar] [CrossRef]
- Battin, T.J.; Besemer, K.; Bengtsson, M.M.; Romani, A.M.; Packmann, A.I. The ecology and biogeochemistry of stream biofilms. Nat. Rev. Microbiol. 2016, 14, 251–263. [Google Scholar] [CrossRef]
- Wahl, M.; Goecke, F.; Labes, A.; Dobretsov, S.; Weinberger, F. The second skin: Ecological role of epibiotic biofilms on marine organisms. Front. Microbiol. 2012, 3, 292. [Google Scholar] [CrossRef]
- Antunes, J.; Leão, P.; Vasconcelos, V. Marine biofilms: Diversity of communities and of chemical cues. Environ. Microbiol. Rep. 2019, 11, 287–305. [Google Scholar] [CrossRef]
- Dobretsov, S.; Abed, R.M.; Voolstra, C.R. The effect of surface colour on the formation of marine micro and macrofouling communities. Biofouling 2013, 29, 617–627. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.-F.; Guo, X.-P.; Chen, Y.-R.; Ding, D.-W.; Yang, J.-L. Comparative analysis of biofilm community on different coloured substrata in relation to mussel settlement. J. Mar. Biol. Assoc. UK 2016, 97, 81–89. [Google Scholar] [CrossRef]
- Arthur, C.; Baker, J.; Bamford, H. (Eds.) NOAA Technical Memorandum NOS-OR&R-30. In Proceedings of the International Research Workshop on the Occurrence, Effects, and Fate of Microplastic Marine Debris, Tacoma, WA, USA, 9–11 September 2008; NOAA: Silver Spring, MD, USA, 2009. [Google Scholar]
- Donlan, R.M. Biofilms: Microbial life on surfaces. Emerg. Infect. Dis. 2002, 8, 881–890. [Google Scholar] [CrossRef]
- Geesey, G.G.; Gillis, R.J.; Avci, R.; Daly, D.; Hamilton, M.; Shope, P.; Harkin, G. The influence of surface features on bacterial colonization and subsequent substratum chemical changes of 316L stainless steel. Corros. Sci. 1996, 38, 73–95. [Google Scholar] [CrossRef]
- Tolker-Nielsen, T.; Molin, S. Spatial organization of microbial biofilm communities. Microb. Ecol. 2000, 40, 75–84. [Google Scholar] [CrossRef]
- Jefferson, K.K. What drives bacteria to produce a biofilm? FEMS Microbiol. Lett. 2004, 236, 163–173. [Google Scholar] [CrossRef]
- Besemer, K.; Singer, G.; Limberger, R.; Chlup, A.K.; Hochedlinger, G.; Hödl, I.; Baranyi, C.; Battin, T.J. Biophysical controls on community succession in stream biofilms. Appl. Environ. Microbiol. 2007, 73, 4966–4974. [Google Scholar] [CrossRef]
- Cowan, M.M.; Warren, T.M.; Fletcher, M. Mixed-species colonization of solid surfaces in laboratory biofilms. Biofouling 1991, 3, 23–34. [Google Scholar] [CrossRef]
- Dang, H.; Lovell, C.R. Bacterial primary colonization and early succession on surfaces in marine waters as determined by amplified rRNA gene restriction analysis and sequence analysis of 16S rRNA genes. Appl. Environ. Microbiol. 2000, 66, 467–475. [Google Scholar] [CrossRef]
- Lappin-Scott, H.M.; Costerton, J.W. Bacterial biofilms and surface fouling. Biofouling 1989, 1, 323–342. [Google Scholar] [CrossRef]
- Di Donato, P.; Poli, A.; Taurisano, V.; Abbamondi, G.R.; Nicolaus, B.; Tommonaro, G. Recent Advances in the Study of Marine Microbial Biofilm: From the Involvement of Quorum Sensing in Its Production up to Biotechnological Application of the Polysaccharide Fractions. J. Mar. Sci. Eng. 2016, 4, 34. [Google Scholar] [CrossRef]
- Stegen, J.C.; Lin, X.; Konopka, A.E.; Fredrickson, J.K. Stochastic and deterministic assembly processes in subsurface microbial communities. ISME J. 2012, 6, 1653–1664. [Google Scholar] [CrossRef] [PubMed]
- Fletcher, M. Attachment of Pseudomonas fluorescens to glass and influence of electrolytes on bacterium-substratum separation distance. J. Bacteriol. 1988, 170, 2027–2030. [Google Scholar] [CrossRef] [PubMed]
- Characklis, W.G.; McFeters, G.A.; Marshall, K.C. Physiological ecology in biofilm systems. In Biofilms; Characklis, W.G., Marshall, K.C., Eds.; John Wiley & Sons: New York, NY, USA, 1990; pp. 341–394. [Google Scholar]
- Chiu, J.M.Y.; Thiyagarajan, V.; Tsoi, M.M.Y.; Qian, P.Y. Qualitative and quantitative changes in marine biofilms as a function of temperature and salinity in summer and winter. Biofilms 2005, 2, 183–195. [Google Scholar] [CrossRef]
- Lau, S.C.K.; Thiyagarajan, V.; Cheung, S.C.K.; Qian, P.Y. Roles of bacterial community composition in biofilms as a mediator for larval settlement of three marine invertebrates. Aquat. Microb. Ecol. 2005, 38, 41–45. [Google Scholar] [CrossRef]
- Webster, N.S.; Smith, L.D.; Heyward, A.J.; Watts, J.E.M.; Webb, R.I.; Blackall, L.L.; Negri, A.P. Metamorphosis of a scleractinian coral in response to microbial biofilms. Appl. Environ. Microbiol. 2004, 70, 1213–1221. [Google Scholar] [CrossRef]
- Chiu, J.; Zhang, R.; Wang, H.; Thiyagarajan, V.; Qian, P. Nutrient effects on intertidal community: From bacteria to invertebrates. Mar. Ecol. Prog. Ser. 2008, 358, 41–50. [Google Scholar] [CrossRef][Green Version]
- Dobretsov, S.; Qian, P.-Y. Facilitation and inhibition of larval attachment of the bryozoan Bugula neritina in association with mono-species and multi-species biofilms. J. Exp. Mar. Biol. Ecol. 2006, 333, 263–274. [Google Scholar] [CrossRef]
- Babin, M.; Roesler, C.S.; Cullen, J.J. Real-time observation systems for ecosystem dynamics and harmful algal blooms: Theory, instrumentation and modelling. In Biofouling and Underwater Measurements; Lehaitre, M., Compère, C., Eds.; Oceanographic Methodology Series; UNESCO: Paris, France, 2008; pp. 463–493. [Google Scholar]
- Oberbeckmann, S.; Osborn, A.M.; Duhaime, M.B. Microbes on a bottle: Substrate, season and geography influence community composition of microbes colonizing marine plastic debris. PLoS ONE 2016, 11, e0159289. [Google Scholar] [CrossRef]
- Scheuerman, T.R.; Camper, A.K.; Hamilton, M.A. Effects of substratum topography on bacterial adhesion. J. Colloid Interf. Sci. 1998, 208, 23–33. [Google Scholar] [CrossRef]
- Crawford, R.J.; Webb, H.K.; Truong, V.K.; Hasan, J.; Ivanova, E.P. Surface topographical factors influencing bacterial attachment. Adv. Colloid Interface Sci. 2012, 179, 142–149. [Google Scholar] [CrossRef] [PubMed]
- Oh, Y.J.; Lee, N.; Jo, W.; Jung, W.K.; Lim, J. Effects of substrates on biofilm formation observed by atomic force microscopy. Ultramicroscopy 2009, 109, 874–880. [Google Scholar] [CrossRef] [PubMed]
- Mitik-Dineva, N.; Wang, J.; Mocanasu, R.C.; Stoddart, P.R.; Crawford, R.J.; Ivanova, E.P. Impact of nano-topography on bacterial attachment. Biotechnol. J. 2008, 3, 536–544. [Google Scholar] [CrossRef] [PubMed]
- Fletcher, M.; Loeb, G.I. Influence of substratum characteristics on the attachment of a marine Pseudomonad to solid surfaces. Appl. Environ. Microbiol. 1979, 37, 67–72. [Google Scholar] [CrossRef]
- von Ammon, U.; Wood, S.A.; Laroche, O.; Zaiko, A.; Tait, L.; Lavery, S.; Inglis, G.; Pochon, X. The impact of artificial surfaces on marine bacterial and eukaryotic biofouling assemblages: A high-throughput sequencing analysis. Mar. Environ. Res. 2018, 133, 57–66. [Google Scholar] [CrossRef]
- Catão, E.C.P.; Pollet, T.; Misson, B.; Garnier, C.; Ghiglione, J.-F.; Barry-Martinet, R.; Maintenay, M.; Bressy, C.; Briand, J.-F. Shear stress as a major driver of marine biofilm communities in the NW Mediterranean Sea. Front. Microbiol. 2019, 10, 1768. [Google Scholar] [CrossRef]
- Stewart, P.S.; Hamilton, M.A.; Goldstein, B.R.; Schneider, B.T. Modeling biocide action against biofilms. Biotechnol. Bioeng. 1996, 49, 445–455. [Google Scholar] [CrossRef]
- Zhang, W.; Wang, Y.; Tian, R.; Bougouffa, S.; Yang, B.; Cao, H.L.; Zhang, G.; Wong, Y.H.; Xu, W.; Batang, Z.; et al. Species sorting during biofilm assembly by artificial substrates deployed in a cold seep system. Sci. Rep. 2015, 4, 6647. [Google Scholar] [CrossRef]
- Zettler, E.R.; Mincer, T.J.; Amaral-Zettler, L.A. Life in the “Plastisphere”: Microbial Communities on Plastic Marine Debris. Environ. Sci. Technol. 2013, 47, 7137–7146. [Google Scholar] [CrossRef]
- Urbanek, A.K.; Rymowicz, W.; Mirończuk, A.M. Degradation of plastics and plastic-degrading bacteria in cold marine habitats. Appl. Microbiol. Biotechnol. 2018, 102, 7669–7678. [Google Scholar] [CrossRef]
- Sudhakar, M.; Priyadarshini, C.; Doble, M.; Murthy, S.; Venkatesan, R. Marine bacteria mediated degradation of nylon 66 and 6. Int. Biodeterior. Biodegrad. 2007, 60, 144–151. [Google Scholar] [CrossRef]
- Terlizzi, A.; Faimali, M. Fouling on artificial substrata. In Biofouling; Dürr, S., Thomason, J.C., Eds.; Blackwell Publishing Ltd.: Oxford, UK, 2010; pp. 170–179. [Google Scholar]
- Cooksey, K.E.; Wigglesworth-Cooksey, B. Adhesion of bacteria and diatoms to surfaces in the sea: A review. Aquat. Microb. Ecol. 1995, 9, 87–96. [Google Scholar] [CrossRef]
- Singh, R.; Paul, D.; Jain, R.K. Biofilms: Implications in bioremediation. Trends Microbiol. 2006, 14, 389–397. [Google Scholar] [CrossRef] [PubMed]
- Donlan, R.M. Biofilms and device-associated infections. Emerg. Infect. Dis. 2001, 7, 277–281. [Google Scholar] [CrossRef] [PubMed]
- Huggett, M.J.; Nedved, B.T.; Hadfield, M.G. Effects of initial surface wettability on biofilm formation and subsequent settlement of Hydroides elegans. Biofouling 2009, 25, 387–399. [Google Scholar] [CrossRef] [PubMed]
- Chung, H.; Lee, O.; Huang, Y.; Mok, S.Y.; Kolter, R.; Qian, P.Y. Bacterial community succession and chemical profiles of subtidal biofilms in relation to larval settlement of the polychaete Hydroides elegans. ISME J. 2010, 4, 817–828. [Google Scholar] [CrossRef]
- Witt, V.; Wild, C.; Anthony, K.R.; Diaz-Pulido, G.; Uthicke, S. Effects of ocean acidification on microbial community composition of, and oxygen fluxes through, biofilms from the Great Barrier Reef. Environ. Microbiol. 2011, 13, 2976–2989. [Google Scholar] [CrossRef]
- Bellou, N.; Papathanassiou, E.; Dobretsov, S.; Lykousis, V.; Colijn, F. The effect of substratum type, orientation and depth on the development of bacterial deep-sea biofilm communities grown on artificial substrata deployed in the Eastern Mediterranean. Biofouling 2012, 28, 199–213. [Google Scholar] [CrossRef]
- Khandekar, N.; Johns, R.B. Marine corrosion studies-II. A biomarker study tracing the early formation of biofilms on steel plates in a marine environment. Org. Geochem. 1990, 15, 531–538. [Google Scholar] [CrossRef]
- Jones, P.R.; Cottrell, M.T.; Kirchman, D.L.; Dexter, S.C. Bacterial community structure of biofilms on artificial surfaces in an estuary. Microb. Ecol. 2007, 53, 153–162. [Google Scholar] [CrossRef]
- Mitik-Dineva, N.; Wang, J.; Truong, V.K.; Stoddart, P.R.; Malherbe, F.; Crawford, R.J.; Ivanova, E.P. Differences in colonisation of five marine bacteria on two types of glass surfaces. Biofouling 2009, 25, 621–631. [Google Scholar] [CrossRef]
- Landoulsi, J.; Cooksey, K.E.; Dupres, V. Review—Interactions between diatoms and stainless steel: Focus on biofouling and biocorrosion. Biofouling 2011, 27, 1105–1124. [Google Scholar] [CrossRef]
- Li, X.; Duan, J.; Xiao, H.; Li, Y.; Liu, H.; Guan, F.; Zhai, X. Analysis of Bacterial Community Composition of Corroded Steel Immersed in Sanya and Xiamen Seawaters in China via Method of Illumina MiSeq Sequencing. Front. Microbiol. 2017, 8, 1737. [Google Scholar] [CrossRef]
- Head, R.; Davenport, J.; Thomason, J.C. The effect of depth on the accrual of marine biofilms on glass substrata deployed in the Clyde Sea, Scotland. Biofouling 2004, 20, 177–180. [Google Scholar] [CrossRef]
- Patil, J.S.; Chandrashekar Anil, A. Biofilm diatom community structure: Influence of temporal and substratum variability. Biofouling 2005, 21, 189–206. [Google Scholar] [CrossRef]
- Mitbavkar, S.; Raghu, C.; Rajaneesh, K.M.; Pavan, D. Picophytoplankton community from tropical marine biofilms. J. Exp. Mar. Biol. Ecol. 2012, 426, 88–96. [Google Scholar] [CrossRef]
- Siboni, N.; Lidor, M.; Kramarsky-Winter, E.; Kushmaro, A. Conditioning film and initial biofilm formation on ceramics tiles in the marine environment. FEMS Microbiol. Lett. 2007, 274, 24–29. [Google Scholar] [CrossRef]
- Kang, J.; Kang, S.; Kim, Y.-T. Study on the control of marine biofouling developed on the surface of porous ceramics. J. Korean Cryst. Growth Cryst. Technol. 2015, 25, 218–224. [Google Scholar] [CrossRef]
- Harrison, H.P.; Sapp, M.; Schratzberger, M.; Osborn, A.M. Interactions between microorganisms and marine microplastics: A call for research. Mar. Technol. Soc. J. 2011, 45, 12–20. [Google Scholar] [CrossRef]
- Auta, H.S.; Emenike, C.U.; Fauziah, S.H. Distribution and importance of microplastics in the marine environment: A review of the sources, fate, effects, and potential solutions. Environ. Int. 2017, 102, 165–176. [Google Scholar] [CrossRef]
- Thompson, R.C.; Olsen, Y.; Mitchell, R.P.; Davis, A.; Rowland, S.J.; John, A.W.; McGonigle, D.; Russell, A.E. Lost at sea: Where is all the plastic? Science 2004, 304, 838. [Google Scholar] [CrossRef]
- Moore, C.J.; Moore, S.L.; Weisberg, S.B.; Lattin, G.L.; Zellers, A.F. A comparison of neustonic plastic and zooplankton abundance in southern California’s coastal waters. Mar. Pollut. Bull. 2002, 44, 1035–1038. [Google Scholar] [CrossRef]
- Lattin, G.L.; Moore, C.J.; Zellers, A.F.; Moore, S.L.; Weisberg, S.B. A comparison of neustonic plastic and zooplankton at different depths near the southern California shore. Mar. Pollut. Bull. 2004, 49, 291–294. [Google Scholar] [CrossRef]
- Cole, M.; Lindeque, P.; Halsband, C.; Galloway, T.S. Microplastics as contaminants in the marine environment: A review. Mar. Pollut. Bull. 2011, 62, 2588–2597. [Google Scholar] [CrossRef]
- Ballent, A.; Pando, S.; Purser, A.; Juliano, M.F.; Thomsen, L. Modelled transport of benthic marine microplastic pollution in the Nazaré Canyon. Biogeosciences 2013, 10, 7957–7970. [Google Scholar] [CrossRef]
- Browne, M.A.; Galloway, T.; Thompson, R. Microplastic—An emerging contaminant of potential concern? Integr. Environ. Assess. Manag. 2007, 3, 559–566. [Google Scholar] [CrossRef]
- Andrady, A.L. Microplastics in the marine environment. Mar. Pollut. Bull. 2011, 62, 1596–1605. [Google Scholar] [CrossRef]
- Lobelle, D.; Cunliffe, M. Early microbial biofilm formation on marine plastic debris. Mar. Pollut. Bull. 2011, 62, 197–200. [Google Scholar] [CrossRef]
- Corcoran, P.L. Benthic plastic debris in marine and fresh water environments. Environ. Sci. Process. Impacts 2015, 8, 1363–1369. [Google Scholar] [CrossRef]
- Corcoran, P.L.; Biesinger, M.C.; Grifi, M. Plastics and beaches: A degrading relationship. Mar. Pollut. Bull. 2009, 58, 80–84. [Google Scholar] [CrossRef]
- Oberbeckmann, S.; Loeder, M.G.; Gerdts, G.; Osborn, A.M. Spatial and seasonal variation in diversity and structure of microbial biofilms on marine plastics in Northern European waters. FEMS Microbiol. Ecol. 2014, 90, 478–492. [Google Scholar] [CrossRef] [PubMed]
- Cai, L.; Wu, D.; Xia, J.; Shi, H.; Kim, H. Influence of physicochemical surface properties on the adhesion of bacteria onto four types of plastics. Sci. Total Environ. 2019, 671, 1101–1107. [Google Scholar] [CrossRef]
- Amaral-Zettler, L.A.; Zettler, E.R.; Slikas, B.; Boyd, G.D.; Melvin, D.W.; Morrall, C.E.; Proskurowski, G.; Mincer, T.J. Biogeography of the Plastisphere: Implications for Policy. Front. Ecol. Environ. 2015, 13, 541–546. [Google Scholar] [CrossRef]
- Oberbeckmann, S.; Kreikemeyer, B.; Labrenz, M. Environmental factors support the formation of specific bacterial assemblages on microplastics. Front. Microbiol. 2018, 8, 2709. [Google Scholar] [CrossRef] [PubMed]
- Briand, J.-F.; Djeridi, I.; Jamet, D.; Coupé, S.; Bressy, C.; Molmeret, M.; Le Berre, B.; Rimet, F.; Bouchez, A.; Blache, Y. Pioneer marine biofilms on artificial surfaces including antifouling coatings immersed in two contrasting French Mediterranean coast sites. Biofouling 2012, 28, 453–463. [Google Scholar] [CrossRef] [PubMed]
- Carson, H.S.; Nerheim, M.S.; Carroll, K.A.; Eriksen, M. The plastic-associated microorganisms of the North Pacific Gyre. Mar. Pollut. Bull. 2013, 75, 126–132. [Google Scholar] [CrossRef]
- Reisser, J.; Shaw, J.; Hallegraeff, G.; Proietti, M.; Barnes, D.K.; Thums, M.; Wilcox, C.; Hardesty, B.D.; Pattiaratchi, C. Millimeter-sized marine plastics: A new pelagic habitat for microorganisms and invertebrates. PLoS ONE 2014, 9, e100289. [Google Scholar] [CrossRef] [PubMed]
- Khandeparker, L.; D’Costa, P.M.; Anil, A.C.; Sawant, S.S. Interactions of bacteria with diatoms: Influence on natural marine biofilms. Mar. Ecol. 2014, 35, 233–248. [Google Scholar] [CrossRef]
- Amin, S.A.; Parker, M.S.; Armbrust, E.V. Interactions between diatoms and bacteria. Microbiol. Mol. Biol. Rev. 2012, 76, 667–684. [Google Scholar] [CrossRef] [PubMed]
- Abell, G.C.; Bowman, J.P. Colonization and community dynamics of class Flavobacteria on diatom detritus in experimental mesocosms based on southern ocean seawater. FEMS Microbiol. Ecol. 2005, 53, 379–391. [Google Scholar] [CrossRef]
- Jacquin, J.; Cheng, J.; Odobel, C.; Pandin, C.; Conan, P.; Pujo-Pay, M.; Barbe, V.; Meistertzheim, A.L.; Ghiglione, J.F. Microbial Ecotoxicology of Marine Plastic Debris: A Review on Colonization and Biodegradation by the “Plastisphere”. Front. Microbiol. 2019, 10, 865. [Google Scholar] [CrossRef] [PubMed]
- Bryant, J.A.; Clemente, T.M.; Viviani, D.A.; Fong, A.A.; Thomas, K.A.; Kemp, P.; Karl, D.M.; White, A.E.; DeLong, E.F. Diversity and activity of communities inhabiting plastic debris in the North Pacific Gyre. mSystems 2016, 1, e00024-16. [Google Scholar] [CrossRef] [PubMed]
- De Tender, C.; Devriese, L.I.; Haegeman, A.; Maes, S.; Vangeyte, J.; Cattrijsse, A.; Dawyndt, P.; Ruttink, T. Temporal dynamics of bacterial and fungal colonization on plastic debris in the North Sea. Environ. Sci. Technol. 2017, 51, 7350–7360. [Google Scholar] [CrossRef] [PubMed]
- Dussud, C.; Meistertzheim, A.L.; Conan, P.; Pujo-Pay, M.; George, M.; Fabre, P.; Coudane, J.; Higgs, P.; Elineau, A.; Pedrotti, M.L.; et al. Evidence of niche partitioning among bacteria living on plastics, organic particles and surrounding seawaters. Environ. Pollut. 2018, 236, 807–816. [Google Scholar] [CrossRef]
- Delacuvellerie, A.; Cyriaque, V.; Gobert, S.; Benali, S.; Wattiez, R. The plastisphere in marine ecosystem hosts potential specific microbial degraders including Alcanivorax borkumensis as a key player for the low-density polyethylene degradation. J. Hazard. Mater. 2019, 380, 120899. [Google Scholar] [CrossRef]
- Hung, O.S.; Thiyagarajan, V.; Zhang, R.; Wu, R.S.S.; Qian, P.Y. Attachment of Balanus amphitrite larvae to biofilms originating from contrasting environments. Mar. Ecol. Prog. Ser. 2007, 333, 229–242. [Google Scholar] [CrossRef][Green Version]
- Balasubramanian, V.; Palanichamy, S.; Subramanian, G.; Rajaram, R. Development of polyvinyl chloride biofilms for succession of selected marine bacterial populations. J. Environ. Biol. 2012, 33, 57–60. [Google Scholar]
- Briand, J.-F.; Barani, A.; Garnieri, C.; Réhel, K.; Urvois, F.; LePoupon, C.; Bouchez, A.; Debroas, D.; Bressy, C. Spatio-temporal variations of marine biofilm communities colonizing artificial substrata including antifouling coatings in contrasted French coastal environments. Microb. Ecol. 2017, 74, 585–598. [Google Scholar] [CrossRef]
- Misic, C.; Harriague, A.C. Development of marine biofilm on plastic: Ecological features in different seasons, temperatures, and light regimes. Hydrobiologia 2019, 835, 129–145. [Google Scholar] [CrossRef]
- O’Connor, N.J.; Richardson, D.L. Attachment of barnacle (Balanus amphitrite Darwin) larvae: Responses to bacterial films and extracellular materials. J. Exp. Mar. Biol. Ecol. 1998, 226, 115–129. [Google Scholar] [CrossRef]
- Leroy, C.; Delbarre-Ladrat, C.; Ghillebaert, F.; Rochet, M.J.; Compère, C.; Combes, D. A marine bacterial adhesion microplate test using the DAPI fluorescent dye: A new method to screen antifouling agents. Lett. Appl. Microbiol. 2007, 44, 372–378. [Google Scholar] [CrossRef] [PubMed]
- Xu, H.; Min, G.-S.; Choi, J.-K.; Jung, J.-H.; Park, M.-H. An approach to analyses of periphytic ciliate colonization for monitoring water quality using a modified artificial substrate in Korean coastal waters. Mar. Pollut. Bull. 2009, 58, 1278–1285. [Google Scholar] [CrossRef] [PubMed]
- Jeong, S.; Kim, J.-A.; Kim, H.; Chung, K.-S.; Yoon, H.-O. Identification of preponderant marine bacteria and their biofouling characteristics on adsorbents of different sizes and shapes in seawater. J. Mar. Sci. Technol. 2018, 26, 458–464. [Google Scholar] [CrossRef]
- Lee, Y.K.; Kwon, K.K.; Cho, K.H.; Park, J.H.; Lee, H.K. Isolation and identification of Bacteria from marine biofilms. Key Eng. Mater. 2005, 277, 612–617. [Google Scholar] [CrossRef]
- Mejdandžić, M.; Ivanković, T.; Pfannkuchen, M.; Godrijan, J.M.; Pfannkuchen, D.M.; Hrenović, J.; Ljubešić, Z. Colonization of diatoms and bacteria on artificial substrates in the northeastern coastal Adriatic Sea. Acta Bot. Croat. 2015, 74, 407–422. [Google Scholar] [CrossRef]
- Guezennec, J.; Ortega-Morales, O.; Raguenes, G.; Geesey, G. Bacterial colonization of artificial substrate in the vicinity of deep-sea hydrothermal vents. FEMS Microbiol. Ecol. 1998, 26, 89–99. [Google Scholar] [CrossRef]
- Artham, T.; Sudhakar, M.; Venkatesan, R.; Madhavan Nair, C.; Murty, K.V.G.K.; Doble, M. Biofouling and stability of synthetic polymers in sea water. Int. Biodeterior. Biodegrad. 2009, 63, 884–890. [Google Scholar] [CrossRef]
- Witt, V.; Wild, C.; Uthicke, S. Effect of substrate type on bacterial community composition in biofilms from the Great Barrier Reef. FEMS Microbiol. Lett. 2011, 323, 188–195. [Google Scholar] [CrossRef]
- Hoellein, T.; Rojas, M.; Pink, A.; Gasior, J.; Kelly, J. Anthropogenic litter in urban freshwater ecosystems: Distribution and microbial interactions. PLoS ONE 2014, 9, e98485. [Google Scholar] [CrossRef]
- Frère, L.; Paul-Pont, I.; Rinnert, E.; Petton, S.; Jaffré, J.; Bihannic, I.; Soudant, P.; Lambert, C.; Huvet, A. Influence of environmental and anthropogenic factors on the composition, concentration and spatial distribution of microplastics: A case study of the Bay of Brest (Brittany, France). Environ. Pollut. 2017, 225, 211–222. [Google Scholar] [CrossRef]
- Dussud, C.; Hudec, C.; George, M.; Fabre, P.; Higgs, P.; Bruzaud, S.; Delort, A.-M.; Eyheraguibel, B.; Meistertzheim, A.-L.; Jacquin, J.; et al. Colonization of Non-biodegradable and Biodegradable Plastics by Marine Microorganisms. Front. Microbiol. 2018, 9, 1571. [Google Scholar] [CrossRef] [PubMed]
- Kirstein, I.V.; Wichels, A.; Krohne, G.; Gerdts, G. Mature biofilm communities on synthetic polymers in seawater—Specific or general? Mar. Environ. Res. 2018, 142, 147–154. [Google Scholar] [CrossRef] [PubMed]
- Ogonowski, M.; Motiei, A.; Ininbergs, K.; Hell, E.; Gerdes, Z.; Udekwu, K.I.; Bacsik, Z.; Gorokhova, E. Evidence for selective bacterial community structuring on microplastics. Environ. Microbiol. 2018, 20, 2796–2808. [Google Scholar] [CrossRef] [PubMed]
- Muthukrishnan, T.; Al Khaburi, M.; Abed, R.M.M. Fouling Microbial Communities on Plastics Compared with Wood and Steel: Are They Substrate- or Location-Specific? Microb. Ecol. 2019, 78, 361–374. [Google Scholar] [CrossRef]
- Pinto, M.; Langer, T.M.; Hüffer, T.; Hofmann, T.; Herndl, G.J. The composition of bacterial communities associated with plastic biofilms differs between different polymers and stages of biofilm succession. PLoS ONE 2019, 14, e0217165. [Google Scholar] [CrossRef]
- Scotto, V.; Alabiso, G.; Marcenaro, G. An example of microbiologically influenced corrosion: The behaviour of stainless steels in natural seawater. Bioelectrochem. Bioenerg. 1986, 16, 347–355. [Google Scholar] [CrossRef]
- Alabiso, G.; Montini, U.; Milillo, M. One year corrosion test of four stainless steel grades at Ny-Alesund (Svalbard Islands): June 2004–June 2005. In CNR-IAMC Technical Report; National Research Council: Taranto, Italy, 2005. [Google Scholar]
- Maki, J.S.; Little, B.J.; Wagner, P.; Mitchell, R. Biofilm formation on metal surfaces in Antarctic waters. Biofouling 1990, 2, 27–38. [Google Scholar] [CrossRef]
- Clark, M.S.; Villota Nieva, L.; Hoffman, J.I.; Davies, A.J.; Trivedi, U.J.; Turner, F.; Ashton, G.V.; Peck, L.S. Lack of long-term acclimation in Antarctic encrusting species suggests vulnerability to warming. Nat. Commun. 2019, 10, 3383. [Google Scholar] [CrossRef]
- Caruso, G.; Azzaro, M.; Dell’Acqua, O.; Lo Giudice, A.; Fazi, S.; Caroppo, C.; Azzaro, F.; La Ferla, R.; Maimone, G.; Laganà, P.; et al. Microbial response to anthropic and natural forcings in two coastal Antarctic sites (Ross Sea). Biol. Mar. Medit. 2019, in press. [Google Scholar]
- Loeb, G.I.; Neihof, R.A. Marine conditioning films. Adv. Chem. 1975, 145, 319–335. [Google Scholar] [CrossRef]
- Bowman, J.P. Bioactive compound synthetic capacity and ecological significance of marine bacterial genus Pseudoalteromonas. Mar. Drugs 2007, 5, 220–241. [Google Scholar] [CrossRef] [PubMed]
- Donlan, R.M.; Costerton, J.W. Biofilms: Survival mechanisms of clinically relevant microorganisms. Clin. Microbiol. Rev. 2002, 15, 167–193. [Google Scholar] [CrossRef] [PubMed]
- Qian, P.-Y.; Rittschof, D.; Sreedhar, B.; Chia, F.S. Macrofouling in unidirectional flow: Miniature pipes as experimental models for studying the effects of hydrodynamics on invertebrate larval settlement. Mar. Ecol. Prog. Ser. 1999, 191, 141–151. [Google Scholar] [CrossRef]
- Qian, P.-Y.; Rittschof, D.; Sreedhar, B. Macrofouling in unidirectional flow: Miniature pipes as experimental models for studying interaction of flow and surface characteristics on the attachment of barnacle, bryozoan and polychaete larvae. Mar. Ecol. Prog. Ser. 2000, 207, 109–121. [Google Scholar] [CrossRef][Green Version]
- Maki, J.S.; Rittschof, D.; Costlow, J.D.; Mitchell, R. Inhibition of attachment of larval barnacles, Balanus amphitrite, by bacterial surface films. Mar. Biol. 1988, 97, 199–206. [Google Scholar] [CrossRef]
- Wieczorek, S.; Todd, C. Inhibition and facilitation of bryozoan and ascidian settlement by natural multi-species biofilms: Effects of film age and the roles of active and passive larval attachment. Mar. Biol. 1997, 128, 463–473. [Google Scholar] [CrossRef]
- Qian, P.-Y.; Thiyagarajan, V.; Lau, S.C.K.; Cheung, S.C.K. Relationship between bacterial community profile in biofilm and attachment of the acorn barnacle Balanus amphitrite. Aquat. Microb. Ecol. 2003, 33, 225–237. [Google Scholar] [CrossRef]
- Chiu, J.M.Y.; Thiyagarajan, V.; Pechenik, J.A.; Hung, O.S.; Qian, P.Y. Influence of bacteria and diatoms in biofilms on metamorphosis of the marine slipper limpet Crepidula onyx. Mar. Biol. 2007, 151, 1417–1431. [Google Scholar] [CrossRef]
- Huang, J.; Shi, Y.; Zeng, G.; Gu, Y.; Chen, G.; Shi, L.; Hu, Y.; Tang, B.; Zhou, J. Acyl-homoserine lactone- based quorum sensing and quorum quenching hold promise to determine the performance of biological wastewater treatments: An overview. Chemosphere 2016, 157, 137–151. [Google Scholar] [CrossRef]
- Dobretsov, S.; Dahms, H.-U.; YiLi, H.; Wahl, M.; Qian, P.-Y. The effect of quorum-sensing blockers on the formation of marine microbial communities and larval attachment. FEMS Microbiol. Ecol. 2007, 60, 177–188. [Google Scholar] [CrossRef]
- Landini, P.; Antoniani, D.; Burgess, J.G.; Nijland, R. Molecular mechanisms of compounds affecting bacterial biofilm formation and dispersal. Appl. Microbiol. Biotechnol. 2010, 86, 813–823. [Google Scholar] [CrossRef]
- de Carvalho, C.C.C.R. Marine biofilms: A successful microbial strategy with economic implications. Front. Mar. Sci. 2018, 5, 126. [Google Scholar] [CrossRef]
- Dobretsov, S.; Coutinho, R.; Rittschof, D.; Salta, M.; Ragazzola, F.; Hellio, C. The oceans are changing: Impact of ocean warming and acidification on biofouling communities. Biofouling 2019, 35, 585–595. [Google Scholar] [CrossRef] [PubMed]
- Zhang, W.; Ding, W.; Li, Y.-X.; Tam, C.; Bougouffa, S.; Wang, R.; Pei, B.; Chiang, H.; Leung, P.; Lu, Y.; et al. Marine biofilms constitute a bank of hidden microbial diversity and functional potential. Nat. Commun. 2019, 10, 517. [Google Scholar] [CrossRef] [PubMed]
- Battin, T.; Kaplan, L.; Denis Newbold, J.; Hansen, C.M.E. Contributions of microbial biofilms to ecosystem processes in stream mesocosms. Nature 2003, 426, 439–442. [Google Scholar] [CrossRef] [PubMed]
- Wilmes, P.; Bond, P.L. Microbial community proteomics: Elucidating the catalysts and metabolic mechanisms that drive the Earth’s biogeochemical cycles. Curr. Opin. Microbiol. 2009, 12, 310–317. [Google Scholar] [CrossRef]
- Moss, J.A.; Nocker, A.; Lepo, J.E.; Snyder, R.A. Stability and change in estuarine biofilm bacterial community diversity. Appl. Environ. Microbiol. 2006, 72, 5679–5688. [Google Scholar] [CrossRef]
- Webster, N.S. and Negri, A.P. Site-specific variation in Antarctic marine biofilms established on artificial surfaces. Environ. Microbiol. 2006, 8, 1177–1190. [Google Scholar] [CrossRef]
- Dang, H.; Li, T.; Chen, M.; Huang, G. Cross-ocean distribution of Rhodobacterales bacteria as primary surface colonizers in temperate coastal marine waters. Appl. Environ. Microbiol. 2008, 74, 52–60. [Google Scholar] [CrossRef]
- Kriwy, P.; Uthicke, S. Microbial diversity in marine biofilms along a water quality gradient on the Great Barrier Reef. Syst. Appl. Microbiol. 2011, 34, 116–126. [Google Scholar] [CrossRef]
- Huang, Y.; Zeng, Y.; Yu, Z.; Zhang, J.; Feng, H.; Lin, X. In silico and experimental methods revealed highly diverse bacteria with quorum sensing and aromatics biodegradation systems—A potential broad application on bioremediation. Bioresour. Technol. 2013, 148, 311–316. [Google Scholar] [CrossRef] [PubMed]
| Biofilm Formation and Development | Biofilm Components | Biofilm Control |
|---|---|---|
| Extracellular polymeric matrix composition | Heterogeneous microhabitats within the biofilms’ structure | Processes related to the synthesis of biofilm matrix |
| Variables and mechanisms that favour and stabilize the formation of biofilm | Intercellular communication/signalling (i.e., quorum sensing) | -Novel and environmentally friendly methods to prevent biofilm formation and/or to remove biofouling -Interactions (inhibition/induction) of microbial biofilm members responsible for antifouling resistance |
| -Application of specific enzymes (i.e., proteases and oxidases); quorum sensing inhibitors; use of bacteriophages | ||
| Biofilm structural and functional diversity | -Nutrient acquisition and recycling by biofilm components -Antibiotic resistance and virulence factors of biofilm components | Mechanisms involved in biofilm tolerance to biocides and other stressors |
| Habitats/substrates supporting biofilm formation | Responses of microbial consortia to surface adhesion |
© 2020 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Caruso, G. Microbial Colonization in Marine Environments: Overview of Current Knowledge and Emerging Research Topics. J. Mar. Sci. Eng. 2020, 8, 78. https://doi.org/10.3390/jmse8020078
Caruso G. Microbial Colonization in Marine Environments: Overview of Current Knowledge and Emerging Research Topics. Journal of Marine Science and Engineering. 2020; 8(2):78. https://doi.org/10.3390/jmse8020078
Chicago/Turabian StyleCaruso, Gabriella. 2020. "Microbial Colonization in Marine Environments: Overview of Current Knowledge and Emerging Research Topics" Journal of Marine Science and Engineering 8, no. 2: 78. https://doi.org/10.3390/jmse8020078
APA StyleCaruso, G. (2020). Microbial Colonization in Marine Environments: Overview of Current Knowledge and Emerging Research Topics. Journal of Marine Science and Engineering, 8(2), 78. https://doi.org/10.3390/jmse8020078






