Phylogenetic Diversity of Diazotrophs along an Experimental Nutrient Gradient in Mangrove Sediments
Abstract
:1. Introduction
2. Material and Methods
2.1. Study Site
2.2. Collection of Environmental Samples
2.3. Distribution of Mangrove Roots
2.4. DNA Extraction
2.5. Nested PCR Amplification
2.6. Terminal Restriction Fragment Length Polymorphism Analysis (TRFLP)
2.7. DNA Cloning and Sequencing
2.8. Statistical Comparisons
2.9. Nucleotide Sequence Accession Numbers
| Clones | Closest match in GenBank | |||||
|---|---|---|---|---|---|---|
| Presumable phylogenetic association | Phylotypes ID | Total | Match | Accession number | % Similarity | Reference |
| Delta-Proteobacteria | N26 | 1 | Isololate from Italian soils | AAS47809 | 87 | |
| N31, P211 | 6 | Paleobacter carbinolicus DSM 2380 | YP357508 | 96 | Mussmann et al. 2005 [60] | |
| P12, P33 | 2 | Geobacter sulfurreducens | NP953865 | 93 | Methe et al. 2003 [61] | |
| P11, C13, C11, P13, N19 | 11 | Microbial mats in the Bahamas | AAZ77144 | 93–97 | Yannarell et al. 2006 [62] | |
| P34, N21, P210 | 3 | Forest soils in Amazon region | AC132206 | 90–92 | ||
| C33 | 1 | Microbial mat in North Carolina | AAA65425 | 88 | Zehr at al. 1995 [63] | |
| C32, P21 | 3 | Microbial mat, dead stems of Spartina alterniflora | AAY85458, AAS57673 | 90–93 | Musat et al. 2006 [64] Moisander et al. 2005 [65] | |
| C14 | 1 | Desulfovibrio vulgaris | YP2437020 | 90 | Heidelberg et al. 2004 [66] | |
| N17 | 2 | Microbial mat in North Carolina | AAA65429 | 99 | Zehr at al. 1995 [63] | |
| P22, N351 | 3 | Desulfomicrobium baculatum | ZP04344086 | 92 | ||
| N35 | 2 | Mangrove sediments in China | ABM66820 | 96 | Zhang et al. 2008 [13] | |
| P15 | 1 | Chesapeake Bay | AAZ06740 | 96 | ||
| C22 | 1 | Microbial mat | AAY85430 | 94 | Musat et al. 2006 [64] | |
| P31 | 1 | Mangrove sediments in China | ABM67091 | 90 | Zhang et al. 2008 [13] | |
| P25 | 8 | Mangrove sediments in China | AAF61027 | 92 | Zhang et al. 2008 [13] | |
| N18 | 8 | Seagrass sediments in the Bahamas | AAL07952 | 89 | Bagwell et al. 2002 [67] | |
| N36, N33, N314, N37, N313 | 8 | Seagrass sediments in the Bahamas | AAL07952 | 90–96 | Bagwell et al. 2002 [67] | |
| P26, C36 | 3 | Microbial mat in North Carolina | AAA65422 | 97 | Zehr et al. 1995 [63] | |
| N213, P27, N32 | 6 | Eastern Mediterranean Sea | ABQ50824 | 93 | ||
| C23, N312, P23, N112,N25, C34 | 20 | Desulfatibacillum alkenivorans AK-01 | YP2430688 | 90–97 | ||
| N211, N13, C31, N310, N27 | 8 | Microbial mat, mangrove sediments | AAY85423, ABM74058 | 90–96 | Musat et al. 2006 [64] | |
| N22, C21, C35, N39, N111 | 6 | Eastern Mediterranean Sea | ABQ50612 | 93–95 | ||
| N23, N14, N110 | 18 | Microbial mat, Eastern Mediterranean Sea | AAY85423, ABQ50807 | 94–95 | Musat et al. 2006 [64] | |
| N38, N24 | 10 | ABQ50691 | 79 | |||
| Gamma-Proteobacteria | C12, P24 | 2 | Vibrio natriegens and Klebsiella pneumoniae | AAD55588, AAO85881 | 94 | |
| N34, N29 | 7 | Pristine marine environment | AAY85422 | 99 | Musat et al. 2006 [64] | |
| N16 | 1 | Thiocapsa roseopersicina | ACC95826 | 93 | ||
| Firmicutes | P14 | 1 | Malaysian soil | ACC95201 | 94 | |
| RP32 | 2 | Forest soils in Amazon region | ACI32162 | 93 | ||
| N311 | 2 | Terrestrial soil | ACI26001 | 91 | ||
| Green sulfur bacteria | N12, P28, C15, N212 | 6 | Rhizosphere of Spartina alterniflora | ABD74331 | 96–97 | Lovell et al. 2008 [18] |
| N28, P29 | 2 | Chlorobium phaeobacteroides | YP1960150 | 98 |
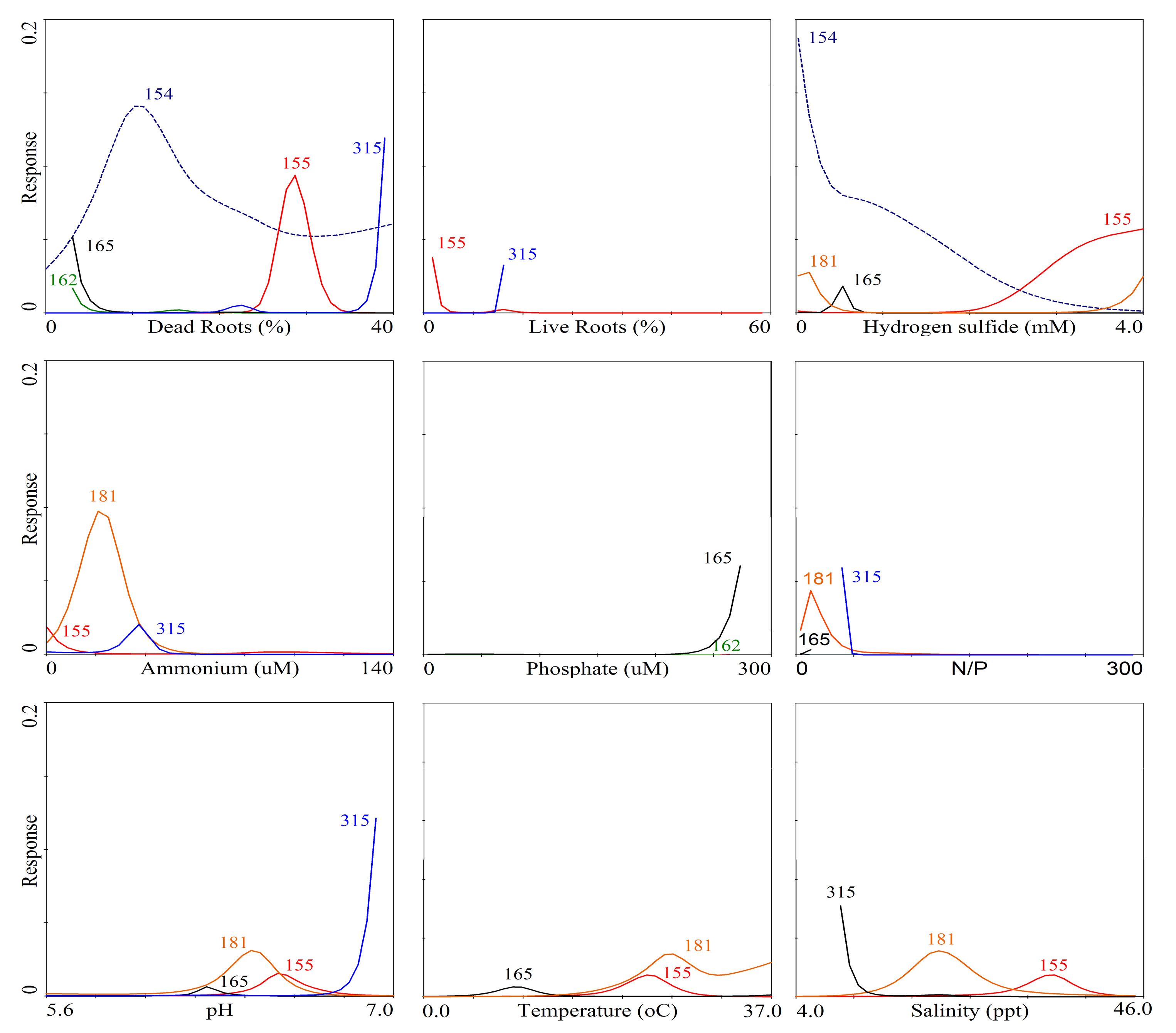
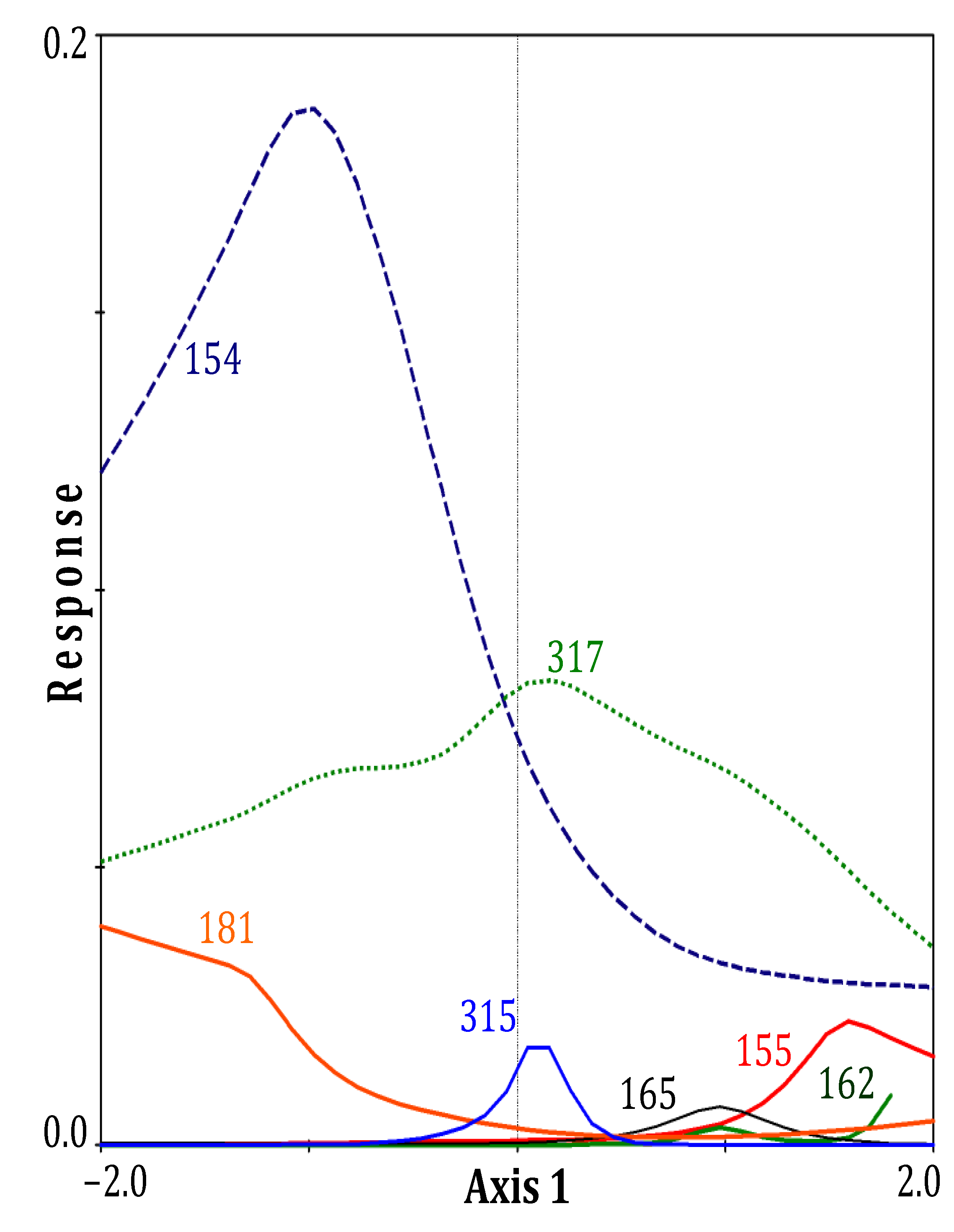
3. Results
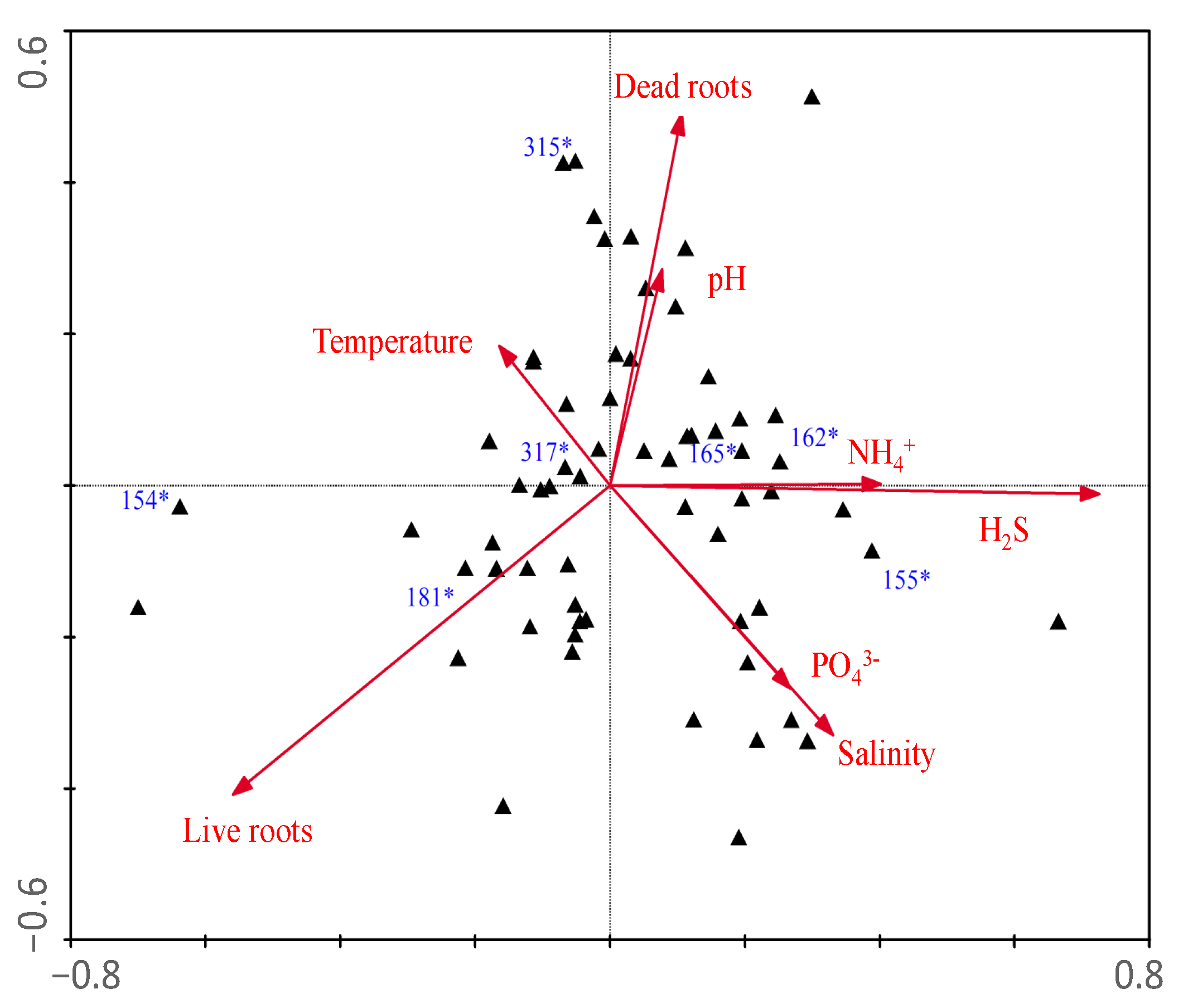
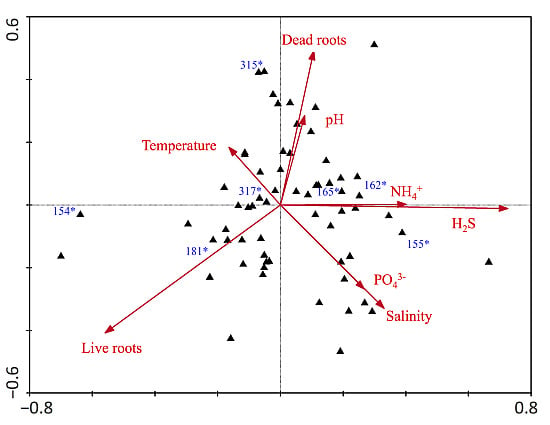
| Axes | ||
|---|---|---|
| Parameters | 1 | 2 |
| Eigenvalues | 0.135 | 0.120 |
| Species-environment Correlations | 0.881 | 0.848 |
| Correlation coefficients | ||
| Dead roots | 0.51 | −0.23 |
| Live roots | −0.47 | 0.02 |
| Temperature | 0.56 | −0.46 |
| pH | 0.56 | −0.48 |
| Salinity | 0.54 | −0.52 |
| NH4+ | 0.44 | −0.44 |
| H2S | 0.50 | −0.40 |
| PO43− | 0.07 | −0.36 |
| N:P (molar) | 0.18 | −0.16 |
| TRFLP | Phylotypes | |||
|---|---|---|---|---|
| OTU # | Total OTUs | ID | Total clones | Presumable microbial process |
| 181 | 9 | P22 | 2 | Sulfate reducer |
| 155 | 6 | N12 | 1 | Sulfur oxidizer |
| N28 | 1 | Sulfur oxidizer | ||
| P13 | 2 | Sulfate reducer | ||
| C11 | 3 | Sulfate reducer | ||
| N211 | 1 | Sulfate reducer | ||
| 154 | 42 | N21 | 1 | Sulfate reducer |
| C14 | 1 | Sulfate reducer | ||
| N111 | 1 | Sulfate reducer | ||
| N18 | 7 | Sulfate reducer | ||
| P11 | 2 | Sulfate reducer | ||
| 153 | 2 | C35 | 2 | Sulfate reducer |
| 321 | 1 | N213 | 2 | Sulfate reducer |
| N110 | 2 | Sulfate reducer | ||
| 227 | 2 | P33 | 1 | Sulfate reducer |
| 117 | 2 | N17 | 1 | Sulfate reducer |
| 315 | 6 | P210 | 1 | Sulfate reducer |
| 317 | 27 | C31 | 1 | Sulfate reducer |
| 162 | 8 | C32 | 1 | Sulfate reducer |
| P32 | 2 | Phototrophic sulfur reducer | ||
| 165 | 6 | N36 | 2 | Sulfate reducer |
4. Discussion
5. Conclusions
Supplementary Materials
Supplementary File 1Acknowledgments
Author Contributions
Conflicts of Interest
References
- Komiyama, A.; Ong, J.E.; Poungparn, S. Allometry, biomass, and productivity of mangrove forests: A review. Aquat. Bot. 2008, 89, 128–137. [Google Scholar] [CrossRef]
- Donato, D.C.; Kauffman, J.B.; Murdiyarso, D.; Kurnianto, S.; Stidham, M.; Kanninen, M. Mangroves among the most carbon-rich forests in the tropics. Nat. Geosci. 2011, 4, 293–297. [Google Scholar] [CrossRef]
- Bouillon, S.; Borges, A.V.; Castañeda-Moya, E.; Diele, K.; Dittmar, T.; Duke, N.C.; Kristensen, E.; Lee, S.Y.; Marchand, C.; Middelburg, J.J.; et al. Mangrove production and carbon sinks: A revision of global budget estimates. Glob. Biogeochem. Cycles 2008, 22, 1–12. [Google Scholar] [CrossRef]
- Alongi, D.M. Carbon sequestration in mangrove forests. Carbon Manag. 2012, 3, 313–322. [Google Scholar] [CrossRef]
- Lovelock, C.E.; Ball, M.C.; Choat, B.; Engelbrecht, B.M.J.; Holbrook, N.M.; Feller, I.C. Linking physiological processes with mangrove forest structure: Phosphorus deficiency limits canopy development, hydraulic conductivity and photosynthetic carbon gain in dwarf Rhizophora mangle. Plant Cell Environ. 2006, 29, 793–802. [Google Scholar] [CrossRef] [PubMed]
- Feller, I.C.; Lovelock, C.E.; Mckee, K.L. Nutrient Addition Differentially Affects Ecological Processes of Avicennia germinans in Nitrogen versus Phosphorus Limited Mangrove Ecosystems. Ecosystems 2007, 10, 347–359. [Google Scholar] [CrossRef]
- Lovelock, C.; Feller, I.; Ellis, J.; Schwarz, A. Mangrove growth in New Zealand estuaries: The role of nutrient enrichment at sites with contrasting rates of sedimentation. Oecologia 2007, 153, 633–641. [Google Scholar] [CrossRef] [PubMed]
- Fogel, M.L.; Wooller, M.J.; Cheeseman, J.; Smallwood, B.J.; Roberts, Q.; Romero, I.; Meyers, M.J. Unusually negative nitrogen isotopic compositions (δ 15 N) of mangroves and lichens in an oligotrophic, microbially-influenced ecosystem. Biogeosciences 2008, 5, 1693–1704. [Google Scholar] [CrossRef]
- Kristensen, E.; Jensen, M.H.; Banta, G.T.; Hansen, K.; Holmer, M.; King, J.M. Transformation and transport of inorganic nitrogen in sediments of a southeast Asian mangrove forest. Aquat. Microbiol. Ecol. 1998, 15, 165–175. [Google Scholar] [CrossRef]
- Holguin, G.; Vazquez, P.; Bashan, Y. The role of sediment microorganisms in the productivity, conservation, and rehabilitation of mangrove ecosystems: An overview. Biol. Fertil. Soils. 2001, 33, 265–278. [Google Scholar] [CrossRef]
- Fernandes, S.O.; Bonin, P.C.; Michotey, V.D.; Garcia, N.; LokaBharathi, P.A. Nitrogen-limited mangrove ecosystems conserve N through dissimilatory nitrate reduction to ammonium. Sci. Rep. 2012, 2, 1–5. [Google Scholar] [CrossRef] [PubMed]
- Lee, R.; Joye, S. Seasonal patterns of nitrogen fixation and denitrification in oceanic mangrove habitats. Mar. Ecol. Prog. Ser. 2006, 307, 127–141. [Google Scholar] [CrossRef]
- Zhang, Y.Y.; Dong, J.J.; Yang, Z.H.; Zhang, S.; Wang, Y.S. Phylogenetic diversity of nitrogen-fixing bacteria in mangrove sediments assessed by PCR-denaturing gradient gel electrophoresis. Arch. Microbiol. 2008, 190, 19–28. [Google Scholar] [CrossRef] [PubMed]
- Romero, I.C.; Jacobson, M.; Fuhrman, J.A.; Fogel, M.; Capone, D.G. Long-term nitrogen and phosphorus fertilization effects on N2 fixation rates and nifH gene community patterns in mangrove sediments. Mar. Ecol. 2011, 33, 1–11. [Google Scholar] [CrossRef]
- Zuberer, D.A.; Silver, W.S. Biological dinitrogen fixation (Acetylene reduction) associated with Florida mangroves. Appl. Environ. Microbiol. 1978, 35, 567–575. [Google Scholar] [PubMed]
- Brown, M.M.; Friez, M.J.; Lovell, C.R. Expression of nifH genes by diazotrophic bacteria in the rhizosphere of short form Spartina alterniflora. FEMS Microbiol. Ecol. 2003, 43, 411–417. [Google Scholar] [CrossRef] [PubMed]
- Zehr, J.; Jenkins, B.; Short, S.; Steward, G. Nitrogenase gene diversity and microbial community structure: A cross-system comparison. Environ. Microbiol. 2003, 5, 539–554. [Google Scholar] [CrossRef] [PubMed]
- Lovell, C.; Decker, P.; Bagwell, C.; Thompson, S.; Matsui, G. Analysis of a diverse assemblage of diazotrophic bacteria from Spartina alterniflora using dgge and clone library screening. J. Microbiol. Methods 2008, 73, 160–171. [Google Scholar] [CrossRef] [PubMed]
- Flores-Mireles, A.; Winans, S.; Holguin, G. Molecular Characterization of Diazotrophic and Denitrifying Bacteria Associated with Mangrove Roots. Appl. Environ. Microbiol. 2007, 73, 7308–7321. [Google Scholar] [CrossRef] [PubMed]
- Dias, A.C.F.; Silva, M.C.P.; Cotta, S.R.; Dini-Andreote, F.; Soares, F.L.; Salles, J.F.; Azevedo, J.L.; van Elsas, J.D.; Andreoted, F.D. Abundance and Genetic Diversity of nifH Gene Sequences in Anthropogenically Affected Brazilian Mangrove Sediments. Appl. Environ. Microbiol. 2012, 78, 7960–7967. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Peng, M.; Li, Y. Phylogenetic diversity of nitrogen-fixing bacteria and the nifH gene from mangrove rhizosphere soil. Can. J. Microbiol. 2012, 58, 531–539. [Google Scholar] [CrossRef] [PubMed]
- Ravikumar, S.; Kathiresan, K.; Ignatiammal, S.; Selvam, M.; Shanthy, S. Nitrogen-fixing azotobacters from mangrove habitat and their utility as marine biofertilizers. J. Exp. Mar. Biol. Ecol. 2004, 312, 5–17. [Google Scholar] [CrossRef]
- Sengupta, A.; Chaudhuri, S. Ecology of heterotrophic dinitrogen fixation in the rhizosphere of mangrove plant community at the Ganges river estuary in India. Oecologia 1991, 87, 560–564. [Google Scholar] [CrossRef]
- Holguin, G.; Guzman, M.A.; Bashan, Y. Two new nitrogen-fixing bacteria from the rhizosphere of mangrove trees: Their isolation, identification and in vitro interaction with rhizosphere Staphylococcus sp. FEMS Microbiol. Lett. 1992, 101, 207–216. [Google Scholar]
- Vitousek, P.M.; Aber, J.D.; Howarth, R.W.; Matson, P.A.; Schindler, D.W.; Schlesinger, D.H.; Likens, G.E.; Tilman, D.G. Human Alteration of the Global Nitrogen Cycle: Sources and consequences. Ecol. Appl. 1997, 7, 737–750. [Google Scholar] [CrossRef]
- Galloway, J.N.; Dentener, F.J.; Capone, D.G.; Boyer, E.W.; Howarth, R.W.; Seitzinger, S.P.; Asner, G.P.; Cleveland, C.C.; Green, P.A.; Holland, E.A.; et al. Nitrogen cycles: Past, present, and future. Biogeochemistry 2004, 70, 153–226. [Google Scholar] [CrossRef]
- Bürgmann, H.; Widmer, F.; Sigler, W.V.; Zeyer, J. mRNA Extraction and Reverse Transcription-PCR Protocol for Detection of nifH Gene Expression by Azotobacter vinelandii in Soil. Appl. Environ. Microbiol. 2003, 69, 1928–1935. [Google Scholar] [CrossRef] [PubMed]
- Fauzi, A.; Skidmore, A.K.; Heitkönig, I.M.A.; van Gils, H.; Schlerf, M. Eutrophication of mangroves linked to depletion of foliar and soil base cations. Environ. Monit. Assess. 2014, 186, 8487–8498. [Google Scholar] [CrossRef] [PubMed]
- Farrar, J.; Hawes, M.; Jones, D.; Lindow, S. How roots control the flux of carbon to the rhizosphere. Ecology 2003, 84, 827–837. [Google Scholar] [CrossRef]
- Chambers, L.G.; Davis, S.E.; Troxler, T.G. Sea Level Rise in the Everglades: Plant-Soil-Microbial Feedbacks in Response to Changing Physical Conditions. In Microbiology of the Everglades Ecosystem; Entry, J.A., Gottlieb, A.D., Jayachandran, K., Ogram, A., Eds.; CRC Press: Boca Raton, FL, USA, 2015; pp. 89–112. [Google Scholar]
- Alongi, D.M.; Wattayakor, G.; Pfitzner, J.; Tirendi, F.; Zarorskis, I.; Brunskill, G.J.; Davidson, A.; Clough, B.F. Organic carbon accumulation and metabolic pathways in sediments of mangrove forest in southern Thailand. Mar. Geol. 2001, 179, 85–103. [Google Scholar] [CrossRef]
- Goneea, M.E.; Paytan, A.; Herrera-Silveira, J.A. Tracing organic matter sources and carbon burial in mangrove sediments over the past 160 years. Estuar. Coast. Shelf Sci. 2004, 61, 211–227. [Google Scholar] [CrossRef]
- McKee, K.L. Biophysical controls on accretion and elevation change in Caribbean mangrove ecosystems. Estuar. Coast. Shelf Sci. 2011, 91, 475–483. [Google Scholar] [CrossRef]
- Breithaupt, J.L.; Smoak, J.M.; Smith, T.J.; Sanders, C.J.; Hoare, A. Organic carbon burial rates in mangrove sediments: Strengthening the global budget. Glob. Biogeochem. Cycles 2012, 26. [Google Scholar] [CrossRef]
- Lovelock, C.E.; Feller, I.C.; Reef, R.; Ruess, R.W. Variable effects of nutrient enrichment on soil respiration in mangrove forests. Plant Soil 2014, 379, 135–148. [Google Scholar] [CrossRef]
- Butler, J.L.; Williams, M.A.; Bottomley, P.J.; Myrold, M.D. Microbial community associated with rhizosphere carbon flux. Appl. Environ. Microbiol. 2003, 69, 6793–6800. [Google Scholar] [CrossRef] [PubMed]
- Kuzyakov, Y.; Domanski, G. Carbon inputs by plants into the soil. Review. J. Plant Nutr. Soil Sci. 2000, 163, 421–431. [Google Scholar] [CrossRef]
- Welsh, D.T. Nitrogen fixation in seagrass meadows: Regulation, plant-bacteria interactions and significance to primary production. Ecol. Lett. 2000, 3, 58–71. [Google Scholar] [CrossRef]
- Walker, T.S.; Bais, H.P.; Grotewold, E.; Vivanco, J.M. Root exudation and rhizosphere biology. Plant Physiol. 2003, 132, 44–51. [Google Scholar] [CrossRef] [PubMed]
- Nedwell, D.B.; Blackburn, T.H.; Wiebe, W.J. Dynamic nature of the turnover of organic carbon, nitrogen and sulphur in the sediments of Jamaican mangrove forest. Mar. Ecol. Prog. Ser. 1994, 110, 223–231. [Google Scholar] [CrossRef]
- Sherman, R.E.; Fahey, T.J.; Howarth, R.W. Soil-plant interactions in a neotropical mangrove forest: Iron, phosphorus and sulfur dynamics. Oecologia 1998, 115, 553–563. [Google Scholar] [CrossRef]
- Alongi, D.M. The role of bacteria in nutrient cycling n tropical mangrove and other coastal benthic ecosystems. Hydrobiologia 1994, 285, 19–32. [Google Scholar] [CrossRef]
- Alongi, D.M. Present State and future of the world’s mangrove forests. Environ. Conserv. 2002, 29, 331–349. [Google Scholar] [CrossRef]
- Miki, T.; Yokokawa, T.; Matsui, K. Biodiversity and multifunctionality in a microbial community: A novel theoretical approach to quantify functional redundancy. Proc. Boil. Soc. 2014, 281, 20132498. [Google Scholar] [CrossRef] [PubMed]
- Nielsen, U.N.; Ayres, E.; Wall, D.H.; Bardgett, R.D. Soil biodiversity and carbon cycling: A review and synthesis of studies examining diversity-function relationships. Eur. J. Soil Sci. 2011, 62, 105–116. [Google Scholar] [CrossRef]
- Hastie, T.; Tibshirani, R. Generalized additive models. Stat. Sci. 1986, 1, 297–310. [Google Scholar] [CrossRef]
- Tao, M.; Xie, P.; Chen, J.; Qin, B.; Zhang, D.; Niu, Y.; Zhang, M.; Wang, Q.; Wu, L. Use of a generalized additive model to investigate key abiotic factors affecting microcystin cellular quotas in heavy bloom areas of Lake Taihu. PLoS ONE 2012, 7, e32020. [Google Scholar] [CrossRef] [PubMed]
- Mckee, K.L.; Cahoon, D.R.; Feller, I.C. Caribbean mangroves adjust to rising sea level through biotic controls on change in soil elevation. Global Ecol. Biogeogr. 2007, 16, 545–556. [Google Scholar] [CrossRef]
- Feller, I.C. Effects of nutrient enrichment on growth and herbivory of dwarf red mangrove (Rhizophora mangle). Ecol. Monogr. 1995, 65, 477–505. [Google Scholar] [CrossRef]
- Rodriguez, W.; Feller, I.C. Mangrove landscape characterization and change in Twin Cays, Belize using aerial photography and Ikonos satellite data. Atoll Res. Bull. 2004, 513, 1–24. [Google Scholar] [CrossRef]
- Cline, J.D. Spectrophotometric determination of hydrogen sulfide in natural waters. Limnol. Oceanogr. 1969, 14, 454–458. [Google Scholar] [CrossRef]
- Solorzano, L. Determination of ammonia in natural waters by the phenol hypochlorite method. Limnol. Oceanogr. 1969, 14, 799–801. [Google Scholar] [CrossRef]
- Strickland, J.D.H.; Parsons, T.R. A Practical Handbook of Sea-Water Analysis; Fisheries Reseach Board of Canada: Ottawa, ON, Canada, 1972; p. 311. [Google Scholar]
- Zehr, J.P.; McReynolds, L.A. Use of degenerate oligonucleotides for amplification of the nifH gene from the marine cyanobacterium Trichodesmiun thiebautii. Appl. Environ. Microbiol. 1989, 55, 2522–2526. [Google Scholar] [PubMed]
- Hewson, I.; Fuhrman, J.A. Spatial and vertical biogeography of coral reef sediment bacterial and diazotroph communities. Mar. Ecol. Prog. Ser. 2006, 306, 79–86. [Google Scholar] [CrossRef]
- Deslippe, J.R.; Egger, K.N.; Henry, G.H.R. Impacts of warming and fertilization on nitrogen-fixing microbial com-munities in the Canadian High Arctic. FEMS Microbiol. Ecol. 2005, 53, 41–50. [Google Scholar] [CrossRef] [PubMed]
- Chao, A. Estimating the population size for capture-recapture data with unequal catchability. Biometrics 1987, 43, 783–791. [Google Scholar] [CrossRef] [PubMed]
- Ter Braak, C.J.F.; Prentice, I.C. A Theory of Gradient Analysis. Adv. Ecol. Res. 1988, 18, 271–339. [Google Scholar]
- Legendre, P.; Legendre, L. Numerical Ecology, 2nd ed.; Elsevier Science: Amsterdam, The Netherlands, 1998; p. 1006. [Google Scholar]
- Mussmann, M.; Ishii, K.; Rabus, R.; Amann, R. Diversity and vertical distribution of cultured and uncultured Deltaproteobacteria in an intertidal mud flat of the Wadden Sea. Environ. Microbiol. 2005, 7, 405–418. [Google Scholar] [CrossRef] [PubMed]
- Methe, B.; Nelson, K.; Eisen, J.; Paulsen, I.; Nelson, W.; Heidelberg, J.; Wu, D.; Wu, M.; Ward, N.; Beanan, M.; et al. Genome of Geobacter sulfurreducens: Metal reduction in subsurface environments. Science 2003, 302, 1967–1969. [Google Scholar] [CrossRef] [PubMed]
- Yannarell, A.; Steppe, T.; Paerl, H. Genetic variance in the composition of two functional groups (diazotrophs and cyanobacteria) from a hypersaline microbial mat. Appl. Environ. Microb. 2006, 72, 1207–1217. [Google Scholar] [CrossRef] [PubMed]
- Zehr, J.P.; Mellon, M.; Braun, S.; Litaker, W.; Steppe, T.F.; Paerl, W.H. Diversity of heterotrophic nitrogen fixation genes in a marine cyanobacterial mat. Appl. Environ. Microb. 2005, 61, 2527–2532. [Google Scholar]
- Musat, F.; Harder, J.; Widdel, F. Study of nitrogen fixation in microbial communities of oil-contaminated marine sediment microcosms. Environ. Microbiol. 2006, 8, 1834–1843. [Google Scholar] [CrossRef] [PubMed]
- Moisander, P.H.; Piehler, M.F.; Paerl, H.W. Diversity and activity of epiphytic nitrogen-fixers on standing dead stems of the salt marsh grass Spartina alterniflora. Aquat. Microb. Ecol. 2005, 39, 271–279. [Google Scholar] [CrossRef]
- Heidelberg, J.; Seshadri, R.; Haveman, S.; Hemme, C.; Paulsen, I.; Kolonay, J.; Eisen, J.; Ward, N.; Methe, B.A.; Brinkac, L.; et al. The genome sequence of the anaerobic, sulfate-reducing bacterium Desulfovibrio vulgaris hildenborough. Nat. biotechnol. 2004, 22, 554–559. [Google Scholar] [CrossRef] [PubMed]
- Bagwell, C.; Rocque, J.; Smith, G.; Polson, S.; Friez, M.; Longshore, J.; Lovell, C. Molecular diversity of diazotrophs in oligotrophic tropical seagrass bed communities. FEMS microbiol. Ecol. 2002, 39, 113–119. [Google Scholar] [CrossRef] [PubMed]
- Ellison, A.M. Macroecology of mangroves: Large-scale patterns and processes in tropical coastal forests. Trees 2002, 16, 181–194. [Google Scholar] [CrossRef]
- Ellison, A.M. Mangrove restoration: Do we know enough? Restor. Ecol. 2000, 8, 219–229. [Google Scholar] [CrossRef]
- Knelman, J.E.; Legg, T.M.; O’Neill, S.P.; Washenberger, C.L.; Gonzalez, A.; Cleveland, C.C.; Nemergut, D.R. Bacterial community structure and function change in association with colonizer plants during early primary succession in a glacier forefield. Soil Biol. Biochem. 2012, 46, 172–180. [Google Scholar] [CrossRef]
- Zou, Y.; Zhang, J.; Yang, D.; Chen, X.; Zhao, J.; Xiu, W.; Lai, X.; Li, G. Effects of different land use patterns on nifH genetic diversity of soil nitrogen-fixing microbial communities in Leymus chinensis steppe. Acta Ecol. Sin. 2011, 31, 150–156. [Google Scholar] [CrossRef]
- Zhan, J.; Sun, Q. Diversity of free-living nitrogen-fixing microorganisms in the rhizosphere and non-rhizosphere of pioneer plants growing on wastelands of copper mine tailings. Microbiol. Res. 2012, 167, 157–165. [Google Scholar] [CrossRef] [PubMed]
- Rico, L.; Ogaya, R.; Terradas, J.; Penuelas, J. Community structures of N2-fixing bacteria associated with the phyllosphere of a Holm oak forest and their response to drought. Plant Biol. 2014, 16, 586–593. [Google Scholar] [CrossRef] [PubMed]
- Alongi, D.M.; Tirendi, F.; Dixon, P.; Trott, L.A.; Brunskill, G.J. Mineralization of organic matter in intertidal sediments of a tropical semi-enclosed delta. Estuar. Coast. Shelf Sci. 1999, 48, 451–467. [Google Scholar] [CrossRef]
- Lee, R.; Porubsky, W.; Feller, I.; McKee, K.; Joye, S. Porewater biogeochemistry and soil metabolism in dwarf red mangrove habitats (Twin Cays, Belize). Biogeochemistry 2008, 87, 181–198. [Google Scholar] [CrossRef]
- Guisan, A.; Zimmermann, N. Predictive habitat distribution models in ecology. Ecol. Model. 2000, 135, 147–186. [Google Scholar] [CrossRef]
- McGill, B.J.; Enquist, B.J.; Weiher, E.; Westoby, M. Rebuilding community ecology from functional traits. Trends Ecol. Evol. 2006, 21, 178–185. [Google Scholar] [CrossRef] [PubMed]
- Trosvik, P.; Rudi, K.; Naes, T.; Kohler, A.; Chan, K.S.; Jakobsen, K.S.; Stenseth, N.C. Characterizing mixed microbial population dynamics using time-series analysis. ISME J. 2008, 2, 707–715. [Google Scholar] [CrossRef] [PubMed]
- Trosvik, P.; Stenseth, N.C.; Rudi, K. Convergent temporal dynamics of the human infant gut microbiota. ISME J. 2010, 4, 151–158. [Google Scholar] [CrossRef] [PubMed]
- Tai, X.S.; Mao, M.L.; Liu, G.X.; Chen, T.; Zhang, W.; Wu, X.K.; Long, H.Z.; Zhang, B.G.; Zhang, Y. High diversity of nitrogen-fixing bacteria in the upper reaches of the Heihe River, northwestern China. Biogeosciences 2013, 10, 5589–5600. [Google Scholar] [CrossRef]
- McKee, K.L. Soil Physicochemical Patterns and Mangrove Species Distribution-Reciprocal Effects? J. Ecol. 1993, 81, 477–487. [Google Scholar] [CrossRef]
- Gallon, J.R. Reconciling the incompatible: N2 fixation and O2. New Phytol. 1992, 122, 571–609. [Google Scholar] [CrossRef]
- Alongi, D.M.; Tirendi, F.; Trott, L.A.; Xuan, T.T. Benthic decomposition rates and pathways in plantations of the mangrove Rhizophora apiculata in the Mekong delta, Vietman. Mar. Ecol. Prog. Ser. 2000, 194, 87–101. [Google Scholar] [CrossRef]
- Hughes, J.B.; Hellmann, J.J.; Ricketts, T.H.; Bohannan, B.J.M. Counting the uncountable: Statistical approaches to estimating microbial diversity. Appl. Environ. Microb. 2001, 67, 4399–4406. [Google Scholar] [CrossRef]
- Lyons, K.G.; Brigham, C.A.; Traut, B.H.; Schwartz, M.W. Rare species and ecosystem functioning. Conserv. Biol. 2005, 19, 1019–1024. [Google Scholar] [CrossRef]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license ( http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Romero, I.C.; Jacobson-Meyers, M.E.; Fuhrman, J.A.; Capone, D.G. Phylogenetic Diversity of Diazotrophs along an Experimental Nutrient Gradient in Mangrove Sediments. J. Mar. Sci. Eng. 2015, 3, 699-719. https://doi.org/10.3390/jmse3030699
Romero IC, Jacobson-Meyers ME, Fuhrman JA, Capone DG. Phylogenetic Diversity of Diazotrophs along an Experimental Nutrient Gradient in Mangrove Sediments. Journal of Marine Science and Engineering. 2015; 3(3):699-719. https://doi.org/10.3390/jmse3030699
Chicago/Turabian StyleRomero, Isabel C., Myrna E. Jacobson-Meyers, Jed A. Fuhrman, and Douglas G. Capone. 2015. "Phylogenetic Diversity of Diazotrophs along an Experimental Nutrient Gradient in Mangrove Sediments" Journal of Marine Science and Engineering 3, no. 3: 699-719. https://doi.org/10.3390/jmse3030699