Microbial Carbon Substrate Utilization Differences among High- and Average-Yield Soybean Areas
Abstract
:1. Introduction
2. Materials and Methods
2.1. “Grow for the Green” Yield Contest
2.2. Study Area Descriptions
2.3. Sample Collection and Processing
2.4. Statistical Analyses
3. Results and Discussion
3.1. Yield Differences
3.2. Treatment Effects
3.3. Principal Component Analysis
3.4. Correlations
4. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Sugiyama, A.; Ueda, Y.; Zushi, T.; Takase, H.; Yazaki, K. Changes in the bacterial community of soybean rhizospheres during growth in the field. PLoS ONE 2014, 9, e100709. [Google Scholar] [CrossRef] [PubMed]
- Balser, T.C.; Kirchner, J.W.; Firestone, M.K. Methodological variability in microbial community level physiological profiles. Soil Sci. Soc. Am. J. 2002, 66, 519–523. [Google Scholar] [CrossRef]
- Larkin, R.P. Characterization of soil microbial communities under different potato cropping systems by microbial population dynamics, substrate utilization, and fatty acid profiles. Soil Biol. Biochem. 2003, 35, 1451–1466. [Google Scholar] [CrossRef]
- Biolog. Microbial Community Analysis with EcoPlatesTM 2007. Available online: http://www.biolog.com/pdf/milit/00A_012_EcoPlate_Sell_Sheet.pdf (accessed on 2 February 2017).
- Bucher, A.E.; Lanyon, L.E. Evaluating soil management with microbial community-level physiological profiles. Appl. Soil Ecol. 2005, 29, 59–71. [Google Scholar] [CrossRef]
- Konopka, A.; Oliver, L.; Turco, R.F., Jr. The use of carbon substrate utilization patterns in environmental and ecological microbiology. Microbial Ecol. 1998, 35, 103–115. [Google Scholar] [CrossRef]
- Haack, S.K.; Garchow, H.; Klug, M.J.; Forney, L.J. Analysis of factors affecting the accuracy, reproducibility, and interpretation of microbial community carbon source utilization patterns. Appl. Environ. Microbiol. 1995, 61, 1458–1468. [Google Scholar] [PubMed]
- Muniz, S.; Lacarta, J.; Pata, M.P.; Jimenez, J.J.; Navarro, E. Analysis of the diversity of substrate utilization of soil bacteria exposed to Cd and earthworm activity using generalized additive models. PLoS ONE 2014, 9, e85057. [Google Scholar] [CrossRef] [PubMed]
- Gryta, A.; Fraq, M.; Oszust, K. The application of the biolog EcoPlate approach in Ecotoxicological evaluation of dairy sewage sludge. Appl. Biochem. Biotechol. 2014, 174, 1434–1443. [Google Scholar] [CrossRef] [PubMed]
- Stefanowicz, A. The biolog plates technique as a tool in ecological studies of microbial communities. Polish J. Environ. Stud. 2006, 15, 669–676. [Google Scholar]
- Tacarindua, C.R.P.; Shiraiwa, T.; Homma, K.; Kumagai, E.; Sameshima, R. The effects of increased temperature on crop growth and yield of soybean grown in a temperature gradient chamber. Field Crops Res. 2013, 154, 74–81. [Google Scholar] [CrossRef]
- Degens, B.P.; Schipper, L.A.; Sparling, G.P.; Duncan, L.C. Is the microbial community in a soil with reduced catabolic diversity less resistant to stress or disturbance? Soil Biol. Biochem. 2001, 33, 1143–1153. [Google Scholar] [CrossRef]
- Nannipieri, P.; Ascher, J.; Ceccherini, M.T.; Landi, L.; Peitramellara, G.; Renella, G. Microbial diversity and soil functions. Eur. J. Soil Sci. 2003, 54, 655–670. [Google Scholar] [CrossRef]
- Janniche, G.S.; Spliid, H.; Albrechtsen, H.J. Microbial community-level physiological profiles (CLPP) and herbicide mineralization potential in groundwater affected by agricultural land use. J. Contam. Hydrol. 2012, 140, 45–55. [Google Scholar] [CrossRef] [PubMed]
- Van Roekel, R.J.; Purcell, L.C.; Salmeron, M. Physiological and management factors contributing to soybean potential yield. Field Crops Res. 2015, 182, 86–97. [Google Scholar] [CrossRef]
- Arkansas Soybean Association (ASA). 2014. Grow for the Green Data 1999–2013. Available online: http://www.arkansassoybean.com/Go_for_the_Green_Data_1999-2013.xlsx (accessed on 2 February 2017).
- United States Department of Agriculture (USDA); Natural Resources Conservation Service (NRCS). Major Land Resource Area (MLRA) Explorer. 2014. NRCS MLRA Explorer. Available online: apps.cei.psu.edu/mlra/ (accessed on 2 February 2017).
- United States Department of Agriculture (USDA); Natural Resources Conservation Service (NRCS). 2014. Official Soil Series Descriptions—View by Name. Available online: https://soilseries.sc.egov.usda.gov/osdname.aspx (accessed on 2 February 2017).
- United States Department of Agriculture (USDA); Natural Resources Conservation Service (NRCS). 2014. Web Soil Survey. Available online: https://websoilsurvey.sc.egov.usda.gov/App/HomePage.htm (accessed on 2 February 2017).
- United States Department of Agriculture (USDA); Natural Resources Conservation Service (NRCS). 2006. Land Resource Regions and Major Land Resource Areas of the United States, the Caribbean, and the Pacific Basin. USDA Handbook 296. Available online: http://www.nrcs.usda.gov/Internet/FSE_DOCUMENTS/nrcs142p2_050898.pdf (accessed on 2 February 2017).
- Southern Regional Climate Center (SRCC). 2014. Station Search. Available online: www.srcc.lsu.edu/ station_search.html (accessed on 2 February 2017).
- Yu, C.; Hu, X.M.; Deng, W.; Li, Y.; Xiong, C.; Ye, C.H.; Han, G.M.; Li, X. Changes in soil microbial community structure and functional diversity in the rhizosphere surrounding mulberry subjected to long-term fertilization. Appl. Soil Ecol. 2015, 86, 30–40. [Google Scholar] [CrossRef]
- Preston-Mafham, J.; Boddy, L.; Randerson, P.F. Analysis of microbial community functional diversity using sole-carbon-source utilization profiles—A critique. FEMS Microbiol. Ecol. 2002, 42, 1–14. [Google Scholar] [PubMed]
- Shannon, C.E. A mathematical theory of communication. Bell Syst. Tech. J. 1948, 27, 379–423, 623–656. [Google Scholar] [CrossRef]
- Beals, M.; Gross, L.; Harrell, S. Diversity Indices: Shannon’s H and E. 2000. Available online: http://www.tiem.utk.edu/~gross/bioed/bealsmodules/shannonDI.html (accessed on 2 February 2017).
- Simpson, E.H. Measurement of diversity. Nature 1949, 163, 688. [Google Scholar] [CrossRef]
- Beals, M.; Gross, L.; Harrell, S. Diversity Indices: Simpson’s D and E. 1999. Available online: http://www.tiem.utk.edu/~gross/bioed/bealsmodules/simpsonDI.html (accessed on 2 February 2017).
- Zak, J.C.; Willig, M.R.; Moorhead, D.L.; Wildman, H.G. Functional diversity of microbial communities: A quantitative approach. Soil Biol. Biochem. 1994, 26, 1101–1108. [Google Scholar] [CrossRef]
- United States Department of Agriculture (USDA); National Agricultural Statistics Service (NASS). 2016. Quick Stats. Available online: https://quickstats.nass.usda.gov/ (accessed on 14 September 2016).
- Deng, H.; Ge, L.; Xu, T.; Zhang, M.; Wang, X.; Zhang, Y.; Peng, H. Analysis of the metabolic utilization of carbon sources and potential functional diversity of the bacterial community in lab-scale horizontal subsurface-flow constructed wetlands. J. Environ. Qual. 2011, 40, 1730–1736. [Google Scholar] [CrossRef] [PubMed]
- Gates, J. Groundwater irrigation and the development of the Grand Prairie rice industry. Ark. Hist. Quart. 2005, 64, 394–413. [Google Scholar]
- Girvan, M.S.; Bullimore, J.; Pretty, J.N.; Osborn, A.M.; Ball, A.S. Soil type is the primary determinant of the composition of the total and active bacterial communities in arable soils. Appl. Environ. Microbiol. 2003, 69, 1800–1809. [Google Scholar] [CrossRef] [PubMed]
- Chakraborty, A.; Chakrabarti, K.; Chakraborty, A.; Ghosh, S. Effect of long-term fertilizers and manure application on microbial biomass and microbial activity of a tropical agricultural soil. Biol. Fertil. Soils 2011, 47, 227–233. [Google Scholar] [CrossRef]
- Paul, E.A.; Clark, F.E. Methods for studying soil microorganisms. In Soil Microbiology and Biochemistry; Paul, E.A., Ed.; Academic Press: San Diego, CA, USA, 1996. [Google Scholar]
- Froliks, A.; Knox, C.; Lim, E.; Jewison, T.; Law, V.; Hau, D.D.; Liu, P.; Gautam, B.; Ly, S.; Guo, A.C.; et al. SMPDB: The small molecule pathway database. Nucleic Acids Res. 2010, 38, D480–D487. [Google Scholar] [CrossRef] [PubMed]
- Bach, B.; Meudec, E.; Lepoutre, J.P.; Rossignol, T.; Blondin, B.; Dequin, S.; Camarasa, C. New insights into γ-Aminobutyric acid catabolism: Evidence for γ-Hydroxybutyric acid and Polyhydrozybutyrate synthesis in Saccharomyces cerevisiae. Appl. Environ. Microbiol. 2009, 75, 4231–4239. [Google Scholar] [CrossRef] [PubMed]
- Berg, J.M.; Tymoczko, J.L.; Stryer, L. Glycogen metabolism. In Biochemistry; Freeman, W.H., Ed.; W.H. Freeman and Company: New York, NY, USA, 2002. [Google Scholar]
- Helferich, B.; Schafer, W. α-Methyl-d-Glucoside. Org. Synth. 1926, 6, 64. [Google Scholar]
- Rais, B.; Chassagnole, C.; Letellier, T.; Fell, D.A.; Mazat, J.P. Threonine synthesis from aspartate in Escherichia coli cell-free extracts: Pathway dynamics. J. Biochem. 2001, 356, 425–432. [Google Scholar] [CrossRef]
- Kurkov, S.V.; Loftsson, T. Cyclodextrins. Int. J. Pharm. 2013, 453, 167–180. [Google Scholar] [CrossRef] [PubMed]
- Chou, D.K.; Krishnamurthy, R.; Randolph, T.W.; Carpenter, J.F.; Manning, M.C. Effects of Tween® 20 and Tween® 80 on the stability of albutropin during agitation. J. Pharm. Sci. 2005, 94, 1368–1381. [Google Scholar] [CrossRef] [PubMed]
- Lyons, M.M.; Dobbs, F.C. Differential utilization of carbon substrates by aggregate-associated and water-associated heterotrophic bacterial communities. Hydrobiologia 2012, 686, 181–193. [Google Scholar] [CrossRef]
- Grayston, S.J.; Campbell, C.D.; Bardgett, R.D.; Mawdsley, J.L.; Clegg, C.D.; Ritz, K.; Griffiths, B.S.; Rodwell, J.S.; Edwards, S.J.; Davies, W.J.; et al. Assessing shifts in microbial community structure across a range of grasslands of differing management intensity using CLPP, PLFA and community DNA techniques. Appl. Soil Ecol. 2004, 25, 63–84. [Google Scholar] [CrossRef]
- Manlay, R.J.; Feller, C.; Swift, M.J. Historical evolution of soil organic matter concepts and their relationships with the fertility and sustainability of cropping systems. Agric. Ecosyst. Environ. 2007, 119, 217–233. [Google Scholar] [CrossRef]
- Baumann, K.; Dignac, M.F.; Rumpel, C.; Bardoux, G.; Sarr, A.; Steffens, M.; Maron, P.A. Soil microbial diversity affects soil organic matter decomposition in a silty grassland soil. Biogeochemistry 2013, 112, 201–212. [Google Scholar] [CrossRef]
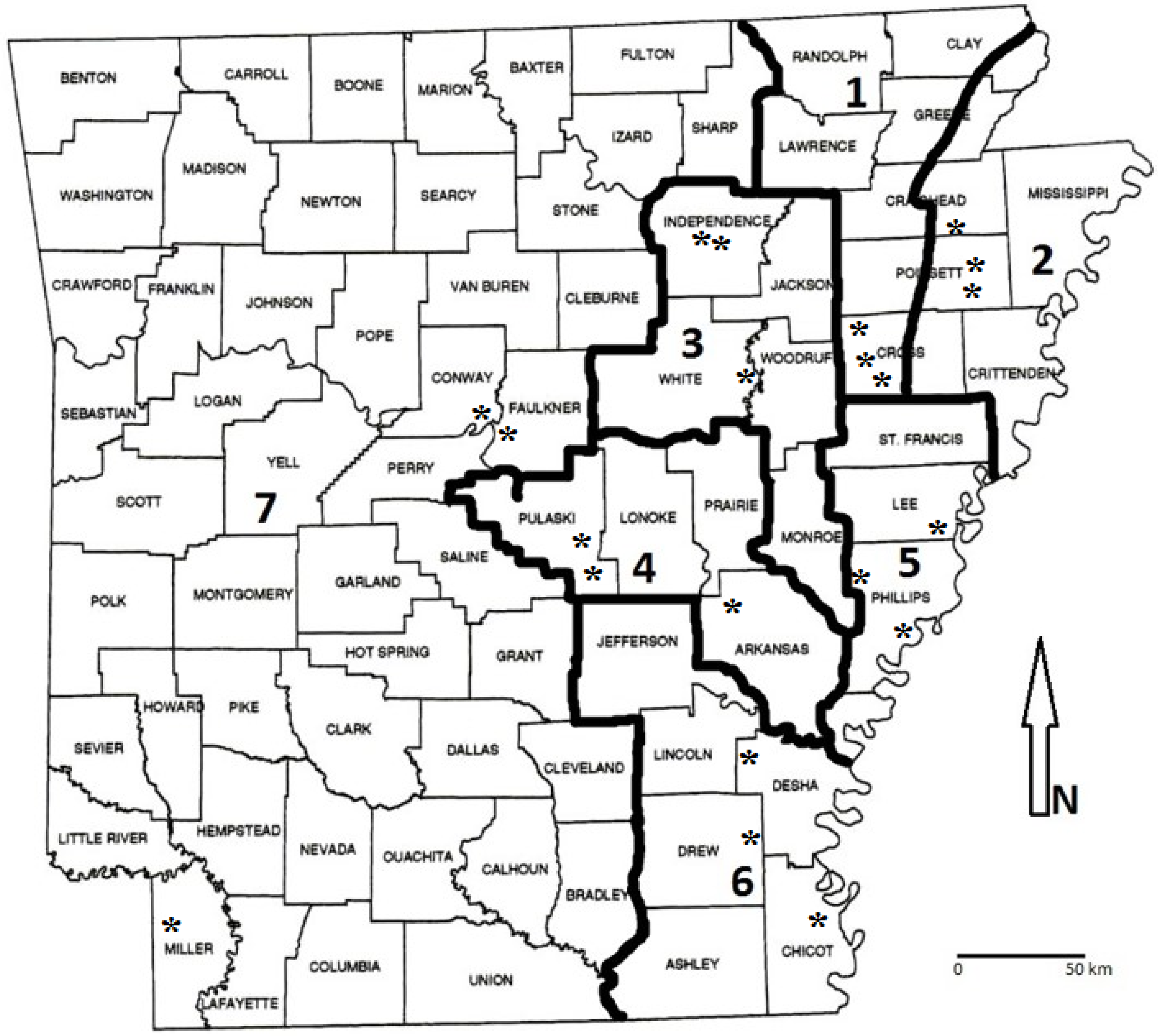
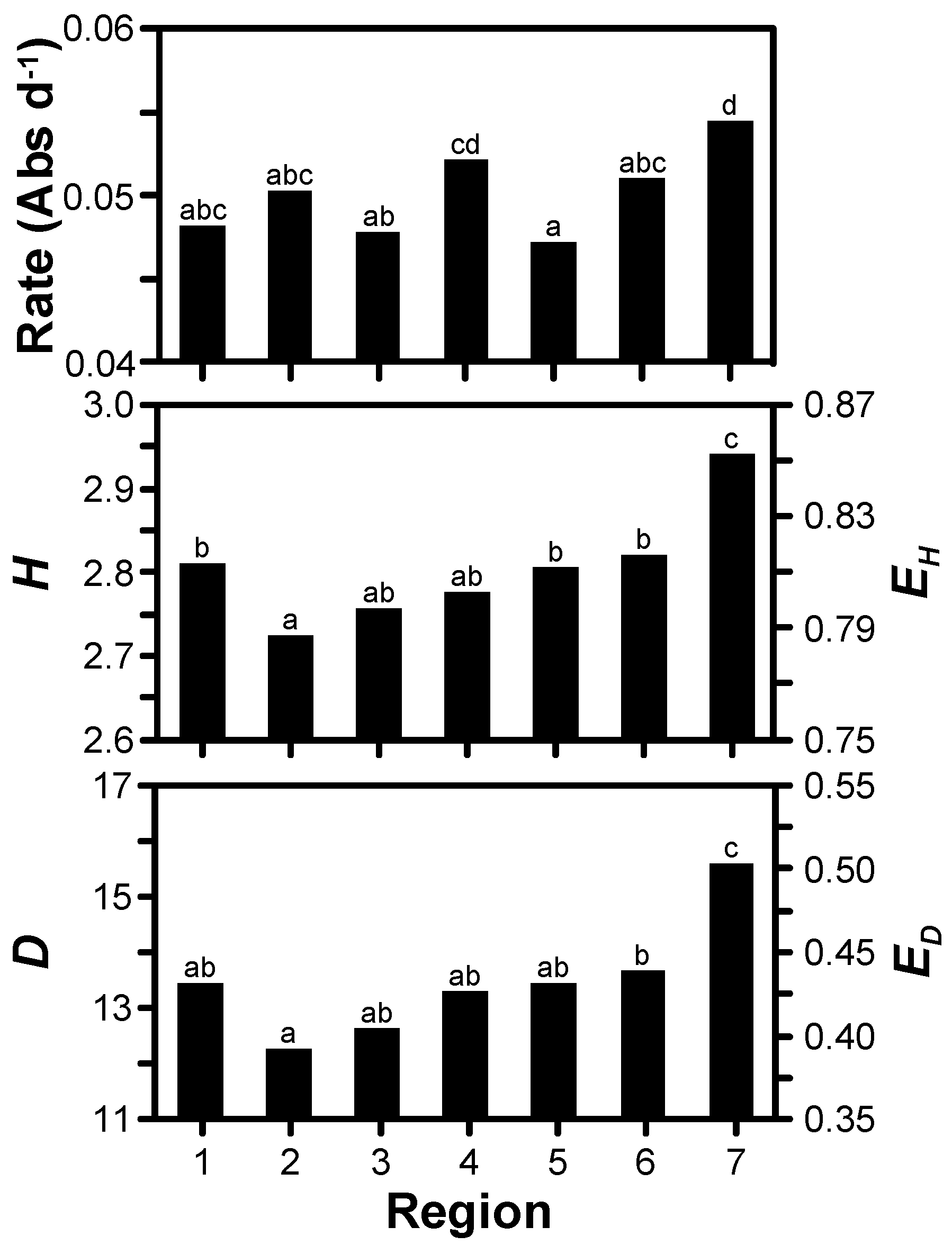
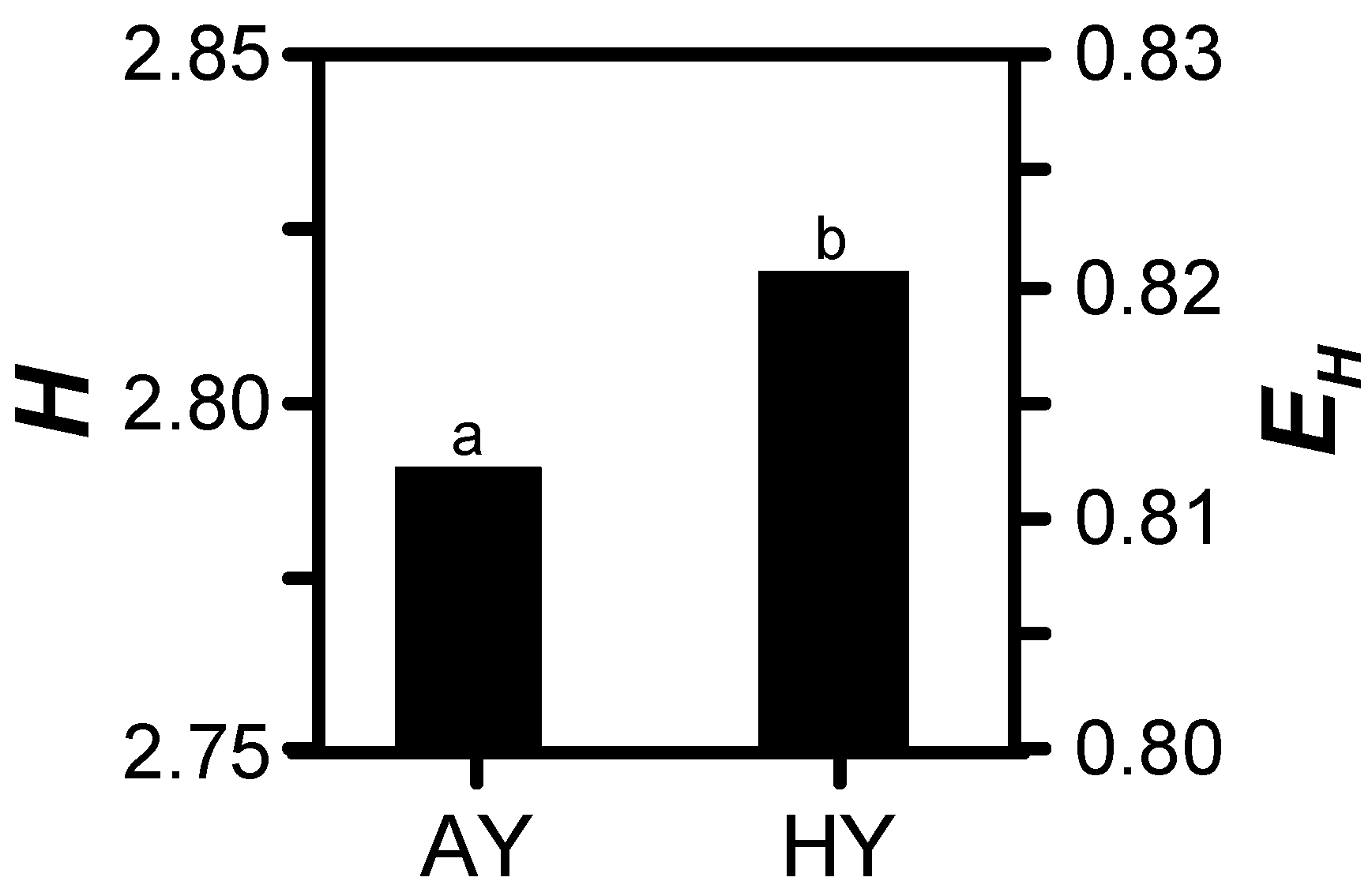
| Year | Region † | Site | MLRA ‡ | Dominant Soil Series (Taxonomic Description) in High-Yielding Area § | Soil Surface Texture § |
|---|---|---|---|---|---|
| 2014 | 1 | 1 | 131A | Dundee (Typic Endoaqualfs) | Fine sandy loam |
| 2 | 131A | Dundee (Typic Endoaqualfs) | Silt loam | ||
| 2 | 1 | 134 | Arkabutla (Fluventic Endoaquepts) | Silt loam | |
| 2 | 134 | Crowley (Typic Albaqualfs) | Silt loam | ||
| 3 | 1 | 131A | Bosket (Mollic Hapludalfs) | Fine sandy loam | |
| 2 | 131A | Wiville (Ultic Hapludalfs) | Fine sandy loam | ||
| 4 | 1 | 131B | Hebert (Aeric Epiaqualfs) | Silt loam | |
| 2 | 131B | Rilla (Typic Hapludalfs) | Silt loam | ||
| 5 | 1 | 131A | Commerce (Fluvaquentic Endoaquepts) | Silt loam | |
| 2 | 134 | Dubbs (Typic Hapludalfs) | Silt loam | ||
| 6 | 1 | 131B | Rilla (Typic Hapludalfs) | Silt loam | |
| 2 | 131B | Hebert (Aeric Epiaqualfs) | Silt loam | ||
| 7 | 1 | 131B | Gallion (Typic Hapludalfs) | Silt loam | |
| 2 | 131C | Bossier (Aeric Epiaquerts) | Clay | ||
| 2015 | 1 | 1 | 131A | Dubbs (Typic Hapludalfs) | Silt loam |
| 2 | 131A | Dundee (Typic Endoaqualfs) | Silt loam | ||
| 2 | 1 | 134 | Arkabutla (Fluventic Endoaquepts) | Silt loam | |
| 2 | 134 | Crowley (Typic Albaqualfs) | Silt loam | ||
| 3 | 1 | 131A | Bosket (Mollic Hapludalfs) | Fine sandy loam | |
| 2 | 131A | Wiville (Ultic Hapludalfs) | Fine sandy loam | ||
| 4 | 1 | 131B | Portland (Chromic Epiaquerts) | Silty clay | |
| 2 | 131D | Rilla (Typic Hapludalfs) | Silt loam | ||
| 5 | 1 | 131A | Henry (Typic Fragiaqualfs) | Silt loam | |
| 2 | 134 | Dundee (Typic Endoaqualfs) | Silt loam | ||
| 6 | 1 | 131B | Dundee (Typic Endoaqualfs) | Silt loam | |
| 2 | 131A | Hebert (Aeric Epiaqualfs) | Silt loam | ||
| 7 | 1 | 131B | Gallion (Typic Hapludalfs) | Silt loam | |
| 2 | 131B | Roxana (Aeric Epiaquerts) | Silt loam |
| Well Number | Compound Type | Compound |
|---|---|---|
| 7 | Carbohydrates | d-cellobiose |
| 8 | α-d-lactose | |
| 9 | β-methyl-d-glucoside | |
| 10 | d-xylose | |
| 11 | i-erythritol | |
| 12 | d-mannitol | |
| 13 | N-acetyl-d-glucosamine | |
| 15 | Glucose-1-phosphate | |
| 16 | d,l-α-glycerol phosphate | |
| 17 | d-galactonic acid γ-lactone | |
| 31 | Amines and amides | Phenylethylamine |
| 32 | Putrescine | |
| 2 | Carboxylic and ketonic Acids | Pyruvic acid methyl ester |
| 18 | d-galacturonic acid | |
| 21 | γ-hydroxybutyric acid | |
| 22 | Itaconic acid | |
| 23 | α-ketobutyric acid | |
| 24 | d-malic acid | |
| 19 | Phenolics | 2-hydroxy benzoic acid |
| 20 | 4-hydroxy benzoic acid | |
| 3 | Polymers | Tween 40 |
| 4 | Tween 80 | |
| 5 | α-cyclodextrin | |
| 6 | Glycogen | |
| 25 | Amino acids | l-arginine |
| 26 | l-asparagine | |
| 27 | l-phenylalanine | |
| 28 | l-serine | |
| 29 | l-threonine | |
| 30 | Glycl-l-glutamic acid |
| Property † | Region | ||||||
|---|---|---|---|---|---|---|---|
| 1 | 2 | 3 | 4 | 5 | 6 | 7 | |
| AWCD rate | 0.036–0.061 | 0.042–0.061 | 0.039−0.063 | 0.033−0.070 | 0.037−0.065 | 0.042−0.062 | 0.023−0.082 |
| Shannon’s diversity | 2.37–2.99 | 2.42−2.97 | 2.35–2.94 | 2.22−3.02 | 2.21−3.25 | 2.65−2.92 | 2.49−3.14 |
| Shannon’s evenness | 0.69−0.87 | 0.70–0.86 | 0.68−0.85 | 0.65−0.88 | 0.64−0.95 | 0.77−0.85 | 0.73−0.92 |
| Simpson’s diversity | 6.66−17.1 | 6.43–16.2 | 6.87−16.0 | 6.22−18.9 | 7.52−21.5 | 11.1−15.6 | 6.79−20.1 |
| Simpson’s evenness | 0.21−0.55 | 0.21–0.52 | 0.22−0.52 | 0.20−0.61 | 0.24−0.69 | 0.36−0.50 | 0.22−0.65 |
| Variable | Region | Yield Area | R x YA |
|---|---|---|---|
| ___________________________ p _______________________ | |||
| Growth rate † | <0.001 | NS ‡ | NS |
| Shannon’s diversity | 0.043 | 0.004 | NS |
| Shannon’s evenness | 0.043 | 0.004 | NS |
| Simpson’s diversity | 0.036 | NS | NS |
| Simpson’s evenness | 0.036 | NS | NS |
| Yield Area | Component | Eigenvalue | Substrate | Type † | Coefficient ‡ |
|---|---|---|---|---|---|
| High-yield | PC 1 | 6.01 (19.4%) | α-d-lactose | C | 0.2899 |
| Glycl-l-glutamic acid | A | 0.2747 | |||
| d-xylose | C | 0.2688 | |||
| α-ketobutyric acid | K | 0.2675 | |||
| 2-hydroxy benzoic acid | P | 0.2660 | |||
| γ-hydroxybutyric acid | K | 0.2632 | |||
| PC 2 | 2.53 (8.1%) | Pyruvic acid methyl ester | K | 0.3350 | |
| d-cellobiose | C | 0.3208 | |||
| N-acetyl-d-glucosamine | C | 0.3004 | |||
| Glucose-1-phosphate | C | 0.2898 | |||
| Glycogen | O | 0.2790 | |||
| Average-yield | PC 1 | 6.89 (22.2%) | d-xylose | C | 0.3089 |
| α-d-lactose | C | 0.3052 | |||
| l-threonine | A | 0.2921 | |||
| i-erythritol | C | 0.2829 | |||
| α-cyclodextrin | O | 0.2676 | |||
| Glycl-l-glutamic acid | A | 0.2667 | |||
| 2-hydroxy benzoic acid | P | 0.2648 | |||
| PC 2 | 2.37 (7.6%) | d-cellobiose | C | 0.4162 | |
| d-mannitol | C | 0.3680 | |||
| Glucose-1-phosphate | C | 0.3353 | |||
| β-methyl-d-glucoside | C | 0.3110 | |||
| Tween 80 | O | 0.2557 |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Adams, T.C.; Brye, K.R.; Savin, M.C.; Lee, J.A.; Gbur, E.E. Microbial Carbon Substrate Utilization Differences among High- and Average-Yield Soybean Areas. Agriculture 2017, 7, 48. https://doi.org/10.3390/agriculture7060048
Adams TC, Brye KR, Savin MC, Lee JA, Gbur EE. Microbial Carbon Substrate Utilization Differences among High- and Average-Yield Soybean Areas. Agriculture. 2017; 7(6):48. https://doi.org/10.3390/agriculture7060048
Chicago/Turabian StyleAdams, Taylor C., Kristofor R. Brye, Mary C. Savin, Jung Ae Lee, and Edward E. Gbur. 2017. "Microbial Carbon Substrate Utilization Differences among High- and Average-Yield Soybean Areas" Agriculture 7, no. 6: 48. https://doi.org/10.3390/agriculture7060048
APA StyleAdams, T. C., Brye, K. R., Savin, M. C., Lee, J. A., & Gbur, E. E. (2017). Microbial Carbon Substrate Utilization Differences among High- and Average-Yield Soybean Areas. Agriculture, 7(6), 48. https://doi.org/10.3390/agriculture7060048





