Sodium Azide as a Chemical Mutagen in Wheat (Triticum aestivum L.): Patterns of the Genetic and Epigenetic Effects with iPBS and CRED-iPBS Techniques
Abstract
1. Introduction
2. Materials and Methods
2.1. Plant Material, Seed Germination and Treatments
2.2. Isolation of Genomic DNA
2.3. iPBS and CRED-iPBS PCR Assays
3. Results
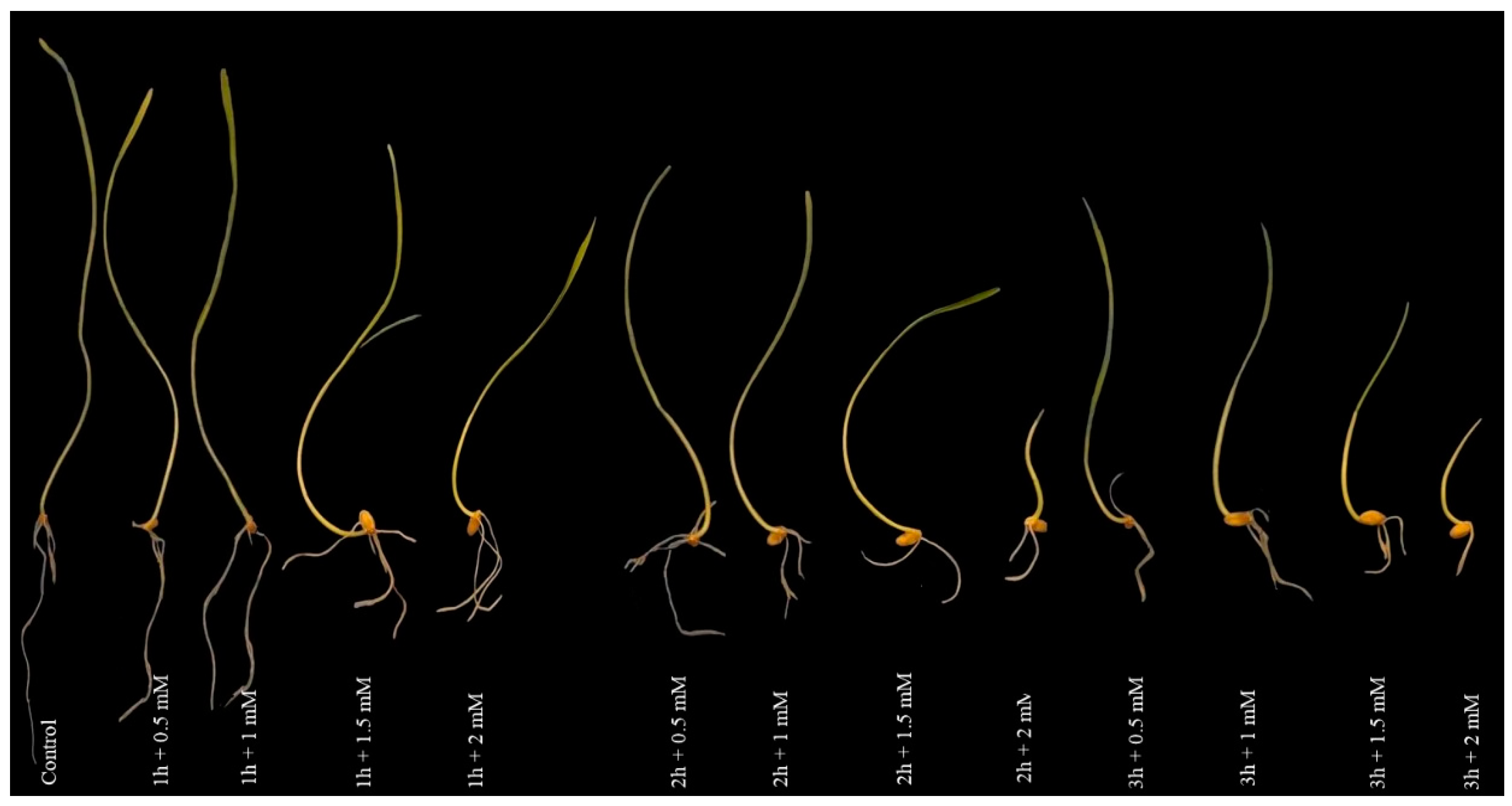
3.1. Morphological Responses under NaN3 Application
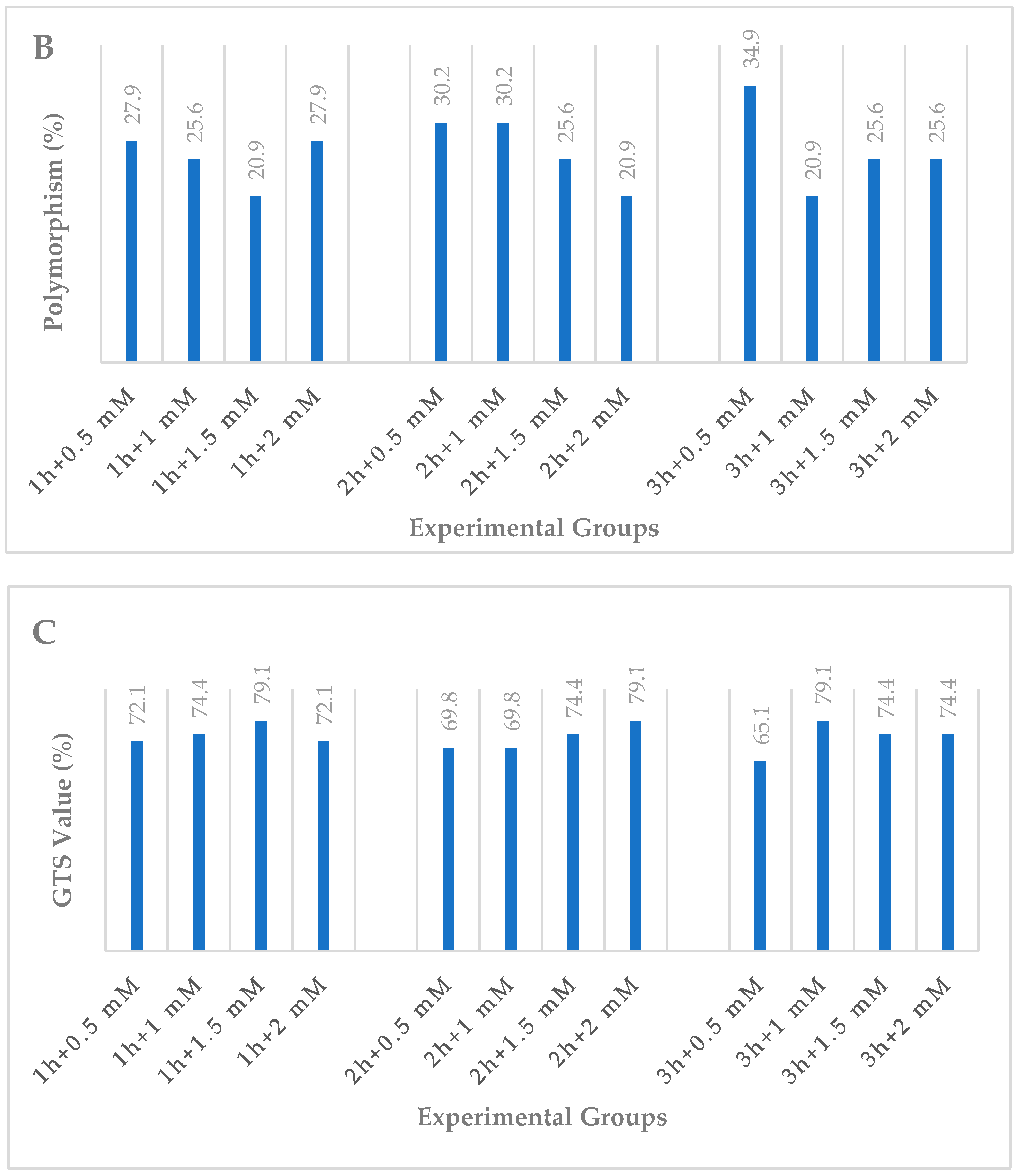
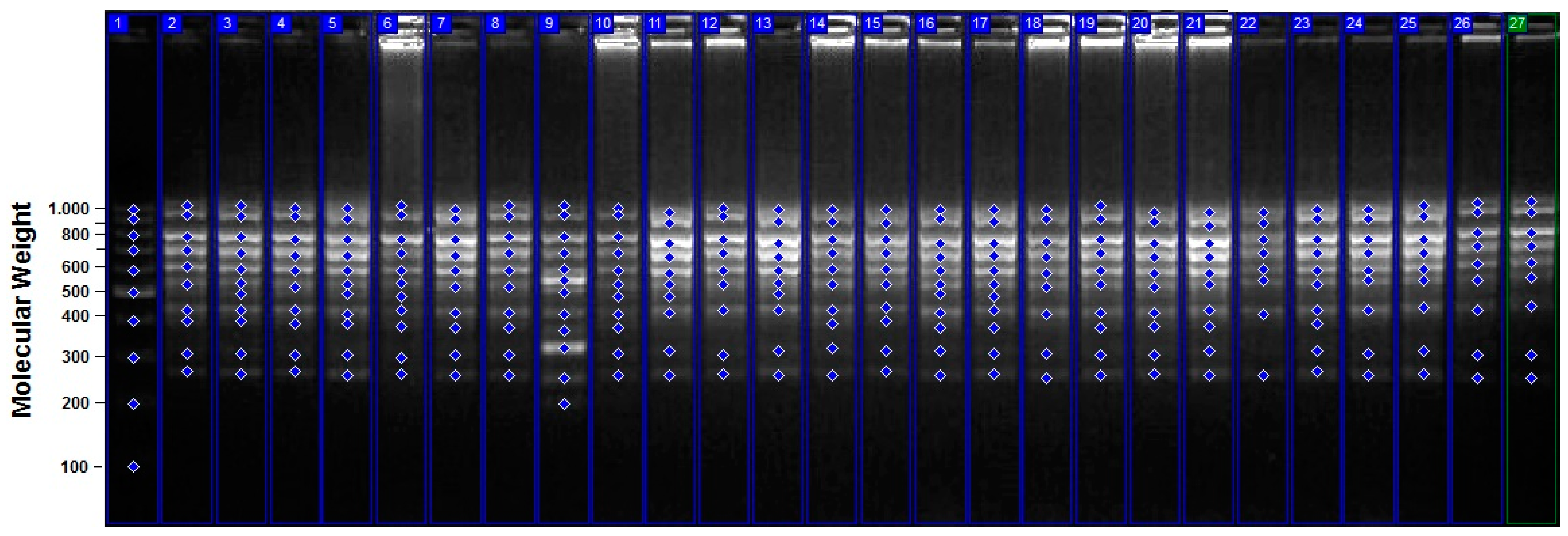
3.2. iPBS Assays
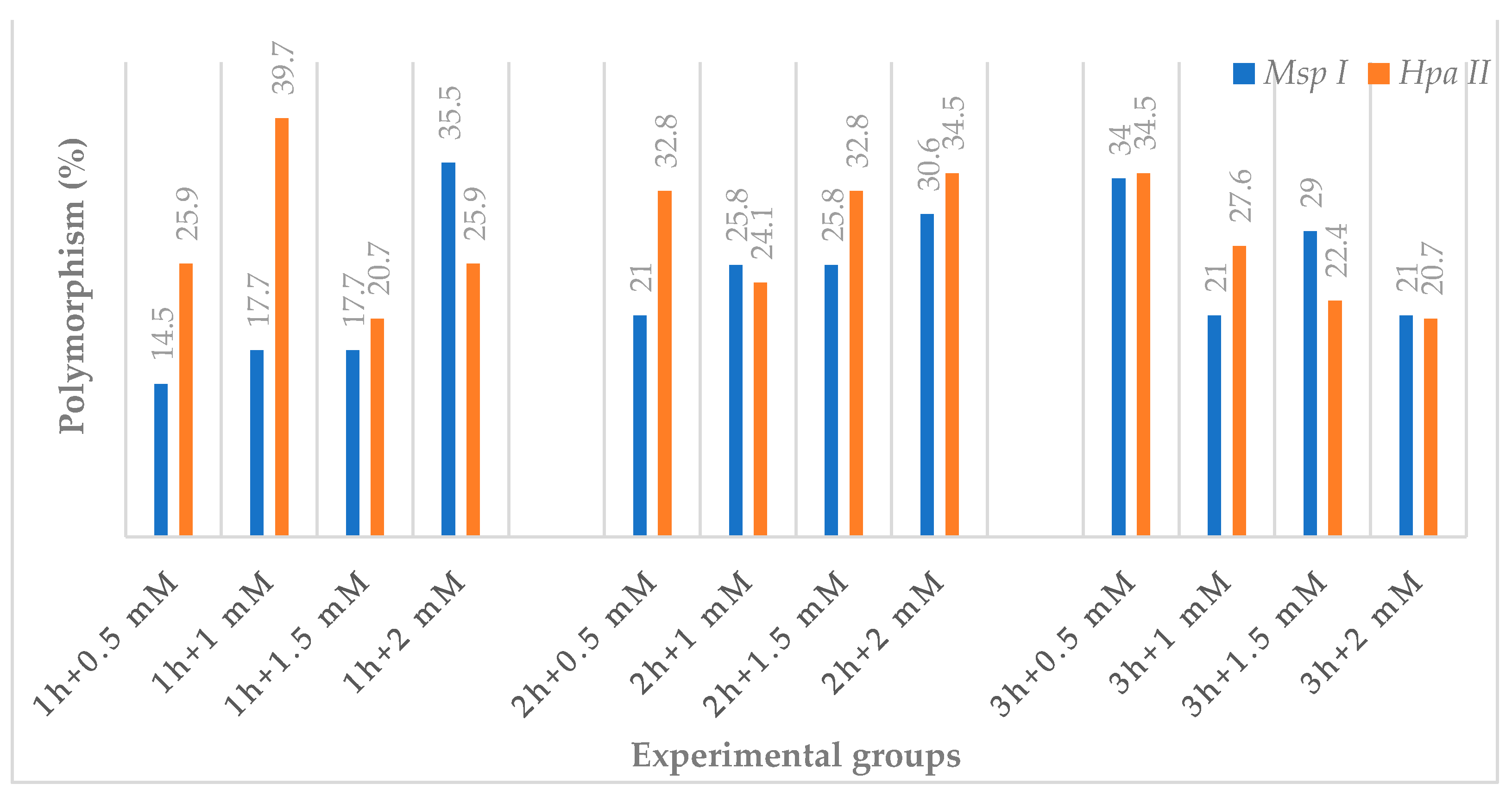
3.3. CRED-iPBS Assays
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Yaman, H.M.; Ordu, B.; Zencirci, N.; Kan, M. Coupling socioeconomic factors and cultural practices in production of einkorn and emmer wheat species in Turkey. Environ. Dev. Sustain. 2020, 22, 8079–8096. [Google Scholar] [CrossRef]
- Tadesse, W.; Bishaw, Z.; Assefa, S. Wheat production and breeding in Sub-Saharan Africa: Challenges and opportunities in the face of climate change. Int. J. Clim. Chang. Strateg. Manag. 2019, 11, 696–715. [Google Scholar] [CrossRef]
- Hasanuzzaman, M.; Bhuyan, M.B.; Parvin, K.; Bhuiyan, T.F.; Anee, T.I.; Nahar, K.; Hossen, M.S.; Zulfiqar, F.; Alam, M.M.; Fujita, M. Regulation of ROS metabolism in plants under environmental stress: A review of recent experimental evidence. Int. J. Mol. Sci. 2020, 21, 8695. [Google Scholar] [CrossRef]
- Raza, A.; Razzaq, A.; Mehmood, S.S.; Zou, X.; Zhang, X.; Lv, Y.; Xu, J. Impact of climate change on crops adaptation and strategies to tackle its outcome: A review. Plants 2019, 8, 34. [Google Scholar] [CrossRef]
- Demirel, F.; Gürcan, K.; Taner, A. Molecular Characterization and Assessment of Population Structure of Hulled Wheats. J. Agric. Nat. 2022, 25, 192–199. [Google Scholar] [CrossRef]
- Kuzmanović, L.; Rossini, F.; Ruggeri, R.; Pagnotta, M.A.; Ceoloni, C. Engineered durum wheat germplasm with multiple alien introgressions: Agronomic and quality performance. Agronomy 2020, 10, 486. [Google Scholar] [CrossRef]
- Qaim, M. Role of new plant breeding technologies for food security and sustainable agricultural development. Appl. Econ. Perspect. Policy 2020, 42, 129–150. [Google Scholar] [CrossRef]
- Hosseinpour, A.; Tosun, M.; Haliloğlu, K. Effects of ethyl methanesulfonate (ems) applied at different times and doses on some germination and seedling characteristics in wheat (Triticum aestivum L.). J. Atatürk Uni. Fac. Agri. 2021, 52, 190–200. [Google Scholar]
- Karakoca, T.A.; Akgün, İ. Determination of the mutagenic effects of different gamma radiation dose applications on some agricultural traits in M2 generation. J. Suleyman Demirel Uni. Grad. Sch. Natur. App. Sci. 2020, 24, 96–104. [Google Scholar]
- de Oliveira Collet, S.A.; Collet, M.A.; Maria de Fátima, P. Differential gene expression for isozymes in somatic mutants of Vitis vinifera L. (Vitaceae). Biochem. Syst. Ecol. 2005, 33, 691–703. [Google Scholar] [CrossRef]
- Pandit, R.; Bhusal, B.; Regmi, R.; Neupane, P.; Bhattarai, K.; Maharjan, B.; Acharya, S.; Kc, B.; Poudel, M.R. Mutation breeding for crop improvement: A review. Rev. Food Agric. 2021, 2, 31–35. [Google Scholar] [CrossRef]
- Saleem, M.; Mukhtar, Z.; Cheema, A.; Atta, B. Induced mutation and in vitro techniques as a method to induce salt tolerance in Basmati rice (Oryza sauva L.). Int. J. Environ. Sci. Technol. 2005, 2, 141–145. [Google Scholar] [CrossRef]
- Öztürk, İ.; Ayşe, Ş.; Kiliç, T.H.; Şahinde, Ş. Selection of advanced mutant wheat (Triticum aestivum L.) lines based on yield and quality parameters. Turkish J. Agric. Nat. Sci. 2020, 7, 87–95. [Google Scholar] [CrossRef]
- Kim, S.H.; Kim, S.W.; Ahn, J.W.; Ryu, J.; Kwon, S.J.; Kang, B.C.; Kim, J.B. Frequency, spectrum, and stability of leaf mutants induced by diverse γ-ray treatments in two Cymbidium hybrids. Plants 2020, 9, 546. [Google Scholar] [CrossRef]
- Hewawasam, W.; Bandara, D.; Aberathne, W. New phenotypes of Crossandra infundibuliformis var. Danica through in vitro culture and induced mutations. Trop. Agric. Res. 2004, 16, 253–270. [Google Scholar]
- Ikhajiagbe, B.; Omoregie, U.E. Growth, yield, genetic parameters and random amplified polymorphic DNA (RAPD) of five rice varieties treated with sodium azide and sown under different saline conditions. Bull. Natl. Res. Cent. 2020, 44, 89. [Google Scholar] [CrossRef]
- Khan, S.; Al-Qurainy, F.; Anwar, F. Sodium azide: A chemical mutagen for enhancement of agronomic traits of crop plants. Environ. We Int. J. Sci. Technol. 2009, 4, 1–21. [Google Scholar]
- Jenks, M.A.; Hasegawa, P.M.; Jain, S.M.; Foolad, M. Advances in Molecular Breeding toward Drought and Salt Tolerant Crops; Springer: New York, NY, USA, 2007. [Google Scholar]
- Wu, C.T.; Morris, J.R. Genes, genetics, and epigenetics: A correspondence. Science 2001, 293, 1103–1105. [Google Scholar] [CrossRef]
- Bird, A. DNA methylation patterns and epigenetic memory. Genes Dev. 2002, 16, 6–21. [Google Scholar] [CrossRef]
- Karaca, M.; Ince, A.; Gocer, E.; Aydin, A. Exonic and intronic DNA methylation differences in a fiber specific gene of Pima cotton (Gossypium barbadense L.). J. Sci. Eng. Res. 2016, 3, 478–486. [Google Scholar]
- Osabe, K.; Clement, J.D.; Bedon, F.; Pettolino, F.A.; Ziolkowski, L.; Llewellyn, D.J.; Finnegan, E.J.; Wilson, I.W. Genetic and DNA methylation changes in cotton (Gossypium) genotypes and tissues. PLoS ONE 2014, 9, e86049. [Google Scholar] [CrossRef] [PubMed]
- Al-Harrasi, I.; Al-Yahyai, R.; Yaish, M.W. Differential DNA methylation and transcription profiles in date palm roots ex-posed to salinity. PLoS ONE 2018, 13, e0191492. [Google Scholar] [CrossRef] [PubMed]
- Fan, H.H.; Wei, J.; Li, T.C.; Li, Z.P.; Guo, N.; Cai, Y.P.; Lin, Y. DNA methylation alterations of upland cotton (Gossypium hirsutum) in response to cold stress. Acta Physiol. Plant. 2013, 35, 2445–2453. [Google Scholar] [CrossRef]
- Feng, S.J.; Liu, X.S.; Tao, H.; Tan, S.K.; Chu, S.S.; Oono, Y.; Zhang, X.D.; Chen, J.; Yang, Z.M. Variation of DNA methylation patterns associated with gene expression in rice (Oryza sativa) exposed to cadmium. Plant Cell Environ. 2016, 39, 2629–2649. [Google Scholar] [CrossRef]
- Li, R.; Hu, F.; Li, B.; Zhang, Y.; Chen, M.; Fan, T.; Wang, T. Whole genome bisulfite sequencing methylome analysis of mulberry (Morus alba) reveals epigenome modifications in response to drought stress. Sci. Rep. 2020, 10, 8013. [Google Scholar] [CrossRef] [PubMed]
- Haliloğlu, K.; Türkoğlu, A.; Balpınar, Ö.; Nadaroğlu, H.; Alaylı, A.; Poczai, P. Effects of Zinc, Copper and Iron Oxide Nanoparticles on Induced DNA Methylation, Genomic Instability and LTR Retrotransposon Polymorphism in Wheat (Triticum aestivum L.). Plants 2022, 11, 2193. [Google Scholar] [CrossRef] [PubMed]
- Oda, M.; Glass, J.L.; Thompson, R.F.; Mo, Y.; Olivier, E.N.; Figueroa, M.E.; Selzer, R.R.; Richmond, T.A.; Zhang, X.; Dannenberg, L.; et al. High-resolution genome-wide cytosine methylation profiling with simultaneous copy number analysis and optimization for limited cell numbers. Nucleic Acids Res. 2009, 37, 3829–3839. [Google Scholar] [CrossRef]
- Olkhov-Mitsel, E.; Bapat, B. Strategies for discovery and validation of methylated and hydroxymethylated DNA biomarkers. Cancer Med. 2012, 1, 237–260. [Google Scholar] [CrossRef]
- Türkoğlu, A.; Tosun, M.; Haliloğlu, K. Mutagenic effects of sodium azide on in vitro mutagenesis, polymorphism and genomic instability in wheat (Triticum aestivum L.). Mol. Biol. Rep. 2022, 49, 10165–10174. [Google Scholar] [CrossRef]
- Zeinalzadehtabrizi, H.; Hosseinpour, A.; Aydin, M.; Haliloglu, K. A modified genomic DNA extraction method from leaves of sunflower for PCR based analyzes. J. Biodivers. Environ. Sci. 2015, 7, 222–225. [Google Scholar]
- Kalendar, R.; Antonius, K.; Smýkal, P.; Schulman, A.H. iPBS: A universal method for DNA fingerprinting and retrotransposon isolation. Theor. Appl. Genet. 2010, 121, 1419–1430. [Google Scholar] [CrossRef] [PubMed]
- Turkoglu, A. Effects of mammalian sex hormones on regeneration capacity, retrotransposon polymorphism and genomic instability in wheat (Triticum aestivum L.). Plant Cell Tissue Organ Cult. 2023, 152, 647–659. [Google Scholar] [CrossRef]
- Türkoğlu, A.; Haliloğlu, K.; Balpinar, Ö.; Öztürk, H.I.; Özkan, G.; Poczai, P. The effect of mammalian sex hormones on polymorphism and genomic instability in the common bean (Phaseolus vulgaris L.). Plants 2022, 11, 2071. [Google Scholar] [CrossRef] [PubMed]
- Türkoğlu, A.; Tosun, M.; Haliloğlu, K. Evaluation of ethyl methanesulfonate-induced in vitro mutagenesis, polymorphism and genomic instability in wheat (Triticum aestivum L.). J. Crop Sci. Biotechnol. 2023, 26, 199–213. [Google Scholar] [CrossRef]
- Moose, S.P.; Mumm, R.H. Molecular plant breeding as the foundation for 21st century crop improvement. Plant Physiol. 2008, 147, 969–977. [Google Scholar] [CrossRef] [PubMed]
- Louwaars, N.P. Plant breeding and diversity: A troubled relationship? Euphytica 2018, 214, 114. [Google Scholar] [CrossRef]
- Holme, I.B.; Gregersen, P.L.; Brinch-Pedersen, H. Induced genetic variation in crop plants by random or targeted mutagenesis: Convergence and differences. Front. Plant Sci. 2019, 10, 1468. [Google Scholar] [CrossRef]
- Oladosu, Y.; Rafii, M.Y.; Abdullah, N.; Malek, M.A.; Rahim, H.A.; Hussin, G.; Ismail, M.R.; Latif, M.A.; Kareem, I. Genetic variability and diversity of mutant rice revealed by quantitative traits and molecular markers. Agrociencia 2015, 49, 249–266. [Google Scholar]
- Sikora, P.; Chawade, A.; Larsson, M.; Olsson, J.; Olsson, O. Mutagenesis as a tool in plant genetics, functional genomics, and breeding. Int. J. Plant Genom. 2011, 2011, 314829. [Google Scholar] [CrossRef]
- Ilhan, E.; Kasapoglu, A.G.; Turkoglu, A.; Aygoren, A.S.; Muslu, S.; Aydin, M.; Aldaif, M.; Haliloglu, K. Evaluation of sodium azide as a chemical mutagen in developing cold-tolerant quinoa (Chenopodium quinoa Willd.) lines. Iran. J. Sci. 2023, 47, 663–671. [Google Scholar] [CrossRef]
- Grúz, P.; Yasui, M.; Ukai, A.; Horibata, K.; Honma, M.; Sugiyama, K.-I. Potent mutagenicity of an azide, 3-azido-1, 2-propanediol, in human TK6 cells. Mutat. Res. Genet. Toxicol. Environ. Mutagen. 2022, 876–877, 503475. [Google Scholar] [CrossRef]
- Deh Sheikh, A.A.; Moradnejad, M. Mutagenesis in olive (Olea europaea L.) calli caused by sodium azide and detection of mutants using ISSR and RAPD markers. J. Hortic. Sci. Biotechnol. 2014, 89, 153–158. [Google Scholar] [CrossRef]
- Grúz, P.; Juříček, M.; Ẑák, P.; Velemínský, J. Mutagenicity of 3-azido-1,2-propanediol and 9-(3-azido-2-hydroxypropyl)-adenine in repair deficient strains of Escherichia coli. Mutat. Res. Lett. 1993, 303, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Ahmed, M.; Abdelaziem, T.; Nower, A. In vitro mutagenesis and propagation of paulownia tomentosa (thumb) for salt tolerance. SABRAO J. Breed. Genet. 2022, 54, 305–320. [Google Scholar] [CrossRef]
- Parry, M.A.; Madgwick, P.J.; Bayon, C.; Tearall, K.; Hernandez-Lopez, A.; Baudo, M.; Rakszegi, M.; Hamada, W.; Al-Yassin, A.; Ouabbou, H.; et al. Mutation discovery for crop improvement. J. Exp. Bot. 2009, 60, 2817–2825. [Google Scholar] [CrossRef] [PubMed]
- Haliloğlu, K.; Türkoğlu, A.; Öztürk, A.; Niedbała, G.; Niazian, M.; Wojciechowski, T.; Piekutowska, M. Genetic Diversity and Population Structure in Bread Wheat Germplasm from Türkiye Using iPBS-Retrotransposons-Based Markers. Agronomy 2023, 13, 255. [Google Scholar] [CrossRef]
- Mansour, A. Water deficit induction of Copia and Gypsy genomic retrotransposons. Plant Stress 2009, 3, 33–39. [Google Scholar]
- Ramakrishnan, M.; Satish, L.; Kalendar, R.; Narayanan, M.; Kandasamy, S.; Sharma, A.; Emamverdian, A.; Wei, Q.; Zhou, M. The dynamism of transposon methylation for plant development and stress adaptation. Int. J. Mol. Sci. 2021, 22, 11387. [Google Scholar] [CrossRef]


| Primer Name | Sequence (5′-3′) | Tm (°C) | CG (%) | Optimal Annealing Ta (°C) |
|---|---|---|---|---|
| 2383 | GCATGGCCTCCA | 50.5 | 66.6 | 53.0 |
| 2384 | GTAATGGGTCCA | 40.9 | 50.0 | 50.0 |
| 2385 | CCATTGGGTCCA | 45.7 | 58.3 | 53.0 |
| 2386 | CTGATCAACCCA | 41.4 | 50.0 | 48.0 |
| 2388 | TTGGAAGACCCA | 43.4 | 50.0 | 51.0 |
| 2389 | ACATCCTTCCCA | 43.0 | 50.0 | 48.0 |
| 2390 | GCAACAACCCCA | 47.6 | 58.3 | 56.4 |
| 2391 | ATCTGTCAGCCA | 43.6 | 50.0 | 52.0 |
| 2392 | TAGATGGTGCCA | 43.1 | 50.0 | 52.2 |
| 2402 | TCTAAGCTCTTGATACCA | 49.0 | 38.9 | 57.0 |
| iPBS Primer | ± 1 | Control * | Experimental Groups | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 h | 2 h | 3 h | ||||||||||||
| 0.5 mM | 1 mM | 1.5 mM | 2 mM | 0.5 mM | 1 mM | 1.5 mM | 2 mM | 0.5 mM | 1 mM | 1.5 mM | 2 mM | |||
| 2383 | + | 8 | - | - | 544 | - | - | 555 | - | - | - | - | - | |
| - | - | - | - | 1466 | 1466; 306 | 1466 | 1466; 600; 306 | 1233 | 1466; 306 | 600 | 1233; 600; 306 | - | ||
| 2384 | + | 2 | 1300; 900; 562; 391 | 1350; 900; 550 | - | - | - | - | - | - | - | - | - | - |
| - | - | - | - | - | - | - | - | - | - | - | ||||
| 2385 | + | 6 | - | - | - | - | - | - | - | - | - | - | - | - |
| - | - | - | 569 | 848; 569; 475; 390 | 848; 569; 475; 390 | 569; 475; 390 | 569; 475; 390 | 569; 475; 390 | 848; 569; 475; 390 | 475 | 475 | 848; 569; 475; 390 | ||
| 2386 | + | 2 | - | - | - | - | - | - | - | - | - | 500 | - | 555 |
| - | - | - | - | - | - | - | - | - | - | - | 900 | - | ||
| 2388 | + | 4 | - | - | - | - | - | - | - | - | - | - | - | - |
| - | 960 | - | 960 | 960 | 960; 649 | 960 | - | 837 | 960; 837; 649 | - | - | 960 | ||
| 2389 | + | 1 | - | - | - | - | - | - | - | - | - | - | - | 766 |
| - | - | - | - | - | - | - | - | - | - | - | - | - | ||
| 2390 | + | 2 | 1357; 1128; 900; 689 | 1342; 1114; 900; 679 | 1357; 1128; 900; 827; 689 | 876; 668 | 1114; 900; 679 | 1342; 1114; 900; 818; 689 | 1142; 876; 658 | 912; 710 | 1157; 900; 700 | 1342; 1157; 900; 700 | 1357; 1128; 900; 710 | 1114; 900 |
| - | - | - | - | - | - | - | - | - | - | - | - | - | ||
| 2391 | + | 10 | - | - | - | - | - | - | - | - | - | - | - | - |
| - | 406 | 406 | - | 508; 406; 262 | 406 | 406 | 406 | 406 | 942; 406 | 406 | 406 | 508; 406 | ||
| 2392 | + | 6 | - | - | - | - | - | - | - | 319 | 319 | 326 | 319 | - |
| - | - | 566 | 519 | 519 | 519 | 519 | 519 | - | - | - | - | - | ||
| 2402 | + | 2 | 1300; 900 | 1350; 900 | - | - | - | - | - | - | - | - | - | - |
| - | - | - | - | - | - | - | - | - | - | - | - | - | ||
| iPBS Primer | M/H 1 | ± 2 | Control 3 | Experimental Groups | |||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 h | 2 h | 3 h | |||||||||||||
| 0.5 mM | 1 mM | 1.5 mM | 2 mM | 0.5 mM | 1 mM | 1.5 mM | 2 mM | 0.5 mM | 1 mM | 1.5 mM | 2 mM | ||||
| 2383 | M | + | 8 | 400 | - | - | 409 | - | - | 381 | - | - | - | - | - |
| - | - | - | 948 | - | 1150; 512 | - | 1150 | 1150; 512 | 1150 | 1150 | - | ||||
| H | + | 8 | - | - | - | - | - | 1375 | - | - | - | - | - | - | |
| - | 487; 419; 373; 321 | 1150; 487; 419; 373; 321 | 373 | 373 | 487; 373 | 373 | 419; 373 | 373; 321 | 373 | 373; 321 | 373 | 373 | |||
| 2384 | M | + | 6 | - | - | 437; 318 | - | 510; 424; 369; 325 | - | - | - | - | 510 | 510 | - |
| - | - | - | - | 923; 725; 632; 550 | - | 1260; 923; 725; 550 | 1260; 923; 725; 550 | 1260; 923; 725; 550 | 1260; 923; 725; 550 | 923 | 923 | 923; 725 | |||
| H | + | 5 | 923 | - | 780 | - | - | - | - | - | - | 923; 500 | - | - | |
| - | - | 1260; 713; 632 | - | 713; 561 | 1260; 713; 561 | 713; 561 | 1260; 713; 561 | 1260; 713; 632 | 1260; 713; 632 | - | - | 713 | |||
| 2385 | M | + | 5 | 565 | 548 | - | - | - | - | - | - | - | - | - | - |
| - | - | - | - | 845; 381 | - | 845; 381 | - | 381 | - | - | - | - | |||
| H | + | 7 | - | - | - | - | - | - | - | - | - | - | - | - | |
| - | 352 | 845; 582; 472; 409; 352 | 352 | 582; 472; 409; 352 | 845; 582; 409; 352 | 845; 582; 409; 352 | 845; 582; 409; 352 | 845; 582; 352 | 845; 582; 409; 352 | 582 | 582; 352 | 352 | |||
| 2386 | M | + | 2 | 964 | - | 964 | 1000 | - | 964 | 1000 | - | 1000 | 1033 | - | - |
| - | - | 614 | - | - | - | 614 | 614 | 614 | 614 | 614 | 614 | - | |||
| H | + | 2 | 1100; 931 | - | 1000 | - | - | 964 | - | - | - | 1000 | - | 1066 | |
| - | - | - | 614 | 614 | - | - | 614 | 614 | 614 | 614 | 614 | 614 | |||
| 2388 | M | + | 6 | - | - | - | - | - | - | - | - | - | - | - | - |
| - | - | 1333; 323 | - | 900; 323 | 1333; 323 | - | 1333; 900; 323 | 1333; 900; 323 | 1333; 323 | 323 | 1333; 900; 323 | 323 | |||
| H | + | 5 | 323 | - | - | - | - | - | 389 | - | - | 338 | 331 | - | |
| - | - | - | 1333 | - | - | 1333; 930 | 1333; 930 | 1333; 930 | 1333; 930; 618 | - | - | 1333 | |||
| 2389 | M | + | 5 | 900; 779 | 900 | 1333; 567 | - | 759 | - | - | 1300; 739 | 1300 | 553 | 1300; 739; 553 | 1366; 739; 567 |
| - | 443 | 1466 | - | 1466; 1033; 443 | - | - | - | - | 1466; 443 | - | - | - | |||
| H | + | 6 | 567 | 871 | - | 845 | 845 | - | - | - | 871 | 553 | 821; 539 | 1366 | |
| - | - | 1500; 1033; 428 | 428 | 1500; 428 | 1500; 1033; 779; 428 | - | 428 | 1500; 779 | 1500; 428 | 779 | 1500 | 779; 428 | |||
| 2390 | M | + | 8 | 1420; 900 | - | - | 1260 | - | - | - | - | - | - | - | - |
| - | - | 936 | 936 | 936 | 936 | 936 | 936 | 936 | 936 | 870 | 870 | 870 | |||
| H | + | 6 | 1460; 1120; 917 | 1440 | 1440; 1100 | - | 1460 | 1480 | 1460 | 1480 | 1480 | 1480 | 1500 | 1480 | |
| - | - | - | - | - | - | - | - | - | - | - | - | - | |||
| 2391 | M | + | 11 | - | - | 181 | - | - | - | - | - | - | - | - | - |
| - | - | 491 | - | 393 | 393 | 491 | - | 491 | 491 | 491 | 491; 393 | 491; 393 | |||
| H | + | 10 | - | 482 | - | 482 | - | - | 474 | - | - | - | - | - | |
| - | - | - | - | - | 610 | - | - | 400 | - | 400; 306 | 400 | 400 | |||
| 2392 | M | + | 5 | - | - | - | - | - | - | - | 627 | 620 | - | - | - |
| - | 670 | 670; 545; 462 | 670; 462 | - | 732; 462 | - | 732; 462 | 732 | 732 | 732; 462 | 732; 545; 462 | 732; 462 | |||
| H | + | 4 | - | - | - | - | - | - | - | - | 627 | 634 | - | - | |
| - | 421 | 421 | 526; 421 | 421 | - | - | - | 526; 421 | - | - | 421 | - | |||
| 2402 | M | + | 6 | - | - | 437; 318 | - | 510; 424 | - | - | - | - | 510 | 510 | - |
| - | - | - | - | 1260; 923; 725; 632; 550 | - | 1260; 923; 725; 550 | 923; 725; 550 | 923; 725; 550 | 1260; 923; 725; 632 | 923 | 923 | 923; 725 | |||
| H | + | 5 | 923 | - | 900 | - | - | - | - | - | - | - | 510 | - | |
| - | - | 1260; 713; 561 | - | 713; 561 | 1260; 713; 561 | 713; 561 | 1260; 713; 561 | 1260; 713; 632 | 1260; 713; 632 | 923; 500 | 713 | 713 | |||
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Türkoğlu, A.; Haliloğlu, K.; Tosun, M.; Szulc, P.; Demirel, F.; Eren, B.; Bujak, H.; Karagöz, H.; Selwet, M.; Özkan, G.; et al. Sodium Azide as a Chemical Mutagen in Wheat (Triticum aestivum L.): Patterns of the Genetic and Epigenetic Effects with iPBS and CRED-iPBS Techniques. Agriculture 2023, 13, 1242. https://doi.org/10.3390/agriculture13061242
Türkoğlu A, Haliloğlu K, Tosun M, Szulc P, Demirel F, Eren B, Bujak H, Karagöz H, Selwet M, Özkan G, et al. Sodium Azide as a Chemical Mutagen in Wheat (Triticum aestivum L.): Patterns of the Genetic and Epigenetic Effects with iPBS and CRED-iPBS Techniques. Agriculture. 2023; 13(6):1242. https://doi.org/10.3390/agriculture13061242
Chicago/Turabian StyleTürkoğlu, Aras, Kamil Haliloğlu, Metin Tosun, Piotr Szulc, Fatih Demirel, Barış Eren, Henryk Bujak, Halit Karagöz, Marek Selwet, Güller Özkan, and et al. 2023. "Sodium Azide as a Chemical Mutagen in Wheat (Triticum aestivum L.): Patterns of the Genetic and Epigenetic Effects with iPBS and CRED-iPBS Techniques" Agriculture 13, no. 6: 1242. https://doi.org/10.3390/agriculture13061242
APA StyleTürkoğlu, A., Haliloğlu, K., Tosun, M., Szulc, P., Demirel, F., Eren, B., Bujak, H., Karagöz, H., Selwet, M., Özkan, G., & Niedbała, G. (2023). Sodium Azide as a Chemical Mutagen in Wheat (Triticum aestivum L.): Patterns of the Genetic and Epigenetic Effects with iPBS and CRED-iPBS Techniques. Agriculture, 13(6), 1242. https://doi.org/10.3390/agriculture13061242










