Abstract
Methanogenesis plays a crucial role in the digestive process of ruminant animals. During this process, methanogenic archaea produce methane as a byproduct of their metabolism. However, the production of methane by ruminants is also a significant contributor to greenhouse gas emissions. Methane is a potent greenhouse gas that has a 28-fold greater global warming potential than carbon dioxide. Around 15% of all anthropogenic greenhouse gas emissions are generated by livestock. Therefore, reducing methane emissions from ruminant livestock is an important goal for reducing the environmental impact of agriculture. There is a variety of strategies that can be used to reduce methane emissions, including dietary modifications, genetic selection, microbiome manipulation, and feed additives, such as plant secondary metabolites, methane inhibitors, lipids, essential oils, and algae. The main and important objective of this paper is to critically discuss the current strategies proposed to reduce methane emissions from livestock. Though many strategies, such as chemical intervention, have remarkably reduced methane emissions from ruminants, their usage remains unappealing because of health and safety concerns as well as consumer decisions. Hence, genetic animal selection and biological feed additives, such as probiotics and secondary plant metabolites, have emerged as promising techniques for mitigating enteric methane emissions. These strategies are highly promising, but more intensive research is needed to validate these approaches and assess their effectiveness in reducing methane production by ruminants.
1. Introduction
In recent years, there has been a growing interest in the greenhouse effect, which is linked to rising global temperatures. Carbon dioxide (CO2), methane (CH4), and nitrous oxide (N2O) are the three most significant greenhouse gases (GHGs). They absorb solar heat and warm the atmosphere by absorbing energy and slowing the rate at which it escapes to space. This increase has serious consequences for humans, animals, and the environment [1,2].
Methane is the second most significant GHG after CO2 and absorbs more energy than CO2 [3]. Furthermore, CH4 has a 28-fold greater global warming potential (GWP) than CO2 over a 100-year period. It also accounts for some indirect effects, as a precursor to ozone, another GHG compound [4]. Rising atmospheric CH4 concentrations are linked to rising human populations, feed production, and anthropogenic sources account for two-thirds of the CH4 production [3,4].
The enteric fermentation process, paddy rice production, and animal waste are significant sources of CH4, contributing about 40% of agriculture’s CH4 emissions [4,5,6]. Domesticated animals, such as cattle, sheep, and goats, naturally produce CH4 during their physiological digestive processes. Cattle and dairy cows were the biggest contributors to CH4 emissions in 2020, accounting for 72% of total sector emissions [7,8]. Buffalos and small ruminants demonstrated only 8.7% and 6.7% of CH4 emissions, respectively [9]. Methane has a shorter lifetime in the atmosphere with a half-life of 8.6 years compared to CO2 [10]. This characteristic makes it a desirable target for mitigating global warming in the short term, as reducing its emissions can lead to immediate benefits. Therefore, researchers and breeders all over the world have been focusing on both increasing animal production and decreasing CH4 emissions from animals, particularly ruminants. In developed countries, it is recommended to cull nonproductive and low-producing animals to reduce CH4 output. This is impractical and difficult in developing countries due to their economic, cultural, and religious backgrounds [5]. Moreover, producers of ruminants must find cost-effective and efficient ways to lower CH4 emissions while still meeting consumer demand for food and health as well as environmental demands.
According to a recent report by the United Nations Environment Programme (UNEP), global methane emissions increased by 9% from 2000 to 2017 [11]. This trend is expected to continue, especially in countries with developing economies. However, the reduction in emission intensity resulting from production efficiency improvements is small (less than 1% per year) and may not be enough to counterbalance the increasing emissions due to the growing demand for animal protein [12]. Therefore, a global effort is necessary to reduce ruminant emissions.
During anaerobic fermentation, CH4 is produced in the rumen and hindgut. Many anaerobic and facultative anaerobic bacteria live in the rumen, forming a large and diverse microbial population. The most dominant bacteria in the rumen are Prevotella, Butyrivibrio, and Ruminococcus, as well as unclassified Lachnospiraceae, Ruminococcaceae, Clostridiales, and Bacteroidales [13,14]. However, the bacterial community structure can be affected by changes in the host’s diet. When animals are fed high-forage diets, the rumen bacterial composition is primarily Gram-negative; in animals fed high-grain diets, more Gram-positive bacteria, such as Lactobacillus, are present [15]. The rumen is also colonised by a diverse population of anaerobic fungi, protozoa, and flagellates, which degrade many polymeric components of the feed into small molecules, such as amino acids and simple sugars [16]. Bacteria and protozoa make up 80% of the microbial mass inside the rumen, which produces a CH4 [2]. Aside from CH4, NH3, CO2, and H2 are produced during fermentation and removed from the rumen by eructation, resulting in a loss of 2–12% of the host’s metabolic energy intake [17].
Methanogens are a group of microorganisms that live in a variety of habitats; they have a wide range of morphology and physiological parameters and produce CH4 as a metabolic byproduct. They belong to the domain Archaea, with minor amounts originating from cyanobacteria and marine microorganisms [18], and are classified into three clades found in the digestive tracts of ruminants based on the substrate used: CH4 derivatives (methylotrophic); H2/CO2 (hydrogenotrophic); and acetate (acetoclastic) [19]. Many methanogens are also highly adaptable and can generate CH4 from a variety of substrates.
In an anaerobic environment, all of the major methanogens in the rumen use H2/CO2, with CO2 acting as the primary H2 sink [20,21]. However, nitrate (NO3−) and sulphate (SO42−) can also act as H2 sinks in addition to CO2 [22]. Despite the low concentration of NO3−/SO42− in the rumen, making it more thermodynamically favourable, the rate of electron flow towards the SO42−/NO3− reduction pathway diverts most of the H2 towards the CH4 formation [23]. The most effective way to remove H2 from the rumen and keep the fermentation process going is methanogenesis. Rumen methanogens are, therefore, a crucial target for CH4 mitigation techniques.
To reduce CH4 emissions in ruminants, various farming practices, dietary modifications, feed additives, chemical methanogenesis modulators, probiotics, immunisation against the rumen microbiome, selective breeding, and genetic approaches are currently being used. The direct and indirect inhibition of methanogens by feed additives plays a significant role in these strategies [24,25,26].
There were almost 9000 papers on rumen methanogenesis published in the literature between 1960 and 2018 [12]. Our goal is to critically analyse the current knowledge and prospects for CH4 mitigation, not to cite the results of individual papers reviewed many times. We are particularly interested in approaches, open questions, and issues that affect researchers, farmers, and the ruminant livestock industries. It is also important to evaluate CH4 mitigation strategies in terms of total GHG and to consider the rationality and possibility of various strategies according to ever-increasing total CH4 emissions. A more basic understanding of the potential strategies, their flaws, and their application would enable us to exploit their potential to decrease CH4 emissions without negative effects on animal and human health, as well as livestock production efficiency.
2. Methanogenesis
Methanogenic archaea are anaerobic microorganisms that produce CH4 for ATP synthesis. There are approximately 155 species that have been identified in nature, divided into 29 genera, 14 families, 6 orders, and 4 classes [27]. Most exist freely in rumen liquid or biofilms adhering to feed particles [28,29], while a small portion is symbionts with protozoa [30]. Methanobrevibacter (Mbb) is the most common rumen methanogen, accounting for 63.2% of all isolates, followed by Methanosphaera (9.8%) and Methanomicrobium (7.7%) [31,32,33,34], while the rest belong to the minority genera, such as Methanimicrococcus, Methanosarcina and Methanobacterium [35].
According to previous research, hydrogenotrophic methanogens dominate in terms of CH4 emissions and community composition [14,36]. Because of their limited substrate range, rumen archaea appear to be much less diverse than rumen bacteria. Henderson et al. [14] discovered that Mbb gottschalkii and Mbb ruminantium account for 74% of the rumen methanogen community in samples from a variety of animals, locations, diets, and environmental conditions. To date, only 13 species of ruminal methanogens have been isolated into pure cultures: Mbb formicicum; Mbb bryantii; Mbb ruminantium; Mbb millerae; Mbb olleyae; Mbb boviskoreani; Methanomicrobium mobile; Methanoculleus olentangyi; Methanosarcina barkeri; Methanobacterium beijingense; Methanoculleus marisnigri; Methanoculleus bourgensis; and Methanosarcina mazei [32,34,37,38,39].
Fermentation processes generate the H2 molecule, which serves as the primary substrate for ruminal methanogenesis. Methanobrevibacter and Methanomicrobium grow in anaerobic conditions at optimal temperatures of 39 to 40 °C, using H2 to reduce CO2 to CH4 via the Wolfe cycle of methanogenesis, according to the stoichiometric equations: 4H2 + CO2 → CH4 + 2H2O and HCO3− + 4H2 → H+ CH4 + 3H2O [40,41]. Thus, they maintain a low H2 partial pressure and limit the hydrogenotrophic acetogenesis [41,42,43]. This pathway is most important and is responsible for about 82% of all CH4 production in the rumen [44]. These methanogens can also use formate as an electron donor [45]. During formatotrophic methanogenesis, four formate molecules are oxidised by formate dehydrogenase to form CO2 according to the stoichiometric equation 4HCOOH → 3CO2 + CH4 + 2H2O [43].
Methanogens, such as Methanosarcinales and Methanobacteriales, oxidise methyl groups, such as methylamines and methanol, into CO2 to obtain electrons used to reduce methyl groups to CH4. This is described by the stoichiometric reaction shown below. 4CH3OH → 3CH4 + CO2 + 2H2O; CH3NH2 + H2 → CH4 + NH3; CH3OH + H2 → CH4 + H2O [32,44,46]. Furthermore, Methanosarcina and Methanosaeta use acetate as the substrate during acetoclastic methanogenesis, [47,48]. Acetate splits to form carboxyl compounds which are oxidised to CO2. The stoichiometric equation states that methyl groups enter the hydrogenotrophic pathway to form CH4: CH3COOH → CO2 + CH4. Nonetheless, methyl groups and acetate play a minor role in rumen methanogenesis because methanogens that rely on these conversions grow very slowly in vitro. As a result of their short retention times, normal rumen conditions would prevent them from thriving [30].
Methane is not the only byproduct of the rumen fermentation of H2. Hydrogen was used by non-methanogenic rumen inhabitants to synthesize acetate and propionate. These bacteria also consume CO2, SO42−, NO3−, and fumarate and compete with methanogens for H2 [41,47,48]. However, they are less important in removing H2 from the rumen ecosystem.
Acetobacterium, a genus of anaerobic bacteria found in the rumen fluid, can convert two CO2 molecules to acetate (4H2 + 2CO2 → CH3COOH + 2H2O) using the reductive acetyl-CoA pathway (Wood-Ljungdahl pathway) [49].
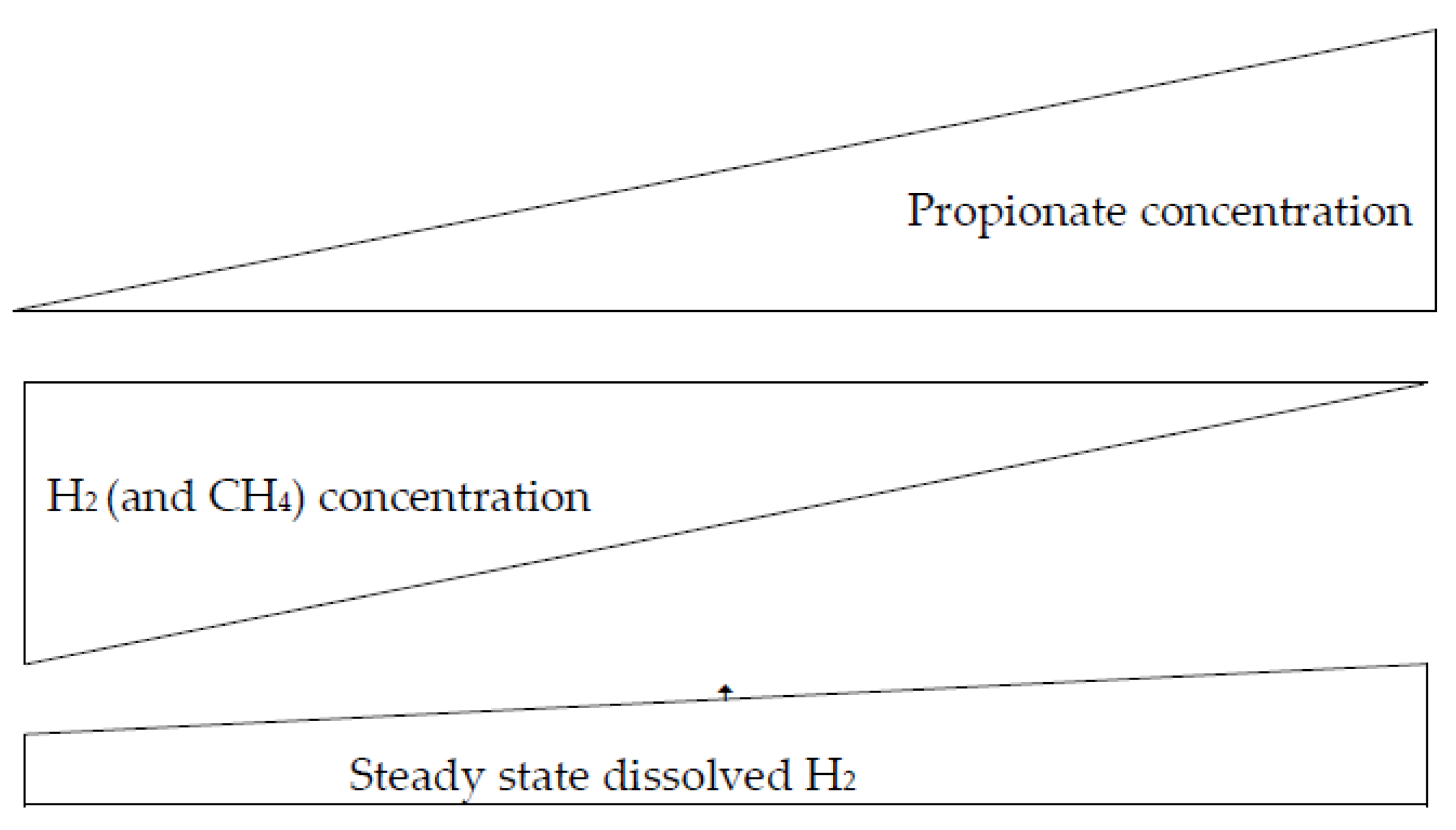
During propionogenesis, H2 in the rumen is redirected to propionate synthesis via the fumarate-succinate pathway or the conversion of pyruvate to lactate and acrylyl-CoA ester and the subsequent reduction to propionate (pyruvate + 4H → propionate (C3) + H2O) [50]. There is an inverse relationship between the amount of propionate in the rumen and the production of CH4 in the rumen; when the concentration of propionate increases, the production of CH4 in the rumen decreases, and vice versa (Figure 1) [21,44,51,52].
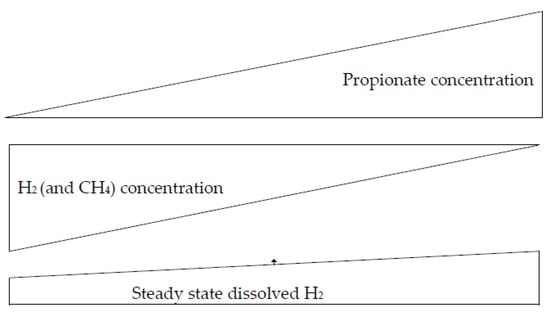
Figure 1.
Relationship between H2, CH4, and propionate concentrations, according to Janssen [18].
The mechanism of NO3− reduction in the rumen is another alternative hydrogen-based pathway. Under anaerobic conditions, NO3− can be reduced in three ways: dissimilatory NO3− reduction to nitrogen (N2) (denitrification); and dissimilatory or assimilatory NO3− reduction to NH3 [53]. However, under physiological conditions, denitrification, in which NO3− is reduced to nitrite (NO2−), then to nitric oxide (NO), further to N2O, and finally, to N2, does not play a significant role in the rumen. In the case of the other two mechanisms, dissimilatory NO3− reduction to ammonia is the dominant rumen pathway of NO3− metabolism described with the following stoichiometric reactions:
NO3− + H2+ + 2H+ → NO2− + 2H2O and NO2− + 3H2 + 2H+ → NH4+ + 2H2O.
The conversion of NO3− to NH3 is thermodynamically more favourable to methanogenesis; however, if sufficient NO3− concentration is available, these processes could be the primary route of H2 elimination in the rumen environment [54,55,56]. Unlike dissimilatory NO3−reduction, NH3, a product of enzymatic respiratory ammonification, remains in the organism to allow microbial protein synthesis [57]. Notably, high NH3 concentrations inhibit assimilatory NO3− reduction; thus, this mechanism is unimportant in the rumen, and microorganisms instead use this pathway for biosynthesis and storage [43].
In some anaerobic environments, hydrogen sulphide can act as an electron donor for the reaction that converts NO3− to NH3 (by reducing NO3− and oxidising sulphides), which increases the H2 utilisation [8,21,44]. The rumen fermentation process also uses SO42− as an alternative electron acceptor. Sulphate has a high reduction potential and serves as an H2 absorber because it has a higher affinity for H2 than CO2 [8].
Bacteria that reduce NO3− can also reduce SO42− through dissimilatory or assimilatory pathways. Both types of bacteria co-exist in the rumen, where they contribute to the reduction of sulphur to hydrogen sulphite (HSO3−) and hydrogen sulphide (H2S) rather than CH4 [58]. Furthermore, both pathways operate concurrently and are not competitive [44]. Sulphate-reducing bacteria reduce SO42− to H2S anaerobically and oxidise organic matter to H2 using sulphate ions as electron acceptors. This reaction is described by the stoichiometric equation (SO42− + 4H2 + H+ → HS− + 4H2O) [8]. However, the rumen SO42− reduction process is severely hampered by the low concentration of various sulphur-containing compounds, which limits the rate of H2 incorporation in their reduction, unless they are supplemented to the diet [44]. Notably, a dissimilatory pathway is used for energy generation, whereas synthesised sulphur compounds are incorporated into biological molecules necessary for bacteria’s survival during assimilatory reduction. Furthermore, a dissimilatory reduction is the primary route of SO42− metabolism in the rumen [59]. Some sulphate-reducing bacteria are adaptable organisms that can both reduce SO42− and oxidise H2S to SO42−, preventing the accumulation of toxic molecules [47,48].
3. Mitigation Strategies
Reduced ruminant enteric CH4 emissions are an important goal for limiting the global temperature rise over the next 30 years. Scientists and breeders are working to identify various strategies that affect methanogenesis in the rumen. As new scientific techniques have been developed, research into methanogenesis has expanded exponentially, and our understanding of the rumen microbiota has grown in depth.
Furthermore, beginning in the early 2000s, we saw a shift in research focus from energy metabolism to the environmental impact of ruminant CH4 and developing strategies for reducing enteric CH4 emissions. Since then, several approaches and technologies have proven to be effective in reducing enteric CH4 emissions. Some, however, are prohibitively expensive, not only for developing countries, and pose environmental and human or animal health risks. As a result, it is critical to understand how and whether new techniques and solutions reduce ruminant CH4 emissions.
Several approaches have been developed to reduce ruminant digestion-related CH4 emissions. Some of them are farm management, feeding strategies, using feed additives and chemical compounds to modify biochemical processes in the rumen, direct manipulation of the rumen microflora, animal immunisation, and breeding strategies, including genetic approaches [3,60]. The most commonly used feed additives that modify rumen biochemical processes towards decreased methanogenesis are saponins, tannins, flavonoids, probiotics, organic acids, fats, and fibre [61,62,63].
3.1. Mitigation through Feed Additives
3.1.1. Saponins
Saponins are surface-active glycosides composed of aglycone-sapogenin and glycon-saccharide. Plants, lower marine animals, and some bacteria are the primary producers [60,64]. The major sources of saponins in ruminant diets are Yucca schidigera, Quillaja saponaria, Camellia sinensis, and Medicago sativa [65].
Saponin structure diversity associated with chemical groups, such as hydroxyl, hydroxymethyl, carboxyl, and acyl in sapogen, causes changes in the bioactivity [64]. Saponins have a significant impact on the rumen ecosystem because they inhibit the population of ciliate protozoa. Ciliate protozoa contribute to proteolysis, particularly ruminal protein recycling and protein passage to the duodenum [66,67]. Furthermore, saponins have the potential to reduce CH4 emissions indirectly through defaunation. The mechanism of defaunation by saponins is thought to involve disruption of the cell membranes of protozoa in the rumen. Saponins have a characteristic structure consisting of a hydrophilic sugar moiety and a hydrophobic steroid or triterpenoid aglycone. This structure allows saponins to form complexes with sterols in cell membranes, leading to membrane destabilisation and lysis of the cell [62,68].
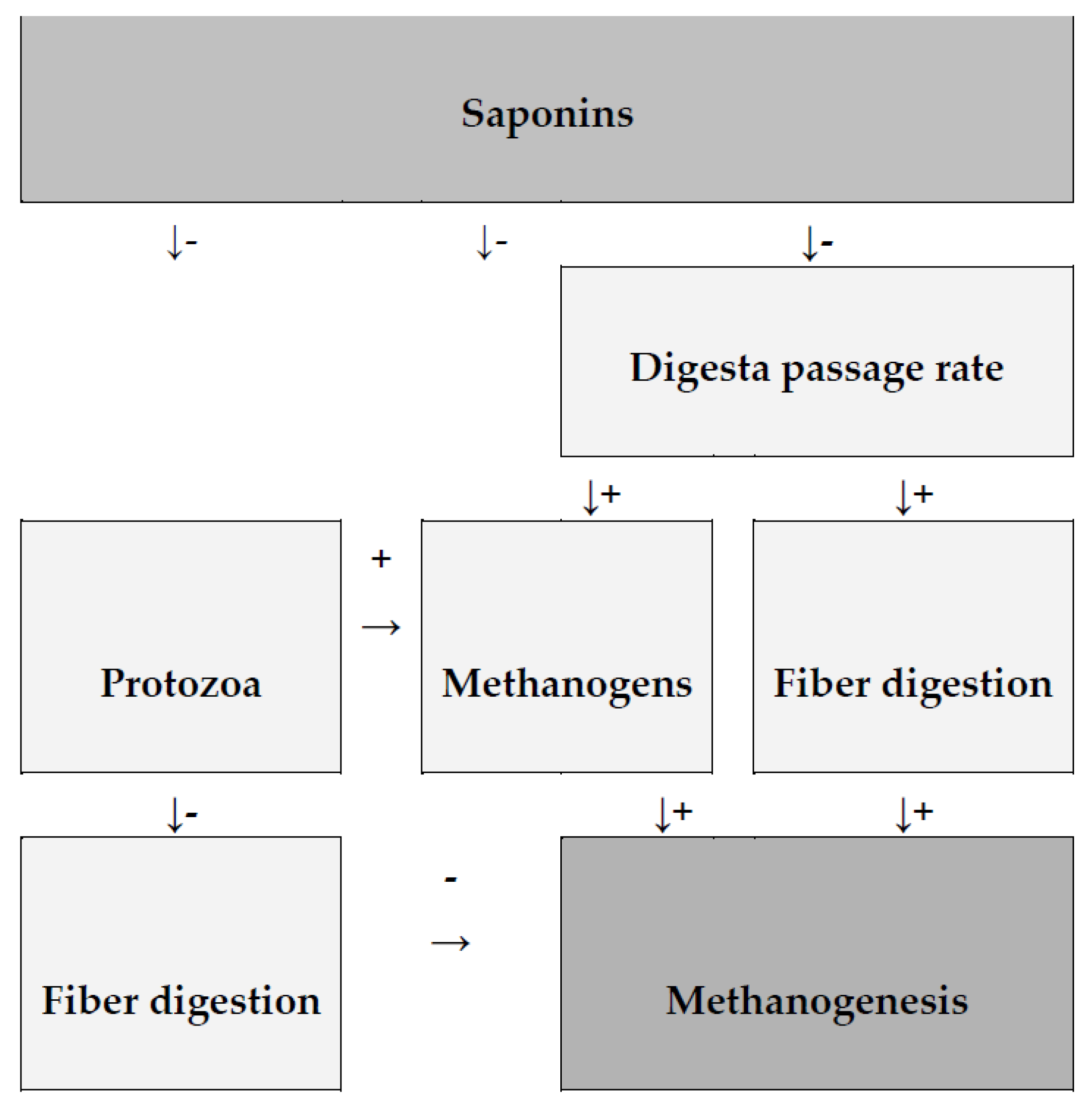
Saponin may also have an indirect effect on CH4 output by slowing methanogenesis, increasing the expression of methanogenesis-related genes, and decreasing the abundance of methanogens. Saponins can affect specific bacteria and fungi in the rumen, altering biochemical processes in the rumen either positively or negatively [62,64]. At this time, there is no clear evidence that saponins directly affect methanogens, but it cannot be ruled out. Nonetheless, some research suggests that saponins may reduce the activity of methanogenesis-related genes or the rate of CH4 production per methanogen cell (Figure 2) [62].
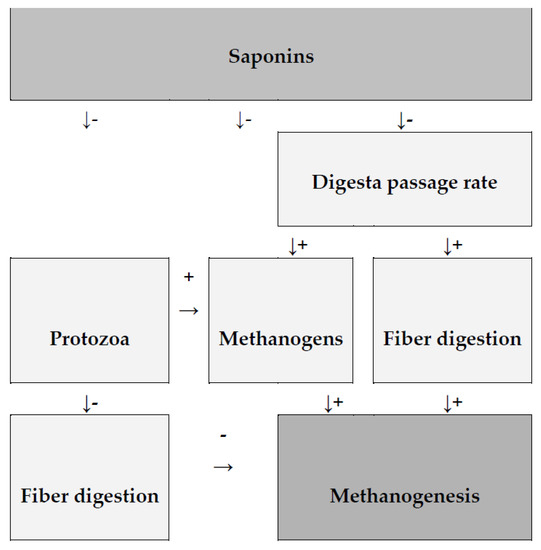
Figure 2.
Diagram shows the impact of saponins on methanogenesis as well as the effects of secondary rumen fermentation modification. (+) increase; (−) decrease, according to Patra and Saxena [62].
Despite the fact that many in vitro studies have used relatively high dose rates, high concentrations of saponins (>5% of dry matter (DM)) and steroidal saponins from certain plants can be toxic to animals. The ability of saponins to suppress CH4 production varies greatly depending on the chemical structure, source, dose, and diet [5]. Furthermore, the type and origin of saponins alter rumen fermentation differently. Methane production was strongly inhibited by Yucca schidigera but not by Quillaja saponaria extract [69]. Saponins isolated from different sources show various results, and most of the results come from in vitro studies [70]. The impact of saponins on the abundance of methanogens and methanogenesis varies considerably across studies, and data are inconsistent [46]. Methane emissions were found to have decreased anywhere between 6 and 50%, depending on the plant source, saponin dose, saponin extraction method, saponin preparation, etc., and combinations of these parameters [46,70].
The antiprotozoal effect of saponins is temporary because they are deemed inactive by rumen microorganisms when they are deglycosylated to sapogenin [59]. This might restrict their use as ruminant feed and as a tool to reduce methane emissions. It was recently proposed that saponins’ antiprotozoal effect could be improved by changing their chemical structure, thereby protecting them from microbial degradation [64].
It is important to note that the level of saponin toxicity in plants can vary depending on several factors, such as the species of the plant, its growing conditions, and the stage of growth. It was well documented that in several plant species, saponin synthesis is induced in response to biotic and abiotic stresses. Such as herbivory and pathogen attacks, humidity, nutrient starvation, light, and temperature [71]. To minimize the risk of saponin toxicity in cattle, it is important to monitor the plants that are present in their grazing areas and to provide a balanced and nutritious diet that meets their dietary needs. So, to minimize the potential negative effects of saponins in animal feed, it is important to carefully select feed ingredients and properly process them to reduce the concentration of saponins [72]. This may include using heat treatment, chemical treatment, or other methods to break down or remove the saponins. For example, fermented soybeans or processed soybean meal have lower saponin levels than raw soybeans [72]. In cases where exposure to high levels of saponins is unavoidable, it is recommended to monitor the animals closely for any signs of toxicity and to provide prompt veterinary care if necessary.
3.1.2. Tannins
Tannins are another class of organic chemicals that affect the rumen environment. They are secondary polyphenolic plant metabolites. They have been linked to plants’ resistance to infection, insects, and herbivores [73]. Tannins are classified into two types: hydrolysed tannins (polyesters of gallic acid and various individual sugars) and condensed tannins (polymers of flavonoids) [74,75].
When hydrolysed tannins are exposed to the diluted acids in the feed, they degrade into simpler phenolic and non-phenolic compounds. The condensed fraction influences feed digestibility, modifies rumen fermentation, and reduces CH4 emissions [73]. Moreover, condensed tannins have a higher antimethanogenic potential, but hydrolysable tannins are more toxic to animals after absorption [76,77].
In general, tannins have been shown to reduce CH4 production in the digestive systems of ruminants by indirect inhibition of hydrogen-producing microflora (and thus, presumably reduced forage digestibility) and direct inhibition of methanogens [61,75]. Supplementation of the tannin-rich plants reduced CH4 emissions per day and per unit of feed and energy intake by up to 24%, but this appears to be primarily to reduced organic matter and fibre digestion [78]. In their valuable review, Jayanegara et al. [79] demonstrated that increasing tannin levels reduced CH4 generation expressed relative to digestible organic matter. Patra and Saxena [62] reported a CH4 reduction in ruminal fermentation by 50% in response to tannin or plant extracts containing these polyphenolic compounds at an average tannin dose of 10–20 g/kg DM. Moreover, Aboagye and Beauchemin [80] suggest that CH4 emission in response to tannin feeding varies greatly depending on the tannin source, type, molecular weight, and the methanogen community present in the rumen.
Tannins inhibit methanogenesis in a bactericidal and bacteriostatic manner by acting on fibrinolytic bacteria and are dependent on their chemical structure as well as the bacteria species [74,81]. In general, the anti-methanogenic activities of tannins involve the binding of tannins to proteins through the interaction of phenolic hydroxyl groups with amino acid residues by hydrogen bonds and hydrophobic interactions; however, low molecular weight tannins could be more effective [76,81,82]. Tannin–protein binding can have a range of effects on protein structure and function, including denaturation, aggregation, and modification of the enzymatic activity [83,84].
Furthermore, they are toxic to ciliate protozoa, which are known to produce a significant amount of CH4 through interactions with archaea [8,61,74]. Malik et al. [85] showed in sheep that diets supplemented with condensed tannin in concentrations of 7.1–10.8 g/kg DM decreased the number of protozoa by up to 23%. A similar reduction in protozoa number (21%) was reported by Salami et al. [86], including 4% of both condensed and hydrolysed tannins in lamb’s diet.
Some tannins decrease fungi but simultaneously increase methanogens [87]. Furthermore, another major limitation of tannins is that CH4 responses highly depend on their concentrations in many forages and feed supplements (≤20 g/kg of dry matter intake, (DMI)). Additionally, dietary tannins can reduce protein digestibility [88]. This is especially important for grazing animals in nitrogen-deficient pastures [89]. The decrease in CH4 caused by tannins could be due to a concurrent reduction in DMI and nutrient digestibility.
It should be noted that rumen microbiota can develop resistance to the adverse effects of tannins through detoxification of tannins, synthesis of tannin-complexing polymers, and formation of glycocalyx [90]. This may hinder the long-term use of tannins to reduce methane emissions.
3.1.3. Flavonoids
Flavonoids are a class of compounds that, similar to tannins, are secondary plant metabolites with a wide range of biological activity, such as antimicrobial activity [46,91]. Probably, flavonoids reduce the population of protozoa and methanogens and inhibit methanogenesis in the rumen by absorbing H2 after the cleavage of their carbon ring structures [34,91]. Flavonoids have been proposed for use in ruminant feed to increase productivity by elevating propionate production relative to the acetate [92]. The flavonoids naringin and quercetin were found to reduce CH4 production, ciliate protozoa, and hydrogenotrophic methanogens in vitro [91,93]. Seradj et al. [94] discovered that a commercial citrus extract of flavonoids (Bioflavex®) or its pure flavonoid components (Neoeriocitrine, Naringine, Isonaringine, Hesperidine, Neohesperidine, Poncirine) in the dose of at 200 μg/g DM decrease CH4 production from 4 to 11% and hydrogenotrophic methanogen (Methanosarcina spp.) population while increasing propionate concentration and lactate-consuming Megasphaera elsdenii population. Megasphaera elsdenii metabolizes lactate and relieves rumen acidosis induced by a high-grain diet [95]. In an in vitro study, Sinz et al. [96] discovered that the flavonoid luteolin-7-glucoside, in a dose-dependent manner (0.5, 5.0, and 50.0 mg/g DM), reduced CH4 (4–11%) and NH3 (12%) concentrations in rumen fluid without lowering fermentation efficiency.
There are currently several flavonoid feed additives on the market that reduce CH4 production. However, these products primarily contain plant crude extracts, and determining the response of rumen microbes to the flavonoids is difficult. Moreover, significant differences in the extraction yields between different high-cost conventional extractions can be observed. There are also differences between the number of compounds obtained and their bioactivity within the same method [97]. Nowadays, non-conventional low-cost extraction techniques using microwaves, ultrasounds, high pressure, supercritical fluids, or digestive enzymes are involved in the effort to extract the flavonoids in a more efficient and/or selective way. Moreover, these techniques decrease the extraction time, increase the compounds’ selectivity and allow the use of solvents that are less harmful to the environment and animal health [97]. Therefore, some of these techniques are green methods, most acceptable to consumers and farmers. In conclusion, flavonoids have a significant potential to mitigate CH4 emissions, according to available data, but more research on in vivo trials is needed.
3.1.4. Lipids
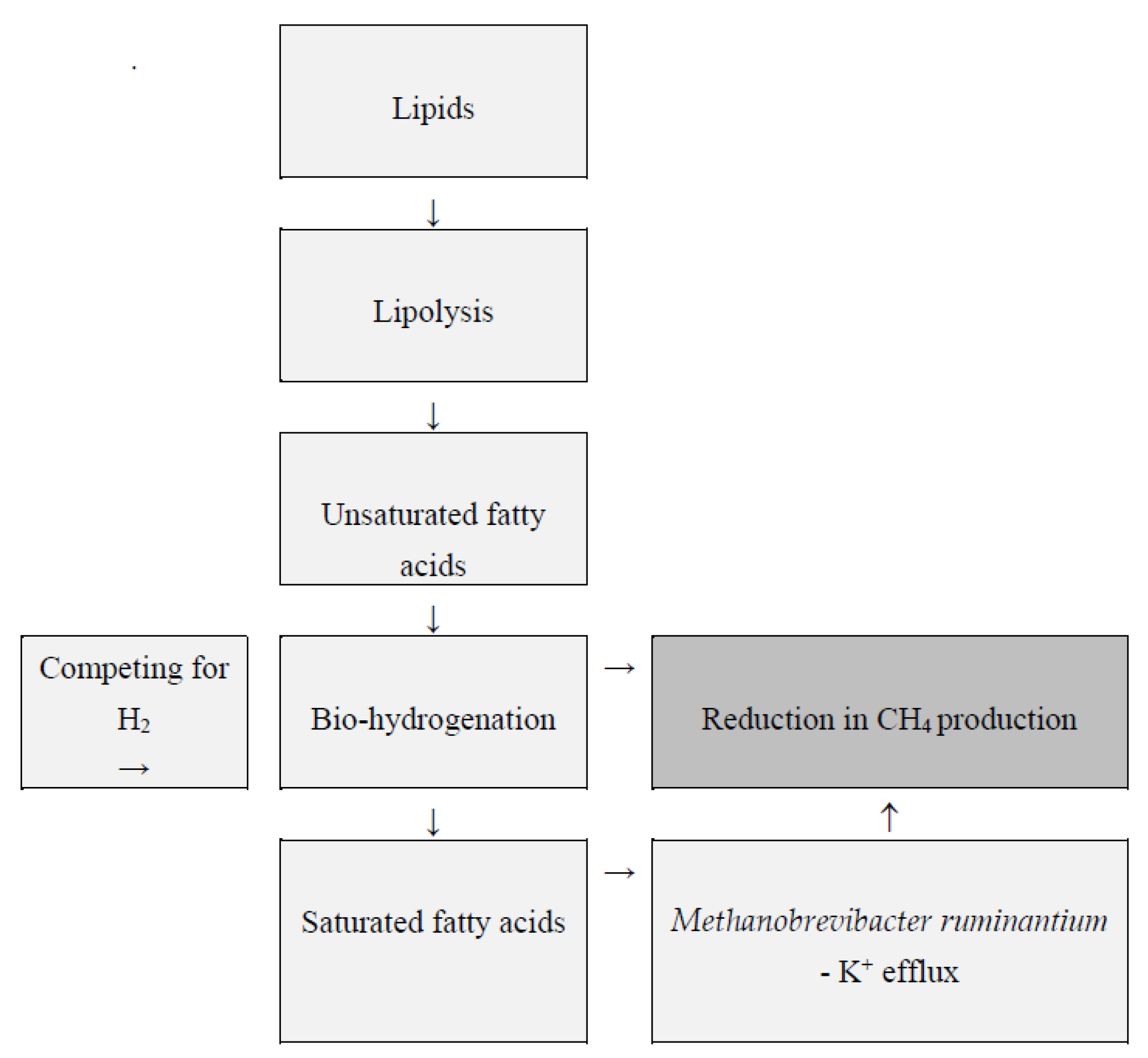
Lipids used as feed additives may be a viable strategy for lowering CH4 emissions (Figure 3) [34]. They reduce the concentration of rumen archaea and Ciliate protozoa through several mechanisms, including toxicity against methanogens and protozoa [98]. However, for some fats, lowering CH4 levels may come at the expense of diet digestibility [42]. Lowering the pH of the rumen content to 5–6 can increase the effectiveness of lipid supplements in reducing methanogenesis [99]. Fatty acids have the potential to damage archaeal cell membranes. The integrity of the archaeal plasma membrane is critical for maintaining chemiosmotic balance, which is required for cell vitality and metabolism. Changes in the structure of the cell membrane cause potassium leakage, electron transport, ion gradients, protein translocations, phosphorylation, and enzyme reactions, which lead to methanogen degradation and CH4 inhibition [5].
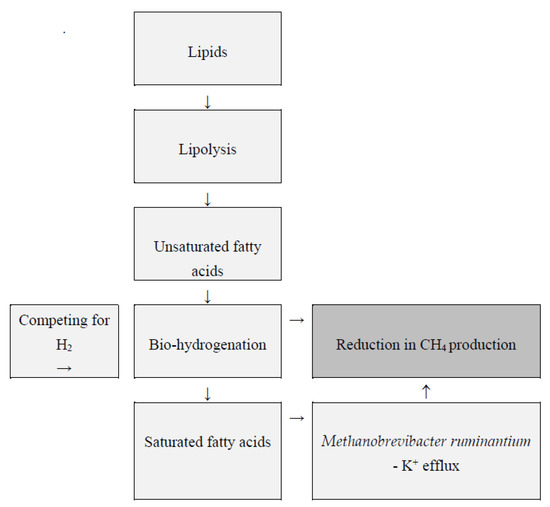
Figure 3.
Rumen lipid metabolism and CH4 production according to Zhou et al. [100] and Klevenhusen et al. [101].
Lipid supplementation could be an effective CH4 mitigation strategy (an average 14% reduction in long-term CH4 emissions when lipid sources were added to supply 34 g fat/kg of DM [102,103]. However, its efficacy is determined by the form, source, amount of supplemental fat, degree of saturation, fatty acid carbon chain lengths, and nutrient and fatty acid composition of the basal diet [104,105]. Dietary lipids have a greater inhibitory effect on CH4 emissions with concentrate-based diets than with forage-based diets, so the application of this supplementation in grazing systems is limited [104,106].
Feeding a high lipid concentration (>6% DM) can reduce feed and fibre digestibility, potentially increasing organic matter and nutrient excretion as well as CH4 emissions from manure [107]. Furthermore, lipid supplementation has been shown to reduce fibre digestibility and DMI, inhibit rumen fermentation, reduce milk fat synthesis, and alter the fatty acid composition of products [108]. Oils supplementation tended to decrease feed intake (from 4% to 7%) and decrease fibre digestibility (from 8% to 11%). Similar to oils, oilseeds did not affect milk yield but decreased weight gain in growing animals (from 13% to 20%) [109]. Therefore, including oils and oilseeds in the diet may be recommended more for lactating animals than for growing animals.
The benefit of lipid supplements is that they are readily available and straightforward to implement in intensive or confined feeding systems. Moreover, their supplementation is not known to pose a risk to animal or human safety; it is widely available and simple to implement in intensive or confined feeding systems. Moreover, maximum oil inclusion in ruminant diets is determined by the animal’s physiological stage, the composition of the basal diet’s lipids and other nutrients, and the fatty acid profile of the supplemental oil [110]. Feeding refined oils is more expensive than feeding processed oilseeds, but the overall cost-effectiveness of feeding oils to reduce CH4 varies by region and country because the prices of oil, meat, and milk vary greatly [109].
The impact of oil production on GHG emissions should be considered in the case of the widespread use of oils as feed additives. When compared to other concentrate feeds, oilseed production is associated with nearly doubling upstream GHG emissions per kg dry matter (1.27 vs. 0.70 CO2 equivalents kg−1 of DM). It is, therefore, critical to consider whether total GHG will increase when concentrates are replaced by oil and fat or whether the reduction in intestinal CH4 emissions will offset GHG emissions from the oil industry.
3.1.5. Plant Essential Oils
Plant essential oils (EO) are volatile and aromatic oily liquids extracted from plant materials such as flowers, seeds, buds, leaves, herbs, wood, fruits, twigs, and roots. Numerous in vitro studies have supported using essential oils derived from plants such as garlic, eucalyptus, clove, rosemary, thyme, paprika, juniper, and ginger. EOs are popular among consumers due to their high volatility, ephemerality, and biodegradability [111]. Some essential oils have been approved as generally safe by Food Drug Administration (FDA) [112]. Some essential oils are capable of affecting the rumen fermentations and decreasing CH4 production in vitro. The results were promising because some of the studies showed impressive results on CH4 emissions, with up to 90% reductions [5,113,114]. Belanche et al. [115] did not show the effect of oregano oil on enteric CH4 emissions when fed (50 mg/kg DM) to dairy cows. Benchaar et al. [116] showed that ruminal microorganisms might adapt to EOs when they are used for a long time, which could diminish their positive effects. Saro et al. [117] found that anti-methanogenic garlic essential oil and linseed oil treatments had no long-term effects on CH4 production in neonatal lambs. Moreover, in vivo studies in ruminants are likely to negatively affect feed intake and ruminal fermentation [118,119]. Furthermore, a large part of the positive effects mentioned during in vitro studies has not yielded the same positive results when applied to in vivo studies and using high doses in ruminants (i.e., >300 mg/L of culture rumen fluid), as would be expected [119].
In general, essential oils appear to work by altering the microbial population in the rumen, the primary site of CH4 production in ruminants and reducing the abundance of methanogens, the microorganisms responsible for producing CH4 [120]. However, the precise mechanism by which essential oils achieve this is not well understood, and more research is needed to determine their optimal use for mitigating CH4 emissions from livestock. It is suggested that the activity is not due to one specific mode of action but involves several targets in the bacterial cell.
It is worth noting that some essential oils increase feed consumption due to the aroma they add to the ration, while others, such as garlic oil, have a negative impact on palatability. Moreover, although feeding garlic or its metabolites may influence the flavour of meat and milk from ruminants, consumers may not accept it. Encapsulation of EOs in wall materials can enhance the long-term stability and antimicrobial ability of EOs in meat products, thereby eliminating or reducing the organoleptic defects of EOs in practical applications [101,121]. Unfortunately, this increases the application costs. Furthermore, with the increasing use of EOs, it is necessary to develop regulations that include the maximum permissible limits, toxicity, and capacity against specific methanogens without affecting other groups of microorganisms in the rumen.
3.1.6. Algae
Recently, algae have become one of the subjects of research aimed at reducing CH4 emissions from ruminants. Particular attention is given to three main taxa of macroalgae, commonly known as seaweed, which represent a large domain of aquatic plants separated into Chlorophyta (green), Phaeophyceae (brown), and Rhodophyta (red). In general, seaweeds contain polysaccharides, proteins, peptides, lipids, phlorotannins, saponins, and alkaloids that are known to reduce CH4 production by suppressing archaea and protozoa. However, the mode of action responsible for the mitigation effect centres around the content of volatile halogenated compounds (bromoform CHBr3) [122].
Dictyota (brown) and Asparagopsis (red) had the greatest potential for decreasing CH4 production among many widely studied species. The best-studied species exhibiting CH4 emission properties are Asparagopsis taxiformis and A. armata. In vivo studies reported dose (from 3.0 to 51.0 mg/kg of DMI) and diet-dependent (high-concentrate vs. high-forage) decreases from 9 to 98% of CH4 production by algae preparation [123,124,125,126]. Additionally, in some small-scale studies on cattle supplemented with algae, researchers have found a significant or numerical decrease in the feed-to-body mass gain ratio, an increase in milk yield, and feed efficiency in addition to the reduction of emission [125,127,128]. Under excess supplementation of A. taxiformis, CHBr3 is detected within milk, making it critical to monitor supplementation levels in lactating animals. Similarly, milk from cows fed seaweed had higher iodine concentrations. Lean et al. [128] assessed an iodine consumption 15-fold higher than the maximum tolerable for children under three years who consumed 1 L/day of milk. Therefore, any concerns of decreased productivity need to be addressed. Algae as a CH4 mitigation additive have received a lot of positive research. However, long-term oral exposure of animals to high concentrations of CHBr3 can cause liver and intestinal tumours. The Environmental Protection Agency (EPA) classified CHBr3 in Group B2 as a probable human carcinogen and toxic substance for the environment (i.e., ozone depletion). Chronic oral exposure of animals to high concentrations of CHBr3 can result in liver and intestinal tumours. EPA recommends that drinking water levels for bromoform should not be more than 0.7 parts per million (ppm) for CHBr3 [129].
Similar to the production of oils for animal feed, the CO2 emissions from growing, harvesting, drying, and transporting algae must be considered when evaluating the efficacy of algae in reducing GHG emissions because they could offset beneficial reductions of CH4 from ruminants. Furthermore, extensive algae aquaculture in ocean farming systems can be harmful to marine life and disrupt biodiversity (i.e., eutrophication) until we provide fully isolated algae cultures. The land-based seaweed production system is still in progress. Unfortunately, land-based cultures pose risks related to increased salinity and CO2 emissions from sources of energy needed to sustain the farm. Of course, it depends on the farms’ location, the salt source for farming (rock or seawater), power energy, and so on [130]. As part of an extensive algae aquaculture, the potential adverse effects of A. taxiformis supplementation on the health of animals, manufacturers, and consumers, as well as the natural environment, should be investigated further.
3.2. Mitigation through Microbiome Manipulation
The primary premise behind microbiome manipulation is that the growing microbial community of a newborn ruminant is more changeable, making it more likely to have long-term impacts than the established microbiome of an adult animal [131]. Furthermore, this method may be beneficial for grazing ruminants when supplementing feed components at the appropriate amount is more problematic.
3.2.1. Probiotics
Probiotics are microbiological feed additives based on selected bacterial or yeast cultures. They affect rumen fermentation and improve animal health by modulating the gastrointestinal microflora [75]. The colonisation of the rumen ecosystem with probiotics supports rumen fermentation and increases feed efficiency, which may reduce CH4 emissions [46]. A well-established relationship exists between increased propionate production and reduced CH4 emissions [8,20] (Figure 2). Microbial additives direct the fermentation pathway towards hydrogen-based propionate production. Thus, the concentration of a key precursor in CH4 production decreases.
The utilisation of H2 for NO3− and NO2− reduction to NH3 in the rumen is more thermodynamically favourable than the formation of CH4 with CO2 as an electron acceptor [53]. Probiotic bacteria are characterised by their capability to reduce NO3−, NO2−, and SO42−, thus competing with CH4 for H2. The yield of sulphate-reducing bacteria is highly dependent on the amount of SO42− delivered in the diet [12,46]. Moreover, probiotics may also stimulate acetogenic bacteria that compete for H2 with methanogens [46,132].
Recent research and applications have focused on three major types of probiotic bacteria. Propionibacteria are Gram-positive bacteria that live in the rumen. These include Propionibacterium acidipropionici, P. freudenreichii, P. propionicus, P. jensenii, P. japonicas, P. japonicas, and P. thoenii [133,134,135]. These in vitro studies demonstrate that some strains of propionibacteria have the potential to lower CH4 production by 20% from mixed ruminal cultures and that this reduction is not always associated with an increase in propionate production. Second, there are homoacetogens, a broad collection of bacterial species that can produce acetate. Third include Acetitomaculum ruminis, Eubacterium limosum, Blautia schinkii, and Blautia producta, which are alternative H2 sinks employed in in vitro experiments to eliminate H2 from the rumen [136]. The last group is methane-oxidising bacteria, which can grow on CH4 as a sole carbon and energy source. These bacteria use a methane monooxygenase to convert CH4 to methanol. Methanol is subsequently oxidised to formaldehyde by methanol dehydrogenase and absorbed into the serine or ribulose-5-monophosphate route for biomass synthesis. This group of bacteria is promising; however, they have not been used in studies to reduce CH4 production in cattle [137].
Other potential H2 consumers present in the rumen are nitrate/nitrite-reducing bacteria and sulphate-reducing bacteria (see section above); however, in normal conditions, their activity is low [22]. NO3−and NO2− reducing bacteria were given to rumen in vitro cultures with NO3− to decrease CH4 synthesis while enhancing NO3− reduction to ammonium and preventing NO2− buildup. This method, however, merely resulted in a quantifiable decrease in plasma NO2− concentrations in sheep [138,139].
3.2.2. Defaunation of the Rumen
The elimination of protozoa from the rumen ecology is known as rumen defaunation. The ciliate protozoan is thought to be a significant methanogen H2 producer. Furthermore, they form intimate associations with methanogens located on the surface and within protozoa cell bodies in structures known as hydrogenosomes. Protozoa, therefore, supply a substrate for methanogenesis while also protecting symbiotic archaea from oxygen toxicity. It is estimated that protozoa-associated methanogens contribute roughly 37% of rumen CH4 emissions [28,44]. Holotrich protozoa are thought to be more efficient H2 producers than entodiniomorphids and, thus, have a greater impact on methanogenesis. Methanogenesis in sheep exclusively faunated with Holotrich rumen was comparable to that in fully faunated animals [28]. Kittelmann et al. [140] found no variations in CH4 emission among sheep with distinct protozoa community types defined by Eadie [141]. They claimed that the structure of the ciliate population had no bearing on methanogenesis. However, Dai et al. [142] showed in a meta-analysis that isotrichids are more critical in methanogenesis than entodiniomorphids. The quantity of protozoa in the rumen corresponds with methanogen CH4 synthesis. Nevertheless, the total protozoa concentration in the rumen linearly correlates with the CH4 production by the methanogens [143,144].
The rumen is defaunated, which results in a 10–13% drop in CH4 production, an increase in propionate concentration, and lower levels of acetate and butyrate in the rumen content [47,67,145]. A meta-analysis of defaunation outcomes indicated no change in ruminal methanogen abundance and a minor increase in the variety [76]. Defaunation boosts bacterial population density, bacterial protein synthesis efficiency, and nitrogen flow to the duodenum, especially when the feed is low in protein relative to its energy content. Furthermore, defaunation reduces carbohydrate digestion of plant cell walls, significantly improves protein supply and livestock productivity, and lowers CH4 production. However, because of the need for rapid reinoculation due to animal cross-contamination maintaining protozoa-free animals in industrial production systems is extremely difficult.
Cattle defaunation is more complicated than sheep defaunation, which is thought to be due to differences in anatomical structures between the ovine and bovine rumens [146]. Thus, if simple but permanent methods of defaunating the animals can be discovered, defaunation has the potential to be a mitigation technique. Furthermore, regardless of the technical complexity of the removal methods, defaunation is not regarded as an appropriate CH4 mitigation strategy. The removal of protozoa from ruminants’ rumen is associated with decreased organic matter digestibility, particularly acid detergent fibre (ADF) and neutral detergent fibre (NDF), and decreased food intake [67,147].
3.2.3. Cellulolytic and Protozoan Activity Modification
The availability of H2, the core substrate for methanogenesis, depends mainly on cellulolytic bacterial and protozoan activity. Microbiome analysis of ruminal bacteria in sheep revealed three main ruminotypes: Q; S; and H. Q and S are associated with low CH4 emissions, but H is associated with high emissions. Q and S types, such as Quinella ovalis, are characterised by a high abundance of propionate-producing and lactate- and succinate-generating anaerobic prokaryotes, respectively. In contrast, sheep emitting more CH4 had more Ruminococcus and Ruminococcae, known for producing large quantities of H2 [140]. Similarly, ruminotypes associated with high and low CH4 emissions were described in dairy cows. Danielsson et al. [31] could distinguish two clusters, H and L, for high and low-emitting bacteria in Holstein and Swedish Red dairy cows, respectively. Cluster L is linked to low CH4 production (291 g CH4/day or 12.4 g CH4/kg DMI), while cluster H is linked to high CH4 production (345 g CH4/day or 14.5 g CH4/kg DMI).
Buffalo are considered low CH4 emitters because they have a different digestive system compared to other ruminants. Specifically, buffalo have a longer digestive tract and a different microbial community in their rumen, which allows for the more efficient digestion of their food and lower production of CH4 as a byproduct. Buffaloes have a naturally large abundance of Fibrobacter succinogenes, Succiniclasticum ruminis, and Syntrophococcus sucromutans rumens. [148]. These bacteria do not produce H2 during the cellulolytic plant fibre fermentation and are capable of producing succinate and propionate, which are non-methanogenic end products of the rumen fermentation [149,150,151]. F. succinogenes has also been shown to reduce CH4 production compared to Ruminococcus albus and Ruminococcus flavefaciens in the rumens of gnotobiotically reared lambs [14].
Moreover, buffaloes have significantly more Bacteroidota species, especially from the genus of the phylum Prevotella [148,152]. Prevotella can degrade noncellulose plant fibres and utilise lactate [153]. Therefore, the higher communities of Bacteroidota and Butyrivibrio found in buffalo rumen compared to the cattle rumen, as well as the comparable levels of Fibrobacter, suggest that buffalo are better adapted to coarse forage than cattle [154].
3.2.4. Vaccination
The development of vaccines for limiting methanogenesis is based on inducing the animal’s immune system to produce antibodies in saliva, which upon entry into the rumen, should suppress the growth of methanogens [155]. Another approach is to use chicken egg antibody (IgY), a rapid, economical, and non-invasive resource for antibodies [156]. Moreover, it does not necessitate changes in agricultural systems; vaccination is a very appealing strategy for reducing enteric CH4 emissions. It appears that this method would be especially beneficial for pasture-based breeding. To be effective, a vaccination must generate sufficiently high quantities of antibodies in the saliva, bind to the appropriate antigens of methanogens in the rumen fluid, and particular antigens over the whole spectrum of target methanogen species. When comparing studies to assess the possibilities of using vaccines against methanogens’ cells, several issues arose, making it difficult to compare results, such as the different adjuvants and vaccination protocols, different immunisation methods, different immunoglobulins (IgG, IgA, and IgY), different approaches (in vivo and in vitro), and samples (blood, saliva, and rumen) used to compare results [157]. There are few reports applying vaccines to mitigate CH4 production from enteric fermentation in ruminants [156,158,159,160,161,162,163]. All the in vitro studies showed a reduction in the amount of CH4 released, ranging from 7 to almost 70%, depending on the type of antibodies and the immunisation protocol [157]. Although antibodies-based in vitro research has shown very positive results, in vivo changes in the rumen methanogen population or CH4 emissions have not been particularly noteworthy or unsuccessful [161,164,165,166]. A vaccine against protozoan antigens has also been reported, but it failed to significantly reduce the ciliate population in Merino sheep [29]. So, it is complicated to evaluate the real effectiveness of this strategy; therefore, more study is needed to reach a firm conclusion on its feasibility, practicality, and long-term viability [157].
3.3. Chemical Intervention
Using chemicals directly inhibiting methanogenesis has been the most often-used strategy [165]. These substances must consistently reduce CH4 emissions while having no negative effects on people, animals, or the environment. They should be cost-effective for producers to accept them while boosting output and profitability. The majority of them are CH4 analogues or methyl-coenzyme M cofactors involved in methyl transfer during methanogenesis. In vivo-used inhibitors include bromochloromethane, trichloroacetamide, chloral hydrate, 9,10-anthraquinone, nitroethane, 3-nitrooxypropanol, also known as 3-nitrooxy-propan-1-ol or 1,3-propanediol mononitrate (3-NOP), and chloroform [62]. 3-NOP is one of the most effective dietary and economically valuable supplements for cattle that have been evaluated [165,166]. 3-Nitrooxypropanol mechanism of action involves inhibiting the enzyme microbial CH4 formation (Methyl-coenzyme M reductase), which is responsible for the final step in CH4 production by methanogenic archaea in the rumen [167]. The additive is intended for use in dairy cow’s feed to reduce enteric CH4 emission at a dose of 60 mg 3-NOP/kg (DM) [168]. 3-NOP is metabolised extensively to 3-nitrooxypropionic acid (NOPA), 3-hydroxy-propionic acid (3-HPA), NO3−/NO2− and CO2 [169]. Besides NO3−, metabolites of 3-NOP contribute to the formation of endogenous compounds, such as lactose and glucose, and are transformed into CO2.
The addition of 60 mg of 3-NOP per kg of DM to the total mixed ratio of dairy cows resulted in a significant reduction in enteric CH4 emissions per day per kg of DM intake, ranging from 20 to 35%. In parallel, there was a high increase in hydrogen emission, resulting in a feed energy waste [168]. Liu et al. [170] suggest that the combination of 3-NOP and fumarate could alleviate the accumulation of H2 and enhance the inhibition of methanogenesis compared with 3-NOP. The 3-NOP reduces CH4 generation; however, high-concentrate meals have been used in other trials to achieve reductions of 80% or more. However, the content of NDF in the diet has a detrimental impact on the 3-NOP’s ability to influence the CH4 generation [171]. Using 3-NOP appears to help lower CH4 emissions, although further studies are needed. The EFSA Panel on Additives and Products or Substances used in animal feed concludes that the genotoxicity potential of 3-NOP cannot be ruled out, as well as the effects of consumer exposure to NOPA.
To decrease the number of rumen microbial population species, organic acids have the potential to replace the antimicrobial compounds that are currently being used [165]. Malate, butyrate, and fumarate are all organic acids that naturally occur in biological tissues and are intermediates in the randomising pathway of propionate production, and they have been the subject of the majority of research on their use in ruminant feeding.
There are many mechanisms by which organic acid affects ruminal fermentation and animal performance. They increase the digestibility of DM and organic matter and, thus, influence CH4 production [172]. Moreover, an increase in the content of digestible energy may be a precursor of propionate and serves as a source of an H2 acceptor [47,173]. Organic acids containing halogens are involved in the methyl group transfer, thereby reducing the methanogenesis [5]. Mohammed et al. [174] and Ebrahimi et al. [175] used fumarate and malate in the presence of methanogenesis inhibitors (β-cyclodextrin iodopropane, 9,10-anthraquinone), with some success, to reduce H2 buildup caused by methanogenesis inhibition. Organic acids typically have only a minor impact on CH4 generation. They could contribute to incorporating electrons into VFAs when another mechanism suppresses methanogenesis. On the other hand, organic acid production is an expensive process. Therefore, dietary supplementation with this group of substances is currently uneconomical [61].
Nitrate is another dietary supplement used in the form of calcium, sodium, or potassium salt that has been used to decrease enteric CH4 in cattle. However, NO3− can exert direct toxic effects on methanogens through its reduction in the intermediate NO2− [176]. Due to the ability of NO3− and NO2− to be absorbed from or passed out of the rumen, the risk of toxicity increases, as does the possibility for NO2− to be unintentionally converted to N2O, a strong GHG. Another reason to stay away from NO3− is that higher rumen NO3− concentrations raise blood levels of NO3− and methemoglobin, which might lead to animal poisoning [177]. Furthermore, it appears to have no on-farm economic benefits, similar to organic acid [166].
Ionophores are commonly used feed additives in beef cattle diets to reduce CH4 emissions. They are carboxylic polyether substances produced by Streptomyces spp. that make the cell membranes of Gram-positive bacteria and protozoa more permeable to ions. Generally, ionophores cause a delay in bacteria growth and shift ruminal fermentation patterns, as well as improve the rumen’s ability to process feed. Several ionophores with a similar mechanism of action in the rumen (lasalocid, monensin, salinomycin, laidlomycin, and narasin) are commercially available. Furthermore, ionophores favour the growth of Gram-negative microorganisms, such as F. succinogenes, while not affecting methanogens, and, thus, indirectly inhibit ruminal methanogenesis by decreasing the availability of hydrogen and formate, the main substrates for methanogenic bacteria [149,178,179].
The most commonly used ionophore is monensin, a medication used as a feed additive for cattle to improve feed efficiency and prevent coccidiosis. As a general guideline, the recommended safe dose of monensin in feed for beef cattle is usually between 20 to 50 mg/kg complete feed depending on the stage of production and the type of diet being fed [180,181]. The effects of ionophores on ruminal fermentation patterns might be related to the diet composition, ionophore type, and dose used. Monensin supplementation to ruminants resulted in no significant effects, with up to a 30% reduction in CH4 emission [182]. For example, monensin (33–36 mg/kg DM) suppressed CH4 production in both high- (10.8% alfalfa silage, 75.2% corn silage, 12.9% canola meal, and 1.1% mineral mix) and low-concentrate diets (22.7% alfalfa silage, 8.3% corn silage, 67.9% barley grain, and 1.1% mineral mix). In the first four weeks of the low-concentrate diet, there was a 27% reduction in CH4 output, but in the first two weeks of the high-concentrate diet, there was a 30% reduction in CH4 output. Unfortunately, the CH4 levels returned to baseline after six weeks [183]. This occurred as a result of microbiota adaptation to the ionophore [184,185,186]. It is important to note that actual monensin dosing may vary based on individual animal and herd health factors.
On the other hand, the use of monensin in beef and dairy animals is prohibited in some countries and permitted in others. For today, the use of ionophores seems acceptable. Ionophores are not considered critically important in human medicine by the World Health Organization (WHO) because they are not used in human medicine. However, some reasons exist for seeking an alternative to continued ionophore use, including mitigating potential negative consequences in the future. Moreover, breeders and feed manufacturers must consider consumer concerns about current food production systems and food safety [187]. The use of chemical additives must be considered in light of potential future governmental legislation on chemical and antibiotic use and possible changes to importing country requirements.
3.4. Reducing Methane Emissions through Genetic Selection
Several studies over the last decade have shown that the heritability of CH4 characteristics in dairy cattle was moderate, ranging from 0.11 to 0.33; however, the heritabilities of CH4 yield in sheep were higher (0.24–0.55) [188,189,190,191,192]. Genetic selection is a very attractive solution because changes are cumulative and permanent; however, it requires multidisciplinary investigation and a large number of animals with CH4 records, but only a few countries actively record CH4 animal emissions. Moreover, animal selection is a very long-term process, and selecting animals with low CH4 emissions looks rather like an excellent future strategy. Here are some examples of multidisciplinary and international CH4 mitigation projects in animals: RuminOmics is a multidisciplinary research project funded by the European Union that aims to develop innovative solutions for reducing CH4 emissions from ruminants. The project brings together experts from various fields, such as genomics, microbiology, nutrition, and environmental science, to study the interactions between rumen microbiota and the host animal; The Animal Selection, Genetics, and Genomics Network (ASGGN) is a collaborative research initiative that formalised protocols for the collection and storage of data (including direct and indirect phenotypes, DNA, and rumen samples (if available)) from all animals measured; The Global Research Alliance is an international partnership that aims to coordinate and fund research on greenhouse gas mitigation in agriculture, including livestock. The alliance involves more than 60 member countries, including many of the world’s major livestock-producing nations, and focuses on a range of research areas, including livestock genomics, nutrition, and manure management.
Bringing together the data from various studies and countries could help create a future and accurate genomic reference database and develop precise genetic parameters for CH4 traits. Unfortunately, we should start reducing CH4 and other GHGs from livestock right now.
When making a genetic selection, we must consider the effect on productivity. There are numerous examples from across the industry of how rising animal performance has, over time, decreased the intensity of CH4 emissions. However, as animal productivity increases, CH4 intensity decreases curvilinearly. Thus, increasing the productivity of lower-producing animals has a relatively significant impact, whereas increasing the productivity of high-producing animals has a relatively small impact [12]. Manzanilla-Pech et al. [193], based on their analyses, find that CH4 concentration is genetically more closely related to CH4 production than any of the other CH4 variables investigated. They conclude that, when compared to CH4 production (MeP; g/d), CH4 yield (MeY; g/kg DMI), and CH4 intensity (MeI; g/kg energy-corrected milk), residual CH4 has the greatest potential for inclusion in the breeding goal because it allows for selecting low methane-emitting animals without compromising other economically important traits. Furthermore, possible relationships between enteric CH4 emissions and feed efficiency must be considered.
The selection of low-emitting animals may reduce the efficiency of fed digestion, particularly NDF, an essential ruminant characteristic in human food production. Many interacting biological and physiological factors, such as digestion rate, passage rate, rumen microbiome, and rumen fermentation, can all have an impact on feed efficiency and CH4 emission.
In a 10-year investigation, Rowe et al. [190,194] demonstrated that CH4 yield is heritable and, consequently, is under host control. Genetic selection has resulted in physiological changes affecting the rumen, feeding behaviour, rumen outputs, and body composition [194,195]. These changes appear to be economically beneficial, as ewes with low CH4 yield wean heavier, leaner lambs that produce more wool. Selected ewes have a 12% difference in CH4 emission yield between high and low emitters [194]. Moreover, the heritability and repeatability of CH4/(CH4 + CO2) for the CH4 yield traits were higher than CH4/DMI, indicating that gas traits are a reliable and accurate substitute for DMI at a constant level of feed intake [196]. On the other hand, preliminary studies show that genetic variation in CH4 emissions is present in Angus cattle; however, they did not identify antagonistic phenotypic or genetic relationships between CH4 and body composition traits.
One of the most difficult challenges in selecting animals with low CH4 production is measuring the CH4 in a large group of animals. Even commercial farms have difficulty performing these measurements [197]. A genetic selection program necessitates thousands of measurements, which should be taken on a weekly basis. Selection is also difficult because grazing systems differ depending on climate, plant species, soil types, and livestock and include season-long continuous grazing, rest-rotation grazing, deferred rotational grazing, and intensively managed grazing. It appears that animal selection is required to develop biomarkers that can reliably estimate CH4 production on all types of farms [193]. Bringing together the data from various studies and countries could help create a future and accurate genomic reference database and develop precise genetic parameters for CH4 traits. Unfortunately, we should start reducing CH4 and other GHGs from livestock right now, not with additional 10 years. If the country governments decide to implement animal breeding strategies to reduce enteric CH4 production and achieve the expected breeding impact, there must be predictability. The only way to accomplish this is to have a sufficient number of animals genotyped and phenotyped, and this data should be made public [197].
3.5. Forage Management
Feeding cellulosic material increases enteric CH4 emissions, with significant variation due to forage source, chemical composition, digestibility, forage selection, grazing management, and other factors. On the other hand, forage development systems are highly variable and dependent on farm site conditions (such as soil type and fertility, water, climate, country, and animal), as well as management practices and country policies regarding animal welfare. This variation opens up possibilities for CH4 reduction via forage management. So, when discussing CH4 mitigation, it is critical to consider the implications of the various options above. Moreover, mitigation of CH4 emissions from forage-based diets can be achieved to some extent by improving forage quality, harvest timing, use of forage species with superior digestibility, use of condensed tannin-containing plants, and storage of forages to conserve digestible nutrient content. Many publications deal with these comprehensive aspects; however, they are outside the scope of this review. It is critical that changes in forage management to reduce enteric CH4 emissions be assessed at the farm level using regionally specific life cycle assessments that account for differences in forage and animal productivity.
4. Perspectives and Conclusions
To meet the Paris Agreement’s projected goals, CH4 emissions from animal production must be reduced by 24 to 47% by 2050 compared to 2010. Livestock, produced worldwide, is an important agricultural product around the world. Globally, the livestock sector (particularly cattle, buffalo, sheep, goats, and camels) contributes 9–25% of anthropogenic GHG emissions, with the range in values attributed to different models and emission sources. Nitrous oxide from manure application and nitrogenous fertilisers, CO2 from fossil fuels and land-use changes, CH4 from enteric fermentation, and manure decomposition are all greenhouse gas emissions from livestock production [198].
Several enteric CH4 mitigation solutions discussed in this review are technically available, but many barriers to their implementation exist. Thus, reducing CH4 emissions on a global scale is still highly challenging. Dietary reformulation and feed additives can reduce enteric CH4 emissions immediately, whereas selective breeding could have effects in the future.
The practical approaches described above reduce CH4 emitted by ruminants with varying degrees of efficiency. However, it should be noted that the methods must be tailored to each of them individually, and a direct translation is unlikely. It is also necessary to consider producers’ economic expectations and consumer attitudes towards feed additives. Studies are still needed to develop strategies to achieve both CH4 mitigation and improvements in animal performance that benefit society and livestock producers. Furthermore, all chemical and biological interventions that may impact the health of ruminants and consumers should be handled with great care. The use of tannins, despite limiting methanogenesis, can be risky due to their toxicity. Probiotics also deserve special attention because natural microorganisms can change the biochemical reactions during methanogenesis and can stimulate the immune system. Moreover, little is known about the distribution of methanogens in gastrointestinal tracts and how additives affect them. Additionally, the net environmental GHG impacts need quantification, especially for algae farming. Another research priority is the implementation of CH4 mitigation strategies in grazing systems and different farming cultures.
Assume we want to adhere to Paris Agreement’s guidelines. In that case, we must intensify in vivo multidirectional research because much of the research data comes from in vitro research and cannot be translated into in vivo conditions (80% reduction vs. 25%, respectively). In vitro data, on the other hand, remain an important scientific result indicating directions for new strategy development.
Many feeding strategies are only now being introduced in ruminant farming to reduce CH4 production. Unfortunately, we generally continue to observe a global increase in CH4 emissions. Between 2000 and 2020, farm-gate GHG emissions increased by 13%; 57% of them are associated with livestock [199]. While CH4 emissions are decreasing in Europe, Australia, and North America, they are rapidly increasing in developing countries, particularly in Africa, followed by South America and Asia [200]. This rise is linked to the ongoing increase in the number of animals since 2000. The United States Department of Agriculture (USDA) reported a 1.0 billion head global cattle inventory in 2021, up 13.2 million heads from the previous year. It was the largest world inventory since 2004 [201]. World sheep numbers reached a new high of 1.266 billion heads in 2021, up from 1.263 billion heads in 2020. In addition, we have seen an increase in the number of goats and camels in recent years [200]. According to the data presented above, more intense research is needed to reduce ruminant CH4 emissions. These studies must take into account both the economic capabilities of developing countries and the animal husbandry methods used.
It is difficult to predict whether farmgate GHG emissions can be reduced using the current strategy, whether the trend of decreasing emissions in developed countries will continue, and whether this will be enough to offset increases in other regions.
We should use all available methods to reduce GHG emissions, but it appears that we should reduce breeding herd growth and increase the proportion of plant- and insect-based protein in food production; however, as long as it does not increase GHG emissions more than animal farming.
Author Contributions
The first draft of the manuscript was written by B.K., E.P.-K. and J.B.; B.K. reviewed and edited the final manuscript. All authors have read and agreed to the published version of the manuscript.
Funding
The APC was funded by Wroclaw University of Environmental and Life Sciences, Poland.
Institutional Review Board Statement
Not applicable.
Data Availability Statement
Not applicable.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Calabrò, P.S. Greenhouse gases emission from municipal waste management: The role of separate collection. Waste Manag. 2009, 29, 2178–2187. [Google Scholar] [CrossRef] [PubMed]
- Moss, A.R.; Jouany, J.-P.; Newbold, J. Methane production by ruminants: Its contribution to global warming. Ann. Zootech. 2000, 49, 231–253. [Google Scholar] [CrossRef]
- Kataria, R.P. Use of feed additives for reducing greenhouse gas emissions from dairy farms. Microbiol. Res. 2015, 6, 6120. [Google Scholar] [CrossRef]
- United States Environmental Protection Agency. Global Mitigation of Non-CO2 Greenhouse Gases: 2010–2030, EPA Report 430R13011; United States Environmental Protection Agency: Washington, DC, USA, 2013. Available online: https://www.epa.gov/global-mitigation-non-co2-greenhouse-gases/global-mitigation-non-co2-ghgs-report-2010-2030 (accessed on 5 February 2023).
- Patra, A.K. Enteric methane mitigation technologies for ruminant livestock: A synthesis of current research and future directions. Environ. Monit. Assess. 2012, 184, 1929–1952. [Google Scholar] [CrossRef] [PubMed]
- Gerber, P.; Steinfeld, H.; Henderson, B.; Mottet, A.; Opio, C.; Dijkman, J.; Falcucci, A.; Tempio, G. Tackling Climate Change Through Livestock: A Global Assessment of Emissions and Mitigation Opportunities; Food and Agriculture Organization of the United Nations (FAO): Rome, Italy, 2013. [Google Scholar]
- FAO. FAOSTAT Database Collections. Available online: http://faostat.fao.org/ (accessed on 10 December 2022).
- Haque, M.N. Dietary manipulation: A sustainable way to mitigate methane emissions from ruminants. J. Anim. Sci. Technol. 2018, 60, 15. [Google Scholar] [CrossRef]
- Opio, C.; Gerber, P.; Mottet, A.; Falcucci, A.; Tempio, G.; MacLeod, M.; Vellinga, T.; Henderson, B.; Steinfeld, H. Greenhouse Gas Emissions from Ruminant Supply Chains—A Global Life Cycle Assessment; Food and Agriculture Organization of the United Nations: Rome, Italy, 2013. [Google Scholar]
- Muller, R.A.; Muller, E.A. Fugitive methane and the role of atmospheric half-life. Geoinfor. Geostat. Overv. 2017, 5, 3. [Google Scholar] [CrossRef]
- Saunois, M.; Stavert, A.R.; Poulter, B.; Bousquet, P.; Canadell, J.G.; Jackson, R.B.; Raymond, P.A.; Dlugokencky, E.J.; Houweling, S.; Patra, P.K.; et al. The global methane budget 2000–2017. Earth Syst. Sci. Data 2020, 12, 1561–1623. [Google Scholar] [CrossRef]
- Beauchemin, K.A.; Ungerfeld, E.M.; Eckard, R.J.; Wang, M. Review: Fifty years of research on rumen methanogenesis: Lessons learned and future challenges for mitigation. Animal 2020, 14, s2–s16. [Google Scholar] [CrossRef]
- Guzman, C.E.; Bereza-Malcolm, L.T.; De Groef, B.; Franks, A.E. Presence of selected methanogens, fibrolytic bacteria, and proteobacteria in the gastrointestinal tract of neonatal dairy calves from birth to 72 hours. PLoS ONE 2015, 10, e0133048. [Google Scholar] [CrossRef]
- Henderson, G.; Cox, F.; Ganesh, S.; Jonker, A.; Young, W.; Abecia, L.; Angarita, E.; Aravena, P.; Nora Arenas, G.; Ariza, C.; et al. Rumen microbial community composition varies with diet and host, but a core microbiome is found across a wide geographical range. Sci. Rep. 2015, 5, 14567. [Google Scholar] [CrossRef]
- Hungate, R.E. The Rumen and Its Microbes; Academic Press: New York, NY, USA, 1966. [Google Scholar]
- Farah Naz, F.; Saba, K. Methanogenic diversity and taxonomy in the gastro intestinal tract of ruminants. In Extremophilic Microbes and Metabolites; Afef, N., Ameur, C., Haïtham, S., Hadda Imene, O., Eds.; IntechOpen: Rijeka, Croatia, 2019; p. 7. [Google Scholar]
- Kumar, S.; Puniya, A.K.; Puniya, M.; Dagar, S.S.; Sirohi, S.K.; Singh, K.; Griffith, G.W. Factors affecting rumen methanogens and methane mitigation strategies. World J. Microbiol. Biotechnol. 2009, 25, 1557–1566. [Google Scholar] [CrossRef]
- Mauerhofer, L.-M.; Zwirtmayr, S.; Pappenreiter, P.; Bernacchi, S.; Seifert, A.H.; Reischl, B.; Schmider, T.; Taubner, R.-S.; Paulik, C.; Rittmann, S.K.M.R. Hyperthermophilic methanogenic archaea act as high-pressure CH4 cell factories. Commun. Biol. 2021, 4, 289. [Google Scholar] [CrossRef] [PubMed]
- De la Fuente, G.; Yañez-Ruiz, D.R.; Seradj, A.R.; Balcells, J.; Belanche, A. Methanogenesis in animals with foregut and hindgut fermentation: A review. Anim. Prod. Sci. 2019, 59, 2109–2122. [Google Scholar] [CrossRef]
- Janssen, P.H. Influence of hydrogen on rumen methane formation and fermentation balances through microbial growth kinetics and fermentation thermodynamics. Anim. Feed Sci. Technol. 2010, 160, 1–22. [Google Scholar] [CrossRef]
- Thauer, R.K.; Kaster, A.K.; Seedorf, H.; Buckel, W.; Hedderich, R. Methanogenic archaea: Ecologically relevant differences in energy conservation. Nat. Rev. Microbiol. 2008, 6, 579–591. [Google Scholar] [CrossRef]
- Van Zijderveld, S.M.; Gerrits, W.J.J.; Apajalahti, J.A.; Newbold, J.R.; Dijkstra, J.; Leng, R.A.; Perdok, H.B. Nitrate and sulfate: Effective alternative hydrogen sinks for mitigation of ruminal methane production in sheep. J. Dairy Sci. 2010, 93, 5856–5866. [Google Scholar] [CrossRef]
- Choudhury, P.K.; Jena, R.; Tomar, S.K.; Puniya, A.K. Reducing enteric methanogenesis through alternate hydrogen sinks in the rumen. Methane 2022, 1, 320–341. [Google Scholar] [CrossRef]
- Buddle, B.M.; Denis, M.; Attwood, G.T.; Altermann, E.; Janssen, P.H.; Ronimus, R.S.; Pinares-Patiño, C.S.; Muetzel, S.; Neil Wedlock, D. Strategies to reduce methane emissions from farmed ruminants grazing on pasture. Vet. J. 2011, 188, 11–17. [Google Scholar] [CrossRef]
- Greening, C.; Geier, R.; Wang, C.; Woods, L.C.; Morales, S.E.; McDonald, M.J.; Rushton-Green, R.; Morgan, X.C.; Koike, S.; Leahy, S.C.; et al. Diverse hydrogen production and consumption pathways influence methane production in ruminants. ISME J. 2019, 13, 2617–2632. [Google Scholar] [CrossRef]
- Malik, P.K.; Bhatta, R.; Gagen, E.J.; Sejian, V.; Soren, N.M.; Prasad, C.S. Alternate H2 sinks for reducing rumen methanogenesis. In Climate Change Impact on Livestock: Adaptation and Mitigation; Sejian, V., Gaughan, J., Baumgard, L., Prasad, C., Eds.; Springer: New Delhi, India, 2015; pp. 303–320. [Google Scholar]
- Holmes, D.E.; Smith, J.A. Biologically produced methane as a renewable energy source. Adv. Appl. Microbiol. 2016, 97, 1–61. [Google Scholar] [CrossRef]
- Belanche, A.; de la Fuente, G.; Newbold, C.J. Study of methanogen communities associated with different rumen protozoal populations. FEMS Microbiol. Ecol. 2014, 90, 663–677. [Google Scholar] [CrossRef] [PubMed]
- Leng, R.A. Interactions between microbial consortia in biofilms: A paradigm shift in rumen microbial ecology and enteric methane mitigation. Anim. Prod. Sci. 2014, 54, 519–543. [Google Scholar] [CrossRef]
- Valle, E.R.; Henderson, G.; Janssen, P.H.; Cox, F.; Alexander, T.W.; McAllister, T.A. Considerations in the use of fluorescence in situ hybridization (FISH) and confocal laser scanning microscopy to characterize rumen methanogens and define their spatial distributions. Can. J. Microbiol. 2015, 61, 417–428. [Google Scholar] [CrossRef] [PubMed]
- Danielsson, R.; Dicksved, J.; Sun, L.; Gonda, H.; Müller, B.; Schnürer, A.; Bertilsson, J. Methane production in dairy cows correlates with rumen methanogenic and bacterial community structure. Front. Microbiol. 2017, 8, 226. [Google Scholar] [CrossRef] [PubMed]
- Janssen, P.H.; Kirs, M. Structure of the archaeal community of the rumen. Appl. Environ. Microbiol. 2008, 74, 3619–3625. [Google Scholar] [CrossRef] [PubMed]
- Leahy, S.C.; Kelly, W.J.; Altermann, E.; Ronimus, R.S.; Yeoman, C.J.; Pacheco, D.M.; Li, D.; Kong, Z.; McTavish, S.; Sang, C.; et al. The genome sequence of the rumen methanogen methanobrevibacter ruminantium reveals new possibilities for controlling ruminant methane emissions. PLoS ONE 2010, 5, e8926. [Google Scholar] [CrossRef] [PubMed]
- Patra, A.; Park, T.; Kim, M.; Yu, Z. Rumen methanogens and mitigation of methane emission by anti-methanogenic compounds and substances. J. Anim. Sci. Biotechnol. 2017, 8, 13. [Google Scholar] [CrossRef]
- Teklebrhan, T.; Tan, Z.; Wang, M.; Wang, R. Rumen methanogens community as drivers of methane emission. J. Vet. Sci. Ani. Husb. 2018, 6, 406. [Google Scholar]
- Hungate, R.E. Hydrogen as an intermediate in the rumen fermentation. Arch. Mikrobiol. 1967, 59, 158–164. [Google Scholar] [CrossRef]
- King, E.E.; Smith, R.P.; St-Pierre, B.; Wright, A.D. Differences in the rumen methanogen populations of lactating Jersey and Holstein dairy cows under the same diet regimen. Appl. Environ. Microbiol. 2011, 77, 5682–5687. [Google Scholar] [CrossRef]
- Lee, J.H.; Kumar, S.; Lee, G.H.; Chang, D.H.; Rhee, M.S.; Yoon, M.H.; Kim, B.C. Methanobrevibacter boviskoreani sp. nov., isolated from the rumen of Korean native cattle. Int. J. Syst. Evol. Microbiol. 2013, 63, 4196–4201. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Whitman, W.B. Metabolic, phylogenetic, and ecological diversity of the methanogenic archaea. Ann. N. Y. Acad. Sci. 2008, 1125, 171–189. [Google Scholar] [CrossRef] [PubMed]
- Thauer, R.K. The Wolfe cycle comes full circle. Proc. Natl. Acad. Sci. USA 2012, 109, 15084–15085. [Google Scholar] [CrossRef] [PubMed]
- Wang, K.; Nan, X.; Chu, K.; Tong, J.; Yang, L.; Zheng, S.; Zhao, G.; Jiang, L.; Xiong, B. Shifts of hydrogen metabolism from methanogenesis to propionate production in response to replacement of forage fiber with non-forage fiber sources in diets in vitro. Front. Microbiol. 2018, 9, 2764. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.; Giller, K.; Kreuzer, M.; Ulbrich, S.E.; Braun, U.; Schwarm, A. Contribution of ruminal fungi, archaea, protozoa, and bacteria to the methane suppression caused by oilseed supplemented diets. Front. Microbiol. 2017, 8, 1864. [Google Scholar] [CrossRef]
- Yang, C.; Rooke, J.A.; Cabeza, I.; Wallace, R.J. Nitrate and inhibition of ruminal methanogenesis: Microbial ecology, obstacles, and opportunities for lowering methane emissions from ruminant livestock. Front. Microbiol. 2016, 7, 132. [Google Scholar] [CrossRef]
- Lan, W.; Yang, C. Ruminal methane production: Associated microorganisms and the potential of applying hydrogen-utilizing bacteria for mitigation. Sci. Total Environ. 2019, 654, 1270–1283. [Google Scholar] [CrossRef]
- Costa, K.C.; Wong, P.M.; Wang, T.; Lie, T.J.; Dodsworth, J.A.; Swanson, I.; Burn, J.A.; Hackett, M.; Leigh, J.A. Protein complexing in a methanogen suggests electron bifurcation and electron delivery from formate to heterodisulfide reductase. Proc. Natl. Acad. Sci. USA 2010, 107, 11050–11055. [Google Scholar] [CrossRef]
- Islam, M.; Lee, S.S. Advanced estimation and mitigation strategies: A cumulative approach to enteric methane abatement from ruminants. J. Anim. Sci. Technol. 2019, 61, 122–137. [Google Scholar] [CrossRef]
- Morgavi, D.P.; Forano, E.; Martin, C.; Newbold, C.J. Microbial ecosystem and methanogenesis in ruminants. Animal 2010, 4, 1024–1036. [Google Scholar] [CrossRef]
- Morgavi, D.P.; Forano, E.; Martin, C.; Newbold, C.J. Microbial ecosystem and methanogenesis in ruminants—CORRIGENDUM. Animal 2012, 6, 871. [Google Scholar] [CrossRef]
- Ragsdale, S.W.; Pierce, E. Acetogenesis and the Wood-Ljungdahl pathway of CO2 fixation. Biochim. Biophys. Acta 2008, 1784, 1873–1898. [Google Scholar] [CrossRef] [PubMed]
- Baldwin, R.L.; Allison, M.J. Rumen metabolism. J. Anim. Sci. 1983, 57 (Suppl. 2), 461–477. [Google Scholar] [PubMed]
- Kobayashi, Y. Abatement of methane production from ruminants: Trends in the manipulation of rumen fermentation. Asian-Australas. J. Anim. Sci. 2010, 23, 410–416. [Google Scholar] [CrossRef]
- Martinez-Fernandez, G.; Denman, S.E.; Cheung, J.; McSweeney, C.S. Phloroglucinol degradation in the rumen promotes the capture of excess hydrogen generated from methanogenesis inhibition. Front. Microbiol. 2017, 8, 1871. [Google Scholar] [CrossRef]
- Latham, E.A.; Anderson, R.C.; Pinchak, W.E.; Nisbet, D.J. Insights on alterations to the rumen ecosystem by nitrate and nitrocompounds. Front. Microbiol. 2016, 7, 228. [Google Scholar] [CrossRef]
- Olijhoek, D.W.; Hellwing, A.L.F.; Brask, M.; Weisbjerg, M.R.; Højberg, O.; Larsen, M.K.; Dijkstra, J.; Erlandsen, E.J.; Lund, P. Effect of dietary nitrate level on enteric methane production, hydrogen emission, rumen fermentation, and nutrient digestibility in dairy cows. J. Dairy Sci. 2016, 99, 6191–6205. [Google Scholar] [CrossRef]
- Ungerfeld, E.M.; Kohn, R.A. The role of thermodynamics in the control of ruminal fermentation. In Ruminant Physiology: Digestion, Metabolism and Impact of Nutrition on Gene Expression, Immunology and Stress; Sejrsen, K., Hvelplund, T., Nielsen, M.O., Eds.; Wageningen Academic Publishers: Wageningen, The Netherlands, 2006; pp. 55–85. [Google Scholar]
- Raleng, A. The Potential of Feeding Nitrate to Reduce Enteric Methane Production in Ruminants; The Department of Climate Change Commonwealth Government of Australia: Canberra, Australia, 2008.
- Simon, J. Enzymology and bioenergetics of respiratory nitrite ammonification. FEMS Microbiol. Rev. 2002, 26, 285–309. [Google Scholar] [CrossRef]
- Uniyal, S.; Mishra, A.K.; Aswin, K.; Sahoo, J.K.; Munde, V.K.; Mishra, G.K. Use of nitrates and sulphates as hydrogen sink in reducing enteric methane production. Research & reviews. J. Vet. Sci. Technol. 2016, 5, 5–8. [Google Scholar]
- Huisingh, J.; McNeill, J.J.; Matrone, G. Sulfate reduction by a Desulfovibrio species isolated from sheep rumen. Appl. Microbiol. 1974, 28, 489–497. [Google Scholar] [CrossRef]
- Negussie, E.; de Haas, Y.; Dehareng, F.; Dewhurst, R.J.; Dijkstra, J.; Gengler, N.; Morgavi, D.P.; Soyeurt, H.; van Gastelen, S.; Yan, T.; et al. Invited review: Large-scale indirect measurements for enteric methane emissions in dairy cattle: A review of proxies and their potential for use in management and breeding decisions. J. Dairy Sci. 2017, 100, 2433–2453. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Choudhury, P.K.; Carro, M.D.; Griffith, G.W.; Dagar, S.S.; Puniya, M.; Calabro, S.; Ravella, S.R.; Dhewa, T.; Upadhyay, R.C.; et al. New aspects and strategies for methane mitigation from ruminants. Appl. Microbiol. Biotechnol. 2014, 98, 31–44. [Google Scholar] [CrossRef] [PubMed]
- Patra, A.K.; Saxena, J. The effect and mode of action of saponins on the microbial populations and fermentation in the rumen and ruminant production. Nutr. Res. Rev. 2009, 22, 204–219. [Google Scholar] [CrossRef] [PubMed]
- Romero-Pérez, G.A.; Ominski, K.H.; McAllister, T.A.; Krause, D.O. Effect of environmental factors and influence of rumen and hindgut biogeography on bacterial communities in steers. Appl. Environ. Microbiol. 2011, 77, 258–268. [Google Scholar] [CrossRef] [PubMed]
- Ramos-Morales, E.; de la Fuente, G.; Duval, S.; Wehrli, C.; Bouillon, M.; Lahmann, M.; Preskett, D.; Braganca, R.; Newbold, C.J. Antiprotozoal effect of saponins in the rumen can be enhanced by chemical modifications in their structure. Front. Microbiol. 2017, 8, 399. [Google Scholar] [CrossRef] [PubMed]
- Jayanegara, A.; Wina, E.; Takahashi, J. Meta-analysis on methane mitigating properties of saponin-rich sources in the rumen: Influence of addition levels and plant sources. Asian-Australas. J. Anim. Sci. 2014, 27, 1426–1435. [Google Scholar] [CrossRef]
- Hartinger, T.; Gresner, N.; Südekum, K.H. Does intra-ruminal nitrogen recycling waste valuable resources? A review of major players and their manipulation. J. Anim. Sci. Biotechnol. 2018, 9, 33. [Google Scholar] [CrossRef]
- Newbold, C.J.; de la Fuente, G.; Belanche, A.; Ramos-Morales, E.; McEwan, N.R. The role of ciliate protozoa in the rumen. Front. Microbiol. 2015, 6, 1313. [Google Scholar] [CrossRef]
- Wina, E.; Muetzel, S.; Becker, K. The impact of saponins or saponin-containing plant materials on ruminant production—A review. J. Agric. Food Chem. 2005, 53, 8093–8105. [Google Scholar] [CrossRef]
- Pen, B.; Sar, C.; Mwenya, B.; Kuwaki, K.; Morikawa, R.; Takahashi, J. Effects of Yucca schidigera and Quillaja saponaria extracts on in vitro ruminal fermentation and methane emission. Anim. Feed Sci. Technol. 2006, 129, 175–186. [Google Scholar] [CrossRef]
- Ku-Vera, J.C.; Jiménez-Ocampo, R.; Valencia-Salazar, S.S.; Montoya-Flores, M.D.; Molina-Botero, I.C.; Arango, J.; Gómez-Bravo, C.A.; Aguilar-Pérez, C.F.; Solorio-Sánchez, F.J. Role of secondary plant metabolites on enteric methane mitigation in ruminants. Front. Vet. Sci. 2020, 7, 584. [Google Scholar] [CrossRef] [PubMed]
- Moses, T.; Papadopoulou, K.K.; Osbourn, A. Metabolic and functional diversity of saponins, biosynthetic intermediates and semi-synthetic derivatives. Crit. Rev. Biochem. Mol. Biol. 2014, 49, 439–462. [Google Scholar] [CrossRef] [PubMed]
- Samtiya, M.; Aluko, R.E.; Dhewa, T. Plant food anti-nutritional factors and their reduction strategies: An overview. Food Prod. Process. Nutr. 2020, 2, 6. [Google Scholar] [CrossRef]
- Broucek, J. Options to methane production abatement in ruminants: A review. J. Anim. Plant Sci. 2018, 28, 348–364. [Google Scholar]
- Liu, H.; Vaddella, V.; Zhou, D. Effects of chestnut tannins and coconut oil on growth performance, methane emission, ruminal fermentation, and microbial populations in sheep. J. Dairy Sci. 2011, 94, 6069–6077. [Google Scholar] [CrossRef]
- Tavendale, M.H.; Meagher, L.P.; Pacheco, D.; Walker, N.; Attwood, G.T.; Sivakumaran, S. Methane production from in vitro rumen incubations with Lotus pedunculatus and Medicago sativa, and effects of extractable condensed tannin fractions on methanogenesis. Anim. Feed Sci. Technol. 2005, 123–124, 403–419. [Google Scholar] [CrossRef]
- Field, J.A.; Kortekaas, S.; Lettinga, G. The tannin theory of methanogenic toxicity. Biol. Wastes 1989, 29, 241–262. [Google Scholar] [CrossRef]
- McSweeney, C.S.; Palmer, B.; McNeill, D.M.; Krause, D.O. Microbial interactions with tannins: Nutritional consequences for ruminants. Anim. Feed Sci. Technol. 2001, 91, 83–93. [Google Scholar] [CrossRef]
- Tiemann, T.T.; Lascano, C.E.; Wettstein, H.R.; Mayer, A.C.; Kreuzer, M.; Hess, H.D. Effect of the tropical tannin-rich shrub legumes Calliandra calothyrsus and Flemingia macrophylla on methane emission and nitrogen and energy balance in growing lambs. Animal 2008, 2, 790–799. [Google Scholar] [CrossRef]
- Jayanegara, A.; Leiber, F.; Kreuzer, M. Meta-analysis of the relationship between dietary tannin level and methane formation in ruminants from in vivo and in vitro experiments. J. Anim. Physiol. Anim. Nutr. 2012, 96, 365–375. [Google Scholar] [CrossRef]
- Aboagye, I.A.; Beauchemin, K.A. Potential of molecular weight and structure of tannins to reduce methane emissions from ruminants: A review. Animals 2019, 9, 856. [Google Scholar] [CrossRef] [PubMed]
- Vasta, V.; Daghio, M.; Cappucci, A.; Buccioni, A.; Serra, A.; Viti, C.; Mele, M. Invited review: Plant polyphenols and rumen microbiota responsible for fatty acid biohydrogenation, fiber digestion, and methane emission: Experimental evidence and methodological approaches. J. Dairy Sci. 2019, 102, 3781–3804. [Google Scholar] [CrossRef] [PubMed]
- McSweeney, C.S.; Palmer, B.; Bunch, R.; Krause, D.O. Effect of the tropical forage calliandra on microbial protein synthesis and ecology in the rumen. J. Appl. Microbiol. 2001, 90, 78–88. [Google Scholar] [CrossRef] [PubMed]
- Obreque-Slier, E.; López-Solís, R.; Peña-Neira, Á.; Zamora-Marín, F. Tannin–protein interaction is more closely associated with astringency than tannin–protein precipitation: Experience with two oenological tannins and a gelatin. Int. J. Food Sci. Technol. 2010, 45, 2629–2636. [Google Scholar] [CrossRef]
- Adamczyk, B.; Simon, J.; Kitunen, V.; Adamczyk, S.; Smolander, A. Tannins and their complex interaction with different organic nitrogen compounds and enzymes: Old paradigms versus recent advances. ChemistryOpen 2017, 6, 610–614. [Google Scholar] [CrossRef]
- Malik, P.K.; Kolte, A.P.; Baruah, L.; Saravanan, M.; Bakshi, B.; Bhatta, R. Enteric methane mitigation in sheep through leaves of selected tanniniferous tropical tree species. Livest. Sci. 2017, 200, 29–34. [Google Scholar] [CrossRef]
- Salami, S.A.; Valenti, B.; Bella, M.; O’Grady, M.N.; Luciano, G.; Kerry, J.P.; Jones, E.; Priolo, A.; Newbold, C.J. Characterisation of the ruminal fermentation and microbiome in lambs supplemented with hydrolysable and condensed tannins. FEMS Microbiol. Ecol. 2018, 94, fiy061. [Google Scholar] [CrossRef]
- Cardoso-Gutierrez, E.; Aranda-Aguirre, E.; Robles-Jimenez, L.E.; Castelán-Ortega, O.A.; Chay-Canul, A.J.; Foggi, G.; Angeles-Hernandez, J.C.; Vargas-Bello-Pérez, E.; González-Ronquillo, M. Effect of tannins from tropical plants on methane production from ruminants: A systematic review. Vet. Anim. Sci. 2021, 14, 100214. [Google Scholar] [CrossRef]
- Animut, G.; Puchala, R.; Goetsch, A.L.; Patra, A.K.; Sahlu, T.; Varel, V.H.; Wells, J. Methane emission by goats consuming diets with different levels of condensed tannins from lespedeza. Anim. Feed Sci. Technol. 2008, 144, 212–227. [Google Scholar] [CrossRef]
- Waghorn, G. Beneficial and detrimental effects of dietary condensed tannins for sustainable sheep and goat production—Progress and challenges. Anim. Feed Sci. Technol. 2008, 147, 116–139. [Google Scholar] [CrossRef]
- Smith, A.H.; Zoetendal, E.; Mackie, R.I. Bacterial mechanisms to overcome inhibitory effects of dietary tannins. Microb. Ecol. 2005, 50, 197–205. [Google Scholar] [CrossRef] [PubMed]
- Oskoueian, E.; Abdullah, N.; Oskoueian, A. Effects of flavonoids on rumen fermentation activity, methane production, and microbial population. BioMed Res. Int. 2013, 2013, 349129. [Google Scholar] [CrossRef] [PubMed]
- Formato, M.; Cimmino, G.; Brahmi-Chendouh, N.; Piccolella, S.; Pacifico, S. Polyphenols for livestock feed: Sustainable perspectives for animal husbandry? Molecules 2022, 27, 7752. [Google Scholar] [CrossRef] [PubMed]
- Olagaray, K.E.; Bradford, B.J. Plant flavonoids to improve productivity of ruminants—A review. Anim. Feed Sci. Technol. 2019, 251, 21–36. [Google Scholar] [CrossRef]
- Seradj, A.R.; Abecia, L.; Crespo, J.; Villalba, D.; Fondevila, M.; Balcells, J. The effect of Bioflavex® and its pure flavonoid components on in vitro fermentation parameters and methane production in rumen fluid from steers given high concentrate diets. Anim. Feed Sci. Technol. 2014, 197, 85–91. [Google Scholar] [CrossRef]
- Chen, L.; Shen, Y.; Wang, C.; Ding, L.; Zhao, F.; Wang, M.; Fu, J.; Wang, H. Megasphaera elsdenii lactate degradation pattern shifts in rumen acidosis models. Front. Microbiol. 2019, 10, 162. [Google Scholar] [CrossRef]
- Sinz, S.; Kunz, C.; Liesegang, A.; Braun, U.; Marquardt, S.; Soliva, R.C.; Kreuzer, M. In vitro bioactivity of various pure flavonoids in ruminal fermentation, with special reference to methane formation. Czech J. Anim. Sci. 2018, 63, 293–304. [Google Scholar] [CrossRef]
- María Carpena, R.; Cristina, C.; Bernabe, N.-E.; Eliana, P.; Maria, F.-C.; Filipa, S.R.; Jesus, S.-G.; Isabel, C.F.R.F.; Miguel, A.P.; Lillian, B. Flavonoids: A group of potential food additives with beneficial health effects. In Natural Food Additives; Miguel, A.P., Paz, O., Eds.; IntechOpen: Rijeka, Croatia, 2021; p. 4. [Google Scholar]
- Machmüller, A.; Soliva, C.R.; Kreuzer, M. Methane-suppressing effect of myristic acid in sheep as affected by dietary calcium and forage proportion. Br. J. Nutr. 2003, 90, 529–540. [Google Scholar] [CrossRef]
- Zhou, X.; Zeitz, J.O.; Meile, L.; Kreuzer, M.; Schwarm, A. Influence of pH and the degree of protonation on the inhibitory effect of fatty acids in the ruminal methanogen Methanobrevibacter ruminantium strain M1. J. Appl. Microbiol. 2015, 119, 1482–1493. [Google Scholar] [CrossRef]
- Zhou, X.; Meile, L.; Kreuzer, M.; Zeitz, J.O. The effect of saturated fatty acids on methanogenesis and cell viability of Methanobrevibacter ruminantium. Archaea 2013, 2013, 106916. [Google Scholar] [CrossRef]
- Klevenhusen, F.; Zeitz, J.O.; Duval, S.; Kreuzer, M.; Soliva, C.R. Garlic oil and its principal component diallyl disulfide fail to mitigate methane, but improve digestibility in sheep. Anim. Feed Sci. Technol. 2011, 166–167, 356–363. [Google Scholar] [CrossRef]
- Beauchemin, K.A.; McGinn, S.M.; Petit, H.V. Methane abatement strategies for cattle: Lipid supplementation of diets. Can. J. Anim. Sci. 2007, 87, 431–440. [Google Scholar] [CrossRef]
- McGinn, S.M.; Beauchemin, K.A.; Coates, T.; Colombatto, D. Methane emissions from beef cattle: Effects of monensin, sunflower oil, enzymes, yeast, and fumaric acid. J. Anim. Sci. 2004, 82, 3346–3356. [Google Scholar] [CrossRef] [PubMed]
- Patra, A.K. The effect of dietary fats on methane emissions, and its other effects on digestibility, rumen fermentation and lactation performance in cattle: A meta-analysis. Livest. Sci. 2013, 155, 244–254. [Google Scholar] [CrossRef]
- Jordan, E.; Lovett, D.K.; Monahan, F.J.; Callan, J.; Flynn, B.; O’Mara, F.P. Effect of refined coconut oil or copra meal on methane output and on intake and performance of beef heifers. J. Anim. Sci. 2006, 84, 162–170. [Google Scholar] [CrossRef]
- Beauchemin, K.A.; Ungerfeld, E.M.; Abdalla, A.L.; Alvarez, C.; Arndt, C.; Becquet, P.; Benchaar, C.; Berndt, A.; Mauricio, R.M.; McAllister, T.A.; et al. Invited review: Current enteric methane mitigation options. J. Dairy Sci. 2022, 105, 9297–9326. [Google Scholar] [CrossRef]
- Hassanat, F.; Benchaar, C. Methane emissions of manure from dairy cows fed red clover- or corn silage-based diets supplemented with linseed oil. J. Dairy Sci. 2019, 102, 11766–11776. [Google Scholar] [CrossRef]
- Grainger, C.; Beauchemin, K.A. Can enteric methane emissions from ruminants be lowered without lowering their production? Anim. Feed Sci. Technol. 2011, 166–167, 308–320. [Google Scholar] [CrossRef]
- Arndt, C.; Hristov, A.N.; Price, W.J.; McClelland, S.C.; Pelaez, A.M.; Cueva, S.F.; Oh, J.; Dijkstra, J.; Bannink, A.; Bayat, A.R.; et al. Full adoption of the most effective strategies to mitigate methane emissions by ruminants can help meet the 1.5 °C target by 2030 but not 2050. Proc. Natl. Acad. Sci. USA 2022, 119, e2111294119. [Google Scholar] [CrossRef]
- Palmquist, D.L.; Jenkins, T.C. A 100-year review: Fat feeding of dairy cows. J. Dairy Sci. 2017, 100, 10061–10077. [Google Scholar] [CrossRef]
- Falleh, H.; Ben Jemaa, M.; Saada, M.; Ksouri, R. Essential oils: A promising eco-friendly food preservative. Food Chem. 2020, 330, 127268. [Google Scholar] [CrossRef] [PubMed]
- Kalogianni, A.I.; Lazou, T.; Bossis, I.; Gelasakis, A.I. Natural phenolic compounds for the control of oxidation, bacterial spoilage, and foodborne pathogens in meat. Foods 2020, 9, 794. [Google Scholar] [CrossRef] [PubMed]
- Busquet, M.; Calsamiglia, S.; Ferret, A.; Carro, M.D.; Kamel, C. Effect of garlic oil and four of its compounds on rumen microbial fermentation. J. Dairy Sci. 2005, 88, 4393–4404. [Google Scholar] [CrossRef] [PubMed]
- Soliva, C.R.; Amelchanka, S.L.; Duval, S.M.; Kreuzer, M. Ruminal methane inhibition potential of various pure compounds in comparison with garlic oil as determined with a rumen simulation technique (Rusitec). Br. J. Nutr. 2011, 106, 114–122. [Google Scholar] [CrossRef] [PubMed]
- Belanche, A.; Newbold, C.J.; Morgavi, D.P.; Bach, A.; Zweifel, B.; Yáñez-Ruiz, D.R. A Meta-analysis describing the effects of the essential oils blend agolin ruminant on performance, rumen fermentation and methane emissions in dairy Cows. Animals 2020, 10, 620. [Google Scholar] [CrossRef] [PubMed]
- Benchaar, C.; Calsamiglia, S.; Chaves, A.V.; Fraser, G.R.; Colombatto, D.; McAllister, T.A.; Beauchemin, K.A. A review of plant-derived essential oils in ruminant nutrition and production. Anim. Feed Sci. Technol. 2008, 145, 209–228. [Google Scholar] [CrossRef]
- Saro, C.; Hohenester, U.M.; Bernard, M.; Lagrée, M.; Martin, C.; Doreau, M.; Boudra, H.; Popova, M.; Morgavi, D.P. Effectiveness of interventions to modulate the rumen microbiota composition and function in pre-ruminant and ruminant lambs. Front. Microbiol. 2018, 9, 1273. [Google Scholar] [CrossRef]
- Mucha, W.; Witkowska, D. The applicability of essential oils in different stages of production of animal-based foods. Molecules 2021, 26, 3798. [Google Scholar] [CrossRef]
- Benchaar, C.; Greathead, H. Essential oils and opportunities to mitigate enteric methane emissions from ruminants. Anim. Feed Sci. Technol. 2011, 166–167, 338–355. [Google Scholar] [CrossRef]
- Wallace, R.J. Antimicrobial properties of plant secondary metabolites. Proc. Nutr. Soc. 2004, 63, 621–629. [Google Scholar] [CrossRef]
- Amin, N.; Tagliapietra, F.; Arango, S.; Guzzo, N.; Bailoni, L. Free and microencapsulated essential oils incubated in vitro: Ruminal stability and fermentation parameters. Animals 2021, 11, 180. [Google Scholar] [CrossRef] [PubMed]
- Machado, L.; Magnusson, M.; Paul, N.A.; Kinley, R.; de Nys, R.; Tomkins, N. Identification of bioactives from the red seaweed Asparagopsis taxiformis that promote antimethanogenic activity in vitro. J. Appl. Phycol. 2016, 28, 3117–3126. [Google Scholar] [CrossRef]
- Li, X.; Norman, H.C.; Kinley, R.D.; Laurence, M.; Wilmot, M.; Bender, H.; de Nys, R.; Tomkins, N. Asparagopsis taxiformis decreases enteric methane production from sheep. Anim. Prod. Sci. 2018, 58, 681–688. [Google Scholar] [CrossRef]
- Roque, B.M.; Salwen, J.K.; Kinley, R.; Kebreab, E. Inclusion of asparagopsis armata in lactating dairy cows’ diet reduces enteric methane emission by over 50 percent. J. Clean. Prod. 2019, 234, 132–138. [Google Scholar] [CrossRef]
- Roque, B.M.; Venegas, M.; Kinley, R.D.; de Nys, R.; Duarte, T.L.; Yang, X.; Kebreab, E. Red seaweed (Asparagopsis taxiformis) supplementation reduces enteric methane by over 80 percent in beef steers. PLoS ONE 2021, 16, e0247820. [Google Scholar] [CrossRef] [PubMed]
- Stefenoni, H.A.; Räisänen, S.E.; Cueva, S.F.; Wasson, D.E.; Lage, C.F.A.; Melgar, A.; Fetter, M.E.; Smith, P.; Hennessy, M.; Vecchiarelli, B.; et al. Effects of the macroalga Asparagopsis taxiformis and oregano leaves on methane emission, rumen fermentation, and lactational performance of dairy cows. J. Dairy Sci. 2021, 104, 4157–4173. [Google Scholar] [CrossRef]
- Kinley, R.D.; Martinez-Fernandez, G.; Matthews, M.K.; de Nys, R.; Magnusson, M.; Tomkins, N.W. Mitigating the carbon footprint and improving productivity of ruminant livestock agriculture using a red seaweed. J. Clean. Prod. 2020, 259, 120836. [Google Scholar] [CrossRef]
- Lean, I.J.; Golder, H.M.; Grant, T.M.D.; Moate, P.J. A meta-analysis of effects of dietary seaweed on beef and dairy cattle performance and methane yield. PLoS ONE 2021, 16, e0249053. [Google Scholar] [CrossRef]
- Agency for Toxic Substances and Disease Registry (ATSDR). Toxicological profile for Bromoform and Chlorodibromomethane; U.S. Department of Health and Human Services, Public Health Service: Atlanta, GA, USA, 2005.
- Nilsson, J.; Martin, M. Exploratory environmental assessment of large-scale cultivation of seaweed used to reduce enteric methane emissions. Sustain. Prod. Consum. 2022, 30, 413–423. [Google Scholar] [CrossRef]
- Yáñez-Ruiz, D.R.; Abecia, L.; Newbold, C.J. Manipulating rumen microbiome and fermentation through interventions during early life: A review. Front. Microbiol. 2015, 6, 1133. [Google Scholar] [CrossRef]
- Chaucheyras, F.; Fonty, G.; Bertin, G.; Gouet, P. In vitro H2 utilization by a ruminal acetogenic bacterium cultivated alone or in association with an archaea methanogen is stimulated by a probiotic strain of Saccharomyces cerevisiae. Appl. Environ. Microbiol. 1995, 61, 3466–3467. [Google Scholar] [CrossRef] [PubMed]
- Alazzeh, A.Y.; Sultana, H.; Beauchemin, K.A.; Wang, Y.; Holo, H.; Harstad, O.M.; McAllister, T.A. Using strains of Propionibacteria to mitigate methane emissions in vitro. Acta Agric. Scand. Sect. A Anim. Sci. 2012, 62, 263–272. [Google Scholar] [CrossRef]
- Vyas, D.; Alazzeh, A.; McGinn, S.M.; McAllister, T.A.; Harstad, O.M.; Holo, H.; Beauchemin, K.A. Enteric methane emissions in response to ruminal inoculation of Propionibacterium strains in beef cattle fed a mixed diet. Anim. Prod. Sci. 2016, 56, 1035–1040. [Google Scholar] [CrossRef]
- Vyas, D.; McGeough, E.J.; Mohammed, R.; McGinn, S.M.; McAllister, T.A.; Beauchemin, K.A. Effects of Propionibacterium strains on ruminal fermentation, nutrient digestibility and methane emissions in beef cattle fed a corn grain finishing diet. Animal 2014, 8, 1807–1815. [Google Scholar] [CrossRef]
- Henderson, G.; Naylor, G.E.; Leahy, S.C.; Janssen, P.H. Presence of novel, potentially homoacetogenic bacteria in the rumen as determined by analysis of formyltetrahydrofolate synthetase sequences from ruminants. Appl. Environ. Microbiol. 2010, 76, 2058–2066. [Google Scholar] [CrossRef]
- Sazinsky, M.H.; Lippard, S.J. Methane monooxygenase: Functionalizing methane at iron and copper. Met. Ions Life Sci. 2015, 15, 205–256. [Google Scholar] [CrossRef]
- De Raphélis-Soissan, V.; Li, L.; Godwin, I.R.; Barnett, M.C.; Perdok, H.B.; Hegarty, R.S. Use of nitrate and Propionibacterium acidipropionici to reduce methane emissions and increase wool growth of Merino sheep. Anim. Prod. Sci. 2014, 54, 1860–1866. [Google Scholar] [CrossRef]
- Jeyanathan, J.; Martin, C.; Morgavi, D.P. The use of direct-fed microbials for mitigation of ruminant methane emissions: A review. Animal 2014, 8, 250–261. [Google Scholar] [CrossRef]
- Kittelmann, S.; Pinares-Patiño, C.S.; Seedorf, H.; Kirk, M.R.; McEwan, J.C.; Janssen, P.H. Natural variation in methane emission of sheep fed on a lucerne pellet diet is unrelated to rumen ciliate community type. Microbiology 2016, 162, 459–465. [Google Scholar] [CrossRef]
- Eadie, J.M. Inter-relationships between certain rumen ciliate protozoa. Microbiology 1962, 29, 579–588. [Google Scholar] [CrossRef]
- Dai, X.; Kalscheur, K.F.; Huhtanen, P.; Faciola, A.P. Effects of ruminal protozoa on methane emissions in ruminants—A meta-analysis. J. Dairy Sci. 2022, 105, 7482–7491. [Google Scholar] [CrossRef] [PubMed]
- Dai, X.; Faciola, A.P. Evaluating strategies to reduce ruminal protozoa and their impacts on nutrient utilization and animal performance in ruminants—A meta-analysis. Front. Microbiol. 2019, 10, 2648. [Google Scholar] [CrossRef] [PubMed]
- Guyader, J.; Eugène, M.; Nozière, P.; Morgavi, D.P.; Doreau, M.; Martin, C. Influence of rumen protozoa on methane emission in ruminants: A meta-analysis approach. Animal 2014, 8, 1816–1825. [Google Scholar] [CrossRef] [PubMed]
- Eugène, M.; Archimède, H.; Sauvant, D. Quantitative meta-analysis on the effects of defaunation of the rumen on growth, intake and digestion in ruminants. Livest. Prod. Sci. 2004, 85, 81–97. [Google Scholar] [CrossRef]
- Nguyen, S.H.; Nguyen, H.D.T.; Hegarty, R.S. Defaunation and its impacts on ruminal fermentation, enteric methane production and animal productivity. Livest. Res. Rural Dev. 2020, 32, 60. [Google Scholar]
- Hristov, A.N.; Oh, J.; Firkins, J.L.; Dijkstra, J.; Kebreab, E.; Waghorn, G.; Makkar, H.P.; Adesogan, A.T.; Yang, W.; Lee, C.; et al. Special topics—Mitigation of methane and nitrous oxide emissions from animal operations: I. A review of enteric methane mitigation options. J. Anim. Sci. 2013, 91, 5045–5069. [Google Scholar] [CrossRef]
- Tong, F.; Wang, T.; Gao, N.L.; Liu, Z.; Cui, K.; Duan, Y.; Wu, S.; Luo, Y.; Li, Z.; Yang, C.; et al. The microbiome of the buffalo digestive tract. Nat. Commun. 2022, 13, 823. [Google Scholar] [CrossRef]
- Morvan, B.; Bonnemoy, F.; Fonty, G.; Gouet, P. Quantitative determination of H2-utilizing acetogenic and sulfate-reducing bacteria and methanogenic archaea from digestive tract of different mammals. Curr. Microbiol. 1996, 32, 129–133. [Google Scholar] [CrossRef]
- Ungerfeld, E.M. Corrigendum: Shifts in metabolic hydrogen sinks in the methanogenesis-inhibited ruminal fermentation: A meta-analysis. Front. Microbiol. 2015, 6, 538. [Google Scholar] [CrossRef]
- Ungerfeld, E.M.; Aedo, M.F.; Martínez, E.D.; Saldivia, M. Inhibiting methanogenesis in rumen batch cultures did not increase the recovery of metabolic hydrogen in microbial amino acids. Microorganisms 2019, 7, 115. [Google Scholar] [CrossRef]
- Aguilar-Marin, S.B.; Betancur-Murillo, C.L.; Isaza, G.A.; Mesa, H.; Jovel, J. Lower methane emissions were associated with higher abundance of ruminal Prevotella in a cohort of Colombian buffalos. BMC Microbiology 2020, 20, 364. [Google Scholar] [CrossRef] [PubMed]
- Bandarupalli, V.V.K.; St-Pierre, B. Identification of a candidate starch utilizing strain of prevotella albensis from bovine rumen. Microorganisms 2020, 8, 2005. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.; Gao, X.; Yang, Y.; Zou, C.; Yang, Y.; Lin, B. A comparative study on rumen ecology of water buffalo and cattle calves under similar feeding regime. Vet. Med. Sci. 2020, 6, 746–754. [Google Scholar] [CrossRef] [PubMed]
- Subharat, S.; Shu, D.; Zheng, T.; Buddle, B.M.; Kaneko, K.; Hook, S.; Janssen, P.H.; Wedlock, D.N. Vaccination of sheep with a methanogen protein provides insight into levels of antibody in saliva needed to target ruminal methanogens. PLoS ONE 2016, 11, e0159861. [Google Scholar] [CrossRef]
- Cook, S.R.; Maiti, P.K.; Chaves, A.V.; Benchaar, C.; Beauchemin, K.A.; McAllister, T.A. Avian (IgY) anti-methanogen antibodies for reducing ruminal methane production: In vitro assessment of their effects. Aust. J. Exp. Agric. 2008, 48, 260–264. [Google Scholar] [CrossRef]
- Baca-González, V.; Asensio-Calavia, P.; González-Acosta, S.; Pérez de la Lastra, J.M.; Morales de la Nuez, A. Are vaccines the solution for methane emissions from ruminants? A systematic review. Vaccines 2020, 8, 460. [Google Scholar] [CrossRef]
- Baker, S.K.; Perth, W. Method for Improving Utilization of Nutrients by Ruminant or Ruminant-like Animals. US6036950A, 14 March 2000. [Google Scholar]
- Clark, H.; Wright, A.-D.; Joblin, K.; Molano, G.; Cavanagh, A.; Peters, J. Field testing an Australian developed anti-methanogen vaccine in growing ewe lambs. In Proceedings of the Workshop on the Science of Atmospheric Trace Gases, Wellington, New Zealand, 18–19 March 2004; Clarkson, T.S., Ed.; pp. 107–108. [Google Scholar]
- Wedlock, D.N.; Janssen, P.H.; Leahy, S.C.; Shu, D.; Buddle, B.M. Progress in the development of vaccines against rumen methanogens. Animal 2013, 7 (Suppl. 2), 244–252. [Google Scholar] [CrossRef]
- Williams, Y.J.; Rea, S.M.; Popovski, S.; Pimm, C.L.; Williams, A.J.; Toovey, A.F.; Skillman, L.C.; Wright, A.D. Reponses of sheep to a vaccination of entodinial or mixed rumen protozoal antigens to reduce rumen protozoal numbers. Br. J. Nutr. 2008, 99, 100–109. [Google Scholar] [CrossRef]
- Wright, A.D.; Kennedy, P.; O’Neill, C.J.; Toovey, A.F.; Popovski, S.; Rea, S.M.; Pimm, C.L.; Klein, L. Reducing methane emissions in sheep by immunization against rumen methanogens. Vaccine 2004, 22, 3976–3985. [Google Scholar] [CrossRef]
- Zhang, L.; Huang, X.; Xue, B.; Peng, Q.; Wang, Z.; Yan, T.; Wang, L. Immunization against rumen methanogenesis by vaccination with a new recombinant protein. PLoS ONE 2015, 10, e0140086. [Google Scholar] [CrossRef]
- Williams, Y.J.; Popovski, S.; Rea, S.M.; Skillman, L.C.; Toovey, A.F.; Northwood, K.S.; Wright, A.D. A vaccine against rumen methanogens can alter the composition of archaeal populations. Appl. Environ. Microbiol. 2009, 75, 1860–1866. [Google Scholar] [CrossRef] [PubMed]
- Khampa, S.; Wanapat, M. Manipulation of rumen fermentation with organic acids supplementation in ruminants raised in the tropics. Pak. J. Nutr. 2007, 6, 20–27. [Google Scholar]
- Alvarez-Hess, P.S.; Little, S.M.; Moate, P.J.; Jacobs, J.L.; Beauchemin, K.A.; Eckard, R.J. A partial life cycle assessment of the greenhouse gas mitigation potential of feeding 3-nitrooxypropanol and nitrate to cattle. Agric. Syst. 2019, 169, 14–23. [Google Scholar] [CrossRef]
- Duin, E.C.; Wagner, T.; Shima, S.; Prakash, D.; Cronin, B.; Yáñez-Ruiz, D.R.; Duval, S.; Rümbeli, R.; Stemmler, R.T.; Thauer, R.K.; et al. Mode of action uncovered for the specific reduction of methane emissions from ruminants by the small molecule 3-nitrooxypropanol. Proc. Natl. Acad. Sci. USA 2016, 113, 6172–6177. [Google Scholar] [CrossRef]
- EFSA Panel on Additives and Products or Substances used in Animal Feed (FEEDAP); Bampidis, V.; Azimonti, G.; Bastos, M.D.L.; Christensen, H.; Dusemund, B.; Fašmon Durjava, M.; Kouba, M.; López-Alonso, M. Safety and efficacy of a feed additive consisting of 3-nitrooxypropanol (Bovaer® 10) for ruminants for milk production and reproduction (DSM Nutritional Products Ltd.). EFSA J. 2021, 19, e06905. [Google Scholar] [CrossRef] [PubMed]
- Yu, G.; Beauchemin, K.A.; Dong, R. A review of 3-nitrooxypropanol for enteric methane mitigation from ruminant livestock. Animals 2021, 11, 3540. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.; Wang, K.; Nan, X.; Cai, M.; Yang, L.; Xiong, B.; Zhao, Y. Synergistic effects of 3-nitrooxypropanol with fumarate in the regulation of propionate formation and methanogenesis in dairy cows in vitro. Appl. Environ. Microbiol. 2022, 88, e0190821. [Google Scholar] [CrossRef]
- Dijkstra, J.; Bannink, A.; France, J.; Kebreab, E.; van Gastelen, S. Short communication: Antimethanogenic effects of 3-nitrooxypropanol depend on supplementation dose, dietary fiber content, and cattle type. J. Dairy Sci. 2018, 101, 9041–9047. [Google Scholar] [CrossRef]
- Hernández, J.; Benedito, J.L.; Abuelo, A.; Castillo, C. Ruminal acidosis in feedlot: From aetiology to prevention. Sci. World J. 2014, 2014, 702572. [Google Scholar] [CrossRef]
- Foley, P.A.; Kenny, D.A.; Callan, J.J.; Boland, T.M.; O’Mara, F.P. Effect of DL-malic acid supplementation on feed intake, methane emission, and rumen fermentation in beef cattle. J. Anim. Sci. 2009, 87, 1048–1057. [Google Scholar] [CrossRef]
- Mohammed, N.; Lila, Z.A.; Ajisaka, N.; Hara, K.; Mikuni, K.; Hara, K.; Kanda, S.; Itabashi, H. Inhibition of ruminal microbial methane production by beta-cyclodextrin iodopropane, malate and their combination in vitro. J. Anim. Physiol. Anim. Nutr. 2004, 88, 188–195. [Google Scholar] [CrossRef] [PubMed]
- Ebrahimia, S.H.; Mohini, M.; Singhal, K.K.; Miri, V.H.; Tyagi, A.K. Evaluation of complementary effects of 9,10-anthraquinone and fumaric acid on methanogenesis and ruminal fermentation in vitro. Arch. Anim. Nutr. 2011, 65, 267–277. [Google Scholar] [CrossRef] [PubMed]
- Van Zijderveld, S.M.; Gerrits, W.J.; Dijkstra, J.; Newbold, J.R.; Hulshof, R.B.; Perdok, H.B. Persistency of methane mitigation by dietary nitrate supplementation in dairy cows. J. Dairy Sci. 2011, 94, 4028–4038. [Google Scholar] [CrossRef] [PubMed]
- Fewtrell, L. Drinking-water nitrate, methemoglobinemia, and global burden of disease: A discussion. Environ. Health Perspect. 2004, 112, 1371–1374. [Google Scholar] [CrossRef] [PubMed]
- Hook, S.E.; Steele, M.A.; Northwood, K.S.; Wright, A.D.; McBride, B.W. Impact of high-concentrate feeding and low ruminal pH on methanogens and protozoa in the rumen of dairy cows. Microb. Ecol. 2011, 62, 94–105. [Google Scholar] [CrossRef] [PubMed]
- Schären, M.; Drong, C.; Kiri, K.; Riede, S.; Gardener, M.; Meyer, U.; Hummel, J.; Urich, T.; Breves, G.; Dänicke, S. Differential effects of monensin and a blend of essential oils on rumen microbiota composition of transition dairy cows. J. Dairy Sci. 2017, 100, 2765–2783. [Google Scholar] [CrossRef]
- National Research Council. Nutrient Requirements of Beef Cattle, 7th ed.; National Academies Press: Washington, DC, USA, 2000.
- EFSA Panel on Additives and Products or Substances used in Animal Feed. Opinion of the scientific panel on additives and products or substances used in animal feed on the safety and efficacy of the coccidiostat Elancoban® (monensin sodium) as a feed additive for calves for rearing and cattle for fattening in accordance with Regulation (EC) No 1831/2003. EFSA J. 2006, 4, 387. [Google Scholar]
- Ranga Niroshan Appuhamy, J.A.D.; Strathe, A.B.; Jayasundara, S.; Wagner-Riddle, C.; Dijkstra, J.; France, J.; Kebreab, E. Anti-methanogenic effects of monensin in dairy and beef cattle: A meta-analysis. J. Dairy Sci. 2013, 96, 5161–5173. [Google Scholar] [CrossRef]
- Guan, H.; Wittenberg, K.M.; Ominski, K.H.; Krause, D.O. Efficacy of ionophores in cattle diets for mitigation of enteric methane1. J. Anim. Sci. 2006, 84, 1896–1906. [Google Scholar] [CrossRef]
- Hook, S.E.; Northwood, K.S.; Wright, A.D.; McBride, B.W. Long-term monensin supplementation does not significantly affect the quantity or diversity of methanogens in the rumen of the lactating dairy cow. Appl. Environ. Microbiol. 2009, 75, 374–380. [Google Scholar] [CrossRef]
- Hook, S.E.; Wright, A.-D.G.; McBride, B.W. Methanogens: Methane producers of the rumen and mitigation strategies. Archaea 2010, 2010, 945785. [Google Scholar] [CrossRef]
- Odongo, N.E.; Bagg, R.; Vessie, G.; Dick, P.; Or-Rashid, M.M.; Hook, S.E.; Gray, J.T.; Kebreab, E.; France, J.; McBride, B.W. Long-term effects of feeding monensin on methane production in lactating dairy cows. J. Dairy Sci. 2007, 90, 1781–1788. [Google Scholar] [CrossRef] [PubMed]
- Committee for Medicinal Products for Veterinary Use. Monensin (Cattle, Including Dairy Cows). European Medicines Agency Veterinary Medicines and Inspections, 2007. Available online: https://www.ema.europa.eu/en/medicines/ema_group_types/ema_document-maximum_residue_limits_report?page=4 (accessed on 10 February 2023).
- Pickering, N.K.; Dodds, K.G.; Blair, H.T.; Hickson, R.E.; Johnson, P.L.; McEwan, J.C. Genetic parameters for production traits in New Zealand dual-purpose sheep, with an emphasis on dagginess1. J. Anim. Sci. 2012, 90, 1411–1420. [Google Scholar] [CrossRef] [PubMed]
- Pszczola, M.; Rzewuska, K.; Mucha, S.; Strabel, T. Heritability of methane emissions from dairy cows over a lactation measured on commercial farms. J. Anim. Sci. 2017, 95, 4813–4819. [Google Scholar] [CrossRef] [PubMed]
- Rowe, S.; Hickey, S.; Jonker, A.; Hess, M.; Janssen, P.; Johnson, T.; Bryson, B.; Knowler, K.; Pinares-Patino, C.; Bain, W.; et al. Selection for divergent methane yield in New Zealand sheep—A ten year perspective. In Proceedings of the 23rd Conference of the Association for the Advancement of Animal Breeding and Genetics (AAABG), Armidale, NSW, Australia, 27 October–1 November 2019; pp. 306–309. [Google Scholar]
- Pinares-Patiño, C.S.; Hickey, S.M.; Young, E.A.; Dodds, K.G.; MacLean, S.; Molano, G.; Sandoval, E.; Kjestrup, H.; Harland, R.; Hunt, C.; et al. Heritability estimates of methane emissions from sheep. Animal 2013, 7 (Suppl. 2), 316–321. [Google Scholar] [CrossRef]
- Sypniewski, M.; Strabel, T.; Pszczola, M. Genetic variability of methane production and concentration measured in the breath of polish holstein-friesian cattle. Animals 2021, 11, 3175. [Google Scholar] [CrossRef]
- Manzanilla-Pech, C.I.V.; L⊘vendahl, P.; Mansan Gordo, D.; Difford, G.F.; Pryce, J.E.; Schenkel, F.; Wegmann, S.; Miglior, F.; Chud, T.C.; Moate, P.J.; et al. Breeding for reduced methane emission and feed-efficient Holstein cows: An international response. J. Dairy Sci. 2021, 104, 8983–9001. [Google Scholar] [CrossRef]
- Rowe, J.B.; Gill, S.; Banks, R.G.; van der Werf, J.H.J. Genomics for the Australian sheep industry: From design to delivery. Proc. Assoc. Adv. Anim. Breed. Genet. 2013, 20, 14–17. [Google Scholar]
- Bain, W.; Bezuidenhout, L.; Jopson, N.; Pinares-Patino, C.; McEwan, J. Rumen differences between sheep identified as being low or high methane emitters. In Proceedings of the 10th World Congress on Genetics Applied to Livestock, Vancouver, BC, Canada, 17–22 August 2014. [Google Scholar]
- Hickey, S.M.; Bain, W.E.; Bilton, T.P.; Greer, G.J.; Elmes, S.; Bryson, B.; Pinares-Patiño, C.S.; Wing, J.; Jonker, A.; Young, E.A.; et al. Impact of breeding for reduced methane emissions in New Zealand sheep on maternal and health traits. Front. Genet. 2022, 13, 2165. [Google Scholar] [CrossRef]
- De Haas, Y.; Veerkamp, R.F.; de Jong, G.; Aldridge, M.N. Selective breeding as a mitigation tool for methane emissions from dairy cattle. Animal 2021, 15, 100294. [Google Scholar] [CrossRef]
- United States Environmental Protection Agency (USEPA). Inventory of US Greenhouse Gas Emissions and Sinks 1990–2019; United States Environmental Protection Agency (USEPA): Washington, DC, USA, 2021.
- FAO. World Food and Agriculture—Statistical Yearbook; FAO: Rome, Italy, 2022. [Google Scholar] [CrossRef]
- FAO. FAOSTAT Statistical Database; FAO: Rome, Italy, 2022. [Google Scholar]
- USDA’s Foreign Agricultural Service (FAS). 2022. Available online: https://gain.fas.usda.gov/#/search (accessed on 5 December 2022).
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).