In-Field Citrus Disease Classification via Convolutional Neural Network from Smartphone Images
Abstract
1. Introduction
2. Dataset Construction
3. Methods
3.1. Network-Based Transfer Learning
3.2. Proposed 13-Layer Convolutional Neural Network (CNN13)
4. Experimental Results and Analysis
4.1. Evaluation Metrics
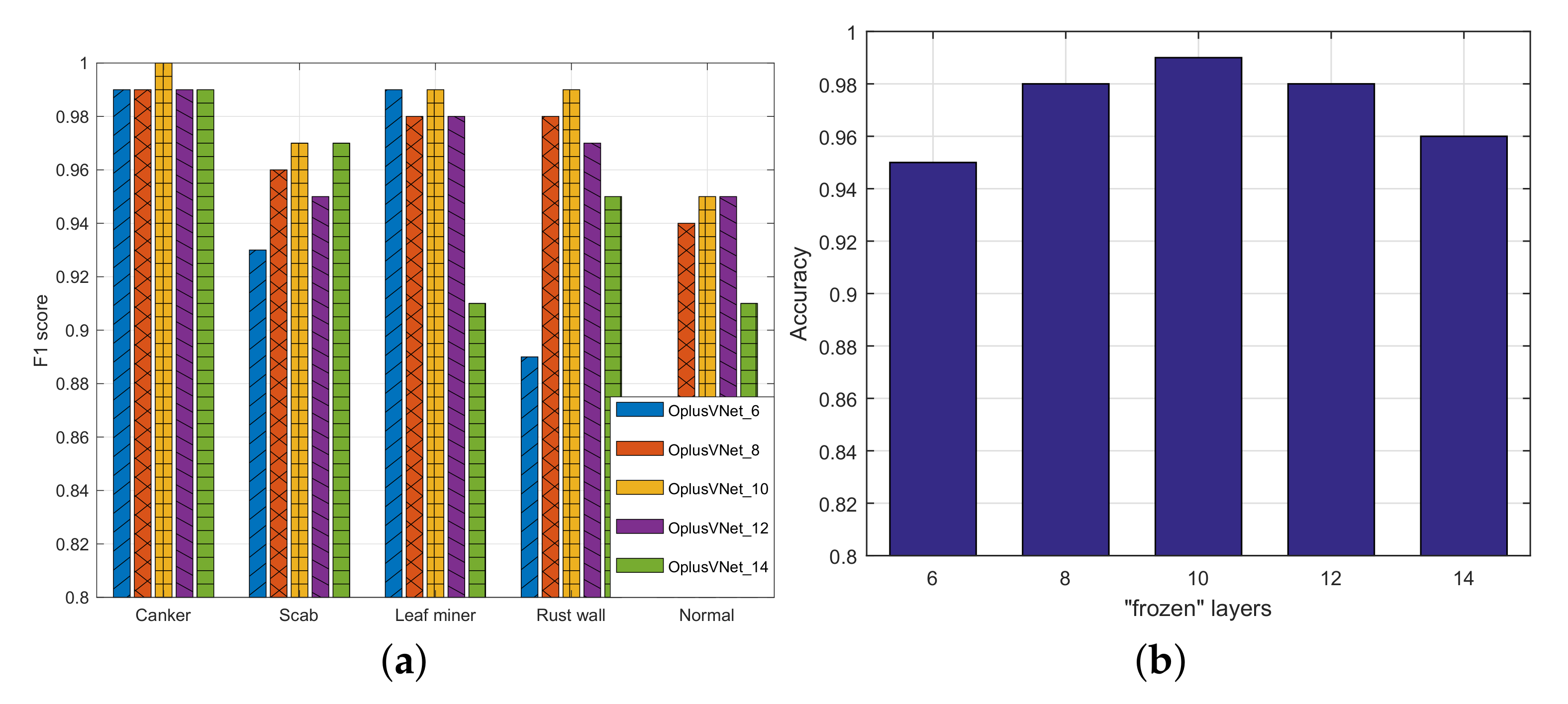
4.2. Experiments with OplusVNet with Different Frozen Mechanisms
4.3. Comparison with State-of-the-Art Networks
4.4. OplusVNet Network Performance Analysis
4.4.1. Experimental Results of Proposed OplusVNet on Small Datasets
4.4.2. Experimental Results of Different Network Models on Small Datasets
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Khan, M.A.; Akram, T.; Sharif, M.; Awais, M.; Javed, K.; Ali, H.; Saba, T. CCDF: Automatic system for segmentation and recognition of fruit crops diseases based on correlation coefficient and deep CNN features. Comput. Electron. Agric. 2018, 155, 220–236. [Google Scholar] [CrossRef]
- Moriya, E.A.S.; Imai, N.N.; Tommaselli, A.M.G.; Berveglieri, A.; Santos, G.H.; Soares, M.A.; Marino, M.; Reis, T.T. Detection and mapping of trees infected with citrus gummosis using UAV hyperspectral data. Comput. Electron. Agric. 2021, 188, 106298. [Google Scholar] [CrossRef]
- Ngugi, L.C.; Abelwahab, M.; Abo-Zahhad, M. Recent advances in image processing techniques for automated leaf pest and disease recognition: A review. Inf. Process. Agric. 2021, 8, 27–51. [Google Scholar] [CrossRef]
- Abdu, A.M.; Mokji, M.M.; Sheikh, U.U. Automatic vegetable disease identification approach using individual lesion features. Comput. Electron. Agric. 2020, 176, 105660. [Google Scholar] [CrossRef]
- Nanehkaran, Y.A.; Zhang, D.; Chen, J.; Tian, Y.; Al-Nabhan, N. Recognition of plant leaf diseases based on computer vision. J. Ambient. Intell. Humaniz. Comput. 2020, 1–18. [Google Scholar] [CrossRef]
- Jagan, K.; Balasubramanian, M.; Palanivel, S. Detection and Recognition of Diseases from Paddy Plant Leaf Images. Int. J. Comput. Appl. 2016, 144, 34–41. [Google Scholar] [CrossRef]
- Hsiao, J.K.; Kang, L.W.; Chang, C.L.; Hsu, C.Y.; Chen, C.Y. Learning sparse representation for leaf image recognition. In Proceedings of the 2014 IEEE International Conference on Consumer Electronics, Taipei, Taiwan, 26–28 May 2014; pp. 209–210. [Google Scholar] [CrossRef]
- Zhang, S.; Wu, X.; You, Z.; Zhang, L. Leaf image based cucumber disease recognition using sparse representation classification. Comput. Electron. Agric. 2017, 134, 135–141. [Google Scholar] [CrossRef]
- Liu, J.; Wang, X. Plant diseases and pests detection based on deep learning: A review. Plant Methods 2021, 17, 22. [Google Scholar] [CrossRef]
- Chen, J.; Zhang, D.; Nanehkaran, Y. Identifying plant diseases using deep transfer learning and enhanced lightweight network. Multimed. Tools Appl. 2020, 79, 31497–31515. [Google Scholar] [CrossRef]
- Chen, J.; Yin, H.; Zhang, D. A self-adaptive classification method for plant disease detection using GMDH-Logistic model. Sustain. Comput. Inform. Syst. 2020, 28, 100415. [Google Scholar] [CrossRef]
- Chen, J.; Chen, J.; Zhang, D.; Sun, Y.; Nanehkaran, Y. Using deep transfer learning for image-based plant disease identification. Comput. Electron. Agric. 2020, 173, 105393. [Google Scholar] [CrossRef]
- Chen, J.; Zhang, D.; Nanehkaran, Y.; Li, D. Detection of rice plant diseases based on deep transfer learning. J. Sci. Food Agric. 2020, 100. [Google Scholar] [CrossRef]
- Haque, M.; Marwaha, S.; Deb, C.; Nigam, S.; Arora, A.; Hooda, K.; Soujanya, P.L.; Aggarwal, S.; Lall, B.; Kumar, M.; et al. Deep learning-based approach for identification of diseases of maize crop. Sci. Rep. 2022, 12, 6334. [Google Scholar] [CrossRef] [PubMed]
- Kundu, N.; Rani, G.; Dhaka, V.S.; Gupta, K.; Nayak, S.C.; Verma, S.; Ijaz, M.F.; Woniak, M. IoT and Interpretable Machine Learning Based Framework for Disease Prediction in Pearl Millet. Sensors 2021, 21, 5386. [Google Scholar] [CrossRef] [PubMed]
- Rehman, M.Z.U.; Ahmed, F.; Khan, M.A.; Tariq, U.; Jamal, S.S.; Ahmad, J.; Hussain, I. Classification of Citrus Plant Diseases Using Deep Transfer Learning. Comput. Mater. Contin. 2022, 70, 1401–1417. [Google Scholar] [CrossRef]
- Nazari, K.; Ebadi, M.; Berahmand, K. Diagnosis of Alternaria disease and Leafminer pest on Tomato Leaves using Image Processing Techniques. J. Sci. Food Agric. 2022. [Google Scholar] [CrossRef] [PubMed]
- Dhiman, P.; Kukreja, V.; Manoharan, P.; Kaur, A.; Kamruzzaman, M.M.; Dhaou, I.B.; Iwendi, C. A Novel Deep Learning Model for Detection of Severity Level of the Disease in Citrus Fruits. Electronics 2022, 11, 495. [Google Scholar] [CrossRef]
- Mohanty, S.P.; Hughes, D.P.; Salath, M. Using Deep Learning for Image-Based Plant Disease Detection. Front. Plant Sci. 2016, 7, 1419. [Google Scholar] [CrossRef]
- Sladojevic, S.; Arsenovic, M.; Anderla, A.; Culibrk, D.; Stefanovic, D. Deep Neural Networks Based Recognition of Plant Diseases by Leaf Image Classification. Comput. Intell. Neurosci. 2016, 2016, 3289801. [Google Scholar] [CrossRef]
- Tan, C.; Sun, F.; Kong, T.; Zhang, W.; Yang, C.; Liu, C. A Survey on Deep Transfer Learning. In Proceedings of the 27th International Conference on Artificial Neural Networks, Rhodes, Greece, 4–7 October 2018; pp. 270–279. [Google Scholar]
- Thenmozhi, K.; Reddy, U.S. Crop pest classification based on deep convolutional neural network and transfer learning. Comput. Electron. Agric. 2019, 164, 104906. [Google Scholar] [CrossRef]
- Selvaraj, M.G.; Vergara, A.; Ruiz, H.; Safari, N.; Elayabalan, S.; Ocimati, W.; Blomme, G. AI-powered banana diseases and pest detection. Plant Methods 2019, 15, 92. [Google Scholar] [CrossRef]
- Coulibaly, S.; Kamsu-Foguem, B.; Kamissoko, D.; Traore, D. Deep neural networks with transfer learning in millet crop images. Comput. Ind. 2019, 108, 115–120. [Google Scholar] [CrossRef]
- Barman, U.; Choudhury, R.D.; Sahu, D.; Barman, G.G. Comparison of convolution neural networks for smartphone image based real time classification of citrus leaf disease. Comput. Electron. Agric. 2020, 177, 105661. [Google Scholar] [CrossRef]
- Morteza, K.; Ezzatollah, A.A.A.; Ehsan, K. Citrus pests classification using an ensemble of deep learning models. Comput. Electron. Agric. 2021, 186, 106192. [Google Scholar]
- Simonyan, K.; Zisserman, A. Very Deep Convolutional Networks for Large-Scale Image Recognition. arXiv 2014, arXiv:1409.1556. [Google Scholar]
- Krizhevsky, A.; Sutskever, I.; Hinton, G.E. ImageNet Classification with Deep Convolutional Neural Networks. In Proceedings of the Annual Conference on Neural Information Processing Systems, Lake Tahoe, NV, USA, 3–8 December 2012; Volume 25. [Google Scholar]
- Krizhevsky, A.; Sutskever, I.; Hinton, G.E. ImageNet Classification with Deep Convolutional Neural Networks. Commun. ACM 2017, 60, 84–90. [Google Scholar] [CrossRef]
- Srivastava, N.; Hinton, G.; Krizhevsky, A.; Sutskever, I.; Salakhutdinov, R. Dropout: A Simple Way to Prevent Neural Networks from Overfitting. J. Mach. Learn. Res. 2014, 15, 1929–1958. [Google Scholar]
- Ding, X.; Zhang, X.; Ma, N.; Han, J.; Ding, G.; Sun, J. Repvgg: Making vgg-style convnets great again. In Proceedings of the IEEE/CVF Conference on Computer Vision and Pattern Recognition, Nashville, TN, USA, 20–25 June 2021; pp. 13733–13742. [Google Scholar]


| Canker | Scab | Leaf Miner | Rust Wall | Normal | |
|---|---|---|---|---|---|
| Total | 1040 | 293 | 320 | 210 | 202 |
| Training set | 624 | 176 | 192 | 126 | 122 |
| Validation set | 208 | 59 | 64 | 42 | 40 |
| Test set | 208 | 58 | 64 | 42 | 40 |
| Number of Layers | VGG16 | Output Shape | Number of Parameters | |
|---|---|---|---|---|
| Frozen layer | 1 | 1792 | ||
| 2 | 36,928 | |||
| 3 | 0 | |||
| 4 | 73,856 | |||
| 5 | 147,584 | |||
| 6 | 0 | |||
| 7 | 295,168 | |||
| 8 | 590,080 | |||
| 9 | 590,080 | |||
| 10 | 0 | |||
| Non-freezing layer | 11 | 1,180,160 | ||
| 12 | 2,359,808 | |||
| 13 | 2,359,808 | |||
| 14 | 0 | |||
| 15 | 2,359,808 | |||
| 16 | 2,359,808 | |||
| 17 | 2,359,808 | |||
| 18 | 0 |
| Layers | Layer Type | Core Size/Number | Convolution Step | Output Shape | Parameters |
|---|---|---|---|---|---|
| 1 | 294,976 | ||||
| 64 | |||||
| 2 | 36,928 | ||||
| 64 | |||||
| 3 | 0 | ||||
| 4 | 73,856 | ||||
| 128 | |||||
| 5 | 147,584 | ||||
| 128 | |||||
| 6 | 0 | ||||
| 7 | 259,168 | ||||
| 256 | |||||
| 8 | 0 | ||||
| 9 | 590,080 | ||||
| 256 | |||||
| 10 | 0 | ||||
| 11 | 256 | 0 | |||
| 12 | 512 | 131,584 | |||
| 256 | |||||
| 13 | 5 | 2565 | |||
| Total number of parameters | 1,538,149 | ||||
| Methods | F1 Score | Accuracy | ||||
|---|---|---|---|---|---|---|
| Canker | Scab | Leaf Miner | Rust Wall | Normal | ||
| AlexNet | 0.98 | 0.88 | 0.93 | 0.87 | 0.83 | 0.93 |
| TL-VGG16 | 0.94 | 0.91 | 0.74 | 0.85 | 0.68 | 0.88 |
| RepVGG | 0.99 | 0.97 | 0.96 | 0.94 | 0.93 | 0.97 |
| OplusVNet | 1.00 | 0.97 | 0.99 | 0.99 | 0.95 | 0.99 |
| Number | Number of Frozen Layers | ||||||
|---|---|---|---|---|---|---|---|
| 4 | 6 | 8 | 10 | 12 | 14 | 16 | |
| 170 | 0.97 | 0.97 | 0.97 | 0.97 | 0.96 | 0.94 | 0.91 |
| 120 | 0.95 | 0.96 | 0.94 | 0.95 | 0.96 | 0.91 | 0.89 |
| 70 | 0.93 | 0.94 | 0.94 | 0.92 | 0.93 | 0.89 | 0.89 |
| Number | Network | F1 Score | Accuracy | ||||
|---|---|---|---|---|---|---|---|
| Canker | Scab | Leaf Miner | Rust Wall | Normal | |||
| 170 | AlexNet | 0.87 | 0.75 | 0.85 | 0.91 | 0.74 | 0.83 |
| TL-VGG16 | 0.92 | 0.90 | 0.87 | 0.85 | 0.83 | 0.87 | |
| OplusVNet_10 | 0.98 | 0.97 | 0.97 | 0.97 | 0.98 | 0.97 | |
| OplusVNet_6 | 0.98 | 0.97 | 0.97 | 0.97 | 0.98 | 0.97 | |
| 120 | AlexNet | 0.81 | 0.75 | 0.85 | 0.87 | 0.70 | 0.80 |
| TL-VGG16 | 0.90 | 0.90 | 0.84 | 0.88 | 0.75 | 0.85 | |
| OplusVNet_10 | 0.93 | 0.94 | 0.97 | 0.96 | 0.96 | 0.95 | |
| OplusVNet_6 | 0.94 | 0.95 | 0.98 | 0.97 | 0.97 | 0.96 | |
| 70 | AlexNet | 0.81 | 0.63 | 0.84 | 0.84 | 0.62 | 0.75 |
| TL-VGG16 | 0.86 | 0.82 | 0.79 | 0.79 | 0.64 | 0.80 | |
| OplusVNet_10 | 0.93 | 0.90 | 0.94 | 0.90 | 0.93 | 0.92 | |
| OplusVNet_6 | 0.95 | 0.92 | 0.97 | 0.91 | 0.95 | 0.94 | |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Yang, C.; Teng, Z.; Dong, C.; Lin, Y.; Chen, R.; Wang, J. In-Field Citrus Disease Classification via Convolutional Neural Network from Smartphone Images. Agriculture 2022, 12, 1487. https://doi.org/10.3390/agriculture12091487
Yang C, Teng Z, Dong C, Lin Y, Chen R, Wang J. In-Field Citrus Disease Classification via Convolutional Neural Network from Smartphone Images. Agriculture. 2022; 12(9):1487. https://doi.org/10.3390/agriculture12091487
Chicago/Turabian StyleYang, Changcai, Zixuan Teng, Caixia Dong, Yaohai Lin, Riqing Chen, and Jian Wang. 2022. "In-Field Citrus Disease Classification via Convolutional Neural Network from Smartphone Images" Agriculture 12, no. 9: 1487. https://doi.org/10.3390/agriculture12091487
APA StyleYang, C., Teng, Z., Dong, C., Lin, Y., Chen, R., & Wang, J. (2022). In-Field Citrus Disease Classification via Convolutional Neural Network from Smartphone Images. Agriculture, 12(9), 1487. https://doi.org/10.3390/agriculture12091487










