Various Perspectives on Microbial Lipase Production Using Agri-Food Waste and Renewable Products
Abstract
1. Structure and Specificity of Lipases
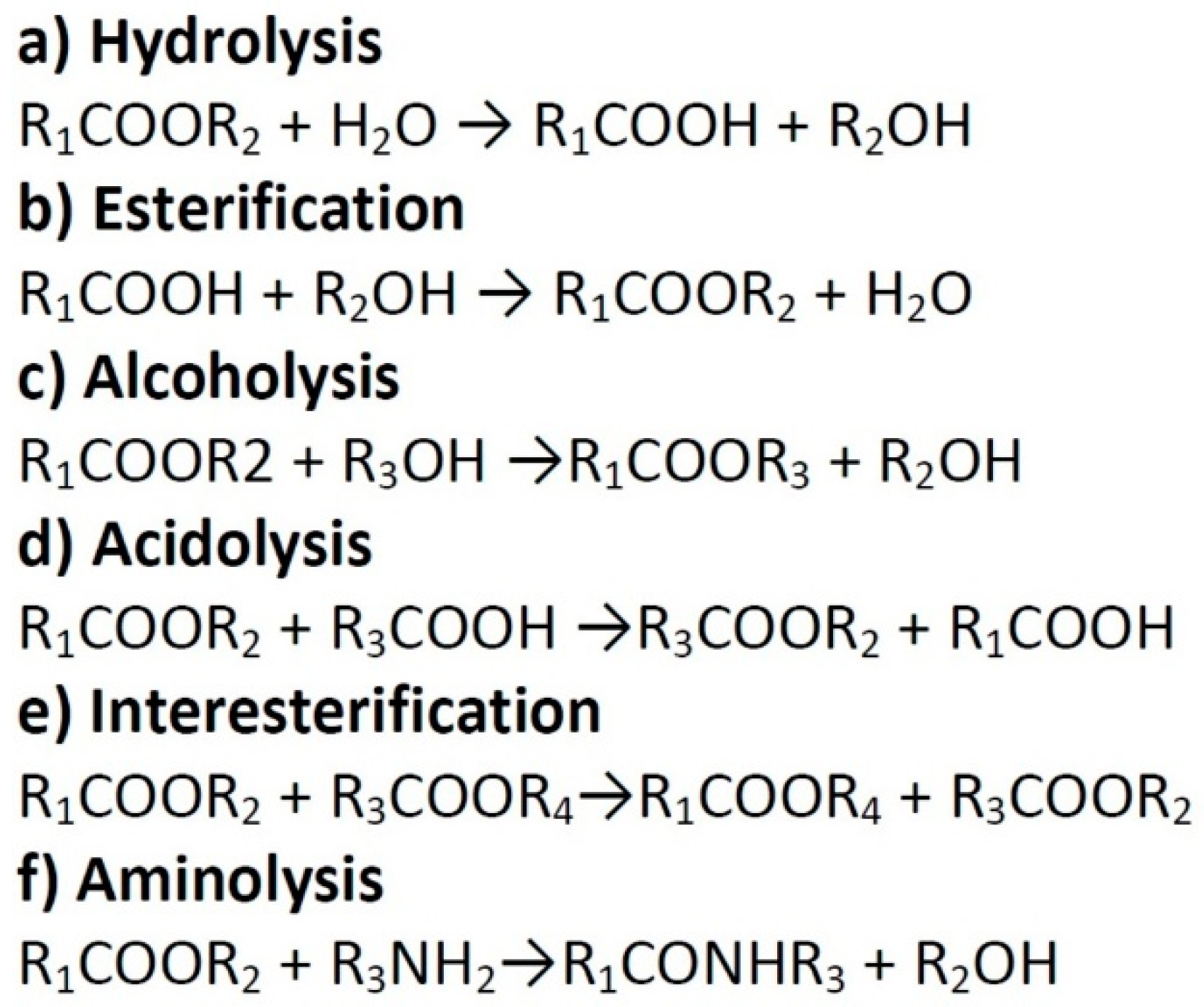
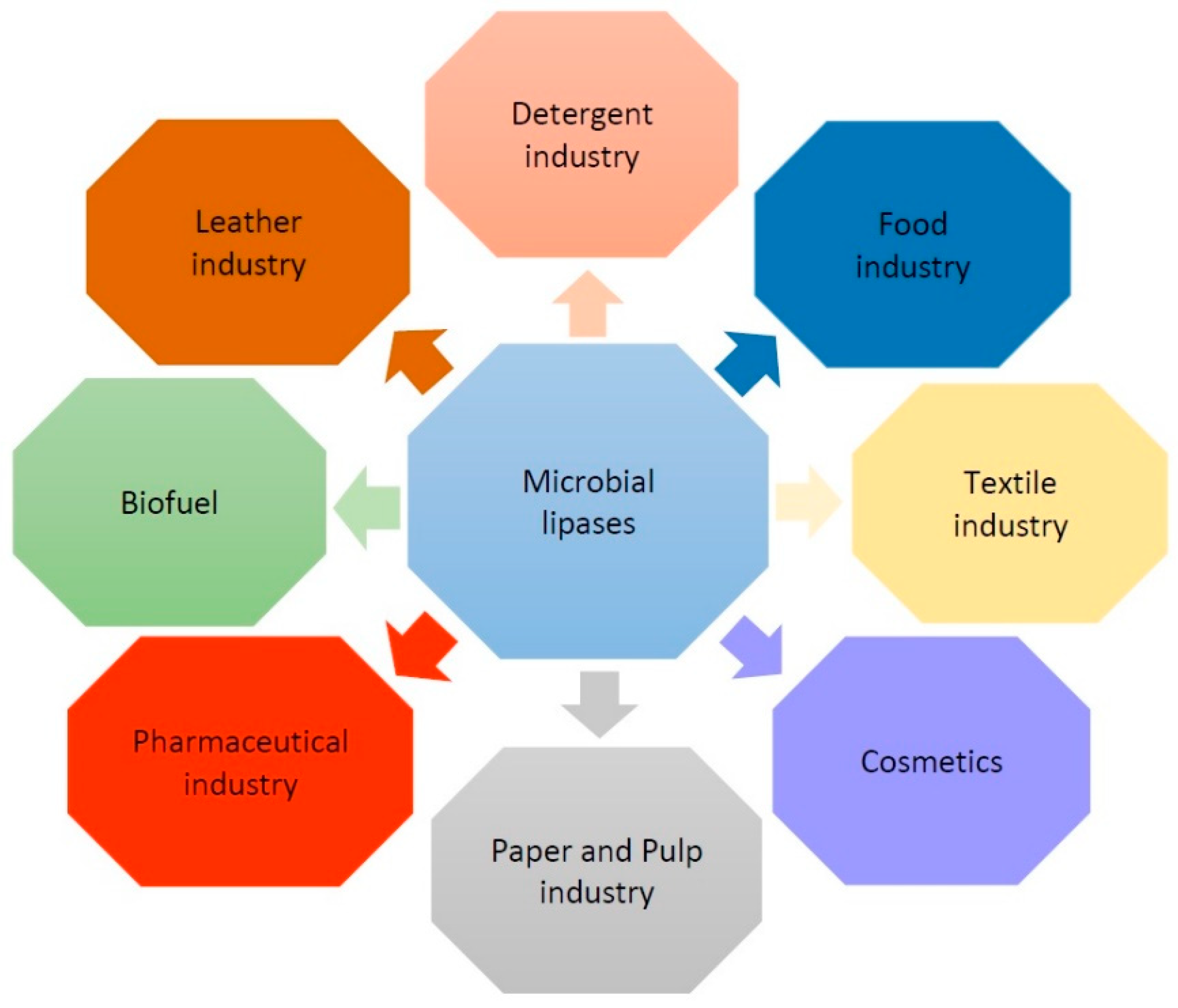
2. Functions and Applications of Lipases
3. Fermentation Techniques
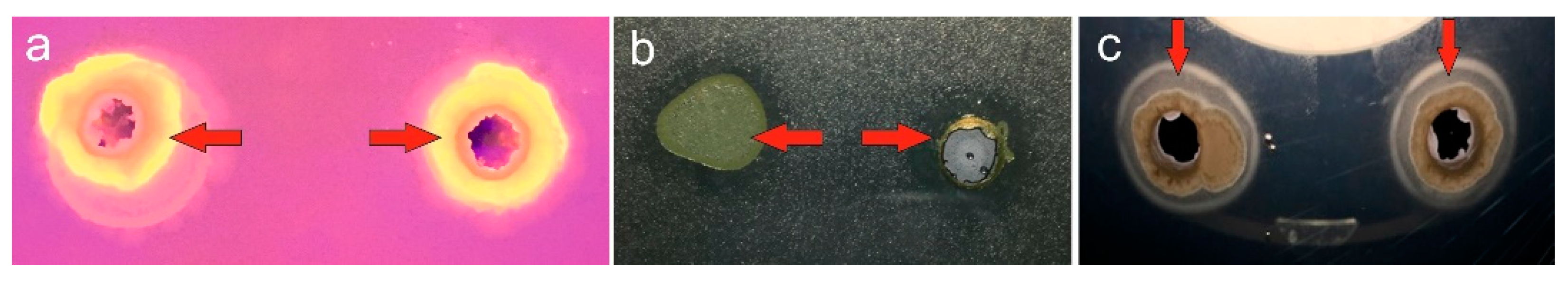
4. Methods of Lipolytic Activity Determination
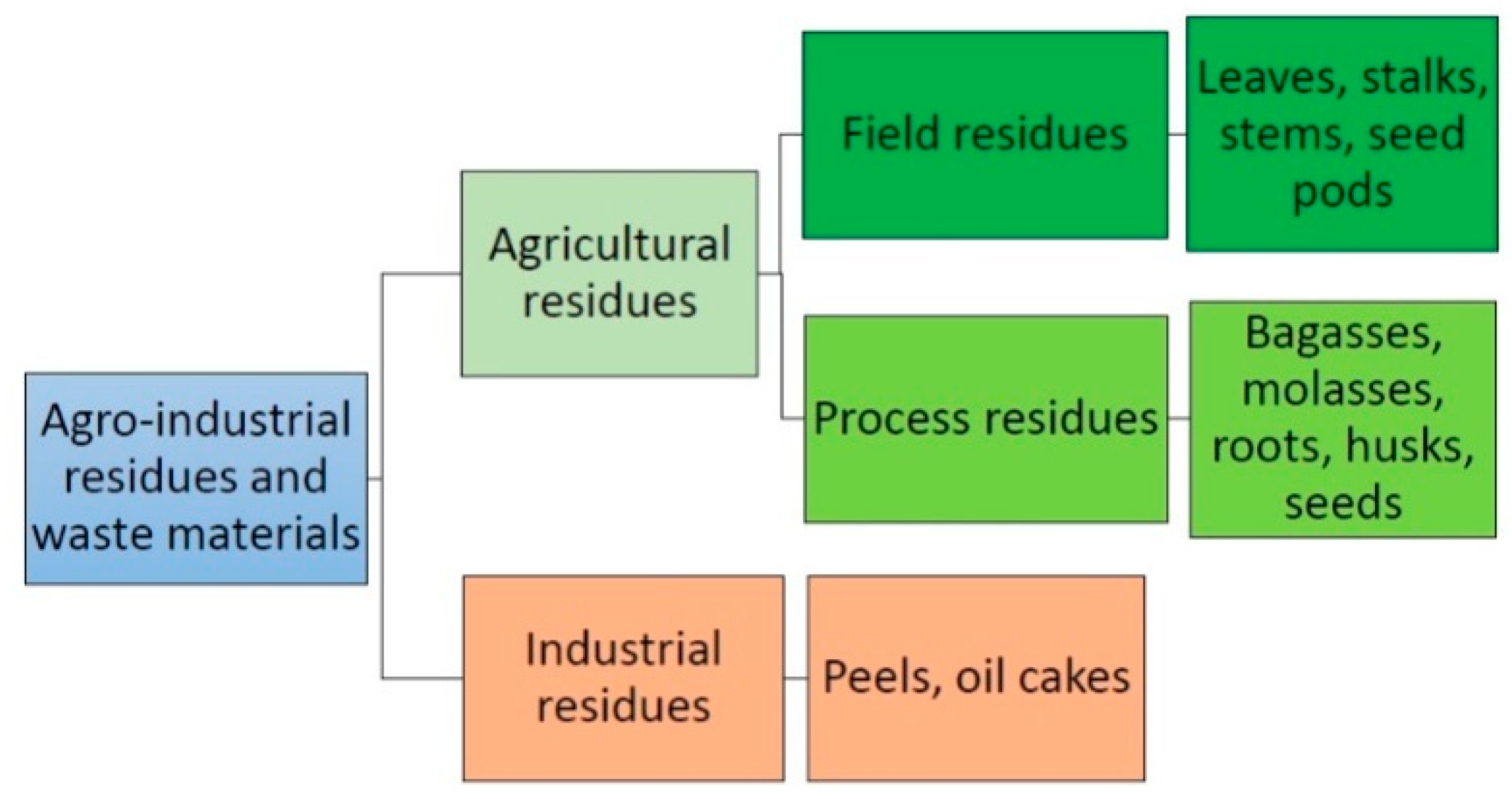
5. Agro-Industrial Residues and Wastes for Lipase Production
6. Oil Cakes
7. Fibrous Residues
8. Industrial Effluents
9. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Rodríguez de Romo, A.C.; Borgstein, J. Claudio Bernard and fat emulsion (or the Sleeping Beauty 150 years later). Gac. Med. Mex. 2000, 136, 379–386. [Google Scholar]
- Pahoja, V.M.; Sethar, M.A. A Review of enzymatic properties of lipase in plants, animals and microorganisms. J. Appl. Sci. 2002, 2, 474–484. [Google Scholar] [CrossRef]
- Javed, S.; Azeem, F.; Hussain, S.; Rasul, I.; Siddique, M.H.; Riazc, M.; Afzal, M.; Kouser, A.; Nadeem, H. Bacterial lipases: A review on purification and characterization. Prog. Biophys. Mol. Biol. 2018, 132, 23–34. [Google Scholar] [CrossRef]
- Mehta, A.; Bodh, U.; Gupta, R. Fungal lipases: A review. J. Biotech Res. 2017, 8, 58–77. [Google Scholar]
- Lock, L.L.; Corbellini, V.A.; Valente, P. Lipases produced by yeasts: Powerful biocatalysts for industrial purposes. Techno-Lógica 2007, 11, 18–25. [Google Scholar] [CrossRef]
- Sharma, R.; Yusuf, C.; Banerjee, U.C. Production, purification, characterization, and applications of lipases. Biotechnol. Adv. 2001, 19, 627–662. [Google Scholar] [CrossRef]
- Tufvesson, P.; Törnvall, U.; Carvalho, J.; Karlsson, A.; Hatti-Kaul, R. Towards a cost-effective immobilized lipase for specialty chemicals. J. Mol. Catal. B Enzym. 2011, 68, 200–205. [Google Scholar] [CrossRef]
- Jaeger, K.E.; Dijkstra, B.W.; Reetz, M.T. Bacterial biocatalysts: Molecular biology, three dimensional structures and biotechnological application of lipases. Annu. Rev. Microbiol. 1999, 53, 315–351. [Google Scholar] [CrossRef]
- Eijkmann, C. Über Enzyme beiBakterien und Schimmelpilzen. Centr. Bakteriol. Parasitenk. I Abt. 1901, 29, 841–848. [Google Scholar]
- van Pouderoyen, G.; Eggert, T.; Jaeger, K.E.; Dijkstra, B.W. The crystal structure of Bacillus subtilis lipase: A minimal alpha/beta hydrolase fold enzyme. J. Mol. Biol. 2001, 309, 215–226. [Google Scholar] [CrossRef]
- Saxena, R.K.; Sheoran, A.; Giri, B.; Davidson, W.S. Purification strategies for microbial lipases. J. Microbiol. Methods 2003, 52, 1–18. [Google Scholar] [CrossRef]
- Brzozowski, A.M.; Savage, H.; Verma, C.S.; Turkenburg, J.P.; Lawson, D.M.; Svendsen, A.; Patkar, S. Structural origins of the interfacial activation in Thermomyces (Humicola) lanuginose lipase. Biochemistry 2000, 39, 15071–15082. [Google Scholar] [CrossRef]
- Pleiss, J.; Fischer, M.; Schmid, R.D. Anatomy of lipase binding sites: The scissile fatty acid binding site. Chem. Phys. Lipids 1998, 93, 67–80. [Google Scholar] [CrossRef]
- Gupta, R.; Kumari, A.; Syal, P.; Singh, Y. Molecular and functional diversity of yeast and fungal lipases: Their role in biotechnology and cellular physiology. Prog. Lipid Res. 2015, 57, 40–54. [Google Scholar] [CrossRef]
- Rajakumara, E.; Acharya, P.; Ahmad, S.; Sankaranaryanan, R.; Rao, N.M. Structural basis for the remarkable stability of Bacillus subtilis lipase (Lip A) at low pH. Biochim.Biophys. Acta 2008, 1784, 302–311. [Google Scholar] [CrossRef]
- BurcuBakir, Z.; Metin, K. Production and characterization of an alkaline lipase from thermophilic Anoxybacillus sp. HBB16. Chem. Biochem. Eng. Q. 2017, 31, 303–312. [Google Scholar] [CrossRef]
- Woolley, P.; Petersen, S.B. Lipases: TheirStructure, Biochemistry and Application; Cambridge University Press: Cambridge, UK, 1994; pp. 315–351. ISBN 9780521207997. [Google Scholar]
- Adlercreutz, P.; Gitlesen, T.; Ncube, I.; Read, J.S. Vernonia lipase: A plant lipase with strong fatty acid selectivity. Methods Enzymol. 1997, 284, 220–232. [Google Scholar] [CrossRef]
- Nagarajan, S. New tools for exploring old friends-microbial lipases. Appl. Biochem. Biotechnol. 2012, 168, 1163–1196. [Google Scholar] [CrossRef]
- Kapoor, M.; Gupta, M.N. Lipase promiscuity and its biochemical applications. Process Biochem. 2012, 47, 555–569. [Google Scholar] [CrossRef]
- Ribeiro, B.D.; Castro, A.M.; Coelho, M.A.Z.; Freire, D.M.G. Production and use of lipases in bioenergy: A review from the feedstocks to biodiesel production. Enzym. Res. 2011, 1, 615803. [Google Scholar] [CrossRef]
- Barros, M.; Fleuri, L.F.; Macedo, G.A. Seed lipases: Sources, applications and properties—A review. Braz. J. Chem. Eng. 2010, 27, 15–29. [Google Scholar] [CrossRef]
- van der Padt, A.; Edema, M.J.; Sewalt, J.J.W.; van’tRiet, K. Enzymatic acylglycerol synthesis in a membrane bioreactor. J. Am. Oil Chem. Soc. 1990, 67, 347–352. [Google Scholar] [CrossRef]
- Marangoni, A.G. Lipases: Structure, function, and properties. Lipid Biotechnol. 2002, 357–386. [Google Scholar] [CrossRef]
- Kuo, C.H.; Chen, H.H.; Chen, J.H.; Liu, Y.C.; Shieh, C.J. High yield of wax ester synthesized from cetyl alcohol and octanoic acid by Lipozyme RMIM and Novozym 435. Int. J. Mol. Sci. 2012, 13, 11694–11704. [Google Scholar] [CrossRef] [PubMed]
- Huang, S.M.; Huang, H.Y.; Chen, Y.M.; Kuo, C.H.; Shieh, C.J. Continuous production of 2-phenylethyl acetate in a solvent-free system using a packed-bed reactor with Novozym® 435. Catalysts 2020, 10, 714. [Google Scholar] [CrossRef]
- Kuo, C.H.; Huang, C.Y.; Lee, C.L.; Kuo, W.C.; Hsieh, S.L.; Shieh, C.J. Synthesis of DHA/EPA ethylesters via lipase-catalyzedacidolysisusingNovozym® 435: A kineticstudy. Catalysts 2020, 10, 565. [Google Scholar] [CrossRef]
- Lee, J.H.; Son, J.M.; Akoh, C.C.; Kim, M.R.; Lee, K.T. Optimized synthesis of 1,3-dioleoyl-2-palmitoylglycerol-rich triacylglycerol via interesterification catalyzed by a lipase from Thermomyceslanuginosus. New Biotechnol. 2010, 27, 38–45. [Google Scholar] [CrossRef]
- Saxena, R.K.; Ghosh, P.K.; Gupta, R.; Davidson, W.S.; Bradoo, S.; Gulati, R. Microbial lipases: Potential biocatalysts for the future. Curr. Sci. 1999, 77, 101–115. [Google Scholar]
- Konkit, M.; Kim, W. Activities of amylase, proteinase, and lipase enzymes from Lactococcus chungangensis and its application in dairy products. J. Dairy Sci. 2016, 99, 4999–5007. [Google Scholar] [CrossRef]
- Unni, K.N.; Priji, P.; Sajith, S.; Faisal, P.A.; Benjamin, S. Pseudomonas aeruginosa strain BUP2, a novel bacterium inhabiting the rumen of Malabari goat, produces an efficient lipase. Biologia 2016, 71, 378–387. [Google Scholar] [CrossRef]
- Dey, A.; Chattopadhyay, A.; Saha, P.; Mukhopadhyay, S. An approach to the identification and characterisation of a psychrotrophic lipase producing Pseudomonas sp. ADT3 from Arctic Region. Adv. Biosci. Biotechnol. 2014, 5, 322–332. [Google Scholar] [CrossRef][Green Version]
- Kalantzi, S.; Mamma, D.; Kalogeris, E.; Kekos, D. Improved properties of cotton fabrics treated with lipase and its combination with pectinase. FIBRES Text. East. Eur. 2010, 18, 86–92. [Google Scholar]
- Sarethy, I.P.; Saxena, Y.; Kapoor, A.; Sharma, M.; Sharma, S.K.; Gupta, V.; Gupta, S. Alkaliphilic bacteria: Applications in industrial biotechnology. J. Ind. Microbiol. Biotechnol. 2011, 38, 769–790. [Google Scholar] [CrossRef]
- Karmee, S.K.; Patria, R.D.; Lin, C.S. Techno-economic evaluation of biodiesel production from waste cooking oil—A case study of Hong Kong. Int. J. Mol. Sci. 2015, 16, 4362–4371. [Google Scholar] [CrossRef]
- MarketInsightsReports (An Ameliorate Digital Consultancy Pvt Ltd. Group Company) Home Page. Available online: htpp://www.marketinsightsreports.com/reports/04071968644/global-microbial-lipase-market-research-report-2020.html (accessed on 1 February 2021).
- Sae-Leaw, T.; Benjakul, S. Lipase from liver of seabass (Lates calcarifer): Characteristics and the use for defatting of fish skin. Food Chem. 2018, 240, 9–15. [Google Scholar] [CrossRef]
- Kheadr, E.E.; Vuillemard, J.C.; El-Deeb, S.A. Impact of liposome-encapsulated enzyme cocktails on cheddar cheese ripening. Food Res. Int. 2003, 36, 241–252. [Google Scholar] [CrossRef]
- Kim, B.H.; Akoh, C.C. Recent research trends on the enzymatic synthesis of structured lipids. J. Food Sci. 2015, 80, 1713–1724. [Google Scholar] [CrossRef] [PubMed]
- Sánchez, M.; Prim, N.; Rández-Gil, F.; Pastor, F.I.; Diaz, P. Engineering of baker’s yeasts, E. coli and Bacillus hosts for the production of Bacillus subtilis Lipase A. Biotechnol. Bioeng. 2002, 78, 339–345. [Google Scholar] [CrossRef] [PubMed]
- Sikora, A.; Siodmiak, T.; Marszałł, M. Kinetic resolution of profens by enantioselective esterification. Chirality 2014, 26, 63–669. [Google Scholar] [CrossRef]
- Ye, L.; Zhang, B.; Seviour, E.G.; Tao, K.X.; Liu, X.H.; Ling, Y.; Chen, J.Y.; Wang, G.B. Monoacylglycerol lipase (MAGL) knockdown inhibits tumor cells growth in colorectal cancer. Cancer Lett. 2011, 307, 6–17. [Google Scholar] [CrossRef]
- Hasan, F.; Shah, A.A.; Hameed, A. Industrial applications of microbial lipases. Enzyme Microb. Technol. 2006, 39, 235–251. [Google Scholar] [CrossRef]
- An, J.D.; Patterson, D.A.; McNeil, S.; Hossain, M.M. Immobilization of lipase on woolen fabrics: Enhanced effectiveness in stain removal. Biotechnol. Prog. 2014, 30, 806–817. [Google Scholar] [CrossRef] [PubMed]
- Thanikaivelan, P.; Rao, J.R.; Nair, B.U.; Ramasami, T. Enzyme applications in leather processing: Current scenario and future outlook. In Enzyme Mixtures and Complex Biosynthesis; Bhattacharya, S.K., Ed.; CRC Press: London, UK, 2007; pp. 37–48. ISBN 9780429089589. [Google Scholar]
- Sangeetha, R.; Arulpandi, I.; Geetha, A. Bacterial lipases as potential industrial biocatalysts: An overview. Res. J. Microbiol. 2011, 6, 1. [Google Scholar] [CrossRef]
- Bhushan, I.; Kumar, A.; Modi, G.; Jamwal, S. Chiral resolution of differently substituted racemic acetyl-1-phenyl ethanol using lipase from Bacillus subtilis. J. Chem. Technol. Biotechnol. 2011, 86, 315–318. [Google Scholar] [CrossRef]
- Bajaj, A.; Lohan, P.; Jha, P.N.; Mehrotra, R. Biodiesel production through lipase catalyzed transesterification: An overview. J. Mol. Catal. B Enzym. 2010, 62, 9–14. [Google Scholar] [CrossRef]
- Sigma-Aldrich Home Page. Available online: http://www.sigmaaldrich.com/catalog/search?term=lipase&interface=All&N=0&mode=match%20partialmax&lang=pl®ion=PL&focus=product.html (accessed on 1 September 2020).
- Haba, E.; Bresco, O.; Ferrer, C.; Marqués, A.; Busquets, M.; Manresa, A. Isolation of lipase secreting bacteria by deploying used frying oil as selective substrate. Enzym. Microb. Technol. 2000, 26, 40–44. [Google Scholar] [CrossRef]
- Brozzoli, V.; Crognale, S.; Sampedro, I.; Federici, F.; D’Annibale, A.; Petruccioli, M. Assessment of olive-mill wastewater as a growth medium for lipase production by Candida cylindracea in bench-top reactor. Bioresour. Technol. 2009, 100, 3395–3402. [Google Scholar] [CrossRef]
- Garlapati, V.K.; Banerjee, R. Optimization of lipase production using differential evolution. Biotechnol. Bioprocess Eng. 2010, 15, 254–260. [Google Scholar] [CrossRef]
- Godoy, M.G.; Gutarra, M.L.E.; Maciel, F.M.; Felix, S.P.; Bevilaqua, J.V.; Machado, O.L.T.; Freire, D.M.G. Use of a low-cost methodology for biodetoxification of castor bean waste and lipase production. Enzym. Microb. Technol. 2009, 44, 317–322. [Google Scholar] [CrossRef]
- Paithankar, A.; Rewatkar, A. Oil cakes as substrate for improved lipase production in solid-state fermentation. IOSR J. Phar. Biol. Sci. 2014, 9, 31–38. [Google Scholar] [CrossRef]
- Rigo, E.; Ninow, J.L.; Polloni, A.E.; Remonatto, D.; Arbter, F.; Vardanega, R.; Oliviera, D.; Treichel, H.; Di Luccio, M. Improved lipase biosynthesis by a newly isolated Penicillium sp. grown on agricultural wastes. Ind. Biotechnol. 2009, 5, 119–126. [Google Scholar] [CrossRef]
- Singhania, R.R.; Patel, K.A.; Thomas, L.; Goswami, M.; Giri, S.G.; Pandey, A. Industrial Enzymes. In Industrial Biorefineries & White Biotechnology; Pandey, A., Höfer, R., Taherzadeh, M., Nampoothiri, K.M., Larroche, C., Eds.; Elsevier: Amsterdam, The Netherlands, 2015; pp. 473–497. ISBN 978-0-444-63453-5. [Google Scholar]
- Pandey, A.; Soccol, C.; Mitchell, D.A. New developments in solid-state fermentation: I-Bioprocess and products. Process Biochem. 2000, 35, 1153–1169. [Google Scholar] [CrossRef]
- Sadh, P.K.; Duhan, S.; Duhan, J.S. Agro-industrial wastes and their utilization using solid-state fermentation: A review. Bioresour. Bioprocess. 2018, 5, 1. [Google Scholar] [CrossRef]
- Hölker, U.; Höfer, M.; Lenz, J. Biotechnological advantages of laboratory-scale solid-state fermentation with fungi. Appl. Microbiol. Biotechnol. 2004, 64, 175–186. [Google Scholar] [CrossRef]
- Krishna, C. Solid-state fermentation systems—An overview. Crit. Rev. Biotechnol. 2005, 25, 1–30. [Google Scholar] [CrossRef] [PubMed]
- Mitchell, D.A.; Berovic, M.; Krieger, N. Biochemical engineering aspects of solid-state bioprocessing. Adv. Biochem. Eng. Biotechnol. 2000, 68, 61–138. [Google Scholar] [CrossRef] [PubMed]
- Sánchez, L.R.; Khanahmadi, M.; Mitchell, D. Continuous solid-state fermentation bioreactors. In Solid-State Fermentation Bioreactors; Mitchell, D.A., Berovič, M., Krieger, N., Eds.; Springer: Berlin, Germany, 2006; pp. 141–158. ISBN 978-3-540-31286-4. [Google Scholar]
- Stoytcheva, M.; Montero, G.; Zlatev, R.; Leon, J.A.; Gochev, V. Analytical methods for lipases activity determination: A review. Curr. Anal. Chem. 2012, 8, 400–407. [Google Scholar] [CrossRef]
- Jaeger, K.E.; Kovacic, F. Determination of lipolytic enzyme activities. Methods Mol. Biol. 2014, 1149, 111–134. [Google Scholar] [CrossRef]
- Gupta, R.; Rathi, P.; Gupta, N.; Bradoo, S. Lipase assays for conventional and molecular screening: An overview. Biotechnol. Appl. Biochem. 2003, 37, 63–71. [Google Scholar] [CrossRef] [PubMed]
- Ramnath, L.; Sithole, B.; Govinden, R. Identification of lipolytic enzymes isolated from bacteria indigenous to Eucalyptus wood species for application in the pulping industry. Biotechnol. Rep. 2017, 15, 114–124. [Google Scholar] [CrossRef]
- Gulomova, K.; Ziomek, E.; Schrag, J.D.; Davranov, K.; Cygler, M. Purification and characterization of a Penicillium sp. lipase which discriminates against diglycerides. Lipids 1996, 31, 379–384. [Google Scholar] [CrossRef]
- Kulkarni, N.; Gadre, R.V. Simple gas chromatography method for lipase assay. Biotechnol. Tech. 1998, 12, 627–628. [Google Scholar] [CrossRef]
- Suocheng, D.; Tong, K.W.; Yuping, W. Municipal solid waste management in China: Using commercial management to solve a growing problem. Util. Policy 2011, 10, 7–11. [Google Scholar] [CrossRef]
- Joardder, U.H.; Mahadi, M.M. Food Preservation in Developing Countries: Challenges and Solutions; Springer International Publishing: London, UK, 2019; pp. 27–55. ISBN 978-3-030-11530-2. [Google Scholar]
- Motte, J.C.; Trably, E.; Escudié, R.; Hamelin, J.; Steyer, J.P.; Bernet, N.; Delgenes, J.P.; Dumas, C. Total solids content: A key parameter of metabolic pathways in dry anaerobic digestion. Biotechnol. Biofuels 2013, 6, 164. [Google Scholar] [CrossRef]
- Oliveira, F.; Souza, C.E.; Peclat, V.R.O.L.; Salgado, J.M.; Ribeiro, B.D.; Coelho, M.A.Z.; Venâncio, A.; Belo, I. Optimization of lipase production by Aspergillus ibericus from oilcakes and its application in esterification reactions. Food Bioprod. Process. 2017, 102, 268–277. [Google Scholar] [CrossRef]
- Kumar, A.; Sadh, P.K.; Kha, S.; Duhan, J.S. Bio-ethanol production from sweet potato using co-culture of saccharolytic molds (Aspergillus spp.) and Saccharomyces cerevisiae MTCC170. J. Adv. Biotechnol. 2016, 6, 822–827. [Google Scholar] [CrossRef]
- Ezejiofor, T.I.N.; Duru, C.I.; Asagbra, A.E.; Ezejiofor, A.N.; Orisakwe, O.E.; Afonne, J.O.; Obi, E. Waste to wealth: Production of oxytetracycline using streptomyces species from house hold kitchen wastes of agricultural produce. Afr. J. Biotechnol. 2012, 11, 10115–10124. [Google Scholar] [CrossRef]
- Subbu Iakshmi, S.; Sornaraj, R. Utilization of see food processing waste for cultivation of the edible mushroom Pleurotusflabellatus. Afr. J. Biotechnol. 2014, 17, 1779–1785. [Google Scholar] [CrossRef]
- Sukan, A.; Roy, I.; Keshavarz, T. Agro-industrial waste materials as substrates for the production of poly(3-hydroxybutyric acid). J. Biomater. Nanobiotechnol. 2014, 5, 229–240. [Google Scholar] [CrossRef]
- Leming, R.; Lember, A. Chemical composition of expeller-extracted and cold-pressed rapeseed cake. Agraarteadus 2005, 16, 96–103. [Google Scholar]
- Boratyński, F.; Szczepańska, E.; Grudniewska, A.; Gniłka, R.; Olejniczak, T. Improving of hydrolases biosynthesis by solid-state fermentation of Penicillium camemberti on rapeseed cake. Sci. Rep. 2018, 8, 10157. [Google Scholar] [CrossRef]
- Sharma, N.K. Chemical composition of oilseed cakes and deoiled cakes in Nepal. Online J. Anim. Feed Res. 2013, 3, 74–76. [Google Scholar]
- Parihar, D.K. Production of lipase utilizing linseed oilcake as fermentation substrate. Int. J. Sci. Environ. Technol. 2012, 1, 135–143. [Google Scholar]
- Studnitz, M. Feeding monogastrics 100% organic and regionally produced feed. Knowledge Synthesis. OK-Net EcoFeed. H2020-project. 2019. Available online: http://orgprints.org/34560/ (accessed on 4 June 2021).
- Tanyol, M.; Uslu, G.; Yönten, V. Optimization of lipase production on agro-industrial residue medium by Pseudomonas fluorescens (NRLL B-2641) using response surface methodology. Biotechnol. Biotechnol. Equip. 2015, 29, 64–71. [Google Scholar] [CrossRef]
- Bhosale, H.J.; Kadam, T.A.; Sukalkar, S.R.; Adekar, S.D. LipaseProduction from Bacillus sp Using Soybean Oil Cake as Substrate. Int. J. Pharm. Biol. Res. 2012, 3, 213–218. [Google Scholar]
- Madruga, M.S.; Camara, F.S. The chemical composition of multimistura as a food supplement. Food Chem. 2000, 68, 41–44. [Google Scholar] [CrossRef]
- Guo, Y.; Rockstraw, D.A. Activated carbons prepared from rice hull by one-step phosphoric acid activation. Microporous Microporous Mater. 2007, 100, 12–19. [Google Scholar] [CrossRef]
- ul-Haq, I.; Idrees, S.; Rajoka, M.I. Production of lipases by Rhizopus oligosporous by solid-state fermentation. Process Biochem. 2002, 37, 637–641. [Google Scholar] [CrossRef]
- Kumar, D.S. Fungal Lipase Production by Solid State Fermentation—An Overview. J. Anal. Bioanal. Tech. 2015, 6, 1. [Google Scholar] [CrossRef]
- Colla, L.M.; Rizzardi, J.; Pinto, M.H.; Reinehr, C.O.; Bertolin, T.E.; Costa, J.A.V. Simultaneous production of lipases and biosurfactants by submerged and solid-state bioprocesses. Bioresour. Technol. 2010, 101, 8308–8314. [Google Scholar] [CrossRef]
- Abid, K.; Jabri, J.; Beckers, Y.; Yaich, H.; Malek, A.; Rekhis, J.; Kamoun, M. Effects of exogenous fibrolytic enzymes on the ruminal fermentation of agro-industrial by-products. S. Afr. J. Anim. Sci. 2019, 49, 612–618. [Google Scholar] [CrossRef]
- Ayahan, V.; Arslan Duru, A.; Ozkaya, S. Possibilities of using dried apple pomace in broiler chicken diet. Kafkas Univ. Vet. Fak. Derg. 2009, 15, 669–672. [Google Scholar]
- Neupane, K.; Khadka, R. Production of Garbage Enzyme from Different Fruit and Vegetable Wastes and Evaluation of its Enzymatic and Antimicrobial Efficacy. Tribhuvan Univ. J. Microbiol. 2019, 6, 113–118. [Google Scholar] [CrossRef]
- Okino-Delgado, C.H.; Prado, D.Z.; Fleuri, L.F. Brazilian fruit processing, wastes as a source of lipase and other biotechnological products: A review. An. Acad. Bras. Ciênc. 2018, 90, 2927–2943. [Google Scholar] [CrossRef]
- Habib, M.A.B.; Yusoff, F.M.; Phang, S.M.; Ang, K.J.; Mohamed, S. Nutritional values of chironomid larvae grown in palm oil mill effluent and algal culture. Aquaculture 1997, 158, 95–105. [Google Scholar] [CrossRef]
- Hermansyah, H.; Maresya, A.; Putri, D.N.; Sahlan, M.; Meyer, M. Production of dry extract lipase from Pseudomonas aeruginosa by the submerged fermentation method in palm oil mill effluent. Int. J. Technol. 2018, 9, 325–334. [Google Scholar] [CrossRef]
- The Commission’s Directorate-General for Agriculture and Rural Development (DG AGRI): Oilseeds and Protein Crops market situation. Committee for the Common Organisation of Agricultural Markets Home Page. Available online: http://ec.europa.eu/info/food-farming-fisheries/key-policies/committees-and-advisory-councils/agricultural-committees_en.html (accessed on 27 August 2020).
- Sivaramakrishnan, S.; Gangadharan, D. Edible Oil Cakes. In Biotechnology for Agro-Industrial Residues Utilisation; Singh-NeeNigam, P., Pandey, A., Eds.; Springer: Dordrecht, The Netherlands, 2009; pp. 253–271. ISBN 978-1-4020-9942-7. [Google Scholar]
- Molina-Alcaide, E.; Yáñez-Ruiz, D.R. Potential use of olive by-products in ruminant feeding: A review. Anim. Feed Sci. Technol. 2008, 147, 247–264. [Google Scholar] [CrossRef]
- Mushtaq, T.; Sarwar, M.; Ahmad, G.; Mirza, M.A.; Ahmad, T.; Noreen, U.; Mushtaq, M.M.H.; Kamran, Z. Influence of sunflower meal based diets supplemented with exogenous enzyme and digestible lysine on performance, digestibility and carcass response of broiler chickens. Anim. Feed Sci. Technol. 2009, 149, 275–286. [Google Scholar] [CrossRef]
- Ramachandran, S.; Singh, S.K.; Larroche, C.; Soccol, C.R.; Pandey, A. Oil cakes and their biotechnological applications—A review. Bioresour. Technol. 2007, 98, 2000–2009. [Google Scholar] [CrossRef]
- Makkar, H.P.S.; Aderibigbe, A.O.; Becker, K. Comparative evaluation of non-toxic and toxic varieties of Jatropha curcas for chemical composition, digestibility, protein degradability and toxic factors. Food Chem. 1998, 62, 207–215. [Google Scholar] [CrossRef]
- Ramakrishnan, C.V.; Banerjee, B.N. Studies on mold lipase. Comparative study of lipases obtained from molds grown on Sesamum indicum. Arch. Biochem. Biophys. 1952, 37, 131–135. [Google Scholar] [CrossRef]
- Shukla, P.; Bhagat, J.; Shrivastava, S. Oil cakes as substrate for improved lipase production in solid state fermentation. Int. J. Microbiol. Res. 2011, 3, 71–73. [Google Scholar]
- Nerurkar, M.; Joshi, M.; Adivarekar, R. Use of sesame oil cake for lipase production from a newly marine isolated Bacillus sonorensis. Innov. Rom. Food Biotechnol. 2013, 13, 11–17. [Google Scholar]
- Sahoo, K.R.; Kumar, M.; Mohanty, S.; Sawyer, M.; Rahman, P.; BehariSukla, L.; Subudhi, E. Statistical optimization for lipase production from solid waste of vegetable oil industry. Prep. Biochem. Biotechnol. 2018, 48, 321–326. [Google Scholar] [CrossRef]
- Rajendran, A.; Thangavelu, V. Utilizing agricultural wastes as substrates for lipase production by Candida rugosa NCIM 3462 in solid-state fermentation: Response surface optimization of fermentation parameters. Waste Biomass Valor. 2013, 4, 347–357. [Google Scholar] [CrossRef]
- Priya, B.S.; Stalin, T.; Selvam, K. Ecofriendly management of mixed coconut oil cake waste for lipase production by marine Streptomyces indiaensis and utilization as detergent additive. Int. J. Water Res. Environ. Eng. 2012, 4, 275–280. [Google Scholar] [CrossRef]
- Chaturvedi, M.; Singh, M.; Man, C.R.; Pandey, S. Lipase production from Bacillus subtilis MTCC 6808 by solid-state fermentation using ground nut oil cakes as substrate. Res. J. Microbiol. 2010, 5, 725–730. [Google Scholar] [CrossRef][Green Version]
- El Aal, R.A.; Shetaia, Y.M.; Shafei, M.S.; Gomaa, S.K.; El Menoufy, H.A.; El-Refai, H.A. Optimization of parameters for lipase production by Aspergillus niger NRRL-599 using response surface methodology. Egypt Pharm. J. 2019, 18, 165–171. [Google Scholar] [CrossRef]
- Graminha, E.B.N.; Gonçalves, A.Z.L.; Pirota, R.D.P.B.; Balsalobre, M.A.A.; Da Silva, R.; Gomes, E. Enzyme production by solid-state fermentation: Application to animal nutrition. Anim. Feed Sci. Technol. 2008, 144, 1–22. [Google Scholar] [CrossRef]
- Colombatto, D.; Mould, F.L.; Bhat, M.K.; Owen, E. Use of fibrolytic enzymes to improve the nutritive value of ruminant diets. A biochemical and in vitro rumen degradation assessment. Anim. Feed Sci. Technol. 2003, 107, 201–209. [Google Scholar] [CrossRef]
- Amin, M.; Bhatti, H.N. Effect of physicochemical parameters on lipase production by Penicillium fellutanum using canola seed oil cake as substrate. Int. J. Agric. Biol. 2014, 16, 118–124. [Google Scholar]
- Mala, J.G.; Edwinoliver, N.G.; Kamini, N.R.; Puvanakrishnan, R. Mixed substrate solid-state fermentation for production and extraction of lipase from Aspergillus niger MTCC 2594. J. Gen. Appl. Microbiol. 2007, 53, 247–253. [Google Scholar] [CrossRef]
- Fleuri, L.F.; de Oliveira, M.C.; de Lara Campos Arcuri, M.; Capoville, B.L.; Pereira, M.S.; Delgado, C.H.O.; Novelli, P.K. Production of fungal lipases using wheat bran and soybean bran and incorporation of sugarcane bagasse as a co-substrate in solid-state fermentation. Food Sci. Biotechnol. 2014, 23, 1199–1205. [Google Scholar] [CrossRef]
- Santos, R.R.; Muruci, L.N.M.; Damaso, M.C.T.; Lima da Silva, J.P.; Santos, L.O. Lipase production by Aspergillus niger 11T53A14 in wheat bran using experimental design methodology. J. Food Nutr. Res. 2014, 2, 659–663. [Google Scholar] [CrossRef][Green Version]
- Dobrev, G.; Strinska, H.; Hambarliiska, A.; Zhekova, B.; Dobreva, V. Optimization of lipase production in solid-state fermentation by Rhizopus arrhizus in nutrient medium containing agroindustrial wastes. Open Biotechnol. J. 2018, 12, 189–203. [Google Scholar] [CrossRef]
- Rekha, K.; Lakshmi, M.C.; Devi, V.S. Production and optimization of lipase from Candida rugosa using groundnut oilcake under solid-state fermentation. Int. J. Res. Eng. Technol. 2012, 1, 571–577. [Google Scholar] [CrossRef]
- Nema, A.; Patnala, S.H.; Mandari, V.; Kota, S.; Devarai, S.K. Production and optimization of lipase using Aspergillus niger MTCC 872 by solid-state fermentation. Bull. Natl. Res. Cent. 2019, 43, 82. [Google Scholar] [CrossRef]
- Pitol, L.O.; Finkler, A.T.; Dias, G.S.; Machado, A.S.; Zanin, G.M.; Mitchell, D.A.; Krieger, N. Optimization studies to develop a low-cost medium for production of the lipases of Rhizopus microspores by solid-state fermentation and scale-up of the process to a pilot packed-bed bioreactor. Process Biochem. 2017, 62, 37–47. [Google Scholar] [CrossRef]
- Awan, U.; Shafiq, K.; Mirza, S.; Ali, S.; Rehman, A.; Haq, I. Mineral constituent of culture medium for lipase production by Rhizopus oligosporous fermentation. Asian J. Plant Sci. 2003, 2, 913–915. [Google Scholar] [CrossRef][Green Version]
- Tariq, M.; Ali, M.; Shah, Z. Characteristics of industrial effluents and their possible impacts on quality of underground water. Soil Environ. 2006, 25, 64–69. [Google Scholar]
- Chillers, J.; Henrik, A. Effect of contaminated water. Glob. Water Hazards 1996, 25, 25–27. [Google Scholar]
- Kanmani, P.; Aravind, J.; Kumaresan, K. An insight into microbial lipases and their environmental facet. Int. J. Environ. Sci. Technol. 2015, 12, 1147–1162. [Google Scholar] [CrossRef]
- Singh, R.P.; Ibrahim, M.I.; Norizan, E. Composting of waste from palm oil mill: A sustainable waste management practice. Rev. Environ. Sci. Bio/Technol. 2010, 9, 331–344. [Google Scholar] [CrossRef]
- Ahmad, A.D.; Ismail, S.; Bhatia, S. Water recycling from palm oil mill effluent (POME) using membrane technology. Desalination 2003, 157, 87–95. [Google Scholar] [CrossRef]
- Roig, A.; Cayuela, M.L.; Sánchez-Monedero, M.A. An overview on olive mill wastes and their valorization methods. Waste Manag. 2006, 26, 960–969. [Google Scholar] [CrossRef]
- Anastopoulos, I.; Massas, I.; Ehaliotis, C. Use of residues and by-products of the olive-oil production chain for the removal of pollutants from environmental media: A review of batch biosorption approaches. J. Environ. Sci. Health 2015, 50, 677–718. [Google Scholar] [CrossRef]
- Salihu, A.; Alam, Z.; AbdulKarim, M.I.; Salleh, H.M. Suitability of using palm oil mill effluent as a medium for lipase production. Afr. J. Biotechnol. 2011, 10, 2044–2052. [Google Scholar]
- Moftah, O.A.S.; Grbavčić, S.Ž.; Moftah, W.A.S.; Luković, N.D.; Prodanović, O.L.; Jakovetić, S.M.; Knežević-Jugović, Z.D. Lipase production by Yarrowialipolytica using olive oil processing wastes as substrates. J. Serb. Chem. Soc. 2013, 78, 781–794. [Google Scholar] [CrossRef]
- D’Annibale, A.; Sermanni, G.G.; Federici, F.; Petruccioli, M. Olive oil wastewaters: A promising substrate for microbial lipase production. Bioresour. Technol. 2006, 97, 1828–1833. [Google Scholar] [CrossRef]
- Kumar, S.; Katiyar, N.; Ingle, P.; Negi, S. Use of evolutionary operation (EVOP) factorial design technique to develop a bioprocess using grease waste as a substrate for lipase production. Bioresour. Technol. 2011, 102, 4909–4912. [Google Scholar] [CrossRef]
- Ertugrul, S.; Dönmez, G.; Takaç, S. Isolation of lipase producing Bacillus sp. from olive mill wastewater and improving its enzyme activity. J. Hazard. Mater. 2007, 149, 720–724. [Google Scholar] [CrossRef] [PubMed]




| Commercial Lipases | Catalogue Number | Form | Specific Activity | Definition of the Unit of Specific Activity | Price |
|---|---|---|---|---|---|
| Lipase from Aspergillus oryzae | 62285 | lyophilized powder | ~50 U/mg | 1 U corresponds to the amount of enzyme that liberates 1 μmol oleic acid per minute at pH 8.0 and 40 °C. | 100 mg/158 USD |
| Lipase from Pseudomonas sp. | L9518 | lyophilized powder | ≥15 units/mg solid | One unit will produce 1.0μmol of glycerol from a triglyceride per minute at pH7.0 at 37 °C in the presence of bovine serum albumin. | 500 units/219 USD |
| Lipase from Candida rugosa | 62302 | lyophilized solid | 15–25 U/mg | 1 U corresponds to the amount of enzyme that liberates 1 μmol oleic acid per minute at pH 8.0 and 40 °C. | 100 mg/207 USD |
| Lipase from Rhizopus oryzae | 62305 | powder (fine) | ~10 U/mg | 1 U corresponds to the amount of enzyme that liberates 1 μmol of butyric acid per minute at pH 8.0 and 40 °C, and 5000 U as described above are equivalent to ~1 U using triolein at pH 8.0 and 40 °C. | 1 g/109 USD |
| Lipase from Aspergillus niger | 62301 | powder (fine) | ~200 U/g | 1 U corresponds to the amount of enzyme that hydrolyses 1 μmol acetic acid per minute at pH 7.4 and 40 °C, and 50 U as described above are equivalent to ~1 U using triolein substrate, at pH 8.0 and 40 °C. | 100 mg/23.90 USD |
| Lipase from Pseudomonas cepacia | 62309 | powder | ≥30 U/mg | 1 U corresponds to the amount of enzyme that liberates 1 μmol oleic acid per minute at pH 8.0 and 40 °C. | 100 mg/89.70 USD |
| Lipase from Candida sp. | L3170 | Aqueous solution | ≥5000 LU/g | Not defined. | 50 mL/77.20 USD |
| Lipase from Mucor miehei | 62298 | powder | ~1 U/mg | 1 U corresponds to the amount of enzyme that liberates 1 μmol oleic acid per minute at pH 8.0 and 40 °C. | 100 mg/86.50 USD |
| Lipase from Mucor javanicus | L8906 | lyophilized powder | ≥300 U/mg | One unit will hydrolyze 1.0 microequivalent of fatty acid from a triglyceride in 1 h at pH 7.7 at 37 °C using olive oil (30 min incubation). | 1 g/271 USD |
| Lipase from Candida antarctica | 02569 | lyophilized powder | ~0.3 U/mg | 1 U corresponds to the amount of enzyme that liberates 1 μmol oleic acid per minute at pH 8.0 and 40 °C. | 100 mg/118 USD |
| Amano Lipase PS from Burkholderiacepacia | 534641 | powder | ≥30,000 U/g, pH 7.0, 50 °C (optimum pH and temperature) | Not defined. | 10 g/60.10 USD |
| Amano Lipase from Pseudomonas fluorescens | 534730 | powder | ≥20,000 U/g, pH 8.0, 55 °C (optimum pH and temperature) | Not defined. | 10 g/51.20 USD |
| Factor | Solid-State Fermentation (SSF) | Submerged Fermentation (SmF) |
|---|---|---|
| Type of process | Batch | Batch or continuous |
| Substrates | Insoluble | Soluble |
| Water consumption in the fermentation process | Low | High |
| Amount of inoculum | Low (usually in liquid form added to medium) | High (usually added on surface of medium) |
| Contamination | High risk of contamination | Low risk of contamination |
| Energy consumption | Low | High |
| pH control | Difficult to control the pH parameter | Easy to control the pH parameter |
| Temperature maintenance | Difficult to control the temperature | Easy to control the temperature |
| Aeration | Better oxygen circulation | Aeration using sparger |
| Agitation | Low rpm (static conditions) | High rpm (homogenisation) |
| Equipment | Low costs and simple equipment | High costs and complex equipment |
| Scale-up | Difficult to scale-up because of engineering and equipment | Engineering and equipment are available |
| Amount of effluents | Low | High |
| Substrates | Dry Matter (%) | Crude Protein (%) | Crude Fat (%) | Crude Fiber (%) | Crude Ash (%) | Source |
|---|---|---|---|---|---|---|
| Rapeseed cake | 95.3 | 36.1 | 12.2 | 13.1 | 7.1 | [77,78] |
| Mustard cake | 91.4 | 30.2 | 6 | 6 | 6.7 | [79,80] |
| Sunflower cake | 93.8 | 33.7 | 8 | 18.5 | 6 | [81,82,83] |
| Soy Bean cake | 93.2 | 44 | 9 | 6 | 6 | |
| Corn germ cake | 87.8 | 18.9 | 2.5 | 7.9 | 5.7 | |
| Wheat bran | n. d. | 18 | 5.3 | 9.4 | 5 | [84,85,86,87,88] |
| Rice hulls | n. d. | 3 | 0.8 | 20 | 22.3 | |
| Mustard meal | n. d. | 34 | n. d. | 1.8 | 7.5 | |
| Coconut meal | n. d. | 21.3 | n. d. | 2 | 6 | |
| Almond meal | n. d. | 22 | n. d. | 4 | 7 | |
| Grape pomace | 52.2 | 14.1 | n. d. | 27.5 | 7.8 | [89,90,91,92] |
| Almond Hull | 31 | 3.2 | n. d. | 13 | 8.4 | |
| Pomegranate peel | 31.3 | 2.6 | n. d. | 13.5 | 5.7 | |
| Dried apple pomace | 89.56 | 5.5 | 4.8 | 17.99 | 3.4 | |
| Palm oil mill effluent | n. d. | 12.7 | 10.2 | n. d. | 14.9 | [93,94] |
| Substrates | Microorganism | Type of Fermentation | Maximum Lipase Activity | Source |
|---|---|---|---|---|
| Andiroba oil cake | Aspergillus ibericusMUM 03.49 | SSF | 1 U/g | [72] |
| Cupuassu oil cake | Aspergillus ibericusMUM 03.49 | SSF | 11 U/g | [72] |
| Canola oil cake | Aspergillus ibericusMUM 03.49 | SSF | 47 U/g | [72] |
| Macauba oil cake | Aspergillus ibericusMUM 03.49 | SSF | 1 U/g | [72] |
| Palm kernel oil cake | Aspergillus ibericusMUM 03.49 | SSF | 127 U/g | [72] |
| Crambe oil cake | Aspergillus ibericusMUM 03.49 | SSF | 44 U/g | [72] |
| Green coffee oil cake | Aspergillus ibericusMUM 03.49 | SSF | 4 U/g | [72] |
| Jatropha oil cake | Rhizopus oryzaeKG-10 | SSF | 80 IU | [102] |
| Teesi oil cake | Rhizopus oryzaeKG-10 | SSF | 60 IU | [102] |
| Mustard oil cake | Rhizopus oryzaeKG-10 | SSF | 170 IU | [102] |
| Bacillus subtilis MTCC 6808 | SSF | 1.68 U/g | [107] | |
| Pseudomonas aeruginosa LB-2 | SSF | 19.2 U/g | [80] | |
| Groundnut oil cake | Rhizopus oryzaeKG-10 | SSF | 60 IU | [102] |
| Bacillus subtilis MTCC 6808 | SSF | 4.5 U/g | [107] | |
| Sesame oilcake | Bacillus sonorensis | SmF | 41.35 U/mL/min | [103] |
| Aspergillus ibericusMUM 03.49 | SSF | 78 U/g | [72] | |
| Candida rugosa NCIM 3462 | SSF | 22.40 U/g | [105] | |
| Rapeseed oil cake | Aspergillus sp. AM31; Aspergillus candidusAM386; Aspergillus glaucusAM211; Aspergillus nidulansAM243; Aspergillus ochraceus AM456; Aspergillus wenthiAM413; Botrytis cinereaAM235; Fusarium avenaceumAM11; Fusarium equiseti AM15; Fusarium oxysporumAM13; Fusarium oxysporumAM21; Fusarium semitectumAM20; Fusarium solaniAM203; Fusarium tricinctumAM16; Mucor spinosusAM398; PapulariaroseaAM17; Penicillumcamembertii AM83; Penicillium chrysogenumAM112; Penicillium frequentansAM351; Penicillium notatum AR904; Penicillium ThomiiAM91; Penicillium vermiculatumAM30; Poria placenta AM38; SclerophomapythiophilaAR55; Spicoria divaricate AM423; SyncephalastrumracemosumAM105 | SSF | 247 U/mg | [78] |
| Olive oil cake | Bacillus licheniformis | SSF | 62.3 IU/g | [104] |
| Coconut oil cake | Bacillus subtilis MTCC 6808 | SSF | 2.35 U/g | [107] |
| Streptomyces indiaensis | SSF | 220.8 IU/mL | [106] | |
| Neem oil cake | Bacillus subtilis MTCC 6808 | SSF | 1.33 U/g | [107] |
| Linseed oil cake | Bacillus subtilis MTCC 6808 | SSF | 1.24 U/g | [107] |
| Wheat germ oil cake | Aspergillus niger NRRL-599 | SSF | 80 U/mL | [108] |
| Jojoba oil cake | Aspergillus niger NRRL-599 | SSF | 60 U/mL | [108] |
| Almond oil cake | Aspergillus niger NRRL-599 | SSF | 90 U/mL | [108] |
| Substrates | Microorganisms | Type of Fermentation | Maximum Lipase Activity | Source |
|---|---|---|---|---|
| Sunflower hulls | Penicillium fellutanum | SSF | 180 U/g | [111] |
| Rice bran | Penicillium fellutanum | SSF | 80 U/g | [111] |
| Rhizopus arrhizus | SSF | 2.6 U/g | [115] | |
| Candida rugosa NCIM 3467 | SSF | 26 U/mL | [116] | |
| Penicillumcitrinum NCIM 765 | SSF | 31 U/mL | [116] | |
| Peanut shells | Penicillium fellutanum | SSF | 150 U/g | [111] |
| Wheat bran | Penicillium fellutanum | SSF | 170 U/g | [111] |
| Aspergillus niger | SmF | 4.52 U | [88] | |
| Aspergillus niger | SSF | 384.3 U/g | [113] | |
| Penicillium sp. | SSF | 6.6 U/mL | [113] | |
| Aspergillus sp. | SSF | 11.35 U/mL | [113] | |
| Fusarium sp. | SSF | 2.55 U/mL | [113] | |
| Aspergillus niger11T51A14 | SSF | 153.4 U/g | [114] | |
| Rhizopus arrhizus | SSF | 5.5 U/g | [115] | |
| Candida rugosa NCIM 3467 | SSF | 21 U/mL | [116] | |
| Penicillumcitrinum NCIM 765 | SSF | 16 U/mL | [116] | |
| Soybean meal | Aspergillus niger | SSF | 25.07 U | [88] |
| Soybean bran | Penicillium sp. | SSF | 6.6 U/mL | [113] |
| Aspergillus sp. | SSF | 11.35 U/mL | [113] | |
| Fusarium sp. | SSF | 2.55 U/mL | [113] | |
| Corn flour | Rhizopus arrhizus | SSF | 1.1 U/g | [115] |
| Wheat flour | Rhizopus arrhizus | SSF | 4.9 U/g | [115] |
| Sunflower meal | Rhizopus arrhizus | SSF | 4.2 U/g | [115] |
| Oat bran | Rhizopus arrhizus | SSF | 3 U/g | [115] |
| Red gram husk | Aspergillus niger MTCC 872 | SSF | 7.17 U/g | [117] |
| Rice husk | Aspergillus niger MTCC 872 | SSF | 4.2 U/g | [117] |
| Sugarcane bagasse | Rhizopus microsporusCPQBA 312-07 DRM | SSF | 264 U/g | [118] |
| Almond meal | Rhizopus oligosporusISU-16 | SSF | 81.22 U/g | [119] |
| Substrates | Microorganisms | Type of Fermentation | Maximum Lipase Activity | Source |
|---|---|---|---|---|
| Palm oil mill effluent | Penicillium spp. E-01PC; Penicillium sppE-04PC; Penicillium spp. E-06PC; Penicillium spp. E-07PC; Penicillium spp. E-08PC; Penicillium sppE-10PC Mucor hemalis; Rhizopusoryzae; Candida cylindracea ATCC 14830; Penicillium citrinum ATCC 42799 | SmF | 4.02 U/mL (Candida cylindracea ATCC 14830) | [127] |
| Olive mill wastewater; Olive oil cake | Yarrowialipolytica NRRL Y-1095 | SmF | 850 IU/dm3 | [128] |
| Palm oil mill effluent | Pseudomonas aeruginosa B2290 | SmF | 1.327 U/mL | [94] |
| Olive mill wastewater | Candida cylindracea NRRL Y-17506 | SmF | 21.6 U/mL | [52] |
| Olive mill wastewater | Geotrichumcandidum NRRLY-552; Geotrichumcandidum Y-553; Rhizopus arrhizus NRRL 2286; Rhizopus arrhizus ISRIM 383; Rhizopus oryzae NRRL 6431; Aspergillus oryzae NRRL 1988; Aspergillus oryzae NRRL 495; Aspergillus niger NRRL 334; Candidacylindracea NRRL Y-17506; Penicillium citrinum NRRL 1841; Penicillium citrinum NRRL 3754; Penicillium citrinum ISRIM 118 | SmF | 9.23 IU/mL (Candida cylindracea NRRL Y-17506) | [129] |
| Grease waste; Wheat bran | Penicilliumchrysogenum | SSF | 46 U/mL | [130] |
| Olive mill wastewater; Whey | Bacillus sp. | SmF | 15 U/mL (extracellular) 168 U/mL (intracellular) | [131] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Szymczak, T.; Cybulska, J.; Podleśny, M.; Frąc, M. Various Perspectives on Microbial Lipase Production Using Agri-Food Waste and Renewable Products. Agriculture 2021, 11, 540. https://doi.org/10.3390/agriculture11060540
Szymczak T, Cybulska J, Podleśny M, Frąc M. Various Perspectives on Microbial Lipase Production Using Agri-Food Waste and Renewable Products. Agriculture. 2021; 11(6):540. https://doi.org/10.3390/agriculture11060540
Chicago/Turabian StyleSzymczak, Tomasz, Justyna Cybulska, Marcin Podleśny, and Magdalena Frąc. 2021. "Various Perspectives on Microbial Lipase Production Using Agri-Food Waste and Renewable Products" Agriculture 11, no. 6: 540. https://doi.org/10.3390/agriculture11060540
APA StyleSzymczak, T., Cybulska, J., Podleśny, M., & Frąc, M. (2021). Various Perspectives on Microbial Lipase Production Using Agri-Food Waste and Renewable Products. Agriculture, 11(6), 540. https://doi.org/10.3390/agriculture11060540








