Characterization of Bacteria and Inducible Phages in an Intensive Care Unit
Abstract
1. Introduction
2. Materials and Methods
2.1. Intensive Care Unit
2.2. Isolation of Bacteria from Environmental Samples
2.3. Lytic Bacteriophage Detection
2.4. Clinical Isolates from Patients
2.5. Prophage Induction
2.6. Bacteriophage Purification, Propagation, and Lysate Preparation
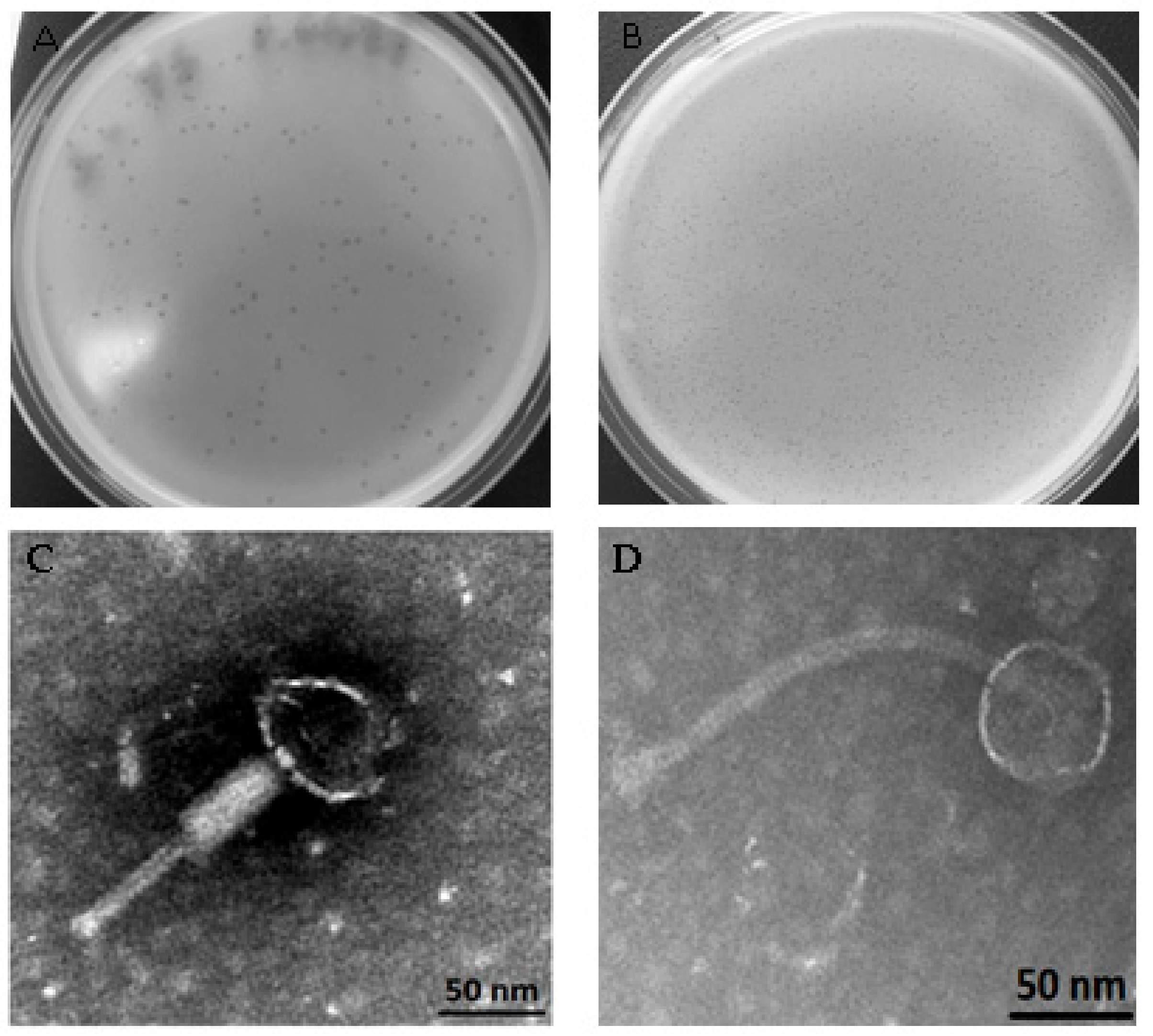
2.7. Transmission Electron Microscopy (TEM)
2.8. Host Range Analysis
2.9. Determination of Efficiency of Plating (EOP)
2.10. Bacteriophage Transduction
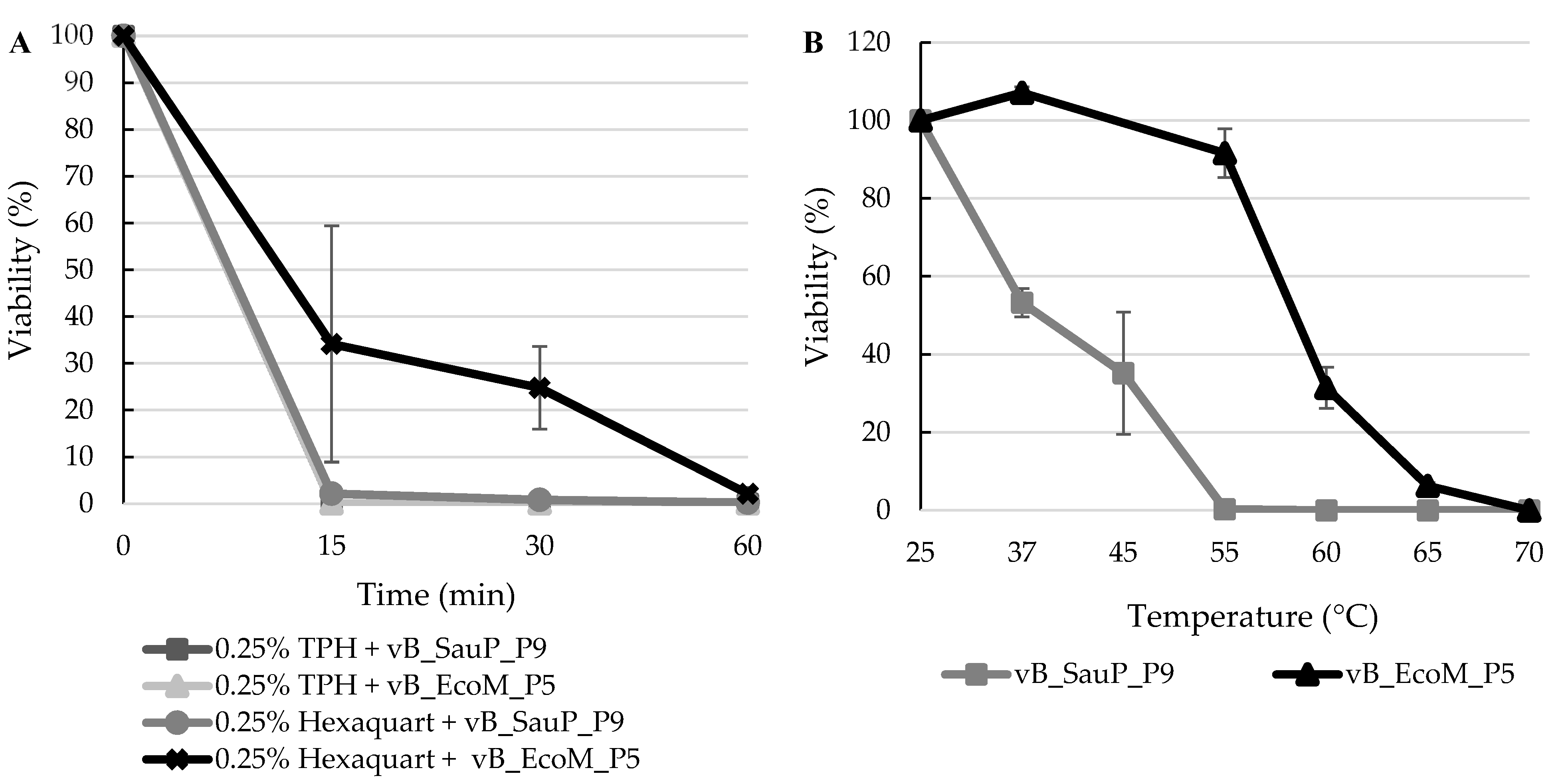
2.11. Disinfectant, Ethanol and Thermal Stability Tests
3. Results
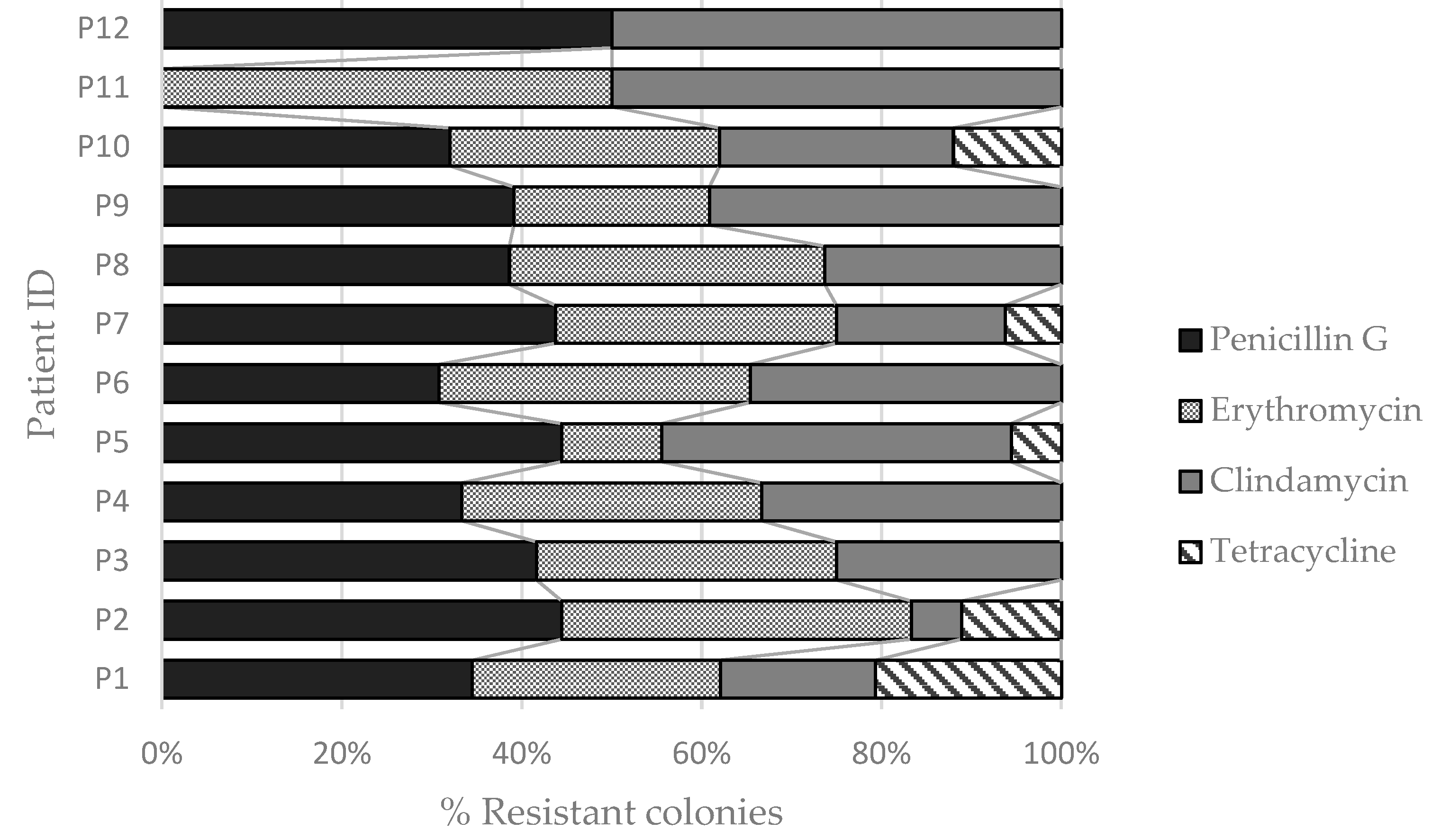
3.1. Bacterial and Viral Contamination
3.2. Temperate Phages
3.3. Lytic Spectrum
3.4. Antimicrobial Resistance Transduction
3.5. Virucidal Effect of Disinfectants and Thermal Stability
4. Discussion
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Brusselaers, N.; Vogelaers, D.; Blot, S. The Rising Problem of Antimicrobial Resistance in the Intensive Care Unit. Ann. Intensive Care 2011, 1, 47. [Google Scholar] [CrossRef] [PubMed]
- Russotto, V.; Cortegiani, A.; Raineri, S.M.; Giarratano, A. Bacterial Contamination of Inanimate Surfaces and Equipment in the Intensive Care Unit. J. Intensive Care 2015, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Del Sorbo, L.; Marshall, J.C. Antibiotic Resistance in the Intensive Care Unit. In Intensive Care Medicine; Vincen, J., Ed.; Springer: New York, NY, USA, 2006; pp. 582–591. [Google Scholar] [CrossRef]
- Vincent, J.; Marshall, J.; Anzueto, A.; Martin, C.D.; Gomersall, C. International Study of the Prevalence and Outcomes of Infection in Intensive Care Units. J. Am. Med. Assoc. 2009, 302, 2323–2329. [Google Scholar] [CrossRef] [PubMed]
- Gastmeier, P.; Loui, A.; Stamm-balderjahn, S.; Hansen, S.; Zuschneid, I. Outbreaks in Neonatal Intensive Care Units—They Are Not Like Others. Am. J. Infect. Control 2007, 35, 172–176. [Google Scholar] [CrossRef] [PubMed]
- European Centre for Disease Prevention and Control E. Annual Epidemiological Report for 2016 Healthcare-Associated Infections in Intensive Care Units; European Centre for Disease Prevention and Control E: Stockholm, Sweden, 2018. [Google Scholar]
- Wertheim, H.F.L.; Melles, D.C.; Vos, M.C.; Leeuwen WVan Belkum AVan Verbrugh, H.A.; Nouwen, J.L. The Role of Nasal Carriage in Staphylococcus aureus Infections. Lancet Infect. Dis. 2005, 5, 751–762. [Google Scholar] [CrossRef]
- Sampedro, G.R.; Wardenburg, J.B. Staphylococcus aureus in the Intensive Care Unit: Are These Golden Grapes Ripe for a New Approach? J. Infect. Dis. 2017, 215, S64–S70. [Google Scholar] [CrossRef]
- Repessé, X.; Artiguenave, M.; Paktoris-Papine, S.; Espinasse, F.; Dinh, A.; Charron, C.; El Sayed, F.; Geri, G.; Vieillard-Baron, A. Epidemiology of Extended-Spectrum Beta-Lactamase-Producing Enterobacteriaceae in an Intensive Care Unit with no Single Rooms. Ann. Intensive Care 2017, 7, 73. [Google Scholar] [CrossRef]
- Henderson, D.K. Managing Methicillin-Resistant Staphylococci: A Paradigm for Preventing Nosocomial Transmission of Resistant Organisms. Am. J. Infect. Control 2006, 119, S45–S52. [Google Scholar] [CrossRef]
- ECDC. Healthcare-Associated Infections—A Threat to Patient Safety in Europe 2018. Available online: https://ecdc.europa.eu/en/publications-data/infographic-healthcare-associated-infections-threat-patient-safety-europe (accessed on 14 May 2019).
- Gaidelyte, A.; Vaara, M.; Bamford, D.H. Bacteria, Phages and Septicemia. PLoS ONE 2007, 2, e1145. [Google Scholar] [CrossRef]
- Sweere, J.M.; Van Belleghem, J.D.; Ishak, H.; Bach, M.S.; Popescu, M.; Sunkari, V.; Kaber, G.; Manasherob, R.; Suh, G.A.; Cao, X.; et al. Bacteriophage Trigger Antiviral Immunity and Prevent Clearance of Bacterial Infection. Science 2019, 363, eaat9691. [Google Scholar] [CrossRef]
- Brussow, H.; Canchaya, C.; Hardt, W.-D. Phages and the Evolution of Bacterial Pathogens: From Genomic Rearrangements to Lysogenic Conversion. Microbiol. Mol. Biol. Rev. 2004, 68, 560–602. [Google Scholar] [CrossRef] [PubMed]
- DeBardeleben, H.K.; Lysenko, E.S.; Dalia, A.B.; Weiser, J. Tolerance of a Phage Element by Streptococcus pneumoniae Leads to a Fitness Defect during Colonization. J. Bacteriol. 2014, 196, 2670–2680. [Google Scholar] [CrossRef] [PubMed]
- Miller-Ensminger, T.; Garretto, A.; Brenner, J.; Thomas-White, K.; Zambom, A.; Wolfe, A.J.; Putonti, C. Bacteriophages of the Urinary Microbiome. J. Bacteriol. 2018, 200, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Navarro, F.; Muniesa, M. Phages in the Human Body. Front. Microbiol. 2017, 8, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Breitbart, M.; Hewson, I.; Felts, B.; Mahaffy, J.M.; Nulton, J.; Salamon, P.; Rohwer, F. Metagenomic Analyses of an Uncultured Viral Community from Human Feces. J. Bacteriol. 2003, 185, 6220–6223. [Google Scholar] [CrossRef] [PubMed]
- Ackermann, H. Tailed Bacteriophages: The Order Caudovirales. Adv. Virus Res. 1999, 51, 135–201. [Google Scholar] [CrossRef]
- King, A.; Adams, M.; Carstens, E.; Lefkowitz, E. Order-Caudovirales. Virus Taxon. 2012, 39–45. [Google Scholar] [CrossRef]
- Krahn, T.; Wibberg, D.; Maus, I.; Winkler, A.; Bontron, S.; Sczyrba, A.; Nordmann, P.; Pühler, A.; Poirel, L.; Schlüter, A. Intraspecies Transfer of the Chromosomally Encoded Acinetobacter baumannii blaNDM-1 Carbapenemase Gene. Antimicrob. Agents Chemother. 2016, 60, 3032–3040. [Google Scholar] [CrossRef]
- Billard-Pomares, T.; Fouteau, S.; Jacquet, M.E.; Roche, D.; Barbe, V.; Castellanos, M.; Bouet, J.Y.; Cruveiller, S.; Médigue, C.; Blanco, J.; et al. Characterization of a P1-Like Bacteriophage Carrying an SHV-2 Extended-Spectrum β-Lactamase from an Escherichia coli Strain. Antimicrob. Agents Chemother. 2014, 58, 6550–6557. [Google Scholar] [CrossRef]
- Sekizuka, T.; Yamamoto, A.; Komiya, T.; Kenri, T.; Takeuchi, F.; Shibayama, K.; Takahashi, M.; Kuroda, M.; Iwaki, M. Corynebacterium ulcerans 0102 Carries the Gene Encoding Diphtheria Toxin on a Prophage Different from the C. diphtheriae NCTC 13129 Prophage. BMC Microbiol. 2012, 12, 72. [Google Scholar] [CrossRef]
- Hendrix, R.W.; Smith, M.C.M.; Burns, R.N.; Ford, M.E.; Hatfull, G.F. Evolutionary Relationships among Diverse Bacteriophages and Prophages: All the world’s a Phage. Proc. Natl. Acad. Sci. USA 1999, 96, 2192–2197. [Google Scholar] [CrossRef] [PubMed]
- Hyman, P.; Abedon, S.T. Bacteriophages in Health and Disease, 1st ed.; CAB International: Oxfordshire, UK, 2012. [Google Scholar]
- Verkaik, N.J. Immune Evasion Cluster-Positive Bacteriophages are Highly Prevalent among Human Staphylococcus aureus Strains, but They Are Not Essential in the First Stages of Nasal Colonization. Clin. Microbiol. Infect. 2011, 17, 343–348. [Google Scholar] [CrossRef] [PubMed]
- Arnold, C. Rethinking Sterile: The Hospital Microbiome. Environ. Health Perspect. 2014, 122, 182–187. [Google Scholar] [CrossRef] [PubMed]
- Mora, M.; Mahnert, A.; Koskinen, K.; Pausan, M.R.; Oberauner-Wappis, L.; Krause, R.; Perras, A.K.; Gorkiewicz, G.; Berg, G.; Moissl-Eichinger, C. Microorganisms in Confined Habitats: Microbial Monitoring and Control of Intensive Care Units, Operating Rooms, Cleanrooms and the International Space Station. Front. Microbiol. 2016, 7. [Google Scholar] [CrossRef] [PubMed]
- Passaretti, C.L.; Otter, J.A.; Reich, N.G.; Myers, J.; Shepard, J.; Ross, T.; Carroll, K.C.; Lipsett, P.; Perl, T.M. An Evaluation of Environmental Decontamination with Hydrogen Peroxide Vapor for Reducing the Risk of Patient Acquisition of Multidrug-Resistant Organisms. Clin. Infect. Dis. 2013, 56, 27–35. [Google Scholar] [CrossRef] [PubMed]
- Salgado, C.D.; Sepkowitz, K.A.; John, J.F.; Cantey, J.R.; Attaway, H.H.; Freeman, K.D.; Sharpe, P.A.; Michels, H.T.; Schmidt, M.G. Copper Surfaces Reduce the Rate of Healthcare-Acquired Infections in the Intensive Care Unit. Infect. Control Hosp. Epidemiol. 2013, 34, 479–486. [Google Scholar] [CrossRef] [PubMed]
- Oberauner, L.; Zachow, C.; Lackner, S.; Högenauer, C.; Smolle, K.; Berg, G. The Ignored Diversity: Complex Bacterial Communities in Intensive Care Units Revealed by 16S Pyrosequencing. Sci. Rep. 2013, 3. [Google Scholar] [CrossRef] [PubMed]
- Shousha, A.; Awaiwanont, N.; Sofka, D.; Smulders, F.J.M.; Paulsen, P.; Szostak, M.P.; Humphrey, T.; Hilbert, F. Bacteriophages Isolated from Chicken Meat and the Horizontal Transfer of Antimicrobial Resistance Genes. Appl. Environ. Microbiol. 2015, 81, 4600–4606. [Google Scholar] [CrossRef]
- The European Committee on Antimicrobial Susceptibility Testing. Breakpoint Tables for Interpretation of MICs and Zone Diameters. Version 71 2017. Available online: http://www.eucast.org (accessed on 2 March 2017).
- International Organization for Standardization. Water Quality—Detection and Enumeration of Bacteriophages. Part 2. Enumeration of Somatic Coliphages; ISO 10705-2:2000; ISO: Geneva, Switzerland, 2000; Volume 91. [Google Scholar]
- Hilbert, M.; Csadek, I.; Auer, U.; Hilbert, F. Antimicrobial Resistance-Transducing Bacteriophages Isolated from Surfaces of Equine Surgery Clinics—A Pilot Study. Eur. J. Microbiol. Immunol. 2017, 7, 296–302. [Google Scholar] [CrossRef]
- García, P.; Madera, C.; Martínez, B.; Rodríguez, A.; Suárez, J.E. Prevalence of Bacteriophages Infecting Staphylococcus aureus in Dairy Samples and Their Potential as Biocontrol Agents. J. Dairy Sci. 2009, 92, 3019–3026. [Google Scholar] [CrossRef]
- Groisman, E.A. In vivo Genetic Engineering with Bacteriophage Mu. Methods Enzymol. 1991, 204, 180–212. [Google Scholar] [CrossRef] [PubMed]
- Salem, M.; Skurnik, M. Genomic Characterization of Sixteen Yersinia enterocolitica-Infecting Podoviruses of Pig Origin. Viruses 2018, 10, 174. [Google Scholar] [CrossRef] [PubMed]
- Maniloff, J.; Ackermann, H. Taxonomy of Bacterial Viruses: Establishment of Tailed Virus Genera and the Order Caudovirales. Arch. Virol. 1998, 143, 2051–2063. [Google Scholar] [CrossRef]
- Briggs, C.E.; Fratamico, P.M. Molecular Characterization of an Antibiotic Resistance Gene Cluster of Salmonella typhimurium DT104. Antimicrob. Agents Chemother. 1999, 43, 846–849. [Google Scholar] [CrossRef]
- Pittet, D.; Allegranzi, B.; Sax, H.; Dharan, S.; Pessoa-Silva, C.L.; Donaldson, L.; Boyce, J. M Evidence-Based Model for Hand Transmission During Patient Care and the Role of Improved Practices. Lancet Infect. Dis. 2006, 6, 641–652. [Google Scholar] [CrossRef]
- Fijan, S.; Turk, S.Š. Hospital Textiles, Are They a Possible Vehicle for Healthcare-Associated Infections? Int. J. Environ. Res. Public Health 2012, 9, 3330–3343. [Google Scholar] [CrossRef]
- Handorean, A.; Robertson, C.E.; Harris, J.K.; Frank, D.; Hull, N.; Kotter, C.; Stevens, M.J.; Baumgardner, D.; Pace, N.R.; Hernandez, M. Microbial Aerosol Liberation from Soiled Textiles Isolated During Routine Residuals Handling in a Modern Health Care Setting. Microbiome 2015, 3. [Google Scholar] [CrossRef] [PubMed]
- Hu, H.; Johani, K.; Gosbell, I.B.; Jacombs, A.S.W.; Almatroudi, A. Intensive Care Unit Environmental Surfaces are Contaminated by Multidrug-Resistant Bacteria in Biofilms: Combined Results of Conventional Culture, Pyrosequencing, Scanning Electron Microscopy, and Confocal Laser Microscopy. J. Hosp. Infect. 2015, 91, 35–44. [Google Scholar] [CrossRef]
- Marti, E.; Variatza, E.; Balcazar, J.L. Bacteriophages as a Reservoir of Extended-Spectrum Beta-Lactamase and Fluoroquinolone Resistance Genes in the Environment. Clin. Microbiol. Infect. 2014, 20, O456–O459. [Google Scholar] [CrossRef]
- Allué-Guardia, A.; Imamovic, L.; Muniesa, M. Evolution of a Self-Inducible Cytolethal Distending Toxin Type V- Encoding Bacteriophage from Escherichia coli O157:H7 to Shigella sonnei. J. Virol. 2013, 87, 13665–13675. [Google Scholar] [CrossRef]
- Górski, A.; Międzybrodzki, R.; Łobocka, M.; Głowacka-Rutkowska, A.; Bednarek, A.; Borysowski, J.; Jończyk-Matysiak, E.; Łusiak-Szelachowska, M.; Weber-Dąbrowska, B.; Bagińska, N.; et al. Phage Therapy: What Have We Learned? Viruses 2018, 10, 288. [Google Scholar] [CrossRef] [PubMed]
- Moye, Z.D.; Woolston, J.; Sulakvelidze, A. Bacteriophage Applications for Food Production and Processing. Viruses 2018, 10, 205. [Google Scholar] [CrossRef] [PubMed]
- Wahida, A.; Ritter, K.; Horz, H. The Janus-Face of Bacteriophages across Human Body Habitats. PLoS Pathog. 2016, 12, e1005634. [Google Scholar] [CrossRef] [PubMed]
- Ho, Y.-H.; Tseng, C.-C.; Wang, L.-S.; Chen, Y.-T.; Ho, G.-J.; Lin, T.-Y.; Wang, L.-Y.; Chen, L.-K. Application of Bacteriophage-Containing Aerosol against Nosocomial Transmission of Carbapenem-Resistant Acinetobacter baumannii in an Intensive Care Unit. PLoS ONE 2016, 11, e0168380. [Google Scholar] [CrossRef] [PubMed]
- Philipson, C.W.; Voegtly, L.J.; Lueder, M.R.; Long, K.A.; Rice, G.K.; Frey, K.G.; Biswas, B.; Cer, R.Z.; Hamilton, T.; Bishop-Lilly, K.A. Characterizing Phage Genomes for Therapeutic Applications. Viruses 2018, 10, 188. [Google Scholar] [CrossRef] [PubMed]
- Hilbert, M. Übertragung von Antibiotikaresistenzen von Phagen auf Bakterien im Veterinärmedizinischen Umfeld; Fachhochschule Wiener Neustadt: Wiener Neustadt, Austria, 2016. [Google Scholar]
- Campagna, C.; Villion, M.; Labrie, S.J.; Duchaine, C.; Moineau, S. Inactivation of Dairy Bacteriophages by Commercial Sanitizers and Disinfectants. Int. J. Food Microbiol. 2014, 171, 41–47. [Google Scholar] [CrossRef] [PubMed]
- Schuelke. TPH Protect n.d. Available online: https://www.schuelke.com/at-de/products/TPH-protect.php (accessed on 12 December 2016).

| Patient ID | Type of Sample | Collection Date | Number of Sites |
|---|---|---|---|
| P1 | Gel swab | 21.02.17 | 5 |
| P2 | Gel swab | 21.02.17 | 5 |
| P3 | Gel swab | 05.04.17 | 6 |
| P4 | Gel swab | 11.05.17 | 7 |
| P5 1 | Gel swab, liquid swab | 21.06.17 | 6, 6 |
| P6 | Liquid swab | 06.07.17 | 6 |
| P7 | Liquid swab | 06.07.17 | 6 |
| P8 | Liquid swab | 06.07.17 | 6 |
| P9 | Liquid swab | 26.07.17 | 5 |
| P10 | Liquid swab | 26.07.17 | 5 |
| P11 2 | Surface wipe | 26.03.18 | 6 |
| P12 2 | Surface wipe | 26.03.18 | 8 |
| Patient ID | Bacterial Isolate | Isolation Source | Colony Count 2 | Antibiogram | Mitomycin C-Inducible Prophage |
|---|---|---|---|---|---|
| P1 | S. aureus | Nose swab | +++ | Sensitive | - |
| P2 | S. aureus | Trachea secretion | +++ | P R, MXF R | - |
| P3 | E. coli1 | Urine transurethral catheter | 107/mL | AM R, SAM R, CPD R, CTX R, CXM R, CIP R, CN R, TMP R | - |
| P4 | E. coli | Urine transurethral catheter | 104/mL | AM R, SAM R | - |
| P5 | E. coli | Urine transurethral catheter | 107/mL | AM R, TMP R | + |
| P6 | K. pneumoniae | Trachea secretion | +++ | AM R | - |
| P7 | K. pneumoniae 1 | Trachea secretion | + | AM R, SAM R, CPD R, CTX R, CXM R, TMP R | - |
| P8 | K. pneumoniae | Trachea secretion | + | AM R | - |
| P9 | S. aureus | Blood culture | Not detectable | Sensitive | + |
| P10 | K. oxytoca | Trachea secretion | + | AM R, SAM R, CXM R | - |
| vB_EcoM_P5 | vB_SauS_P9 | ||||||
|---|---|---|---|---|---|---|---|
| Bacteria | Lysis | E.O.P. | Reference and/or Source | Bacteria | Lysis | E.O.P. | Reference and/or Source |
| Culture collection strains | Staphylococcus aureus from surface samples | ||||||
| Escherichia coli DSM 12242 | + | 1.000 ± 0.000 | DSMZ | Surface P1 (0/9) 1 | - | - | Environment, this study |
| Escherichia coli W3110 (ATCC 27325) | - | - | ATCC | Surface P2 (0/5) | - | - | Environment, this study |
| Escherichia coli JM109 (DSM 3423) | + | 0.005 ± 0.001 | DSM | Surface P3 (0/4) | - | - | Environment, this study |
| Escherichia coli DH5α (DSM 6897) | - | - | DSM | Surface P4 (0/2) | - | - | Environment, this study |
| Escherichia coli ATCC 11303 | - | - | ATCC | Surface P5 (7/11) | + | 0.142 ± 0.080–0.781 ± 0.219 | Environment, this study |
| Escherichia coli MC1061 (ATCC 53338) | + | 0.009 ± 0.001 | ATCC | Surface P6 (0/8) | - | - | Environment, this study |
| Klebsiella pneumoniae sub. pneumoniae ATCC 13883 | - | - | ATCC | Surface P7 (7/7) | + | 0.000 ± 0.000–0.001 ± 0.000 | Environment, this study |
| Yersinia enterocolitica sub. palearctica DSM11502 | - | - | DSM | Surface P8 (7/11) | + | 0.000 ± 0.000–0.013 ± 0.009 | Environment, this study |
| Salmonella enterica sub. enterica ATCC 14028 | - | - | ATCC | Surface P9 (0/5) | - | - | Environment, this study |
| Salmonella typhimurium DT104 isolate H3380 | - | - | Human [40] | Surface P10 (0/7) | - | - | Environment, this study |
| Patient isolates | Surface P11 (0/1) | - | - | Environment, this study | |||
| E. coli isolate 19 (P3) | - | - | Human, this study | Surface P12 (0/3) | - | - | Environment, this study |
| E. coli isolate 33 (P4) | - | - | Human, this study | Culture collection | |||
| E. coli isolate 46 (P5) | - | - | Human, this study | Staphylococcus aureus Sa9 | + | 1.0 ± 0.00 | Food, [36] |
| K. pneumoniae isolate 77 (P6) | - | - | Human, this study | Staphylococcus aureus ATCC 33862 | - | - | ATCC |
| K. pneumoniae isolate 76 (P7) | - | - | Human, this study | Staphylococcus aureus NCTC 6571 | - | - | NCTC |
| K. pneumoniae isolate 75 (P8) | - | - | Human, this study | Patient isolates | |||
| K. oxytoca isolate 89 (P10) | - | - | Human, this study | S. aureus isolate 11 (P1) | - | - | Human, this study |
| Clinical isolates | S. aureus isolate 12 (P2) | - | - | Human, this study | |||
| E. coli (n = 20) | - | - | Human, this study | S. aureus isolate 83 (P9) | - | - | Human, this study |
| Clinical isolates | |||||||
| S. aureus isolate 24 | + | 0.017 ± 0.013 | Human, this study | ||||
| S. aureus isolates (n = 16) | - | - | Human, this study | ||||
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Pacífico, C.; Hilbert, M.; Sofka, D.; Dinhopl, N.; Pap, I.-J.; Aspöck, C.; Hilbert, F. Characterization of Bacteria and Inducible Phages in an Intensive Care Unit. J. Clin. Med. 2019, 8, 1433. https://doi.org/10.3390/jcm8091433
Pacífico C, Hilbert M, Sofka D, Dinhopl N, Pap I-J, Aspöck C, Hilbert F. Characterization of Bacteria and Inducible Phages in an Intensive Care Unit. Journal of Clinical Medicine. 2019; 8(9):1433. https://doi.org/10.3390/jcm8091433
Chicago/Turabian StylePacífico, Cátia, Miriam Hilbert, Dmitrij Sofka, Nora Dinhopl, Ildiko-Julia Pap, Christoph Aspöck, and Friederike Hilbert. 2019. "Characterization of Bacteria and Inducible Phages in an Intensive Care Unit" Journal of Clinical Medicine 8, no. 9: 1433. https://doi.org/10.3390/jcm8091433
APA StylePacífico, C., Hilbert, M., Sofka, D., Dinhopl, N., Pap, I.-J., Aspöck, C., & Hilbert, F. (2019). Characterization of Bacteria and Inducible Phages in an Intensive Care Unit. Journal of Clinical Medicine, 8(9), 1433. https://doi.org/10.3390/jcm8091433





