Meta-Analysis of Differential miRNA Expression after Bariatric Surgery
Abstract
:1. Introduction
2. Methods
2.1. Search Strategies
2.2. Study Selection
2.3. Data Collection Process
2.4. Synthesis of Results
3. Results
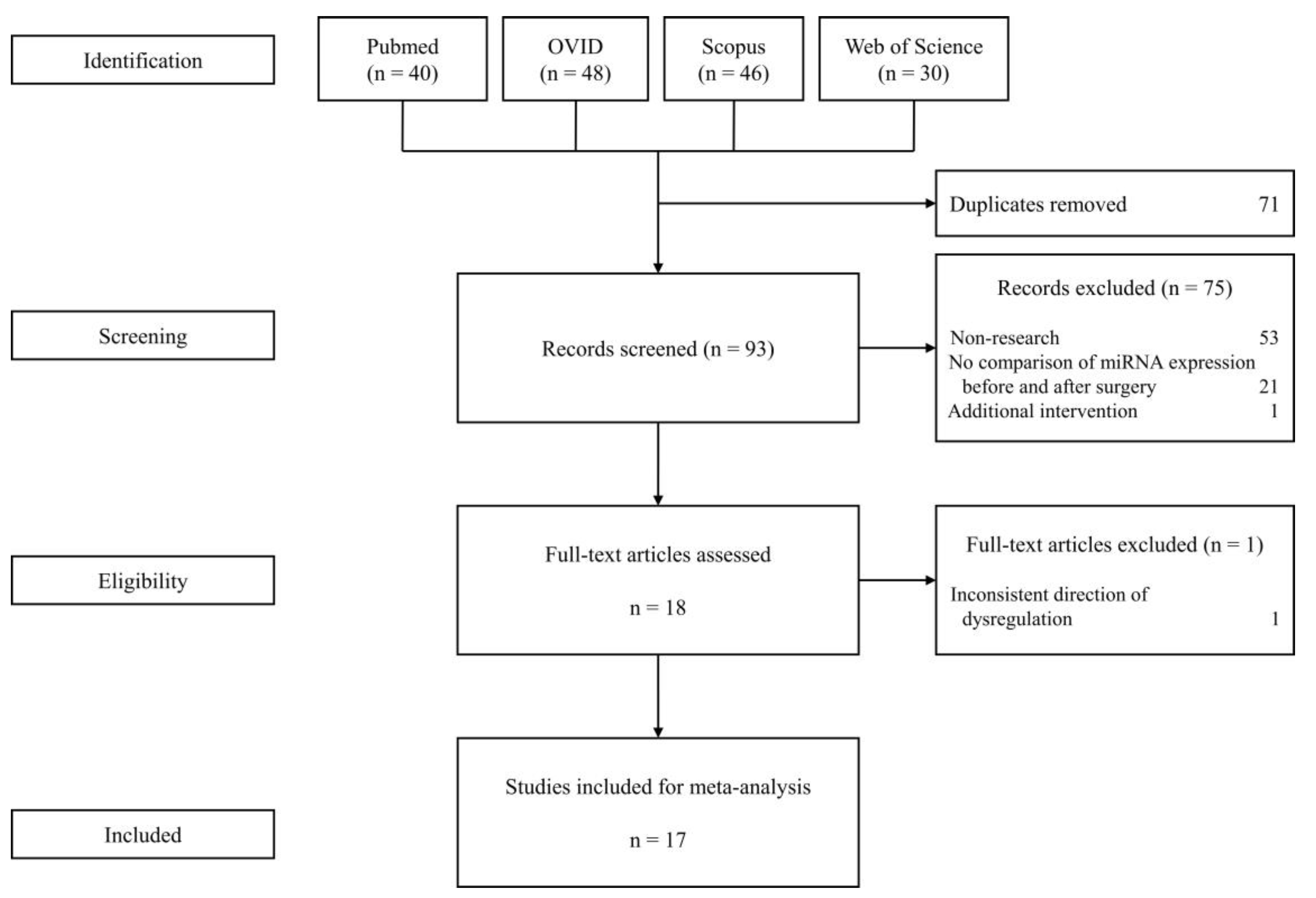
3.1. Selected Studies for the Meta-Analysis
3.2. Differential Expression of miRNA before and after Surgery
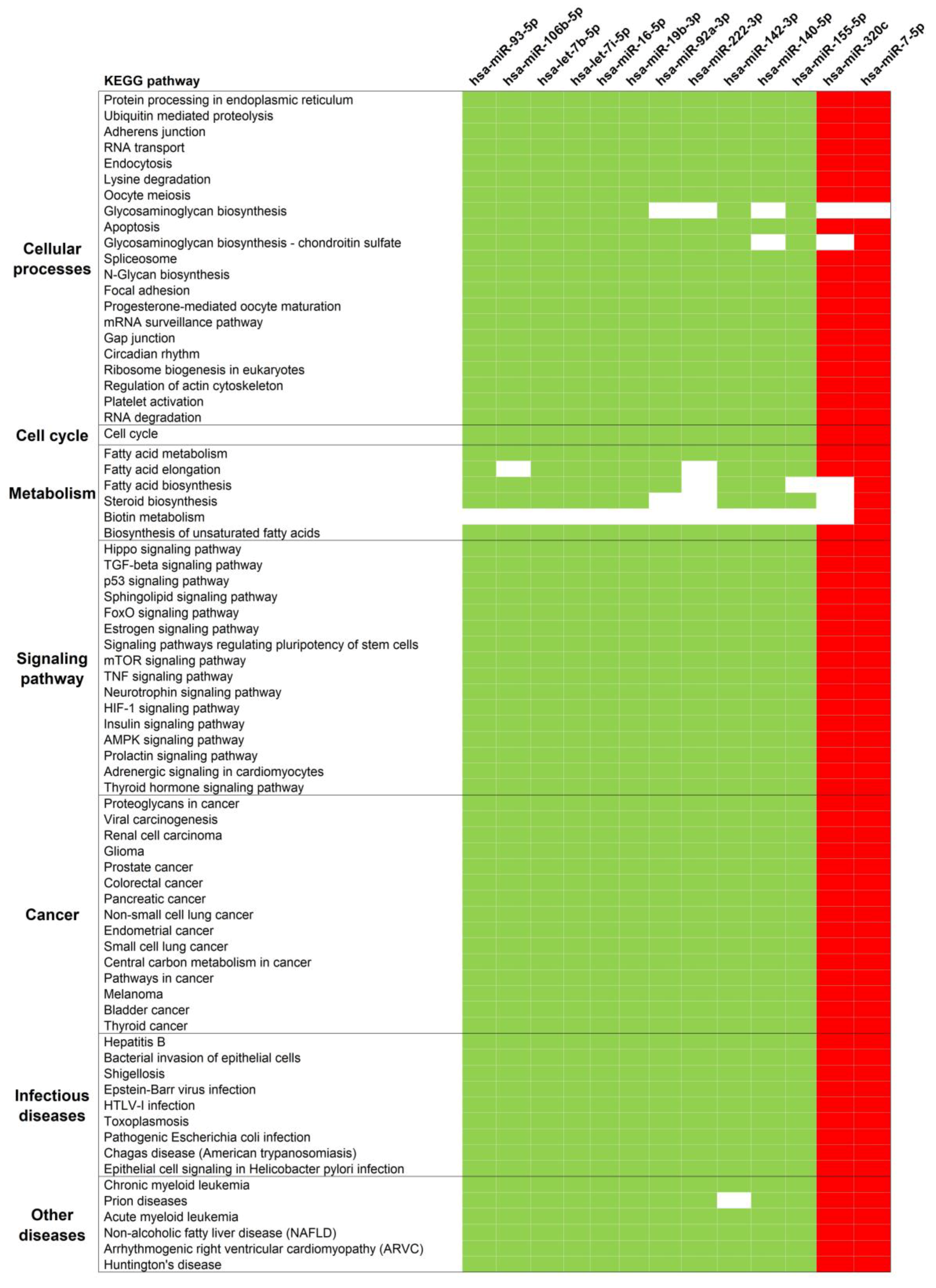
3.3. Pathway Analysis
4. Discussion
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Moshiri, M.; Osman, S.; Robinson, T.J.; Khandelwal, S.; Bhargava, P.; Rohrmann, C.A. Evolution of Bariatric Surgery: A Historical Perspective. Am. J. Roentgenol. 2013, 201, W40–W48. [Google Scholar] [CrossRef] [PubMed]
- Arterburn, D.E.; Courcoulas, A.P. Bariatric Surgery for Obesity and Metabolic Conditions in Adults. BMJ 2014, 349, g3961. [Google Scholar] [CrossRef] [PubMed]
- Maciejewski, M.L.; Arterburn, D.E.; Van Scoyoc, L.; Smith, V.A.; Yancy, W.S.; Weidenbacher, H.J.; Livingston, E.H.; Olsen, M.K. Bariatric Surgery and Long-Term Durability of Weight Loss. Jama Surg. 2016, 151, 1046. [Google Scholar] [CrossRef] [PubMed]
- Courcoulas, A.P.; King, W.C.; Belle, S.H.; Berk, P.; Flum, D.R.; Garcia, L.; Gourash, W.; Horlick, M.; Mitchell, J.E.; Pomp, A.; et al. Seven-Year Weight Trajectories and Health Outcomes in the Longitudinal Assessment of Bariatric Surgery (LABS) Study. Jama Surg. 2018, 153, 427. [Google Scholar] [CrossRef] [PubMed]
- Inge, T.H.; Courcoulas, A.P.; Jenkins, T.M.; Michalsky, M.P.; Helmrath, M.A.; Brandt, M.L.; Harmon, C.M.; Zeller, M.H.; Chen, M.K.; Xanthakos, S.A.; et al. Weight Loss and Health Status 3 Years after Bariatric Surgery in Adolescents. N. Engl. J. Med. 2016, 374, 113–123. [Google Scholar] [CrossRef] [PubMed]
- ASMBS. Estimate of Bariatric Surgery Numbers, 2011–2017 | American Society for Metabolic and Bariatric Surgery. Available online: https://asmbs.org/resources/estimate-of-bariatric-surgery-numbers (accessed on 29 July 2019).
- Angrisani, L.; Santonicola, A.; Iovino, P.; Formisano, G.; Buchwald, H.; Scopinaro, N. Bariatric Surgery Worldwide 2013. Obes. Surg. 2015, 25, 1822–1832. [Google Scholar] [CrossRef]
- Janik, M.R.; Stanowski, E.; Paśnik, K. Present Status of Bariatric Surgery in Poland. Videosurgery Miniinvasive Tech. 2016, 1, 22–25. [Google Scholar] [CrossRef]
- Chang, S.-H.; Stoll, C.R.T.; Song, J.; Varela, J.E.; Eagon, C.J.; Colditz, G.A. The Effectiveness and Risks of Bariatric Surgery: An Updated Systematic Review and Meta-Analysis, 2003–2012. Jama Surg. 2014, 149, 275. [Google Scholar] [CrossRef]
- Inge, T.H.; Jenkins, T.M.; Xanthakos, S.A.; Dixon, J.B.; Daniels, S.R.; Zeller, M.H.; Helmrath, M.A. Long-Term Outcomes of Bariatric Surgery in Adolescents with Severe Obesity (FABS-5+): A Prospective Follow-up Analysis. Lancet Diabetes Endocrinol. 2017, 5, 165–173. [Google Scholar] [CrossRef]
- Olbers, T.; Beamish, A.J.; Gronowitz, E.; Flodmark, C.-E.; Dahlgren, J.; Bruze, G.; Ekbom, K.; Friberg, P.; Göthberg, G.; Järvholm, K.; et al. Laparoscopic Roux-En-Y Gastric Bypass in Adolescents with Severe Obesity (AMOS): A Prospective, 5-Year, Swedish Nationwide Study. Lancet Diabetes Endocrinol. 2017, 5, 174–183. [Google Scholar] [CrossRef]
- Ibrahim, A.M.; Thumma, J.R.; Dimick, J.B. Reoperation and Medicare Expenditures After Laparoscopic Gastric Band Surgery. Jama Surg. 2017, 152, 835. [Google Scholar] [CrossRef]
- Lee, W.-J.; Chong, K.; Aung, L.; Chen, S.-C.; Ser, K.-H.; Lee, Y.-C. Metabolic Surgery for Diabetes Treatment: Sleeve Gastrectomy or Gastric Bypass? World J. Surg. 2017, 41, 216–223. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Lai, D.; Wu, D. Laparoscopic Roux-En-Y Gastric Bypass Versus Laparoscopic Sleeve Gastrectomy to Treat Morbid Obesity-Related Comorbidities: A Systematic Review and Meta-Analysis. Obes. Surg. 2016, 26, 429–442. [Google Scholar] [CrossRef] [PubMed]
- Sjöström, L.; Lindroos, A.-K.; Peltonen, M.; Torgerson, J.; Bouchard, C.; Carlsson, B.; Dahlgren, S.; Larsson, B.; Narbro, K.; Sjöström, C.D.; et al. Lifestyle, Diabetes, and Cardiovascular Risk Factors 10 Years after Bariatric Surgery. N. Engl. J. Med. 2004, 351, 2683–2693. [Google Scholar] [CrossRef] [PubMed]
- Poirier, P.; Cornier, M.-A.; Mazzone, T.; Stiles, S.; Cummings, S.; Klein, S.; McCullough, P.A.; Ren Fielding, C.; Franklin, B.A. Bariatric Surgery and Cardiovascular Risk Factors: A Scientific Statement from the American Heart Association. Circulation 2011, 123, 1683–1701. [Google Scholar] [CrossRef] [PubMed]
- Schauer, D.P.; Feigelson, H.S.; Koebnick, C.; Caan, B.; Weinmann, S.; Leonard, A.C.; Powers, J.D.; Yenumula, P.R.; Arterburn, D.E. Bariatric Surgery and the Risk of Cancer in a Large Multisite Cohort. Ann. Surg. 2019, 269, 95–101. [Google Scholar] [CrossRef]
- Dixon, J.B.; Zimmet, P.; Alberti, K.G.; Rubino, F. Bariatric Surgery: An IDF Statement for Obese Type 2 Diabetes. Diabet. Med. 2011, 28, 628–642. [Google Scholar] [CrossRef]
- Courcoulas, A.P.; Belle, S.H.; Neiberg, R.H.; Pierson, S.K.; Eagleton, J.K.; Kalarchian, M.A.; DeLany, J.P.; Lang, W.; Jakicic, J.M. Three-Year Outcomes of Bariatric Surgery vs. Lifestyle Intervention for Type 2 Diabetes Mellitus Treatment: A Randomized Clinical Trial. Jama Surg. 2015, 150, 931. [Google Scholar] [CrossRef]
- Cummings, D.E.; Arterburn, D.E.; Westbrook, E.O.; Kuzma, J.N.; Stewart, S.D.; Chan, C.P.; Bock, S.N.; Landers, J.T.; Kratz, M.; Foster-Schubert, K.E.; et al. Gastric Bypass Surgery vs Intensive Lifestyle and Medical Intervention for Type 2 Diabetes: The CROSSROADS Randomised Controlled Trial. Diabetologia 2016, 59, 945–953. [Google Scholar] [CrossRef]
- Ikramuddin, S.; Korner, J.; Lee, W.-J.; Connett, J.E.; Inabnet, W.B.; Billington, C.J.; Thomas, A.J.; Leslie, D.B.; Chong, K.; Jeffery, R.W.; et al. Roux-En-Y Gastric Bypass vs Intensive Medical Management for the Control of Type 2 Diabetes, Hypertension, and Hyperlipidemia: The Diabetes Surgery Study Randomized Clinical Trial. JAMA 2013, 309, 2240. [Google Scholar] [CrossRef]
- Mingrone, G.; Panunzi, S.; De Gaetano, A.; Guidone, C.; Iaconelli, A.; Nanni, G.; Castagneto, M.; Bornstein, S.; Rubino, F. Bariatric–Metabolic Surgery versus Conventional Medical Treatment in Obese Patients with Type 2 Diabetes: 5 Year Follow-up of an Open-Label, Single-Centre, Randomised Controlled Trial. Lancet 2015, 386, 964–973. [Google Scholar] [CrossRef]
- Rubino, F.; Nathan, D.M.; Eckel, R.H.; Schauer, P.R.; Alberti, K.G.M.M.; Zimmet, P.Z.; Del Prato, S.; Ji, L.; Sadikot, S.M.; Herman, W.H.; et al. Metabolic Surgery in the Treatment Algorithm for Type 2 Diabetes: A Joint Statement by International Diabetes Organizations. Diabetes Care 2016, 39, 861–877. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Aminian, A.; Brethauer, S.A.; Andalib, A.; Nowacki, A.S.; Jimenez, A.; Corcelles, R.; Hanipah, Z.N.; Punchai, S.; Bhatt, D.L.; Kashyap, S.R.; et al. Individualized Metabolic Surgery Score: Procedure Selection Based on Diabetes Severity. Ann. Surg. 2017, 266, 650–657. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.-C.; Hsu, N.-Y.; Lee, W.-J.; Chen, S.-C.; Ser, K.-H.; Lee, Y.-C. Prediction of Type 2 Diabetes Remission after Metabolic Surgery: A Comparison of the Individualized Metabolic Surgery Score and the ABCD Score. Surg. Obes. Relat. Dis. 2018, 14, 640–645. [Google Scholar] [CrossRef] [PubMed]
- Ashrafian, H.; Bueter, M.; Ahmed, K.; Suliman, A.; Bloom, S.R.; Darzi, A.; Athanasiou, T. Metabolic Surgery: An Evolution through Bariatric Animal Models: Metabolic and Bariatric Surgery Animal Models. Obes. Rev. 2010, 11, 907–920. [Google Scholar] [CrossRef] [PubMed]
- Li, J.V.; Ashrafian, H.; Bueter, M.; Kinross, J.; Sands, C.; le Roux, C.W.; Bloom, S.R.; Darzi, A.; Athanasiou, T.; Marchesi, J.R.; et al. Metabolic Surgery Profoundly Influences Gut Microbial-Host Metabolic Cross-Talk. Gut 2011, 60, 1214–1223. [Google Scholar] [CrossRef] [PubMed]
- Izquierdo, A.G.; Crujeiras, A.B. Obesity-Related Epigenetic Changes After Bariatric Surgery. Front. Endocrinol. 2019, 10, 232. [Google Scholar] [CrossRef]
- Ortega, F.J.; Mercader, J.M.; Catalan, V.; Moreno-Navarrete, J.M.; Pueyo, N.; Sabater, M.; Gomez-Ambrosi, J.; Anglada, R.; Fernandez-Formoso, J.A.; Ricart, W.; et al. Targeting the Circulating MicroRNA Signature of Obesity. Clin. Chem. 2013, 59, 781–792. [Google Scholar] [CrossRef] [Green Version]
- Alkandari, A.; Ashrafian, H.; Sathyapalan, T.; Sedman, P.; Darzi, A.; Holmes, E.; Athanasiou, T.; Atkin, S.L.; Gooderham, N.J. Improved Physiology and Metabolic Flux after Roux-En-Y Gastric Bypass Is Associated with Temporal Changes in the Circulating MicroRNAome: A Longitudinal Study in Humans. Bmc Obes. 2018, 5, 20. [Google Scholar] [CrossRef]
- Atkin, S.L.; Ramachandran, V.; Yousri, N.A.; Benurwar, M.; Simper, S.C.; McKinlay, R.; Adams, T.D.; Najafi-Shoushtari, S.H.; Hunt, S.C. Changes in Blood MicroRNA Expression and Early Metabolic Responsiveness 21 Days Following Bariatric Surgery. Front. Endocrinol. 2019, 9, 773. [Google Scholar] [CrossRef]
- Bae, Y.; Kim, Y.; Lee, H.; Kim, H.; Jeon, J.S.; Noh, H.; Han, D.C.; Ryu, S.; Kwon, S.H. Bariatric Surgery Alters MicroRNA Content of Circulating Exosomes in Patients with Obesity. Obesity 2019, 27, 264–271. [Google Scholar] [CrossRef] [PubMed]
- Blum, A.; Yehuda, H.; Geron, N.; Meerson, A. Elevated Levels of MiR-122 in Serum May Contribute to Improved Endothelial Function and Lower Oncologic Risk Following Bariatric Surgery. Isr. Med. Assoc. J. Imaj 2017, 19, 620–624. [Google Scholar] [PubMed]
- Guo, W.; Han, H.; Wang, Y.; Zhang, X.; Liu, S.; Zhang, G.; Hu, S. MiR-200a Regulates Rheb-Mediated Amelioration of Insulin Resistance after Duodenal–Jejunal Bypass. Int. J. Obes. 2016, 40, 1222–1232. [Google Scholar] [CrossRef] [PubMed]
- Wei, G.; Yi, S.; Yong, D.; Shaozhuang, L.; Guangyong, Z.; Sanyuan, H. MiR-320 Mediates Diabetes Amelioration after Duodenal-Jejunal Bypass via Targeting AdipoR1. Surg. Obes. Relat. Dis. 2018, 14, 960–971. [Google Scholar] [CrossRef] [PubMed]
- Hohensinner, P.J.; Kaun, C.; Ebenbauer, B.; Hackl, M.; Demyanets, S.; Richter, D.; Prager, M.; Wojta, J.; Rega-Kaun, G. Reduction of Premature Aging Markers After Gastric Bypass Surgery in Morbidly Obese Patients. Obes. Surg. 2018, 28, 2804–2810. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hubal, M.J.; Nadler, E.P.; Ferrante, S.C.; Barberio, M.D.; Suh, J.-H.; Wang, J.; Dohm, G.L.; Pories, W.J.; Mietus-Snyder, M.; Freishtat, R.J. Circulating Adipocyte-Derived Exosomal MicroRNAs Associated with Decreased Insulin Resistance after Gastric Bypass: Gastric Bypass Alters Exosomal MicroRNAs. Obesity 2017, 25, 102–110. [Google Scholar] [CrossRef] [PubMed]
- Hulsmans, M.; Sinnaeve, P.; Van der Schueren, B.; Mathieu, C.; Janssens, S.; Holvoet, P. Decreased MiR-181a Expression in Monocytes of Obese Patients Is Associated with the Occurrence of Metabolic Syndrome and Coronary Artery Disease. J. Clin. Endocrinol. Metab. 2012, 97, E1213–E1218. [Google Scholar] [CrossRef] [PubMed]
- Kwon, I.G.; Ha, T.K.; Ryu, S.-W.; Ha, E. Roux-En-Y Gastric Bypass Stimulates Hypothalamic MiR-122 and Inhibits Cardiac and Hepatic MiR-122 Expressions. J. Surg. Res. 2015, 199, 371–377. [Google Scholar] [CrossRef]
- Lirun, K.; Sewe, M.; Yong, W. A Pilot Study: The Effect of Roux-En-Y Gastric Bypass on the Serum MicroRNAs of the Type 2 Diabetes Patient. Obes. Surg. 2015, 25, 2386–2392. [Google Scholar] [CrossRef]
- Mysore, R.; Ortega, F.J.; Latorre, J.; Ahonen, M.; Savolainen-Peltonen, H.; Fischer-Posovszky, P.; Wabitsch, M.; Olkkonen, V.M.; Fernández-Real, J.M.; Haridas, P.A.N. MicroRNA-221-3p Regulates Angiopoietin-Like 8 (ANGPTL8) Expression in Adipocytes. J. Clin. Endocrinol. Metab. 2017, 102, 4001–4012. [Google Scholar] [CrossRef] [Green Version]
- Ortega, F.J.; Mercader, J.M.; Moreno-Navarrete, J.M.; Nonell, L.; Puigdecanet, E.; Rodriquez-Hermosa, J.I.; Rovira, O.; Xifra, G.; Guerra, E.; Moreno, M.; et al. Surgery-Induced Weight Loss Is Associated With the Downregulation of Genes Targeted by MicroRNAs in Adipose Tissue. J. Clin. Endocrinol. Metab. 2015, 100, E1467–E1476. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ortega, F.J.; Moreno, M.; Mercader, J.M.; Moreno-Navarrete, J.M.; Fuentes-Batllevell, N.; Sabater, M.; Ricart, W.; Fernández-Real, J.M. Inflammation Triggers Specific MicroRNA Profiles in Human Adipocytes and Macrophages and in Their Supernatants. Clin. Epigenetics 2015, 7, 49. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Wang, D.-S.; Cheng, Y.-S.; Jia, B.-L.; Yu, G.; Yin, X.-Q.; Wang, Y. Expression of MicroRNA-448 and SIRT1 and Prognosis of Obese Type 2 Diabetic Mellitus Patients After Laparoscopic Bariatric Surgery. Cell. Physiol. Biochem. 2018, 45, 935–950. [Google Scholar] [CrossRef] [PubMed]
- Wu, Q.; Li, J.V.; Seyfried, F.; le Roux, C.W.; Ashrafian, H.; Athanasiou, T.; Fenske, W.; Darzi, A.; Nicholson, J.K.; Holmes, E.; et al. Metabolic Phenotype-MicroRNA Data Fusion Analysis of the Systemic Consequences of Roux-En-Y Gastric Bypass Surgery. Int. J. Obes. 2015, 39, 1126–1134. [Google Scholar] [CrossRef] [PubMed]
- Lee, Y.; Kim, M.; Han, J.; Yeom, K.-H.; Lee, S.; Baek, S.H.; Kim, V.N. MicroRNA Genes Are Transcribed by RNA Polymerase II. Embo J. 2004, 23, 4051–4060. [Google Scholar] [CrossRef] [PubMed]
- Bartel, D.P. MicroRNAs. Cell 2004, 116, 281–297. [Google Scholar] [CrossRef]
- Kim, V.N.; Han, J.; Siomi, M.C. Biogenesis of Small RNAs in Animals. Nat. Rev. Mol. Cell Biol. 2009, 10, 126–139. [Google Scholar] [CrossRef]
- Krol, J.; Loedige, I.; Filipowicz, W. The Widespread Regulation of MicroRNA Biogenesis, Function and Decay. Nat. Rev. Genet. 2010, 11, 597–610. [Google Scholar] [CrossRef]
- Lim, L.P.; Lau, N.C.; Garrett-Engele, P.; Grimson, A.; Schelter, J.M.; Castle, J.; Bartel, D.P.; Linsley, P.S.; Johnson, J.M. Microarray Analysis Shows That Some MicroRNAs Downregulate Large Numbers of Target MRNAs. Nature 2005, 433, 769–773. [Google Scholar] [CrossRef]
- Hausser, J.; Zavolan, M. Identification and Consequences of MiRNA–Target Interactions—Beyond Repression of Gene Expression. Nat. Rev. Genet. 2014, 15, 599–612. [Google Scholar] [CrossRef]
- Ebert, M.S.; Sharp, P.A. Roles for MicroRNAs in Conferring Robustness to Biological Processes. Cell 2012, 149, 515–524. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Karbiener, M.; Fischer, C.; Nowitsch, S.; Opriessnig, P.; Papak, C.; Ailhaud, G.; Dani, C.; Amri, E.-Z.; Scheideler, M. MicroRNA MiR-27b Impairs Human Adipocyte Differentiation and Targets PPARγ. Biochem. Biophys. Res. Commun. 2009, 390, 247–251. [Google Scholar] [CrossRef] [PubMed]
- Plaisance, V.; Abderrahmani, A.; Perret-Menoud, V.; Jacquemin, P.; Lemaigre, F.; Regazzi, R. MicroRNA-9 Controls the Expression of Granuphilin/Slp4 and the Secretory Response of Insulin-Producing Cells. J. Biol. Chem. 2006, 281, 26932–26942. [Google Scholar] [CrossRef] [PubMed]
- Lu, H.; Buchan, R.J.; Cook, S.A. MicroRNA-223 Regulates Glut4 Expression and Cardiomyocyte Glucose Metabolism. Cardiovasc. Res. 2010, 86, 410–420. [Google Scholar] [CrossRef] [PubMed]
- Esau, C.; Davis, S.; Murray, S.F.; Yu, X.X.; Pandey, S.K.; Pear, M.; Watts, L.; Booten, S.L.; Graham, M.; McKay, R.; et al. MiR-122 Regulation of Lipid Metabolism Revealed by in Vivo Antisense Targeting. Cell Metab. 2006, 3, 87–98. [Google Scholar] [CrossRef] [PubMed]
- Oger, F.; Gheeraert, C.; Mogilenko, D.; Benomar, Y.; Molendi-Coste, O.; Bouchaert, E.; Caron, S.; Dombrowicz, D.; Pattou, F.; Duez, H.; et al. Cell-Specific Dysregulation of MicroRNA Expression in Obese White Adipose Tissue. J. Clin. Endocrinol. Metab. 2014, 99, 2821–2833. [Google Scholar] [CrossRef] [Green Version]
- Hilton, C.; Neville, M.J.; Karpe, F. MicroRNAs in Adipose Tissue: Their Role in Adipogenesis and Obesity. Int. J. Obes. 2013, 37, 325–332. [Google Scholar] [CrossRef]
- Trajkovski, M.; Hausser, J.; Soutschek, J.; Bhat, B.; Akin, A.; Zavolan, M.; Heim, M.H.; Stoffel, M. MicroRNAs 103 and 107 Regulate Insulin Sensitivity. Nature 2011, 474, 649–653. [Google Scholar] [CrossRef]
- Arner, P.; Kulyté, A. MicroRNA Regulatory Networks in Human Adipose Tissue and Obesity. Nat. Rev. Endocrinol. 2015, 11, 276–288. [Google Scholar] [CrossRef]
- Guay, C.; Regazzi, R. Circulating MicroRNAs as Novel Biomarkers for Diabetes Mellitus. Nat. Rev. Endocrinol. 2013, 9, 513–521. [Google Scholar] [CrossRef]
- Rottiers, V.; Näär, A.M. MicroRNAs in Metabolism and Metabolic Disorders. Nat. Rev. Mol. Cell Biol. 2012, 13, 239–250. [Google Scholar] [CrossRef] [PubMed]
- Xie, H.; Lim, B.; Lodish, H.F. MicroRNAs Induced During Adipogenesis That Accelerate Fat Cell Development Are Downregulated in Obesity. Diabetes 2009, 58, 1050–1057. [Google Scholar] [CrossRef] [PubMed]
- Hulsmans, M.; De Keyzer, D.; Holvoet, P. MicroRNAs Regulating Oxidative Stress and Inflammation in Relation to Obesity and Atherosclerosis. Faseb J. 2011, 25, 2515–2527. [Google Scholar] [CrossRef] [PubMed]
- Bramer, W.M.; Rethlefsen, M.L.; Kleijnen, J.; Franco, O.H. Optimal Database Combinations for Literature Searches in Systematic Reviews: A Prospective Exploratory Study. Syst. Rev. 2017, 6, 245. [Google Scholar] [CrossRef] [PubMed]
- Zhu, H.; Leung, S. Identification of Potential MicroRNA Biomarkers by Meta-Analysis. In Computational Drug Discovery and Design; Gore, M., Jagtap, U.B., Eds.; Springer: New York, NY, USA, 2018; Volume 1762, pp. 473–484. [Google Scholar] [CrossRef]
- Vlachos, I.S.; Zagganas, K.; Paraskevopoulou, M.D.; Georgakilas, G.; Karagkouni, D.; Vergoulis, T.; Dalamagas, T.; Hatzigeorgiou, A.G. DIANA-MiRPath v3.0: Deciphering MicroRNA Function with Experimental Support. Nucleic Acids Res. 2015, 43, W460–W466. [Google Scholar] [CrossRef] [PubMed]
- The FANTOM Consortium; de Rie, D.; Abugessaisa, I.; Alam, T.; Arner, E.; Arner, P.; Ashoor, H.; Åström, G.; Babina, M.; Bertin, N.; et al. An Integrated Expression Atlas of MiRNAs and Their Promoters in Human and Mouse. Nat. Biotechnol. 2017, 35, 872–878. [Google Scholar] [CrossRef]
- Martín-Núñez, G.M.; Cabrera-Mulero, A.; Alcaide-Torres, J.; García-Fuentes, E.; Tinahones, F.J.; Morcillo, S. No Effect of Different Bariatric Surgery Procedures on LINE-1 DNA Methylation in Diabetic and Nondiabetic Morbidly Obese Patients. Surg. Obes. Relat. Dis. 2017, 13, 442–450. [Google Scholar] [CrossRef]
- Balasar, Ö.; Çakır, T.; Erkal, Ö.; Aslaner, A.; Çekiç, B.; Uyar, M.; Bülbüller, N.; Oruç, M.T. The Effect of Rs9939609 FTO Gene Polymorphism on Weight Loss after Laparoscopic Sleeve Gastrectomy. Surg. Endosc. 2016, 30, 121–125. [Google Scholar] [CrossRef]
- Käkelä, P.; Jääskeläinen, T.; Torpström, J.; Ilves, I.; Venesmaa, S.; Pääkkönen, M.; Gylling, H.; Paajanen, H.; Uusitupa, M.; Pihlajamäki, J. Genetic Risk Score Does Not Predict the Outcome of Obesity Surgery. Obes. Surg. 2014, 24, 128–133. [Google Scholar] [CrossRef]
- Zhang, W.; Lu, W.; Ananthan, S.; Suto, M.J.; Li, Y. Discovery of Novel Frizzled-7 Inhibitors by Targeting the Receptor’s Transmembrane Domain. Oncotarget 2017, 8. [Google Scholar] [CrossRef]
- Christodoulides, C.; Scarda, A.; Granzotto, M.; Milan, G.; Dalla Nora, E.; Keogh, J.; De Pergola, G.; Stirling, H.; Pannacciulli, N.; Sethi, J.K.; et al. WNT10B Mutations in Human Obesity. Diabetologia 2006, 49, 678–684. [Google Scholar] [CrossRef] [PubMed]
- Mayas, M.D.; Ortega, F.J.; Macías-González, M.; Bernal, R.; Gómez-Huelgas, R.; Fernández-Real, J.M.; Tinahones, F.J. Inverse Relation between FASN Expression in Human Adipose Tissue and the Insulin Resistance Level. Nutr. Metab. 2010, 7, 3. [Google Scholar] [CrossRef] [PubMed]
- Wang, D.; DuBois, R.N. Associations Between Obesity and Cancer: The Role of Fatty Acid Synthase. Jnci J. Natl. Cancer Inst. 2012, 104, 343–345. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- O’Connell, R.M.; Rao, D.S.; Baltimore, D. MicroRNA Regulation of Inflammatory Responses. Annu. Rev. Immunol. 2012, 30, 295–312. [Google Scholar] [CrossRef] [PubMed]
- Moore, C.S.; Rao, V.T.S.; Durafourt, B.A.; Bedell, B.J.; Ludwin, S.K.; Bar-Or, A.; Antel, J.P. MiR-155 as a Multiple Sclerosis–Relevant Regulator of Myeloid Cell Polarization. Ann. Neurol. 2013, 74, 709–720. [Google Scholar] [CrossRef]
- Faraoni, I.; Antonetti, F.R.; Cardone, J.; Bonmassar, E. MiR-155 Gene: A Typical Multifunctional MicroRNA. Biochim. Biophys. Acta Bba Mol. Basis Dis. 2009, 1792, 497–505. [Google Scholar] [CrossRef]
- Ding, S.; Huang, H.; Xu, Y.; Zhu, H.; Zhong, C. MiR-222 in Cardiovascular Diseases: Physiology and Pathology. Biomed Res. Int. 2017, 2017. [Google Scholar] [CrossRef]
- Li, Y.; Kowdley, K.V. Method for MicroRNA Isolation from Clinical Serum Samples. Anal. Biochem. 2012, 431, 69–75. [Google Scholar] [CrossRef]
- Kroh, E.M.; Parkin, R.K.; Mitchell, P.S.; Tewari, M. Analysis of Circulating MicroRNA Biomarkers in Plasma and Serum Using Quantitative Reverse Transcription-PCR (QRT-PCR). Methods 2010, 50, 298–301. [Google Scholar] [CrossRef]
- Chugh, P.; Dittmer, D.P. Potential Pitfalls in MicroRNA Profiling. Wiley Interdiscip. Rev. Rna 2012, 3, 601–616. [Google Scholar] [CrossRef]
- Git, A.; Dvinge, H.; Salmon-Divon, M.; Osborne, M.; Kutter, C.; Hadfield, J.; Bertone, P.; Caldas, C. Systematic Comparison of Microarray Profiling, Real-Time PCR, and next-Generation Sequencing Technologies for Measuring Differential MicroRNA Expression. RNA 2010, 16, 991–1006. [Google Scholar] [CrossRef] [PubMed]
- Sato, F.; Tsuchiya, S.; Terasawa, K.; Tsujimoto, G. Intra-Platform Repeatability and Inter-Platform Comparability of MicroRNA Microarray Technology. PLoS ONE 2009, 4, e5540. [Google Scholar] [CrossRef] [PubMed]
- Schwarzenbach, H.; da Silva, A.M.; Calin, G.; Pantel, K. Which Is the Accurate Data Normalization Strategy for MicroRNA Quantification? Clin. Chem. 2015, 61, 1333–1342. [Google Scholar] [CrossRef] [PubMed]
- Vandesompele, J.; De Preter, K.; Pattyn, F.; Poppe, B.; Van Roy, N.; De Paepe, A.; Speleman, F. Accurate Normalization of Real-Time Quantitative RT-PCR Data by Geometric Averaging of Multiple Internal Control Genes. Genome Biol. 2002, 3, RESEARCH0034. [Google Scholar] [CrossRef] [PubMed]
- Huang, R.S.; Gamazon, E.R.; Ziliak, D.; Wen, Y.; Im, H.K.; Zhang, W.; Wing, C.; Duan, S.; Bleibel, W.K.; Cox, N.J.; et al. Population Differences in MicroRNA Expression and Biological Implications. Rna Biol. 2011, 8, 692–701. [Google Scholar] [CrossRef] [PubMed]
- Rawlings-Goss, R.A.; Campbell, M.C.; Tishkoff, S.A. Global Population-Specific Variation in MiRNA Associated with Cancer Risk and Clinical Biomarkers. Bmc Med. Genom. 2014, 7, 53. [Google Scholar] [CrossRef] [PubMed]
- Cui, C.; Yang, W.; Shi, J.; Zhou, Y.; Yang, J.; Cui, Q.; Zhou, Y. Identification and Analysis of Human Sex-Biased MicroRNAs. Genom. Proteom. Bioinform. 2018, 16, 200–211. [Google Scholar] [CrossRef] [PubMed]
- Guo, L.; Zhang, Q.; Ma, X.; Wang, J.; Liang, T. MiRNA and MRNA Expression Analysis Reveals Potential Sex-Biased MiRNA Expression. Sci. Rep. 2017, 7, 39812. [Google Scholar] [CrossRef]
| Study | Year | Country | Sample Size | Sex (Males/Females) |
|---|---|---|---|---|
| Human studies (comparing before vs. after bariatric surgery) | ||||
| Ortega et al. [29] | 2013 | Spain | 22 | 5/17 |
| Alkandari et al. [30] | 2018 | UK | 9 | 4/5 |
| Atkin et al. [31] | 2019 | USA | 29 | 9/20 |
| Bae et al. [32] | 2019 | South Korea | 12 | Unspecified |
| Blum et al. [33] | 2017 | Israel | 21 | 14/7 |
| Hohensinner et al. [36] | 2018 | Austria | 58 | 17/41 |
| Hubal et al. [37] | 2017 | USA | 6 | 0/6 |
| Hulsmans et al. [38] | 2012 | Belgium | 21 | 7/14 |
| Lirun et al. [40] | 2015 | China | 18 | 4/11 |
| Mysore et al. [41] | 2017 | Spain | 22 | 0/22 |
| Ortega et al. [42] | 2015 | Spain | 25 | 0/25 |
| Ortega et al. [43] | 2015 | Spain | 9 | 0/9 |
| Wang et al. [44] | 2018 | China | 124 | 46/78 |
| Animal studies (comparing bariatric vs sham surgery) | ||||
| Guo et al. [34] | 2016 | China | 35 | 35/0 |
| Wei et al. [35] | 2018 | China | 45 | 45/0 |
| Kwon et al. [39] | 2015 | South Korea | 25 | 25/0 |
| Wu et al. [45] | 2015 | UK | 12 | 12/0 |
| Study | Tissue | Isolation | Platform | Normalization |
|---|---|---|---|---|
| Human studies | ||||
| Ortega et al. [29] | Plasma | mirVana PARIS Isolation Kit | TaqMan array miRNA cards in a subset and qPCR in the final sample | Geometric mean of six miRNAs (hsa-miR-106a-5p, hsa-miR-146a-5p, hsa-miR-19b-3p, hsa-miR-223-3p, hsa-miR-186-5p, hsa-miR-199a-3p) |
| Alkandari et al. [30] | Plasma | mirVana PARIS Isolation Kit | miRCURY qPCR panel | Four miRNAs (hsa-miR-223-3p, hsa-miR-26a-5p, hsa-miR-101-3p, and hsa-miR-19a-3p) |
| Atkin et al. [31] | Plasma | miRCURY RNA Isolation kit | qPCR and a FANTOM miRNA atlas [68] | Global mean |
| Bae et al. [32] | Exosome | miRNeasy Mini Kit | Small RNA sequencing | Relative log expression using DESeq2 |
| Blum et al. [33] | Serum | miRNeasy serum/plasma kit | RNA sequencing in a subset and qPCR in the final sample | hsa-miR-451a |
| Hohensinner et al. [36] | Plasma | miRNA tissue lysis kit | qPCR | RNA spike-in |
| Hubal et al. [37] | Exosome | mirVANA miRNA Isolation Kit | GeneChip miRNA 4.0 Array | RMA algorithm |
| Hulsmans et al. [38] | Monocytes | TRIzol reagent | qPCR | RNU5G |
| Lirun et al. [40] | Plasma | mirVana RNA Isolation Kit | GeneChip miRNA 3.0 Array | RMA algorithm |
| Mysore et al. [41] | Subcutaneous Adipose Tissue (SAT) | miRNeasy Mini kit | qPCR | RNU44 |
| Ortega et al. [42] | SAT | miRNeasy Mini Kit | GeneChip miRNA 3.0 array in a subset and qPCR in the final sample | RMA algorithm and RNU48 |
| Ortega et al. [43] | SAT | miRNeasy Mini Kit | qPCR | RNU6B |
| Wang et al. [44] | Circulating Endothelial Progenitor Cells | High Pure RNA kit | qPCR | RNU6 |
| Animal studies | ||||
| Guo et al. [34] | Liver | TRIzol reagent | miProfile Customized Rat qPCR arrays | 5S rRNA and RsnRNA U6 |
| Wei et al. [35] | Liver | TRIzol reagent | miProfile Customized Rat qPCR arrays | 5S rRNA, RsnRNA U6, rno-miR-25, and rno-miR-186 |
| Kwon et al. [39] | Hypothalamus, Heart, and Liver | Unspecified | Agilent Rat miRNA 8x15k microarray for hypothalamus and heart samples, then qPCR for liver and validation | Whole-array and RNU6 |
| Wu et al. [45] | Plasma and Liver | mirVANA PARIS RNA Isolation kit | TaqMan Array Rodent Card | RNU6-1, RNU6-2, rno-miR-16-5p, rno-miR-223-3p, mmu-miR-1937b |
| Study | Year | Bariatric Surgery Type | Time of Observation after Surgery |
|---|---|---|---|
| Human studies | |||
| Ortega et al. [29] | 2013 | RYGB | 12 months |
| Alkandari et al. [30] | 2018 | RYGB | 1, 3, 6, 9, and 12 months |
| Atkin et al. [31] | 2019 | RYGB | 21 days |
| Bae et al. [32] | 2019 | RYGB and SG | 6 months |
| Blum et al. [33] | 2017 | SG | 3 months |
| Hohensinner et al. [36] | 2018 | RYGB | 24 months |
| Hubal et al. [37] | 2017 | RYGB | 12 months |
| Hulsmans et al. [38] | 2012 | RYGB | 3 months |
| Lirun et al. [40] | 2015 | RYGB | 3 months |
| Mysore et al. [41] | 2017 | RYGB | 24 months |
| Ortega et al. [42] | 2015 | RYGB | 24 months |
| Ortega et al. [43] | 2015 | RYGB | 24 months |
| Wang et al. [44] | 2018 | Not specified | 3 months |
| Animal studies | |||
| Guo et al. [34] | 2016 | DJB and SG | 2, 4, 8 weeks |
| Wei et al. [35] | 2018 | DJB | 2, 4, 8 weeks |
| Kwon et al. [39] | 2015 | RYGB | 25 days |
| Wu et al. [45] | 2015 | RYGB | 53 days |
| miRNA | miRBase | References | Direction of Expression | No. of Subjects | Tissue | Time of Observation | |
|---|---|---|---|---|---|---|---|
| Group 1 miRNAs (same direction of expression after surgery in two or more studies) | |||||||
| 1 | hsa-miR-93-5p | MIMAT0000093 | Lirun [40] | − | 15 | Plasma | 3 months |
| Alkandari [30] | 9 | Plasma | 3 months | ||||
| 2 | hsa-miR-106b-5p | MIMAT0000680 | Lirun [40] | − | 15 | Plasma | 3 months |
| Alkandari [30] | 9 | Plasma | 3, 12 months | ||||
| 3 | hsa-let-7b-5p | MIMAT0000063 | Lirun [40] | − | 15 | Plasma | 3 months |
| Alkandari [30] | 9 | Plasma | 3 months | ||||
| 4 | hsa-let-7i-5p | MIMAT0000415 | Lirun [40] | − | 15 | Plasma | 3 months |
| Alkandari [30] | 9 | Plasma | 6, 9 months | ||||
| Atkin [31] | 29 | Plasma | 21 days | ||||
| 5 | hsa-miR-16-5p | MIMAT0000069 | Lirun [40] | − | 15 | Plasma | 3 months |
| Hubal [37] | 6 | Exosomes | 12 months | ||||
| 6 | hsa-miR-19b-3p | MIMAT0000074 | Ortega [43] | − | 9 | SAT | 24 months |
| Lirun [40] | 15 | Plasma | 3 months | ||||
| Ortega [29] | 22 | Plasma | 12 months | ||||
| 7 | hsa-miR-92a-3p | MIMAT0000092 | Lirun [40] | − | 15 | Plasma | 3 months |
| Alkandari [30] | 9 | Plasma | 9, 12 months | ||||
| 8 | hsa-miR-222-3p | MIMAT0000279 | Ortega [29] | − | 22 | Plasma | 12 months |
| Ortega [43] | 9 | SAT | 24 months | ||||
| 9 | hsa-miR-142-3p | MIMAT0000434 | Bae [32] | − | 12 | Exosome | 6 months |
| Ortega [29] | 22 | Plasma | 12 months | ||||
| 10 | hsa-miR-140-5p | MIMAT0000431 | Bae [32] | − | 12 | Exosome | 6 months |
| Ortega [29] | 22 | Plasma | 12 months | ||||
| 11 | hsa-miR-155-5p | MIMAT0000646 | Ortega [43] | − | 9 | SAT | 24 months |
| Ortega [42] | 25 | SAT | 24 months | ||||
| 12 | rno-miR-320-3p | MIMAT0000903 | Wu [45] | − | 4 | Plasma | 53 days |
| Wei [35] | 5 | liver | 2 months | ||||
| 13 | hsa-miR-320c | MIMAT0005793 | Atkin [31] | + | 29 | Plasma | 21 days |
| Lirun [40] | 15 | Plasma | 3 months | ||||
| 14 | hsa-miR-7-5p | MIMAT0000252 | Atkin [31] | + | 29 | Plasma | 21 days |
| Bae [32] | 12 | Exosome | 6 months | ||||
| Group 2 miRNAs (overall same direction of expression after surgery in two or more studies) | |||||||
| 1 | hsa-miR-125b-5p | MIMAT0000423 | Ortega [29] | − | 22 | Plasma | 12 months |
| Alkandari [30] | − | 9 | Plasma | 6, 9, 12 months | |||
| Hubal [37] | + | 6 | Exosomes | 12 months | |||
| 2 | hsa-miR-130b-3p | MIMAT0000691 | Ortega [42] | − | 25 | SAT | 24 months |
| Alkandari [30] | − | 9 | Plasma | 12 months | |||
| Ortega [29] | + | 22 | Plasma | 12 months | |||
| 3 | hsa-miR-221-3p | MIMAT0000278 | Ortega [43] | − | 9 | SAT | 24 months |
| Ortega [42] | − | 25 | SAT | 24 months | |||
| Mysore [41] | − | 22 | SAT | 24 months | |||
| Lirun [40] | − | 15 | Plasma | 3 months | |||
| Ortega [29] | + | 22 | Plasma | 12 months | |||
| 4 | rno-miR-122-5p | MIMAT0000827 | Kwon [39] | − | 25 | heart | 25 days |
| Kwon [39] | − | 25 | liver | 25 days | |||
| Wu [45] | − | 4 | Plasma | 53 days | |||
| Wu [45] | − | 8 | Liver | 53 days | |||
| Kwon [39] | + | 25 | hypothalamus | 25 days | |||
| 5 | hsa-miR-146a-5p | MIMAT0000449 | Lirun [40] | − | 15 | Plasma | 3 months |
| Ortega [43] | − | 9 | SAT | 24 months | |||
| Ortega [29] | + | 22 | Plasma | 12 months | |||
| 6 | rno-miR-503-5p | MIMAT0003213 | Kwon [39] | + | 25 | hypothalamus | 25 days |
| Kwon [39] | + | 25 | heart | 25 days | |||
| Wei [35] | − | 4 | liver | 2 months | |||
| Group 3 miRNAs (reported in at least two studies, but with no agreement in direction of expression) | |||||||
| 1 | hsa-miR-21-5p | MIMAT0000076 | Alkandari [30] | − | 9 | Plasma | 9, 12 months |
| Ortega [29] | + | 22 | Plasma | 12 months | |||
| 2 | hsa-miR-33a-5p | MIMAT0000091 | Bae [32] | − | 12 | Exosome | 6 months |
| Alkandari [30] | + | 9 | Plasma | 6 months | |||
| 3 | hsa-miR-320a-3p | MIMAT0000510 | Alkandari [30] | − | 9 | Plasma | 6, 9, 12 months |
| Lirun [40] | + | 15 | Plasma | 3 months | |||
| 4 | hsa-miR-320b | MIMAT0005792 | Alkandari [30] | − | 9 | Plasma | 9 months |
| Lirun [40] | + | 15 | Plasma | 3 months | |||
| 5 | hsa-miR-378a-3p | MIMAT0000732 | Alkandari [30] | − | 9 | Plasma | 6, 9, 12 months |
| Lirun [40] | + | 15 | Plasma | 3 months | |||
| 6 | hsa-miR-103-3p | MIMAT0000101 | Lirun [40] | − | 15 | Plasma | 3 months |
| Hubal [37] | + | 6 | Exosomes | 12 months | |||
| 7 | rno-miR-133b-3p | MIMAT0003126 | Wei [35] | − | 4 | liver | 2 months |
| Kwon [39] | + | 25 | hypothalamus | 25 days | |||
| 8 | rno-miR-194-5p | MIMAT0000869 | Kwon [39] | − | 25 | heart | 25 days |
| Guo [34] | + | 4 | liver | 2 months | |||
| 9 | hsa-miR-122-5p | MIMAT0000421 | Ortega [29] | − | 22 | Plasma | 12 months |
| Blum [33] | + | 21 | Serum | 3 months | |||
| 10 | rno-miR-146a-5p | MIMAT0000852 | Wu [45] | − | 4 | Plasma | 53 days |
| Kwon [39] | + | 25 | hypothalamus | 25 days | |||
| 11 | rno-miR-542-3p | MIMAT0003179 | Wei [35] | − | 4 | liver | 2 months |
| Kwon [39] | + | 25 | hypothalamus | 25 days | |||
| 12 | hsa-miR-191-5p | MIMAT0000440 | Lirun [40] | − | 15 | Plasma | 3 months |
| Bae [32] | + | 12 | Exosome | 6 months | |||
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Langi, G.; Szczerbinski, L.; Kretowski, A. Meta-Analysis of Differential miRNA Expression after Bariatric Surgery. J. Clin. Med. 2019, 8, 1220. https://doi.org/10.3390/jcm8081220
Langi G, Szczerbinski L, Kretowski A. Meta-Analysis of Differential miRNA Expression after Bariatric Surgery. Journal of Clinical Medicine. 2019; 8(8):1220. https://doi.org/10.3390/jcm8081220
Chicago/Turabian StyleLangi, Gladys, Lukasz Szczerbinski, and Adam Kretowski. 2019. "Meta-Analysis of Differential miRNA Expression after Bariatric Surgery" Journal of Clinical Medicine 8, no. 8: 1220. https://doi.org/10.3390/jcm8081220
APA StyleLangi, G., Szczerbinski, L., & Kretowski, A. (2019). Meta-Analysis of Differential miRNA Expression after Bariatric Surgery. Journal of Clinical Medicine, 8(8), 1220. https://doi.org/10.3390/jcm8081220





