Potential Role of the Microbiome in Acne: A Comprehensive Review
Abstract
1. Introduction
2. Methods
3. Skin Microbiota (Overview)
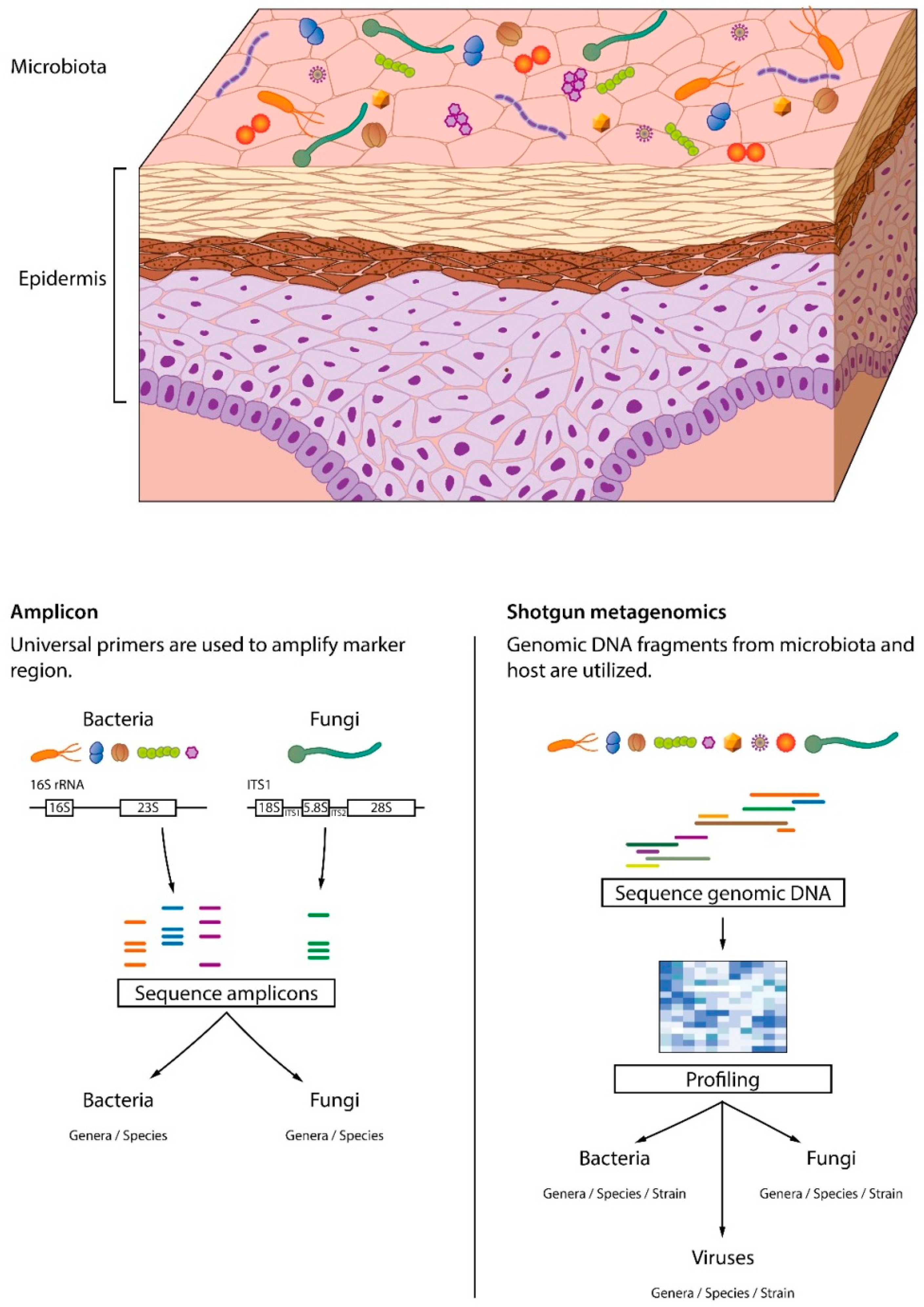
3.1. Skin Microbiome Sampling
3.2. Skin Microbiome Analysis
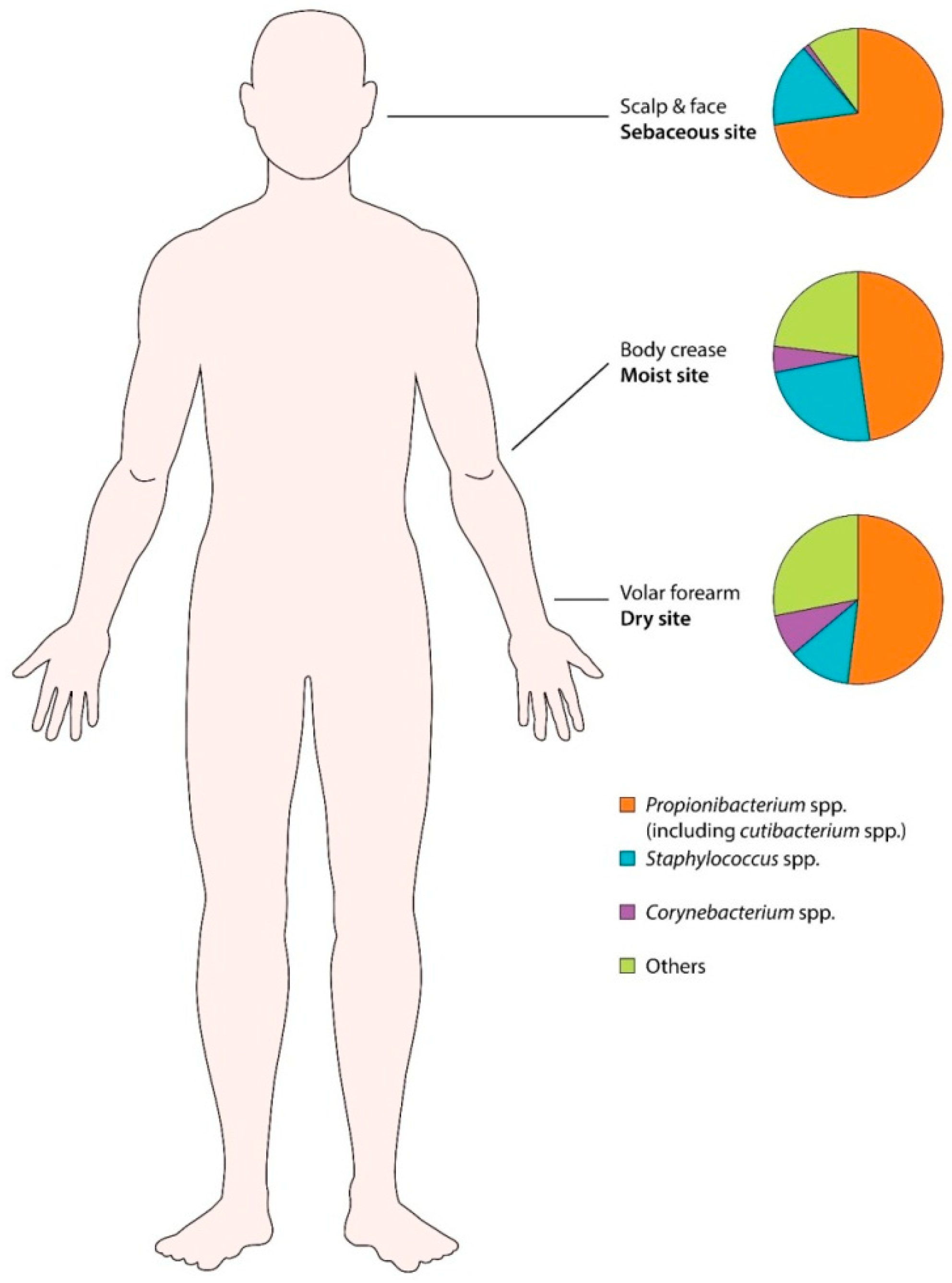
3.3. Human Skin Microbiota (Healthy Skin)
4. Skin Microbiota and Acne
4.1. Classification of C. acnes
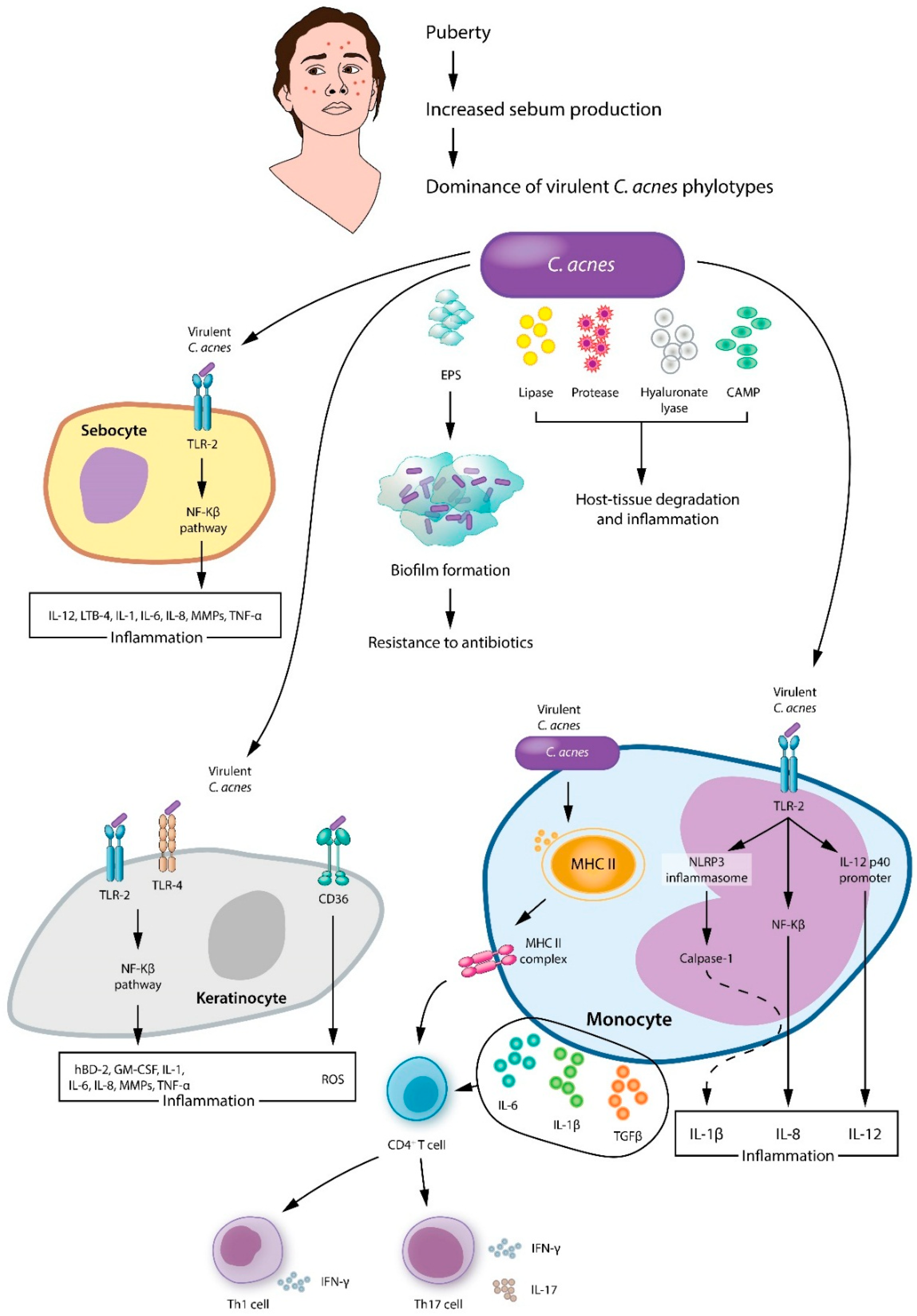
4.2. Cutibacterium acnes in Acne
4.3. Other Acne-Associated Microbiota
5. Acne Treatment and Skin Microbes
5.1. Antibiotics and the Acne Skin Microbiota
5.2. Antibiotic Resistance in the Microbiota of Skin with Acne
5.3. Skin Microbiota As a Biomarker for Acne Drug Development
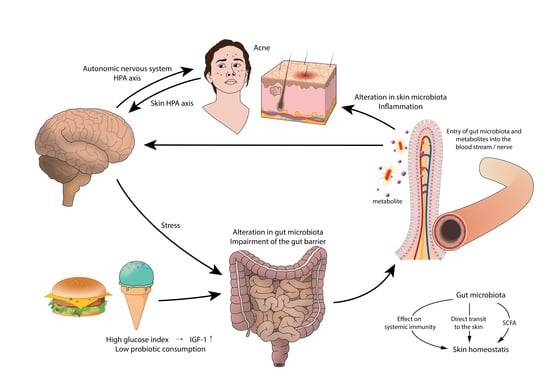
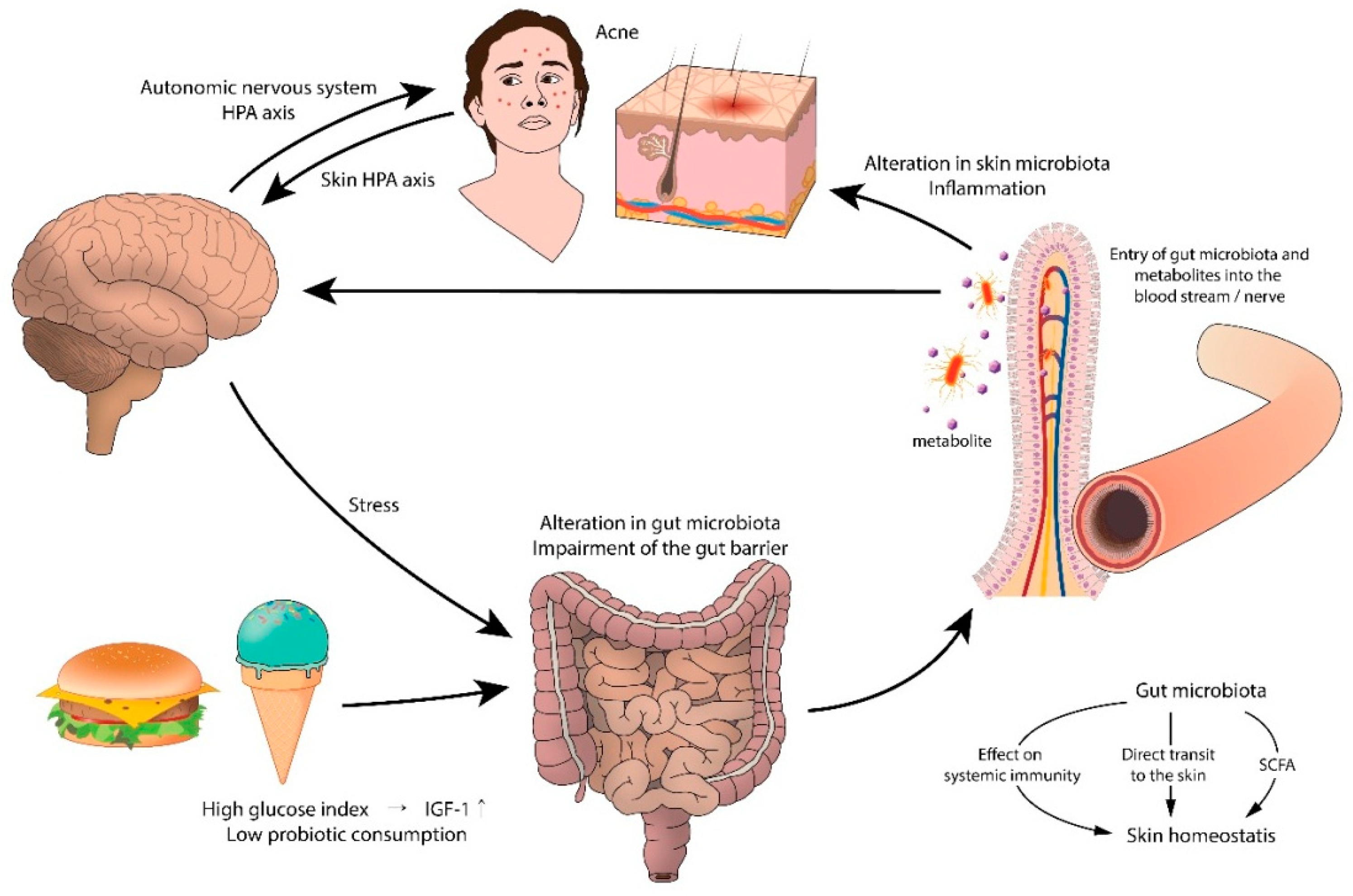
6. Gut Microbiota and the Skin
6.1. Gut Microbiota and Acne
6.2. Gut Microbiota in Acne
7. Probiotics and the Skin
Probiotics and Acne
8. Conclusions
Author Contributions
Funding
Acknowledgements
Conflicts of Interest
References
- Salvucci, E. Microbiome, holobiont and the net of life. Crit. Rev. Microbiol. 2016, 42, 485–494. [Google Scholar] [CrossRef] [PubMed]
- Hooper, L.V.; Littman, D.R.; Macpherson, A.J. Interactions between the microbiota and the immune system. Science 2012, 336, 1268–1273. [Google Scholar] [CrossRef] [PubMed]
- Adebamowo, C.A.; Spiegelman, D.; Berkey, C.S.; Danby, F.W.; Rockett, H.H.; Colditz, G.A.; Willett, W.C.; Holmes, M.D. Milk consumption and acne in teenaged boys. J. Am. Acad. Dermatol. 2008, 58, 787–793. [Google Scholar] [CrossRef] [PubMed]
- Adebamowo, C.A.; Spiegelman, D.; Berkey, C.S.; Danby, F.W.; Rockett, H.H.; Colditz, G.A.; Willett, W.C.; Holmes, M.D. Milk consumption and acne in adolescent girls. Dermatol. Online J. 2006, 12, 1. [Google Scholar] [PubMed]
- Agamia, N.F.; Abdallah, D.M.; Sorour, O.; Mourad, B.; Younan, D.N. Skin expression of mammalian target of rapamycin and forkhead box transcription factor O1, and serum insulin-like growth factor-1 in patients with acne vulgaris and their relationship with diet. Br. J. Dermatol. 2016, 174, 1299–1307. [Google Scholar] [CrossRef]
- Kim, H.; Moon, S.Y.; Sohn, M.Y.; Lee, W.J. Insulin-Like Growth Factor-1 Increases the Expression of Inflammatory Biomarkers and Sebum Production in Cultured Sebocytes. Ann. Dermatol. 2017, 29, 20–25. [Google Scholar] [CrossRef] [PubMed]
- Bowe, W.; Patel, N.B.; Logan, A.C. Acne vulgaris, probiotics and the gut-brain-skin axis: From anecdote to translational medicine. Benef. Microbes 2014, 5, 185–199. [Google Scholar] [CrossRef]
- Pasparakis, M.; Haase, I.; Nestle, F.O. Mechanisms regulating skin immunity and inflammation. Nat. Rev. Immunol. 2014, 14, 289–301. [Google Scholar] [CrossRef]
- Naik, S.; Bouladoux, N.; Wilhelm, C.; Molloy, M.J.; Salcedo, R.; Kastenmuller, W.; Deming, C.; Quinones, M.; Koo, L.; Conlan, S.; et al. Compartmentalized control of skin immunity by resident commensals. Science 2012, 337, 1115–1119. [Google Scholar] [CrossRef]
- Belkaid, Y.; Segre, J.A. Dialogue between skin microbiota and immunity. Science 2014, 346, 954–959. [Google Scholar] [CrossRef]
- Palm, N.W.; de Zoete, M.R.; Flavell, R.A. Immune-microbiota interactions in health and disease. Clin. Immunol. 2015, 159, 122–127. [Google Scholar] [CrossRef] [PubMed]
- Grice, E.A.; Kong, H.H.; Renaud, G.; Young, A.C.; Bouffard, G.G.; Blakesley, R.W.; Wolfsberg, T.G.; Turner, M.L.; Segre, J.A. A diversity profile of the human skin microbiota. Genome Res. 2008, 18, 1043–1050. [Google Scholar] [CrossRef] [PubMed]
- Hall, J.B.; Cong, Z.; Imamura-Kawasawa, Y.; Kidd, B.A.; Dudley, J.T.; Thiboutot, D.M.; Nelson, A.M. Isolation and Identification of the Follicular Microbiome: Implications for Acne Research. J. Investig. Dermatol. 2018, 138, 2033–2040. [Google Scholar] [CrossRef] [PubMed]
- Prast-Nielsen, S.; Tobin, A.M.; Adamzik, K.; Powles, A.; Hugerth, L.; Sweeney, C.; Kirby, B.; Engstrand, L.; Fry, L. Investigation of the Skin Microbiome: Swabs vs Biopsies. Br. J. Dermatol. 2019. [Google Scholar] [CrossRef] [PubMed]
- Harris, J.K.; Wagner, B.D. Bacterial identification and analytic challenges in clinical microbiome studies. J. Allergy Clin. Immunol. 2012, 129, 441–442. [Google Scholar] [CrossRef] [PubMed]
- Grogan, M.D.; Bartow-McKenney, C.; Flowers, L.; Knight, S.A.B.; Uberoi, A.; Grice, E.A. Research Techniques Made Simple: Profiling the Skin Microbiota. J. Investig. Dermatol. 2019, 139, 747–752. [Google Scholar] [CrossRef] [PubMed]
- Hamady, M.; Knight, R. Microbial community profiling for human microbiome projects: Tools, techniques, and challenges. Genome Res. 2009, 19, 1141–1152. [Google Scholar] [CrossRef]
- Dreno, B.; Martin, R.; Moyal, D.; Henley, J.B.; Khammari, A.; Seite, S. Skin microbiome and acne vulgaris: Staphylococcus, a new actor in acne. Exp. Derm. 2017, 26, 798–803. [Google Scholar] [CrossRef]
- Leyden, J.J.; McGinley, K.J.; Mills, O.H.; Kligman, A.M. Age-related changes in the resident bacterial flora of the human face. J. Investig. Dermatol. 1975, 65, 379–381. [Google Scholar] [CrossRef]
- Marples, R.R. Sex, constancy, and skin bacteria. Arch. Dermatol. Res. 1982, 272, 317–320. [Google Scholar] [CrossRef]
- Giacomoni, P.U.; Mammone, T.; Teri, M. Gender-linked differences in human skin. J. Derm. Sci. 2009, 55, 144–149. [Google Scholar] [CrossRef] [PubMed]
- Faergemann, J.; Larko, O. The effect of UV-light on human skin microorganisms. Acta Derm. Venereol. 1987, 67, 69–72. [Google Scholar]
- Fierer, N.; Hamady, M.; Lauber, C.L.; Knight, R. The influence of sex, handedness, and washing on the diversity of hand surface bacteria. Proc. Natl. Acad. Sci. USA 2008, 105, 17994–17999. [Google Scholar] [CrossRef] [PubMed]
- McBride, M.E.; Duncan, W.C.; Knox, J.M. The environment and the microbial ecology of human skin. Appl. Environ. Microbiol. 1977, 33, 603–608. [Google Scholar]
- Grice, E.A.; Segre, J.A. The skin microbiome. Nat. Rev. Microbiol. 2011, 9, 244–253. [Google Scholar] [CrossRef] [PubMed]
- Capone, K.A.; Dowd, S.E.; Stamatas, G.N.; Nikolovski, J. Diversity of the human skin microbiome early in life. J. Investig. Dermatol. 2011, 131, 2026–2032. [Google Scholar] [CrossRef] [PubMed]
- Human Microbiome Project Consortium. Structure, function and diversity of the healthy human microbiome. Nature 2012, 486, 207–214. [Google Scholar] [CrossRef]
- Staudinger, T.; Pipal, A.; Redl, B. Molecular analysis of the prevalent microbiota of human male and female forehead skin compared to forearm skin and the influence of make-up. J. Appl. Microbiol. 2011, 110, 1381–1389. [Google Scholar] [CrossRef]
- Xu, H.; Li, H. Acne, the Skin Microbiome, and Antibiotic Treatment. Am. J. Clin. Dermatol. 2019, 20, 335–344. [Google Scholar] [CrossRef]
- Grice, E.A.; Kong, H.H.; Conlan, S.; Deming, C.B.; Davis, J.; Young, A.C.; Program, N.C.S.; Bouffard, G.G.; Blakesley, R.W.; Murray, P.R.; et al. Topographical and temporal diversity of the human skin microbiome. Science 2009, 324, 1190–1192. [Google Scholar] [CrossRef]
- Patel, A.; Calfee, R.P.; Plante, M.; Fischer, S.A.; Green, A. Propionibacterium acnes colonization of the human shoulder. J. Shoulder Elb. Surg. 2009, 18, 897–902. [Google Scholar] [CrossRef] [PubMed]
- Christensen, G.J.; Bruggemann, H. Bacterial skin commensals and their role as host guardians. Benef. Microbes 2014, 5, 201–215. [Google Scholar] [CrossRef] [PubMed]
- Fitz-Gibbon, S.; Tomida, S.; Chiu, B.H.; Nguyen, L.; Du, C.; Liu, M.; Elashoff, D.; Erfe, M.C.; Loncaric, A.; Kim, J.; et al. Propionibacterium acnes strain populations in the human skin microbiome associated with acne. J. Investig. Dermatol. 2013, 133, 2152–2160. [Google Scholar] [CrossRef] [PubMed]
- Barnard, E.; Shi, B.; Kang, D.; Craft, N.; Li, H. The balance of metagenomic elements shapes the skin microbiome in acne and health. Sci. Rep. 2016, 6, 39491. [Google Scholar] [CrossRef] [PubMed]
- Johnson, J.L.; Cummins, C.S. Cell wall composition and deoxyribonucleic acid similarities among the anaerobic coryneforms, classical propionibacteria, and strains of Arachnia propionica. J. Bacteriol. 1972, 109, 1047–1066. [Google Scholar] [PubMed]
- McDowell, A.; Perry, A.L.; Lambert, P.A.; Patrick, S. A new phylogenetic group of Propionibacterium acnes. J. Med. Microbiol. 2008, 57, 218–224. [Google Scholar] [CrossRef] [PubMed]
- Scholz, C.F.; Kilian, M. The natural history of cutaneous propionibacteria, and reclassification of selected species within the genus Propionibacterium to the proposed novel genera Acidipropionibacterium gen. nov., Cutibacterium gen. nov. and Pseudopropionibacterium gen. nov. Int. J. Syst. Evol. Microbiol. 2016, 66, 4422–4432. [Google Scholar] [CrossRef] [PubMed]
- McDowell, A.; Barnard, E.; Liu, J.; Li, H.; Patrick, S. Corrigendum: Proposal to reclassify Propionibacterium acnes type I as Propionibacterium acnes subsp. acnes subsp. nov. and Propionibacterium acnes type II as Propionibacterium acnes subsp. defendens subsp. nov. Int. J. Syst. Evol. Microbiol. 2017, 67, 4880. [Google Scholar] [CrossRef] [PubMed][Green Version]
- McDowell, A.; Barnard, E.; Liu, J.; Li, H.; Patrick, S. Emendation of Propionibacterium acnes subsp. acnes (Deiko et al. 2015) and proposal of Propionibacterium acnes type II as Propionibacterium acnes subsp. defendens subsp. nov. Int. J. Syst. Evol. Microbiol. 2016, 66, 5358–5365. [Google Scholar] [CrossRef]
- Dekio, I.; Culak, R.; Misra, R.; Gaulton, T.; Fang, M.; Sakamoto, M.; Ohkuma, M.; Oshima, K.; Hattori, M.; Klenk, H.P.; et al. Dissecting the taxonomic heterogeneity within Propionibacterium acnes: Proposal for Propionibacterium acnes subsp. acnes subsp. nov. and Propionibacterium acnes subsp. elongatum subsp. nov. Int. J. Syst. Evol. Microbiol. 2015, 65, 4776–4787. [Google Scholar] [CrossRef]
- McDowell, A.; Barnard, E.; Nagy, I.; Gao, A.; Tomida, S.; Li, H.; Eady, A.; Cove, J.; Nord, C.E.; Patrick, S. An expanded multilocus sequence typing scheme for propionibacterium acnes: Investigation of ‘pathogenic’, ‘commensal’ and antibiotic resistant strains. PLoS ONE 2012, 7, e41480. [Google Scholar] [CrossRef] [PubMed]
- Lomholt, H.B.; Kilian, M. Population genetic analysis of Propionibacterium acnes identifies a subpopulation and epidemic clones associated with acne. PLoS ONE 2010, 5, e12277. [Google Scholar] [CrossRef] [PubMed]
- Tomida, S.; Nguyen, L.; Chiu, B.H.; Liu, J.; Sodergren, E.; Weinstock, G.M.; Li, H. Pan-genome and comparative genome analyses of propionibacterium acnes reveal its genomic diversity in the healthy and diseased human skin microbiome. mBio 2013, 4, e00003–00013. [Google Scholar] [CrossRef] [PubMed]
- McDowell, A.; Nagy, I.; Magyari, M.; Barnard, E.; Patrick, S. The opportunistic pathogen Propionibacterium acnes: Insights into typing, human disease, clonal diversification and CAMP factor evolution. PLoS ONE 2013, 8, e70897. [Google Scholar] [CrossRef] [PubMed]
- Barnard, E.; Liu, J.; Yankova, E.; Cavalcanti, S.M.; Magalhaes, M.; Li, H.; Patrick, S.; McDowell, A. Strains of the Propionibacterium acnes type III lineage are associated with the skin condition progressive macular hypomelanosis. Sci. Rep. 2016, 6, 31968. [Google Scholar] [CrossRef] [PubMed]
- Petersen, R.L.; Scholz, C.F.; Jensen, A.; Bruggemann, H.; Lomholt, H.B. Propionibacterium Acnes Phylogenetic Type III is Associated with Progressive Macular Hypomelanosis. Eur. J. Microbiol. Immunol. 2017, 7, 37–45. [Google Scholar] [CrossRef] [PubMed]
- Johnson, T.; Kang, D.; Barnard, E.; Li, H. Strain-Level Differences in Porphyrin Production and Regulation in Propionibacterium acnes Elucidate Disease Associations. mSphere 2016, 1. [Google Scholar] [CrossRef]
- Schaller, M.; Loewenstein, M.; Borelli, C.; Jacob, K.; Vogeser, M.; Burgdorf, W.H.; Plewig, G. Induction of a chemoattractive proinflammatory cytokine response after stimulation of keratinocytes with Propionibacterium acnes and coproporphyrin III. Br. J. Dermatol. 2005, 153, 66–71. [Google Scholar] [CrossRef]
- Kang, D.; Shi, B.; Erfe, M.C.; Craft, N.; Li, H. Vitamin B12 modulates the transcriptome of the skin microbiota in acne pathogenesis. Sci. Transl. Med. 2015, 7, 293ra103. [Google Scholar] [CrossRef]
- Nagy, I.; Pivarcsi, A.; Koreck, A.; Szell, M.; Urban, E.; Kemeny, L. Distinct strains of Propionibacterium acnes induce selective human beta-defensin-2 and interleukin-8 expression in human keratinocytes through toll-like receptors. J. Investig. Dermatol. 2005, 124, 931–938. [Google Scholar] [CrossRef]
- Nagy, I.; Pivarcsi, A.; Kis, K.; Koreck, A.; Bodai, L.; McDowell, A.; Seltmann, H.; Patrick, S.; Zouboulis, C.C.; Kemeny, L. Propionibacterium acnes and lipopolysaccharide induce the expression of antimicrobial peptides and proinflammatory cytokines/chemokines in human sebocytes. Microbes Infect. 2006, 8, 2195–2205. [Google Scholar] [CrossRef] [PubMed]
- Agak, G.W.; Kao, S.; Ouyang, K.; Qin, M.; Moon, D.; Butt, A.; Kim, J. Phenotype and Antimicrobial Activity of Th17 Cells Induced by Propionibacterium acnes Strains Associated with Healthy and Acne Skin. J. Investig. Dermatol. 2018, 138, 316–324. [Google Scholar] [CrossRef] [PubMed]
- Yu, Y.; Champer, J.; Agak, G.W.; Kao, S.; Modlin, R.L.; Kim, J. Different Propionibacterium acnes Phylotypes Induce Distinct Immune Responses and Express Unique Surface and Secreted Proteomes. J. Investig. Dermatol. 2016, 136, 2221–2228. [Google Scholar] [CrossRef] [PubMed]
- Iinuma, K.; Sato, T.; Akimoto, N.; Noguchi, N.; Sasatsu, M.; Nishijima, S.; Kurokawa, I.; Ito, A. Involvement of Propionibacterium acnes in the augmentation of lipogenesis in hamster sebaceous glands in vivo and in vitro. J. Investig. Dermatol. 2009, 129, 2113–2119. [Google Scholar] [CrossRef] [PubMed]
- Saint-Leger, D.; Bague, A.; Cohen, E.; Chivot, M. A possible role for squalene in the pathogenesis of acne. I. In vitro study of squalene oxidation. Br. J. Dermatol. 1986, 114, 535–542. [Google Scholar] [CrossRef] [PubMed]
- Saint-Leger, D.; Bague, A.; Lefebvre, E.; Cohen, E.; Chivot, M. A possible role for squalene in the pathogenesis of acne. II. In vivo study of squalene oxides in skin surface and intra-comedonal lipids of acne patients. Br. J. Dermatol. 1986, 114, 543–552. [Google Scholar] [CrossRef] [PubMed]
- Jarrousse, V.; Castex-Rizzi, N.; Khammari, A.; Charveron, M.; Dreno, B. Modulation of integrins and filaggrin expression by Propionibacterium acnes extracts on keratinocytes. Arch. Dermatol. Res. 2007, 299, 441–447. [Google Scholar] [CrossRef] [PubMed]
- Isard, O.; Knol, A.C.; Aries, M.F.; Nguyen, J.M.; Khammari, A.; Castex-Rizzi, N.; Dreno, B. Propionibacterium acnes activates the IGF-1/IGF-1R system in the epidermis and induces keratinocyte proliferation. J. Investig. Dermatol. 2011, 131, 59–66. [Google Scholar] [CrossRef] [PubMed]
- Graham, G.M.; Farrar, M.D.; Cruse-Sawyer, J.E.; Holland, K.T.; Ingham, E. Proinflammatory cytokine production by human keratinocytes stimulated with Propionibacterium acnes and P. acnes GroEL. Br. J. Dermatol. 2004, 150, 421–428. [Google Scholar] [CrossRef]
- Jugeau, S.; Tenaud, I.; Knol, A.C.; Jarrousse, V.; Quereux, G.; Khammari, A.; Dreno, B. Induction of toll-like receptors by Propionibacterium acnes. Br. J. Dermatol. 2005, 153, 1105–1113. [Google Scholar] [CrossRef]
- Schmuth, M.; Ortegon, A.M.; Mao-Qiang, M.; Elias, P.M.; Feingold, K.R.; Stahl, A. Differential expression of fatty acid transport proteins in epidermis and skin appendages. J. Investig. Dermatol. 2005, 125, 1174–1181. [Google Scholar] [CrossRef] [PubMed]
- Grange, P.A.; Chereau, C.; Raingeaud, J.; Nicco, C.; Weill, B.; Dupin, N.; Batteux, F. Production of superoxide anions by keratinocytes initiates P. acnes-induced inflammation of the skin. PLoS Pathog. 2009, 5, e1000527. [Google Scholar] [CrossRef] [PubMed]
- Cong, T.X.; Hao, D.; Wen, X.; Li, X.H.; He, G.; Jiang, X. From pathogenesis of acne vulgaris to anti-acne agents. Arch. Dermatol. Res. 2019, 311, 337–349. [Google Scholar] [CrossRef] [PubMed]
- Qin, M.; Pirouz, A.; Kim, M.H.; Krutzik, S.R.; Garban, H.J.; Kim, J. Propionibacterium acnes Induces IL-1beta secretion via the NLRP3 inflammasome in human monocytes. J. Investig. Dermatol. 2014, 134, 381–388. [Google Scholar] [CrossRef] [PubMed]
- Jappe, U.; Ingham, E.; Henwood, J.; Holland, K.T. Propionibacterium acnes and inflammation in acne; P. acnes has T-cell mitogenic activity. Br. J. Dermatol. 2002, 146, 202–209. [Google Scholar] [CrossRef]
- Mouser, P.E.; Baker, B.S.; Seaton, E.D.; Chu, A.C. Propionibacterium acnes-reactive T helper-1 cells in the skin of patients with acne vulgaris. J. Investig. Dermatol. 2003, 121, 1226–1228. [Google Scholar] [CrossRef]
- Agak, G.W.; Qin, M.; Nobe, J.; Kim, M.H.; Krutzik, S.R.; Tristan, G.R.; Elashoff, D.; Garban, H.J.; Kim, J. Propionibacterium acnes Induces an IL-17 Response in Acne Vulgaris that Is Regulated by Vitamin A and Vitamin D. J. Investig. Dermatol. 2014, 134, 366–373. [Google Scholar] [CrossRef] [PubMed]
- Kistowska, M.; Meier, B.; Proust, T.; Feldmeyer, L.; Cozzio, A.; Kuendig, T.; Contassot, E.; French, L.E. Propionibacterium acnes promotes Th17 and Th17/Th1 responses in acne patients. J. Investig. Dermatol. 2015, 135, 110–118. [Google Scholar] [CrossRef]
- Ross, J.I.; Snelling, A.M.; Eady, E.A.; Cove, J.H.; Cunliffe, W.J.; Leyden, J.J.; Collignon, P.; Dreno, B.; Reynaud, A.; Fluhr, J.; et al. Phenotypic and genotypic characterization of antibiotic-resistant Propionibacterium acnes isolated from acne patients attending dermatology clinics in Europe, the U.S.A., Japan and Australia. Br. J. Dermatol. 2001, 144, 339–346. [Google Scholar] [CrossRef]
- Higaki, S.; Kitagawa, T.; Kagoura, M.; Morohashi, M.; Yamagishi, T. Correlation between Propionibacterium acnes biotypes, lipase activity and rash degree in acne patients. J. Dermatol. 2000, 27, 519–522. [Google Scholar] [CrossRef]
- Katsuta, Y.; Iida, T.; Hasegawa, K.; Inomata, S.; Denda, M. Function of oleic acid on epidermal barrier and calcium influx into keratinocytes is associated with N-methyl D-aspartate-type glutamate receptors. Br. J. Dermatol. 2009, 160, 69–74. [Google Scholar] [CrossRef] [PubMed]
- Lee, W.L.; Shalita, A.R.; Suntharalingam, K.; Fikrig, S.M. Neutrophil chemotaxis by Propionibacterium acnes lipase and its inhibition. Infect. Immun. 1982, 35, 71–78. [Google Scholar] [PubMed]
- Holland, C.; Mak, T.N.; Zimny-Arndt, U.; Schmid, M.; Meyer, T.F.; Jungblut, P.R.; Bruggemann, H. Proteomic identification of secreted proteins of Propionibacterium acnes. BMC Microbiol. 2010, 10, 230. [Google Scholar] [CrossRef] [PubMed]
- Makris, G.; Wright, J.D.; Ingham, E.; Holland, K.T. The hyaluronate lyase of Staphylococcus aureus—A virulence factor? Microbiology (Read. Engl.) 2004, 150, 2005–2013. [Google Scholar] [CrossRef] [PubMed]
- Steiner, B.; Romero-Steiner, S.; Cruce, D.; George, R. Cloning and sequencing of the hyaluronate lyase gene from Propionibacterium acnes. Can. J. Microbiol. 1997, 43, 315–321. [Google Scholar] [CrossRef] [PubMed]
- Rocha, M.A.; Bagatin, E. Skin barrier and microbiome in acne. Arch. Dermatol. Res. 2018, 310, 181–185. [Google Scholar] [CrossRef]
- Kasimatis, G.; Fitz-Gibbon, S.; Tomida, S.; Wong, M.; Li, H. Analysis of complete genomes of Propionibacterium acnes reveals a novel plasmid and increased pseudogenes in an acne associated strain. Biomed Res. Int. 2013, 2013, 11. [Google Scholar] [CrossRef]
- Jahns, A.C.; Lundskog, B.; Ganceviciene, R.; Palmer, R.H.; Golovleva, I.; Zouboulis, C.C.; McDowell, A.; Patrick, S.; Alexeyev, O.A. An increased incidence of Propionibacterium acnes biofilms in acne vulgaris: A case-control study. Br. J. Dermatol. 2012, 167, 50–58. [Google Scholar] [CrossRef]
- Donlan, R.M.; Costerton, J.W. Biofilms: Survival mechanisms of clinically relevant microorganisms. Clin. Microbiol. Rev. 2002, 15, 167–193. [Google Scholar] [CrossRef]
- Donlan, R.M. Biofilm formation: A clinically relevant microbiological process. Clin. Infect. Dis. Off. Publ. Infect. Dis. Soc. Am. 2001, 33, 1387–1392. [Google Scholar] [CrossRef]
- Rajiv, P.; Nitesh, K.; Raj, K.; Hemant, G. Staphylococcus epidermidis in Human Skin Microbiome associated with Acne: A Cause of Disease or Defence? Res. J. Biotechnol. 2013, 8, 78–82. [Google Scholar]
- Iwase, T.; Uehara, Y.; Shinji, H.; Tajima, A.; Seo, H.; Takada, K.; Agata, T.; Mizunoe, Y. Staphylococcus epidermidis Esp inhibits Staphylococcus aureus biofilm formation and nasal colonization. Nature 2010, 465, 346–349. [Google Scholar] [CrossRef] [PubMed]
- Otto, M.; Echner, H.; Voelter, W.; Gotz, F. Pheromone cross-inhibition between Staphylococcus aureus and Staphylococcus epidermidis. Infect. Immun. 2001, 69, 1957–1960. [Google Scholar] [CrossRef] [PubMed]
- Bierbaum, G.; Gotz, F.; Peschel, A.; Kupke, T.; van de Kamp, M.; Sahl, H.G. The biosynthesis of the lantibiotics epidermin, gallidermin, Pep5 and epilancin K7. Antonie Van Leeuwenhoek 1996, 69, 119–127. [Google Scholar] [CrossRef] [PubMed]
- Cogen, A.L.; Nizet, V.; Gallo, R.L. Skin microbiota: A source of disease or defence? Br. J. Dermatol. 2008, 158, 442–455. [Google Scholar] [CrossRef] [PubMed]
- Ekkelenkamp, M.B.; Hanssen, M.; Danny Hsu, S.T.; de Jong, A.; Milatovic, D.; Verhoef, J.; van Nuland, N.A. Isolation and structural characterization of epilancin 15X, a novel lantibiotic from a clinical strain of Staphylococcus epidermidis. FEBS Lett. 2005, 579, 1917–1922. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Dai, A.; Huang, S.; Kuo, S.; Shu, M.; Tapia, C.P.; Yu, J.; Two, A.; Zhang, H.; Gallo, R.L.; et al. Propionic acid and its esterified derivative suppress the growth of methicillin-resistant Staphylococcus aureus USA300. Benef. Microbes 2014, 5, 161–168. [Google Scholar] [CrossRef]
- Christensen, G.J.; Scholz, C.F.; Enghild, J.; Rohde, H.; Kilian, M.; Thurmer, A.; Brzuszkiewicz, E.; Lomholt, H.B.; Bruggemann, H. Antagonism between Staphylococcus epidermidis and Propionibacterium acnes and its genomic basis. BMC Genom. 2016, 17, 152. [Google Scholar] [CrossRef]
- Xia, X.; Li, Z.; Liu, K.; Wu, Y.; Jiang, D.; Lai, Y. Staphylococcal LTA-Induced miR-143 Inhibits Propionibacterium acnes-Mediated Inflammatory Response in Skin. J. Investig. Dermatol. 2016, 136, 621–630. [Google Scholar] [CrossRef]
- Gehse, M.; Hoffler, U.; Gloor, M.; Pulverer, G. Propionibacteria in patients with acne vulgaris and in healthy persons. Arch. Dermatol. Res. 1983, 275, 100–104. [Google Scholar] [CrossRef]
- Whiteside, J.A.; Voss, J.G. Incidence and lipolytic activity of Propionibacterium acnes (Corynebacterium acnes group I) and P. granulosum (C. acnes group II) in acne and in normal skin. J. Investig. Dermatol. 1973, 60, 94–97. [Google Scholar] [CrossRef] [PubMed][Green Version]
- O’Neill, C.A.; Monteleone, G.; McLaughlin, J.T.; Paus, R. The gut-skin axis in health and disease: A paradigm with therapeutic implications. Bioessays News Rev. Mol. Cell. Dev. Biol. 2016, 38, 1167–1176. [Google Scholar] [CrossRef] [PubMed]
- Mak, T.N.; Schmid, M.; Brzuszkiewicz, E.; Zeng, G.; Meyer, R.; Sfanos, K.S.; Brinkmann, V.; Meyer, T.F.; Bruggemann, H. Comparative genomics reveals distinct host-interacting traits of three major human-associated propionibacteria. BMC Genom. 2013, 14, 640. [Google Scholar] [CrossRef] [PubMed]
- Akaza, N.; Akamatsu, H.; Numata, S.; Yamada, S.; Yagami, A.; Nakata, S.; Matsunaga, K. Microorganisms inhabiting follicular contents of facial acne are not only Propionibacterium but also Malassezia spp. J. Dermatol. 2016, 43, 906–911. [Google Scholar] [CrossRef] [PubMed]
- Gang, H.; Wei, Y.-p.; Jie, F. Malasseziainfection: Is there any chance or necessity in refractory acne? Chin. Med. J. 2010, 123, 628–632. [Google Scholar]
- Song, Y.C.; Hahn, H.J.; Kim, J.Y.; Ko, J.H.; Lee, Y.W.; Choe, Y.B.; Ahn, K.J. Epidemiologic Study of Malassezia Yeasts in Acne Patients by Analysis of 26S rDNA PCR-RFLP. Ann. Dermatol. 2011, 23, 321–328. [Google Scholar] [CrossRef] [PubMed]
- Numata, S.; Akamatsu, H.; Akaza, N.; Yagami, A.; Nakata, S.; Matsunaga, K. Analysis of facial skin-resident microbiota in Japanese acne patients. Dermatology (Baselswitz.) 2014, 228, 86–92. [Google Scholar] [CrossRef] [PubMed]
- Akaza, N.; Akamatsu, H.; Takeoka, S.; Sasaki, Y.; Mizutani, H.; Nakata, S.; Matsunaga, K. Malassezia globosa tends to grow actively in summer conditions more than other cutaneous Malassezia species. J. Dermatol. 2012, 39, 613–616. [Google Scholar] [CrossRef]
- Katsuta, Y.; Iida, T.; Inomata, S.; Denda, M. Unsaturated fatty acids induce calcium influx into keratinocytes and cause abnormal differentiation of epidermis. J. Investig. Dermatol. 2005, 124, 1008–1013. [Google Scholar] [CrossRef]
- Kesavan, S.; Walters, C.E.; Holland, K.T.; Ingham, E. The effects of Malassezia on pro-inflammatory cytokine production by human peripheral blood mononuclear cells in vitro. Med. Mycol. 1998, 36, 97–106. [Google Scholar] [CrossRef]
- Akaza, N.; Akamatsu, H.; Takeoka, S.; Mizutani, H.; Nakata, S.; Matsunaga, K. Increased hydrophobicity in Malassezia species correlates with increased proinflammatory cytokine expression in human keratinocytes. Med. Mycol. 2012, 50, 802–810. [Google Scholar] [CrossRef] [PubMed]
- Zaenglein, A.L.; Pathy, A.L.; Schlosser, B.J.; Alikhan, A.; Baldwin, H.E.; Berson, D.S.; Bowe, W.P.; Graber, E.M.; Harper, J.C.; Kang, S.; et al. Guidelines of care for the management of acne vulgaris. J. Am. Acad. Dermatol. 2016, 74, 945–973. [Google Scholar] [CrossRef] [PubMed]
- Dreno, B. Topical antibacterial therapy for acne vulgaris. Drugs 2004, 64, 2389–2397. [Google Scholar] [CrossRef] [PubMed]
- Eichenfield, L.F.; Krakowski, A.C.; Piggott, C.; Del Rosso, J.; Baldwin, H.; Friedlander, S.F.; Levy, M.; Lucky, A.; Mancini, A.J.; Orlow, S.J.; et al. Evidence-based recommendations for the diagnosis and treatment of pediatric acne. Pediatrics 2013, 131, S163–186. [Google Scholar] [CrossRef] [PubMed]
- Sardana, K.G.T.; Kumar, B.; Gautam, H.K.; Goh, C.L.; Abad-Casintahan, F.; Aw, D.C.; Baba, R.; Chan, L.C.; Hung, N.T.; Kulthanan, K.; et al. South-East Asia study alliance guidelines on the management of acne vulgaris in South-East Asian patients. J. Dermatol. 2015, 42, 945–953. [Google Scholar] [CrossRef]
- Gollnick, H.P.; Bettoli, V.; Lambert, J.; Araviiskaia, E.; Binic, I.; Dessinioti, C.; Galadari, I.; Ganceviciene, R.; Ilter, N.; Kaegi, M.; et al. A consensus-based practical and daily guide for the treatment of acne patients. J. Eur. Acad. Dermatol. Venereol. JEADV 2016, 30, 1480–1490. [Google Scholar] [CrossRef] [PubMed]
- Thiboutot, D.M.; Dreno, B.; Abanmi, A.; Alexis, A.F.; Araviiskaia, E.; Barona Cabal, M.I.; Bettoli, V.; Casintahan, F.; Chow, S.; da Costa, A.; et al. Practical management of acne for clinicians: An international consensus from the Global Alliance to Improve Outcomes in Acne. J. Am. Acad. Dermatol. 2018, 78, S1–S23. [Google Scholar] [CrossRef] [PubMed]
- Kircik, L.H. The role of benzoyl peroxide in the new treatment paradigm for acne. J. Drugs Dermatol. JDD 2013, 12, s73–s76. [Google Scholar]
- Bojar, R.A.; Cunliffe, W.J.; Holland, K.T. Disruption of the transmembrane pH gradient--a possible mechanism for the antibacterial action of azelaic acid in Propionibacterium acnes and Staphylococcus epidermidis. J. Antimicrob. Chemother. 1994, 34, 321–330. [Google Scholar] [CrossRef]
- Layton, A. The use of isotretinoin in acne. Derm. Endocrinol. 2009, 1, 162–169. [Google Scholar] [CrossRef]
- Dispenza, M.C.; Wolpert, E.B.; Gilliland, K.L.; Dai, J.P.; Cong, Z.; Nelson, A.M.; Thiboutot, D.M. Systemic isotretinoin therapy normalizes exaggerated TLR-2-mediated innate immune responses in acne patients. J. Investig. Dermatol. 2012, 132, 2198–2205. [Google Scholar] [CrossRef] [PubMed]
- Kelhala, H.L.; Aho, V.T.E.; Fyhrquist, N.; Pereira, P.A.B.; Kubin, M.E.; Paulin, L.; Palatsi, R.; Auvinen, P.; Tasanen, K.; Lauerma, A. Isotretinoin and lymecycline treatments modify the skin microbiota in acne. Exp. Dermatol. 2018, 27, 30–36. [Google Scholar] [CrossRef] [PubMed]
- Ryan-Kewley, A.E.; Williams, D.R.; Hepburn, N.; Dixon, R.A. Non-antibiotic Isotretinoin Treatment Differentially Controls Propionibacterium acnes on Skin of Acne Patients. Front. Microbiol. 2017, 8, 1381. [Google Scholar] [CrossRef] [PubMed]
- Oprica, C.; Emtestam, L.; Hagstromer, L.; Nord, C.E. Clinical and microbiological comparisons of isotretinoin vs. tetracycline in acne vulgaris. Acta Derm. Venereol. 2007, 87, 246–254. [Google Scholar] [CrossRef] [PubMed]
- Coughlin, C.C.; Swink, S.M.; Horwinski, J.; Sfyroera, G.; Bugayev, J.; Grice, E.A.; Yan, A.C. The preadolescent acne microbiome: A prospective, randomized, pilot study investigating characterization and effects of acne therapy. Pediatr. Dermatol. 2017, 34, 661–664. [Google Scholar] [CrossRef] [PubMed]
- McCoy, W.H.; Otchere, E.; Rosa, B.A.; Martin, J.; Mann, C.M.; Mitreva, M. Skin Ecology during Sebaceous Drought-How Skin Microbes Respond to Isotretinoin. J. Investig. Dermatol. 2019, 139, 732–735. [Google Scholar] [CrossRef] [PubMed]
- Charakida, A.; Seaton, E.D.; Charakida, M.; Mouser, P.; Avgerinos, A.; Chu, A.C. Phototherapy in the treatment of acne vulgaris: What is its role? Am. J. Clin. Dermatol. 2004, 5, 211–216. [Google Scholar] [CrossRef]
- Rassai, S.; Rafeie, E.; Ramirez-Fort, M.K.; Feily, A. Adjuvant Narrow Band UVB Improves the Efficacy of Oral Azithromycin for the Treatment of Moderate to Severe Inflammatory Facial Acne Vulgaris. J. Cutan. Aesthet. Surg. 2014, 7, 151–154. [Google Scholar] [CrossRef]
- de Annunzio, S.R.; de Freitas, L.M.; Blanco, A.L.; da Costa, M.M.; Carmona-Vargas, C.C.; de Oliveira, K.T.; Fontana, C.R. Susceptibility of Enterococcus faecalis and Propionibacterium acnes to antimicrobial photodynamic therapy. J. Photochem. Photobiol. B Biol. 2018, 178, 545–550. [Google Scholar] [CrossRef]
- Eady, E.A.; Cove, J.H.; Holland, K.T.; Cunliffe, W.J. Superior antibacterial action and reduced incidence of bacterial resistance in minocycline compared to tetracycline-treated acne patients. Br. J. Dermatol. 1990, 122, 233–244. [Google Scholar] [CrossRef]
- Chien, A.L.; Tsai, J.; Leung, S.; Mongodin, E.F.; Nelson, A.M.; Kang, S.; Garza, L.A. Association of Systemic Antibiotic Treatment of Acne With Skin Microbiota Characteristics. JAMA Dermatol. 2019. [Google Scholar] [CrossRef] [PubMed]
- Walsh, T.R.; Efthimiou, J.; Dreno, B. Systematic review of antibiotic resistance in acne: An increasing topical and oral threat. Lancet Infect. Dis. 2016, 16, e23–33. [Google Scholar] [CrossRef]
- Sardana, K.; Gupta, T.; Kumar, B.; Gautam, H.K.; Garg, V.K. Cross-sectional Pilot Study of Antibiotic Resistance in Propionibacterium Acnes Strains in Indian Acne Patients Using 16S-RNA Polymerase Chain Reaction: A Comparison Among Treatment Modalities Including Antibiotics, Benzoyl Peroxide, and Isotretinoin. Indian J. Dermatol. 2016, 61, 45–52. [Google Scholar] [CrossRef] [PubMed]
- Gonzalez, R.; Welsh, O.; Ocampo, J.; Hinojosa-Robles, R.M.; Vera-Cabrera, L.; Delaney, M.L.; Gomez, M. In vitro antimicrobial susceptibility of Propionibacterium acnes isolated from acne patients in northern Mexico. Int. J. Dermatol. 2010, 49, 1003–1007. [Google Scholar] [CrossRef] [PubMed]
- Grech, I. Susceptibility profiles of Propionibacterium acnes isolated from patients with acne vulgaris. J. Glob. Antimicrob. Resist. 2014, 2, 35–38. [Google Scholar] [CrossRef] [PubMed]
- Luk, N.M.; Hui, M.; Lee, H.C.; Fu, L.H.; Liu, Z.H.; Lam, L.Y.; Eastel, M.; Chan, Y.K.; Tang, L.S.; Cheng, T.S.; et al. Antibiotic-resistant Propionibacterium acnes among acne patients in a regional skin centre in Hong Kong. J. Eur. Acad. Dermatol. Venereol. JEADV 2013, 27, 31–36. [Google Scholar] [CrossRef] [PubMed]
- Ross, J.I.; Snelling, A.M.; Carnegie, E.; Coates, P.; Cunliffe, W.J.; Bettoli, V.; Tosti, G.; Katsambas, A.; Galvan Perez Del Pulgar, J.I.; Rollman, O.; et al. Antibiotic-resistant acne: Lessons from Europe. Br. J. Dermatol. 2003, 148, 467–478. [Google Scholar] [CrossRef] [PubMed]
- Giannopoulos, L.; Papaparaskevas, J.; Refene, E.; Daikos, G.; Stavrianeas, N.; Tsakris, A. MLST typing of antimicrobial-resistant Propionibacterium acnes isolates from patients with moderate to severe acne vulgaris. Anaerobe 2015, 31, 50–54. [Google Scholar] [CrossRef]
- El-Mahdy, T.S.; Abdalla, S.; El-Domany, R.; Mohamed, M.S.; Ross, J.I.; Snelling, A.M. Detection of a new erm(X)-mediated antibiotic resistance in Egyptian cutaneous propionibacteria. Anaerobe 2010, 16, 376–379. [Google Scholar] [CrossRef]
- Nakase, K.; Nakaminami, H.; Takenaka, Y.; Hayashi, N.; Kawashima, M.; Noguchi, N. Relationship between the severity of acne vulgaris and antimicrobial resistance of bacteria isolated from acne lesions in a hospital in Japan. J. Med. Microbiol. 2014, 63, 721–728. [Google Scholar] [CrossRef]
- Nakase, K.; Nakaminami, H.; Takenaka, Y.; Hayashi, N.; Kawashima, M.; Noguchi, N. Propionibacterium acnes is developing gradual increase in resistance to oral tetracyclines. J. Med. Microbiol. 2017, 66, 8–12. [Google Scholar] [CrossRef] [PubMed]
- Lomholt, H.B.; Kilian, M. Clonality and anatomic distribution on the skin of antibiotic resistant and sensitive Propionibacterium acnes. Acta Derm. Venereol. 2014, 94, 534–538. [Google Scholar] [CrossRef]
- Nishijima, S.; Kurokawa, I.; Katoh, N.; Watanabe, K. The bacteriology of acne vulgaris and antimicrobial susceptibility of Propionibacterium acnes and Staphylococcus epidermidis isolated from acne lesions. J. Dermatol. 2000, 27, 318–323. [Google Scholar] [CrossRef] [PubMed]
- Nast, A.; Dreno, B.; Bettoli, V.; Bukvic Mokos, Z.; Degitz, K.; Dressler, C.; Finlay, A.Y.; Haedersdal, M.; Lambert, J.; Layton, A.; et al. European evidence-based (S3) guideline for the treatment of acne—Update 2016—Short version. J. Eur. Acad. Dermatol. Venereol. JEADV 2016, 30, 1261–1268. [Google Scholar] [CrossRef]
- Feneran, A.N.; Kaufman, W.S.; Dabade, T.S.; Feldman, S.R. Retinoid plus antimicrobial combination treatments for acne. Clin. Cosmet. Investig. Dermatol. 2011, 4, 79–92. [Google Scholar] [CrossRef] [PubMed]
- Thiboutot, D.; Gollnick, H.; Bettoli, V.; Dreno, B.; Kang, S.; Leyden, J.J.; Shalita, A.R.; Lozada, V.T.; Berson, D.; Finlay, A.; et al. New insights into the management of acne: An update from the Global Alliance to Improve Outcomes in Acne group. J. Am. Acad. Dermatol. 2009, 60, S1–50. [Google Scholar] [CrossRef]
- Niemeyer-van der Kolk, T.; van der Wall, H.E.C.; Balmforth, C.; Van Doorn, M.B.A.; Rissmann, R. A systematic literature review of the human skin microbiome as biomarker for dermatological drug development. Br. J. Clin. Pharmacol. 2018, 84, 2178–2193. [Google Scholar] [CrossRef]
- Levkovich, T.; Poutahidis, T.; Smillie, C.; Varian, B.J.; Ibrahim, Y.M.; Lakritz, J.R.; Alm, E.J.; Erdman, S.E. Probiotic bacteria induce a ‘glow of health’. PLoS ONE 2013, 8, e53867. [Google Scholar] [CrossRef]
- Ipci, K.; Altintoprak, N.; Muluk, N.B.; Senturk, M.; Cingi, C. The possible mechanisms of the human microbiome in allergic diseases. Eur. Arch. Oto-Rhino-Laryngol. 2017, 274, 617–626. [Google Scholar] [CrossRef]
- Wu, G.D.; Lewis, J.D. Analysis of the human gut microbiome and association with disease. Clin. Gastroenterol. Hepatol. Off. Clin. Pract. J. Am. Gastroenterol. Assoc. 2013, 11, 774–777. [Google Scholar] [CrossRef]
- Moore-Connors, J.M.; Dunn, K.A.; Bielawski, J.P.; Van Limbergen, J. Novel Strategies for Applied Metagenomics. Inflamm. Bowel Dis. 2016, 22, 709–718. [Google Scholar] [CrossRef] [PubMed]
- Clark, A.K.; Haas, K.N.; Sivamani, R.K. Edible Plants and Their Influence on the Gut Microbiome and Acne. Int. J. Mol. Sci. 2017, 18, 1070. [Google Scholar] [CrossRef] [PubMed]
- Salem, I.; Ramser, A.; Isham, N.; Ghannoum, M.A. The Gut Microbiome as a Major Regulator of the Gut-Skin Axis. Front. Microbiol. 2018, 9, 1459. [Google Scholar] [CrossRef] [PubMed]
- Bowe, W.P.; Logan, A.C. Acne vulgaris, probiotics and the gut-brain-skin axis - Back to the future? Gut Pathog. 2011, 3, 1. [Google Scholar] [CrossRef] [PubMed]
- Samuelson, D.R.; Welsh, D.A.; Shellito, J.E. Regulation of lung immunity and host defense by the intestinal microbiota. Front. Microbiol. 2015, 6, 1085. [Google Scholar] [CrossRef] [PubMed]
- Schwarz, A.; Bruhs, A.; Schwarz, T. The Short-Chain Fatty Acid Sodium Butyrate Functions as a Regulator of the Skin Immune System. J. Investig. Dermatol. 2017, 137, 855–864. [Google Scholar] [CrossRef] [PubMed]
- Jung, M.J.; Lee, J.; Shin, N.R.; Kim, M.S.; Hyun, D.W.; Yun, J.H.; Kim, P.S.; Whon, T.W.; Bae, J.W. Chronic Repression of mTOR Complex 2 Induces Changes in the Gut Microbiota of Diet-induced Obese Mice. Sci. Rep. 2016, 6, 30887. [Google Scholar] [CrossRef]
- Ryu, J.H.; Kim, S.H.; Lee, H.Y.; Bai, J.Y.; Nam, Y.D.; Bae, J.W.; Lee, D.G.; Shin, S.C.; Ha, E.M.; Lee, W.J. Innate immune homeostasis by the homeobox gene caudal and commensal-gut mutualism in Drosophila. Science 2008, 319, 777–782. [Google Scholar] [CrossRef]
- Sommer, F.; Adam, N.; Johansson, M.E.; Xia, L.; Hansson, G.C.; Backhed, F. Altered mucus glycosylation in core 1 O-glycan-deficient mice affects microbiota composition and intestinal architecture. PLoS ONE 2014, 9, e85254. [Google Scholar] [CrossRef]
- Kimura, I.; Ozawa, K.; Inoue, D.; Imamura, T.; Kimura, K.; Maeda, T.; Terasawa, K.; Kashihara, D.; Hirano, K.; Tani, T.; et al. The gut microbiota suppresses insulin-mediated fat accumulation via the short-chain fatty acid receptor GPR43. Nat. Commun. 2013, 4, 1829. [Google Scholar] [CrossRef]
- Feng, Y.; Ralls, M.W.; Xiao, W.; Miyasaka, E.; Herman, R.S.; Teitelbaum, D.H. Loss of enteral nutrition in a mouse model results in intestinal epithelial barrier dysfunction. Ann. N. Y. Acad. Sci. 2012, 1258, 71–77. [Google Scholar] [CrossRef] [PubMed]
- Thomson, A.W.; Turnquist, H.R.; Raimondi, G. Immunoregulatory functions of mTOR inhibition. Nat. Rev. Immunol. 2009, 9, 324–337. [Google Scholar] [CrossRef] [PubMed]
- Arpaia, N.; Campbell, C.; Fan, X.; Dikiy, S.; van der Veeken, J.; deRoos, P.; Liu, H.; Cross, J.R.; Pfeffer, K.; Coffer, P.J.; et al. Metabolites produced by commensal bacteria promote peripheral regulatory T-cell generation. Nature 2013, 504, 451–455. [Google Scholar] [CrossRef] [PubMed]
- Smith, P.M.; Howitt, M.R.; Panikov, N.; Michaud, M.; Gallini, C.A.; Bohlooly, Y.M.; Glickman, J.N.; Garrett, W.S. The microbial metabolites, short-chain fatty acids, regulate colonic Treg cell homeostasis. Science 2013, 341, 569–573. [Google Scholar] [CrossRef] [PubMed]
- Furusawa, Y.; Obata, Y.; Fukuda, S.; Endo, T.A.; Nakato, G.; Takahashi, D.; Nakanishi, Y.; Uetake, C.; Kato, K.; Kato, T.; et al. Commensal microbe-derived butyrate induces the differentiation of colonic regulatory T cells. Nature 2013, 504, 446–450. [Google Scholar] [CrossRef] [PubMed]
- Monfrecola, G.; Lembo, S.; Caiazzo, G.; De Vita, V.; Di Caprio, R.; Balato, A.; Fabbrocini, G. Mechanistic target of rapamycin (mTOR) expression is increased in acne patients’ skin. Exp. Dermatol. 2016, 25, 153–155. [Google Scholar] [CrossRef] [PubMed]
- Melnik, B.C. Linking diet to acne metabolomics, inflammation, and comedogenesis: An update. Clin. Cosmet. Investig. Dermatol. 2015, 8, 371–388. [Google Scholar] [CrossRef]
- Marcason, W. Milk consumption and acne--is there a link? J. Am. Diet. Assoc. 2010, 110, 152. [Google Scholar] [CrossRef]
- Deng, Y.; Wang, H.; Zhou, J.; Mou, Y.; Wang, G.; Xiong, X. Patients with Acne Vulgaris Have a Distinct Gut Microbiota in Comparison with Healthy Controls. Acta Derm. Venereol. 2018, 98, 783–790. [Google Scholar] [CrossRef]
- Morales, P.; Fujio, S.; Navarrete, P.; Ugalde, J.A.; Magne, F.; Carrasco-Pozo, C.; Tralma, K.; Quezada, M.; Hurtado, C.; Covarrubias, N.; et al. Impact of Dietary Lipids on Colonic Function and Microbiota: An Experimental Approach Involving Orlistat-Induced Fat Malabsorption in Human Volunteers. Clin. Transl. Gastroenterol. 2016, 7, e161. [Google Scholar] [CrossRef]
- Reddymasu, S.C.; Sostarich, S.; McCallum, R.W. Small intestinal bacterial overgrowth in irritable bowel syndrome: Are there any predictors? BMC Gastroenterol. 2010, 10, 23. [Google Scholar] [CrossRef] [PubMed]
- Lombardo, L.; Foti, M.; Ruggia, O.; Chiecchio, A. Increased incidence of small intestinal bacterial overgrowth during proton pump inhibitor therapy. Clin. Gastroenterol. Hepatol. Off. Clin. Pract. J. Am. Gastroenterol. Assoc. 2010, 8, 504–508. [Google Scholar] [CrossRef] [PubMed]
- Loveman, D.E.; Noojin, R.O.; Winkler, C.H., Jr. Comparative studies of enteric bacterial flora in acne vulgaris. J. Investig. Dermatol. 1955, 25, 135–137. [Google Scholar] [CrossRef] [PubMed]
- Volkova, L.A.; Khalif, I.L.; Kabanova, I.N. Impact of the impaired intestinal microflora on the course of acne vulgaris. Klin. Meditsina 2001, 79, 39–41. [Google Scholar]
- Yan, H.M.; Zhao, H.J.; Guo, D.Y.; Zhu, P.Q.; Zhang, C.L.; Jiang, W. Gut microbiota alterations in moderate to severe acne vulgaris patients. J. Dermatol. 2018, 45, 1166–1171. [Google Scholar] [CrossRef] [PubMed]
- Lee, D.K.; Kim, M.J.; Ham, J.W.; An, H.M.; Cha, M.K.; Lee, S.W.; Park, C.I.; Shin, S.H.; Lee, K.O.; Kim, K.J.; et al. In vitro evaluation of antibacterial activities and anti-inflammatory effects of Bifidobacterium spp. addressing acne vulgaris. Arch. Pharmacal. Res. 2012, 35, 1065–1071. [Google Scholar] [CrossRef] [PubMed]
- Madsen, K.; Cornish, A.; Soper, P.; McKaigney, C.; Jijon, H.; Yachimec, C.; Doyle, J.; Jewell, L.; De Simone, C. Probiotic bacteria enhance murine and human intestinal epithelial barrier function. Gastroenterology 2001, 121, 580–591. [Google Scholar] [CrossRef]
- Kwon, H.K.; Lee, C.G.; So, J.S.; Chae, C.S.; Hwang, J.S.; Sahoo, A.; Nam, J.H.; Rhee, J.H.; Hwang, K.C.; Im, S.H. Generation of regulatory dendritic cells and CD4+Foxp3+ T cells by probiotics administration suppresses immune disorders. Proc. Natl. Acad. Sci. USA 2010, 107, 2159–2164. [Google Scholar] [CrossRef]
- Mahowald, M.A.; Rey, F.E.; Seedorf, H.; Turnbaugh, P.J.; Fulton, R.S.; Wollam, A.; Shah, N.; Wang, C.; Magrini, V.; Wilson, R.K.; et al. Characterizing a model human gut microbiota composed of members of its two dominant bacterial phyla. Proc. Natl. Acad. Sci. USA 2009, 106, 5859–5864. [Google Scholar] [CrossRef]
- Becker, E.; Schmidt, T.S.B.; Bengs, S.; Poveda, L.; Opitz, L.; Atrott, K.; Stanzel, C.; Biedermann, L.; Rehman, A.; Jonas, D.; et al. Effects of oral antibiotics and isotretinoin on the murine gut microbiota. Int. J. Antimicrob. Agents 2017, 50, 342–351. [Google Scholar] [CrossRef]
- Servin, A.L.; Coconnier, M.H. Adhesion of probiotic strains to the intestinal mucosa and interaction with pathogens. Best Pract. Res. Clin. Gastroenterol. 2003, 17, 741–754. [Google Scholar] [CrossRef]
- Gibson, G.R.; Roberfroid, M.B. Dietary modulation of the human colonic microbiota: Introducing the concept of prebiotics. J. Nutr. 1995, 125, 1401–1412. [Google Scholar] [CrossRef] [PubMed]
- Ouwehand, A.; Isolauri, E.; Salminen, S. The role of the intestinal microflora for the development of the immune system in early childhood. Eur. J. Nutr. 2002, 41, I32–37. [Google Scholar] [CrossRef] [PubMed]
- Kober, M.M.; Bowe, W.P. The effect of probiotics on immune regulation, acne, and photoaging. Int. J. Womens Dermatol. 2015, 1, 85–89. [Google Scholar] [CrossRef] [PubMed]
- Hsieh, F.C.; Lee, C.L.; Chai, C.Y.; Chen, W.T.; Lu, Y.C.; Wu, C.S. Oral administration of Lactobacillus reuteri GMNL-263 improves insulin resistance and ameliorates hepatic steatosis in high fructose-fed rats. Nutr. Metab. 2013, 10, 35. [Google Scholar] [CrossRef] [PubMed]
- Hacini-Rachinel, F.; Gheit, H.; Le Luduec, J.B.; Dif, F.; Nancey, S.; Kaiserlian, D. Oral probiotic control skin inflammation by acting on both effector and regulatory T cells. PLoS ONE 2009, 4, e4903. [Google Scholar] [CrossRef] [PubMed]
- Benson, K.F.; Redman, K.A.; Carter, S.G.; Keller, D.; Farmer, S.; Endres, J.R.; Jensen, G.S. Probiotic metabolites from Bacillus coagulans GanedenBC30 support maturation of antigen-presenting cells in vitro. World J. Gastroenterol. 2012, 18, 1875–1883. [Google Scholar] [CrossRef]
- Jensen, G.S.; Benson, K.F.; Carter, S.G.; Endres, J.R. GanedenBC30 cell wall and metabolites: Anti-inflammatory and immune modulating effects in vitro. BMC Immunol. 2010, 11, 15. [Google Scholar] [CrossRef]
- Bowe, W.P.; Filip, J.C.; DiRienzo, J.M.; Volgina, A.; Margolis, D.J. Inhibition of propionibacterium acnes by bacteriocin-like inhibitory substances (BLIS) produced by Streptococcus salivarius. J. Drugs Dermatol. JDD 2006, 5, 868–870. [Google Scholar]
- Oh, S.; Kim, S.H.; Ko, Y.; Sim, J.H.; Kim, K.S.; Lee, S.H.; Park, S.; Kim, Y.J. Effect of bacteriocin produced by Lactococcus sp. HY 449 on skin-inflammatory bacteria. Food Chem. Toxicol. Int. J. Publ. Br. Ind. Biol. Res. Assoc. 2006, 44, 1184–1190. [Google Scholar] [CrossRef]
- Di Marzio, L.; Centi, C.; Cinque, B.; Masci, S.; Giuliani, M.; Arcieri, A.; Zicari, L.; De Simone, C.; Cifone, M.G. Effect of the lactic acid bacterium Streptococcus thermophilus on stratum corneum ceramide levels and signs and symptoms of atopic dermatitis patients. Exp. Derm. 2003, 12, 615–620. [Google Scholar] [CrossRef] [PubMed]
- Di Marzio, L.; Cinque, B.; Cupelli, F.; De Simone, C.; Cifone, M.G.; Giuliani, M. Increase of skin-ceramide levels in aged subjects following a short-term topical application of bacterial sphingomyelinase from Streptococcus thermophilus. Int. J. Immunopathol. Pharmacol. 2008, 21, 137–143. [Google Scholar] [CrossRef] [PubMed]
- Di Marzio, L.; Cinque, B.; De Simone, C.; Cifone, M.G. Effect of the lactic acid bacterium Streptococcus thermophilus on ceramide levels in human keratinocytes in vitro and stratum corneum in vivo. J. Investig. Dermatol. 1999, 113, 98–106. [Google Scholar] [CrossRef] [PubMed]
- Pavicic, T.; Wollenweber, U.; Farwick, M.; Korting, H.C. Anti-microbial and -inflammatory activity and efficacy of phytosphingosine: An in vitro and in vivo study addressing acne vulgaris. Int. J. Cosmet. Sci. 2007, 29, 181–190. [Google Scholar] [CrossRef] [PubMed]
- Cosseau, C.; Devine, D.A.; Dullaghan, E.; Gardy, J.L.; Chikatamarla, A.; Gellatly, S.; Yu, L.L.; Pistolic, J.; Falsafi, R.; Tagg, J.; et al. The commensal Streptococcus salivarius K12 downregulates the innate immune responses of human epithelial cells and promotes host-microbe homeostasis. Infect. Immun. 2008, 76, 4163–4175. [Google Scholar] [CrossRef]
- Gueniche, A.; Bastien, P.; Ovigne, J.M.; Kermici, M.; Courchay, G.; Chevalier, V.; Breton, L.; Castiel-Higounenc, I. Bifidobacterium longum lysate, a new ingredient for reactive skin. Exp. Derm. 2010, 19, e1–e8. [Google Scholar] [CrossRef] [PubMed]
- Gueniche, A.; Benyacoub, J.; Philippe, D.; Bastien, P.; Kusy, N.; Breton, L.; Blum, S.; Castiel-Higounenc, I. Lactobacillus paracasei CNCM I-2116 (ST11) inhibits substance P-induced skin inflammation and accelerates skin barrier function recovery in vitro. Eur. J. Dermatol. EJD 2010, 20, 731–737. [Google Scholar] [CrossRef]
- Lee, W.J.; Jung, H.D.; Lee, H.J.; Kim, B.S.; Lee, S.J.; Kim, D.W. Influence of substance-P on cultured sebocytes. Arch. Dermatol. Res. 2008, 300, 311–316. [Google Scholar] [CrossRef]
- Bowe, W.P.; Logan, A.C. Clinical implications of lipid peroxidation in acne vulgaris: Old wine in new bottles. Lipids Health Dis. 2010, 9, 141. [Google Scholar] [CrossRef]
- Quadros, E.; Landzert, N.M.; LeRoy, S.; Gasparini, F.; Worosila, G. Colonic absorption of insulin-like growth factor I in vitro. Pharm. Res. 1994, 11, 226–230. [Google Scholar] [CrossRef]
- Kang, B.S.; Seo, J.G.; Lee, G.S.; Kim, J.H.; Kim, S.Y.; Han, Y.W.; Kang, H.; Kim, H.O.; Rhee, J.H.; Chung, M.J.; et al. Antimicrobial activity of enterocins from Enterococcus faecalis SL-5 against Propionibacterium acnes, the causative agent in acne vulgaris, and its therapeutic effect. J. Microbiol. (Seoulkorea) 2009, 47, 101–109. [Google Scholar] [CrossRef] [PubMed]
- Muizzuddin, N.; Maher, W.; Sullivan, M.; Schnittger, S.; Mammone, T. Physiological effect of a probiotic on skin. J. Cosmet. Sci. 2012, 63, 385–395. [Google Scholar] [PubMed]
- Marchetti, F.; Capizzi, R.; Tulli, A. Efficacy of regulators of the intestinal bacterial flora in the therapy of acne vulgaris. Clin. Ter. 1987, 122, 339–343. [Google Scholar] [PubMed]
- Jung, G.W.; Tse, J.E.; Guiha, I.; Rao, J. Prospective, randomized, open-label trial comparing the safety, efficacy, and tolerability of an acne treatment regimen with and without a probiotic supplement and minocycline in subjects with mild to moderate acne. J. Cutan. Med. Surg. 2013, 17, 114–122. [Google Scholar] [CrossRef] [PubMed]
| Sampling Method | ||||
|---|---|---|---|---|
| Swab | Scrape | Pore Strip | Biopsy | |
| C. acnes populations | ||||
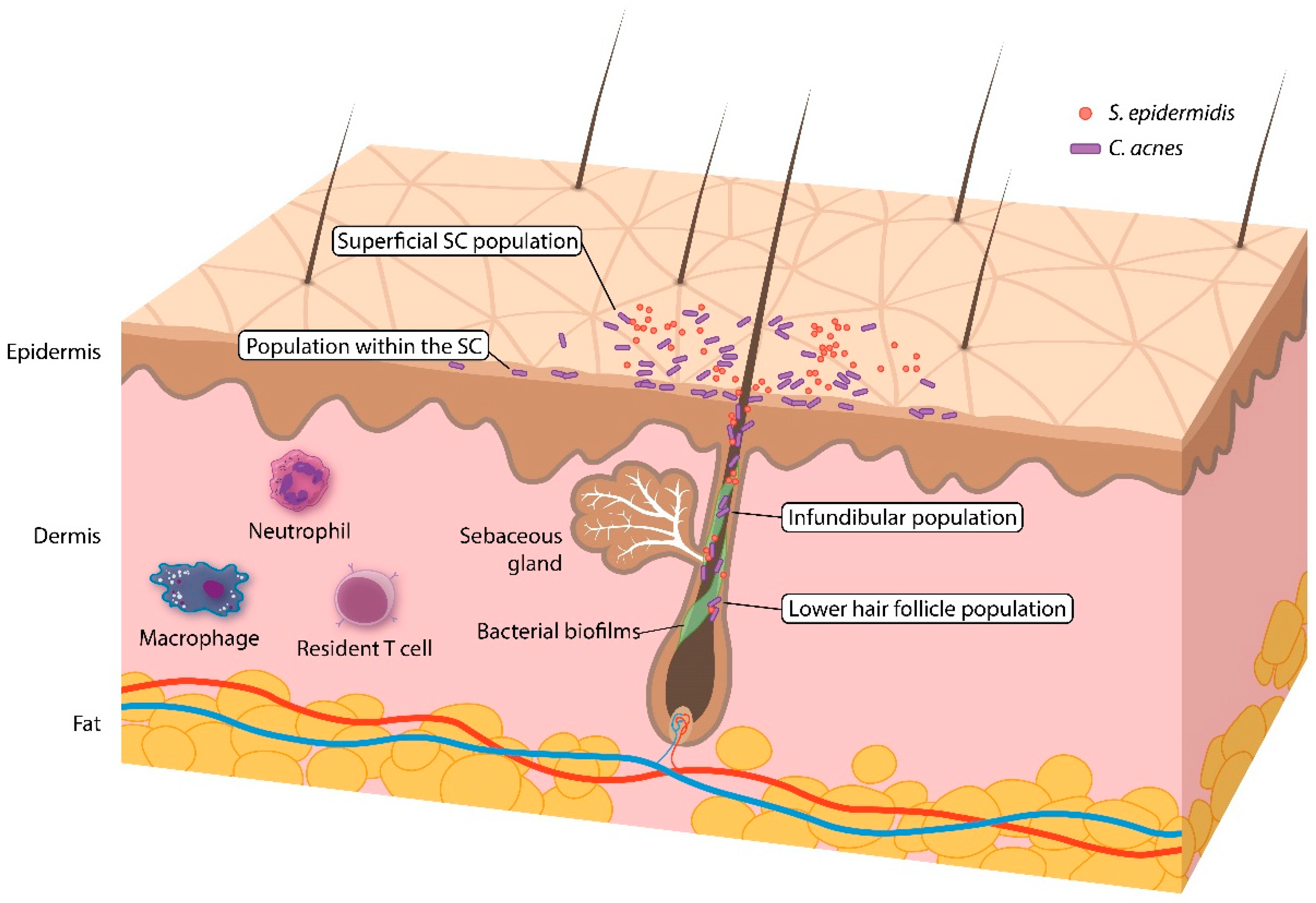
| Superficial stratum corneum | + | + | + | ± a |
| Within stratum corneum | – | + | + | + |
| Infundibulum | – | – | + | + |
| Lower hair follicle | – | – | – | + |
| Follicular biofilms | – | – | – | ± b |
| Advantages and disadvantages | ||||
| Pros | Simple, quick, and noninvasive. | Enables collection of skin cells and their associated microbes. | Collects follicular contents. | Samples all layers of the skin. |
| Cons | Might not correctly reflect the microbiota across all skin layers. | Might not correctly reflect the microbiota across all skin layers. | Might not reflect the microbiota in the lower hair follicles. | Is invasive and covers a smaller surface area than the other sampling methods. |
| Clade (Based on Whole-Genome Sequencing) | Clade (Based on Belfast eMLST [38]) | Clade (Based on Aarhus MLST [39]) | RT [30] | Acne | Healthy Skin |
|---|---|---|---|---|---|
| IA-1 | IA1 | I-1a | RT1 | √ | √ |
| IA-2 | IA1 | I-1a | RT4, RT5 | √ | |
| IB-1 | IA1 | I-1b | RT8 | √ | |
| IB-2 | IA2 | I-1a | RT3 | √ | √ |
| IB-3 | IB | I-2 | RT1 | √ | √ |
| IC | IC | NA | RT5 | √ | |
| II | II | II | RT2, RT6 | √ | |
| III | III | III | NA |
| Significant Changes in Skin Microbiota | Significant Changes in Gut Microbiota |
|---|---|
| ↑Cutibacterium acnes ↑Cutibacterium granulosum ↑Staphylococcus epidermidis ↑Proteobacteria and Firmicutes ↓Actinobacteria ↑Sterptococcus (pre-adolescent) ↑Malassezia species | ↑Bacteroides |
| Key Microbes Involved | Potentially Beneficial Microorganisms | Main Mechanism of Action | Experimental Model |
|---|---|---|---|
| C. acne (hyper-colonization and dominance of virulent strains) | Staphylococcus epidermidis [18] | Fermentation of glycerol (inhibition of C. acnes growth) | In vitro |
| Streptococcus salivarius [166] | Production of bacteriocin-like inhibitory substance (inhibition of C. acnes growth) | In vitro | |
| Lactococcus sp. HY449 [167] | Release of bacteriocin (inhibition of C. acnes growth) | In vitro | |
| Streptococcus thermophiles [169,170] | Increase in ceramide production, secondary antimicrobial activity (restoration of the skin barrier, inhibition of C. acnes growth) | In vivo, In vitro | |
| Lactobacillus paracasei [173,174] | Suppression of substance P-induced inflammation (reduction of inflammation) | Ex vivo | |
| Enterococcus faecalis [178] | Production of enterocins (inhibition of C. acnes growth) | In vivo | |
| Lactobacillus plantarum [179] | Production of antimicrobial peptides (inhibition of C. acnes growth) | In vivo |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lee, Y.B.; Byun, E.J.; Kim, H.S. Potential Role of the Microbiome in Acne: A Comprehensive Review. J. Clin. Med. 2019, 8, 987. https://doi.org/10.3390/jcm8070987
Lee YB, Byun EJ, Kim HS. Potential Role of the Microbiome in Acne: A Comprehensive Review. Journal of Clinical Medicine. 2019; 8(7):987. https://doi.org/10.3390/jcm8070987
Chicago/Turabian StyleLee, Young Bok, Eun Jung Byun, and Hei Sung Kim. 2019. "Potential Role of the Microbiome in Acne: A Comprehensive Review" Journal of Clinical Medicine 8, no. 7: 987. https://doi.org/10.3390/jcm8070987
APA StyleLee, Y. B., Byun, E. J., & Kim, H. S. (2019). Potential Role of the Microbiome in Acne: A Comprehensive Review. Journal of Clinical Medicine, 8(7), 987. https://doi.org/10.3390/jcm8070987