H. pylori Eradication Treatment Alters Gut Microbiota and GLP-1 Secretion in Humans
Abstract
1. Introduction
2. Materials and Methods
2.1. Study Design and Population
2.2. Bioinformatic Analysis
2.3. Statistical Analysis
3. Results
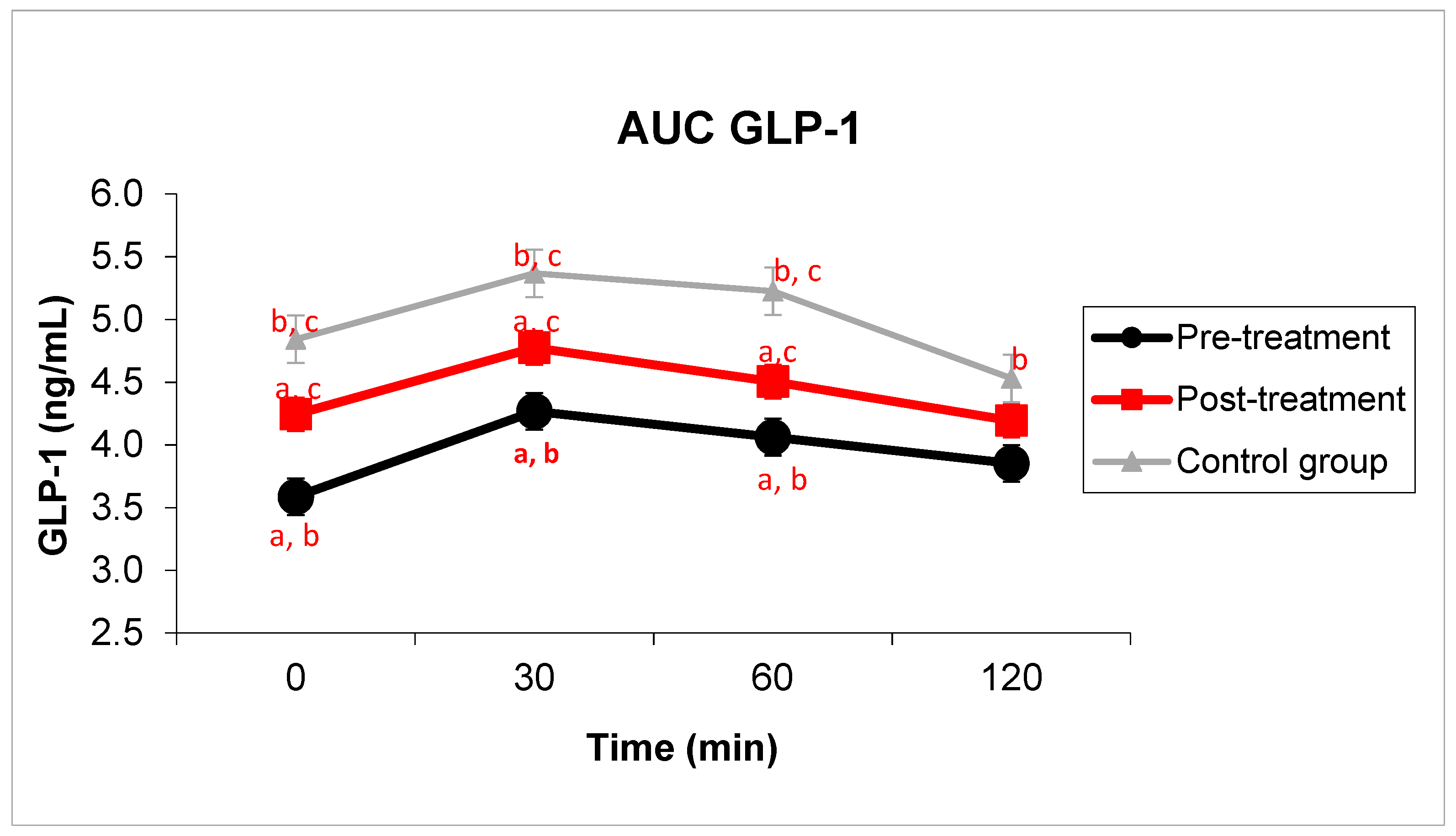
3.1. Carbohydrate Metabolism
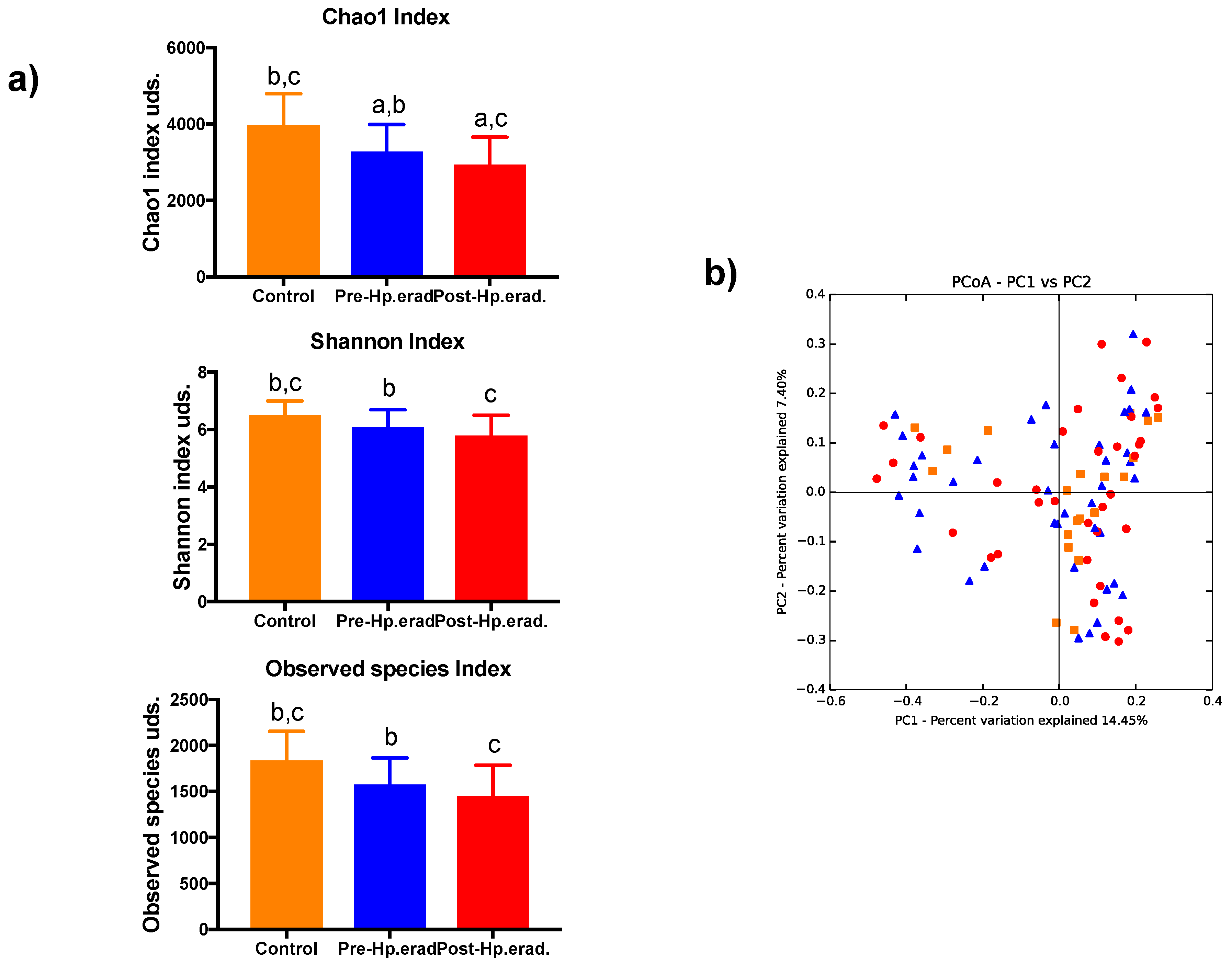
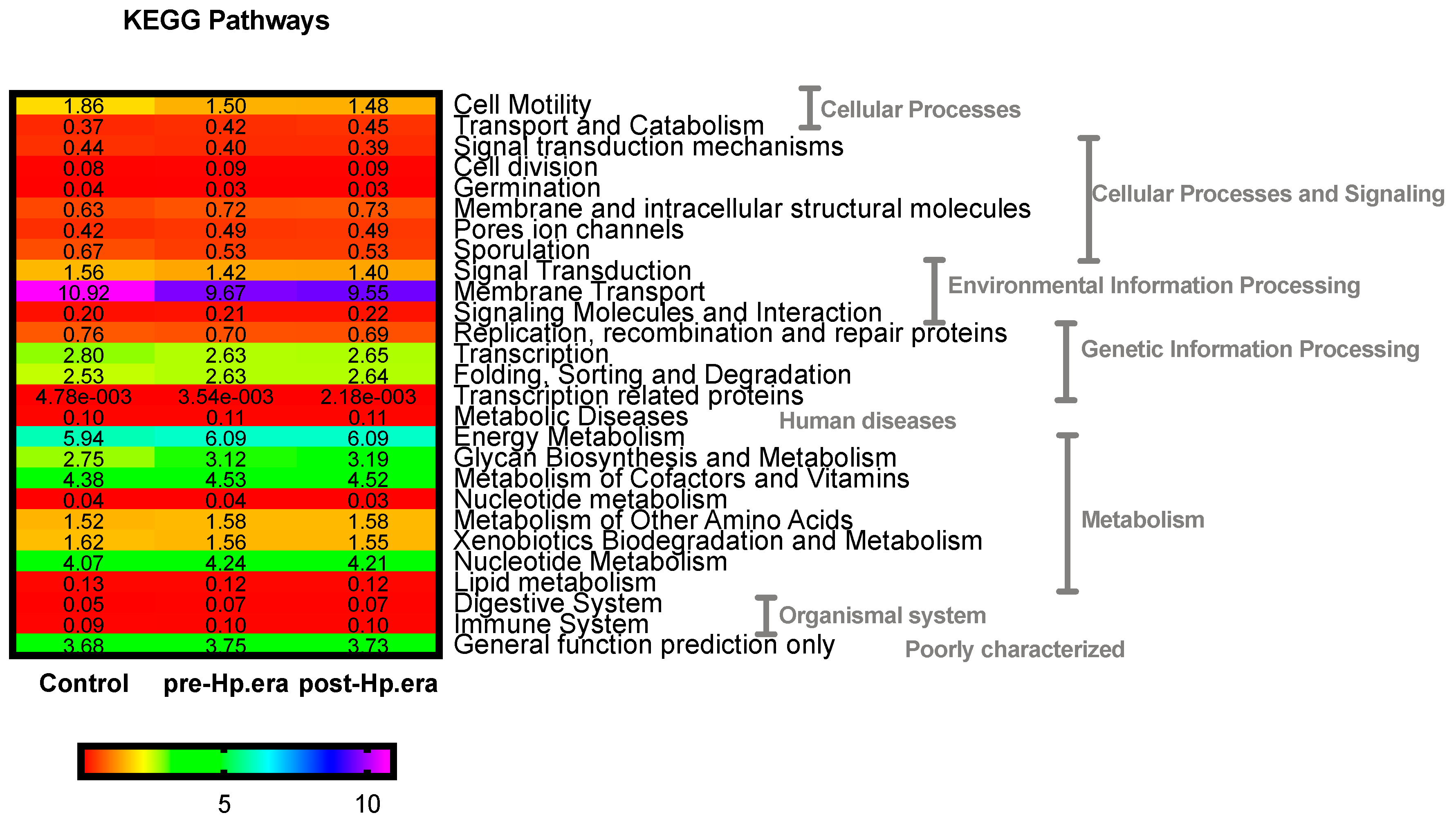
3.2. Gut Microbiota
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Arumugam, M.; Raes, J.; Pelletier, E.; Le Paslier, D.; Yamada, T.; Mende, D.R.; Fernandes, G.R.; Tap, J.; Bruls, T.; Batto, J.-M.; et al. Enterotypes of the human gut microbiome. Nature 2011, 473, 174–180. [Google Scholar] [CrossRef] [PubMed]
- Ridaura, V.K.; Faith, J.J.; Rey, F.E.; Cheng, J.; Duncan, A.E.; Kau, A.L.; Griffin, N.W.; Lombard, V.; Henrissat, B.; Bain, J.R.; et al. Gut microbiota from twins discordant for obesity modulate metabolism in mice. Science 2013, 341, 1241214. [Google Scholar] [CrossRef] [PubMed]
- Antunes, L.C.M.; Finlay, B.B. A comparative analysis of the effect of antibiotic treatment and enteric infection on intestinal homeostasis. Gut Microbes 2011, 2, 105–108. [Google Scholar] [CrossRef] [PubMed]
- Koenig, J.E.; Spor, A.; Scalfone, N.; Fricker, A.D.; Stombaugh, J.; Knight, R.; Angenent, L.T.; Ley, R.E. Succession of microbial consortia in the developing infant gut microbiome. Proc. Natl. Acad. Sci. USA 2011, 108 (Suppl. 1), 4578–4585. [Google Scholar] [CrossRef]
- Karlsson, F.H.; Tremaroli, V.; Nookaew, I.; Bergström, G.; Behre, C.J.; Fagerberg, B.; Nielsen, J.; Bäckhed, F. Gut metagenome in European women with normal, impaired and diabetic glucose control. Nature 2013, 498, 99–103. [Google Scholar] [CrossRef] [PubMed]
- Turnbaugh, P.J.; Ley, R.E.; Mahowald, M.A.; Magrini, V.; Mardis, E.R.; Gordon, J.I. An obesity-associated gut microbiome with increased capacity for energy harvest. Nature 2006, 444, 1027–1031. [Google Scholar] [CrossRef] [PubMed]
- Rask, E.; Olsson, T.; Söderberg, S.; Johnson, O.; Seckl, J.; Holst, J.J.; Ahrén, B. Northern Sweden Monitoring of Trends and Determinants in Cardiovascular Disease (MONICA) Impaired incretin response after a mixed meal is associated with insulin resistance in nondiabetic men. Diabetes Care 2001, 24, 1640–1645. [Google Scholar] [CrossRef]
- Holst, J.J. The physiology of glucagon-like peptide 1. Physiol. Rev. 2007, 87, 1409–1439. [Google Scholar] [CrossRef]
- Farilla, L.; Bulotta, A.; Hirshberg, B.; Li Calzi, S.; Khoury, N.; Noushmehr, H.; Bertolotto, C.; Di Mario, U.; Harlan, D.M.; Perfetti, R. Glucagon-like peptide 1 inhibits cell apoptosis and improves glucose responsiveness of freshly isolated human islets. Endocrinology 2003, 144, 5149–5158. [Google Scholar] [CrossRef]
- Zietek, T.; Rath, E. Inflammation Meets Metabolic Disease: Gut Feeling Mediated by GLP-1. Front. Immunol. 2016, 7, 154. [Google Scholar] [CrossRef]
- Freeland, K.R.; Wilson, C.; Wolever, T.M.S. Adaptation of colonic fermentation and glucagon-like peptide-1 secretion with increased wheat fibre intake for 1 year in hyperinsulinaemic human subjects. Br. J. Nutr. 2010, 103, 82–90. [Google Scholar] [CrossRef] [PubMed]
- Zhou, J.; Martin, R.J.; Tulley, R.T.; Raggio, A.M.; McCutcheon, K.L.; Shen, L.; Danna, S.C.; Tripathy, S.; Hegsted, M.; Keenan, M.J. Dietary resistant starch upregulates total GLP-1 and PYY in a sustained day-long manner through fermentation in rodents. Am. J. Physiol. Endocrinol. Metab. 2008, 295, E1160–E1166. [Google Scholar] [CrossRef]
- Martín-Núñez, G.M.; Cornejo-Pareja, I.; Coin-Aragüez, L.; Roca-Rodríguez, M.D.M.; Muñoz-Garach, A.; Clemente-Postigo, M.; Cardona, F.; Moreno-Indias, I.; Tinahones, F.J.H. H. pylori eradication with antibiotic treatment causes changes in glucose homeostasis related to modifications in the gut microbiota. PLoS ONE 2019, 14, e0213548. [Google Scholar] [CrossRef] [PubMed]
- Cani, P.D.; Everard, A.; Duparc, T. Gut microbiota, enteroendocrine functions and metabolism. Curr. Opin. Pharmacol. 2013, 13, 935–940. [Google Scholar] [CrossRef]
- Bala, V.; Rajagopal, S.; Kumar, D.P.; Nalli, A.D.; Mahavadi, S.; Sanyal, A.J.; Grider, J.R.; Murthy, K.S. Release of GLP-1 and PYY in response to the activation of G protein-coupled bile acid receptor TGR5 is mediated by Epac/PLC-ε pathway and modulated by endogenous H2S. Front. Physiol. 2014, 5, 420. [Google Scholar] [CrossRef]
- Puddu, A.; Sanguineti, R.; Montecucco, F.; Viviani, G.L. Evidence for the gut microbiota short-chain fatty acids as key pathophysiological molecules improving diabetes. Mediat. Inflamm. 2014, 2014, 162021. [Google Scholar] [CrossRef] [PubMed]
- Moreno-Indias, I.; Sanchez-Alcoholado, L.; Garcia-Fuentes, E.; Cardona, F.; Queipo-Ortuno, M.I.; Tinahones, F.J. Insulin resistance is associated with specific gut microbiota in appendix samples from morbidly obese patients. Am. J. Transl. Res. 2016, 8, 5672–5684. [Google Scholar] [PubMed]
- Langille, M.G.I.; Zaneveld, J.; Caporaso, J.G.; McDonald, D.; Knights, D.; Reyes, J.A.; Clemente, J.C.; Burkepile, D.E.; Vega Thurber, R.L.; Knight, R.; et al. Predictive functional profiling of microbial communities using 16S rRNA marker gene sequences. Nat. Biotechnol. 2013, 31, 814–821. [Google Scholar] [CrossRef]
- Parks, D.H.; Tyson, G.W.; Hugenholtz, P.; Beiko, R.G. STAMP: Statistical analysis of taxonomic and functional profiles. Bioinform. Oxf. Engl. 2014, 30, 3123–3124. [Google Scholar] [CrossRef]
- Polansky, O.; Sekelova, Z.; Faldynova, M.; Sebkova, A.; Sisak, F.; Rychlik, I. Important Metabolic Pathways and Biological Processes Expressed by Chicken Cecal Microbiota. Appl. Environ. Microbiol. 2015, 82, 1569–1576. [Google Scholar] [CrossRef] [PubMed]
- Burcelin, R.; Cani, P.D.; Knauf, C. Glucagon-like peptide-1 and energy homeostasis. J. Nutr. 2007, 137, 2534S–2538S. [Google Scholar] [CrossRef] [PubMed]
- Pani, L.N.; Korenda, L.; Meigs, J.B.; Driver, C.; Chamany, S.; Fox, C.S.; Sullivan, L.; D’Agostino, R.B.; Nathan, D.M. Effect of aging on A1C levels in individuals without diabetes: Evidence from the Framingham Offspring Study and the National Health and Nutrition Examination Survey 2001–2004. Diabetes Care 2008, 31, 1991–1996. [Google Scholar] [CrossRef] [PubMed]
- Kaji, I.; Karaki, S.; Kuwahara, A. Short-chain fatty acid receptor and its contribution to glucagon-like peptide-1 release. Digestion 2014, 89, 31–36. [Google Scholar] [CrossRef] [PubMed]
- Qin, J.; Li, Y.; Cai, Z.; Li, S.; Zhu, J.; Zhang, F.; Liang, S.; Zhang, W.; Guan, Y.; Shen, D.; et al. A metagenome-wide association study of gut microbiota in type 2 diabetes. Nature 2012, 490, 55–60. [Google Scholar] [CrossRef] [PubMed]
- Larsen, N.; Vogensen, F.K.; van den Berg, F.W.J.; Nielsen, D.S.; Andreasen, A.S.; Pedersen, B.K.; Al-Soud, W.A.; Sørensen, S.J.; Hansen, L.H.; Jakobsen, M. Gut microbiota in human adults with type 2 diabetes differs from non-diabetic adults. PLoS ONE 2010, 5, e9085. [Google Scholar] [CrossRef] [PubMed]
- Jakobsson, H.E.; Jernberg, C.; Andersson, A.F.; Sjölund-Karlsson, M.; Jansson, J.K.; Engstrand, L. Short-term antibiotic treatment has differing long-term impacts on the human throat and gut microbiome. PLoS ONE 2010, 5, e9836. [Google Scholar] [CrossRef] [PubMed]
- Ubeda, C.; Pamer, E.G. Antibiotics, microbiota, and immune defense. Trends Immunol. 2012, 33, 459–466. [Google Scholar] [CrossRef]
- Tian, P.; Xu, B.; Sun, H.; Li, X.; Li, Z.; Wei, P. Isolation and gut microbiota modulation of antibiotic-resistant probiotics from human feces. Diagn. Microbiol. Infect. Dis. 2014, 79, 405–412. [Google Scholar] [CrossRef]
- Dethlefsen, L.; Huse, S.; Sogin, M.L.; Relman, D.A. The pervasive effects of an antibiotic on the human gut microbiota, as revealed by deep 16S rRNA sequencing. PLoS Biol. 2008, 6, e280. [Google Scholar] [CrossRef]
- Robinson, C.J.; Young, V.B. Antibiotic administration alters the community structure of the gastrointestinal micobiota. Gut Microbes 2010, 1, 279–284. [Google Scholar] [CrossRef] [PubMed]
- Sjölund, M.; Wreiber, K.; Andersson, D.I.; Blaser, M.J.; Engstrand, L. Long-term persistence of resistant Enterococcus species after antibiotics to eradicate Helicobacter pylori. Ann. Intern. Med. 2003, 139, 483–487. [Google Scholar] [CrossRef]
- Dethlefsen, L.; Relman, D.A. Incomplete recovery and individualized responses of the human distal gut microbiota to repeated antibiotic perturbation. Proc. Natl. Acad. Sci. USA 2011, 108 (Suppl. 1), 4554–4561. [Google Scholar] [CrossRef]
- Yap, T.W.-C.; Gan, H.-M.; Lee, Y.-P.; Leow, A.H.-R.; Azmi, A.N.; Francois, F.; Perez-Perez, G.I.; Loke, M.-F.; Goh, K.-L.; Vadivelu, J. Helicobacter pylori Eradication Causes Perturbation of the Human Gut Microbiome in Young Adults. PLoS ONE 2016, 11, e0151893. [Google Scholar] [CrossRef] [PubMed]
- Jernberg, C.; Löfmark, S.; Edlund, C.; Jansson, J.K. Long-term impacts of antibiotic exposure on the human intestinal microbiota. Microbiol. Read. Engl. 2010, 156, 3216–3223. [Google Scholar] [CrossRef] [PubMed]
- Moreno-Indias, I.; Cardona, F.; Tinahones, F.J.; Queipo-Ortuño, M.I. Impact of the gut microbiota on the development of obesity and type 2 diabetes mellitus. Front. Microbiol. 2014, 5, 190. [Google Scholar] [CrossRef] [PubMed]
- Muñoz-Garach, A.; Diaz-Perdigones, C.; Tinahones, F.J. Gut microbiota and type 2 diabetes mellitus. Endocrinol. Nutr. Organol. Soc. Espanola Endocrinol. Nutr. 2016, 63, 560–568. [Google Scholar] [CrossRef]
- Le Chatelier, E.; Nielsen, T.; Qin, J.; Prifti, E.; Hildebrand, F.; Falony, G.; Almeida, M.; Arumugam, M.; Batto, J.-M.; Kennedy, S.; et al. Richness of human gut microbiome correlates with metabolic markers. Nature 2013, 500, 541–546. [Google Scholar] [CrossRef]
- Cho, I.; Yamanishi, S.; Cox, L.; Methé, B.A.; Zavadil, J.; Li, K.; Gao, Z.; Mahana, D.; Raju, K.; Teitler, I.; et al. Antibiotics in early life alter the murine colonic microbiome and adiposity. Nature 2012, 488, 621–626. [Google Scholar] [CrossRef]
- Gao, J.-J.; Zhang, Y.; Gerhard, M.; Mejias-Luque, R.; Zhang, L.; Vieth, M.; Ma, J.-L.; Bajbouj, M.; Suchanek, S.; Liu, W.-D.; et al. Association Between Gut Microbiota and Helicobacter pylori-Related Gastric Lesions in a High-Risk Population of Gastric Cancer. Front. Cell. Infect. Microbiol. 2018, 8, 202. [Google Scholar] [CrossRef]
- Schmidt, T.S.; Hayward, M.R.; Coelho, L.P.; Li, S.S.; Costea, P.I.; Voigt, A.Y.; Wirbel, J.; Maistrenko, O.M.; Alves, R.J.; Bergsten, E.; et al. Extensive transmission of microbes along the gastrointestinal tract. eLife 2019, 8, e42693. [Google Scholar] [CrossRef] [PubMed]
- Ruggiero, P. Helicobacter pylori and inflammation. Curr. Pharm. Des. 2010, 16, 4225–4236. [Google Scholar] [CrossRef] [PubMed]
- Khosravi, Y.; Seow, S.W.; Amoyo, A.A.; Chiow, K.H.; Tan, T.L.; Wong, W.Y.; Poh, Q.H.; Sentosa, I.M.D.; Bunte, R.M.; Pettersson, S.; et al. Helicobacter pylori infection can affect energy modulating hormones and body weight in germ free mice. Sci. Rep. 2015, 5, 8731. [Google Scholar] [CrossRef] [PubMed]
- Shade, A. Diversity is the question, not the answer. ISME J. 2017, 11, 1–6. [Google Scholar] [CrossRef] [PubMed]
- Vrieze, A.; Out, C.; Fuentes, S.; Jonker, L.; Reuling, I.; Kootte, R.S.; van Nood, E.; Holleman, F.; Knaapen, M.; Romijn, J.A.; et al. Impact of oral vancomycin on gut microbiota, bile acid metabolism, and insulin sensitivity. J. Hepatol. 2014, 60, 824–831. [Google Scholar] [CrossRef] [PubMed]
- Murphy, E.F.; Cotter, P.D.; Hogan, A.; O’Sullivan, O.; Joyce, A.; Fouhy, F.; Clarke, S.F.; Marques, T.M.; O’Toole, P.W.; Stanton, C.; et al. Divergent metabolic outcomes arising from targeted manipulation of the gut microbiota in diet-induced obesity. Gut 2013, 62, 220–226. [Google Scholar] [CrossRef] [PubMed]
- Mikkelsen, K.H.; Frost, M.; Bahl, M.I.; Licht, T.R.; Jensen, U.S.; Rosenberg, J.; Pedersen, O.; Hansen, T.; Rehfeld, J.F.; Holst, J.J.; et al. Effect of Antibiotics on Gut Microbiota, Gut Hormones and Glucose Metabolism. PLoS ONE 2015, 10, e0142352. [Google Scholar] [CrossRef] [PubMed]
- Hwang, I.; Park, Y.J.; Kim, Y.-R.; Kim, Y.N.; Ka, S.; Lee, H.Y.; Seong, J.K.; Seok, Y.-J.; Kim, J.B. Alteration of gut microbiota by vancomycin and bacitracin improves insulin resistance via glucagon-like peptide 1 in diet-induced obesity. FASEB J. Off. Publ. Fed. Am. Soc. Exp. Biol. 2015, 29, 2397–2411. [Google Scholar] [CrossRef]
- Rajpal, D.K.; Klein, J.-L.; Mayhew, D.; Boucheron, J.; Spivak, A.T.; Kumar, V.; Ingraham, K.; Paulik, M.; Chen, L.; Van Horn, S.; et al. Selective Spectrum Antibiotic Modulation of the Gut Microbiome in Obesity and Diabetes Rodent Models. PLoS ONE 2015, 10, e0145499. [Google Scholar] [CrossRef]
- Grasset, E.; Puel, A.; Charpentier, J.; Collet, X.; Christensen, J.E.; Tercé, F.; Burcelin, R. A Specific Gut Microbiota Dysbiosis of Type 2 Diabetic Mice Induces GLP-1 Resistance through an Enteric NO-Dependent and Gut-Brain Axis Mechanism. Cell Metab. 2017, 25, 1075–1090. [Google Scholar] [CrossRef]
- Greiner, T.U.; Bäckhed, F. Microbial regulation of GLP-1 and L-cell biology. Mol. Metab. 2016, 5, 753–758. [Google Scholar] [CrossRef]
- Whitford, M.F.; Yanke, L.J.; Forster, R.J.; Teather, R.M. Lachnobacterium bovis gen. nov., sp. nov., a novel bacterium isolated from the rumen and faeces of cattle. Int. J. Syst. Evol. Microbiol. 2001, 51, 1977–1981. [Google Scholar] [CrossRef] [PubMed]
- Morrison, D.J.; Preston, T. Formation of short chain fatty acids by the gut microbiota and their impact on human metabolism. Gut Microbes 2016, 7, 189–200. [Google Scholar] [CrossRef] [PubMed]
- Yadav, H.; Lee, J.-H.; Lloyd, J.; Walter, P.; Rane, S.G. Beneficial metabolic effects of a probiotic via butyrate-induced GLP-1 hormone secretion. J. Biol. Chem. 2013, 288, 25088–25097. [Google Scholar] [CrossRef] [PubMed]
- Tolhurst, G.; Heffron, H.; Lam, Y.S.; Parker, H.E.; Habib, A.M.; Diakogiannaki, E.; Cameron, J.; Grosse, J.; Reimann, F.; Gribble, F.M. Short-chain fatty acids stimulate glucagon-like peptide-1 secretion via the G-protein-coupled receptor FFAR2. Diabetes 2012, 61, 364–371. [Google Scholar] [CrossRef] [PubMed]
- Cani, P.D.; Possemiers, S.; Van de Wiele, T.; Guiot, Y.; Everard, A.; Rottier, O.; Geurts, L.; Naslain, D.; Neyrinck, A.; Lambert, D.M.; et al. Changes in gut microbiota control inflammation in obese mice through a mechanism involving GLP-2-driven improvement of gut permeability. Gut 2009, 58, 1091–1103. [Google Scholar] [CrossRef] [PubMed]
- Neish, A.S. Microbes in gastrointestinal health and disease. Gastroenterology 2009, 136, 65–80. [Google Scholar] [CrossRef] [PubMed]
- Amaretti, A.; Tamburini, E.; Bernardi, T.; Pompei, A.; Zanoni, S.; Vaccari, G.; Matteuzzi, D.; Rossi, M. Substrate preference of Bifidobacterium adolescentis MB 239: Compared growth on single and mixed carbohydrates. Appl. Microbiol. Biotechnol. 2006, 73, 654–662. [Google Scholar] [CrossRef] [PubMed]
- Rios-Covian, D.; Gueimonde, M.; Duncan, S.H.; Flint, H.J.; de los Reyes-Gavilan, C.G. Enhanced butyrate formation by cross-feeding between Faecalibacterium prausnitzii and Bifidobacterium adolescentis. FEMS Microbiol. Lett. 2015, 362. [Google Scholar] [CrossRef]
- Cani, P.D.; Delzenne, N.M. The role of the gut microbiota in energy metabolism and metabolic disease. Curr. Pharm. Des. 2009, 15, 1546–1558. [Google Scholar] [CrossRef]
- Velasquez-Manoff, M. Gut microbiome: The peacekeepers. Nature 2015, 518, S3–S11. [Google Scholar] [CrossRef]
- Lê, K.-A.; Li, Y.; Xu, X.; Yang, W.; Liu, T.; Zhao, X.; Tang, Y.G.; Cai, D.; Go, V.L.W.; Pandol, S.; et al. Alterations in fecal Lactobacillus and Bifidobacterium species in type 2 diabetic patients in Southern China population. Front. Physiol. 2012, 3, 496. [Google Scholar] [CrossRef] [PubMed]
- Clavel, T.; Desmarchelier, C.; Haller, D.; Gérard, P.; Rohn, S.; Lepage, P.; Daniel, H. Intestinal microbiota in metabolic diseases: From bacterial community structure and functions to species of pathophysiological relevance. Gut Microbes 2014, 5, 544–551. [Google Scholar] [CrossRef] [PubMed]
- Palau-Rodriguez, M.; Tulipani, S.; Isabel Queipo-Ortuño, M.; Urpi-Sarda, M.; Tinahones, F.J.; Andres-Lacueva, C. Metabolomic insights into the intricate gut microbial-host interaction in the development of obesity and type 2 diabetes. Front. Microbiol. 2015, 6, 1151. [Google Scholar] [CrossRef] [PubMed]
- Allin, K.H.; Nielsen, T.; Pedersen, O. Mechanisms in endocrinology: Gut microbiota in patients with type 2 diabetes mellitus. Eur. J. Endocrinol. 2015, 172, R167–R177. [Google Scholar] [CrossRef] [PubMed]
- Gomez-Arango, L.F.; Barrett, H.L.; McIntyre, H.D.; Callaway, L.K.; Morrison, M.; Dekker Nitert, M. SPRING Trial Group Connections Between the Gut Microbiome and Metabolic Hormones in Early Pregnancy in Overweight and Obese Women. Diabetes 2016, 65, 2214–2223. [Google Scholar] [CrossRef] [PubMed]
- Liu, L.; Aigner, A.; Schmid, R.D. Identification, cloning, heterologous expression, and characterization of a NADPH-dependent 7β-hydroxysteroid dehydrogenase from Collinsella aerofaciens. Appl. Microbiol. Biotechnol. 2011, 90, 127–135. [Google Scholar] [CrossRef] [PubMed]
- Tsuchida, T.; Shiraishi, M.; Ohta, T.; Sakai, K.; Ishii, S. Ursodeoxycholic acid improves insulin sensitivity and hepatic steatosis by inducing the excretion of hepatic lipids in high-fat diet-fed KK-Ay mice. Metabolism 2012, 61, 944–953. [Google Scholar] [CrossRef]

| Pre-H. pylori Eradication | Post-H. pylori Eradication | Control Group | |
|---|---|---|---|
| VARIABLES | Mean ± SEM | Mean ± SEM | Mean ± SEM |
| Sex (female/male) | (24/16) | __ | (12/8) |
| Age (years) | 47.0 ± 2.0 | __ | 43.6 ± 2.69 |
| Previous treatment with Omeprazole (%) | 35 | __ | 20 |
| Anthropometry | |||
| Weight (kg) | 72.8 ± 2.0 | 72.6 ± 2.0 | 72.5 ± 3.3 |
| BMI (Kg/m2) | 26.9 ± 0.7 | 26.2 ± 0.7 | 25.8 ± 1.0 |
| Waist (cm) | 92.3 ± 1.9 | 91.3 ± 1.9 | 89.3 ± 3.0 |
| Hip (cm) | 102.0 ± 1.3 | 102.9 ± 1.7 | 101.9 ± 2.3 |
| Blood pressure | |||
| Systolic (mmHg) | 123.9 ± 2.6 | 125.4 ± 3.5 | 120.3 ± 3.0 |
| Diastolic (mmHg) | 77.8 ± 1.5 | 80.5 ± 1.8 | 75.6 ± 2.3 |
| Lipid Profile | |||
| TGs (mg/dL) | 97.2 ± 6.3 | 93.5 ± 5.9 | 89.7 ± 9.3 |
| TC (mg/dL) | 194.2 ± 6.5 | 191.3 ± 6.3 | 177.1 ± 8.8 |
| LDL-c (mg/dL) | 121.5 ± 5.7 b | 118.0 ± 5.2 | 102.1 ± 7.6 b |
| HDL-c (mg/dL) | 53.0 ± 2.0 a | 55.4 ± 2.7 a | 57.0 ± 3.5 |
| Inflammation | |||
| CRP (mg/dL) | 4.1 ± 0.4 | 3.6 ± 0.3 | 2.2 ± 0.7 |
| Glycemic status | |||
| HbA1c (%) | 5.5 ± 0.08 a | 5.29 ± 0.06 a | 5.3 ± 0.1 |
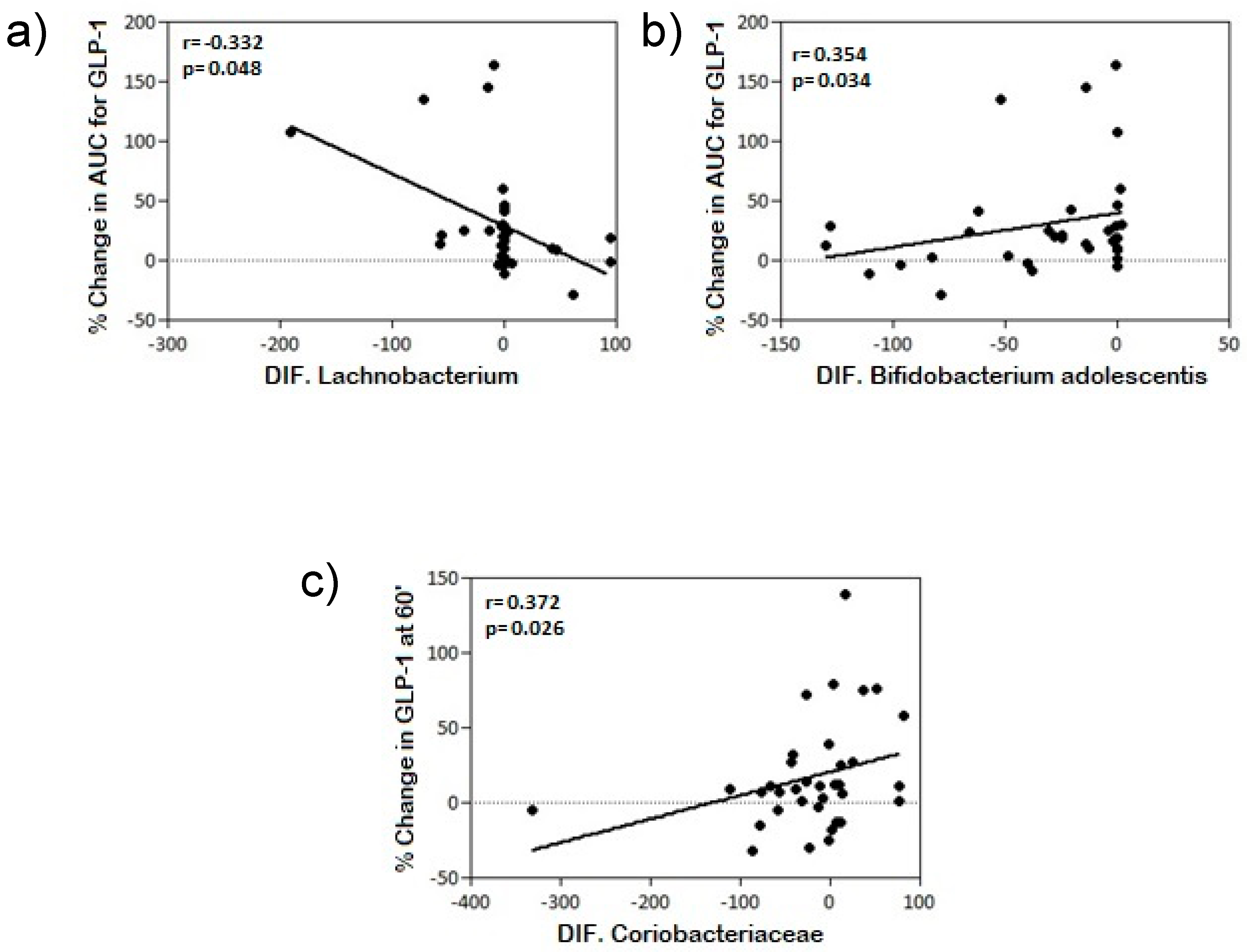
| PHYLA | FAMILY/GENUS/SPECIES | % Change AUC GLP-1 | % Change GLP-1 60’ |
|---|---|---|---|
| Actinobacteria differential | Coriobacteriaceae | NS | r = 0.372 * p = 0.026 |
| Bifidobacterium adolescentis | r = 0.354 * p = 0.034 | NS | |
| Firmicutes differential | Lachnobacterium | r = −0.332 * p = 0.048 | NS |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Cornejo-Pareja, I.; Martín-Núñez, G.M.; Roca-Rodríguez, M.M.; Cardona, F.; Coin-Aragüez, L.; Sánchez-Alcoholado, L.; Gutiérrez-Repiso, C.; Muñoz-Garach, A.; Fernández-García, J.C.; Moreno-Indias, I.; et al. H. pylori Eradication Treatment Alters Gut Microbiota and GLP-1 Secretion in Humans. J. Clin. Med. 2019, 8, 451. https://doi.org/10.3390/jcm8040451
Cornejo-Pareja I, Martín-Núñez GM, Roca-Rodríguez MM, Cardona F, Coin-Aragüez L, Sánchez-Alcoholado L, Gutiérrez-Repiso C, Muñoz-Garach A, Fernández-García JC, Moreno-Indias I, et al. H. pylori Eradication Treatment Alters Gut Microbiota and GLP-1 Secretion in Humans. Journal of Clinical Medicine. 2019; 8(4):451. https://doi.org/10.3390/jcm8040451
Chicago/Turabian StyleCornejo-Pareja, Isabel, Gracia M. Martín-Núñez, M. Mar Roca-Rodríguez, Fernando Cardona, Leticia Coin-Aragüez, Lidia Sánchez-Alcoholado, Carolina Gutiérrez-Repiso, Araceli Muñoz-Garach, José C. Fernández-García, Isabel Moreno-Indias, and et al. 2019. "H. pylori Eradication Treatment Alters Gut Microbiota and GLP-1 Secretion in Humans" Journal of Clinical Medicine 8, no. 4: 451. https://doi.org/10.3390/jcm8040451
APA StyleCornejo-Pareja, I., Martín-Núñez, G. M., Roca-Rodríguez, M. M., Cardona, F., Coin-Aragüez, L., Sánchez-Alcoholado, L., Gutiérrez-Repiso, C., Muñoz-Garach, A., Fernández-García, J. C., Moreno-Indias, I., & Tinahones, F. J. (2019). H. pylori Eradication Treatment Alters Gut Microbiota and GLP-1 Secretion in Humans. Journal of Clinical Medicine, 8(4), 451. https://doi.org/10.3390/jcm8040451






