Expression of EEF1A1 Is Associated with Prognosis of Patients with Colon Adenocarcinoma
Abstract
:1. Introduction
2. Methods
2.1. Identification of Prognostic Biomarker Genes Using the Wx Algorithm with TCGA Database
2.2. Patients and Tissue Samples
2.3. Quantitative Reverse Transcription PCR (qRT-PCR)
2.4. Tissue Microarray Construction and Immunohistochemistry
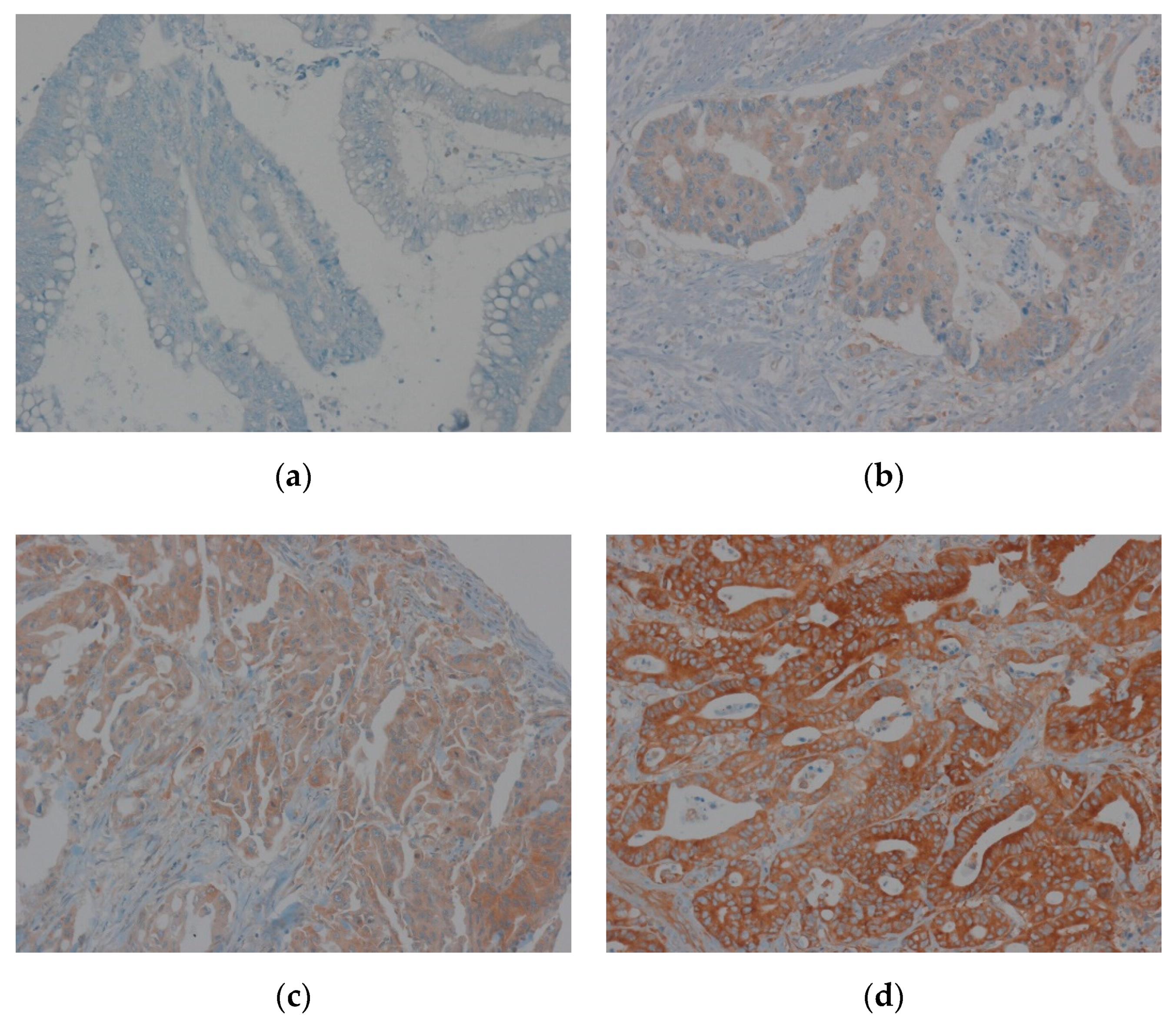
2.5. Evaluation of Immunohistochemical Staining of EEF1A1
2.6. Statistical Analysis
3. Results
3.1. Baseline Characteristics of the Study Population
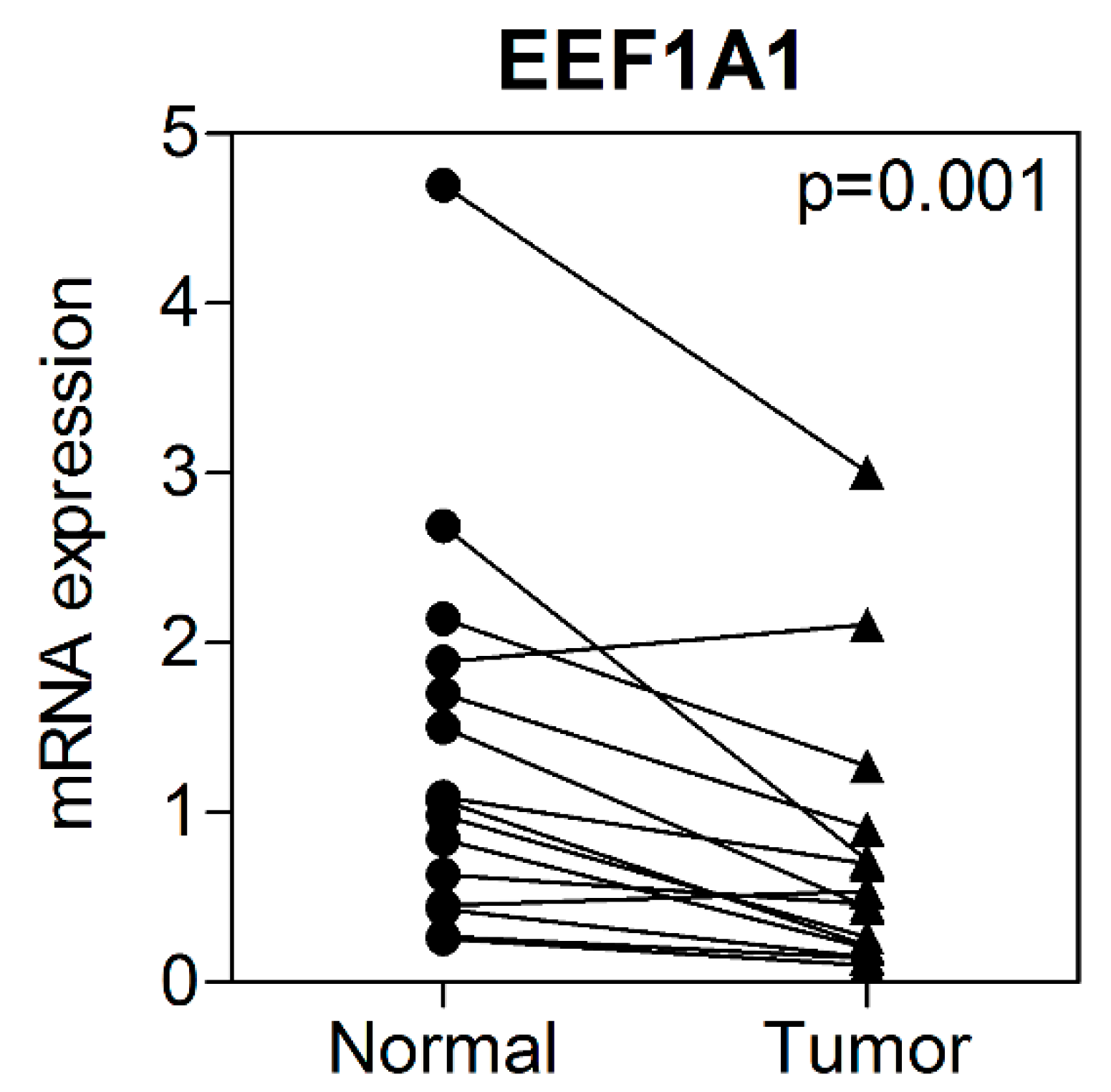
3.2. mRNA Expression of EEF1A1 in Human Colon Cancer Tissues Compared to That in Normal Adjacent Tissues
3.3. Correlation between EEF1A1 Expression and Clinicopathological Characteristics
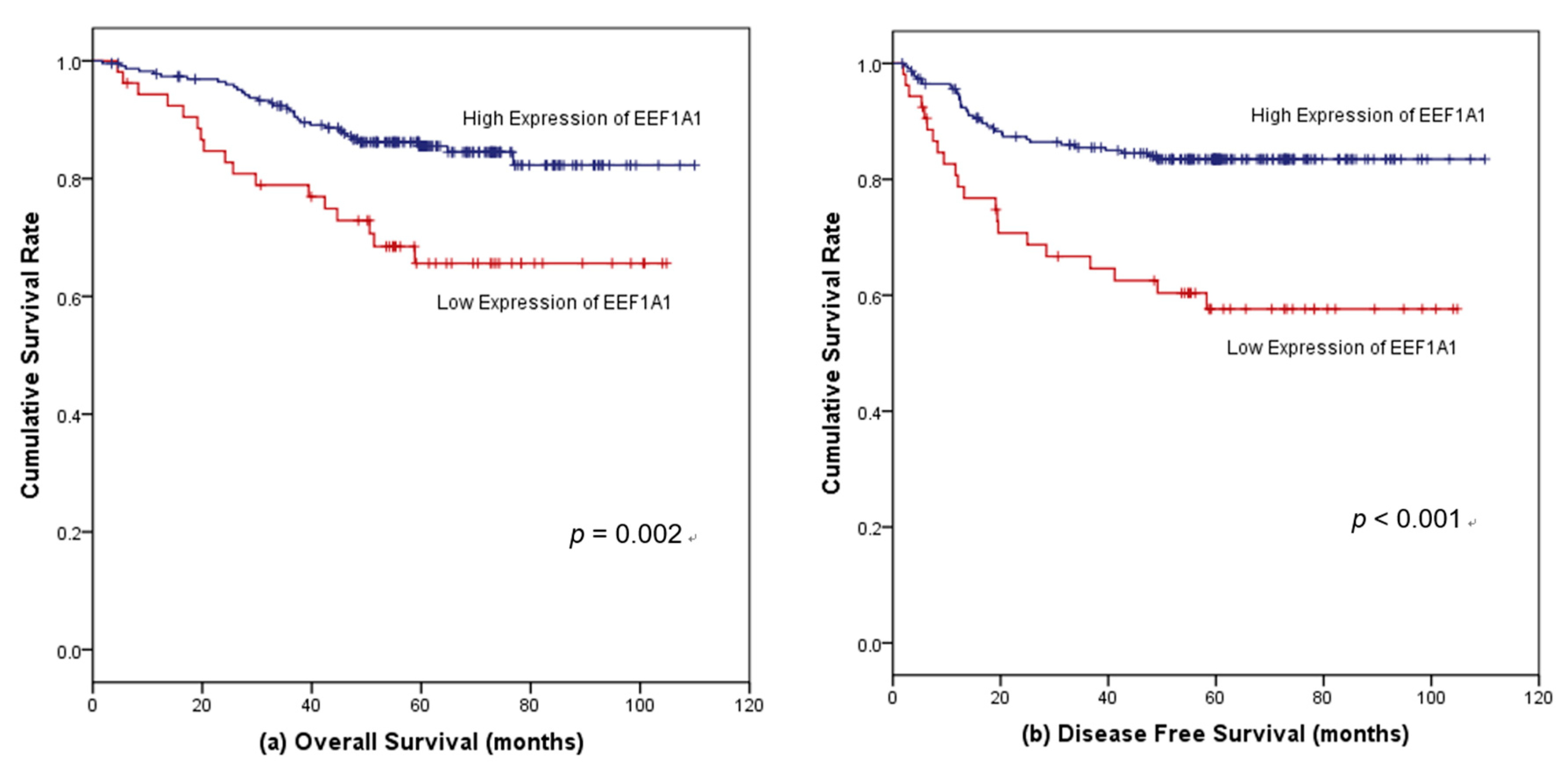
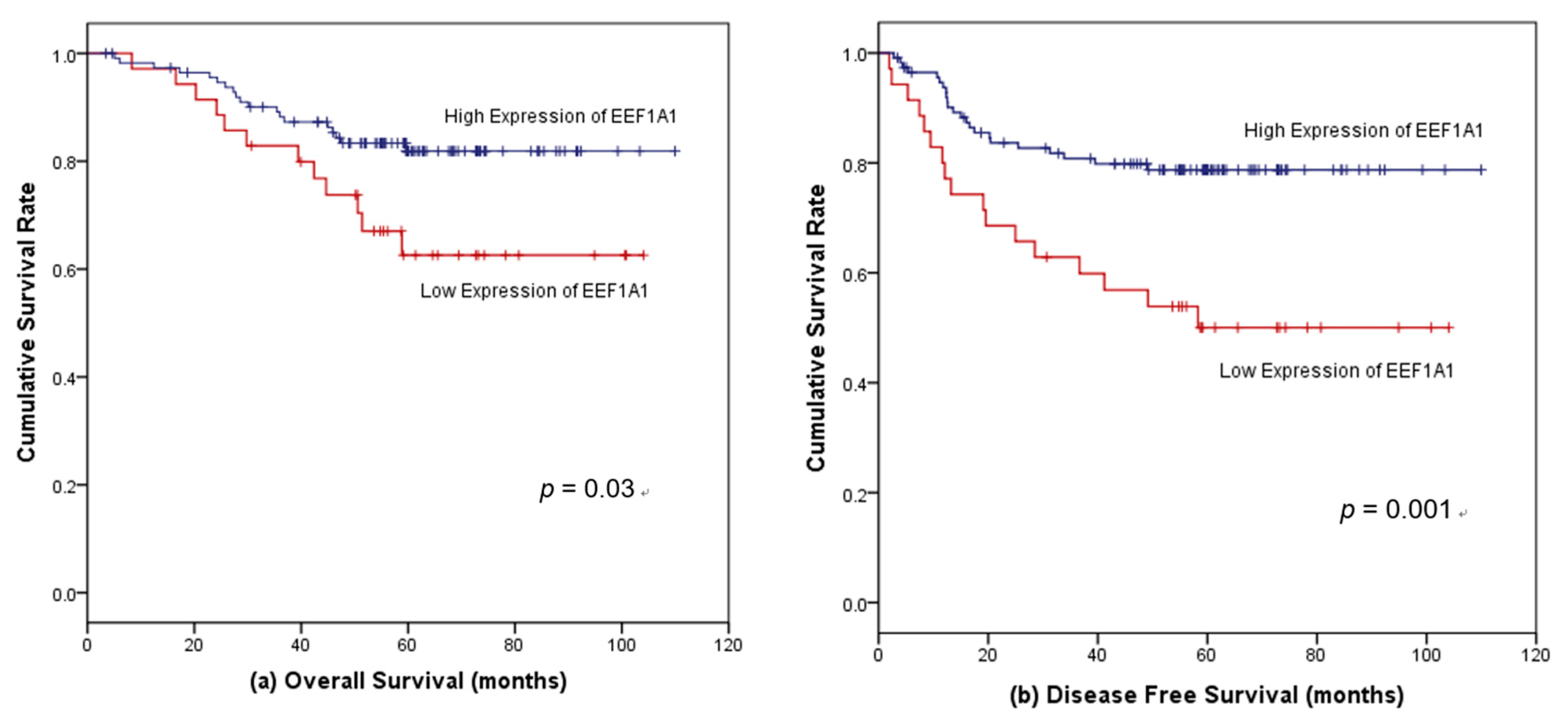
3.4. Survival Outcomes According to EEF1A1 Expression
3.5. Factors Associated with Survival in Patients with Stages II and III Colon Cancer
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Conflicts of Interest
References
- Bray, F.; Ferlay, J.; Soerjomataram, I.; Siegel, R.L.; Torre, L.A.; Jemal, A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 2018, 68, 394–424. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Puccini, A.; Berger, M.D.; Zhang, W.; Lenz, H.J. What We Know about Stage II and III Colon Cancer: It’s Still Not Enough. Target. Oncol. 2017, 12, 265–275. [Google Scholar] [CrossRef] [PubMed]
- Auclin, E.; Zaanan, A.; Vernerey, D.; Douard, R.; Gallois, C.; Laurent-Puig, P.; Bonnetain, F.; Taieb, J. Subgroups and prognostication in stage III colon cancer: Future perspectives for adjuvant therapy. Ann. Oncol. 2017, 28, 958–968. [Google Scholar] [CrossRef] [PubMed]
- Park, S.; Shin, B.; Shim, S.W.; Choi, Y.; Kang, K.; Kang, K. Wx: A neural network-based feature selection algorithm for transcriptomic data. Sci. Rep. 2019, 9, 10500. [Google Scholar] [CrossRef] [PubMed]
- Martinez-Ledesma, E.; Verhaak, R.G.; Treviño, V. Identification of a multi-cancer gene expression biomarker for cancer clinical outcomes using a network-based algorithm. Sci. Rep. 2015, 5, 11966. [Google Scholar] [CrossRef]
- Lee, S.; Francoeur, A.M.; Liu, S.; Wang, E. Tissue-specific expression in mammalian brain, heart, and muscle of S1, a member of the elongation factor-1 alpha gene family. J. Biol. Chem. 1992, 267, 24064–24068. [Google Scholar] [PubMed]
- Condeelis, J. Elongation factor 1 alpha, translation and the cytoskeleton. Trends Biochem. Sci. 1995, 20, 169–170. [Google Scholar] [CrossRef]
- Lund, A.; Knudsen, S.M.; Vissing, H.; Clark, B.; Tommerup, N. Assignment of human elongation factor 1alpha genes: EEF1A maps to chromosome 6q14 and EEF1A2 to 20q13.3. Genomics 1996, 36, 359–361. [Google Scholar] [CrossRef]
- Thornton, S.; Anand, N.; Purcell, D.; Lee, J. Not just for housekeeping: Protein initiation and elongation factors in cell growth and tumorigenesis. J. Mol. Med. 2003, 81, 536–548. [Google Scholar] [CrossRef]
- Abbas, W.; Kumar, A.; Herbein, G. The eEF1A Proteins: At the Crossroads of Oncogenesis, Apoptosis, and Viral Infections. Front. Oncol. 2015, 5, 75. [Google Scholar] [CrossRef]
- Anand, N.; Murthy, S.; Amann, G.; Wernick, M.; Porter, L.A.; Cukier, I.H.; Collins, C.; Gray, J.W.; Diebold, J.; Demetrick, D.J.; et al. Protein elongation factor EEF1A2 is a putative oncogene in ovarian cancer. Nat. Genet. 2002, 31, 301–305. [Google Scholar] [CrossRef] [PubMed]
- Ejiri, S. Moonlighting functions of polypeptide elongation factor 1: From actin bundling to zinc finger protein R1-associated nuclear localization. Biosci. Biotechnol. Biochem. 2002, 66, 1–21. [Google Scholar] [CrossRef] [PubMed]
- Lamberti, A.; Caraglia, M.; Longo, O.; Marra, M.; Abbruzzese, A.; Arcari, P. The translation elongation factor 1A in tumorigenesis, signal transduction and apoptosis: Review article. Amino Acids 2004, 26, 443–448. [Google Scholar] [CrossRef] [PubMed]
- Pinke, D.E.; Kalloger, S.E.; Francetic, T.; Huntsman, D.G.; Lee, J.M. The prognostic significance of elongation factor eEF1A2 in ovarian cancer. Gynecol. Oncol. 2008, 108, 561–568. [Google Scholar] [CrossRef] [PubMed]
- Kulkarni, G.; Turbin, D.A.; Amiri, A.; Jeganathan, S.; Andrade-Navarro, M.A.; Wu, T.D.; Huntsman, D.G.; Lee, J.M. Expression of protein elongation factor eEF1A2 predicts favorable outcome in breast cancer. Breast Cancer Res. Treat. 2007, 102, 31–41. [Google Scholar] [CrossRef] [PubMed]
- Pecorari, L.; Marin, O.; Silvestri, C.; Candini, O.; Rossi, E.; Guerzoni, C.; Cattelani, S.; Mariani, S.A.; Corradini, F.; Ferrari-Amorotti, G.; et al. Elongation Factor 1 alpha interacts with phospho-Akt in breast cancer cells and regulates their proliferation, survival and motility. Mol. Cancer 2009, 8, 58. [Google Scholar] [CrossRef]
- Tomlinson, V.A.; Newbery, H.J.; Wray, N.R.; Jackson, J.; Larionov, A.; Miller, W.R.; Dixon, J.M.; Abbott, C.M. Translation elongation factor eEF1A2 is a potential oncoprotein that is overexpressed in two-thirds of breast tumours. BMC Cancer 2005, 5, 113. [Google Scholar] [CrossRef]
- Kawamura, M.; Endo, C.; Sakurada, A.; Hoshi, F.; Notsuda, H.; Kondo, T. The prognostic significance of eukaryotic elongation factor 1 alpha-2 in non-small cell lung cancer. Anticancer Res. 2014, 34, 651–658. [Google Scholar]
- Cao, H.; Zhu, Q.; Huang, J.; Li, B.; Zhang, S.; Yao, W.; Zhang, Y. Regulation and functional role of eEF1A2 in pancreatic carcinoma. Biochem. Biophys. Res. Commun. 2009, 380, 11–16. [Google Scholar] [CrossRef]
- Duanmin, H.; Chao, X.; Qi, Z. eEF1A2 protein expression correlates with lymph node metastasis and decreased survival in pancreatic ductal adenocarcinoma. Hepatogastroenterology 2013, 60, 870–875. [Google Scholar] [CrossRef]
- Li, X.; Li, J.; Li, F. P21 activated kinase 4 binds translation elongation factor eEF1A1 to promote gastric cancer cell migration and invasion. Oncol. Rep. 2017, 37, 2857–2864. [Google Scholar] [CrossRef]
- Yang, S.; Lu, M.; Chen, Y.; Meng, D.; Sun, R.; Yun, D.; Zhao, Z.; Lu, D.; Li, Y. Overexpression of eukaryotic elongation factor 1 alpha-2 is associated with poorer prognosis in patients with gastric cancer. J. Cancer Res. Clin. Oncol. 2015, 141, 1265–1275. [Google Scholar] [CrossRef]
- Scaggiante, B.; Dapas, B.; Bonin, S.; Grassi, M.; Zennaro, C.; Farra, R.; Cristiano, L.; Siracusano, S.; Zanconati, F.; Giansante, C.; et al. Dissecting the expression of EEF1A1/2 genes in human prostate cancer cells: The potential of EEF1A2 as a hallmark for prostate transformation and progression. Br. J. Cancer 2012, 106, 166–173. [Google Scholar] [CrossRef]
- Chen, S.L.; Lu, S.X.; Liu, L.L.; Wang, C.H.; Yang, X.; Zhang, Z.Y.; Zhang, H.Z.; Yun, J.P. eEF1A1 Overexpression Enhances Tumor Progression and Indicates Poor Prognosis in Hepatocellular Carcinoma. Transl. Oncol. 2018, 11, 125–131. [Google Scholar] [CrossRef] [PubMed]
- Huang, J.; Zheng, C.; Shao, J.; Chen, L.; Liu, X.; Shao, J. Overexpression of eEF1A1 regulates G1-phase progression to promote HCC proliferation through the STAT1-cyclin D1 pathway. Biochem. Biophys. Res. Commun. 2017, 494, 542–549. [Google Scholar] [CrossRef]
- Ruest, L.B.; Marcotte, R.; Wang, E. Peptide elongation factor eEF1A-2/S1 expression in cultured differentiated myotubes and its protective effect against caspase-3-mediated apoptosis. J. Biol. Chem. 2002, 277, 5418–5425. [Google Scholar] [CrossRef]
- Borradaile, N.M.; Buhman, K.K.; Listenberger, L.L.; Magee, C.J.; Morimoto, E.T.; Ory, D.S.; Schaffer, J.E. A critical role for eukaryotic elongation factor 1A-1 in lipotoxic cell death. Mol. Biol. Cell 2006, 17, 770–778. [Google Scholar] [CrossRef]
- Bosutti, A.; Scaggiante, B.; Grassi, G.; Guarnieri, G.; Biolo, G. Overexpression of the elongation factor 1A1 relates to muscle proteolysis and proapoptotic p66(ShcA) gene transcription in hypercatabolic trauma patients. Metabolism 2007, 56, 1629–1634. [Google Scholar] [CrossRef]
- Liu, S.; Hausmann, S.; Carlson, S.M.; Fuentes, M.E.; Francis, J.W.; Pillai, R.; Lofgren, S.M.; Hulea, L.; Tandoc, K.; Lu, J.; et al. METTL13 Methylation of eEF1A Increases Translational Output to Promote Tumorigenesis. Cell 2019, 176, 491–504. [Google Scholar] [CrossRef] [PubMed]
- Lin, C.Y.; Beattie, A.; Baradaran, B.; Dray, E.; Duijf, P.H.G. Contradictory mRNA and protein misexpression of EEF1A1 in ductal breast carcinoma due to cell cycle regulation and cellular stress. Sci. Rep. 2018, 8, 13904. [Google Scholar] [CrossRef] [PubMed]
- Hassan, M.K.; Kumar, D.; Naik, M.; Dixit, M. The expression profile and prognostic significance of eukaryotic translation elongation factors in different cancers. PLoS ONE 2018, 13, e0191377. [Google Scholar] [CrossRef] [PubMed]
| EEF1A1-Low (n = 42) | EEF1A1-High (n = 239) | p-Value | |
|---|---|---|---|
| Age (years) | |||
| <65 | 22 (52.4%) | 113 (47.3%) | 0.616 |
| ≥65 | 20 (47.6%) | 126 (52.7%) | |
| Sex | |||
| Male | 26 (61.9%) | 129 (54.0%) | 0.402 |
| Female | 16 (38.1%) | 110 (46.0%) | |
| Body mass index (BMI) (kg/m2) | |||
| <18.5 | 2 (4.8%) | 17 (7.1%) | 0.839 |
| 18.5–25.0 | 23 (54.7%) | 124 (51.9%) | |
| >25.0 | 17 (40.5%) | 98 (41.0%) | |
| Number of comorbid diseases | |||
| 0 | 23 (54.8%) | 101 (42.3%) | 0.265 |
| 1 | 10 (23.8%) | 84 (35.1%) | |
| ≥2 | 9 (21.4%) | 54 (22.6%) | |
| Location of primary cancer | |||
| Left | 20 (47.6%) | 152 (63.6%) | 0.059 |
| Right | 22 (52.4%) | 87 (36.4%) | 0.059 |
| Pre-operative carcinoembryonic antigen (CEA) | |||
| <5 ng/mL | 27 (64.3%) | 151 (63.4%) | 1.000 |
| >5 ng/mL | 15 (35.7%) | 87 (36.6%) | |
| Bowel obstruction or perforation | |||
| No | 32 (76.2%) | 215 (90.0%) | 0.019 |
| Yes | 10 (23.8%) | 24 (10.0%) | |
| Adjuvant chemotherapy | |||
| No | 11 (76.2%) | 50 (20.9%) | 0.424 |
| Yes | 31 (73.8%) | 189 (79.1%) | |
| Stage (tumor/node/metastasis, TNM) | |||
| II | 15 (35.7%) | 117 (49.0%) | 0.132 |
| III | 27 (64.3%) | 122 (51.0%) | |
| pT stage | |||
| 2 | 1 (2.4%) | 10 (4.2%) | 0.097 |
| 3 | 30 (71.4%) | 197 (82.4%) | |
| 4 | 11 (26.2%) | 32 (13.4%) | |
| pN stage | 0.076 | ||
| 0 | 15 (35.7%) | 118 (49.4%) | |
| 1 | 18 (42.9%) | 62 (25.9%) | |
| 2 | 9 (21.4%) | 59 (24.7%) | |
| Differentiation | 0.319 | ||
| Well | 1 (2.4%) | 6 (2.5%) | |
| Moderate | 36 (87.8%) | 223 (93.3%) | |
| Poor | 4 (9.8%) | 10 (4.2%) | |
| Perineural invasion | 0.083 | ||
| No | 21 (50.0%) | 155 (64.9%) | |
| Yes | 21 (50.0%) | 84 (35.1%) | |
| Venous invasion | 0.503 | ||
| No | 37 (88.1%) | 199 (83.3%) | |
| Yes | 5 (11.9%) | 40 (16.7%) | |
| Lymphatic invasion | 0.610 | ||
| No | 27 (64.3%) | 141 (59.0%) | |
| Yes | 15 (35.7%) | 98 (41.0%) | |
| Number of harvested lymoh nodes | 23.5 (10–63) | 22 (4–115) | 0.259 |
| Number of metastatic lymph nodes | 1 (0–15) | 1 (0–18) | 0.560 |
| Metastatic lymphnodes/harvested lymph nodesratio | 4.9 (0–66.7) | 2.6 (0–100) | 0.943 |
| KRAS1 | 0.811 | ||
| Wild-type | 14 (66.7%) | 77 (63.1%) | |
| Mutant | 7 (33.3%) | 45 (36.9%) |
| Crude Hazard Ratio (95% CI) | p-Value | Adjusted Hazard Ratio (95% CI) | p-Value | |
|---|---|---|---|---|
| Age ≥ 65 (years) | 5.08 (2.468–10.462) | <0.001 | 5.93 (2.810–12.493) | <0.001 |
| Male | 1.31 (0.738–2.316) | 0.358 | ||
| BMI (reference: <18.5 kg/m2) | ||||
| 18.5–25.0 (kg/m2) | 1.17 (0.355–3.853) | 0.796 | ||
| >25.0 (kg/m2) | 0.98 (0.290–3.330) | 0.978 | ||
| Number of comorbid diseases (reference: 0) | ||||
| 1 | 1.02 (0.520–1.985) | 0.964 | ||
| ≥2 | 1.02 (0.776–2.963) | 0.223 | ||
| Right-sided location of primary cancer | 1.45 (0.827–2.527) | 0.196 | ||
| Pre-operative CEA ≥ 5 ng/mL | 1.794 (1.030–3.126) | 0.039 | 1.76 (1.008–3.027) | 0.058 |
| Combined bowel obstruction or perforation at initial diagnosis | 1.286 (0.578–2.859) | 0.537 | ||
| Adjuvant chemotherapy | 2.240 (1.247–4.024) | 0.007 | 0.034 | |
| Stage III | 1.509 (0.852–2.672) | 0.158 | ||
| Differentiation (reference: well) | ||||
| Moderate | 0.508 (0.123–2.096) | 0.349 | ||
| Poor | 0.531 (0.075–3.773) | 0.527 | ||
| Perineural invasion | 1.443 (0.827–2.516) | 0.197 | ||
| Venous invasion | 1.545 (0.791–3.017) | 0.203 | ||
| Lymphatic invasion | 1.542 (0.886–2.685) | 0.120 | ||
| Number of harvested lymph nodes | 0.996 (0.792–1.021) | 0.756 | ||
| Number of metastatic lymph nodes | 1.078 (1.004–1.156) | 0.037 | 1.350 (0.718–2.539) | 0.351 |
| KRAS1 mutated cancer | 2.020 (0.779–5.236) | 0.148 | ||
| Low EEF1A1 expression | 2.560 (1.380–4.749) | 0.003 | 3.01 (1.636–5.543) | <0.001 |
| Crude Hazard Ratio (95% CI) | p-Value | Adjusted Hazard Ratio (95% CI) | p-Value | |
|---|---|---|---|---|
| Age ≥ 65 (years-old) | 1.44 (0.852–2.442) | 0.172 | ||
| Male | 1.18 (0.697–2.008) | 0.533 | ||
| BMI (reference: <18.5 kg/m2) | 0.651 | |||
| 18.5–25.0 (kg/m2) | 0.69 (0.271–1.800) | 0.457 | ||
| >25.0 (kg/m2) | 0.98 (0.290–3.330) | 0.978 | ||
| Number of comorbid diseases (reference: 0) | ||||
| 1 | 1.04 (0.567–1.891) | 0.909 | ||
| ≥2 | 1.15 (0.593–2.216) | 0.685 | ||
| Right-sided location of primary cancer | 1.24 (0.730–2.103) | 0.428 | ||
| Pre-operative CEA ≥ 5 ng/mL | 2.35 (1.397–3.966) | 0.001 | 2.23 (1.308–3.806) | 0.003 |
| Combined bowel obstruction or perforation at initial diagnosis | 0.88 (0.379–2.057) | 0.773 | ||
| Adjuvant chemotherapy | 1.09 (0.578–2.066) | 0.785 | ||
| Stage III | 2.22 (1.256–3.908) | 0.006 | 25.80 (3.087–215.562) | 0.003 |
| Differentiation (reference: well) | 0.645 | |||
| Moderate | 0.51 (0.123–2.096) | 0.349 | ||
| Poor | 0.53 (0.075–3.773) | 0.527 | ||
| Perineural invasion | 1.34 (0.796–2.268) | 0.268 | ||
| Venous invasion | 1.19 (0.604–2.364) | 0.610 | ||
| Lymphatic invasion | 1.52 (0.901–2.546) | 0.117 | ||
| Number of harvested lymph nodes | 1.01 (1.002–1.026) | 0.021 | 1.02 (0.994–1.042) | 0.150 |
| Number of metastatic lymph nodes | 1.10 (1.034–1.171) | 0.003 | 0.24 (0.029–1.953) | 0.182 |
| KRAS 1 mutated cancer | 1.42 (0.665–3.036) | 0.364 | ||
| Low EEF1A1 expression | 2.86 (1.619–5.042) | <0.001 | 2.54 (1.459–4.434) | <0.001 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Joung, E.k.; Kim, J.; Yoon, N.; Maeng, L.-s.; Kim, J.H.; Park, S.; Kang, K.; Kim, J.S.; Ahn, Y.-H.; Ko, Y.H.; et al. Expression of EEF1A1 Is Associated with Prognosis of Patients with Colon Adenocarcinoma. J. Clin. Med. 2019, 8, 1903. https://doi.org/10.3390/jcm8111903
Joung Ek, Kim J, Yoon N, Maeng L-s, Kim JH, Park S, Kang K, Kim JS, Ahn Y-H, Ko YH, et al. Expression of EEF1A1 Is Associated with Prognosis of Patients with Colon Adenocarcinoma. Journal of Clinical Medicine. 2019; 8(11):1903. https://doi.org/10.3390/jcm8111903
Chicago/Turabian StyleJoung, Eun kyo, Jiyoung Kim, Nara Yoon, Lee-so Maeng, Ji Hoon Kim, Sungsoo Park, Keunsoo Kang, Jeong Seon Kim, Young-Ho Ahn, Yoon Ho Ko, and et al. 2019. "Expression of EEF1A1 Is Associated with Prognosis of Patients with Colon Adenocarcinoma" Journal of Clinical Medicine 8, no. 11: 1903. https://doi.org/10.3390/jcm8111903
APA StyleJoung, E. k., Kim, J., Yoon, N., Maeng, L.-s., Kim, J. H., Park, S., Kang, K., Kim, J. S., Ahn, Y.-H., Ko, Y. H., Byun, J. H., & Hong, J. H. (2019). Expression of EEF1A1 Is Associated with Prognosis of Patients with Colon Adenocarcinoma. Journal of Clinical Medicine, 8(11), 1903. https://doi.org/10.3390/jcm8111903






