MicroRNA Processing and Human Cancer
Abstract
:1. Introduction
2. MicroRNA Processing Machinery
3. Pri- to Pre-miRNA Processing in Cancer
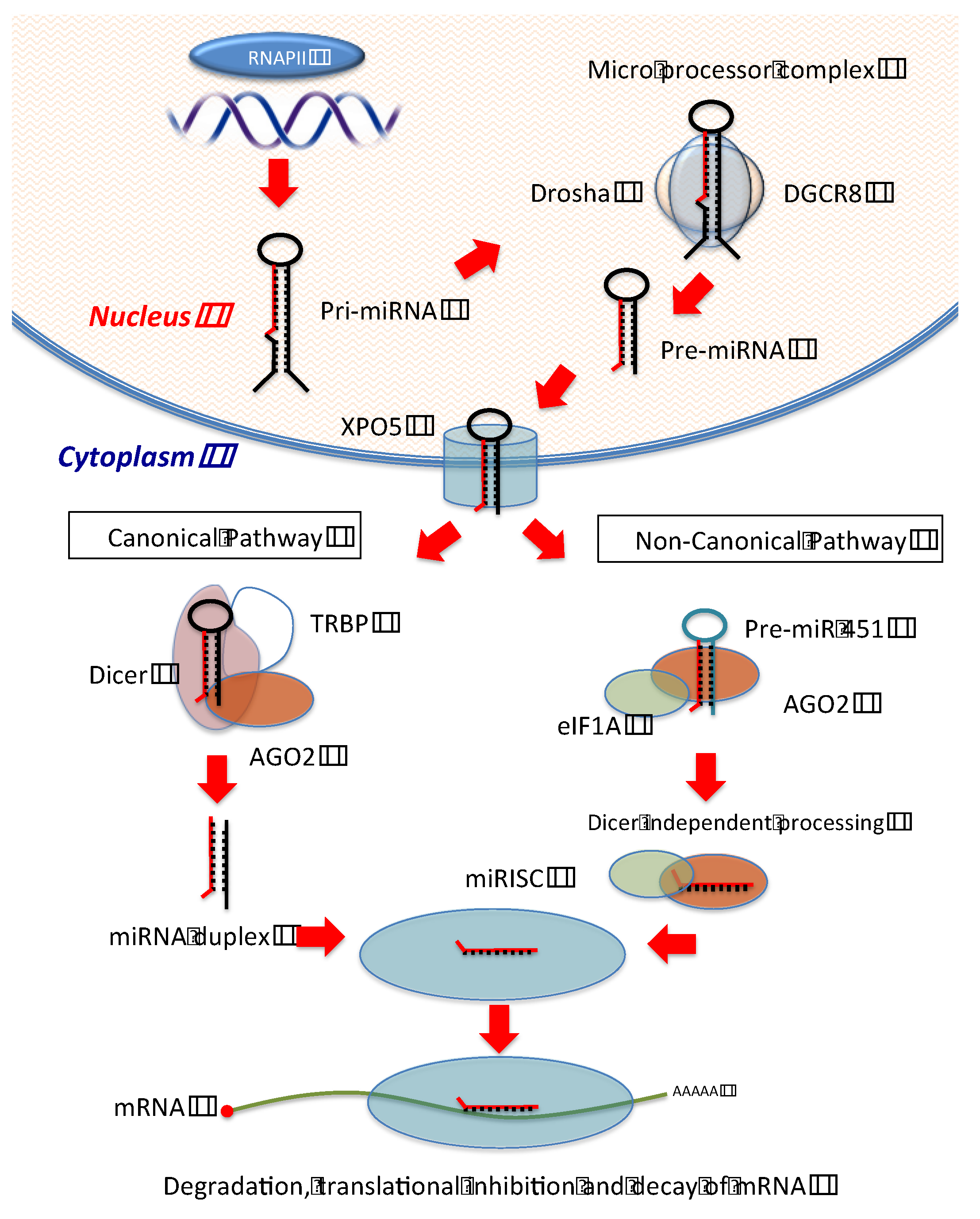
4. MiRNA Transportation and Cancer
5. Pre-miR to Mature MiRNA Processing and Cancer
| Location | MiRNAs Processing Related Factors | Clinical Relevance | Referrence |
|---|---|---|---|
| Nucleus | DGCR8 | Deletion of DGCR8 induces DiGeorge syndrome (22q.2 deletion syndrome). | [34] |
| Deletion of DGCR8 reduces stem cell development and cell proliferation in mice. | [35] | ||
| In colorectal cancer, DGCR8 expression is increased in tumors compared with normal tissue. | [36] | ||
| Down-regulation of DGCR8 enhances cellular transformation and tumor gurowth in lung cancer. | [19] | ||
| Drosha | Up-regulation of Drosha regulates cell proliferation; associated with poor prognosis of esophageal cancer and non-small cell lung cancer. | [37,38] | |
| Low expression of Drosha is associated with poor prognosis of ovarian cancer and neurobastoma. | [39,40] | ||
| Mutations of DGCR8 and Drosha | Together with the mutations in SIX1/2, mutations of DGCR8 and Drosha are associated with Wilms tumor. | [20,21] | |
| METTL3 | METTL3 regulates the recognition of stem-loop structure of pri-miRNAs by DGCR8. | [41] | |
| P68 and P72; RBPs | p68 and p72 are highly expressed in cancer and associated with miRNAs processing by Drosha. | [42,43,44,45,46] | |
| SMAD and SNIP1 | By regulating Drosha, SMAD and SNIP1 block the maturation of oncomiRs. | [47,48,49] | |
| KSRP, DDX1; DNA damage | ATM phosphorylation regulates the binding of KSRP and DDX1 to Drosha. | [53,54,55] | |
| Trans Nuclear Membrane | XP05 | XP05 increases the expression levels of tumor-suppressor miRNAs, indicating that XP05 has tumor-suppressive features. | [58,59] |
| RMND5A | RMNDA5A regulates XP05 stability together with miR-138. | [60,61] | |
| AKM-AKT signal; DNA damage | The activation of ATM-AKT signal after DNA damage, Nup153 binds to XP05, which induces nuclear export of pre-miRNAs. | [62] | |
| Cytoplasm | Dicer | High levels of Dicer expression are associated with god prognosis in ovarian cancer, breast cancer and CLL. | [39] |
| Up-regulation of Dicer is associated with poor progonosis in colorectal and prostate cancer. | [65,66,67] | ||
| Mutation of Dicer | Dicer mutation incuce a Dicer-related disorders including PPB. | [69,70] | |
| TRBP and TARBP2 | TRBP and TARBP2 destabilize Dicer, impairing miRNAs processing in human cancer. | [74,75] | |
| AG02 | AG02 regulates Dicer independent miRNA-451 through the non-canonical pathway. | [88,89] | |
| EGFR | In hypoxic condition, EGFR binds to AG02 and blocks miRNAs maturation. | [90,91] |
6. RISC-Related Defects in Cancer
7. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Orom, U.A.; Nielsen, F.C.; Lund, A.H. Microrna-10a binds the 5′utr of ribosomal protein mRNAs and enhances their translation. Mol. Cell 2008, 30, 460–471. [Google Scholar] [CrossRef] [PubMed]
- Baek, D.; Villen, J.; Shin, C.; Camargo, F.D.; Gygi, S.P.; Bartel, D.P. The impact of MicroRNAs on protein output. Nature 2008, 455, 64–71. [Google Scholar] [CrossRef] [PubMed]
- Selbach, M.; Schwanhausser, B.; Thierfelder, N.; Fang, Z.; Khanin, R.; Rajewsky, N. Widespread changes in protein synthesis induced by MicroRNAs. Nature 2008, 455, 58–63. [Google Scholar] [CrossRef] [PubMed]
- Kozomara, A.; Griffiths-Jones, S. Mirbase: Annotating high confidence MicroRNAs using deep sequencing data. Nucleic Acids Res. 2014, 42, D68–D73. [Google Scholar] [CrossRef] [PubMed]
- Krol, J.; Loedige, I.; Filipowicz, W. The widespread regulation of MicroRNA biogenesis, function and decay. Nat. Rev. Genet. 2010, 11, 597–610. [Google Scholar] [CrossRef] [PubMed]
- Huang, Y.; Shen, X.J.; Zou, Q.; Wang, S.P.; Tang, S.M.; Zhang, G.Z. Biological functions of MicroRNAs: A review. J. Physiol. Biochem. 2011, 67, 129–139. [Google Scholar] [CrossRef] [PubMed]
- Reinhart, B.J.; Slack, F.J.; Basson, M.; Pasquinelli, A.E.; Bettinger, J.C.; Rougvie, A.E.; Horvitz, H.R.; Ruvkun, G. The 21-nucleotide let-7 RNA regulates developmental timing in Caenorhabditis elegans. Nature 2000, 403, 901–906. [Google Scholar] [PubMed]
- Calin, G.A.; Dumitru, C.D.; Shimizu, M.; Bichi, R.; Zupo, S.; Noch, E.; Aldler, H.; Rattan, S.; Keating, M.; Rai, K.; et al. Frequent deletions and down-regulation of micro- RNA genes mir15 and mir16 at 13q14 in chronic lymphocytic leukemia. Proc. Natl. Acad. Sci. USA 2002, 99, 15524–15529. [Google Scholar] [CrossRef] [PubMed]
- Ota, A.; Tagawa, H.; Karnan, S.; Tsuzuki, S.; Karpas, A.; Kira, S.; Yoshida, Y.; Seto, M. Identification and characterization of a novel gene, c13orf25, as a target for 13q31-q32 amplification in malignant lymphoma. Cancer Res. 2004, 64, 3087–3095. [Google Scholar] [CrossRef] [PubMed]
- He, L.; Thomson, J.M.; Hemann, M.T.; Hernando-Monge, E.; Mu, D.; Goodson, S.; Powers, S.; Cordon-Cardo, C.; Lowe, S.W.; Hannon, G.J.; et al. A MicroRNA polycistron as a potential human oncogene. Nature 2005, 435, 828–833. [Google Scholar] [CrossRef] [PubMed]
- Patel, K.; Kollory, A.; Takashima, A.; Sarkar, S.; Faller, D.V.; Ghosh, S.K. MicroRNA let-7 downregulates stat3 phosphorylation in pancreatic cancer cells by increasing socs3 expression. Cancer Lett. 2014, 347, 54–64. [Google Scholar] [CrossRef] [PubMed]
- Lu, J.; Getz, G.; Miska, E.A.; Alvarez-Saavedra, E.; Lamb, J.; Peck, D.; Sweet-Cordero, A.; Ebert, B.L.; Mak, R.H.; Ferrando, A.A.; et al. MicroRNA expression profiles classify human cancers. Nature 2005, 435, 834–838. [Google Scholar] [CrossRef] [PubMed]
- Lee, E.J.; Baek, M.; Gusev, Y.; Brackett, D.J.; Nuovo, G.J.; Schmittgen, T.D. Systematic evaluation of MicroRNA processing patterns in tissues, cell lines, and tumors. RNA 2008, 14, 35–42. [Google Scholar] [CrossRef] [PubMed]
- Michael, M.Z.; SM, O.C.; van Holst Pellekaan, N.G.; Young, G.P.; James, R.J. Reduced accumulation of specific MicroRNAs in colorectal neoplasia. Mol. Cancer Res. 2003, 1, 882–891. [Google Scholar] [PubMed]
- Gaur, A.; Jewell, D.A.; Liang, Y.; Ridzon, D.; Moore, J.H.; Chen, C.; Ambros, V.R.; Israel, M.A. Characterization of MicroRNA expression levels and their biological correlates in human cancer cell lines. Cancer Res. 2007, 67, 2456–2468. [Google Scholar] [CrossRef] [PubMed]
- Visone, R.; Pallante, P.; Vecchione, A.; Cirombella, R.; Ferracin, M.; Ferraro, A.; Volinia, S.; Coluzzi, S.; Leone, V.; Borbone, E.; et al. Specific MicroRNAs are downregulated in human thyroid anaplastic carcinomas. Oncogene 2007, 26, 7590–7595. [Google Scholar] [CrossRef] [PubMed]
- Kim, M.S.; Oh, J.E.; Kim, Y.R.; Park, S.W.; Kang, M.R.; Kim, S.S.; Ahn, C.H.; Yoo, N.J.; Lee, S.H. Somatic mutations and losses of expression of MicroRNA regulation-related genes ago2 and tnrc6a in gastric and colorectal cancers. J. Pathol. 2010, 221, 139–146. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Huang, J.; Yang, N.; Greshock, J.; Megraw, M.S.; Giannakakis, A.; Liang, S.; Naylor, T.L.; Barchetti, A.; Ward, M.R.; et al. MicroRNAs exhibit high frequency genomic alterations in human cancer. Proc. Natl. Acad. Sci. USA 2006, 103, 9136–9141. [Google Scholar] [CrossRef] [PubMed]
- Kumar, M.S.; Lu, J.; Mercer, K.L.; Golub, T.R.; Jacks, T. Impaired MicroRNA processing enhances cellular transformation and tumorigenesis. Nat. Genet. 2007, 39, 673–677. [Google Scholar] [CrossRef] [PubMed]
- Wegert, J.; Ishaque, N.; Vardapour, R.; Georg, C.; Gu, Z.; Bieg, M.; Ziegler, B.; Bausenwein, S.; Nourkami, N.; Ludwig, N.; et al. Mutations in the six1/2 pathway and the drosha/dgcr8 miRNA microprocessor complex underlie high-risk blastemal type wilms tumors. Cancer Cell 2015, 27, 298–311. [Google Scholar] [CrossRef] [PubMed]
- Walz, A.L.; Ooms, A.; Gadd, S.; Gerhard, D.S.; Smith, M.A.; Guidry Auvil, J.M.; Meerzaman, D.; Chen, Q.R.; Hsu, C.H.; Yan, C.; et al. Recurrent dgcr8, drosha, and six homeodomain mutations in favorable histology wilms tumors. Cancer Cell 2015, 27, 286–297. [Google Scholar] [CrossRef] [PubMed]
- Bartel, D.P. MicroRNAs: Genomics, biogenesis, mechanism, and function. Cell 2004, 116, 281–297. [Google Scholar] [CrossRef]
- Kim, V.N. MicroRNA biogenesis: Coordinated cropping and dicing. Nat. Rev. Mol. Cell Biol. 2005, 6, 376–385. [Google Scholar] [CrossRef] [PubMed]
- Han, J.; Lee, Y.; Yeom, K.H.; Kim, Y.K.; Jin, H.; Kim, V.N. The drosha-dgcr8 complex in primary MicroRNA processing. Genes Dev. 2004, 18, 3016–3027. [Google Scholar] [CrossRef] [PubMed]
- Grund, S.E.; Polycarpou-Schwarz, M.; Luo, C.; Eichmuller, S.B.; Diederichs, S. Rare drosha splice variants are deficient in MicroRNA processing but do not affect general MicroRNA expression in cancer cells. Neoplasia 2012, 14, 238–248. [Google Scholar] [CrossRef] [PubMed]
- Yi, R.; Qin, Y.; Macara, I.G.; Cullen, B.R. Exportin-5 mediates the nuclear export of pre-MicroRNAs and short hairpin RNAs. Genes Dev. 2003, 17, 3011–3016. [Google Scholar] [CrossRef] [PubMed]
- Zeng, Y.; Cullen, B.R. Structural requirements for pre-MicroRNA binding and nuclear export by exportin 5. Nucleic Acids Res. 2004, 32, 4776–4785. [Google Scholar] [CrossRef] [PubMed]
- Chendrimada, T.P.; Gregory, R.I.; Kumaraswamy, E.; Norman, J.; Cooch, N.; Nishikura, K.; Shiekhattar, R. Trbp recruits the dicer complex to ago2 for MicroRNA processing and gene silencing. Nature 2005, 436, 740–744. [Google Scholar] [CrossRef] [PubMed]
- Feng, Y.; Zhang, X.; Graves, P.; Zeng, Y. A comprehensive analysis of precursor MicroRNA cleavage by human dicer. RNA 2012, 18, 2083–2092. [Google Scholar] [CrossRef] [PubMed]
- Maniataki, E.; Mourelatos, Z. A human, atp-independent, risc assembly machine fueled by pre-miRNA. Genes Dev. 2005, 19, 2979–2990. [Google Scholar] [CrossRef] [PubMed]
- Ghildiyal, M.; Zamore, P.D. Small silencing RNAs: An expanding universe. Nat. Rev. Genet. 2009, 10, 94–108. [Google Scholar] [CrossRef] [PubMed]
- Filipowicz, W.; Bhattacharyya, S.N.; Sonenberg, N. Mechanisms of post-transcriptional regulation by MicroRNAs: Are the answers in sight? Nat. Rev. Genet. 2008, 9, 102–114. [Google Scholar] [CrossRef] [PubMed]
- Marco, A.; Macpherson, J.I.; Ronshaugen, M.; Griffiths-Jones, S. MicroRNAs from the same precursor have different targeting properties. Silence 2012, 3, 8. [Google Scholar] [CrossRef] [PubMed]
- Shiohama, A.; Sasaki, T.; Noda, S.; Minoshima, S.; Shimizu, N. Molecular cloning and expression analysis of a novel gene dgcr8 located in the digeorge syndrome chromosomal region. Biochem. Biophys. Res. Commun. 2003, 304, 184–190. [Google Scholar] [CrossRef]
- Wang, Y.; Medvid, R.; Melton, C.; Jaenisch, R.; Blelloch, R. Dgcr8 is essential for MicroRNA biogenesis and silencing of embryonic stem cell self-renewal. Nat. Genet. 2007, 39, 380–385. [Google Scholar] [CrossRef] [PubMed]
- Kim, B.; Lee, J.H.; Park, J.W.; Kwon, T.K.; Baek, S.K.; Hwang, I.; Kim, S. An essential MicroRNA maturing microprocessor complex component dgcr8 is up-regulated in colorectal carcinomas. Clin. Exp. Med. 2014, 14, 331–336. [Google Scholar] [CrossRef] [PubMed]
- Sugito, N.; Ishiguro, H.; Kuwabara, Y.; Kimura, M.; Mitsui, A.; Kurehara, H.; Ando, T.; Mori, R.; Takashima, N.; Ogawa, R.; et al. Rnasen regulates cell proliferation and affects survival in esophageal cancer patients. Clin. Cancer Res. 2006, 12, 7322–7328. [Google Scholar] [CrossRef] [PubMed]
- Diaz-Garcia, C.V.; Agudo-Lopez, A.; Perez, C.; Lopez-Martin, J.A.; Rodriguez-Peralto, J.L.; de Castro, J.; Cortijo, A.; Martinez-Villanueva, M.; Iglesias, L.; Garcia-Carbonero, R.; et al. Dicer1, drosha and miRNAs in patients with non-small cell lung cancer: Implications for outcomes and histologic classification. Carcinogenesis 2013, 34, 1031–1038. [Google Scholar] [CrossRef] [PubMed]
- Merritt, W.M.; Lin, Y.G.; Han, L.Y.; Kamat, A.A.; Spannuth, W.A.; Schmandt, R.; Urbauer, D.; Pennacchio, L.A.; Cheng, J.F.; Nick, A.M.; et al. Dicer, drosha, and outcomes in patients with ovarian cancer. N. Engl. J. Med. 2008, 359, 2641–2650. [Google Scholar] [CrossRef] [PubMed]
- Lin, R.J.; Lin, Y.C.; Chen, J.; Kuo, H.H.; Chen, Y.Y.; Diccianni, M.B.; London, W.B.; Chang, C.H.; Yu, A.L. Microrna signature and expression of dicer and drosha can predict prognosis and delineate risk groups in neuroblastoma. Cancer Res. 2010, 70, 7841–7850. [Google Scholar] [CrossRef] [PubMed]
- Alarcon, C.R.; Lee, H.; Goodarzi, H.; Halberg, N.; Tavazoie, S.F. N6-methyladenosine marks primary MicroRNAs for processing. Nature 2015, 519, 482–485. [Google Scholar] [CrossRef] [PubMed]
- Shiohama, A.; Sasaki, T.; Noda, S.; Minoshima, S.; Shimizu, N. Nucleolar localization of dgcr8 and identification of eleven dgcr8-associated proteins. Exp. Cell Res. 2007, 313, 4196–4207. [Google Scholar] [CrossRef] [PubMed]
- Fuller-Pace, F.V.; Moore, H.C. RNA helicases p68 and p72: Multifunctional proteins with important implications for cancer development. Future Oncol. 2011, 7, 239–251. [Google Scholar] [CrossRef] [PubMed]
- Gregory, R.I.; Yan, K.P.; Amuthan, G.; Chendrimada, T.; Doratotaj, B.; Cooch, N.; Shiekhattar, R. The microprocessor complex mediates the genesis of MicroRNAs. Nature 2004, 432, 235–240. [Google Scholar] [CrossRef] [PubMed]
- Van Kouwenhove, M.; Kedde, M.; Agami, R. MicroRNA regulation by RNA-binding proteins and its implications for cancer. Nat. Rev. Cancer 2011, 11, 644–656. [Google Scholar] [CrossRef] [PubMed]
- Suzuki, H.I.; Yamagata, K.; Sugimoto, K.; Iwamoto, T.; Kato, S.; Miyazono, K. Modulation of MicroRNA processing by p53. Nature 2009, 460, 529–533. [Google Scholar] [CrossRef] [PubMed]
- Davis, B.N.; Hilyard, A.C.; Lagna, G.; Hata, A. Smad proteins control drosha-mediated MicroRNA maturation. Nature 2008, 454, 56–61. [Google Scholar] [CrossRef] [PubMed]
- Davis, B.N.; Hilyard, A.C.; Nguyen, P.H.; Lagna, G.; Hata, A. Smad proteins bind a conserved RNA sequence to promote MicroRNA maturation by drosha. Mol. Cell 2010, 39, 373–384. [Google Scholar] [CrossRef] [PubMed]
- Yu, B.; Bi, L.; Zheng, B.; Ji, L.; Chevalier, D.; Agarwal, M.; Ramachandran, V.; Li, W.; Lagrange, T.; Walker, J.C.; et al. The FHA domain proteins dawdle in arabidopsis and snip1 in humans act in small RNA biogenesis. Proc. Natl. Acad. Sci. USA 2008, 105, 10073–10078. [Google Scholar] [CrossRef] [PubMed]
- Diaz-Moreno, I.; Hollingworth, D.; Frenkiel, T.A.; Kelly, G.; Martin, S.; Howell, S.; Garcia-Mayoral, M.; Gherzi, R.; Briata, P.; Ramos, A. Phosphorylation-mediated unfolding of a KH domain regulates KSRP localization via 14–3-3 binding. Nat. Struct. Mol. Boil. 2009, 16, 238–246. [Google Scholar] [CrossRef] [PubMed]
- Banin, S.; Moyal, L.; Shieh, S.; Taya, Y.; Anderson, C.W.; Chessa, L.; Smorodinsky, N.I.; Prives, C.; Reiss, Y.; Shiloh, Y.; et al. Enhanced phosphorylation of p53 by atm in response to DNA damage. Science 1998, 281, 1674–1677. [Google Scholar] [CrossRef] [PubMed]
- Garcia-Mayoral, M.F.; Hollingworth, D.; Masino, L.; Diaz-Moreno, I.; Kelly, G.; Gherzi, R.; Chou, C.F.; Chen, C.Y.; Ramos, A. The structure of the C-terminal KH domains of KSRP reveals a noncanonical motif important for MRNA degradation. Structure 2007, 15, 485–498. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.; Wan, G.; Berger, F.G.; He, X.; Lu, X. The ATM kinase induces MicroRNA biogenesis in the DNA damage response. Mol. Cell 2011, 41, 371–383. [Google Scholar] [CrossRef] [PubMed]
- Yang, D.; Sun, Y.; Hu, L.; Zheng, H.; Ji, P.; Pecot, C.V.; Zhao, Y.; Reynolds, S.; Cheng, H.; Rupaimoole, R.; et al. Integrated analyses identify a master MicroRNA regulatory network for the mesenchymal subtype in serous ovarian cancer. Cancer Cell 2013, 23, 186–199. [Google Scholar] [CrossRef] [PubMed]
- Han, C.; Liu, Y.; Wan, G.; Choi, H.J.; Zhao, L.; Ivan, C.; He, X.; Sood, A.K.; Zhang, X.; Lu, X. The RNA-binding protein ddx1 promotes primary MicroRNA maturation and inhibits ovarian tumor progression. Cell Rep. 2014, 8, 1447–1460. [Google Scholar] [CrossRef] [PubMed]
- Urbach, A.; Yermalovich, A.; Zhang, J.; Spina, C.S.; Zhu, H.; Perez-Atayde, A.R.; Shukrun, R.; Charlton, J.; Sebire, N.; Mifsud, W.; et al. Lin28 sustains early renal progenitors and induces wilms tumor. Genes Dev. 2014, 28, 971–982. [Google Scholar] [CrossRef] [PubMed]
- Viswanathan, S.R.; Powers, J.T.; Einhorn, W.; Hoshida, Y.; Ng, T.L.; Toffanin, S.; O’Sullivan, M.; Lu, J.; Phillips, L.A.; Lockhart, V.L.; et al. Lin28 promotes transformation and is associated with advanced human malignancies. Nat. Genet. 2009, 41, 843–848. [Google Scholar] [CrossRef] [PubMed]
- Melo, S.A.; Moutinho, C.; Ropero, S.; Calin, G.A.; Rossi, S.; Spizzo, R.; Fernandez, A.F.; Davalos, V.; Villanueva, A.; Montoya, G.; et al. A genetic defect in exportin-5 traps precursor MicroRNAs in the nucleus of cancer cells. Cancer Cell 2010, 18, 303–315. [Google Scholar] [CrossRef] [PubMed]
- Melo, S.A.; Esteller, M. A precursor MicroRNA in a cancer cell nucleus: Get me out of here! Cell Cycle 2011, 10, 922–925. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Wang, C.; Chen, Z.; Jin, Y.; Wang, Y.; Kolokythas, A.; Dai, Y.; Zhou, X. Microrna-138 suppresses epithelial-mesenchymal transition in squamous cell carcinoma cell lines. Biochem. J. 2011, 440, 23–31. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Chen, Y.; Qin, X.; Wen, J.; Ding, H.; Xia, W.; Li, S.; Su, X.; Wang, W.; Li, H.; et al. Mir-138 downregulates miRNA processing in hela cells by targeting rmnd5a and decreasing exportin-5 stability. Nucleic Acids Res. 2014, 42, 458–474. [Google Scholar] [CrossRef] [PubMed]
- Wan, G.; Zhang, X.; Langley, R.R.; Liu, Y.; Hu, X.; Han, C.; Peng, G.; Ellis, L.M.; Jones, S.N.; Lu, X. DNA-damage-induced nuclear export of precursor MicroRNAs is regulated by the atm-akt pathway. Cell Rep. 2013, 3, 2100–2112. [Google Scholar] [CrossRef] [PubMed]
- Bernstein, E.; Caudy, A.A.; Hammond, S.M.; Hannon, G.J. Role for a bidentate ribonuclease in the initiation step of RNA interference. Nature 2001, 409, 363–366. [Google Scholar] [CrossRef] [PubMed]
- Hutvagner, G.; McLachlan, J.; Pasquinelli, A.E.; Balint, E.; Tuschl, T.; Zamore, P.D. A cellular function for the RNA-interference enzyme dicer in the maturation of the let-7 small temporal RNA. Science 2001, 293, 834–838. [Google Scholar] [CrossRef] [PubMed]
- Chiosea, S.; Jelezcova, E.; Chandran, U.; Acquafondata, M.; McHale, T.; Sobol, R.W.; Dhir, R. Up-regulation of dicer, a component of the MicroRNA machinery, in prostate adenocarcinoma. Am. J. Pathol. 2006, 169, 1812–1820. [Google Scholar] [CrossRef] [PubMed]
- Faber, C.; Horst, D.; Hlubek, F.; Kirchner, T. Overexpression of dicer predicts poor survival in colorectal cancer. Eur. J. Cancer 2011, 47, 1414–1419. [Google Scholar] [CrossRef] [PubMed]
- Stratmann, J.; Wang, C.J.; Gnosa, S.; Wallin, A.; Hinselwood, D.; Sun, X.F.; Zhang, H. Dicer and mirna in relation to clinicopathological variables in colorectal cancer patients. BMC Cancer 2011, 11, 345. [Google Scholar] [CrossRef] [PubMed]
- Zhu, D.X.; Fan, L.; Lu, R.N.; Fang, C.; Shen, W.Y.; Zou, Z.J.; Wang, Y.H.; Zhu, H.Y.; Miao, K.R.; Liu, P.; et al. Downregulated dicer expression predicts poor prognosis in chronic lymphocytic leukemia. Cancer Sci. 2012, 103, 875–881. [Google Scholar] [CrossRef] [PubMed]
- Hill, D.A.; Ivanovich, J.; Priest, J.R.; Gurnett, C.A.; Dehner, L.P.; Desruisseau, D.; Jarzembowski, J.A.; Wikenheiser-Brokamp, K.A.; Suarez, B.K.; Whelan, A.J.; et al. Dicer1 mutations in familial pleuropulmonary blastoma. Science 2009, 325, 965. [Google Scholar] [CrossRef] [PubMed]
- Doros, L.; Schultz, K.A.; Stewart, D.R.; Bauer, A.J.; Williams, G.; Rossi, C.T.; Carr, A.; Yang, J.; Dehner, L.P.; Messinger, Y.; et al. Dicer1-related disorders. GeneReviews® 2014, 1993–2015. [Google Scholar]
- Heyam, A.; Lagos, D.; Plevin, M. Dissecting the roles of TRBP and pact in double-stranded RNA recognition and processing of noncoding RNAs. Wiley Interdiscip Rev. RNA 2015, 6, 271–289. [Google Scholar] [CrossRef] [PubMed]
- Lee, Y.; Hur, I.; Park, S.Y.; Kim, Y.K.; Suh, M.R.; Kim, V.N. The role of pact in the RNA silencing pathway. EMBO J. 2006, 25, 522–532. [Google Scholar] [CrossRef] [PubMed]
- Lee, H.Y.; Zhou, K.; Smith, A.M.; Noland, C.L.; Doudna, J.A. Differential roles of human dicer-binding proteins TRBP and pact in small RNA processing. Nucleic Acids Res. 2013, 41, 6568–6576. [Google Scholar] [CrossRef] [PubMed]
- Melo, S.A.; Ropero, S.; Moutinho, C.; Aaltonen, L.A.; Yamamoto, H.; Calin, G.A.; Rossi, S.; Fernandez, A.F.; Carneiro, F.; Oliveira, C.; et al. A tarbp2 mutation in human cancer impairs microrna processing and dicer1 function. Nat. Genet. 2009, 41, 365–370. [Google Scholar] [CrossRef] [PubMed]
- De Vito, C.; Riggi, N.; Cornaz, S.; Suva, M.L.; Baumer, K.; Provero, P.; Stamenkovic, I. A tarbp2-dependent miRNA expression profile underlies cancer stem cell properties and provides candidate therapeutic reagents in ewing sarcoma. Cancer Cell 2012, 21, 807–821. [Google Scholar] [CrossRef] [PubMed]
- Bohmert, K.; Camus, I.; Bellini, C.; Bouchez, D.; Caboche, M.; Benning, C. Ago1 defines a novel locus of arabidopsis controlling leaf development. EMBO J. 1998, 17, 170–180. [Google Scholar] [CrossRef] [PubMed]
- Van Grootheest, D.S.; van den Berg, S.M.; Cath, D.C.; Willemsen, G.; Boomsma, D.I. Marital resemblance for obsessive-compulsive, anxious and depressive symptoms in a population-based sample. Psychol. Med. 2008, 38, 1731–1740. [Google Scholar] [CrossRef] [PubMed]
- Hutvagner, G.; Simard, M.J. Argonaute proteins: Key players in RNA silencing. Nat. Rev. Mol. Cell Biol. 2008, 9, 22–32. [Google Scholar] [CrossRef] [PubMed]
- Song, J.J.; Smith, S.K.; Hannon, G.J.; Joshua-Tor, L. Crystal structure of argonaute and its implications for risc slicer activity. Science 2004, 305, 1434–1437. [Google Scholar] [CrossRef] [PubMed]
- Frank, F.; Sonenberg, N.; Nagar, B. Structural basis for 5′-nucleotide base-specific recognition of guide RNA by human ago2. Nature 2010, 465, 818–822. [Google Scholar] [CrossRef] [PubMed]
- Yan, K.S.; Yan, S.; Farooq, A.; Han, A.; Zeng, L.; Zhou, M.M. Structure and conserved RNA binding of the Paz domain. Nature 2003, 426, 468–474. [Google Scholar] [CrossRef] [PubMed]
- Parker, J.S.; Roe, S.M.; Barford, D. Structural insights into MRNA recognition from a piwi domain-siRNA guide complex. Nature 2005, 434, 663–666. [Google Scholar] [CrossRef] [PubMed]
- Ma, J.B.; Yuan, Y.R.; Meister, G.; Pei, Y.; Tuschl, T.; Patel, D.J. Structural basis for 5′-end-specific recognition of guide RNA by the a. Fulgidus piwi protein. Nature 2005, 434, 666–670. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Carmell, M.A.; Rivas, F.V.; Marsden, C.G.; Thomson, J.M.; Song, J.J.; Hammond, S.M.; Joshua-Tor, L.; Hannon, G.J. Argonaute2 is the catalytic engine of mammalian RNAI. Science 2004, 305, 1437–1441. [Google Scholar] [CrossRef] [PubMed]
- Meister, G.; Landthaler, M.; Patkaniowska, A.; Dorsett, Y.; Teng, G.; Tuschl, T. Human argonaute2 mediates RNA cleavage targeted by miRNAs and siRNAs. Mol. Cell 2004, 15, 185–197. [Google Scholar] [CrossRef] [PubMed]
- Diederichs, S.; Haber, D.A. Dual role for argonautes in MicroRNA processing and posttranscriptional regulation of MicroRNA expression. Cell 2007, 131, 1097–1108. [Google Scholar] [CrossRef] [PubMed]
- Winter, J.; Diederichs, S. Argonaute proteins regulate microrna stability: Increased MicroRNA abundance by argonaute proteins is due to MicroRNA stabilization. RNA Biol. 2011, 8, 1149–1157. [Google Scholar] [CrossRef] [PubMed]
- Yi, T.; Arthanari, H.; Akabayov, B.; Song, H.; Papadopoulos, E.; Qi, H.H.; Jedrychowski, M.; Guttler, T.; Guo, C.; Luna, R.E.; et al. Eif1a augments ago2-mediated dicer-independent miRNA biogenesis and RNA interference. Nat. Commun. 2015, 6, 7194. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.S.; Lai, E.C. Alternative miRNA biogenesis pathways and the interpretation of core miRNA pathway mutants. Mol. Cell 2011, 43, 892–903. [Google Scholar] [CrossRef] [PubMed]
- Shen, J.; Xia, W.; Khotskaya, Y.B.; Huo, L.; Nakanishi, K.; Lim, S.O.; Du, Y.; Wang, Y.; Chang, W.C.; Chen, C.H.; et al. Egfr modulates MicroRNA maturation in response to hypoxia through phosphorylation of ago2. Nature 2013, 497, 383–387. [Google Scholar] [CrossRef] [PubMed]
- Wu, C.; So, J.; Davis-Dusenbery, B.N.; Qi, H.H.; Bloch, D.B.; Shi, Y.; Lagna, G.; Hata, A. Hypoxia potentiates MicroRNA-mediated gene silencing through posttranslational modification of argonaute2. Mol. Cell. Biol. 2011, 31, 4760–4774. [Google Scholar] [CrossRef] [PubMed]
- Ling, H.; Fabbri, M.; Calin, G.A. MicroRNAs and other non-coding RNAs as targets for anticancer drug development. Nat. Rev. Drug Discov. 2013, 12, 847–865. [Google Scholar] [CrossRef] [PubMed]
- Machlin, E.S.; Sarnow, P.; Sagan, S.M. Masking the 5′ terminal nucleotides of the hepatitis C virus genome by an unconventional MicroRNA-target RNA complex. Proc. Natl. Acad. Sci. USA 2011, 108, 3193–3198. [Google Scholar] [CrossRef] [PubMed]
- Janssen, H.L.; Reesink, H.W.; Lawitz, E.J.; Zeuzem, S.; Rodriguez-Torres, M.; Patel, K.; van der Meer, A.J.; Patick, A.K.; Chen, A.; Zhou, Y.; et al. Treatment of HCV infection by targeting MicroRNA. N. Engl. J. Med. 2013, 368, 1685–1694. [Google Scholar] [CrossRef] [PubMed]
- Valadi, H.; Ekstrom, K.; Bossios, A.; Sjostrand, M.; Lee, J.J.; Lotvall, J.O. Exosome-mediated transfer of mRNAs and MicroRNAs is a novel mechanism of genetic exchange between cells. Nat. Cell Biol. 2007, 9, 654–659. [Google Scholar] [CrossRef] [PubMed]
- Fleischhacker, M.; Schmidt, B. Circulating nucleic acids (cnas) and cancer—A survey. Biochim. Biophys. Acta 2007, 1775, 181–232. [Google Scholar] [CrossRef] [PubMed]
- Skog, J.; Wurdinger, T.; van Rijn, S.; Meijer, D.H.; Gainche, L.; Sena-Esteves, M.; Curry, W.T., Jr.; Carter, B.S.; Krichevsky, A.M.; Breakefield, X.O. Glioblastoma microvesicles transport rna and proteins that promote tumour growth and provide diagnostic biomarkers. Nat. Cell Biol. 2008, 10, 1470–1476. [Google Scholar] [CrossRef] [PubMed]
- Melo, S.A.; Sugimoto, H.; O’Connell, J.T.; Kato, N.; Villanueva, A.; Vidal, A.; Qiu, L.; Vitkin, E.; Perelman, L.T.; Melo, C.A.; et al. Cancer exosomes perform cell-independent microrna biogenesis and promote tumorigenesis. Cancer Cell 2014, 26, 707–721. [Google Scholar] [CrossRef] [PubMed]
- Kim, D.H.; Behlke, M.A.; Rose, S.D.; Chang, M.S.; Choi, S.; Rossi, J.J. Synthetic dsrna dicer substrates enhance rnai potency and efficacy. Nat. Biotechnol. 2005, 23, 222–226. [Google Scholar] [CrossRef] [PubMed]
- Amarzguioui, M.; Lundberg, P.; Cantin, E.; Hagstrom, J.; Behlke, M.A.; Rossi, J.J. Rational design and in vitro and in vivo delivery of dicer substrate siRNA. Nat. Protoc. 2006, 1, 508–517. [Google Scholar] [CrossRef] [PubMed]
- Dore-Savard, L.; Roussy, G.; Dansereau, M.A.; Collingwood, M.A.; Lennox, K.A.; Rose, S.D.; Beaudet, N.; Behlke, M.A.; Sarret, P. Central delivery of dicer-substrate sirna: A direct application for pain research. Mol. Ther. 2008, 16, 1331–1339. [Google Scholar] [CrossRef] [PubMed]
- Dudek, H.; Wong, D.H.; Arvan, R.; Shah, A.; Wortham, K.; Ying, B.; Diwanji, R.; Zhou, W.; Holmes, B.; Yang, H.; et al. Knockdown of beta-catenin with dicer-substrate siRNAs reduces liver tumor burden in vivo. Mol. Ther. 2014, 22, 92–101. [Google Scholar]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ohtsuka, M.; Ling, H.; Doki, Y.; Mori, M.; Calin, G.A. MicroRNA Processing and Human Cancer. J. Clin. Med. 2015, 4, 1651-1667. https://doi.org/10.3390/jcm4081651
Ohtsuka M, Ling H, Doki Y, Mori M, Calin GA. MicroRNA Processing and Human Cancer. Journal of Clinical Medicine. 2015; 4(8):1651-1667. https://doi.org/10.3390/jcm4081651
Chicago/Turabian StyleOhtsuka, Masahisa, Hui Ling, Yuichiro Doki, Masaki Mori, and George Adrian Calin. 2015. "MicroRNA Processing and Human Cancer" Journal of Clinical Medicine 4, no. 8: 1651-1667. https://doi.org/10.3390/jcm4081651
APA StyleOhtsuka, M., Ling, H., Doki, Y., Mori, M., & Calin, G. A. (2015). MicroRNA Processing and Human Cancer. Journal of Clinical Medicine, 4(8), 1651-1667. https://doi.org/10.3390/jcm4081651






