MicroRNA Expression Relating to Dietary-Induced Liver Steatosis and NASH
Abstract
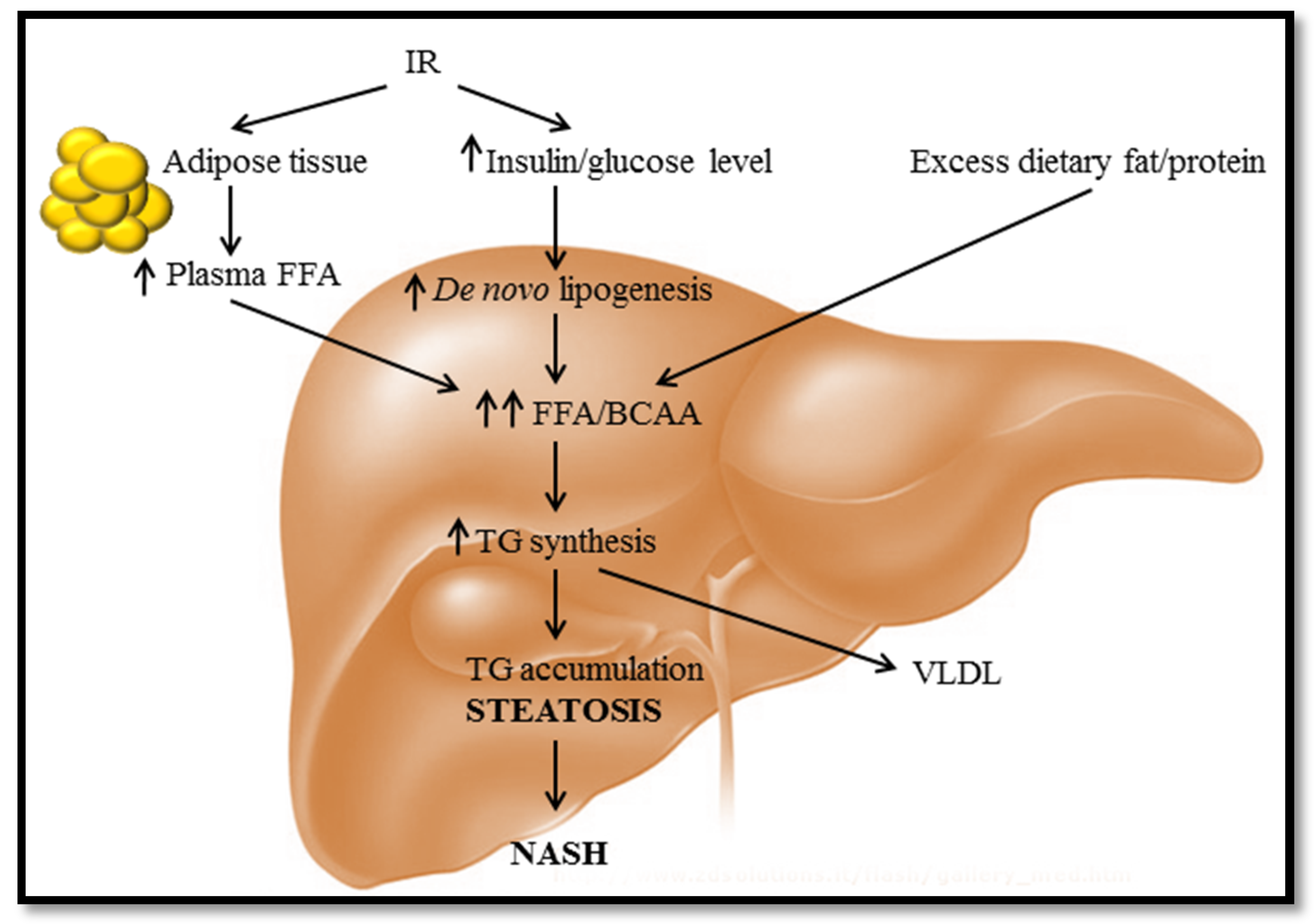
:1. Diet-Induced Fatty Liver Disease
2. MicroRNAs in Metabolic Syndrome and NAFLD
3. The Clinical Relevance of Exosomal Signalling in NAFLD
| Biological Samples | miRNA(s) | Outcome | Target * | References |
|---|---|---|---|---|
| Plasma | miR-122 | Inhibition of miR-122 reduced hepatic cholesterol and FA biosynthesis and elevated FA oxidation in humans | SREBP-1c, SREBP-2 | [26,27,32] |
| Intracellular | miR-122, miR-451, miR-27a | Downregulated in the liver of high fat/fructose-fed rats | miR-451→NFκB | [34,74,75] |
| miR-27a→PPARγ | ||||
| Plasma/Intracellular | miR-33a/b, miR-143, miR-92b | Inhibition of miR-33a/b increased HDL and lowers VLDL | SREBP-1, SREBP-2, AMPKα, IRS2, MSTN, FOXO1 | [29,30,55] |
| Overexpression of miR-33a/b increased HDL and reduces VLDL | ||||
| Plasma/Intracellular | miR-34a, miR-16, miR-21, miR-27b, miR-122, miR-192, miR-375, miR-19a/b, miR-146b, miR-181a, miR-200 | Overexpressed in circulation of steatotic, NASH and T2DM humans/rats | miR-34a→p53 | [32,33,61,62,76,77,78,79] |
| miR-21→HMG-CoA | ||||
| miR-27b→ PPARγ | ||||
| miR-146b→IL-6, TNF-α | ||||
| Intracellular | miR-200a/b, miR-429 | Upregulated in the liver of high-fat/fructose-fed rats | [34] | |
| Intracellular | miR-29c | Downregulated in the liver of ob/ob mice on a lipogenic diet | [35] | |
| Plasma | miR-192, miR-375, miR-122 | Increased particularly in NASH, suppression of glucose-induced insulin secretion | miR-375→HMG-CoA | [32,62,80] |
| Plasma | miR-155, miR-122 | Upregulated in rats with ASH | miR-155→LXRα | [67,81] |
4. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Kopelman, P.G. Obesity as a medical problem. Nature 2000, 404, 635–643. [Google Scholar] [PubMed]
- Mokdad, A.H.; Ford, E.S.; Bowman, B.A.; Dietz, W.H.; Vinicor, F.; Bales, V.S.; Marks, J.S. Prevalence of obesity, diabetes, and obesity-related health risk factors, 2001. JAMA 2003, 289, 76–79. [Google Scholar] [CrossRef] [PubMed]
- Spiegelman, B.M.; Flier, J.S. Obesity and the regulation of energy balance. Cell 2001, 104, 531–543. [Google Scholar] [CrossRef]
- Goldman, L.; Ausiello, D. Cecil Textbook of Medicine; Arjomand, M.; Setudenia, A.H.; Qasemi, Sh., Translators; Nasle Farda: Tehran, Iran, 2003; Volume 11. [Google Scholar]
- Lara-Castro, C.; Garvey, W.T. Intracellular lipid accumulation in liver and muscle and the insulin resistance syndrome. Endocrinol. Metab. Clin. N. Am. 2008, 37, 841–856. [Google Scholar] [CrossRef] [PubMed]
- Bullock, R.E.; Zaitoun, A.M.; Aithal, G.P.; Ryder, S.D.; Beckingham, I.J.; Lobo, D.N. Association of non-alcoholic steatohepatitis without significant fibrosis with hepatocellular carcinoma. J. Hepatol. 2004, 41, 685–686. [Google Scholar] [CrossRef] [PubMed]
- Marrero, J.A.; Fontana, R.J.; Su, G.L.; Conjeevaram, H.S.; Emick, D.M.; Lok, A.S. NAFLD may be a common underlying liver disease in patients with hepatocellular carcinoma in the United States. Hepatology 2002, 36, 1349–1354. [Google Scholar] [CrossRef] [PubMed]
- Rottiers, V.; Näär, A.M. MicroRNAs in metabolism and metabolic disorders. Nat. Rev. Mol. Cell Biol. 2012, 13, 239–250. [Google Scholar] [CrossRef] [PubMed]
- Yahagi, N.; Shimano, H.; Matsuzaka, T.; Sekiya, M.; Najima, Y.; Okazaki, S.; Okazaki, H.; Tamura, Y.; Iizuka, Y.; Inoue, N. p53 involvement in the pathogenesis of fatty liver disease. J. Biol. Chem. 2004, 279, 20571–20575. [Google Scholar] [CrossRef] [PubMed]
- Sethi, J.K.; Vidal-Puig, A.J. Thematic review series: Adipocyte biology. Adipose tissue function and plasticity orchestrate nutritional adaptation. J. Lipid Res. 2007, 48, 1253–1262. [Google Scholar] [CrossRef] [PubMed]
- Stefan, N.; Kantartzis, K.; Machann, J.; Schick, F.; Thamer, C.; Rittig, K.; Balletshofer, B.; Machicao, F.; Fritsche, A.; Häring, H.-U. Identification and characterization of metabolically benign obesity in humans. Arch. Intern. Med. 2008, 168, 1609–1616. [Google Scholar] [CrossRef] [PubMed]
- Fabbrini, E.; Magkos, F.; Mohammed, B.S.; Pietka, T.; Abumrad, N.A.; Patterson, B.W.; Okunade, A.; Klein, S. Intrahepatic fat, not visceral fat, is linked with metabolic complications of obesity. Proc. Natl. Acad. Sci. USA 2009, 106, 15430–15435. [Google Scholar] [CrossRef] [PubMed]
- Misu, H.; Takamura, T.; Takayama, H.; Hayashi, H.; Matsuzawa-Nagata, N.; Kurita, S.; Ishikura, K.; Ando, H.; Takeshita, Y.; Ota, T. A liver-derived secretory protein, selenoprotein P causes insulin resistance. Cell Metab. 2010, 12, 483–495. [Google Scholar] [CrossRef] [PubMed]
- Yang, S.; Hwang, S.; Choi, H.; Yoo, H.; Seo, J.; Kim, S.; Kim, N.; Baik, S.; Choi, D.; Choi, K.; Choi, K. Serum selenoprotein P levels in patients with type 2 diabetes and prediabetes: Implications for insulin resistance, inflammation, and atherosclerosis. J. Clin. Endocrinol. Metab. 2011, 96, E1325–E1329. [Google Scholar] [CrossRef] [PubMed]
- Misu, H.; Ishikura, K.; Kurita, S.; Takeshita, Y.; Ota, T.; Saito, Y.; Takahashi, K.; Kaneko, S.; Takamura, T. Inverse correlation between serum levels of selenoprotein P and adiponectin in patients with type 2 diabetes. PLoS ONE 2012, 7, e34952. [Google Scholar] [CrossRef] [PubMed]
- Williams, M.D.; Mitchell, G.M. MicroRNAs in insulin resistance and obesity. Exp. Diabetes Res. 2012, 2012. [Google Scholar] [CrossRef] [PubMed]
- Etheridge, A.; Lee, I.; Hood, L.; Galas, D.; Wang, K. Extracellular microRNA: A new source of biomarkers. Mutat. Res. Fundam. Mol. Mech. Mutagen. 2011, 717, 85–90. [Google Scholar] [CrossRef] [PubMed]
- Aravin, A.A.; Lagos-Quintana, M.; Yalcin, A.; Zavolan, M.; Marks, D.; Snyder, B.; Gaasterland, T.; Meyer, J.; Tuschl, T. The small RNA profile during Drosophila melanogaster development. Dev. Cell 2003, 5, 337–350. [Google Scholar] [CrossRef]
- Ketting, R.F. MicroRNA biogenesis and function. In Regulation of microRNAs; Springer: Basel, Switzerland, 2010; pp. 1–14. [Google Scholar]
- Lim, L.P.; Lau, N.C.; Garrett-Engele, P.; Grimson, A.; Schelter, J.M.; Castle, J.; Bartel, D.P.; Linsley, P.S.; Johnson, J.M. Microarray analysis shows that some microRNAs downregulate large numbers of target mRNAs. Nature 2005, 433, 769–773. [Google Scholar] [CrossRef] [PubMed]
- Pillai, R.S.; Bhattacharyya, S.N.; Artus, C.G.; Zoller, T.; Cougot, N.; Basyuk, E.; Bertrand, E.; Filipowicz, W. Inhibition of translational initiation by Let-7 microRNA in human cells. Science 2005, 309, 1573–1576. [Google Scholar] [CrossRef] [PubMed]
- Valinezhad Orang, A.; Safaralizadeh, R.; Kazemzadeh-Bavili, M. Mechanisms of miRNA-mediated gene regulation from common downregulation to mRNA-specific upregulation. Int. J. Genomics 2014, 2014. [Google Scholar] [CrossRef] [PubMed]
- Gauthier, B.R.; Wollheim, C.B. MicroRNAs: “ribo-regulators” of glucose homeostasis. Nat. Med. 2006, 12, 36–38. [Google Scholar] [CrossRef] [PubMed]
- He, A.; Zhu, L.; Gupta, N.; Chang, Y.; Fang, F. Overexpression of micro ribonucleic acid 29, highly up-regulated in diabetic rats, leads to insulin resistance in 3T3-L1 adipocytes. Mol. Endocrinol. 2007, 21, 2785–2794. [Google Scholar] [CrossRef] [PubMed]
- Poy, M.; Spranger, M.; Stoffel, M. microRNAs and the regulation of glucose and lipid metabolism. Diabetes Obes. Metab. 2007, 9, 67–73. [Google Scholar] [CrossRef] [PubMed]
- Esau, C.; Davis, S.; Murray, S.F.; Yu, X.X.; Pandey, S.K.; Pear, M.; Watts, L.; Booten, S.L.; Graham, M.; McKay, R. miR-122 regulation of lipid metabolism revealed by in vivo antisense targeting. Cell Metab. 2006, 3, 87–98. [Google Scholar] [CrossRef] [PubMed]
- Krützfeldt, J.; Rajewsky, N.; Braich, R.; Rajeev, K.G.; Tuschl, T.; Manoharan, M.; Stoffel, M. Silencing of microRNAs in vivo with “antagomirs”. Nature 2005, 438, 685–689. [Google Scholar] [CrossRef] [PubMed]
- Rottiers, V.; Najafi-Shoushtari, S.H.; Kristo, F.; Gurumurthy, S.; Zhong, L.; Li, Y.; et al. MicroRNAs in metabolism and metabolic diseases. Cold Spring Harb. Symp. Quant. Biol. 2011, 76, 225–233. [Google Scholar] [CrossRef] [PubMed]
- Dávalos, A.; Goedeke, L.; Smibert, P.; Ramírez, C.M.; Warrier, N.P.; Andreo, U.; Cirera-Salinas, D.; Rayner, K.; Suresh, U.; Pastor-Pareja, J.C. miR-33a/b contribute to the regulation of fatty acid metabolism and insulin signaling. Proc. Natl. Acad. Sci. USA 2011, 108, 9232–9237. [Google Scholar] [CrossRef] [PubMed]
- Rayner, K.J.; Esau, C.C.; Hussain, F.N.; McDaniel, A.L.; Marshall, S.M.; van Gils, J.M.; Ray, T.D.; Sheedy, F.J.; Goedeke, L.; Liu, X. Inhibition of miR-33a/b in non-human primates raises plasma HDL and lowers VLDL triglycerides. Nature 2011, 478, 404–407. [Google Scholar] [CrossRef] [PubMed]
- Trajkovski, M.; Hausser, J.; Soutschek, J.; Bhat, B.; Akin, A.; Zavolan, M.; Heim, M.H.; Stoffel, M. MicroRNAs 103 and 107 regulate insulin sensitivity. Nature 2011, 474, 649–653. [Google Scholar] [CrossRef] [PubMed]
- Cheung, O.; Puri, P.; Eicken, C.; Contos, M.J.; Mirshahi, F.; Maher, J.W.; Kellum, J.M.; Min, H.; Luketic, V.A.; Sanyal, A.J. Nonalcoholic steatohepatitis is associated with altered hepatic MicroRNA expression. Hepatology 2008, 48, 1810–1820. [Google Scholar] [CrossRef] [PubMed]
- Kong, L.; Zhu, J.; Han, W.; Jiang, X.; Xu, M.; Zhao, Y.; Dong, Q.; Pang, Z.; Guan, Q.; Gao, L. Significance of serum microRNAs in pre-diabetes and newly diagnosed type 2 diabetes: A clinical study. Acta Diabetol. 2011, 48, 61–69. [Google Scholar] [CrossRef] [PubMed]
- Alisi, A.; da Sacco, L.; Bruscalupi, G.; Piemonte, F.; Panera, N.; De Vito, R.; Leoni, S.; Bottazzo, G.F.; Masotti, A.; Nobili, V. Mirnome analysis reveals novel molecular determinants in the pathogenesis of diet-induced nonalcoholic fatty liver disease. Lab. Investig. 2011, 91, 283–293. [Google Scholar] [CrossRef] [PubMed]
- Pogribny, I.P.; Starlard-Davenport, A.; Tryndyak, V.P.; Han, T.; Ross, S.A.; Rusyn, I.; Beland, F.A. Difference in expression of hepatic microRNAs miR-29c, miR-34a, miR-155, and miR-200b is associated with strain-specific susceptibility to dietary nonalcoholic steatohepatitis in mice. Lab. Investig. 2010, 90, 1437–1446. [Google Scholar] [CrossRef] [PubMed]
- Li, S.; Chen, X.; Zhang, H.; Liang, X.; Xiang, Y.; Yu, C.; Zen, K.; Li, Y.; Zhang, C.-Y. Differential expression of microRNAs in mouse liver under aberrant energy metabolic status. J. Lipid Res. 2009, 50, 1756–1765. [Google Scholar] [CrossRef] [PubMed]
- Castro, R.E.; Ferreira, D.M.; Afonso, M.B.; Borralho, P.M.; Machado, M.V.; Cortez-Pinto, H.; Rodrigues, C.M. miR-34a/SIRT1/p53 is suppressed by ursodeoxycholic acid in the rat liver and activated by disease severity in human non-alcoholic fatty liver disease. J. Hepatol. 2013, 58, 119–125. [Google Scholar] [CrossRef] [PubMed]
- Kemper, J.K.; Xiao, Z.; Ponugoti, B.; Miao, J.; Fang, S.; Kanamaluru, D.; Tsang, S.; Wu, S.-Y.; Chiang, C.-M.; Veenstra, T.D. FXR acetylation is normally dynamically regulated by p300 and SIRT1 but constitutively elevated in metabolic disease states. Cell Metab. 2009, 10, 392–404. [Google Scholar] [CrossRef] [PubMed]
- Motta, M.C.; Divecha, N.; Lemieux, M.; Kamel, C.; Chen, D.; Gu, W.; Bultsma, Y.; McBurney, M.; Guarente, L. Mammalian SIRT1 represses forkhead transcription factors. Cell 2004, 116, 551–563. [Google Scholar] [CrossRef]
- Ponugoti, B.; Kim, D.-H.; Xiao, Z.; Smith, Z.; Miao, J.; Zang, M.; Wu, S.-Y.; Chiang, C.-M.; Veenstra, T.D.; Kemper, J.K. SIRT1 deacetylates and inhibits SREBP-1C activity in regulation of hepatic lipid metabolism. J. Biol. Chem. 2010, 285, 33959–33970. [Google Scholar] [CrossRef] [PubMed]
- Nakanishi, N.; Nakagawa, Y.; Tokushige, N.; Aoki, N.; Matsuzaka, T.; Ishii, K.; Yahagi, N.; Kobayashi, K.; Yatoh, S.; Takahashi, A. The up-regulation of microRNA-335 is associated with lipid metabolism in liver and white adipose tissue of genetically obese mice. Biochem. Biophys. Res. Commun. 2009, 385, 492–496. [Google Scholar] [CrossRef] [PubMed]
- Dohi, O.; Yasui, K.; Gen, Y.; Takada, H.; Endo, M.; Tsuji, K.; Konishi, C.; Yamada, N.; Mitsuyoshi, H.; Yagi, N. Epigenetic silencing of miR-335 and its host gene MEST in hepatocellular carcinoma. Int. J. Oncol. 2013, 42, 411–418. [Google Scholar] [PubMed]
- Takahashi, M.; Kamei, Y.; Ezaki, O. Mest/Peg1 imprinted gene enlarges adipocytes and is a marker of adipocyte size. Am. J. Physiol. Endocrinol. Metab. 2005, 288, E117–E124. [Google Scholar] [CrossRef] [PubMed]
- Zhu, L.; Chen, L.; Shi, C.-M.; Xu, G.-F.; Xu, L.-L.; Zhu, L.-L.; Ni, Y.; Cui, Y.; Ji, C. MiR-335, an adipogenesis-related microRNA, is involved in adipose tissue inflammation. Cell Biochem. Biophys. 2014, 68, 283–290. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.; Wu, C.-Q.; Zhang, Z.-Q.; Yao, D.-K.; Zhu, L. Loss of expression of miR-335 is implicated in hepatic stellate cell migration and activation. Exp. Cell Res. 2011, 317, 1714–1725. [Google Scholar] [CrossRef] [PubMed]
- Zarfeshani, A.; Ngo, S.; Murphy, R.; Sheppard, A.M. MiRNAs as bariatric surgery outcome biomarkers and hepatokine regulators after GBP but not SG. Mol. End. 2015. Submitted. [Google Scholar]
- Pal, D.; Dasgupta, S.; Kundu, R.; Maitra, S.; Das, G.; Mukhopadhyay, S.; Ray, S.; Majumdar, S.S.; Bhattacharya, S. Fetuin-A acts as an endogenous ligand of TLR4 to promote lipid-induced insulin resistance. Nat. Med. 2012, 18, 1279–1285. [Google Scholar] [CrossRef] [PubMed]
- Jüllig, M.; Yip, S.; Xu, A.; Smith, G.; Middleditch, M.; Booth, M.; Babor, R.; Beban, G.; Murphy, R. Lower fetuin-A retinol binding protein 4 and several metabolites after gastric bypass compared to sleeve gastrectomy in patients with type 2 diabetes. PLoS ONE 2014, 9, e96489. [Google Scholar] [CrossRef] [PubMed]
- Hittel, D.S.; Berggren, J.R.; Shearer, J.; Boyle, K.; Houmard, J.A. Increased secretion and expression of myostatin in skeletal muscle from extremely obese women. Diabetes 2009, 58, 30–38. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Cao, L.; Ye, J.; Zhu, D. Upregulation of myostatin gene expression in streptozotocin-induced type 1 diabetes mice is attenuated by insulin. Biochem. Biophys. Res. Commun. 2009, 388, 112–116. [Google Scholar] [CrossRef] [PubMed]
- Milan, G.; Dalla Nora, E.; Pilon, C.; Pagano, C.; Granzotto, M.; Manco, M.; Mingrone, G.; Vettor, R. Changes in muscle myostatin expression in obese subjects after weight loss. J. Clin. Endocrinol. Metab. 2004, 89, 2724–2727. [Google Scholar] [CrossRef] [PubMed]
- Schuelke, M.; Wagner, K.R.; Stolz, L.E.; Hübner, C.; Riebel, T.; Kömen, W.; Braun, T.; Tobin, J.F.; Lee, S.-J. Myostatin mutation associated with gross muscle hypertrophy in a child. N. Engl. J. Med. 2004, 350, 2682–2688. [Google Scholar] [CrossRef] [PubMed]
- Zhang, C.; McFarlane, C.; Lokireddy, S.; Masuda, S.; Ge, X.; Gluckman, P.D.; Sharma, M.; Kambadur, R. Inhibition of myostatin protects against diet-induced obesity by enhancing fatty acid oxidation and promoting a brown adipose phenotype in mice. Diabetologia 2012, 55, 183–193. [Google Scholar] [CrossRef] [PubMed]
- Zhao, B.; Wall, R.J.; Yang, J. Transgenic expression of myostatin propeptide prevents diet-induced obesity and insulin resistance. Biochem. Biophys. Res. Commun. 2005, 337, 248–255. [Google Scholar] [CrossRef] [PubMed]
- Zarfeshani, A.; Ngo, S.; Sheppard, A.M. Leucine alters hepatic glucose/lipid homeostasis via the myostatin-AMP-activated protein kinase pathway-potential implications for nonalcoholic fatty liver disease. Clin. Epigenet. 2014, 6, 27. [Google Scholar] [CrossRef] [PubMed]
- Lake, A.D.; Novak, P.; Shipkova, P.; Aranibar, N.; Robertson, D.G.; Reily, M.D.; Lehman-McKeeman, L.D.; Vaillancourt, R.R.; Cherrington, N.J. Branched chain amino acid metabolism profiles in progressive human nonalcoholic fatty liver disease. Amino Acids 2015, 47, 603–615. [Google Scholar] [CrossRef] [PubMed]
- Castera, L.; Vilgrain, V.; Angulo, P. Noninvasive evaluation of NAFLD. Nat. Rev. Gastroenterol. Hepatol. 2013, 10, 666–675. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.-K. Exosomes and Microvesicles as Biomarkers in Metabolic Diseases. Korean J. Obes. 2014, 23, 150–155. [Google Scholar] [CrossRef]
- Pirola, C.J.; Gianotti, T.F.; Castaño, G.O.; Mallardi, P.; San Martino, J.; Ledesma, M.M.G.L.; Flichman, D.; Mirshahi, F.; Sanyal, A.J.; Sookoian, S. Circulating microRNA signature in non-alcoholic fatty liver disease: From serum non-coding RNAs to liver histology and disease pathogenesis. Gut 2014. [Google Scholar] [CrossRef] [PubMed]
- McArdle, M.A.; Finucane, O.M.; Connaughton, R.M.; McMorrow, A.M.; Roche, H.M. Mechanisms of obesity-induced inflammation and insulin resistance: Insights into the emerging role of nutritional strategies. Front. Endocrinol. 2013, 4. [Google Scholar] [CrossRef] [PubMed]
- Gori, M.; Arciello, M.; Balsano, C. MicroRNAs in nonalcoholic fatty liver disease: Novel biomarkers and prognostic tools during the transition from steatosis to hepatocarcinoma. BioMed. Res. Int. 2014, 2014. [Google Scholar] [CrossRef] [PubMed]
- Tryndyak, V.P.; Latendresse, J.R.; Montgomery, B.; Ross, S.A.; Beland, F.A.; Rusyn, I.; Pogribny, I.P. Plasma microRNAs are sensitive indicators of inter-strain differences in the severity of liver injury induced in mice by a choline-and folate-deficient diet. Toxicol. Appl. Pharmacol. 2012, 262, 52–59. [Google Scholar] [CrossRef] [PubMed]
- Celikbilek, M.; Baskol, M.; Taheri, S.; Deniz, K.; Dogan, S.; Zararsiz, G.; Gursoy, S.; Guven, K.; Ozbakır, O.; Dundar, M.; Yucesoy, M. Circulating microRNAs in patients with non-alcoholic fatty liver disease. World J. Hepatol. 2014, 6, 613. [Google Scholar] [PubMed]
- Gallo, A.; Tandon, M.; Alevizos, I.; Illei, G.G. The majority of microRNAs detectable in serum and saliva is concentrated in exosomes. PLoS ONE 2012, 7, e30679. [Google Scholar] [CrossRef] [PubMed]
- Arroyo, J.D.; Chevillet, J.R.; Kroh, E.M.; Ruf, I.K.; Pritchard, C.C.; Gibson, D.F.; Mitchell, P.S.; Bennett, C.F.; Pogosova-Agadjanyan, E.L.; Stirewalt, D.L.; et al. Argonaute2 complexes carry a population of circulating microRNAs independent of vesicles in human plasma. Proc. Natl. Acad. Sci. USA 2011, 108, 5003–5008. [Google Scholar] [CrossRef] [PubMed]
- Berger, F.; Reiser, M.F. Micro-RNAs as potential new molecular biomarkers in oncology: Have they reached relevance for the clinical imaging sciences? Theranostics 2013, 3, 943. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.-P.; Jin, X.; Xiang, Z.; Chen, S.-H.; Li, Y.-M. Circulating MicroRNAs as potential biomarkers for alcoholic steatohepatitis. Liver Int. 2013, 33, 1257–1265. [Google Scholar] [CrossRef] [PubMed]
- Boeri, M.; Verri, C.; Conte, D.; Roz, L.; Modena, P.; Facchinetti, F.; Calabrò, E.; Croce, C.M.; Pastorino, U.; Sozzi, G. MicroRNA signatures in tissues and plasma predict development and prognosis of computed tomography detected lung cancer. Proc. Natl. Acad. Sci. USA 2011, 108, 3713–3718. [Google Scholar] [CrossRef] [PubMed]
- Lu, Z.; Liu, M.; Stribinskis, V.; Klinge, C.; Ramos, K.; Colburn, N.; Li, Y. MicroRNA-21 promotes cell transformation by targeting the programmed cell death 4 gene. Oncogene 2008, 27, 4373–4379. [Google Scholar] [CrossRef] [PubMed]
- Morrissey, D.V.; Lockridge, J.A.; Shaw, L.; Blanchard, K.; Jensen, K.; Breen, W.; Hartsough, K.; Polisky, B. Potent and persistent in vivo anti-HBV activity of chemically modified siRNAs. Nat. Biotechnol. 2005, 23, 1002–1007. [Google Scholar] [CrossRef] [PubMed]
- Wolfrum, C.; Shi, S.; Jayaprakash, K.N.; Jayaraman, M.; Wang, G.; Pandey, R.K.; Rajeev, K.G.; Nakayama, T.; Charrise, K.; Ndungo, E.M. Mechanisms and optimization of in vivo delivery of lipophilic siRNAs. Nat. Biotechnol. 2007, 25, 1149–1157. [Google Scholar] [CrossRef] [PubMed]
- Lanford, R.E.; Hildebrandt-Eriksen, E.S.; Petri, A.; Persson, R.; Lindow, M.; Munk, M.E.; Kauppinen, S.; Ørum, H. Therapeutic silencing of microRNA-122 in primates with chronic hepatitis C virus infection. Science 2010, 327, 198–201. [Google Scholar] [CrossRef] [PubMed]
- Janssen, H.L.; Reesink, H.W.; Lawitz, E.J.; Zeuzem, S.; Rodriguez-Torres, M.; Patel, K.; van der Meer, A.J.; Patick, A.K.; Chen, A.; Zhou, Y. Treatment of HCV infection by targeting microRNA. N. Engl. J. Med. 2013, 368, 1685–1694. [Google Scholar] [CrossRef] [PubMed]
- Hur, W.; Lee, J.H.; Kim, S.W.; Kim, J.-H.; Bae, S.H.; Kim, M.; Park, T.; Um, S.-J. Downregulation of microRNA-451 in non-alcoholic steatohepatitis inhibits fatty acid-induced proinflammatory cytokine production through the AMPK/AKT pathway. Int. J. Biochem. Cell Biol. 2015, 64, 265–276. [Google Scholar] [CrossRef] [PubMed]
- Lin, Q.; Gao, Z.; Alarcon, R.M.; Ye, J.; Yun, Z. A role of miR-27 in the regulation of adipogenesis. FEBS J. 2009, 276, 2348–2358. [Google Scholar] [CrossRef] [PubMed]
- Raver-Shapira, N.; Marciano, E.; Meiri, E.; Spector, Y.; Rosenfeld, N.; Moskovits, N.; Bentwich, Z.; Oren, M. Transcriptional activation of miR-34a contributes to p53-mediated apoptosis. Mol. Cell 2007, 26, 731–743. [Google Scholar] [CrossRef] [PubMed]
- Cermelli, S.; Ruggieri, A.; Marrero, J.A.; Ioannou, G.N.; Beretta, L. Circulating microRNAs in patients with chronic hepatitis C and non-alcoholic fatty liver disease. PLoS ONE 2011, 6, e23937. [Google Scholar] [CrossRef] [PubMed]
- Montanini, L.; Lazzeroni, P.; Sartori, C.; Nobili, V.; Crafa, P.; Bernasconi, S.; Street, M.E. Mir-146a and-155 are involved in FOXO1 regulation and non alcoholic fatty liver disease in childhood obesity. In Proceedings of European Society For Paediatric Endocrinology, Dublin, Ireland, 18–20 September 2014.
- Jiang, W.; Liu, J.; Dai, Y.; Zhou, N.; Ji, C.; Li, X. MiR-146b attenuates high-fat diet-induced non-alcoholic steatohepatitis in mice. J. Gastroenterol. Hepatol. 2015, 30, 933–943. [Google Scholar] [CrossRef] [PubMed]
- Cheung, O.; J Sanyal, A. Role of microRNAs in non-alcoholic steatohepatitis. Curr. Pharm. Design 2010, 16, 1952–1957. [Google Scholar] [CrossRef]
- Miller, A.M.; Gilchrist, D.S.; Nijjar, J.; Araldi, E.; Ramirez, C.M.; Lavery, C.A.; Fernandez-Hernando, C.; McInnes, I.B.; Kurowska-Stolarska, M. MiR-155 has a protective role in the development of non-alcoholic hepatosteatosis in mice. PLoS ONE 2013, 8, e72324. [Google Scholar] [CrossRef] [PubMed]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zarfeshani, A.; Ngo, S.; Sheppard, A.M. MicroRNA Expression Relating to Dietary-Induced Liver Steatosis and NASH. J. Clin. Med. 2015, 4, 1938-1950. https://doi.org/10.3390/jcm4111938
Zarfeshani A, Ngo S, Sheppard AM. MicroRNA Expression Relating to Dietary-Induced Liver Steatosis and NASH. Journal of Clinical Medicine. 2015; 4(11):1938-1950. https://doi.org/10.3390/jcm4111938
Chicago/Turabian StyleZarfeshani, Aida, Sherry Ngo, and Allan M. Sheppard. 2015. "MicroRNA Expression Relating to Dietary-Induced Liver Steatosis and NASH" Journal of Clinical Medicine 4, no. 11: 1938-1950. https://doi.org/10.3390/jcm4111938
APA StyleZarfeshani, A., Ngo, S., & Sheppard, A. M. (2015). MicroRNA Expression Relating to Dietary-Induced Liver Steatosis and NASH. Journal of Clinical Medicine, 4(11), 1938-1950. https://doi.org/10.3390/jcm4111938





