Elemental Dynamics in Hair Accurately Predict Future Autism Spectrum Disorder Diagnosis: An International Multi-Center Study
Abstract
1. Introduction
2. Materials and Methods
2.1. Study Populations
2.2. Laboratory Methods
2.3. Computational Methods
2.4. Feature Engineering
2.5. Quality Analysis/Quality Control
2.6. Statistical Analysis
2.7. Predictive Modeling
2.8. Software Implementation
3. Results
3.1. Demographic and Diagnostic Criteria
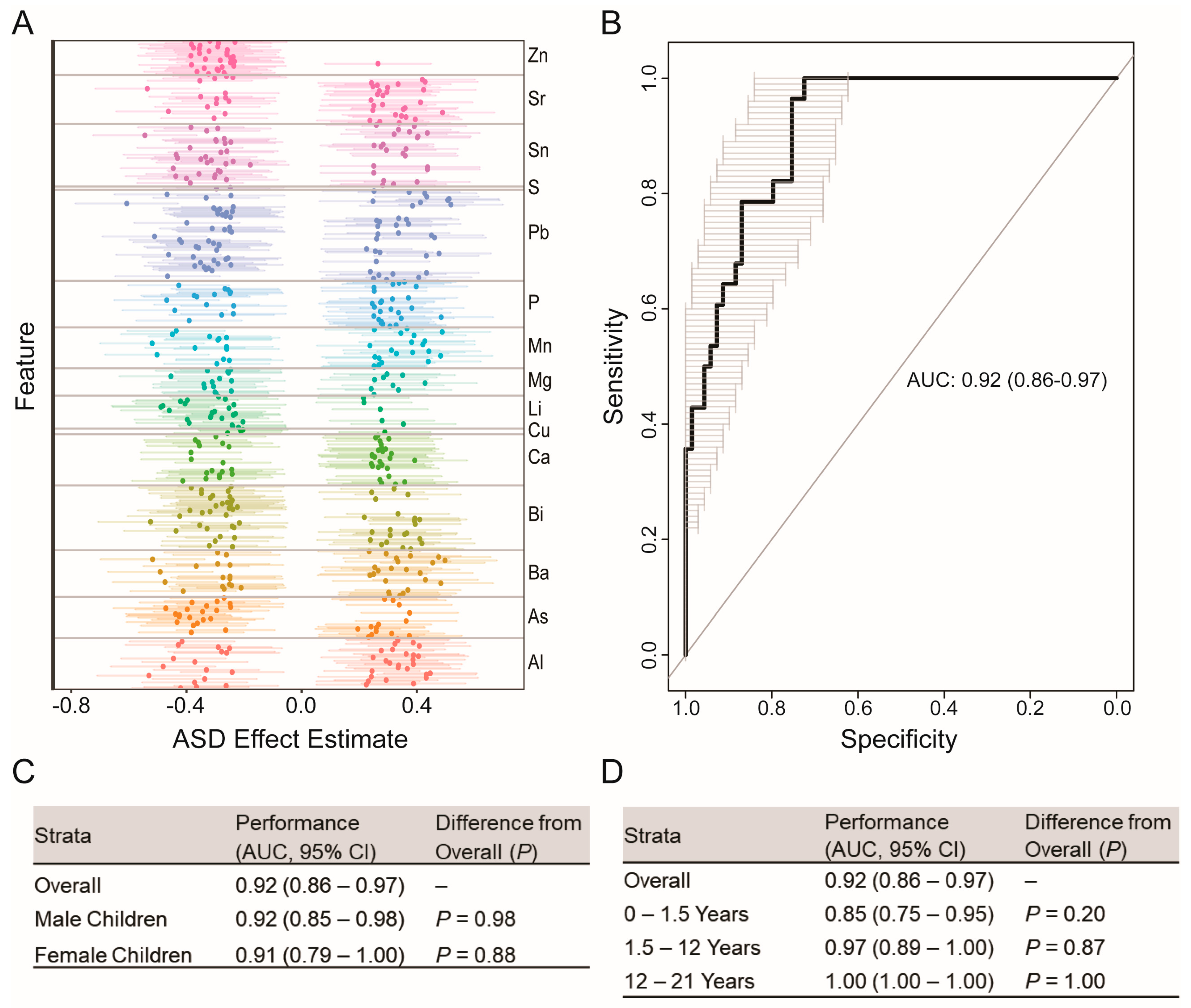
3.2. Elemental Signatures of ASD Phenotype
3.3. Prediction of ASD Case Status
4. Discussion
5. Conclusions
6. Patents
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- de Schipper, E.; Lundequist, A.; Coghill, D.; de Vries, P.J.; Granlund, M.; Holtmann, M.; Jonsson, U.; Karande, S.; Robison, J.E.; Shulman, C.; et al. Ability and Disability in Autism Spectrum Disorder: A Systematic Literature Review Employing the International Classification of Functioning, Disability and Health-Children and Youth Version. Autism Res. 2015, 8, 782–794. [Google Scholar] [CrossRef] [PubMed]
- Simonoff, E.; Pickles, A.; Charman, T.; Chandler, S.; Loucas, T.; Baird, G. Psychiatric disorders in children with autism spectrum disorders: Prevalence, comorbidity, and associated factors in a population-derived sample. J. Am. Acad. Child Adolesc. Psychiatry 2008, 47, 921–929. [Google Scholar] [CrossRef]
- Maenner, M.J.; Shaw, K.A.; Baio, J. Prevalence of Autism Spectrum Disorder Among Children Aged 8 Years-Autism and Developmental Disabilities Monitoring Network, 11 Sites, United States, 2016. MMWR Surveill. Summ. 2020, 69, 1–12. [Google Scholar] [CrossRef] [PubMed]
- van ’t Hof, M.; Tisseur, C.; van Berckelear-Onnes, I.; van Nieuwenhuyzen, A.; Daniels, A.M.; Deen, M.; Hoek, H.W.; Ester, W.A. Age at autism spectrum disorder diagnosis: A systematic review and meta-analysis from 2012 to 2019. Autism 2021, 25, 862–873. [Google Scholar] [CrossRef] [PubMed]
- Shonkoff, J.P.; Phillips, D.A. From Neurons to Neighborhoods: The Science of Early Childhood Development; Shonkoff, J.P., Phillips, D.A., Eds.; National Academies Press (US): Washington, DC, USA, 2000. [Google Scholar] [CrossRef]
- Hyman, S.L.; Levy, S.E.; Myers, S.M.; Council On Children With Disabilities, S.O.D.; Behavioral, P. Identification, Evaluation, and Management of Children With Autism Spectrum Disorder. Pediatrics 2020, 145, e20193447. [Google Scholar] [CrossRef]
- Dawson, G.; Rogers, S.; Munson, J.; Smith, M.; Winter, J.; Greenson, J.; Donaldson, A. Randomized, controlled trial of an intervention for toddlers with autism: The Early Start Denver Model. Pediatric 2010, 125, e17–e23. [Google Scholar] [CrossRef]
- Green, J.; Charman, T.; McConachie, H.; Aldred, C.; Slonims, V.; Howlin, P.; Le Couteur, A.; Leadbitter, K.; Hudry, K.; Byford, S.; et al. Parent-mediated communication-focused treatment in children with autism (PACT): A randomised controlled trial. Lancet 2010, 375, 2152–2160. [Google Scholar] [CrossRef]
- Pickles, A.; Le Couteur, A.; Leadbitter, K.; Salomone, E.; Cole-Fletcher, R.; Tobin, H.; Gammer, I. Parent-mediated social communication therapy for young children with autism (PACT): Long-term follow-up of a randomised controlled trial. Lancet 2016, 388, 2501–2509. [Google Scholar] [CrossRef]
- Sinai-Gavrilov, Y.; Gev, T.; Mor-Snir, I.; Vivanti, G.; Golan, O. Integrating the Early Start Denver Model into Israeli community autism spectrum disorder preschools: Effectiveness and treatment response predictors. Autism 2020, 24, 2081–2093. [Google Scholar] [CrossRef]
- Frazier, T.W.; Coury, D.L.; Sohl, K.; Wagner, K.E.; Uhlig, R.; Hicks, S.D.; Middleton, F.A. Evidence-based use of scalable biomarkers to increase diagnostic efficiency and decrease the lifetime costs of autism. Autism. Res. 2021, 14, 1271–1283. [Google Scholar] [CrossRef]
- Branca, M. Slivers of the spectrum. Nat. Biotechnol. 2021, 39, 540–545. [Google Scholar] [CrossRef] [PubMed]
- Grabrucker, A.M. Environmental factors in autism. Front. Psychiatry 2012, 3, 118. [Google Scholar] [CrossRef] [PubMed]
- Ha, H.T.T.; Leal-Ortiz, S.; Lalwani, K.; Kiyonaka, S.; Hamachi, I.; Mysore, S.P.; Montgomery, J.M.; Garner, C.C.; Huguenard, J.R.; Kim, S.A. Shank and Zinc Mediate an AMPA Receptor Subunit Switch in Developing Neurons. Front. Mol. Neurosci. 2018, 11, 405. [Google Scholar] [CrossRef] [PubMed]
- Shih, P.Y.; Hsieh, B.Y.; Lin, M.H.; Huang, T.N.; Tsai, C.Y.; Pong, W.L.; Lee, S.P.; Hsueh, Y.P. CTTNBP2 Controls Synaptic Expression of Zinc-Related Autism-Associated Proteins and Regulates Synapse Formation and Autism-like Behaviors. Cell Rep. 2020, 31, 107700. [Google Scholar] [CrossRef] [PubMed]
- Modabbernia, A.; Arora, M.; Reichenberg, A. Environmental exposure to metals, neurodevelopment, and psychosis. Curr. Opin. Pediatr. 2016, 28, 243–249. [Google Scholar] [CrossRef]
- Austin, C.; Curtin, P.; Curtin, A.; Gennings, C.; Arora, M.; Tammimies, K.; Isaksson, J.; Willfors, C.; Bolte, S. Dynamical properties of elemental metabolism distinguish attention deficit hyperactivity disorder from autism spectrum disorder. Transl. Psychiatry 2019, 9, 238. [Google Scholar] [CrossRef]
- Curtin, P.; Austin, C.; Curtin, A.; Gennings, C.; Arora, M.; Tammimies, K.; Willfors, C.; Berggren, S.; Siper, P.; Rai, D.; et al. Dynamical features in fetal and postnatal zinc-copper metabolic cycles predict the emergence of autism spectrum disorder. Sci. Adv. 2018, 4, eaat1293. [Google Scholar] [CrossRef]
- Robbins, C.R. Chemical and Physical Behavior of Human Hair, 5th ed.; Springer: Berlin/Heidelberg, Germany, 2012. [Google Scholar]
- Michikawa, T.; Nitta, H.; Nakayama, S.F.; Yamazaki, S.; Isobe, T.; Tamura, K.; Suda, E.; Ono, M.; Yonemoto, J.; Iwai-Shimada, M.; et al. Baseline Profile of Participants in the Japan Environment and Children’s Study (JECS). J. Epidemiol. 2018, 28, 99–104. [Google Scholar] [CrossRef]
- Kawamoto, T.; Nitta, H.; Murata, K.; Toda, E.; Tsukamoto, N.; Hasegawa, M.; Yamagata, Z.; Kayama, F.; Kishi, R.; Ohya, Y.; et al. Rationale and study design of the Japan environment and children’s study (JECS). BMC Public Health 2014, 14, 25. [Google Scholar] [CrossRef]
- Bolte, S.; Willfors, C.; Berggren, S.; Norberg, J.; Poltrago, L.; Mevel, K.; Coco, C.; Fransson, P.; Borg, J.; Sitnikov, R.; et al. The Roots of Autism and ADHD Twin Study in Sweden (RATSS). Twin Res. Hum. Genet. 2014, 17, 164–176. [Google Scholar] [CrossRef]
- Anckarsater, H.; Lundstrom, S.; Kollberg, L.; Kerekes, N.; Palm, C.; Carlstrom, E.; Langstrom, N.; Magnusson, P.K.; Halldner, L.; Bolte, S.; et al. The Child and Adolescent Twin Study in Sweden (CATSS). Twin Res. Hum. Genet. 2011, 14, 495–508. [Google Scholar] [CrossRef] [PubMed]
- Lichtenstein, P.; De Faire, U.; Floderus, B.; Svartengren, M.; Svedberg, P.; Pedersen, N.L. The Swedish Twin Registry: A unique resource for clinical, epidemiological and genetic studies. J. Intern. Med. 2002, 252, 184–205. [Google Scholar] [CrossRef] [PubMed]
- Pedersen, N.L.; Lichtenstein, P.; Svedberg, P. The Swedish Twin Registry in the third millennium. Twin Res. 2002, 5, 427–432. [Google Scholar] [CrossRef] [PubMed]
- Purcell, S.; Neale, B.; Todd-Brown, K.; Thomas, L.; Ferreira, M.A.; Bender, D.; Maller, J.; Sklar, P.; de Bakker, P.I.; Daly, M.J.; et al. PLINK: A tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 2007, 81, 559–575. [Google Scholar] [CrossRef]
- Siper, P.M.; Kolevzon, A.; Wang, A.T.; Buxbaum, J.D.; Tavassoli, T. A clinician-administered observation and corresponding caregiver interview capturing DSM-5 sensory reactivity symptoms in children with ASD. Autism Res. 2017, 10, 1133–1140. [Google Scholar] [CrossRef] [PubMed]
- Webber, C.L.; Zbilut, J.P. Dynamical Assessment of Physiological Systems and States Using Recurrence Plot Strategies. J. Appl. Physiol. 1994, 76, 965–973. [Google Scholar] [CrossRef]
- Marwan, N.; Romano, M.C.; Thiel, M.; Kurths, J. Recurrence plots for the analysis of complex systems. Phys. Rep. 2007, 438, 237–329. [Google Scholar] [CrossRef]
- Curtin, P.; Austin, C.; Curtin, A.; Gennings, C.; Figueroa-Romero, C.; Mikhail, K.A.; Botero, T.M.; Goutman, S.A.; Feldman, E.L.; Arora, M. Dysregulated biodynamics in metabolic attractor systems precede the emergence of amyotrophic lateral sclerosis. PLoS Comput. Biol. 2020, 16, e1007773. [Google Scholar] [CrossRef]
- Curtin, P.; Curtin, A.; Austin, C.; Gennings, C.; Tammimies, K.; Bolte, S.; Arora, M. Recurrence quantification analysis to characterize cyclical components of environmental elemental exposures during fetal and postnatal development. PLoS ONE 2017, 12, e0187049. [Google Scholar] [CrossRef]
- Zbilut, J.P.; Giuliani, A.; Webber, C.L. Recurrence quantification analysis and principal components in the detection of short complex signals. Phys. Lett. A 1998, 237, 131–135. [Google Scholar] [CrossRef]
- Zhang, Y.; Parmigiani, G.; Johnson, W.E. ComBat-seq: Batch effect adjustment for RNA-seq count data. NAR Genom. Bioinform. 2020, 2, lqaa078. [Google Scholar] [CrossRef] [PubMed]
- Chen, T.; Guestrin, C. XGBoost: A Scalable Tree Boosting System. In Proceedings of the KDD ’16: The 22nd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining, San Francisco, CA, USA, 13–17 August 2016; pp. 785–794. [Google Scholar]
- Arora, M.; Curtin, P.; Austin, C. Systems and Methods for Diagnostics for Biological Disorders Associated with Periodic Variations in Metal Metabolism. U.S. Patent 17,616,626, 2022. [Google Scholar]
- Bischl, B.; Richter, J.; Bossek, J.; Horn, D.; Thomas, J.; Lang, M. mlrMBO: A Modular Framework for Model-Based Optimization of Expensive Black-Box Functions. arXiv 2018, arXiv:1703.03373 [stat.ML]. [Google Scholar]
- Datseris, G. DynamicalSystems.jl: A Julia software library for chaos and nonlinear dynamics. J. Open Source Softw. 2018, 3, 598. [Google Scholar] [CrossRef]
- Petrocchi, S.; Levante, A.; Lecciso, F. Systematic Review of Level 1 and Level 2 Screening Tools for Autism Spectrum Disorders in Toddlers. Brain Sci. 2020, 10, 180. [Google Scholar] [CrossRef]
- Arora, M.; Reichenberg, A.; Willfors, C.; Austin, C.; Gennings, C.; Berggren, S.; Lichtenstein, P.; Anckarsater, H.; Tammimies, K.; Bolte, S. Fetal and postnatal metal dysregulation in autism. Nat. Commun. 2017, 8, 15493. [Google Scholar] [CrossRef]
- Grabrucker, A.M. A role for synaptic zinc in ProSAP/Shank PSD scaffold malformation in autism spectrum disorders. Dev. Neurobiol. 2014, 74, 136–146. [Google Scholar] [CrossRef]
- Grabrucker, S.; Jannetti, L.; Eckert, M.; Gaub, S.; Chhabra, R.; Pfaender, S.; Mangus, K.; Reddy, P.P.; Rankovic, V.; Schmeisser, M.J.; et al. Zinc deficiency dysregulates the synaptic ProSAP/Shank scaffold and might contribute to autism spectrum disorders. Brain 2014, 137, 137–152. [Google Scholar] [CrossRef]
- Youden, W.J. Index for rating diagnostic tests. Cancer 1950, 3, 32–35. [Google Scholar] [CrossRef]
- Pugsley, K.; Scherer, S.W.; Bellgrove, M.A.; Hawi, Z. Environmental exposures associated with elevated risk for autism spectrum disorder may augment the burden of deleterious de novo mutations among probands. Mol. Psychiatry 2021, 27, 710–730. [Google Scholar] [CrossRef]
- Hansen, S.N.; Schendel, D.E.; Francis, R.W.; Windham, G.C.; Bresnahan, M.; Levine, S.Z.; Reichenberg, A.; Gissler, M.; Kodesh, A.; Bai, D.; et al. Recurrence Risk of Autism in Siblings and Cousins: A Multinational, Population-Based Study. J. Am. Acad. Child Adolesc. Psychiatry 2019, 58, 866–875. [Google Scholar] [CrossRef]
- Rao, P.A.; Landa, R.J. Association between severity of behavioral phenotype and comorbid attention deficit hyperactivity disorder symptoms in children with autism spectrum disorders. Autism 2014, 18, 272–280. [Google Scholar] [CrossRef] [PubMed]
- Sandin, S.; Schendel, D.; Magnusson, P.; Hultman, C.; Surén, P.; Susser, E.; Grønborg, T.; Gissler, M.; Gunnes, N.; Gross, R.; et al. Autism risk associated with parental age and with increasing difference in age between the parents. Mol. Psychiatry 2016, 21, 693–700. [Google Scholar] [CrossRef] [PubMed]
- Sandin, S.; Hultman, C.M.; Kolevzon, A.; Gross, R.; MacCabe, J.H.; Reichenberg, A. Advancing maternal age is associated with increasing risk for autism: A review and meta-analysis. J. Am. Acad. Child Adolesc. Psychiatry 2012, 51, 477–486.e471. [Google Scholar] [CrossRef] [PubMed]
- Matson, J.L.; Mahan, S.; Kozlowski, A.M.; Shoemaker, M. Developmental milestones in toddlers with autistic disorder, pervasive developmental disorder--not otherwise specified and atypical development. Dev. Neurorehabil. 2010, 13, 239–247. [Google Scholar] [CrossRef] [PubMed]
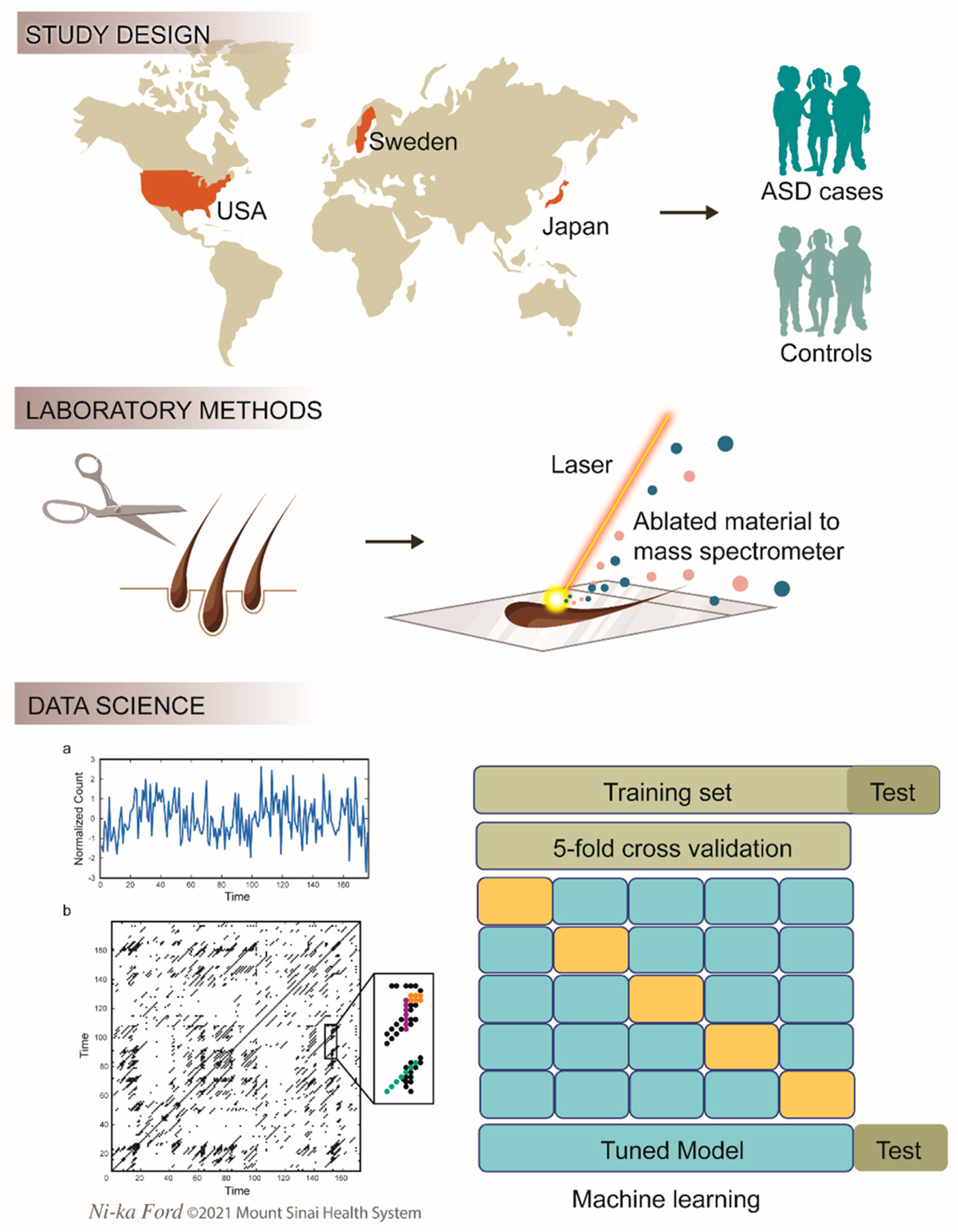
| Study | Location | Design | N (Cases) | Male/Female | Age, Months (SD) |
|---|---|---|---|---|---|
| JECS | Japan | National prospective study | 220 (110) | 110/110 | 1 (0) |
| RATSS | Sweden | National cross-sectional study | 138 (42) | 76/62 | 170.8 (36.9) |
| Seaver | USA | Cross-sectional study | 128 (23) | 80/48 | 61.6 (33.4) |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Austin, C.; Curtin, P.; Arora, M.; Reichenberg, A.; Curtin, A.; Iwai-Shimada, M.; Wright, R.O.; Wright, R.J.; Remnelius, K.L.; Isaksson, J.; et al. Elemental Dynamics in Hair Accurately Predict Future Autism Spectrum Disorder Diagnosis: An International Multi-Center Study. J. Clin. Med. 2022, 11, 7154. https://doi.org/10.3390/jcm11237154
Austin C, Curtin P, Arora M, Reichenberg A, Curtin A, Iwai-Shimada M, Wright RO, Wright RJ, Remnelius KL, Isaksson J, et al. Elemental Dynamics in Hair Accurately Predict Future Autism Spectrum Disorder Diagnosis: An International Multi-Center Study. Journal of Clinical Medicine. 2022; 11(23):7154. https://doi.org/10.3390/jcm11237154
Chicago/Turabian StyleAustin, Christine, Paul Curtin, Manish Arora, Abraham Reichenberg, Austen Curtin, Miyuki Iwai-Shimada, Robert O. Wright, Rosalind J. Wright, Karl Lundin Remnelius, Johan Isaksson, and et al. 2022. "Elemental Dynamics in Hair Accurately Predict Future Autism Spectrum Disorder Diagnosis: An International Multi-Center Study" Journal of Clinical Medicine 11, no. 23: 7154. https://doi.org/10.3390/jcm11237154
APA StyleAustin, C., Curtin, P., Arora, M., Reichenberg, A., Curtin, A., Iwai-Shimada, M., Wright, R. O., Wright, R. J., Remnelius, K. L., Isaksson, J., Bölte, S., & Nakayama, S. F. (2022). Elemental Dynamics in Hair Accurately Predict Future Autism Spectrum Disorder Diagnosis: An International Multi-Center Study. Journal of Clinical Medicine, 11(23), 7154. https://doi.org/10.3390/jcm11237154








