Radiomics Combined with Multiple Machine Learning Algorithms in Differentiating Pancreatic Ductal Adenocarcinoma from Pancreatic Neuroendocrine Tumor: More Hands Produce a Stronger Flame
Abstract
1. Introduction
2. Materials and Methods
2.1. Patient Selection
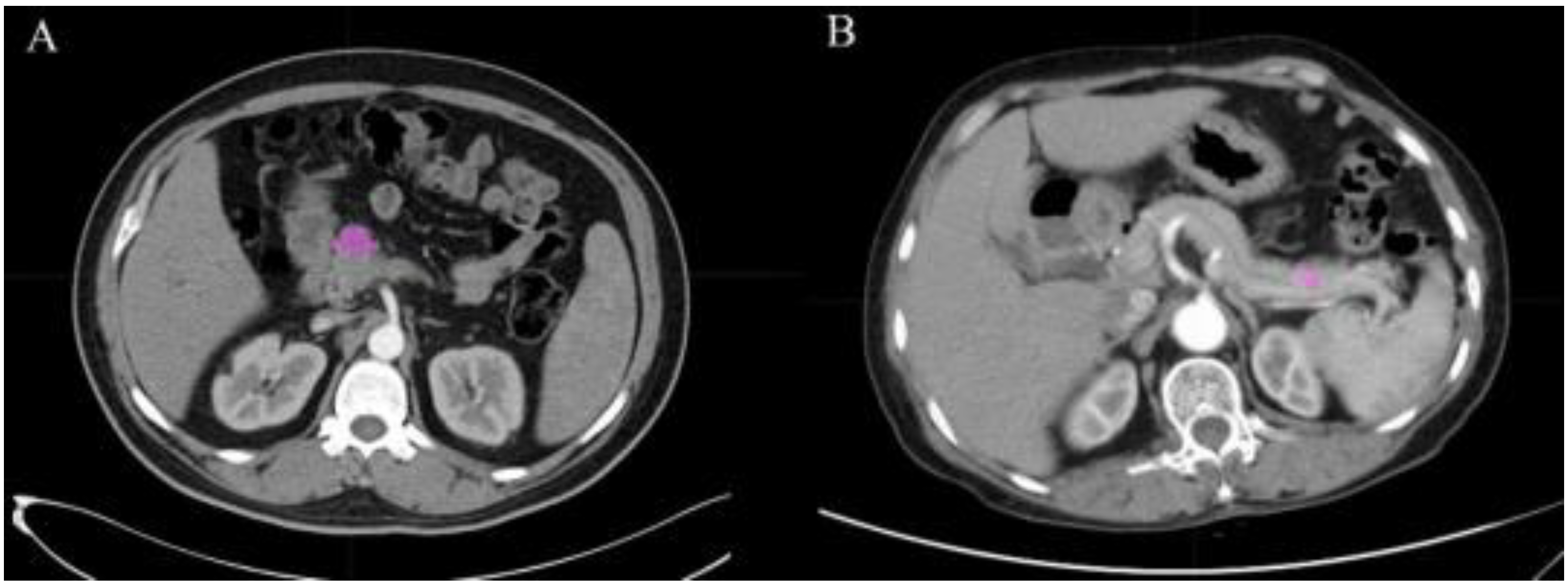
2.2. Image Acquisition and Texture Feature Extraction
2.3. Discriminative Model Establishment and Data Analysis
3. Results
3.1. Patient Characters
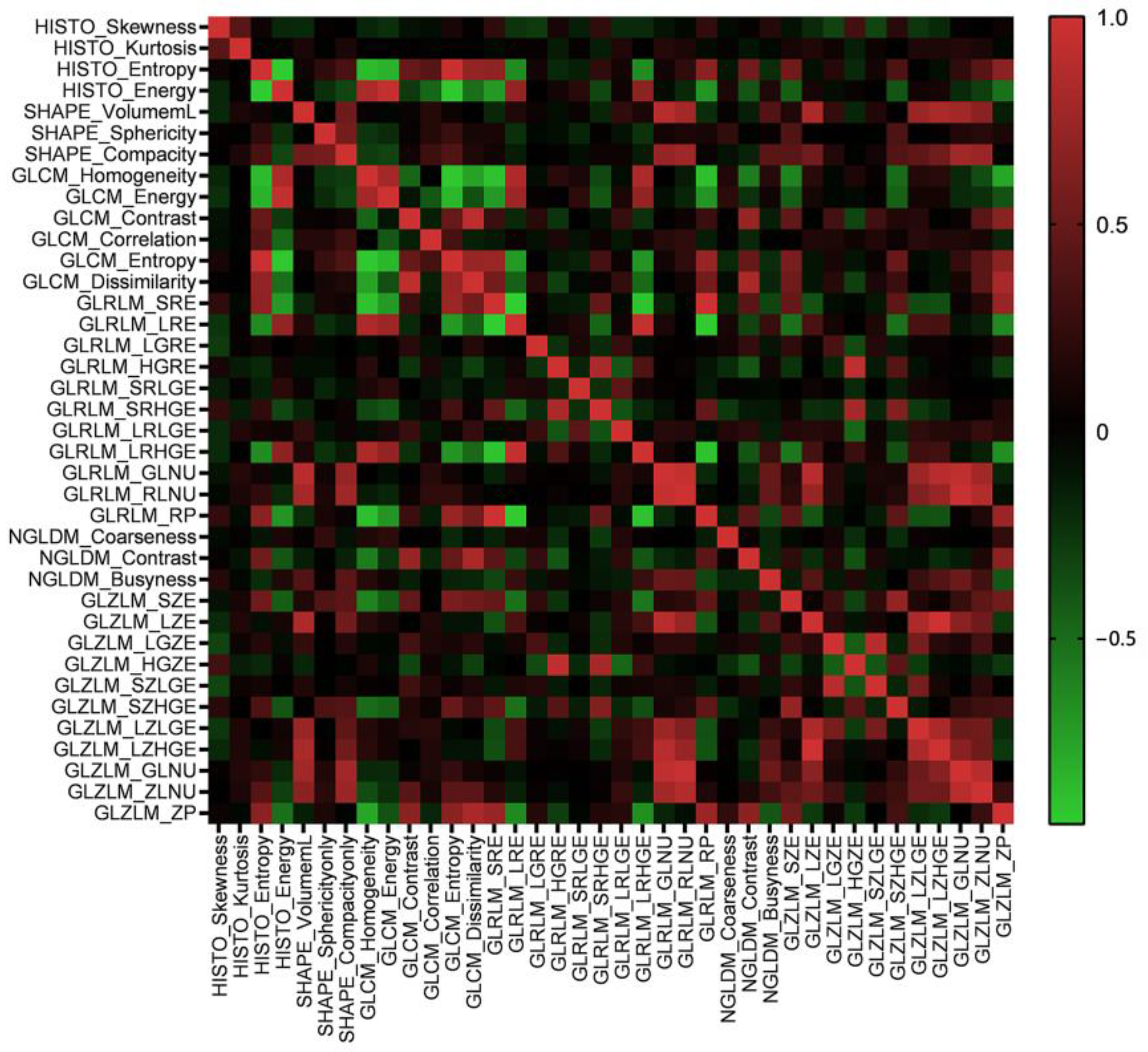
3.2. Radiomics Features
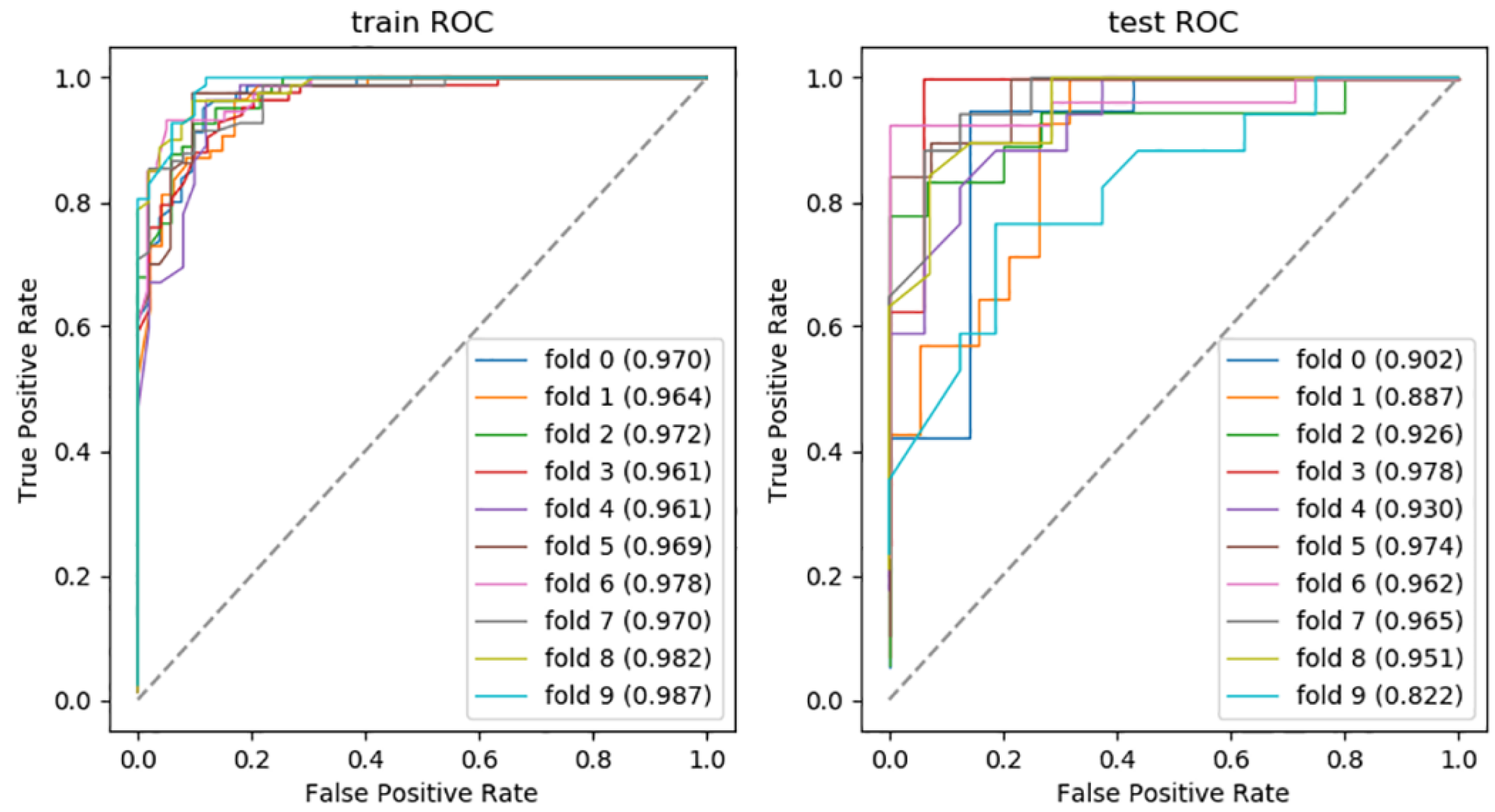
3.3. Model Assessment
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Bray, F.; Ferlay, J.; Soerjomataram, I.; Siegel, R.L.; Torre, L.A.; Jemal, A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA A Cancer J. Clin. 2018, 68, 394–424. [Google Scholar] [CrossRef] [PubMed]
- Mizrahi, J.D.; Surana, R.; Valle, J.W.; Shroff, R.T. Pancreatic cancer. Lancet 2020, 395, 2008–2020. [Google Scholar] [CrossRef]
- Sung, H.; Ferlay, J.; Siegel, R.L.; Laversanne, M.; Soerjomataram, I.; Jemal, A.; Bray, F. Global Cancer Statistics 2020: GLOBOCAN Estimates of Incidence and Mortality Worldwide for 36 Cancers in 185 Countries. CA A Cancer J. Clin. 2021, 71, 209–249. [Google Scholar] [CrossRef] [PubMed]
- Luo, G.; Fan, Z.; Gong, Y.; Jin, K.; Yang, C.; Cheng, H.; Huang, D.; Ni, Q.; Liu, C.; Yu, X. Characteristics and Outcomes of Pancreatic Cancer by Histological Subtypes. Pancreas 2019, 48, 817–822. [Google Scholar] [CrossRef] [PubMed]
- Bengtsson, A.; Andersson, R.; Ansari, D. The actual 5-year survivors of pancreatic ductal adenocarcinoma based on real-world data. Sci. Rep. 2020, 10, 16425. [Google Scholar] [CrossRef]
- Sarantis, P.; Koustas, E.; Papadimitropoulou, A.; Papavassiliou, A.G.; Karamouzis, M.V. Pancreatic ductal adenocarcinoma: Treatment hurdles, tumor microenvironment and immunotherapy. World J. Gastrointest. Oncol. 2020, 12, 173–181. [Google Scholar] [CrossRef]
- Somani, V.K.; Zhang, D.; Dodhiawala, P.B.; Lander, V.E.; Liu, X.; Kang, L.I.; Chen, H.P.; Knolhoff, B.L.; Li, L.; Grierson, P.M.; et al. IRAK4 Signaling Drives Resistance to Checkpoint Immunotherapy in Pancreatic Ductal Adenocarcinoma. Gastroenterology 2022, 162, 2047–2062. [Google Scholar] [CrossRef]
- Yadav, S.; Sharma, P.; Zakalik, D. Comparison of Demographics, Tumor Characteristics, and Survival Between Pancreatic Adenocarcinomas and Pancreatic Neuroendocrine Tumors: A Population-based Study. Am. J. Clin. Oncol. 2018, 41, 485–491. [Google Scholar] [CrossRef]
- Ro, C.; Chai, W.; Yu, V.E.; Yu, R. Pancreatic neuroendocrine tumors: Biology, diagnosis, and treatment. Chin. J. Cancer 2013, 32, 312–324. [Google Scholar] [CrossRef]
- Cejas, P.; Drier, Y.; Dreijerink, K.M.A.; Brosens, L.A.A.; Deshpande, V.; Epstein, C.B.; Conemans, E.B.; Morsink, F.H.M.; Graham, M.K.; Valk, G.D.; et al. Enhancer signatures stratify and predict outcomes of non-functional pancreatic neuroendocrine tumors. Nat. Med. 2019, 25, 1260–1265. [Google Scholar] [CrossRef]
- O’Grady, H.L.; Conlon, K.C. Pancreatic neuroendocrine tumours. Eur. J. Surg. Oncol. 2008, 34, 324–332. [Google Scholar] [CrossRef] [PubMed]
- Tempero, M.A.; Malafa, M.P.; Al-Hawary, M.; Asbun, H.; Bain, A.; Behrman, S.W.; Benson, A.B., 3rd; Binder, E.; Cardin, D.B.; Cha, C.; et al. Pancreatic Adenocarcinoma, Version 2.2017, NCCN Clinical Practice Guidelines in Oncology. J. Natl. Compr. Cancer Netw. JNCCN 2017, 15, 1028–1061. [Google Scholar] [CrossRef] [PubMed]
- Kimura, T.; Miyamoto, H.; Fukuya, A.; Kitamura, S.; Okamoto, K.; Kimura, M.; Muguruma, N.; Ikemoto, T.; Shimada, M.; Yoneda, A.; et al. Neuroendocrine carcinoma of the pancreas with similar genetic alterations to invasive ductal adenocarcinoma. Clin. J. Gastroenterol. 2016, 9, 261–265. [Google Scholar] [CrossRef] [PubMed]
- Raman, S.P.; Hruban, R.H.; Cameron, J.L.; Wolfgang, C.L.; Fishman, E.K. Pancreatic imaging mimics: Part 2, pancreatic neuroendocrine tumors and their mimics. AJR Am. J. Roentgenol. 2012, 199, 309–318. [Google Scholar] [CrossRef]
- Kumar, V.; Gu, Y.; Basu, S.; Berglund, A.; Eschrich, S.A.; Schabath, M.B.; Forster, K.; Aerts, H.J.; Dekker, A.; Fenstermacher, D.; et al. Radiomics: The process and the challenges. Magn. Reson. Imaging 2012, 30, 1234–1248. [Google Scholar] [CrossRef]
- Lambin, P.; Leijenaar, R.T.H.; Deist, T.M.; Peerlings, J.; de Jong, E.E.C.; van Timmeren, J.; Sanduleanu, S.; Larue, R.; Even, A.J.G.; Jochems, A.; et al. Radiomics: The bridge between medical imaging and personalized medicine. Nat. Rev. Clin. Oncol. 2017, 14, 749–762. [Google Scholar] [CrossRef]
- Rogers, W.; Thulasi Seetha, S.; Refaee, T.A.G.; Lieverse, R.I.Y.; Granzier, R.W.Y.; Ibrahim, A.; Keek, S.A.; Sanduleanu, S.; Primakov, S.P.; Beuque, M.P.L.; et al. Radiomics: From qualitative to quantitative imaging. Br. J. Radiol. 2020, 93, 20190948. [Google Scholar] [CrossRef]
- Machine learning in cancer diagnostics. EBioMedicine 2019, 45, 1–2. [CrossRef]
- Hyun, S.H.; Ahn, M.S.; Koh, Y.W.; Lee, S.J. A Machine-Learning Approach Using PET-Based Radiomics to Predict the Histological Subtypes of Lung Cancer. Clin. Nucl. Med. 2019, 44, 956–960. [Google Scholar] [CrossRef]
- Park, Y.W.; Oh, J.; You, S.C.; Han, K.; Ahn, S.S.; Choi, Y.S.; Chang, J.H.; Kim, S.H.; Lee, S.K. Radiomics and machine learning may accurately predict the grade and histological subtype in meningiomas using conventional and diffusion tensor imaging. Eur. Radiol. 2019, 29, 4068–4076. [Google Scholar] [CrossRef]
- Sun, W.; Jiang, M.; Dang, J.; Chang, P.; Yin, F.F. Effect of machine learning methods on predicting NSCLC overall survival time based on Radiomics analysis. Radiat. Oncol. 2018, 13, 197. [Google Scholar] [CrossRef] [PubMed]
- Park, S.; Chu, L.C.; Hruban, R.H.; Vogelstein, B.; Kinzler, K.W.; Yuille, A.L.; Fouladi, D.F.; Shayesteh, S.; Ghandili, S.; Wolfgang, C.L.; et al. Differentiating autoimmune pancreatitis from pancreatic ductal adenocarcinoma with CT radiomics features. Diagn. Interv. Imaging 2020, 101, 555–564. [Google Scholar] [CrossRef] [PubMed]
- Dalal, V.; Carmicheal, J.; Dhaliwal, A.; Jain, M.; Kaur, S.; Batra, S.K. Radiomics in stratification of pancreatic cystic lesions: Machine learning in action. Cancer Lett. 2020, 469, 228–237. [Google Scholar] [CrossRef] [PubMed]
- Yang, J.; Guo, X.; Ou, X.; Zhang, W.; Ma, X. Discrimination of Pancreatic Serous Cystadenomas From Mucinous Cystadenomas With CT Textural Features: Based on Machine Learning. Front. Oncol. 2019, 9, 494. [Google Scholar] [CrossRef] [PubMed]
- He, M.; Liu, Z.; Lin, Y.; Wan, J.; Li, J.; Xu, K.; Wang, Y.; Jin, Z.; Tian, J.; Xue, H. Differentiation of atypical non-functional pancreatic neuroendocrine tumor and pancreatic ductal adenocarcinoma using CT based radiomics. Eur. J. Radiol. 2019, 117, 102–111. [Google Scholar] [CrossRef] [PubMed]
- Xu, J.; Yang, J.; Feng, Y.; Zhang, J.; Zhang, Y.; Chang, S.; Jin, J.; Du, X. MRI Feature-Based Nomogram Model for Discrimination Between Non-Hypervascular Pancreatic Neuroendocrine Tumors and Pancreatic Ductal Adenocarcinomas. Front. Oncol. 2022, 12, 856306. [Google Scholar] [CrossRef]
- Yu, H.; Huang, Z.; Li, M.; Wei, Y.; Zhang, L.; Yang, C.; Zhang, Y.; Song, B. Differential Diagnosis of Nonhypervascular Pancreatic Neuroendocrine Neoplasms From Pancreatic Ductal Adenocarcinomas, Based on Computed Tomography Radiological Features and Texture Analysis. Acad. Radiol. 2020, 27, 332–341. [Google Scholar] [CrossRef]
- Al-Hawary, M.M.; Francis, I.R.; Chari, S.T.; Fishman, E.K.; Hough, D.M.; Lu, D.S.; Macari, M.; Megibow, A.J.; Miller, F.H.; Mortele, K.J.; et al. Pancreatic ductal adenocarcinoma radiology reporting template: Consensus statement of the society of abdominal radiology and the american pancreatic association. Gastroenterology 2014, 146, 291–304.e291. [Google Scholar] [CrossRef]
- Nioche, C.; Orlhac, F.; Boughdad, S.; Reuzé, S.; Goya-Outi, J.; Robert, C.; Pellot-Barakat, C.; Soussan, M.; Frouin, F.; Buvat, I. LIFEx: A Freeware for Radiomic Feature Calculation in Multimodality Imaging to Accelerate Advances in the Characterization of Tumor Heterogeneity. Cancer Res. 2018, 78, 4786–4789. [Google Scholar] [CrossRef]
- Savic, M.; Kurbalija, V.; Ivanovic, M.; Bosnic, Z. A Feature Selection Method Based on Feature Correlation Networks. In Proceedings of the 7th International Conference on Model and Data Engineering (MEDI), Barcelona, Spain, 4–6 October 2017; pp. 248–261. [Google Scholar]
- Zhang, Y.; Chen, C.; Cheng, Y.; Teng, Y.; Guo, W.; Xu, H.; Ou, X.; Wang, J.; Li, H.; Ma, X.; et al. Ability of Radiomics in Differentiation of Anaplastic Oligodendroglioma from Atypical Low-Grade Oligodendroglioma Using Machine-Learning Approach. Front. Oncol. 2019, 9, 1371. [Google Scholar] [CrossRef]
- Siegel, R.L.; Miller, K.D.; Fuchs, H.E.; Jemal, A. Cancer Statistics, 2021. CA A Cancer J. Clin. 2021, 71, 7–33. [Google Scholar] [CrossRef] [PubMed]
- Zhuge, X.; Guo, C.; Chen, Y.; Feng, L.; Jia, R.; Zhao, Y.; Sun, K.; Wang, Z.; Chen, X. The Levels of Tumor Markers in Pancreatic Neuroendocrine Carcinoma and Their Values in Differentiation Between Pancreatic Neuroendocrine Carcinoma and Pancreatic Ductal Adenocarcinoma. Pancreas 2018, 47, 1290–1295. [Google Scholar] [CrossRef] [PubMed]
- Álvarez, R.; Carrato, A.; Adeva, J.; Alés, I.; Prados, S.; Valladares, M.; Macarulla, T.; Muñoz, A.; Hidalgo, M. Management of hyperbilirubinaemia in pancreatic cancer patients. Eur. J. Cancer 2018, 94, 26–36. [Google Scholar] [CrossRef] [PubMed]
- Zhang, S.; Huang, X.; Tian, Y.; Aimaiti, S.; Zhang, J.; Zhao, J.; Chen, Y.; Wang, C. Clinicopathologic characteristics, laboratory parameters, treatment protocols, and outcomes of pancreatic cancer: A retrospective cohort study of 1433 patients in China. PeerJ 2018, 6, e4893. [Google Scholar] [CrossRef] [PubMed]
- Guo, C.; Zhuge, X.; Wang, Q.; Xiao, W.; Wang, Z.; Wang, Z.; Feng, Z.; Chen, X. The differentiation of pancreatic neuroendocrine carcinoma from pancreatic ductal adenocarcinoma: The values of CT imaging features and texture analysis. Cancer Imaging 2018, 18, 37. [Google Scholar] [CrossRef]
- Park, S.; Sham, J.G.; Kawamoto, S.; Blair, A.B.; Rozich, N.; Fouladi, D.F.; Shayesteh, S.; Hruban, R.H.; He, J.; Wolfgang, C.L.; et al. CT Radiomics-Based Preoperative Survival Prediction in Patients With Pancreatic Ductal Adenocarcinoma. AJR Am. J. Roentgenol. 2021, 217, 1104–1112. [Google Scholar] [CrossRef]
- Meng, Y.; Zhang, H.; Li, Q.; Liu, F.; Fang, X.; Li, J.; Yu, J.; Feng, X.; Zhu, M.; Li, N.; et al. CT Radiomics and Machine-Learning Models for Predicting Tumor-Stroma Ratio in Patients With Pancreatic Ductal Adenocarcinoma. Front. Oncol. 2021, 11, 707288. [Google Scholar] [CrossRef]
- Chu, L.C.; Goggins, M.G.; Fishman, E.K. Diagnosis and Detection of Pancreatic Cancer. Cancer J. 2017, 23, 333–342. [Google Scholar] [CrossRef]
- Ryozawa, S.; Kitoh, H.; Gondo, T.; Urayama, N.; Yamashita, H.; Ozawa, H.; Yanai, H.; Okita, K. Usefulness of endoscopic ultrasound-guided fine-needle aspiration biopsy for the diagnosis of pancreatic cancer. J. Gastroenterol. 2005, 40, 907–911. [Google Scholar] [CrossRef]
- Gao, H.L.; Wang, W.Q.; Yu, X.J.; Liu, L. Molecular drivers and cells of origin in pancreatic ductal adenocarcinoma and pancreatic neuroendocrine carcinoma. Exp. Hematol. Oncol. 2020, 9, 28. [Google Scholar] [CrossRef]
- Pulvirenti, A.; Rao, D.; McIntyre, C.A.; Gonen, M.; Tang, L.H.; Klimstra, D.S.; Fleisher, M.; Ramanathan, L.V.; Reidy-Lagunes, D.; Allen, P.J. Limited role of Chromogranin A as clinical biomarker for pancreatic neuroendocrine tumors. HPB 2019, 21, 612–618. [Google Scholar] [CrossRef] [PubMed]
- Goldberg, R.; Meirovitz, A.; Abecassis, A.; Hermano, E.; Rubinstein, A.M.; Nahmias, D.; Grinshpun, A.; Peretz, T.; Elkin, M. Regulation of Heparanase in Diabetes-Associated Pancreatic Carcinoma. Front. Oncol. 2019, 9, 1405. [Google Scholar] [CrossRef] [PubMed]
- Wang, W.Z.; Xu, L.J.; Hu, D.M.; Tang, W. Case of an abnormal procalcitonin-producing metastatic pancreatic neuroendocrine tumor. Clin. Case Rep. 2020, 8, 2269–2272. [Google Scholar] [CrossRef] [PubMed]
- Hassani, C.; Varghese, B.A.; Nieva, J.; Duddalwar, V. Radiomics in Pulmonary Lesion Imaging. AJR Am. J. Roentgenol. 2019, 212, 497–504. [Google Scholar] [CrossRef]
- Goldenberg, S.L.; Nir, G.; Salcudean, S.E. A new era: Artificial intelligence and machine learning in prostate cancer. Nat. Rev. Urol. 2019, 16, 391–403. [Google Scholar] [CrossRef]
- Li, J.; Lu, J.; Liang, P.; Li, A.; Hu, Y.; Shen, Y.; Hu, D.; Li, Z. Differentiation of atypical pancreatic neuroendocrine tumors from pancreatic ductal adenocarcinomas: Using whole-tumor CT texture analysis as quantitative biomarkers. Cancer Med. 2018, 7, 4924–4931. [Google Scholar] [CrossRef]
- Yoo, J.; Kim, S.H.; Jeon, S.K.; Bae, J.S.; Han, J.K. Added value of [(68)Ga]Ga-DOTA-TOC PET/CT for characterizing pancreatic neuroendocrine neoplasms: A comparison with contrast-enhanced CT and/or MRI in a large study cohort. Eur. Radiol. 2021, 31, 7734–7745. [Google Scholar] [CrossRef]
- Li, J.; Liang, L.; Yu, H.; Shen, Y.; Hu, Y.; Hu, D.; Tang, H.; Li, Z. Whole-tumor histogram analysis of non-Gaussian distribution DWI parameters to differentiation of pancreatic neuroendocrine tumors from pancreatic ductal adenocarcinomas. Magn. Reson. Imaging 2019, 55, 52–59. [Google Scholar] [CrossRef]
- Arefan, D.; Chai, R.; Sun, M.; Zuley, M.L.; Wu, S. Machine learning prediction of axillary lymph node metastasis in breast cancer: 2D versus 3D radiomic features. Med. Phys. 2020, 47, 6334–6342. [Google Scholar] [CrossRef]
- Kawakami, E.; Tabata, J.; Yanaihara, N.; Ishikawa, T.; Koseki, K.; Iida, Y.; Saito, M.; Komazaki, H.; Shapiro, J.S.; Goto, C.; et al. Application of Artificial Intelligence for Preoperative Diagnostic and Prognostic Prediction in Epithelial Ovarian Cancer Based on Blood Biomarkers. Clin. Cancer Res. 2019, 25, 3006–3015. [Google Scholar] [CrossRef]
- Chen, C.; Ou, X.; Wang, J.; Guo, W.; Ma, X. Radiomics-Based Machine Learning in Differentiation Between Glioblastoma and Metastatic Brain Tumors. Front. Oncol. 2019, 9, 806. [Google Scholar] [CrossRef] [PubMed]
- Zhu, S.-L.; Dong, J.; Zhang, C.; Huang, Y.-B.; Pan, W. Application of machine learning in the diagnosis of gastric cancer based on noninvasive characteristics. PLoS ONE 2020, 15, e0244869. [Google Scholar] [CrossRef] [PubMed]
- Gao, J.; Zhang, L.; Yu, G.; Qu, G.; Li, Y.; Yang, X. Model with the GBDT for Colorectal Adenoma Risk Diagnosis. Curr. Bioinform. 2020, 15, 971–979. [Google Scholar] [CrossRef]
- Al-Quraishi, T.; Abawajy, J.H.; Chowdhury, M.U.; Rajasegarar, S.; Abdalrada, A.S. Breast Cancer Recurrence Prediction Using Random Forest Model. In Proceedings of the 3rd International Conference on Soft Computing and Data Mining (SCDM), Johor, Malaysia, 6–7 February 2018; pp. 318–329. [Google Scholar]
- Okun, O.; Priisalu, H. Random forest for gene expression based cancer classification: Overlooked issues. In Proceedings of the 3rd Iberian Conference on Pattern Recognition and Image Analysis, Girona, Spain, 6–8 June 2007; pp. 483–490. [Google Scholar]
- Xu, S.; Pan, Z. A novel ensemble of random forest for assisting diagnosis of Parkinson’s disease on small handwritten dynamics dataset. Int. J. Med. Inform. 2020, 144, 104283. [Google Scholar] [CrossRef] [PubMed]
| Characteristics | PDAC (n = 156) | pNET (n = 82) | p Value |
|---|---|---|---|
| Male gender, n (%) | 97 (62.2%) | 48 (58.5%) | 0.307 |
| Mean age (year) | 59.69 ± 11.57 | 53.39 ± 13.01 | <0.001 * |
| BMI (kg/m2) | 25.26 ± 5.67 | 21.64 ± 4.25 | 0.701 |
| Glucose (mmol/L) | 6.83 ± 3.47 (n = 137) | 5.45 ± 3.39 (n = 76) | <0.001 * |
| Procalcitonin (ng/mL) | 1.06 ± 2.22 (n = 115) | 0.92 ± 1.18 (n = 58) | 0.812 |
| AFP (ng/mL) | 3.90 ± 8.58 (n = 151) | 3.114 ± 1.70 (n = 68) | 0.434 |
| CEA (U/mL) | 6.24 ± 10.01 (n = 153) | 3.78 ± 6.68 (n = 77) | 0.092 |
| CA19-9 (U/mL) | 380.10 ± 380.60 (n = 155) | 137.91 ± 282.90 (n = 79) | <0.001 * |
| CA125 (U/mL) | 48.96 ± 49.41 (n = 134) | 20.54 ± 15.04 (n = 66) | 0.184 |
| TBIL (umol/L) | 115.31 ± 144.51 (n = 156) | 26.07 ± 51.74 (n = 82) | <0.001* |
| Amylase (IU/L) | 111.64 ± 107.14 (n = 140) | 225.13 ± 651.94 (n = 70) | 0.208 |
| Lipase (IU/L) | 183.38 ± 270.75 (n = 127) | 309.11 ± 1058.07 (n = 65) | 0.312 |
| Training Dataset | Validation Dataset | |||||
|---|---|---|---|---|---|---|
| Sensitivity | Specificity | AUC | Sensitivity | Specificity | AUC | |
| DC + LDA | 0.839 | 0.903 | 0.953 | 0.775 | 0.884 | 0.938 |
| DC + SVM | 0.972 | 0.563 | 0.931 | 0.987 | 0.528 | 0.910 |
| DC + RF | 0.730 | 0.963 | 0.935 | 0.653 | 0.929 | 0.879 |
| DC + AdaBoost | 1.000 | 1.000 | 1.000 | 0.753 | 0.905 | 0.919 |
| DC + KNN | 0.834 | 0.924 | 0.961 | 0.769 | 0.872 | 0.884 |
| DC + GaussianNB | 0.810 | 0.851 | 0.922 | 0.781 | 0.796 | 0.896 |
| DC + LR | 0.759 | 0.903 | 0.926 | 0.703 | 0.863 | 0.901 |
| DC + GBDT | 1.000 | 1.000 | 1.000 | 0.758 | 0.879 | 0.918 |
| DC + DT | 1.000 | 1.000 | 1.000 | 0.793 | 0.841 | 0.817 |
| RF + LDA | 0.797 | 0.95 | 0.954 | 0.775 | 0.929 | 0.927 |
| RF + SVM | 1.00 | 1.00 | - | 0.00 | 1.00 | - |
| RF + RF | 0.745 | 0.974 | 0.957 | 0.692 | 0.931 | 0.902 |
| RF + AdaBoost | 1.000 | 1.000 | 1.000 | 0.759 | 0.912 | 0.921 |
| RF + KNN | 0.608 | 0.917 | 0.886 | 0.395 | 0.830 | 0.707 |
| RF + GaussianNB | 0.632 | 0.956 | 0.917 | 0.604 | 0.921 | 0.883 |
| RF + LR | 0.796 | 0.897 | 0.948 | 0.816 | 0.878 | 0.944 |
| RF + GBDT | 1.000 | 1.000 | 1.000 | 0.746 | 0.884 | 0.927 |
| RF + DT | 1.000 | 1.000 | 1.000 | 0.732 | 0.832 | 0.782 |
| LASSO + LDA | 0.672 | 0.916 | 0.878 | 0.633 | 0.883 | 0.811 |
| LASSO + SVM | 0.718 | 0.846 | 0.814 | 0.712 | 0.803 | 0.784 |
| LASSO + RF | 0.709 | 0.929 | 0.917 | 0.626 | 0.859 | 0.847 |
| LASSO + AdaBoost | 1.000 | 1.000 | 1.000 | 0.660 | 0.852 | 0.828 |
| LASSO + KNN | 0.659 | 0.955 | 0.915 | 0.601 | 0.894 | 0.796 |
| LASSO + GaussianNB | 0.328 | 0.961 | 0.820 | 0.357 | 0.929 | 0.766 |
| LASSO + LR | 0.364 | 0.986 | 0.819 | 0.340 | 0.963 | 0.788 |
| LASSO + GBDT | 1.000 | 1.000 | 1.000 | 0.677 | 0.833 | 0.849 |
| LASSO + DT | 1.000 | 1.000 | 1.000 | 0.663 | 0.842 | 0.753 |
| Xgboost + LDA | 0.808 | 0.881 | 0.945 | 0.817 | 0.876 | 0.943 |
| Xgboost + SVM | 0.914 | 0.761 | 0.939 | 0.936 | 0.713 | 0.951 |
| Xgboost + RF | 0.750 | 0.980 | 0.960 | 0.711 | 0.968 | 0.925 |
| Xgboost + AdaBoost | 1.000 | 1.000 | 1.000 | 0.821 | 0.916 | 0.932 |
| Xgboost + KNN | 0.804 | 0.941 | 0.967 | 0.777 | 0.928 | 0.940 |
| Xgboost + GaussianNB | 0.848 | 0.801 | 0.918 | 0.840 | 0.739 | 0.896 |
| Xgboost + LR | 0.815 | 0.866 | 0.938 | 0.851 | 0.855 | 0.948 |
| Xgboost + GBDT | 1.000 | 1.000 | 1.000 | 0.758 | 0.925 | 0.931 |
| Xgboost + DT | 1.000 | 1.000 | 1.000 | 0.783 | 0.868 | 0.825 |
| GBDT + LDA | 0.861 | 0.914 | 0.966 | 0.827 | 0.886 | 0.945 |
| GBDT + SVM | 1.000 | 1.000 | - | 0.000 | 1.000 | - |
| GBDT + RF | 0.804 | 0.973 | 0.971 | 0.742 | 0.934 | 0.930 |
| GBDT + AdaBoost | 1.000 | 1.000 | 1.000 | 0.799 | 0.890 | 0.929 |
| GBDT + KNN | 0.620 | 0.942 | 0.881 | 0.429 | 0.866 | 0.730 |
| GBDT + GaussianNB | 0.329 | 0.954 | 0.907 | 0.373 | 0.928 | 0.884 |
| GBDT + LR | 0.809 | 0.920 | 0.953 | 0.782 | 0.882 | 0.927 |
| GBDT + GBDT | 1.000 | 1.000 | 1.000 | 0.746 | 0.893 | 0.927 |
| GBDT + DT | 1.000 | 1.000 | 1.000 | 0.735 | 0.828 | 0.781 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhang, T.; Xiang, Y.; Wang, H.; Yun, H.; Liu, Y.; Wang, X.; Zhang, H. Radiomics Combined with Multiple Machine Learning Algorithms in Differentiating Pancreatic Ductal Adenocarcinoma from Pancreatic Neuroendocrine Tumor: More Hands Produce a Stronger Flame. J. Clin. Med. 2022, 11, 6789. https://doi.org/10.3390/jcm11226789
Zhang T, Xiang Y, Wang H, Yun H, Liu Y, Wang X, Zhang H. Radiomics Combined with Multiple Machine Learning Algorithms in Differentiating Pancreatic Ductal Adenocarcinoma from Pancreatic Neuroendocrine Tumor: More Hands Produce a Stronger Flame. Journal of Clinical Medicine. 2022; 11(22):6789. https://doi.org/10.3390/jcm11226789
Chicago/Turabian StyleZhang, Tao, Yu Xiang, Hang Wang, Hong Yun, Yichun Liu, Xing Wang, and Hao Zhang. 2022. "Radiomics Combined with Multiple Machine Learning Algorithms in Differentiating Pancreatic Ductal Adenocarcinoma from Pancreatic Neuroendocrine Tumor: More Hands Produce a Stronger Flame" Journal of Clinical Medicine 11, no. 22: 6789. https://doi.org/10.3390/jcm11226789
APA StyleZhang, T., Xiang, Y., Wang, H., Yun, H., Liu, Y., Wang, X., & Zhang, H. (2022). Radiomics Combined with Multiple Machine Learning Algorithms in Differentiating Pancreatic Ductal Adenocarcinoma from Pancreatic Neuroendocrine Tumor: More Hands Produce a Stronger Flame. Journal of Clinical Medicine, 11(22), 6789. https://doi.org/10.3390/jcm11226789





