HGF, IL-1α, and IL-27 Are Robust Biomarkers in Early Severity Stratification of COVID-19 Patients
Abstract
1. Introduction
2. Material and Methods
2.1. Patient Selection
2.2. Biological Samples
2.3. Degrees of Severity
2.4. Cytokines and Chemokines Analysis
2.5. Variables
2.6. Statistical Analysis
3. Results
4. Discussion
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Yang, Y.; Peng, F.; Wang, R.; Guan, K.; Jiang, T.; Xu, G.; Sun, J.; Chang, C. The Deadly Coronaviruses: The 2003 SARS Pandemic and the 2020 Novel Coronavirus Epidemic in China. J. Autoimmun. 2020, 109, 102434. [Google Scholar] [CrossRef] [PubMed]
- Zhu, N.; Zhang, D.; Wang, W.; Li, X.; Yang, B.; Song, J.; Zhao, X.; Huang, B.; Shi, W.; Lu, R.; et al. A Novel Coronavirus from Patients with Pneumonia in China, 2019. N. Engl. J. Med. 2020. [Google Scholar] [CrossRef]
- Li, R.; Pei, S.; Chen, B.; Song, Y.; Zhang, T.; Yang, W.; Shaman, J. Substantial Undocumented Infection Facilitates the Rapid Dissemination of Novel Coronavirus (SARS-CoV2). Science 2020. [Google Scholar] [CrossRef]
- Lauer, S.A.; Grantz, K.H.; Bi, Q.; Jones, F.K.; Zheng, Q.; Meredith, H.R.; Azman, A.S.; Reich, N.G.; Lessler, J. The Incubation Period of Coronavirus Disease 2019 (COVID-19) From Publicly Reported Confirmed Cases: Estimation and Application. Ann. Intern. Med. 2020. [Google Scholar] [CrossRef] [PubMed]
- Yang, X.; Yu, Y.; Xu, J.; Shu, H.; Xia, J.; Liu, H.; Wu, Y.; Zhang, L.; Yu, Z.; Fang, M.; et al. Clinical Course and Outcomes of Critically Ill Patients with SARS-CoV-2 Pneumonia in Wuhan, China: A Single-Centered, Retrospective, Observational Study. Lancet Respir. Med. 2020. [Google Scholar] [CrossRef]
- Clinical Management of Severe Acute Respiratory Infection When COVID-19 Is Suspected. Available online: https://www.who.int/publications-detail/clinical-management-of-severe-acute-respiratory-infection-when-novel-coronavirus-(ncov)-infection-is-suspected (accessed on 26 April 2020).
- Li, G.; Fan, Y.; Lai, Y.; Han, T.; Li, Z.; Zhou, P.; Pan, P.; Wang, W.; Hu, D.; Liu, X.; et al. Coronavirus Infections and Immune Responses. J. Med. Virol. 2020, 92, 424–432. [Google Scholar] [CrossRef] [PubMed]
- Schultze, J.L.; Aschenbrenner, A.C. COVID-19 and the Human Innate Immune System. Cell 2021, 184, 1671–1692. [Google Scholar] [CrossRef]
- Schett, G.; Sticherling, M.; Neurath, M.F. COVID-19: Risk for Cytokine Targeting in Chronic Inflammatory Diseases? Nat. Rev. Immunol. 2020. [Google Scholar] [CrossRef] [PubMed]
- Qin, C.; Zhou, L.; Hu, Z.; Zhang, S.; Yang, S.; Tao, Y.; Xie, C.; Ma, K.; Shang, K.; Wang, W.; et al. Dysregulation of Immune Response in Patients with COVID-19 in Wuhan, China. Clin. Infect. Dis. 2020. [Google Scholar] [CrossRef]
- Li, Y.X.; Wu, W.; Yang, T.; Zhou, W.; Fu, Y.M.; Feng, Q.M.; Ye, J.M. Characteristics of peripheral blood leukocyte differential counts in patients with COVID-19. Zhonghua Nei Ke Za Zhi 2020, 59, E003. [Google Scholar] [CrossRef]
- Chen, G.; Wu, D.; Guo, W.; Cao, Y.; Huang, D.; Wang, H.; Wang, T.; Zhang, X.; Chen, H.; Yu, H.; et al. Clinical and Immunological Features of Severe and Moderate Coronavirus Disease 2019. J. Clin. Investig. 2020. [Google Scholar] [CrossRef] [PubMed]
- Ruan, Q.; Yang, K.; Wang, W.; Jiang, L.; Song, J. Clinical Predictors of Mortality Due to COVID-19 Based on an Analysis of Data of 150 Patients from Wuhan, China. Intensive Care Med. 2020, 1–3. [Google Scholar] [CrossRef] [PubMed]
- Induction of Pro-Inflammatory Cytokines (IL-1 and IL-6) and Lung Inflammation by Coronavirus-19 (COVI-19 or SARS-CoV-2): Anti-Inflammatory Strategies. Available online: https://www.ncbi.nlm.nih.gov/pubmed/?term=Induction+of+pro-inflammatory+cytokines+(IL-1+and+IL-6)+and+lung+inflammation+by+COVID-19%3A+anti-inflammatory+strategies (accessed on 26 April 2020).
- Bizzarri, M.; Laganà, A.S.; Aragona, D.; Unfer, V. Inositol and Pulmonary Function. Could Myo-Inositol Treatment Downregulate Inflammation and Cytokine Release Syndrome in SARS-CoV-2? Eur. Rev. Med. Pharmacol. Sci. 2020, 24, 3426–3432. [Google Scholar] [CrossRef]
- Chen, L.; Liu, H.G.; Liu, W.; Liu, J.; Liu, K.; Shang, J.; Deng, Y.; Wei, S. Analysis of clinical features of 29 patients with 2019 novel coronavirus pneumonia. Zhonghua Jie He He Hu Xi Za Zhi 2020, 43, E005. [Google Scholar] [CrossRef] [PubMed]
- Wan, S.; Yi, Q.; Fan, S.; Lv, J.; Zhang, X.; Guo, L.; Lang, C.; Xiao, Q.; Xiao, K.; Yi, Z.; et al. Relationships among Lymphocyte Subsets, Cytokines, and the Pulmonary Inflammation Index in Coronavirus (COVID-19) Infected Patients. Br. J. Haematol. 2020. [Google Scholar] [CrossRef] [PubMed]
- Huang, C.; Wang, Y.; Li, X.; Ren, L.; Zhao, J.; Hu, Y.; Zhang, L.; Fan, G.; Xu, J.; Gu, X.; et al. Clinical Features of Patients Infected with 2019 Novel Coronavirus in Wuhan, China. Lancet 2020, 395, 497–506. [Google Scholar] [CrossRef]
- Moore, B.J.B.; June, C.H. Cytokine Release Syndrome in Severe COVID-19. Science 2020. [Google Scholar] [CrossRef]
- Karakike, E.; Giamarellos-Bourboulis, E.J. Macrophage Activation-Like Syndrome: A Distinct Entity Leading to Early Death in Sepsis. Front. Immunol. 2019, 10. [Google Scholar] [CrossRef]
- WHO-2019-NCoV-Clinical-2021. Available online: https://apps.who.int/iris/bitstream/handle/10665/338882/WHO-2019-nCoV-clinical-2021.1-eng.pdf?sequence=1&isAllowed=y (accessed on 25 January 2021).
- Lee, L. NADA: Nondetects and Data Analysis for Environmental Data. 2020. Available online: https://cran.r-project.org/web/packages/NADA/NADA.pdf (accessed on 25 January 2021).
- Bindoli, S.; Felicetti, M.; Sfriso, P.; Doria, A. The Amount of Cytokine-Release Defines Different Shades of Sars-Cov2 Infection. Exp. Biol. Med. 2020, 245, 970–976. [Google Scholar] [CrossRef]
- Liu, J.; Li, S.; Liu, J.; Liang, B.; Wang, X.; Wang, H.; Li, W.; Tong, Q.; Yi, J.; Zhao, L.; et al. Longitudinal Characteristics of Lymphocyte Responses and Cytokine Profiles in the Peripheral Blood of SARS-CoV-2 Infected Patients. EBioMedicine 2020, 55, 102763. [Google Scholar] [CrossRef]
- Zhang, X.; Tan, Y.; Ling, Y.; Lu, G.; Liu, F.; Yi, Z.; Jia, X.; Wu, M.; Shi, B.; Xu, S.; et al. Viral and Host Factors Related to the Clinical Outcome of COVID-19. Nature 2020, 583, 437–440. [Google Scholar] [CrossRef]
- Han, H.; Ma, Q.; Li, C.; Liu, R.; Zhao, L.; Wang, W.; Zhang, P.; Liu, X.; Gao, G.; Liu, F.; et al. Profiling Serum Cytokines in COVID-19 Patients Reveals IL-6 and IL-10 Are Disease Severity Predictors. Emerg. Microbes. Infect. 2020, 9, 1123–1130. [Google Scholar] [CrossRef]
- Meizlish, M.L.; Pine, A.B.; Bishai, J.D.; Goshua, G.; Nadelmann, E.R.; Simonov, M.; Chang, C.-H.; Zhang, H.; Shallow, M.; Bahel, P.; et al. A Neutrophil Activation Signature Predicts Critical Illness and Mortality in COVID-19. medRxiv 2020. [Google Scholar] [CrossRef]
- Wang, C.; Xie, J.; Zhao, L.; Fei, X.; Zhang, H.; Tan, Y.; Nie, X.; Zhou, L.; Liu, Z.; Ren, Y.; et al. Alveolar Macrophage Dysfunction and Cytokine Storm in the Pathogenesis of Two Severe COVID-19 Patients. EBioMedicine 2020, 57. [Google Scholar] [CrossRef] [PubMed]
- Abassi, Z.; Knaney, Y.; Karram, T.; Heyman, S.N. The Lung Macrophage in SARS-CoV-2 Infection: A Friend or a Foe? Front. Immunol. 2020, 11. [Google Scholar] [CrossRef] [PubMed]
- Lucas, C.; Wong, P.; Klein, J.; Castro, T.B.R.; Silva, J.; Sundaram, M.; Ellingson, M.K.; Mao, T.; Oh, J.E.; Israelow, B.; et al. Longitudinal Analyses Reveal Immunological Misfiring in Severe COVID-19. Nature 2020, 584, 463–469. [Google Scholar] [CrossRef] [PubMed]
- Galván-Román, J.M.; Rodríguez-García, S.C.; Roy-Vallejo, E.; Marcos-Jiménez, A.; Sánchez-Alonso, S.; Fernández-Díaz, C.; Alcaraz-Serna, A.; Mateu-Albero, T.; Rodríguez-Cortes, P.; Sánchez-Cerrillo, I.; et al. IL-6 Serum Levels Predict Severity and Response to Tocilizumab in COVID-19: An Observational Study. J. Allergy Clin. Immunol. 2021, 147, 72–80.e8. [Google Scholar] [CrossRef] [PubMed]
- Utrero-Rico, A.; Ruiz-Hornillos, J.; González-Cuadrado, C.; Rita, C.G.; Almoguera, B.; Minguez, P.; Herrero-González, A.; Fernández-Ruiz, M.; Carretero, O.; Taracido-Fernández, J.C.; et al. IL-6-Based Mortality Prediction Model for COVID-19: Validation and Update in Multicenter and Second Wave Cohorts. J. Allergy Clin. Immunol. 2021. [Google Scholar] [CrossRef]
- Stone, J.H.; Frigault, M.J.; Serling-Boyd, N.J.; Fernandes, A.D.; Harvey, L.; Foulkes, A.S.; Horick, N.K.; Healy, B.C.; Shah, R.; Bensaci, A.M.; et al. Efficacy of Tocilizumab in Patients Hospitalized with Covid-19. N. Engl. J. Med. 2020. [Google Scholar] [CrossRef]
- REMAP-CAP Investigators; Gordon, A.C.; Mouncey, P.R.; Al-Beidh, F.; Rowan, K.M.; Nichol, A.D.; Arabi, Y.M.; Annane, D.; Beane, A.; van Bentum-Puijk, W.; et al. Interleukin-6 Receptor Antagonists in Critically Ill Patients with Covid-19. N. Engl. J. Med. 2021, 384, 1491–1502. [Google Scholar] [CrossRef]
- Group, R.C.; Horby, P.W.; Pessoa-Amorim, G.; Peto, L.; Brightling, C.E.; Sarkar, R.; Thomas, K.; Jeebun, V.; Ashish, A.; Tully, R.; et al. Tocilizumab in Patients Admitted to Hospital with COVID-19 (RECOVERY): Preliminary Results of a Randomised, Controlled, Open-Label, Platform Trial. MedRxiv 2021. [Google Scholar] [CrossRef]
- RECOVERY Collaborative, Group; Horby, P.; Lim, W.S.; Emberson, J.R.; Mafham, M.; Bell, J.L.; Linsell, L.; Staplin, N.; Brightling, C.; Ustianowski, A.; et al. Dexamethasone in Hospitalized Patients with Covid-19—Preliminary Report. N. Engl. J. Med. 2020. [Google Scholar] [CrossRef]
- McElvaney, O.J.; Curley, G.F.; Rose-John, S.; McElvaney, N.G. Interleukin-6: Obstacles to Targeting a Complex Cytokine in Critical Illness. Lancet Respir. Med. 2021. [Google Scholar] [CrossRef]
- Chen, L.Y.C.; Biggs, C.M.; Jamal, S.; Stukas, S.; Wellington, C.L.; Sekhon, M.S. Soluble Interleukin-6 Receptor in the COVID-19 Cytokine Storm Syndrome. Cell Rep. Med. 2021, 100269. [Google Scholar] [CrossRef] [PubMed]

| Mild (n = 34) | Moderate (n = 26) | Severe (n = 16) | Critical (n = 32) | p Value | |
|---|---|---|---|---|---|
| Age [median (IQR)] | 68 (18) | 65 (17) | 75 (14) | 70 (16) | 0.121 |
| Male [%(n)] | 45.2% (14) | 61.5% (16) | 62.5% (10) | 54.8% (17) | 0.568 |
| -Comorbidities, [%(n)] | |||||
| Use of tobacco | 8.80% (3) | 3.80% (1) | 6.3% (1) | 12.5% (4) | 0.679 |
| Use of alcohol | 5.90% (2) | 0% (0) | 0% (0) | 3.1% (1) | 0.488 |
| Coronary cardiopathy | 8.8% (3) | 11.5% (3) | 12.5% (2) | 6.30% (2) | 0.870 |
| Valvular disease | 5.90% (2) | 0% (0) | 12.5% (2) | 0% (0) | 0.104 |
| Atrial fibrillation | 17.6% (6) | 3.80% (1) | 18.8% (3) | 6.3% (2) | 0.206 |
| Diabetes | 11.8% (4) | 11.5% (3) | 18.8% (3) | 25% (8) | 0.435 |
| Hypertension | 50% (17) | 34.6% (9) | 56.3% (9) | 46.9% (15) | 0.521 |
| Liver disease | 0% (0) | 0% (0) | 0% (0) | 6.3% (2) | - |
| COPD | 0% (0) | 7.7% (2) | 18.8% (3) | 6.3% (2) | 0.094 |
| Kidney disease | 2.90% (1) | 0% (0) | 0% (0) | 6.3% (2) | 0.452 |
| Asthma | 11.8% (4) | 3.80% (1) | 0% (0) | 3.1% (1) | 0.268 |
| -Laboratory, [median (IQR)] | |||||
| Glucemia (mg/dL) | 90 (13) | 109 (56) | 120 (59) | 209 (99) | <0.001 |
| Leukocytes (n º/mL) | 4620 (2880) | 6990 (3020) | 6630 (3480) | 7900 (8680) | <0.001 |
| Lymphocytes (n º/mL) | 1000 (430) | 1000 (1000) | 1120 (531) | 440 (455) | <0.001 |
| Neutrophil (n º/mL) | 3215 (2420) | 4945 (2380) | 5315 (3450) | 7045 (7800) | <0.001 |
| Procalcitonin (ng/mL) | 0.06 (0) | 0.05 (0) | 0.15 (1) | 0.24 (0) | <0.001 |
| CRP (mg/L) | 76.5 (88) | 73.5 (106) | 127.0 (113) | 97.0 (153) | 0.250 |
| Creatinine (mg/dL) | 0.81 (0) | 0.78 (0) | 0.88 (0) | 0.89 (1) | 0.242 |
| Total bilirubin (mg/dL) | 0.40 (0) | 0.5 (0) | 0.65 (0) | 0.50 (1) | 0.187 |
| Platelet (cell/mm3) | (82,000) | 232,500 (171,000) | 198,500 (108,500) | 216,500 (108,000) | 0.005 |
| Ferritin (ng/mL) | 587 (600) | 674 (906) | 1025 (938) | 1700 (1093) | <0.001 |
| D-dimer (ng/mL) | 547 (333) | 693 (702) | 1083 (1398) | 1847 (1823) | <0.001 |
| PaO2/FiO2 | 371 (48) | 304 (94) | 238 (102) | 127 (44) | <0.001 |
| -Hospital meters, [median (IQR)] | |||||
| Length of hospital stay (days) | 8 (4) | 8 (6) | 13.5 (10) | 26.5 (39) | <0.001 |
| Length of ICU stay (days) | 0 (0) | 0 (0) | 0 (0) | 18.5 (14) | 0.172 |
| Intubation time (days) | 0 (0) | 0 (0) | 0 (0) | 14 (12) | 0.172 |
| -Mortality, [%(n)] | |||||
| 90-days mortality | 2.9% (1) | 3.8% (1) | 50% (8) | 43.8% (14) | <0.001 |
| 28-days mortality | 0% (0) | 3.8% (1) | 43.8% (7) | 37.5% (12) | <0.001 |
| Int. | Age | Sex | HGF | IL-1α | IL-15 | IL-2 | IL-27 | IL-5 | MCP1 | PDGFBB | PIGF1 | VEGFA | AIC | |
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| M0 | √ | √ | √ | 301.7077 | ||||||||||
| M1 | √ | √ | √ | √ | 268.1021 | |||||||||
| M2 | √ | √ | √ | √ | √ | 268.3859 | ||||||||
| M3 | √ | √ | √ | √ | √ | √ | 265.8642 | |||||||
| M4 | √ | √ | √ | √ | √ | √ | √ | 264.8347 | ||||||
| M5 | √ | √ | √ | √ | √ | √ | √ | √ | 265.6192 | |||||
| M6 | √ | √ | √ | √ | √ | √ | √ | √ | √ | 267.9954 | ||||
| M7 | √ | √ | √ | √ | √ | √ | √ | √ | √ | √ | 271.669 | |||
| M8 | √ | √ | √ | √ | √ | √ | √ | √ | √ | √ | √ | 274.8803 | ||
| M9 | √ | √ | √ | √ | √ | √ | √ | √ | √ | √ | √ | √ | 278.3977 | |
| M10 | √ | √ | √ | √ | √ | √ | √ | √ | √ | √ | √ | √ | √ | 282.1787 |
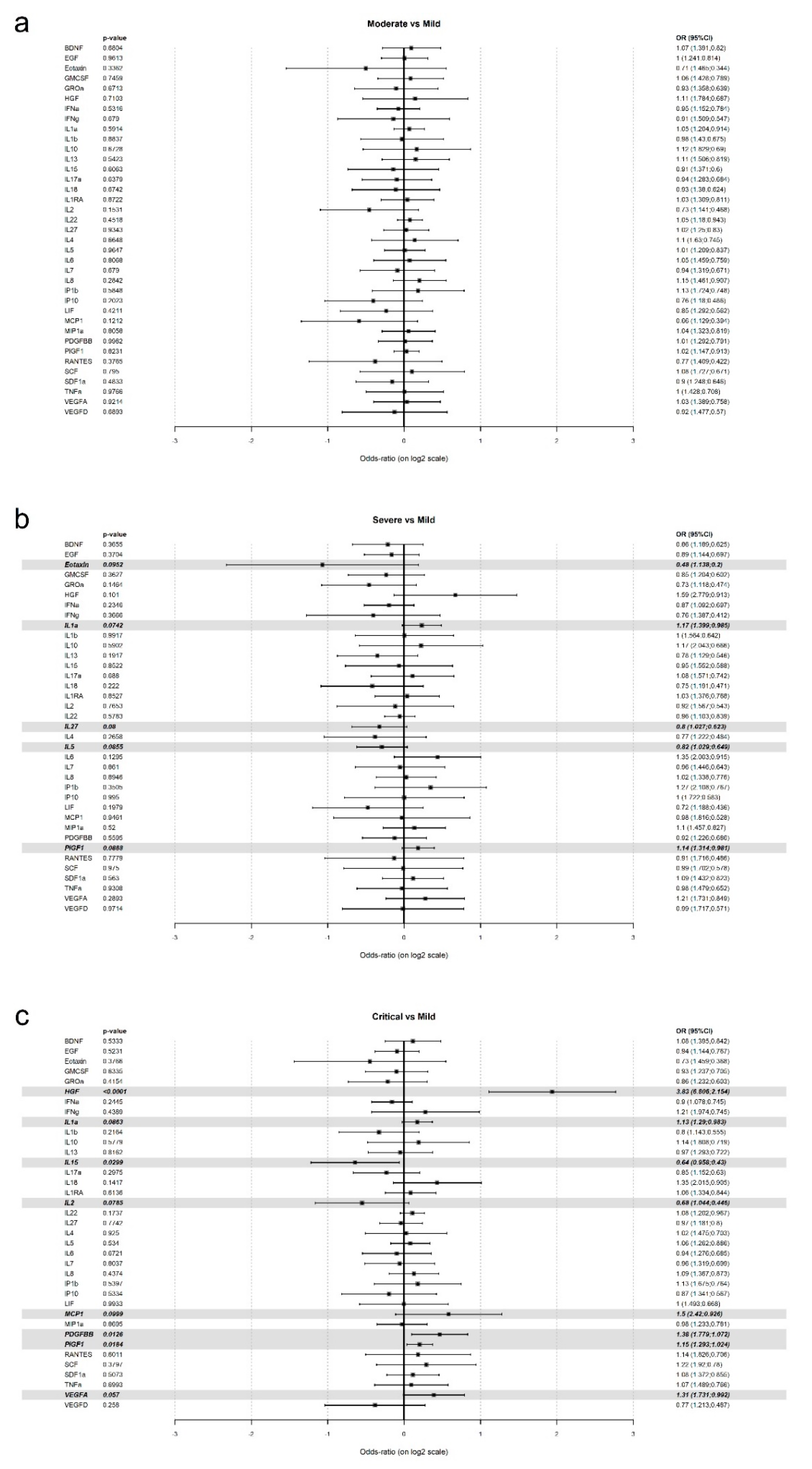
| Severity | Effect | p Value | OR | CI 95% | |
|---|---|---|---|---|---|
| Low | High | ||||
| Moderate | Age | 0.573 | 0.9883 | 0.9486 | 1.0296 |
| Sex = Female | 0.1648 | 0.4618 | 0.1553 | 1.3735 | |
| HGF | 0.7528 | 1.0853 | 0.652 | 1.8066 | |
| IL1a | 0.4346 | 1.081 | 0.8891 | 1.3144 | |
| IL2 | 0.067 | 0.57 | 0.3124 | 1.0401 | |
| IL27 | 0.487 | 1.1148 | 0.8206 | 1.5144 | |
| Severe | Age | 0.0452 | 1.0687 | 1.0014 | 1.1405 |
| Sex = Female | 0.1504 | 0.3517 | 0.0847 | 1.4611 | |
| HGF | 0.2144 | 1.5301 | 0.7818 | 2.9946 | |
| IL1a | 0.0109 | 1.3634 | 1.0741 | 1.7308 | |
| IL2 | 0.4125 | 1.4144 | 0.6172 | 3.2414 | |
| IL27 | 0.0057 | 0.5753 | 0.3888 | 0.8511 | |
| Critical | Age | 0.13 | 0.9615 | 0.9139 | 1.0116 |
| Sex = Female | 0.758 | 0.8242 | 0.241 | 2.8192 | |
| HGF | <0.0001 | 3.5122 | 1.9495 | 6.3276 | |
| IL1a | 0.1977 | 1.134 | 0.9365 | 1.3731 | |
| IL2 | 0.1105 | 0.5776 | 0.2943 | 1.1334 | |
| IL27 | 0.8571 | 0.9677 | 0.6772 | 1.383 | |
| Mild Threshold: 0.3597126 | Moderate Threshold: 0.2513263 | Severe Threshold: 0.1438022 | Critical Threshold: 0.2084408 | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Value | CI 95% | Value | CI 95% | Value | CI 95% | Value | CI 95% | |||||
| Lower | Higher | Lower | Higher | Lower | Higher | Lower | Higher | |||||
| AUC | 0.647 | 0.535 | 0.759 | 0.602 | 0.477 | 0.727 | 0.730 | 0.624 | 0.837 | 0.794 | 0.701 | 0.888 |
| Sensitivity (%) | 58.82 | 42.28 | 75.37 | 53.85 | 34.68 | 73.01 | 62.5 | 38.78 | 86.22 | 81.25 | 67.73 | 94.77 |
| Specificity (%) | 70.27 | 59.86 | 80.68 | 65.85 | 55.59 | 76.12 | 73.91 | 64.94 | 82.89 | 69.74 | 59.41 | 80.07 |
| Accuracy (%) | 66.67 | 57.78 | 75.56 | 62.96 | 53.86 | 72.07 | 72.22 | 63.77 | 80.67 | 73.15 | 64.79 | 81.51 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Tamayo-Velasco, Á.; Martínez-Paz, P.; Peñarrubia-Ponce, M.J.; de la Fuente, I.; Pérez-González, S.; Fernández, I.; Dueñas, C.; Gómez-Sánchez, E.; Lorenzo-López, M.; Gómez-Pesquera, E.; et al. HGF, IL-1α, and IL-27 Are Robust Biomarkers in Early Severity Stratification of COVID-19 Patients. J. Clin. Med. 2021, 10, 2017. https://doi.org/10.3390/jcm10092017
Tamayo-Velasco Á, Martínez-Paz P, Peñarrubia-Ponce MJ, de la Fuente I, Pérez-González S, Fernández I, Dueñas C, Gómez-Sánchez E, Lorenzo-López M, Gómez-Pesquera E, et al. HGF, IL-1α, and IL-27 Are Robust Biomarkers in Early Severity Stratification of COVID-19 Patients. Journal of Clinical Medicine. 2021; 10(9):2017. https://doi.org/10.3390/jcm10092017
Chicago/Turabian StyleTamayo-Velasco, Álvaro, Pedro Martínez-Paz, María Jesús Peñarrubia-Ponce, Ignacio de la Fuente, Sonia Pérez-González, Itziar Fernández, Carlos Dueñas, Esther Gómez-Sánchez, Mario Lorenzo-López, Estefanía Gómez-Pesquera, and et al. 2021. "HGF, IL-1α, and IL-27 Are Robust Biomarkers in Early Severity Stratification of COVID-19 Patients" Journal of Clinical Medicine 10, no. 9: 2017. https://doi.org/10.3390/jcm10092017
APA StyleTamayo-Velasco, Á., Martínez-Paz, P., Peñarrubia-Ponce, M. J., de la Fuente, I., Pérez-González, S., Fernández, I., Dueñas, C., Gómez-Sánchez, E., Lorenzo-López, M., Gómez-Pesquera, E., Heredia-Rodríguez, M., Carnicero-Frutos, I., Muñoz-Moreno, M. F., Bernardo, D., Álvarez, F. J., Tamayo, E., & Gonzalo-Benito, H. (2021). HGF, IL-1α, and IL-27 Are Robust Biomarkers in Early Severity Stratification of COVID-19 Patients. Journal of Clinical Medicine, 10(9), 2017. https://doi.org/10.3390/jcm10092017







