A Deep Learning Ensemble Approach for Automated COVID-19 Detection from Chest CT Images
Abstract
:1. Introduction
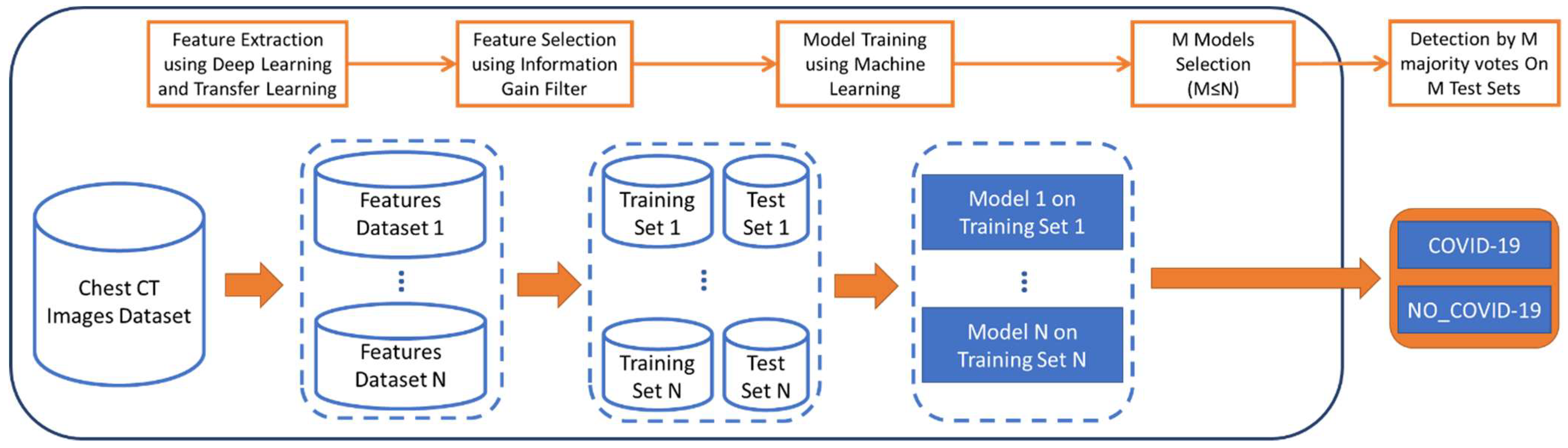
- A feature extraction step using a set of convolutional neural networks that have been pretrained on the ImageNet dataset [30]. ImageNet is a large, publicly available database of natural images with 1000 object classes that was specifically created for computer vision and, more recently, has been widely used for deep learning and transfer learning research. There are more than 1.2 million training images, 50,000 validation images, and 100,000 test images that are available in the database with relative annotations. The images are mostly in the JPG file format and vary in size.
- A feature selection step that uses the information gain filter.
- Training of the generated models using machine learning approaches.
- A model selection step.
- Classification of CT images into one of two classes using a majority voting approach.
2. Materials and Methods
2.1. Dataset
2.2. Proposed Approach
2.3. Transfer Learning
- Instance-based: Mainly refers to instance weighting strategy.
- Feature-based: Transforms the original features to create a new feature representation.
- Parameter-based: Transfers the knowledge at the model/parameter level.
- Relational-based: Focuses on the problems in relational domains. This approach transfers the logical relationship or rules learned in the source domain to the target domain.
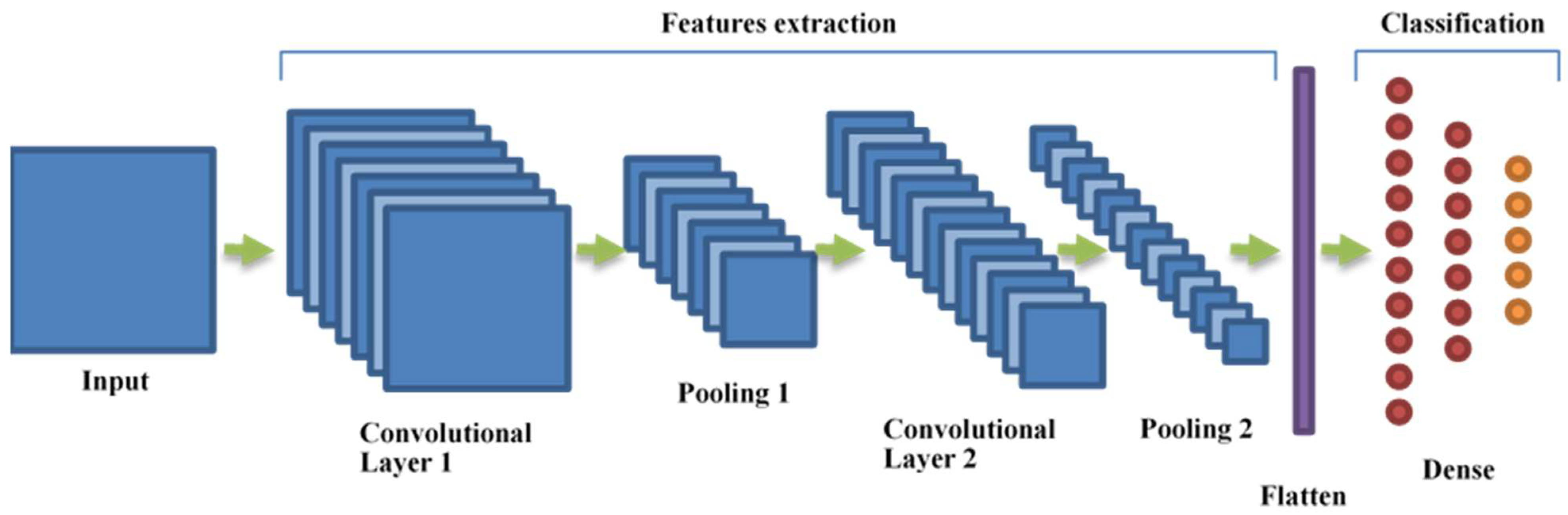
2.4. Convolutional Neural Networks (CNNs)
2.5. K-Nearest Neighbor and Majority Voting Approach
3. Results
3.1. Feature Extraction and Selection
3.2. Model Training
3.3. Model Selection
3.4. Classification
4. Discussion
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- World Health Organization. Available online: https://covid19.who.int/ (accessed on 11 October 2021).
- Armocida, B.; Formenti, B.; Ussai, S.; Palestra, F.; Missoni, E. The Italian health system and the COVID-19 challenge. Lancet Public Health 2020, 5, e253. [Google Scholar] [CrossRef]
- Narin, A.; Kaya, C.; Pamuk, Z. Automatic detection of coronavirus disease (COVID-19) using X-ray images and deep convolutional neural networks. arXiv 2020, arXiv:2003.10849. [Google Scholar] [CrossRef] [PubMed]
- Huang, P.; Liu, T.; Huang, L.; Liu, H.; Lei, M.; Xu, W.; Hu, X.; Chen, J.; Liu, B. Use of chest CT in combination with negative RT-PCR assay for the 2019 novel coronavirus but high clinical suspicion. Radiology 2020, 295, 22–23. [Google Scholar] [CrossRef]
- Winichakoon, P.; Chaiwarith, R.; Liwsrisakun, C.; Salee, P.; Goonna, A.; Limsukon, A.; Kaewpoowat, Q. Negative Nasopharyngeal and Oropharyngeal Swabs Do Not Rule Out COVID-19. J. Clin. Microbiol. 2020, 58, e00297-20. [Google Scholar] [CrossRef] [Green Version]
- Chen, Z.; Li, Y.; Wu, B.; Hou, Y.; Bao, J.; Deng, X. A patient with covid-19 presenting a false-negative reverse transcriptase polymerase chain reaction result. Korean J. Radiol. 2020, 21, 623–624. [Google Scholar] [CrossRef] [Green Version]
- Sethuraman, N.; Jeremiah, S.S.; Ryo, A. Interpreting Diagnostic Tests for SARS-CoV-2. JAMA 2020, 323, 2249–2251. [Google Scholar] [CrossRef]
- American Society for Microbiology. ASM Expresses Concern about Coronavirus Test Reagent Shortages. 2020. Available online: https://asm.org/Articles/Policy/2020/March/ASM-Expresses-Concern-about-Test-Reagent-Shortages (accessed on 11 October 2021).
- Liu, H.; Liu, F.; Li, J.; Zhang, T.; Wang, D.; Lan, W. Clinical and CT imaging features of the COVID-19 pneumonia: Focus on pregnant women and children. J. Infect. 2020, 80, e7–e13. [Google Scholar] [CrossRef]
- Chung, M.; Bernheim, A.; Mei, X.; Zhang, N.; Cui, J.; Jacobi, A.; Li, K.; Li, S.; Shan, H.; Xu, W.; et al. CT imaging features of 2019 novel coronavirus (2019-nCoV). Radiology 2020, 295, 202–207. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chen, S.-G.; Chen, J.-Y.; Yang, Y.-P.; Chien, C.-S.; Wang, M.-L.; Lin, L.-T. Use of radiographic features in COVID-19 diagnosis: Challenges and perspectives. J. Chin. Med. Assoc. 2020, 83, 644–647. [Google Scholar] [CrossRef]
- Borakati, A.; Perera, A.; Johnson, J.; Sood, T. Diagnostic accuracy of X-ray versus CT in COVID-19: A propensity-matched database study. BMJ Open 2020, 10, e042946. [Google Scholar] [CrossRef] [PubMed]
- Khazaei, M.; Asgari, R.; Zarei, E.; Moharramzad, Y.; Haghighatkhah, H.; Taheri, M.S. Incidentally Diagnosed COVID-19 Infection in Trauma Patients; a Clinical Experience. Arch. Acad. Emerg. Med. 2020, 8, e31. [Google Scholar] [PubMed]
- Bai, H.X.; Hsieh, B.; Xiong, Z.; Halsey, K.; Choi, J.W.; Tran, T.M.L.; Pan, I.; Shi, L.-B.; Wang, D.-C.; Mei, J.; et al. Performance of radiologists in differentiating COVID-19 from viral pneumonia on chest CT. Radiology 2020, 296, E46–E54. [Google Scholar] [CrossRef]
- Gao, L.; Zhang, L.; Liu, C.; Wu, S. Handling imbalanced medical image data: A deep-learning-based one-class classification approach. Artif. Intell. Med. 2020, 108, 101935. [Google Scholar] [CrossRef]
- Apostolopoulos, I.D.; Mpesiana, T.A. Covid-19: Automatic detection from X-ray images utilizing transfer learning with convolutional neural networks. Phys. Eng. Sci. Med. 2020, 43, 635–640. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ohata, E.F.; Bezerra, G.M.; das Chagas, J.V.S.; Neto, A.V.L.; Albuquerque, A.B.; de Albuquerque, V.H.C.; Filho, P.P.R. Automatic detection of COVID-19 infection using chest X-ray images through transfer learning. IEEE/CAA J. Autom. Sin. 2020, 8, 239–248. [Google Scholar] [CrossRef]
- Jain, R.; Gupta, M.; Taneja, S.; Hemanth, D.J. Deep learning based detection and analysis of COVID-19 on chest X-ray images. Appl. Intell. 2020, 51, 1690–1700. [Google Scholar] [CrossRef]
- Kassani, H.; Sara, P.H.K.; Wesolowski, M.J.; Schneider, K.A.; Deters, R. Automatic detection of coronavirus disease (COVID-19) in X-ray and CT images: A machine learning based approach. arXiv 2004, arXiv:2004.10641. [Google Scholar] [CrossRef]
- Maghdid, H.S.; Asaad, A.T.; Ghafoor, K.Z.G.; Sadiq, A.S.; Mirjalili, S.; Khan, M.K.K. Diagnosing COVID-19 Pneumonia from X-Ray and CT Images using Deep Learning and Transfer Learning Algorithms. SPIE 2021, 11734, 117340E. [Google Scholar] [CrossRef]
- Shan, F.; Gao, Y.; Wang, J.; Shi, W.; Shi, N.; Han, M.; Xue, J.; Shen, D.; Shi, Y. Lung Infection Quantification of COVID-19 in CT Images with Deep Learning. arXiv 2020, arXiv:2003.04655. [Google Scholar]
- Alshazly, H.; Linse, C.; Barth, E.; Martinetz, T. Explainable COVID-19 Detection Using Chest CT Scans and Deep Learning. Sensors 2021, 21, 455. [Google Scholar] [CrossRef]
- Xu, X.; Jiang, X.; Ma, C.; Du, P.; Li, X.; Lv, S.; Yu, L.; Ni, Q.; Chen, Y.; Su, J.; et al. A Deep Learning System to Screen Novel Coronavirus Disease 2019 Pneumonia. Engineering 2020, 6, 1122–1129. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.; Kang, B.; Ma, J.; Zeng, X.; Xiao, M.; Guo, J.; Cai, M.; Yang, J.; Li, Y.; Meng, X.; et al. A deep learning algorithm using CT images to screen for Corona virus disease (COVID-19). Eur. Radiol. 2021, 31, 6096–6104. [Google Scholar] [CrossRef]
- Gozes, O.; Frid-Adar, M.; Greenspan, H.; Browning, P.D.; Zhang, H.; Ji, W.; Bernheim, A.; Siegel, E. Rapid AI development cycle for the coronavirus (COVID-19) pandemic: Initial results for automated detection & patient monitoring using deep learning ct image analysis. arXiv 2020, arXiv:2003.05037. [Google Scholar]
- Hasan, N.; Bao, Y.; Shawon, A.; Huang, Y. DenseNet Convolutional Neural Networks Application for Predicting COVID-19 Using CT Image. SN Comput. Sci. 2021, 2, 389. [Google Scholar] [CrossRef]
- Rohila, V.S.; Gupta, N.; Kaul, A.; Sharma, D.K. Deep learning assisted COVID-19 detection using full CT-scans. Internet Things 2021, 14, 100377. [Google Scholar] [CrossRef]
- Soares, E.; Angelov, P.; Biaso, S.; Froes, M.H.; Abe, D.K. SARS-CoV-2 CT-scan dataset: A large dataset of real patients CT scans for SARS-CoV-2 identification. medRxiv 2020. [Google Scholar] [CrossRef]
- Loddo, A.; Pili, F.; Di Ruberto, C. Deep Learning for COVID-19 Diagnosis from CT Images. Appl. Sci. 2021, 11, 8227. [Google Scholar] [CrossRef]
- Deng, J.; Dong, W.; Socher, R.; Li, L.J.; Li, K.; Li, F.-F. Imagenet: A large-scale hierarchical image database. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Miami, FL, USA, 20–25 June 2009; pp. 248–255. [Google Scholar]
- Abadi, M.; Barham, P.; Chen, J.; Chen, Z.; Davis, A.; Dean, J.; Devin, M.; Ghemawat, S.; Irving, G.; Isard, M.; et al. TensorFlow: A System for Large-Scale Machine Learning. In Proceedings of the 12th USENIX Symposium on Operating Systems Design and Implementation (OSDI ’16), Savannah, GA, USA, 2–4 November 2016. [Google Scholar]
- Tan, P.; Steinbach, M.; Kumar, V. Introduction to Data Mining; Pearson Addison Wesley: Boston, MA, USA, 2005. [Google Scholar]
- Pan, S.J.; Yang, Q. A Survey on Transfer Learning. IEEE Trans. Knowl. Data Eng. 2010, 22, 1345–1359. [Google Scholar] [CrossRef]
- Sarkar, D.; Bali, R.; Ghosh, T. Hands-On Transfer Learning with Python; Packt Publishing Ltd.: Birmingham, UK, 2018. [Google Scholar]
- Zhuang, F.; Qi, Z.; Duan, K.; Xi, D.; Zhu, Y.; Zhu, H.; Xiong, H.; He, Q. A Comprehensive Survey on Transfer Learning. Proc. IEEE 2021, 109, 43–76. [Google Scholar] [CrossRef]
- Yang, Q.; Zhang, Y.; Dai, W.; Pan, S.J. Transfer Learning; Cambridge University Press: Cambridge, UK, 2020. [Google Scholar]
- Huh, M.; Agrawal, P.; Efros, A.A. What makes imagenet good for transfer learning? arXiv 2016, arXiv:1608.08614. [Google Scholar]
- Huang, G.; Liu, Z.; Van Der Maaten, L.; Weinberger, K.Q. Densely Connected Convolutional Networks. In Proceedings of the 2017 IEEE Conference on Computer Vision and Pattern Recognition (CVPR), Honolulu, HI, USA, 21–26 July 2017; pp. 2261–2269. [Google Scholar] [CrossRef] [Green Version]
- Tan, M.; Le, Q. EfficientNet: Rethinking Model Scaling for Convolutional Neural Networks. In Proceedings of the 36th International Conference on Machine Learning, Long Beach, CA, USA, 9–15 June 2019; Volume 97, pp. 6105–6114. Available online: http://proceedings.mlr.press/v97/tan19a.html (accessed on 11 October 2021).
- Szegedy, C.; Ioffe, S.; Vanhoucke, V.; Alemi, A.A. Inception-v4, inception-ResNet and the impact of residual connections on learning. In Proceedings of the Thirty-First AAAI Conference on Artificial Intelligence (AAAI’17), San Francisco, CA, USA, 4–9 February 2017; AAAI Press: Cambridge, MA, USA, 2017; pp. 4278–4284. [Google Scholar]
- Szegedy, C.; Liu, W.; Jia, Y.; Sermanet, P.; Reed, S.; Anguelov, D.; Erhan, D.; Vanhoucke, V.; Rabinovich, A. Going deeper with convolutions. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Boston, MA, USA, 7–12 June 2015; pp. 1–9. [Google Scholar] [CrossRef] [Green Version]
- Howard, A.G.; Zhu, M.; Chen, B.; Kalenichenko, D.; Wang, W.; Weyand, T.; Andreetto, M.; Adam, H. MobileNets: Efficient Convolutional Neural Networks for Mobile Vision Applications. arXiv 2017, arXiv:1704.04861. [Google Scholar]
- He, K.; Zhang, X.; Ren, S.; Sun, J. Deep Residual Learning for Image Recognition. In Proceedings of the 2016 IEEE Conference on Computer Vision and Pattern Recognition (CVPR), Las Vegas, NV, USA, 27–30 June 2016; pp. 770–778. [Google Scholar] [CrossRef] [Green Version]
- Simonyan, K.; Zisserman, A. Very Deep Convolutional Networks for Large-Scale Image Recognition. In Proceedings of the International Conference on Learning Representations, San Diego, CA, USA, 7–9 May 2015. [Google Scholar]
- Chollet, F. Xception: Deep Learning with Depthwise Separable Convolutions. In Proceedings of the 2017 IEEE Conference on Computer Vision and Pattern Recognition (CVPR), Honolulu, HI, USA, 21–26 July 2017; pp. 1800–1807. [Google Scholar] [CrossRef] [Green Version]
- Leon, F.; Floria, S.; Bădică, C. Evaluating the effect of voting methods on ensemble-based classification. In Proceedings of the 2017 IEEE International Conference on INnovations in Intelligent SysTems and Applications (INISTA), Gdynia, Poland, 3–5 July 2017; pp. 1–6. [Google Scholar] [CrossRef]
- Kuncheva, L.I. Combining Pattern Classifiers: Methods and Algorithms, 2nd ed.; John Wiley & Sons, Inc.: Hoboken, NJ, USA, 2014. [Google Scholar]
- Aha, D.; Kibler, D. Instance-based learning algorithms. Mach. Learn. 1991, 6, 37–66. [Google Scholar] [CrossRef] [Green Version]
- Bhatia, N.; Vandana. Survey of Nearest Neighbor Techniques. IJCSIS Int. J. Comput. Sci. Inf. Secur. 2010, 8, 302–305. [Google Scholar]
- Lubis, A.R.; Lubis, M.; Al-Khowarizmi, A.-K. Optimization of distance formula in K-Nearest Neighbor method. Bull. Electr. Eng. Inform. 2020, 9, 326–338. [Google Scholar] [CrossRef]
| Subsetting Name | Description | Label Information | |
|---|---|---|---|
| 1 | Inductive TL | the target task is different, but related, from the source task | comes from the target domain |
| 2 | Transductive TL | the source and target tasks are the same, while the source and target domains are different | comes from the source domain |
| 3 | Unsupervised TL | the target is different from, but related to, the source task, and the focus is on solving unsupervised learning tasks in the target domain | is always unknown for both the source and the target domains |
| N | Deep Neural Network | Maximum Value of Information Gain | N. of Original Features | N. of Selected Features | Percentage Reduction of Features by IG |
|---|---|---|---|---|---|
| 1 | DenseNet121 | 0.145 | 1025 | 930 | 9.27% |
| 2 | DenseNet169 | 0.162 | 1665 | 1488 | 10.63% |
| 3 | DenseNet201 | 0.165 | 1921 | 1669 | 13.12% |
| 4 | EfficientNetB0 | 0.203 | 1281 | 1159 | 9.52% |
| 5 | EfficientNetB1 | 0.160 | 1281 | 1202 | 6.17% |
| 6 | EfficientNetB2 | 0.187 | 1409 | 1314 | 6.74% |
| 7 | EfficientNetB3 | 0.207 | 1537 | 1431 | 6.90% |
| 8 | EfficientNetB4 | 0.141 | 1793 | 1598 | 10.88% |
| 9 | EfficientNetB5 | 0.164 | 2049 | 1903 | 7.13% |
| 10 | EfficientNetB6 | 0.181 | 2305 | 2074 | 10.02% |
| 11 | EfficientNetB7 | 0.157 | 2561 | 2315 | 9.61% |
| 12 | InceptionResNetV2 | 0.126 | 1737 | 1533 | 11.74% |
| 13 | InceptionV3 | 0.186 | 2049 | 1899 | 7.32% |
| 14 | MobileNet | 0.175 | 1025 | 829 | 19.12% |
| 15 | MobileNetV2 | 0.144 | 1281 | 889 | 30.60% |
| 16 | MobileNetV3Large | 0.116 | 1281 | 1080 | 15.69% |
| 17 | MobileNetV3Small | 0.111 | 1025 | 849 | 17.17% |
| 18 | ResNet50 | 0.333 | 2049 | 1735 | 15.32% |
| 19 | ResNet50V2 | 0.150 | 2049 | 955 | 53.39% |
| 20 | ResNet101 | 0.204 | 2049 | 1717 | 16.20% |
| 21 | ResNet101V2 | 0.183 | 2049 | 797 | 61.10% |
| 22 | ResNet152 | 0.284 | 2049 | 1665 | 18.74% |
| 23 | ResNet152V2 | 0.206 | 2049 | 1518 | 25.92% |
| 24 | VGG16 | 0.156 | 513 | 404 | 21.25% |
| 25 | VGG19 | 0.184 | 513 | 440 | 14.23% |
| 26 | Xception | 0.138 | 2049 | 1071 | 47.73% |
| Average Accuracy = 90.4548% | Average Accuracy = 91.5502% (without InceptionResNetV2) | ||
|---|---|---|---|
| N | Name | Accuracy of k-NN (k = 1) 10-Fold Cross Validated | Accuracy of k-NN (k = 1) on 10% Test Set |
| 1 | DenseNet121 | 93.4186% | 93.3014% |
| 2 | DenseNet169 | 92.5926% | 94.7368% |
| 3 | DenseNet201 | 91.3403% | 90.1914% |
| 4 | EfficientNetB0 | 95.2038% | 95.933% |
| 5 | EfficientNetB1 | 96.7493% | 96.6507% |
| 6 | EfficientNetB2 | 93.6318% | 94.7368% |
| 7 | EfficientNetB3 | 93.7393% | 96.1722% |
| 8 | EfficientNetB4 | 92.8058% | 93.5407% |
| 9 | EfficientNetB5 | 91.8732% | 91.3876% |
| 10 | EfficientNetB6 | 88.5159% | 87.5598% |
| 11 | EfficientNetB7 | 92.3261% | 94.0191% |
| 12 | InceptionResNetV2 | 78.737% | 77.2727% |
| 13 | InceptionV3 | 88.5425% | 88.756% |
| 14 | MobileNet | 90.9406% | 90.6699% |
| 15 | MobileNetV2 | 91.3669% | 93.7799% |
| 16 | MobileNetV3Large | 88.5425% | 90.1914% |
| 17 | MobileNetV3Small | 84.3858% | 83.7321% |
| 18 | ResNet50 | 95.3637% | 96.1722% |
| 19 | ResNet50V2 | 81.1617% | 81.5789% |
| 20 | ResNet101 | 94.1913% | 93.5407% |
| 21 | ResNet101V2 | 86.2776% | 84.9282% |
| 22 | ResNet152 | 94.1114% | 96.89% |
| 23 | ResNet152V2 | 84.0927% | 85.6459% |
| 24 | VGG16 | 94.7509% | 93.3014% |
| 25 | VGG19 | 94.5377% | 95.933% |
| 26 | Xception | 82.6272% | 85.4067% |
| YES | NO | Classified as | |
|---|---|---|---|
| YES | TP = 215 | FN = 2 | |
| NO | FP = 2 | TN = 199 | |
| Meta-Classifier Accuracy 99.04% | |||
| Symbol | Performance Metric | Definition as | What Does It Measure? | Value |
|---|---|---|---|---|
| CCR | Correctly Classified instance Rate—Accuracy | (TP + TN)/(TP + TN + FP + FN) | How good the model is at correctly predicting both positive and negative cases | 0.9904 |
| TPR | True Positive Rate—Sensitivity—Recall | TP/(TP + FN) | How good the model is at correctly predicting positive cases | 0.9908 |
| FPR | False Positive Rate—Fall-out | FP/(FP + TN) | Proportion of incorrectly classified negative cases | 0.010 |
| PPV | Positive Predictive Value—Precision | TP/(TP + FP) | Proportion of correctly classified positive cases out of total positive predictions | 0.9908 |
| AUC | ROC Area | Area under the ROC curve | Area under plot of TPR against FPR | 0.997 |
| Author | ML Approach | Data Source | Transfer Learning | Achieved Performance |
|---|---|---|---|---|
| Alshazly et al. [22] | Pre-trained SqueezeNet, Inception, ResNet, ResNeXt, Xception, ShuffleNet and DenseNet CNN with fine tuning | 2482 CT images + 746 CT images | Not Declared | Accuracy: 99.4% and 92.9% on the two datasets |
| Xu et al. [23] | ROI segmentation with 3D CNN + Classification with ad hoc ResNet-18 CNN | 618 CT images | No | Accuracy: 86.7% |
| Wang et al. [24] | Pre-trained Inception CNN with fine tuning | 1065 CT images | Not Declared | Accuracy: 79.3% Recall: 83% Specificity: 67% AUC: 0.81 |
| Gozes et al. [25] | Pre-trained ResNet-50 CNN with fine tuning | 206 patients CT scans | ImageNet | AUC: 0.996 |
| Hasan et al. [26] | DenseNet-121 CNN | 2482 CT images | No | Accuracy: 92% Recall: 95% |
| Rohila et al. [27] | Ad hoc deep learning network based on ResNet-101 | 1110 patients CT scans | Yes, but no ImageNet | Accuracy: 94.9% |
| Soares et al. [28] | xDNN (eXplainable Deep Neural Network) | 2482 CT images | ImageNet | Accuracy: 97.4% Recall: 95.53% Precision: 99.16% AUC: 0.9736 |
| Loddo et al. [29] | Pre-trained AlexNet, Residual Networks, ResNet18, ResNet50, ResNet101, GoogLeNet, ShuffleNet, MobileNetV2, InceptionV3, VGG16 and VGG19 | 470 + 194,122 Chest CT images | No | Accuracy: 98.87% (nets comparison) 95.91% (patient status classification) |
| Our approach | Pre-trained CNNs, k Nearest Neighbors with 10-fold cross validation, majority voting approach | 2482 CT images | Yes ImageNet | Accuracy: 99.04% Recall: 99.08% Precision: 99.08% AUC: 0.997 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zazzaro, G.; Martone, F.; Romano, G.; Pavone, L. A Deep Learning Ensemble Approach for Automated COVID-19 Detection from Chest CT Images. J. Clin. Med. 2021, 10, 5982. https://doi.org/10.3390/jcm10245982
Zazzaro G, Martone F, Romano G, Pavone L. A Deep Learning Ensemble Approach for Automated COVID-19 Detection from Chest CT Images. Journal of Clinical Medicine. 2021; 10(24):5982. https://doi.org/10.3390/jcm10245982
Chicago/Turabian StyleZazzaro, Gaetano, Francesco Martone, Gianpaolo Romano, and Luigi Pavone. 2021. "A Deep Learning Ensemble Approach for Automated COVID-19 Detection from Chest CT Images" Journal of Clinical Medicine 10, no. 24: 5982. https://doi.org/10.3390/jcm10245982
APA StyleZazzaro, G., Martone, F., Romano, G., & Pavone, L. (2021). A Deep Learning Ensemble Approach for Automated COVID-19 Detection from Chest CT Images. Journal of Clinical Medicine, 10(24), 5982. https://doi.org/10.3390/jcm10245982







