Identification of Immunological Parameters as Predictive Biomarkers of Relapse in Patients with Chronic Myeloid Leukemia on Treatment-Free Remission
Abstract
1. Introduction
2. Experimental Methods
2.1. Participants
2.2. Flow Cytometry Analysis
2.3. HLA-E Genotyping
2.4. KIR Haplotyping
2.5. Random Forest Algorithm
2.6. Statistical Analysis
3. Results
3.1. Patients’ Characteristics
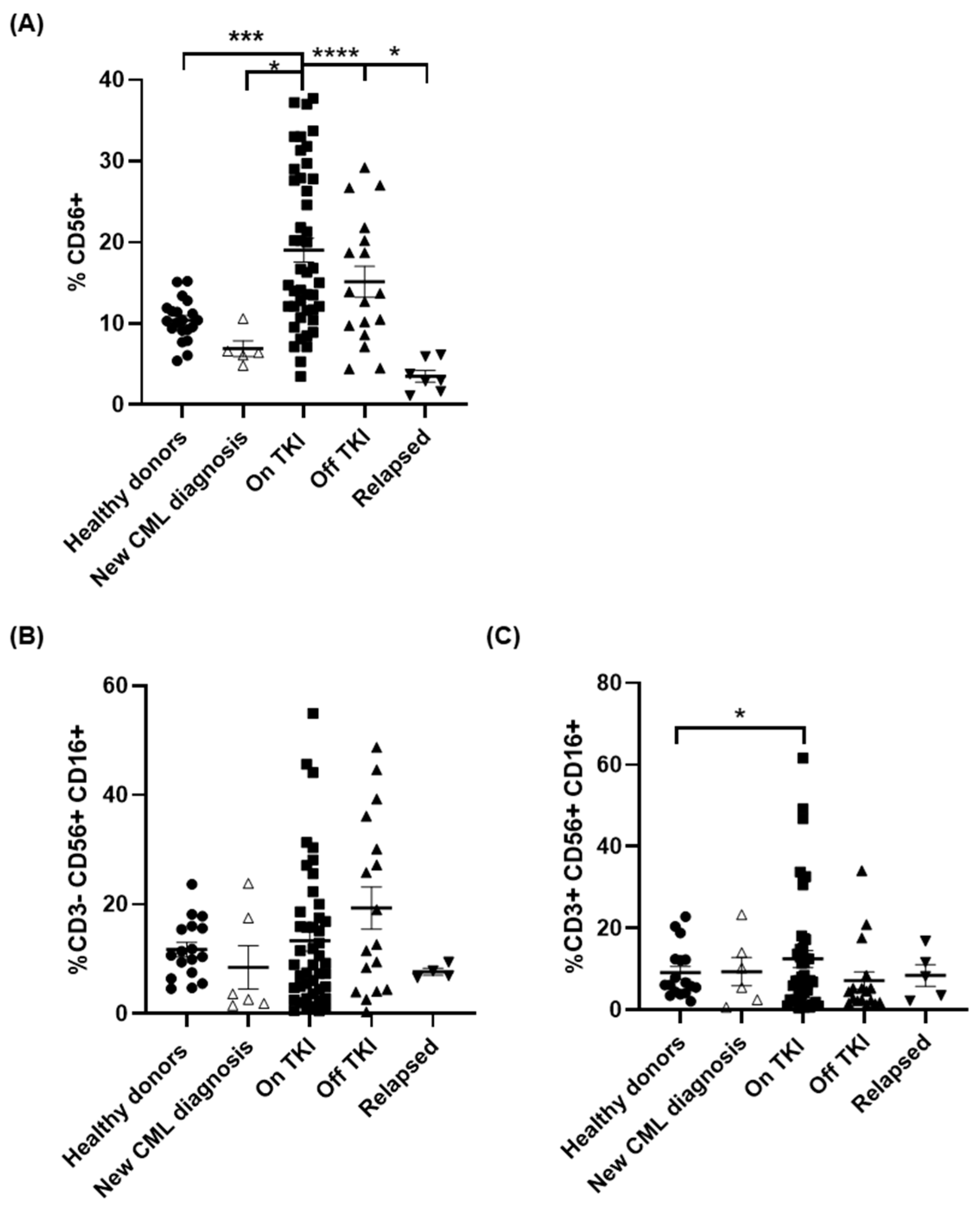
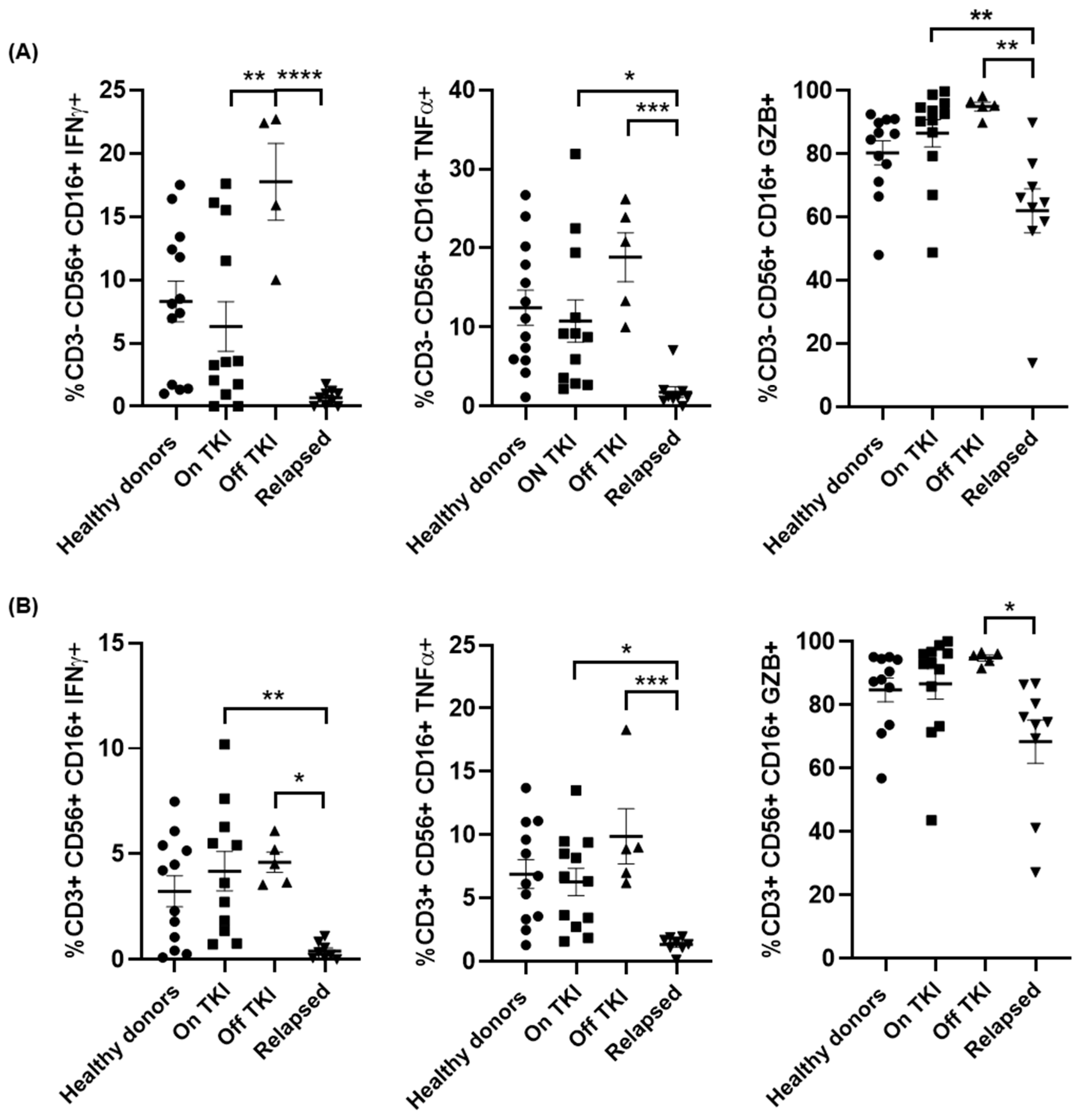
3.2. Profile and Cytokine Production of NK Cell Populations Induced by Treatment with TKIs
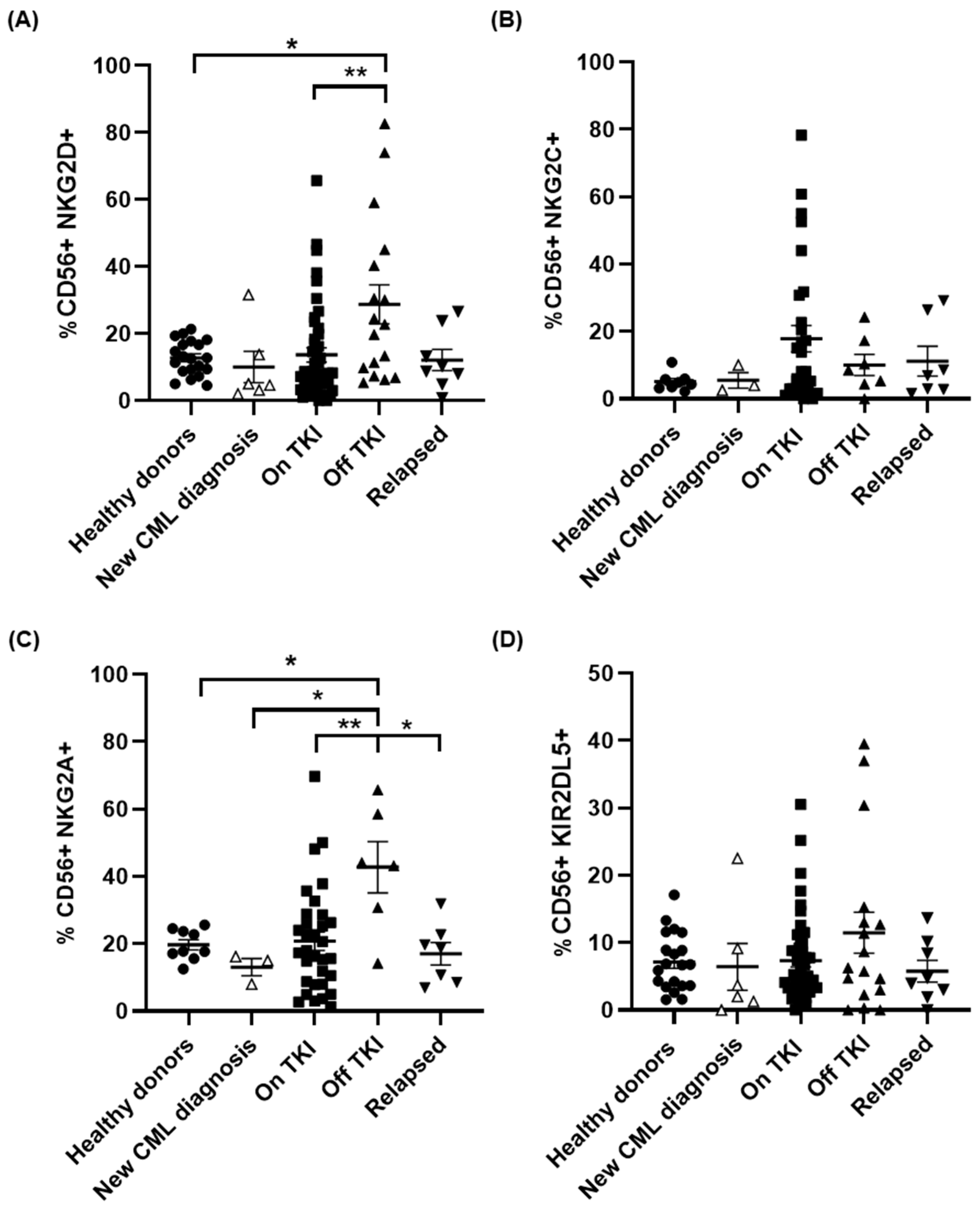
3.3. Differential Expression of Stimulatory and Inhibitory NK Markers
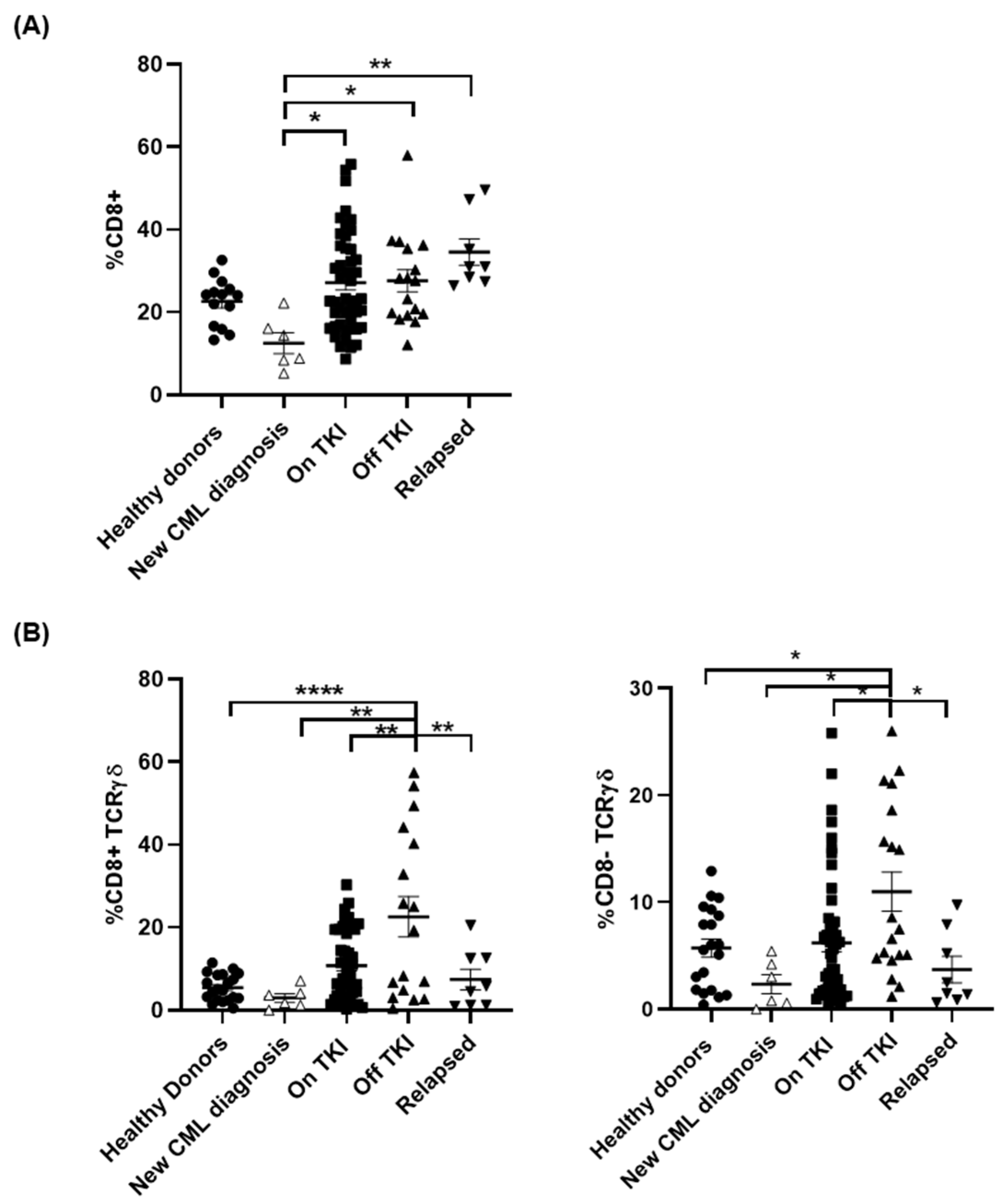
3.4. Treatment with TKIs Increased CD8± TCRγβ+ T Cell Populations
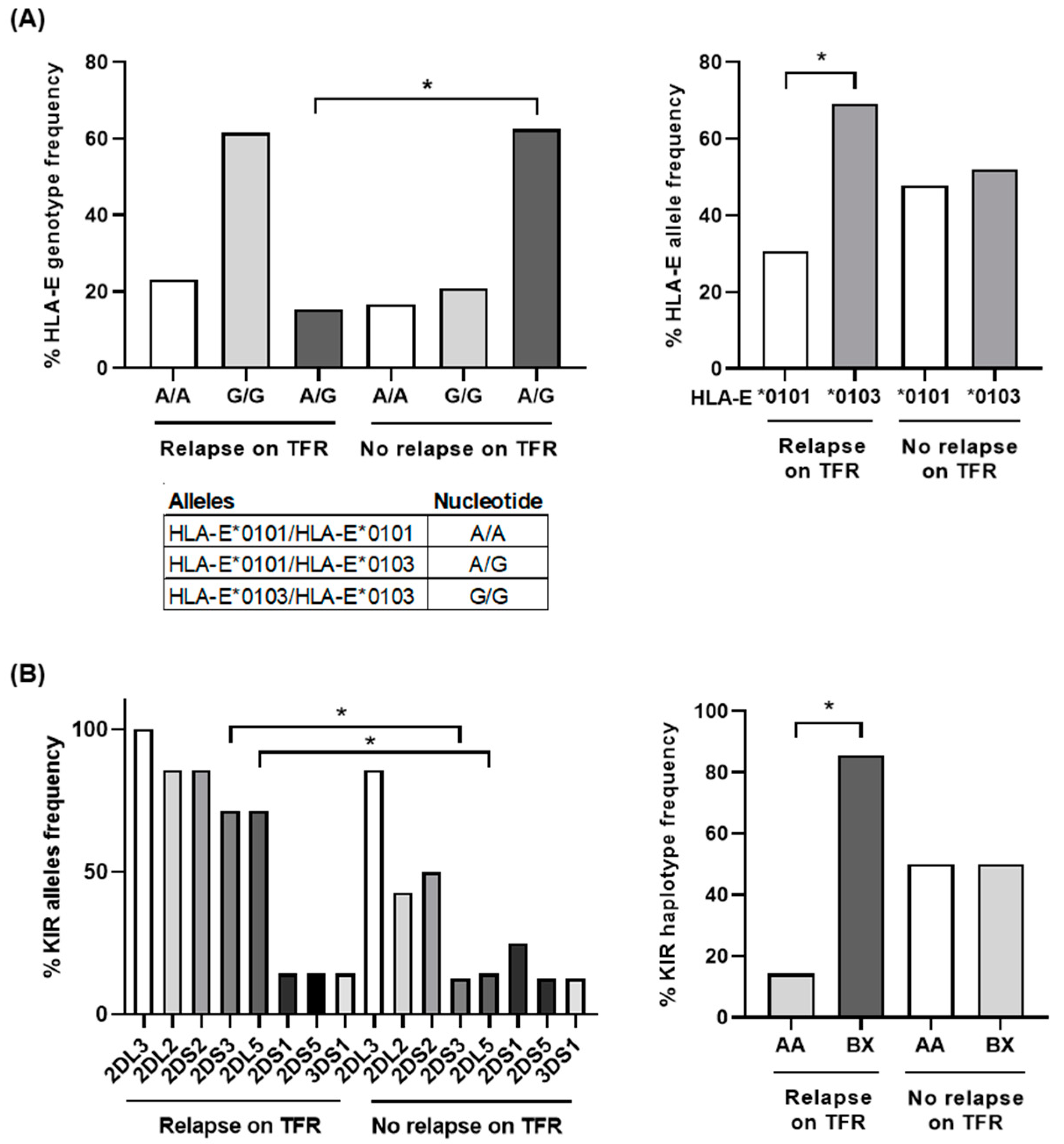
3.5. Predominant HLA-E Homozygosis and KIR Haplotype BX in Patients Who Relapsed during TFR
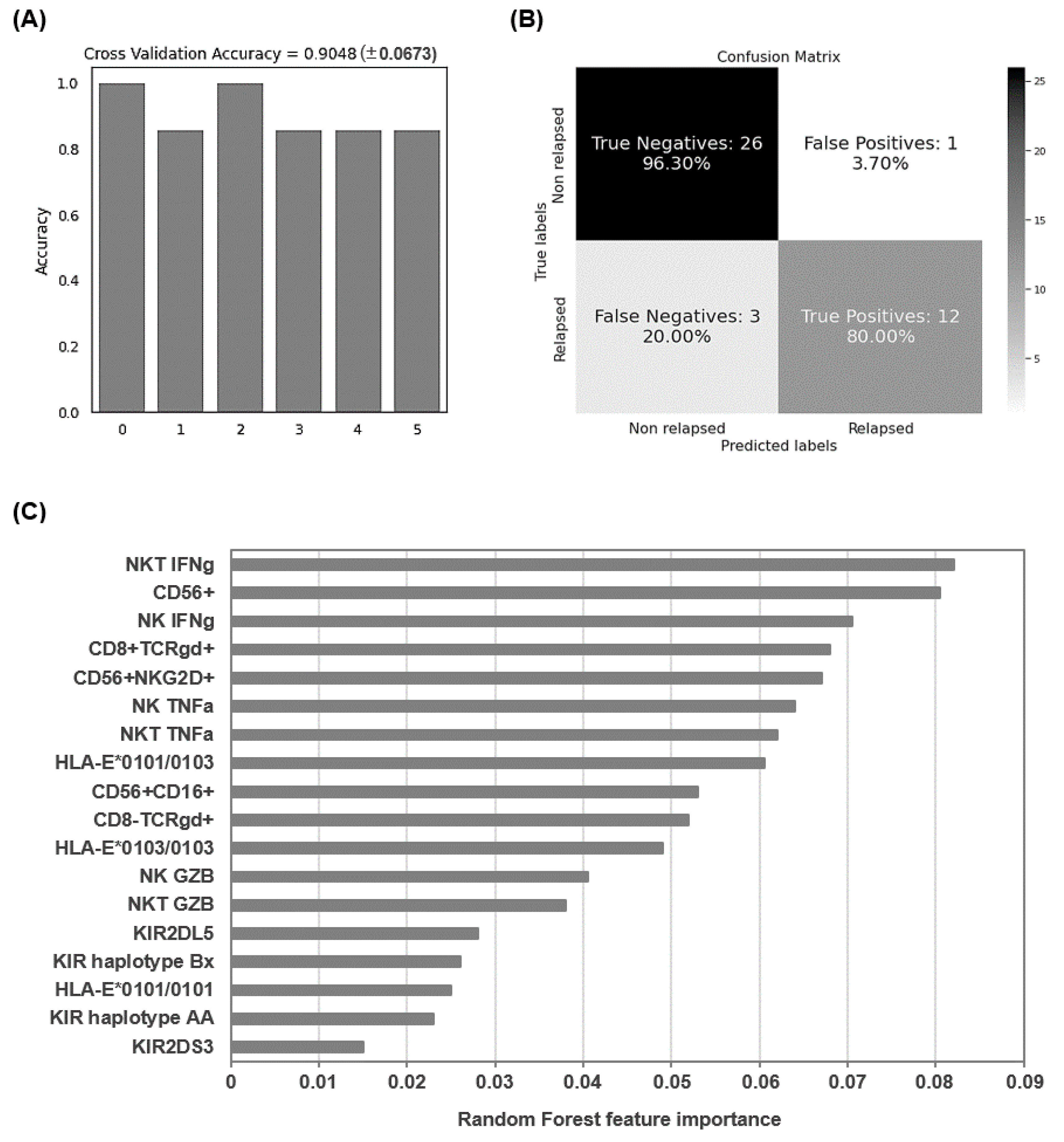
3.6. Application of Random Forest Algorithm
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Acknowledgments
Conflicts of Interest
References
- Rowley, J.D. The Philadelphia chromosome translocation. A paradigm for understanding leukemia. Cancer 1990, 65, 2178–2184. [Google Scholar] [CrossRef]
- D’Antonio, J. Chronic myelogenous leukemia. Clin. J. Oncol. Nurs. 2005, 9, 535–538. [Google Scholar] [CrossRef]
- Siegel, R.L.; Fedewa, S.A.; Miller, K.D.; Goding-Sauer, A.; Pinheiro, P.S.; Martinez-Tyson, D.; Jemal, A. Cancer statistics for Hispanics/Latinos, 2015. CA Cancer J. Clin. 2015, 65, 457–480. [Google Scholar] [CrossRef] [PubMed]
- Efficace, F.; Baccarani, M.; Breccia, M.; Alimena, G.; Rosti, G.; Cottone, F.; Lambertenghi Deliliers, G.; Baratè, C.; Russo Rossi, A.; Fioritoni, G.; et al. Health-related quality of life in chronic myeloid leukemia patients receiving long-term therapy with imatinib compared with the general population. Blood 2011, 118, 4554–4560. [Google Scholar] [CrossRef] [PubMed]
- Nowell, P.C.H.D. A minute chromosome in human chronic granulocytic leukemia. Science 1960, 132, 1497. [Google Scholar]
- Etienne, G.; Guilhot, J.; Rea, D.; Rigal-Huguet, F.; Nicolini, F.; Charbonnier, A.; Guerci-Bresler, A.; Legros, L.; Varet, B.; Gardembas, M.; et al. Long-Term Follow-Up of the French Stop Imatinib (STIM1) Study in Patients with Chronic Myeloid Leukemia. J. Clin. Oncol. 2017, 35, 298–305. [Google Scholar] [CrossRef]
- Saussele, S.; Richter, J.; Guilhot, J.; Gruber, F.X.; Hjorth-Hansen, H.; Almeida, A.; Janssen, J.; Mayer, J.; Koskenvesa, P.; Panayiotidis, P.; et al. Discontinuation of tyrosine kinase inhibitor therapy in chronic myeloid leukaemia (EURO-SKI): A prespecified interim analysis of a prospective, multicentre, non-randomised, trial. Lancet Oncol. 2018, 19, 747–757. [Google Scholar] [CrossRef]
- Okada, M.; Imagawa, J.; Tanaka, H.; Nakamae, H.; Hino, M.; Murai, K.; Ishida, Y.; Kumagai, T.; Sato, S.; Ohashi, K.; et al. Final 3-year Results of the Dasatinib Discontinuation Trial in Patients With Chronic Myeloid Leukemia Who Received Dasatinib as a Second-line Treatment. Clin. Lymphoma Myeloma Leuk. 2018, 18, 353–360 e1. [Google Scholar] [CrossRef]
- Mahon, F.X.; Réa, D.; Guilhot, J.; Guilhot, F.; Huguet, F.; Nicolini, F.; Legros, L.; Charbonnier, A.; Guerci, A.; Varet, B.; et al. Discontinuation of imatinib in patients with chronic myeloid leukaemia who have maintained complete molecular remission for at least 2 years: The prospective, multicentre Stop Imatinib (STIM) trial. Lancet Oncol. 2010, 11, 1029–1035. [Google Scholar] [CrossRef]
- Rousselot, P.; Charbonnier, A.; Cony-Makhoul, P.; Agape, P.; Nicolini, F.E.; Varet, B.; Gardembas, M.; Etienne, G.; Réa, D.; Roy, L.; et al. Loss of major molecular response as a trigger for restarting tyrosine kinase inhibitor therapy in patients with chronic-phase chronic myelogenous leukemia who have stopped imatinib after durable undetectable disease. J. Clin. Oncol. 2014, 32, 424–430. [Google Scholar] [CrossRef]
- Sweet, K.; Zhang, L.; Pinilla-Ibarz, J. Biomarkers for determining the prognosis in chronic myelogenous leukemia. J. Hematol. Oncol. 2013, 6, 54. [Google Scholar] [CrossRef] [PubMed]
- Hahnel, T.; Baldow, C.; Guilhot, J.; Guilhot, F.; Saussele, S.; Mustjoki, S.; Jilg, S.; Jost, P.J.; Dulucq, S.; Xavier Mahon, F.X.; et al. Model-Based Inference and Classification of Immunologic Control Mechanisms from TKI Cessation and Dose Reduction in Patients with CML. Cancer Res. 2020, 80, 2394–2406. [Google Scholar]
- Bernardi, S.; Malagola, M.; Zanaglio, C.; Polverelli, N.; Dereli Eke, E.; D’Adda, M.; Farina, M.; Bucelli, C.; Scaffidi, L.; Toffoletti, E.; et al. Digital PCR improves the quantitation of DMR and the selection of CML candidates to TKIs discontinuation. Cancer Med. 2019, 8, 2041–2055. [Google Scholar] [CrossRef] [PubMed]
- Zanaglio, C.; Bernardi, S.; Gandolfi, L.; Farina, M.; Re, F.; Polverelli, N.; Zollner, T.; Turra, A.; Morello, E.; Malagola, M.; et al. RT-qPCR versus Digital PCR: How Do They Impact Differently on Clinical Management of Chronic Myeloid Leukemia Patients? Case Rep. Oncol. 2020, 13, 1263–1269. [Google Scholar] [CrossRef] [PubMed]
- Bocchia, M.; Defina, M.; Aprile, L.; Ippoliti, M.; Crupi, R.; Rondoni, M.; Gozzetti, A.; Lauria, F. Complete molecular response in CML after p210 BCR-ABL1-derived peptide vaccination. Nat. Rev. Clin. Oncol. 2010, 7, 600–603. [Google Scholar] [CrossRef]
- Hayashi, Y.; Nakamae, H.; Katayama, T.; Nakane, T.; Koh, H.; Nakamae, M.; Hirose, A.; Hagihara, K.; Terada, Y.; Nakao, Y.; et al. Different immunoprofiles in patients with chronic myeloid leukemia treated with imatinib, nilotinib or dasatinib. Leuk. Lymphoma 2012, 53, 1084–1089. [Google Scholar] [CrossRef] [PubMed]
- Ilander, M.; Olsson-Stromberg, U.; Schlums, H.; Guilhot, J.; Bruck, O.; Lahteenmaki, H.; Kasanen, T.; Koskenvesa, P.; Söderlund, S.; Höglund, M.; et al. Increased proportion of mature NK cells is associated with successful imatinib discontinuation in chronic myeloid leukemia. Leukemia 2017, 31, 1108–1116. [Google Scholar] [CrossRef]
- Ishiyama, K.; Kitawaki, T.; Sugimoto, N.; Sozu, T.; Anzai, N.; Okada, M.; Nohgawa, M.; Hatanaka, K.; Arima, N.; Ishikawa, T.; et al. Principal component analysis uncovers cytomegalovirus-associated NK cell activation in Ph(+) leukemia patients treated with dasatinib. Leukemia 2017, 31, 268. [Google Scholar] [CrossRef]
- Climent, N.; Plana, M. Immunomodulatory Activity of Tyrosine Kinase Inhibitors to Elicit Cytotoxicity against Cancer and Viral Infection. Front. Pharm. 2019, 10, 1232. [Google Scholar] [CrossRef]
- El Missiry, M.; Adnan Awad, S.; Rajala, H.L.; Al-Samadi, A.; Ekblom, M.; Markevan, B.; Åstrand-Grundström, I.; Wold, M.; Rabben Svedahl, E.; Ravn Juhl, B.; et al. Assessment of bone marrow lymphocytic status during tyrosine kinase inhibitor therapy and its relation to therapy response in chronic myeloid leukaemia. J. Cancer Res. Clin. Oncol. 2016, 142, 1041–1050. [Google Scholar] [CrossRef]
- Dulucq, S.; Mahon, F.X. Deep molecular responses for treatment-free remission in chronic myeloid leukemia. Cancer Med. 2016, 5, 2398–2411. [Google Scholar] [CrossRef] [PubMed]
- Ross, D.M.; Masszi, T.; Gomez Casares, M.T.; Hellmann, A.; Stentoft, J.; Conneally, E.; Garcia-Gutierrez, V.; Gattermann, N.; D le Coutre, P.; Martino, B.; et al. Durable treatment-free remission in patients with chronic myeloid leukemia in chronic phase following frontline nilotinib: 96-week update of the ENESTfreedom study. J. Cancer Res. Clin. Oncol. 2018, 144, 945–954. [Google Scholar] [CrossRef] [PubMed]
- Lanier, L.L. NK cell recognition. Annu. Rev. Immunol. 2005, 23, 225–274. [Google Scholar] [CrossRef] [PubMed]
- Chang, M.C.; Cheng, H.I.; Hsu, K.; Hsu, Y.N.; Kao, C.W.; Chang, Y.F.; Lim, K.H.; Chen, C.G. NKG2A Down-Regulation by Dasatinib Enhances Natural Killer Cytotoxicity and Accelerates Effective Treatment Responses in Patients With Chronic Myeloid Leukemia. Front. Immunol. 2018, 9, 3152. [Google Scholar] [CrossRef] [PubMed]
- Cichocki, F.; Cooley, S.; Davis, Z.; DeFor, T.E.; Schlums, H.; Zhang, B.; Brunstein, C.G.; Blazar, B.R.; Wagner, J.; Diamond, D.J.; et al. CD56dimCD57+NKG2C+ NK cell expansion is associated with reduced leukemia relapse after reduced intensity HCT. Leukemia 2016, 30, 456–463. [Google Scholar] [CrossRef]
- Wu, G.Q.; Zhao, Y.M.; Lai, X.Y.; Luo, Y.; Tan, Y.M.; Shi, J.M.; Li, L.; Zheng, W.Y.; Zhang, J.; Hu, X.R.; et al. The beneficial impact of missing KIR ligands and absence of donor KIR2DS3 gene on outcome following unrelated hematopoietic SCT for myeloid leukemia in the Chinese population. Bone Marrow. Transpl. 2010, 45, 1514–1521. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Sugioka, D.K.; Gonçalves, C.E.; Bicalho, M.D. KIR repertory in patients with hematopoietic diseases and healthy family members. BMC Hematol. 2016, 16, 25. [Google Scholar] [CrossRef]
- Farstad, I.N.; Halstensen, T.S.; Lien, B.; Kilshaw, P.J.; Lazarovits, A.I.; Brandtzaeg, P. Distribution of beta 7 integrins in human intestinal mucosa and organized gut-associated lymphoid tissue. Immunology 1996, 89, 227–237. [Google Scholar] [CrossRef]
- Grant, E.J.; Nguyen, A.T.; Lobos, C.A.; Szeto, C.; Chatzileontiadou, D.S.M.; Gras, S. The unconventional role of HLA-E: The road less traveled. Mol. Immunol. 2020, 120, 101–112. [Google Scholar] [CrossRef]
- D’Asaro, M.; La Mendola, C.; Di Liberto, D.; Orlando, V.; Todaro, M.; Spina, M.; Guggino, G.; Meraviglia, S.; Caccamo, N.; Messina, A.; et al. V gamma 9V delta 2 T lymphocytes efficiently recognize and kill zoledronate-sensitized, imatinib-sensitive, and imatinib-resistant chronic myelogenous leukemia cells. J. Immunol. 2010, 184, 3260–3268. [Google Scholar] [CrossRef]
- Poccia, F.; Cipriani, B.; Vendetti, S.; Colizzi, V.; Poquet, Y.; Battistini, L.; López-Botet, M.; Fournié, J.J.; Gougeon, M.L. CD94/NKG2 inhibitory receptor complex modulates both anti-viral and anti-tumoral responses of polyclonal phosphoantigen-reactive V gamma 9V delta 2 T lymphocytes. J. Immunol. 1997, 159, 6009–6017. [Google Scholar] [PubMed]
- Irani, Y.D.; Hughes, A.; Clarson, J.; Kok, C.H.; Shanmuganathan, N.; White, D.L.; Yeung, D.T.; Ross, D.M.; Hughes, T.P.; Yong, A.S.M. Successful treatment-free remission in chronic myeloid leukaemia and its association with reduced immune suppressors and increased natural killer cells. Br. J. Haematol. 2020, 191, 433–441. [Google Scholar] [CrossRef] [PubMed]
- Smirnikhina, S.A.; Lavrov, A.V.; Chelysheva, E.Y.; Adilgereeva, E.P.; Shukhov, O.A.; Turkina, A.; Kutsev, S.I. Whole-exome sequencing reveals potential molecular predictors of relapse after discontinuation of the targeted therapy in chronic myeloid leukemia patients. Leuk. Lymphoma 2016, 57, 1669–1676. [Google Scholar] [CrossRef] [PubMed]
- Söderlund, S.; Persson, I.; Ilander, M.; Guilhot, J.; Hjorth-Hansen, H.; Koskenvesa, P.; Adilgereeva, E.P.; Kutsev, S.I. Plasma proteomics of biomarkers for inflammation or cancer cannot predict relapse in chronic myeloid leukaemia patients stopping tyrosine kinase inhibitor therapy. Leuk. Res. 2020, 90, 106310. [Google Scholar] [CrossRef] [PubMed]
- Takahashi, N.; Kyo, T.; Maeda, Y.; Sugihara, T.; Usuki, K.; Kawaguchi, T.; Usui, N.; Okamoto, S.; Ohe, Y.; Ohtake, S.; et al. Discontinuation of imatinib in Japanese patients with chronic myeloid leukemia. Haematologica 2012, 97, 903–906. [Google Scholar] [CrossRef]
- Ross, D.M.; Branford, S.; Seymour, J.F.; Schwarer, A.P.; Arthur, C.; Yeung, D.T.; Dang, P.; Goyne, J.M.; Slader, C.; Filshie, R.J.; et al. Safety and efficacy of imatinib cessation for CML patients with stable undetectable minimal residual disease: Results from the TWISTER study. Blood 2013, 122, 515–522. [Google Scholar] [CrossRef]
- Gonzalez-Galarza, F.F.; McCabe, A.; Santos, E.; Jones, J.; Takeshita, L.; Ortega-Rivera, N.D.; Del Cid-Pavon, G.G.; Ramsbottom, K.; Ghattaoraya, G.; Alfirevic, A.; et al. Allele frequency net database (AFND) 2020 update: Gold-standard data classification, open access genotype data and new query tools. Nucleic Acids Res. 2020, 48, D783–D788. [Google Scholar] [CrossRef]
- Breiman, L. Random forests. Mach. Learn. 1991, 45, 5–32. [Google Scholar] [CrossRef]
- Mayer, J.R.R.; Ghosh, S.; Pal, R. Sequential feature selection and inference using multi-variate random forests. Bioinformatics 2018, 34, 1336–1344. [Google Scholar] [CrossRef]
- Breiman, L. Random Forests; Statistics Department, University of California: Berkely, Calif, USA, 2001; Available online: http://ozberkeleyedu/users/breiman/randomforest2001pdf (accessed on 21 September 2020).
- Hastie, T.T.R.; Friedman, J. The Elements of Statistical Learning; Springer: Berlin/Heidelberg, Germany, 2009. [Google Scholar]
- Sarica, A.; Cerasa, A.; Quattrone, A. Random Forest Algorithm for the Classification of Neuroimaging Data in Alzheimer’s Disease: A Systematic Review. Front. Aging Neurosci. 2017, 9, 329. [Google Scholar] [CrossRef]
- Nembrini, S.; Konig, I.R.; Wright, M.N. The revival of the Gini importance? Bioinformatics 2018, 34, 3711–3718. [Google Scholar] [CrossRef] [PubMed]
- Gastpar, R.; Gehrmann, M.; Bausero, M.A.; Asea, A.; Gross, C.; Schroeder, J.A.; Multhoff, G. Heat shock protein 70 surface-positive tumor exosomes stimulate migratory and cytolytic activity of natural killer cells. Cancer Res. 2005, 65, 5238–5247. [Google Scholar] [CrossRef] [PubMed]
- Khoury, H.J.; Williams, L.A.; Atallah, E.; Hehlmann, R. Chronic Myeloid Leukemia: What Every Practitioner Needs to Know in 2017. Am. Soc. Clin. Oncol. Educ. Book 2017, 37, 468–479. [Google Scholar] [CrossRef] [PubMed]
- Kalmanti, L.; Saussele, S.; Lauseker, M.; Proetel, U.; Müller, M.C.; Hanfstein, B.; Schreiber, A.; Fabarius, A.; Pfirrmann, M.; Schnittger, S.; et al. Younger patients with chronic myeloid leukemia do well in spite of poor prognostic indicators: Results from the randomized CML study IV. Ann. Hematol. 2013, 93, 71–80. [Google Scholar] [CrossRef]
- Fava, C.; Rege-Cambrin, G.; Dogliotti, I.; Cerrano, M.; Berchialla, P.; Dragani, M.; Rosti, G.; Castagnetti, F.; Gugliotta, G.; Martino, B.; et al. Observational study of chronic myeloid leukemia Italian patients who discontinued tyrosine kinase inhibitors in clinical practice. Haematologica 2019, 104, 1589–1596. [Google Scholar] [CrossRef]
- Shah, N.P.; Garcia-Gutierrez, V.; Jimenez-Velasco, A.; Larson, S.; Saussele, S.; Rea, D.; Mahon, F.X.; Levy, M.Y.; Gómez-Casares, M.T.; Pane, F.; et al. Dasatinib discontinuation in patients with chronic-phase chronic myeloid leukemia and stable deep molecular response: The DASFREE study. Leuk. Lymphoma 2020, 61, 650–659. [Google Scholar] [CrossRef]
- Baccarani, M.; Abruzzese, E.; Accurso, V.; Albano, F.; Annunziata, M.; Barulli, S.; Beltrami, G.; Bergamaschi, M.; Binotto, G.; Bocchia, M.; et al. Managing chronic myeloid leukemia for treatment-free remission: A proposal from the GIMEMA CML WP. Blood Adv. 2019, 3, 4280–4290. [Google Scholar] [CrossRef]
- Kreutzman, A.; Juvonen, V.; Kairisto, V.; Ekblom, M.; Stenke, L.; Seggewiss, R.; Porkka, K.; Mustjoki, S. Mono/oligoclonal T and NK cells are common in chronic myeloid leukemia patients at diagnosis and expand during dasatinib therapy. Blood 2010, 116, 772–782. [Google Scholar] [CrossRef]
- Kreutzman, A.; Ladell, K.; Koechel, C.; Gostick, E.; Ekblom, M.; Stenke, L.; Melo, T.; Einsele, H.; Porkka, K.; Price, D.A.; et al. Expansion of highly differentiated CD8+ T-cells or NK-cells in patients treated with dasatinib is associated with cytomegalovirus reactivation. Leukemia 2011, 25, 1587–1597. [Google Scholar] [CrossRef]
- Mustjoki, S.; Auvinen, K.; Kreutzman, A.; Rousselot, P.; Hernesniemi, S.; Melo, T.; Lahesmaa-Korpinen, A.M.; Hautaniemi, S.; Bouchet, S.; Molimard, M.; et al. Rapid mobilization of cytotoxic lymphocytes induced by dasatinib therapy. Leukemia 2013, 27, 914–924. [Google Scholar] [CrossRef]
- Schiffer, C.A.; Cortes, J.E.; Hochhaus, A.; Saglio, G.; le Coutre, P.; Porkka, K.; Mustjoki, S.; Mohamed, H.; Shah, N.P. Lymphocytosis after treatment with dasatinib in chronic myeloid leukemia: Effects on response and toxicity. Cancer 2016, 122, 1398–1407. [Google Scholar] [CrossRef] [PubMed]
- Rivera-Torres, J.; San José, E. Src Tyrosine Kinase Inhibitors: New Perspectives on Their Immune, Antiviral, and Senotherapeutic Potential. Front. Pharm. 2019, 10, 1011. [Google Scholar] [CrossRef] [PubMed]
- Hassold, N.; Seystahl, K.; Kempf, K.; Urlaub, D.; Zekl, M.; Einsele, H.; Watzl, C.; Wischhusen, J.; Seggewiss-Bernhardt, R. Enhancement of natural killer cell effector functions against selected lymphoma and leukemia cell lines by dasatinib. Int. J. Cancer 2012, 131, E916–E927. [Google Scholar] [CrossRef] [PubMed]
- Bauer, S.; Groh, V.; Wu, J.; Steinle, A.; Phillips, J.H.; Lanier, L.L.; Spies, T. Activation of NK cells and T cells by NKG2D, a receptor for stress-inducible MICA. Science 1999, 285, 727–729. [Google Scholar] [CrossRef] [PubMed]
- Danier, A.C.; de Melo, R.P.; Napimoga, M.H.; Laguna-Abreu, M.T. The role of natural killer cells in chronic myeloid leukemia. Rev. Bras. Hematol. Hemoter 2011, 33, 216–220. [Google Scholar] [CrossRef][Green Version]
- Molldrem, J.J.; Lee, P.P.; Kant, S.; Wieder, E.; Jiang, W.; Lu, S.; Wang, C.; Davis, M.M. Chronic myelogenous leukemia shapes host immunity by selective deletion of high-avidity leukemia-specific T cells. J. Clin. Investig. 2003, 111, 639–647. [Google Scholar] [CrossRef]
- Mumprecht, S.; Schürch, C.; Scherrer, S.; Claus, C.; Ochsenbein, A.F. Chronic myelogenous leukemia maintains specific CD8(+) T cells through IL-7 signaling. Eur. J. Immunol. 2010, 40, 2720–2730. [Google Scholar] [CrossRef]
- Halim, L.; Parente-Pereira, A.C.; Maher, J. Prospects for immunotherapy of acute myeloid leukemia using γδ T cells. Immunotherapy 2017, 9, 111–114. [Google Scholar] [CrossRef]
- Pietra, G.; Romagnani, C.; Moretta, L.; Mingari, M.C. HLA-E and HLA-E-bound peptides: Recognition by subsets of NK and T cells. Curr. Pharm. Des. 2009, 15, 3336–3344. [Google Scholar] [CrossRef]
- Saikh, K.U.; Kissner, T.; Ulrich, R.G. Regulation of HLA-DR and co-stimulatory molecule expression on natural killer T cells by granulocyte-macrophage colony-stimulating factor. Immunology 2002, 106, 363–372. [Google Scholar] [CrossRef]
- Auton, A.; Brooks, L.D.; Durbin, R.M.; Garrison, E.P.; Kang, H.M.; Korbel, J.O.; Marchini, J.L.; McCarthy, S.; McVean, G.A. A global reference for human genetic variation. Nature 2015, 526, 68–74. [Google Scholar] [PubMed]
- Xu, Y.P.; Wieten, L.; Wang, S.X.; Cai, Y.; Olieslagers, T.; Zhang, L.; He, L.L.; Tilanus, M.G.J.; Hong, W.X. Clinical significance of HLA-E genotype and surface/soluble expression levels between healthy individuals and patients with acute leukemia. Leuk. Lymphoma 2019, 60, 208–215. [Google Scholar] [CrossRef] [PubMed]
- Zheng, H.; Lu, R.; Xie, S.; Wen, X.; Wang, H.; Gao, X.; Guo, L. Human leukocyte antigen-E alleles and expression in patients with serous ovarian cancer. Cancer Sci. 2015, 106, 522–528. [Google Scholar] [CrossRef] [PubMed]
- Wagner, B.; da Silva Nardi, F.; Schramm, S.; Kraemer, T.; Celik, A.A.; Dürig, J.; Horn, P.A.; Dührsen, U.; Nückel, H.; Rebmann, V.L. HLA-E allelic genotype correlates with HLA-E plasma levels and predicts early progression in chronic lymphocytic leukemia. Cancer 2017, 123, 814–823. [Google Scholar] [CrossRef]
- Kanevskiy, L.; Erokhina, S.; Kobyzeva, P.; Streltsova, M.; Sapozhnikov, A.; Kovalenko, E. Dimorphism of HLA-E and its Disease Association. Int. J. Mol. Sci. 2019, 20, 5496. [Google Scholar] [CrossRef]
- Strong, R.K.; Holmes, M.A.; Li, P.; Braun, L.; Lee, N.; Geraghty, D.E. HLA-E allelic variants. Correlating differential expression, peptide affinities, crystal structures, and thermal stabilities. J. Biol. Chem. 2003, 278, 5082–5090. [Google Scholar] [CrossRef]
- Kraemer, T.; Celik, A.A.; Huyton, T.; Kunze-Schumacher, H.; Blasczyk, R.; Bade-Döding, C. HLA-E: Presentation of a Broader Peptide Repertoire Impacts the Cellular Immune Response-Implications on HSCT Outcome. Stem. Cells Int. 2015, 2015, 346714. [Google Scholar] [CrossRef]
- Hirankarn, N.; Kimkong, I.; Mutirangura, A. HLA-E polymorphism in patients with nasopharyngeal carcinoma. Tissue Antigens 2004, 64, 588–592. [Google Scholar] [CrossRef]
- Carrington, M.; O’Brien, S.J. The influence of HLA genotype on AIDS. Annu. Rev. Med. 2003, 54, 535–551. [Google Scholar] [CrossRef]
- Coupel, S.; Moreau, A.; Hamidou, M.; Horejsi, V.; Soulillou, J.P.; Charreau, B. Expression and release of soluble HLA-E is an immunoregulatory feature of endothelial cell activation. Blood 2007, 109, 2806–2814. [Google Scholar] [CrossRef]
- Cisneros, E.; Moraru, M.; Gómez-Lozano, N.; López-Botet, M.; Vilches, C. KIR2DL5: An Orphan Inhibitory Receptor Displaying Complex Patterns of Polymorphism and Expression. Front. Immunol. 2012, 3, 289. [Google Scholar] [CrossRef] [PubMed]
- Yeung, D.T.; Tang, C.; Vidovic, L.; White, D.L.; Branford, S.; Hughes, T.P.; Yong, A.S. KIR2DL5B genotype predicts outcomes in CML patients treated with response-directed sequential imatinib/nilotinib strategy. Blood 2015, 126, 2720–2723. [Google Scholar] [CrossRef] [PubMed]
- Wilson, M.J.; Torkar, M.; Haude, A.; Milne, S.; Jones, T.; Sheer, D.; Beck, S.; Trowsdale, J. Plasticity in the organization and sequences of human KIR/ILT gene families. Proc. Natl. Acad. Sci. USA 2000, 97, 4778–4783. [Google Scholar] [CrossRef] [PubMed]
- Caocci, G.; Martino, B.; Greco, M.; Abruzzese, E.; Trawinska, M.M.; Lai, S.; Lai, S.; Ragatzu, P.; Galimberti, S.; Baratè, C.; et al. Killer immunoglobulin-like receptors can predict TKI treatment-free remission in chronic myeloid leukemia patients. Exp. Hematol. 2015, 43, 1015–1018.e1. [Google Scholar] [CrossRef] [PubMed]
- Dumas, P.Y.; Bérard, E.; Bréal, C.; Dulucq, S.; Réa, D.; Nicolini, F.; Forcade, E.; Dufossée, M.; Pasquet, J.M.; Turcq, B.; et al. Killer immunoglobulin-like receptor genotypes and chronic myeloid leukemia outcomes after imatinib cessation for treatment-free remission. Cancer Med. 2019, 8, 4976–4985. [Google Scholar] [CrossRef]
- Hughes, A.; Clarson, J.; Tang, C.; Vidovic, L.; White, D.L.; Hughes, T.P.; Yong, A.S.M. CML patients with deep molecular responses to TKI have restored immune effectors and decreased PD-1 and immune suppressors. Blood 2017, 129, 1166–1176. [Google Scholar] [CrossRef]
| New CML Diagnosis | CML on TKI | CML off TKI | CML Relapsed | ||
|---|---|---|---|---|---|
| Patients (n) | 6 | 45 | 27 | 15 | |
| Male/female (n) | 3/3 | 22/23 | 18/9 | 7/8 | |
| Median age (years) | 54.5y (IQR 44.7 to 64) | 49.0y (IQR 37.5 to 59) | 48.5y (IQR 39.7 to 65) | 38.0y (IQR 27 to 57) | |
| Sokal Risk Score (L/I/H/UD) * (n) | 3/2/1/0 | 34/6/5/0 | 17/7/0/3 | 13/2/0/0 | |
| Response at sampling ** (n) | CCyR | N/A | 4 | 0 | 10 |
| MMR | N/A | 8 | 0 | 0 | |
| DMR | N/A | 33 | 27 | 5 | |
| UD | N/A | 0 | 0 | 0 | |
| Patient’ Code | Gender (M/F) | Age at Diagnosis (Years) | Sokal Risk | CML Phase: Chronic Accelerated Blast | Hematological Remission (Y/N) | Cytogenetic Remission (Y/N) | Molecular Response (Log) | Last TKI | Time on Treatment with Last TKI (Months) | Time on TFR (Months) | |
|---|---|---|---|---|---|---|---|---|---|---|---|
| New CML diagnosis | 1 | F | 44 | Low | Chronic | N/A | N/A | N/A | - | - | - |
| 2 | M | 56 | High | Chronic | N/A | N/A | N/A | - | - | - | |
| 3 | F | 45 | Low | Chronic | N/A | N/A | N/A | - | - | - | |
| 4 | M | 57 | Intermediate | Chronic | N/A | N/A | N/A | - | - | - | |
| 5 | M | 85 | Intermediate | Chronic | N/A | N/A | N/A | - | - | - | |
| 6 | F | 53 | Low | Chronic | N/A | N/A | N/A | - | - | - | |
| CML On treatment with TKI | 7 | M | 35 | Low | Chronic | Yes | Yes | 5 | Imatinib | 96 | - |
| 8 | F | 38 | Low | Chronic | Yes | Yes | 4 | Imatinib | 14.4 | - | |
| 9 | F | 62 | Low | Chronic | Yes | Yes | 4 | Imatinib | 24 | - | |
| 10 | F | 15 | Low | Chronic | Yes | Yes | 4.5 | Imatinib | 12 | - | |
| 11 | M | 52 | Low | Chronic | Yes | Yes | 5 | Imatinib | 168 | - | |
| 12 | F | 56 | Low | Chronic | Yes | Yes | 5 | Imatinib | 156 | - | |
| 13 | F | 52 | Intermediate | Chronic | Yes | Yes | 5 | Imatinib | 32.4 | - | |
| 14 | M | 75 | Intermediate | Chronic | Yes | Yes | 3 | Imatinib | 6 | - | |
| 15 | M | 65 | Low | Chronic | Yes | Yes | 5 | Imatinib | 15.6 | - | |
| 16 | M | 38 | Low | Chronic | Yes | Yes | 5 | Imatinib | 132 | - | |
| 17 | M | 34 | Low | Chronic | Yes | Yes | 5 | Imatinib | 40.8 | - | |
| 18 | F | 49 | Low | Chronic | Yes | Yes | 4 | Dasatinib | 30 | - | |
| 19 | F | 27 | Low | Chronic | Yes | Yes | 4.5 | Dasatinib | 9.6 | - | |
| 20 | M | 55 | Low | Chronic | Yes | Yes | 3 | Dasatinib | 30 | - | |
| 21 | F | 41 | Low | Chronic | Yes | Yes | No | Dasatinib | 1.2 | - | |
| 22 | M | 24 | Low | Chronic | Yes | Yes | 3 | Dasatinib | 84 | - | |
| 23 | F | 57 | Intermediate | Chronic | Yes | Yes | 3 | Dasatinib | 34.8 | - | |
| 24 | F | 59 | Low | Chronic | Yes | Yes | 4 | Dasatinib | 30 | - | |
| 25 | F | 57 | Low | Chronic | Yes | Yes | 3 | Dasatinib | 72 | - | |
| 26 | M | 39 | High | Chronic | Yes | Yes | No | Dasatinib | 1.2 | - | |
| 27 | F | 56 | High | Chronic | Yes | Yes | 5 | Dasatinib | 48 | - | |
| 28 | F | 55 | High | Chronic | Yes | Yes | 5 | Dasatinib | 4.8 | - | |
| 29 | M | 43 | Low | Chronic | Yes | Yes | 4.5 | Dasatinib | 12 | - | |
| 30 | M | 44 | Intermediate | Chronic | Yes | Yes | 4.5 | Dasatinib | 84 | - | |
| 31 | F | 60 | Intermediate | Chronic | Yes | Yes | 5 | Dasatinib | 12 | - | |
| 32 | F | 43 | Low | Chronic | Yes | Yes | 5 | Dasatinib | 42 | - | |
| 33 | F | 59 | Intermediate | Chronic | Yes | Yes | 5 | Dasatinib | 12 | - | |
| 34 | M | 72 | Low | Chronic | Yes | Yes | 4 | Dasatinib | 48 | - | |
| 35 | M | 26 | Low | Chronic | Yes | Yes | 5 | Dasatinib | 80.4 | - | |
| 36 | F | 35 | Low | Chronic | Yes | Yes | 4.5 | Nilotinib | 48 | - | |
| 37 | M | 35 | Low | Chronic | Yes | Yes | 5 | Nilotinib | 63.6 | - | |
| 38 | F | 43 | Low | Chronic | Yes | Yes | No | Nilotinib | 72 | - | |
| 39 | F | 59 | High | Chronic | Yes | Yes | 4 | Nilotinib | 30 | - | |
| 40 | M | 41 | Low | Chronic | Yes | Yes | 4 | Nilotinib | 79.2 | - | |
| 41 | M | 37 | Low | Chronic | Yes | Yes | 5 | Nilotinib | 60 | - | |
| 42 | F | 67 | High | Chronic | Yes | Yes | 5 | Nilotinib | 72 | - | |
| 43 | M | 47 | Low | Chronic | Yes | Yes | 5 | Nilotinib | 66 | - | |
| 44 | M | 49 | Low | Chronic | Yes | Yes | 5 | Nilotinib | 39.6 | - | |
| 45 | M | 72 | Low | Chronic | Yes | Yes | 4 | Bosutinib | 54 | - | |
| 46 | F | 54 | Low | Chronic | Yes | Yes | 4 | Bosutinib | 9.6 | - | |
| 47 | M | 29 | Low | Chronic | Yes | Yes | 3 | Bosutinib | 39.6 | - | |
| 48 | F | 62 | Low | Chronic | Yes | Yes | 3 | Bosutinib | 26.4 | - | |
| 49 | M | 29 | Low | Chronic | Yes | Yes | No | Ponatinib | UD | - | |
| 50 | F | 47 | Low | Chornic | Yes | Yes | 4 | Asciminib | 9.6 | - | |
| 51 | M | 60 | Low | Chronic | Yes | Yes | 3 | Asciminib | 1.2 | - | |
| Treatment withdrawal without CML relapse | 52 | M | 39 | Intermediate | Chronic | Yes | Yes | 4 | Imatinib | 20.4 | 12 |
| 53 | M | 49 | Low | Chronic | Yes | Yes | 4 | Imatinib | 180 | 13.2 | |
| 54 | F | 38 | Low | Chronic | Yes | Yes | 5 | Imatinib | 171.6 | 16.8 | |
| 55 | F | 55 | Low | Chronic | Yes | Yes | 5 | Imatinib | 168 | 7.2 | |
| 56 | M | 41 | Low | Chronic | Yes | Yes | 5 | Imatinib | 178.8 | 24 | |
| 57 | F | 42 | Low | Chronic | Yes | Yes | 5 | Imatinib | 182.4 | 24 | |
| 58 | F | 61 | Low | Chronic | Yes | Yes | 4 | Imatinib | 156 | 28.8 | |
| 59 | M | 71 | Low | Chronic | Yes | Yes | 5 | Imatinib | 39.6 | 7.2 | |
| 60 | M | 34 | Low | Chronic | Yes | Yes | 5 | Imatinib | 40.8 | 8.4 | |
| 61 | M | 46 | Intermediate | Chronic | Yes | Yes | 5 | Dasatinib | 54 | 13.2 | |
| 62 | M | 26 | Low | Chronic | Yes | Yes | 5 | Dasatinib | 60 | 21.6 | |
| 63 | M | 31 | Intermediate | Chronic | Yes | Yes | 5 | Dasatinib | 85.2 | 58.8 | |
| 64 | M | U/D | UD | Chronic | Yes | Yes | 4–5 | Dasatinib | UD | 24 | |
| 65 | M | 72 | Low | Chronic | Yes | Yes | 4 | Dasatinib | 48 | 8.4 | |
| 66 | M | 61 | Low | Chronic | Yes | Yes | 4 | Nilotinib | 52.8 | 54 | |
| 67 | F | 48 | Low | Chronic | Yes | Yes | 5 | Nilotinib | 15.6 | 15.6 | |
| 68 | F | 71 | Intermediate | Chronic | Yes | Yes | 5 | Nilotinib | 51.6 | 44.4 | |
| 69 | M | 74 | Intermediate | Chronic | Yes | Yes | 5 | Nilotinib | 39.6 | 40.8 | |
| 70 | M | 53 | Low | Chronic | Yes | Yes | 4.5 | Nilotinib | 43.2 | 42 | |
| 71 | F | 33 | Low | Chronic | Yes | Yes | 5 | Nilotinib | 44.4 | 40.8 | |
| 72 | F | 83 | UD | Chronic | Yes | Yes | 4–5 | Nilotinib | 24 | 6 | |
| 73 | M | 61 | UD | Chronic | Yes | Yes | 4–5 | Nilotinib | 24 | 6 | |
| 74 | M | 40 | Low | Chronic | Yes | Yes | 4–5 | Nilotinib | 60 | 12 | |
| 75 | F | 63 | Intermediate | Chronic | Yes | Yes | 4–5 | Nilotinib | 44.4 | 26.4 | |
| 76 | M | 74 | Intermediate | Chronic | Yes | Yes | 4–5 | Nilotinib | 44.4 | 26.4 | |
| 77 | M | 41 | Low | Chronic | Yes | Yes | 5 | Nilotinib | 79.2 | 9.6 | |
| 78 | M | 47 | Low | Chronic | Yes | Yes | 5 | Nilotinib | 66 | 27.6 | |
| CML relapsed | 79 | F | 61 | Intermediate | Chronic | Yes | Yes | No | Imatinib | 40.8 | 14.4 |
| 80 | M | 35 | Low | Chronic | Yes | Yes | No | Imatinib | 39.6 | 4.8 | |
| 81 | M | 71 | Low | Chronic | Yes | Yes | No | Imatinib | 39.6 | 9.6 | |
| 82 | F | 28 | Low | Chronic | Yes | Yes | 4 | Imatinib | 60 | 8.4 | |
| 83 | M | 48 | Low | Chronic | Yes | Yes | No | Dasatinib | 43.2 | 3.6 | |
| 84 | M | 26 | Low | Chronic | Yes | Yes | No | Dasatinib | 80.4 | 3.6 | |
| 85 | F | 25 | Low | Chronic | Yes | Yes | 4 | Dasatinib | 60 | 7.2 | |
| 86 | F | 36 | Low | Chronic | Yes | Yes | 4.5 | Dasatinib | 60 | 4.8 | |
| 87 | F | 49 | Low | Chronic | Yes | Yes | 4 | Dasatinib | 180 | 2.4 | |
| 88 | M | 45 | Low | Chronic | Yes | Yes | No | Nilotinib | 228 | UD | |
| 89 | M | 22 | Low | Chronic | Yes | Yes | No | Nilotinib | 50.4 | 3.6 | |
| 90 | M | 57 | Intermediate | Chronic | Yes | Yes | No | Nilotinib | 48 | 6 | |
| 91 | F | 38 | Low | Chronic | Yes | Yes | 4.5 | Nilotinib | 72 | 3.6 | |
| 92 | F | 27 | Low | Chronic | Yes | No | No | Bosutinib | 48 | 1.2 | |
| 93 | F | 57 | Low | Chronic | No | No | No | Bosutinib | UD | 7.2 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Vigón, L.; Luna, A.; Galán, M.; Rodríguez-Mora, S.; Fuertes, D.; Mateos, E.; Piris-Villaespesa, M.; Bautista, G.; San José, E.; Rivera-Torres, J.; et al. Identification of Immunological Parameters as Predictive Biomarkers of Relapse in Patients with Chronic Myeloid Leukemia on Treatment-Free Remission. J. Clin. Med. 2021, 10, 42. https://doi.org/10.3390/jcm10010042
Vigón L, Luna A, Galán M, Rodríguez-Mora S, Fuertes D, Mateos E, Piris-Villaespesa M, Bautista G, San José E, Rivera-Torres J, et al. Identification of Immunological Parameters as Predictive Biomarkers of Relapse in Patients with Chronic Myeloid Leukemia on Treatment-Free Remission. Journal of Clinical Medicine. 2021; 10(1):42. https://doi.org/10.3390/jcm10010042
Chicago/Turabian StyleVigón, Lorena, Alejandro Luna, Miguel Galán, Sara Rodríguez-Mora, Daniel Fuertes, Elena Mateos, Miguel Piris-Villaespesa, Guiomar Bautista, Esther San José, José Rivera-Torres, and et al. 2021. "Identification of Immunological Parameters as Predictive Biomarkers of Relapse in Patients with Chronic Myeloid Leukemia on Treatment-Free Remission" Journal of Clinical Medicine 10, no. 1: 42. https://doi.org/10.3390/jcm10010042
APA StyleVigón, L., Luna, A., Galán, M., Rodríguez-Mora, S., Fuertes, D., Mateos, E., Piris-Villaespesa, M., Bautista, G., San José, E., Rivera-Torres, J., Steegmann, J. L., de Ory, F., Pérez-Olmeda, M., Alcamí, J., Planelles, V., López-Huertas, M. R., García-Gutiérrez, V., & Coiras, M. (2021). Identification of Immunological Parameters as Predictive Biomarkers of Relapse in Patients with Chronic Myeloid Leukemia on Treatment-Free Remission. Journal of Clinical Medicine, 10(1), 42. https://doi.org/10.3390/jcm10010042






