Next-Generation Bioinformatics Approaches and Resources for Coronavirus Vaccine Discovery and Development—A Perspective Review
Abstract
:1. Introduction
2. Strategies for Vaccine Development
2.1. Biology and Structure of the Pathogen
2.2. Initiation of Immune Response
2.3. Vaccine Formulation and Manufacture
2.4. Pre-Clinical and Clinical Evaluation of Vaccines
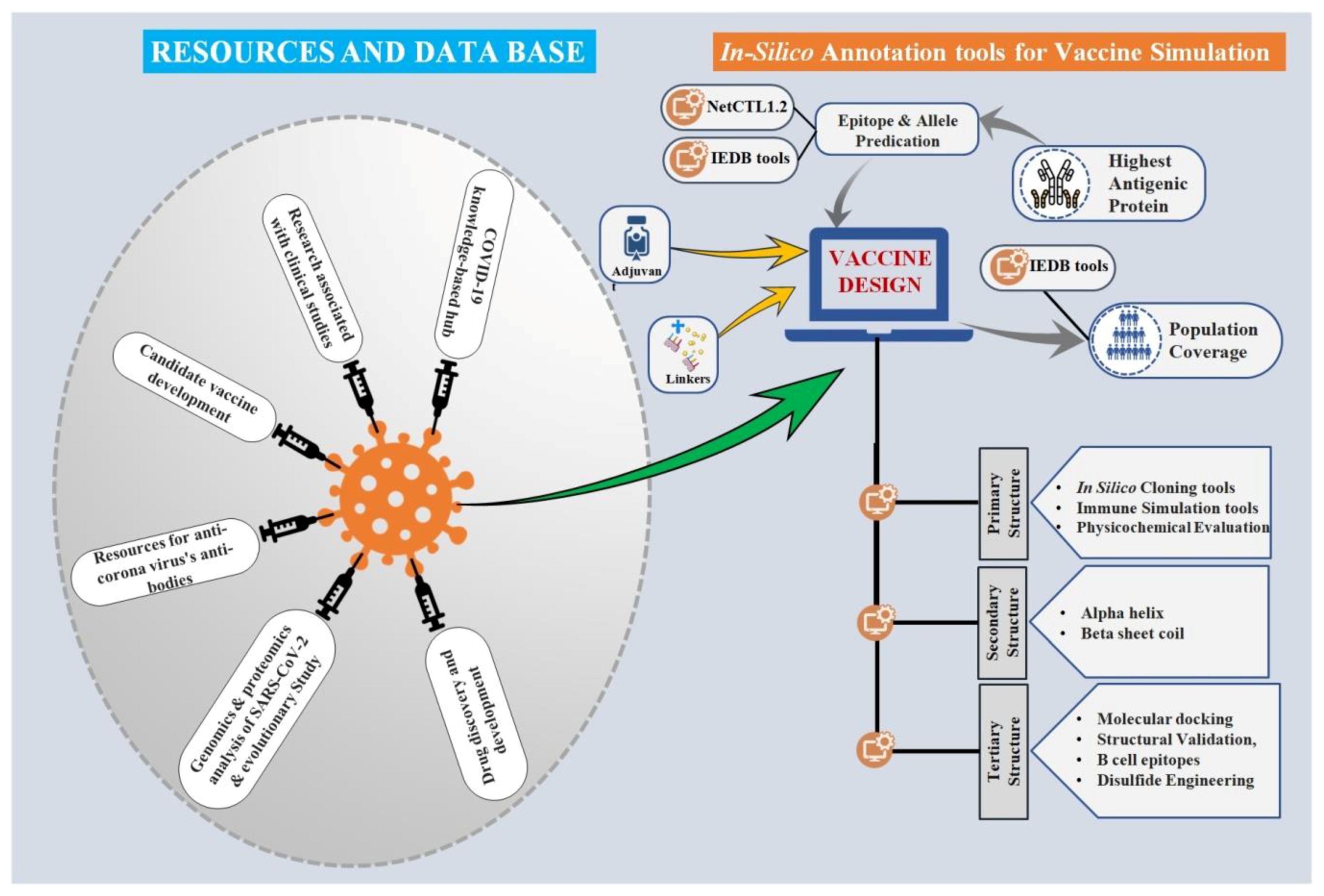
3. Computation Approaches and Online Databases: Scope for the Pre-Validations
3.1. Resources for Candidate Vaccine Development
3.1.1. DBCOVP: Resources for Cross-Genome Comparison and Structural Virulent Glycoproteins
3.1.2. CoronaVIR: Multi-Omics Resource for Literature and Internet Resources
3.1.3. CoVdb: Annotation Data Resources of Coronavirus Genes and Genomes
3.1.4. hCoronavirusesDB: Comprehensive Resources for Genetic and Proteomic Data
3.2. Resources for Genomics Proteomics and Evolutionary Analysis of SARS-CoV-2
3.2.1. SARS-CoV-2 3D: Comprehensive Resource for Proteome and Computational Modeling
3.2.2. COVIEdb: Resources for Immune Epitopes of Coronaviruses
3.2.3. GISAID: Rapid Sharing of Data from Influenza Viruses and Visualization
3.2.4. CoV3D: Structure-Based Design of Vaccines and Therapeutics against SARS-CoV-2
3.2.5. COVID-Profiler: Visualization of Multiple Genomic and Immunoinformatic Meta-Analyses
3.2.6. VirHostNet 2.0: Analysis and Visualization of Virus/Host Protein–Protein Interactions Network
3.3. Resources for Drug Discovery and Development
3.3.1. CUReD: Web-Based Resources of Currently Available Drugs against SARS-CoV-2
3.3.2. CORDITE: Database on Drug Interactions Based on Literature Aggregation with Web Interface
3.3.3. LSHTM VaC Tracker: Database on Up-To-Date Information on All the COVID-19 Vaccine Candidates
3.4. Resources Associated with Clinical Studies
3.5. Resources for Anti-Coronavirus Anti-Bodies
CoV-AbDab: Resources for the Sequence–Structural Information and Metadata
3.6. COVID-19 Knowledge-Based Hub
COVID-19 Disease Map: Assembly of Molecular Interaction Diagrams
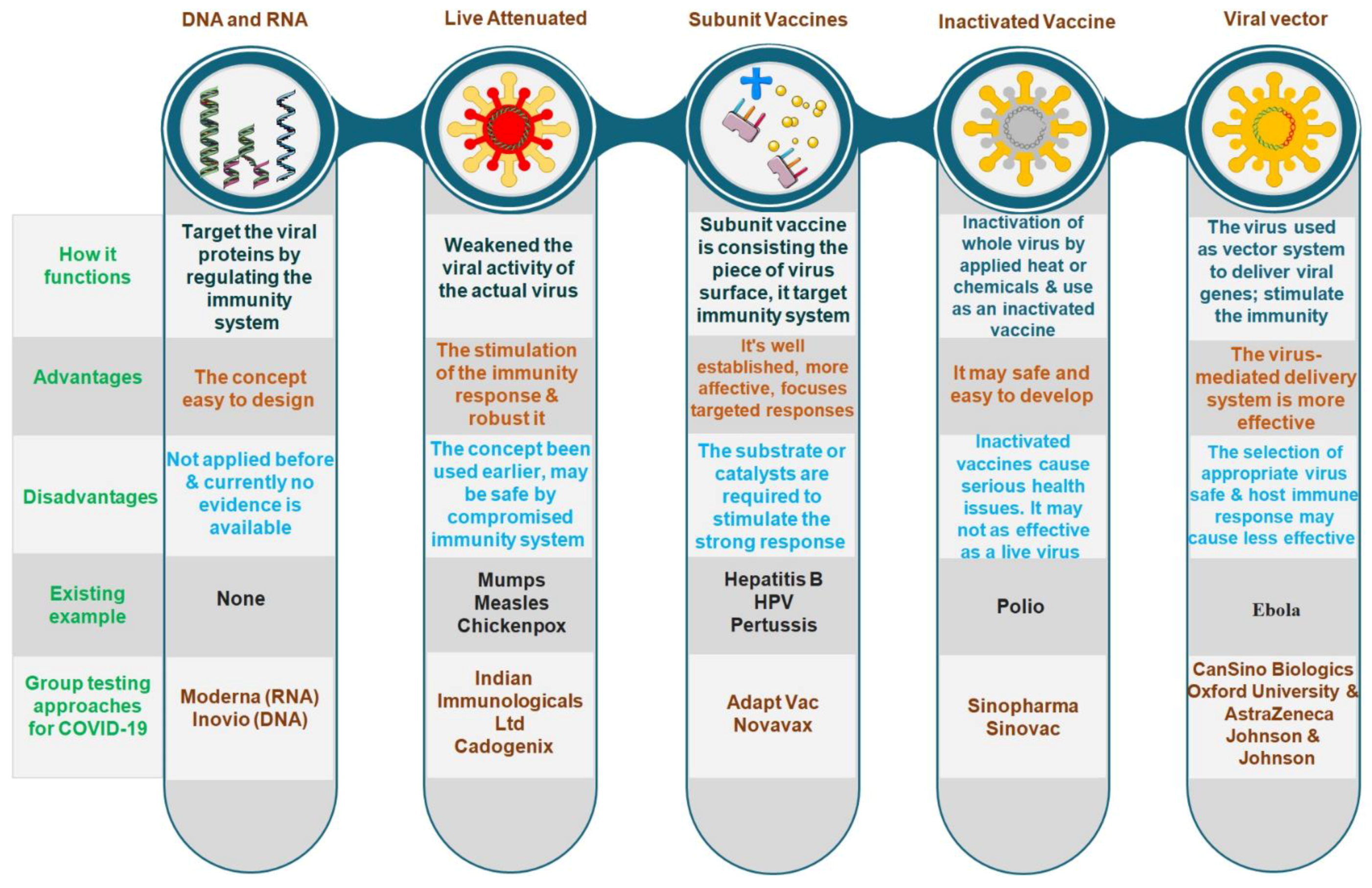
4. Vaccines for COVID-19: Efficacy and Prospective
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Ahn, D.G.; Shin, H.J.; Kim, M.H.; Lee, S.; Kim, H.S.; Myoung, J.; Kim, B.T.; Kim, S.J. Current status of epidemiology, diagnosis, therapeutics, and vaccines for novel coronavirus disease 2019 (COVID-19). J. Microbiol. Biotechnol. 2020, 30, 313–324. [Google Scholar] [CrossRef]
- Lim, Y.X.; Ng, Y.L.; Tam, J.P.; Liu, D.X. Human coronaviruses: A review of virus–host interactions. Diseases 2016, 4, 26. [Google Scholar] [CrossRef]
- Rouse, B.T.; Sehrawat, S. Immunity and immunopathology to viruses: What decides the outcome? Nat. Rev. Immunol. 2010, 10, 514–526. [Google Scholar] [CrossRef] [PubMed]
- O’Hagan, D.T.; Valiante, N.M. Recent advances in the discovery and delivery of vaccine adjuvants. Nat. Rev. Drug. Discov. 2003, 2, 727–735. [Google Scholar] [CrossRef]
- Amanat, F.; Krammer, F. SARS-CoV-2 vaccines: Status report. Immunity 2020, 52, 583–589. [Google Scholar] [CrossRef]
- Bosch, F.X.; Broker, T.R.; Forman, D.; Moscicki, A.B.; Gillison, M.L.; Doorbar, J.; Stern, P.L.; Stanley, M.; Arbyn, M.; Poljak, M.; et al. Comprehensive control of human papillomavirus infections and related diseases. Vaccine 2013, 31, H1–H31. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chu, H.; Mazmanian, S.K. Innate immune recognition of the microbiota promotes host-microbial symbiosis. Nat. Immunol. 2013, 14, 668–675. [Google Scholar] [CrossRef]
- Calina, D.; Docea, A.O.; Petrakis, D.; Egorov, A.M.; Ishmukhametov, A.A.; Gabibov, A.G.; Shtilman, M.I.; Kostoff, R.; Carvalho, F.; Vinceti, M.; et al. Towards effective COVID 19 vaccines: Updates, perspectives and challenges (Review). Int. J. Mol. Med. 2020, 46, 3–16. [Google Scholar] [CrossRef]
- Rappuoli, R. Reverse vaccinology, a genome-based approach to vaccine development. Vaccine 2001, 19, 2688–2691. [Google Scholar] [CrossRef]
- Conway, D.J. Paths to a malaria vaccine illuminated by parasite genomics. Trends Genet. 2015, 31, 97–107. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hoffman, S.L.; Vekemans, J.; Richie, T.L.; Duffy, P.E. The march toward malaria vaccines. Vaccine 2015, 33, D13–D23. [Google Scholar] [CrossRef] [Green Version]
- Berger, G. Escape of pathogens from the host immune response by mutations and mimicry. Possible means to improve vaccine performance. Med. Hypotheses 2015, 85, 664–669. [Google Scholar] [CrossRef]
- Corey, L.; Gilbert, P.B.; Tomaras, G.D.; Haynes, B.F.; Pantaleo, G.; Fauci, A.S. Immune correlates of vaccine protection against HIV-1 acquisition. Sci. Transl. Med. 2015, 7, 310rv7. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pelton, S.I. The global evolution of meningococcal epidemiology following the introduction of meningococcal vaccines. J. Adolesc. Health 2016, 59, S3–S11. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bennekov, T.; Dietrich, J.; Rosenkrands, I.; Stryhn, A.; Doherty, T.M.; Andersen, P. Alteration of epitope recognition pattern in Ag85B and ESAT-6 has a profound influence on vaccine-induced protection against Mycobacterium tuberculosis. Eur. J. Immunol. 2006, 36, 3346–3355. [Google Scholar] [CrossRef]
- Pinto, V.V.; Salanti, A.; Joergensen, L.M.; Dahlbäck, M.; Resende, M.; Ditlev, S.B.; Agger, E.M.; Arnot, D.E.; Theander, T.G.; Nielsen, M.A. The effect of adjuvants on the immune response induced by a DBL4ɛ-ID4 VAR2CSA based Plasmodium falciparum vaccine against placental malaria. Vaccine 2012, 30, 572–579. [Google Scholar] [CrossRef]
- Rappuoli, R.; Aderem, A.A. 2020 vision for vaccines against HIV, tuberculosis and malaria. Nature 2011, 473, 463–469. [Google Scholar] [CrossRef] [PubMed]
- Pollard, A.J.; Bijker, E.M. A guide to vaccinology: From basic principles to new developments. Nat. Rev. Immunol. 2021, 21, 83–100. [Google Scholar] [CrossRef] [PubMed]
- Kahn, J.P.; Henry, L.M.; Mastroianni, A.C.; Chen, W.H.; Macklin, R. Opinion: For now, it’s unethical to use human challenge studies for SARS-CoV-2 vaccine development. Proc. Natl. Acad. Sci. USA 2020, 117, 28538–28542. [Google Scholar] [CrossRef]
- Mastelic, B.; Garçon, N.; Del Giudice, G.; Golding, H.; Gruber, M.; Neels, P.; Fritzell, B. Predictive markers of safety and immunogenicity of adjuvanted vaccines. Biologicals 2013, 41, 458–468. [Google Scholar] [CrossRef]
- Wu, S.C. Progress and concept for COVID-19 vaccine development. Biotechnol. J. 2020, 15, e2000147. [Google Scholar] [CrossRef] [PubMed]
- Sahoo, S.; Mahapatra, S.R.; Parida, B.K.; Rath, S.; Dehury, B.; Raina, V.; Mohakud, N.K.; Misra, N.; Suar, M. DBCOVP: A database of coronavirus virulent glycoproteins. Comput. Biol. Med. 2021, 129, 104131. [Google Scholar] [CrossRef]
- Patiyal, S.; Kaur, D.; Kaur, H.; Sharma, N.; Dhall, A.; Sahai, S.; Agrawal, P.; Maryam, L.; Arora, C.; Raghava, G.P. A Web-Based Platform on Coronavirus Disease-19 to Maintain Predicted Diagnostic, Drug, and Vaccine Candidates. Monoclon. Antibodies Immunodiagn. Immunother. 2020, 39, 204–216. [Google Scholar] [CrossRef]
- Zhu, Z.; Meng, K.; Meng, G.A. Database resource for Genome-wide dynamics analysis of Coronaviruses on a historical and global scale. Database 2020, baaa070. [Google Scholar] [CrossRef]
- Zhu, Z.; Meng, K.; Meng, G. Genomic recombination events may reveal the evolution of coronavirus and the origin of SARS-CoV-2. Sci. Rep. 2020, 10, 21617. [Google Scholar] [CrossRef]
- Zhu, Z.; Liu, G.; Meng, K.; Yang, L.; Liu, D.; Meng, G. Rapid spread of mutant alleles in worldwide SARS-CoV-2 strains revealed by genome-wide single nucleotide polymorphism and variation analysis. Genome Biol. Evol. 2021, 13, evab015. [Google Scholar] [CrossRef]
- Alsulami, A.F.; Thomas, S.E.; Jamasb, A.R.; Beaudoin, C.A.; Moghul, I.; Bannerman, B.; Copoiu, L.; Vedithi, S.C.; Torres, P.; Blundell, T.L. SARS-CoV-2 3D database: Understanding the coronavirus proteome and evaluating possible drug targets. Brief. Bioinform. 2020. [Google Scholar] [CrossRef] [Green Version]
- Rezaei, S.; Pereira, F. In silico tracking of SARS-CoV-2 Nsp1 structural variants in helix-turn-helix motif. Authorea Prepr. 2021. [Google Scholar] [CrossRef]
- Wu, J.; Chen, W.; Zhou, J.; Zhao, W.; Sun, Y.; Zhu, H.; Yao, P.; Chen, S.; Jiang, J.; Zhou, Z. COVIEdb: A database for potential immune epitopes of coronaviruses. Front. Pharmacol. 2020, 11, 1401. [Google Scholar] [CrossRef]
- Shu, Y.; McCauley, J. GISAID: Global initiative on sharing all influenza data–from vision to reality. Eurosurveillance 2017, 22, 30494. [Google Scholar] [CrossRef] [Green Version]
- Ceraolo, C.; Giorgi, F.M. Genomic variance of the 2019-nCoV coronavirus. J. Med. Virol. 2020, 92, 522–528. [Google Scholar] [CrossRef] [Green Version]
- Shen, Z.; Xiao, Y.; Kang, L.; Ma, W.; Shi, L.; Zhang, L.; Zhou, Z.; Yang, J.; Zhong, J.; Yang, D.; et al. Genomic diversity of severe acute respiratory syndrome–coronavirus 2 in patients with coronavirus disease 2019. Clin. Infect. Dis. 2020, 71, 713–720. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Korber, B.; Fischer, W.M.; Gnanakaran, S.; Yoon, H.; Theiler, J.; Abfalterer, W.; Hengartner, N.; Giorgi, E.E.; Bhattacharya, T.; Foley, B.; et al. Tracking changes in SARS-CoV-2 Spike: Evidence that D614G increases infectivity of the COVID-19 virus. Cell 2020, 182, 812–827. [Google Scholar] [CrossRef] [PubMed]
- Gowthaman, R.; Guest, J.D.; Yin, R.; Adolf-Bryfogle, J.; Schief, W.R.; Pierce, B.G. CoV3D: A database of high-resolution coronavirus protein structures. Nucleic Acids Res. 2020, 49, D282–D287. [Google Scholar] [CrossRef]
- Ward, D.; Higgins, M.; Phelan, J.E.; Hibberd, M.L.; Campino, S.; Clark, T.G. An integrated in silico immuno-genetic analytical platform provides insights into COVID-19 serological and vaccine targets. Genome Med. 2021, 13, 1–12. [Google Scholar] [CrossRef]
- Guirimand, T.; Delmotte, S.; Navratil, V. VirHostNet 2.0: Surfing on the web of virus/host molecular interactions data. Nucleic Acids Res. 2015, 43, D583–D587. [Google Scholar] [CrossRef] [Green Version]
- Martin, R.; Löchel, H.F.; Welzel, M.; Hattab, G.; Hauschild, A.C.; Heider, D. CORDITE: The curated CORona drug In TERactions database for SARS-CoV-2. I Science 2020, 23, 101297. [Google Scholar]
- Shrotri, M.; Swinner, T.; Kampmann, B.; Parker, E.P.K. An interactive website tracking COVID-19 vaccine development. Lancet Glob. Health. 2021, 9, E590–E592. [Google Scholar] [CrossRef]
- Tzou, P.L.; Tao, K.; Nouhin, J.; Rhee, S.Y.; Hu, B.D.; Pai, S.; Parkin, N.; Shafer, R.W. Coronavirus antiviral research database (CoV-RDB): An online database designed to facilitate comparisons between candidate anti-coronavirus Compounds. Viruses 2020, 12, 1006. [Google Scholar] [CrossRef]
- Raybould, M.I.; Kovaltsuk, A.; Marks, C.; Deane, C.M. CoV-AbDab: The coronavirus antibody database. Bioinformatics 2020, 5, 734–735. [Google Scholar] [CrossRef] [PubMed]
- Iwasaki, A.; Yang, Y. The potential danger of suboptimal antibody responses in COVID-19. Nat. Rev. Immunol. 2020, 20, 339–341. [Google Scholar] [CrossRef] [Green Version]
- Hansen, J.; Baum, A.; Pascal, K.E.; Russo, V.; Giordano, S.; Wloga, E.; Fulton, B.O.; Yan, Y.; Koon, K.; Patel, K.; et al. Studies in humanized mice and convalescent humans yield a SARS-CoV-2 antibody cocktail. Science 2020, 369, 1010–1014. [Google Scholar] [CrossRef]
- Ostaszewski, M.; Mazein, A.; Gillespie, M.E.; Kuperstein, I.; Niarakis, A.; Hermjakob, H.; Pico, A.R.; Willighagen, E.L.; Evelo, C.T.; Hasenauer, J.; et al. COVID-19 Disease Map, building a computational repository of SARS-CoV-2 virus-host interaction mechanisms. Sci. Data 2020, 7, 1–4. [Google Scholar]
- Dhama, K.; Sharun, K.; Tiwari, R.; Dadar, M.; Malik, Y.S.; Singh, K.P.; Chaicumpa, W. COVID-19, an emerging coronavirus infection: Advances and prospects in designing and developing vaccines, immunotherapeutics, and therapeutics. Hum. Vaccines Immunother. 2020, 16, 1232–1238. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hwang, T.J.; Ross, J.S.; Vokinger, K.N.; Kesselheim, A.S. Association between FDA and EMA expedited approval programs and therapeutic value of new medicines: Retrospective cohort study. BMJ 2020, 371, M3434. [Google Scholar] [CrossRef] [PubMed]
- Kaur, S.P.; Gupta, V. COVID-19 Vaccine: A comprehensive status report. Virus Res. 2020, 288, 198114. [Google Scholar] [CrossRef] [PubMed]
- Rawat, K.; Kumari, P.; Saha, L. COVID-19 vaccine: A recent update in pipeline vaccines, their design and development strategies. Eur. J. Pharmacol. 2021, 892, 173751. [Google Scholar] [CrossRef] [PubMed]
- Krammer, F. SARS-CoV-2 vaccines in development. Nature 2020, 586, 516–527. [Google Scholar] [CrossRef] [PubMed]
- Jeyanathan, M.; Afkhami, S.; Smaill, F.; Miller, M.S.; Lichty, B.D.; Xing, Z. Immunological considerations for COVID-19 vaccine strategies. Nat. Rev. Immunol. 2020, 20, 615–632. [Google Scholar] [CrossRef]

| Categories | Resources | Web Link & Accessed Details | Utility |
|---|---|---|---|
| Resources for Candidate vaccine development | DBCOVP | http://covp.immt.res.in/ accessed on 21 July 2021 | This web resource provides comprehensive knowledge on the complete repertoire of structural virulent glycoproteins, viz., spike, envelope, membrane, and nucleocapsid protein from the 137 coronaviruses strain belonging to βcoronavirus genera. |
| CoronaVIR | https://webs.iiitd.edu.in/raghava/coronavir/ accessed on 21 July 2021 | This multi-omics resource includes valuable insights into the genomic, proteomic, therapeutic, and diagnostic knowledge of novel SARS-CoV-2 coronaviruses curated from the literature and existing databases. | |
| COVIEdb | http://biopharm.zju.edu.cn/coviedb/ accessed on 21 July 2021 | This database provides details on potential B/T-cell epitopes for SARS-CoV, SARS-CoV-2, and MERS-CoV to provide potential targets for pan-coronaviruses vaccine development. | |
| hCoronavirusesDB | http://hcoronaviruses.net/#/ accessed on 21 July 2021 | An integrated database and analysis resource covering the highly pathogenic human coronaviruses of SARS-CoV, MERS-CoV, and SARS-CoV-2. | |
| Resources focused on genomics proteomics and evolutionary analysis of SARS-CoV-2 | SARS-CoV-2 3D database | https://sars3d.com/ accessed on 21 July 2021 | A comprehensive database of the structures of all gene products and their higher-order assemblies, i.e., homo- and hetero-oligomers, and trans-membrane regions, as well as ligand and metal–ion interactions, with an acceptable assessment score. |
| CoVdb | http://covdb.popgenetics.net accessed on 21 July 2021 | An online genomic, proteomic, and evolutionary analysis platform that includes 5709 coronavirus strains (From 1941 to 2020) belonging to various host species from approximately 60 countries. | |
| GISAID | https://www.gisaid.org/ accessed on 21 July 2021 | This consortium mainly supports rapid sharing of all influenza virus data, including the recently detected SARS-CoV-2 coronavirus. | |
| CoV3D | https://cov3d.ibbr.umd.edu/ accessed on 21 July 2021 | This database presents comprehensive sets of 3D structures of coronavirus proteins and their interactive complexes with antibodies, receptors, and small molecules. | |
| COVID-Profiler | http://genomics.lshtm.ac.uk/ accessed on 21 July 2021 | This web server includes a plethora of tools that enable users to analyze SARS-CoV-2 sequencing and immunological data. | |
| VirHostNet 2.0 | http://virhostnet.prabi.fr accessed on 21 July 2021 | A knowledgebase resource specific to the network-based exploration of virus–host protein–protein interactions. | |
| Resources for drug discovery and development | CUReD | https://iiim.res.in/cured/ accessed on 21 July 2021 | A web server that provides information regarding the drugs, diagnostics, and devices including the current stage of the trials on COVID-19. |
| CORDITE | https://cordite.mathematik.uni-marburg.de accessed on 21 July 2021 | This database combines and represents information from various published articles as well as preprints about potential drugs, targets, and their interactions. It enables users to access, sort, and download relevant data to conduct meta-analyses, to design new clinical trials, or to conduct a curated literature search. | |
| LSHTM VaC tracker | https://vac-lshtm.shinyapps.io/ncov_vaccine_landscape/ accessed on 21 July 2021 | This database hosted by the Vaccine Centre (VaC) at the School of Hygiene and Tropical Medicine provides a user-friendly up-to-date view of the global vaccine landscape. | |
| Resources associated with clinical studies | CoV-RDB | https://covdb.stanford.edu accessed on 21 July 2021 | This database includes details of over 1800 cell cultures, 465 entry assays, 519 biochemical experiments, 259 animal model studies, and 71 clinical studies from over 400 published papers. SARS-CoV-2, SARS-CoV, and MERS-CoV account for 85% of the data. |
| Resources for anti-coronavirus anti-bodies | CoV-AbDab | http://opig.stats.ox.ac.uk/webapps/covabdab/ accessed on 21 July 2021 | This portal presents sequence–structural information and metadata on all pre-printed, published, and patented anti-coronavirus antibodies. |
| COVID-19 knowledge-based hub | COVID-19 disease map | https://covid.pages.uni.lu/ accessed on 21 July 2021 | A consortium of SARS-CoV-2 virus–host interaction mechanisms based on the input from domain experts and guided by various published works. |
| Name of the Vaccine Candidate | Name of Developers | Platform Utilized | Country of Origin |
|---|---|---|---|
| Moderna COVID-19 Vaccine (mRNA-1273) | Moderna, BARDA, NIAID | mRNA-based vaccine | USA |
| Covaxin | Bharat Biotech, ICMR | Inactivated vaccine | India |
| Comirnaty (BNT162b2) | Pfizer, BioNTech; Fosun Pharma | mRNA-based vaccine | Multinational |
| Covishield | Oxford University and AstraZeneca | Adenovirus vaccine | UK |
| Sputnik V | Gamaleya Research Institute, Acellena Contract Drug Research, and Development | Recombinant adenovirus vaccine (rAd26 and rAd5) | Russia |
| CoronaVac | Sinovac | Inactivated vaccine (formalin with alum adjuvant) | China |
| COVID-19 Vaccine Janssen (JNJ-78436735; Ad26.COV2.S) | Janssen Vaccines (Johnson & Johnson) | Non-replicating viral vector | The Netherlands, USA |
| EpiVacCorona | Federal Budgetary Research Institution State Research Center of Virology and Biotechnology | Peptide vaccine | Russia |
| BBIBP-CorV | Beijing Institute of Biological Products; China National Pharmaceutical Group (Sinopharm) | Inactivated vaccine | China |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chatterjee, R.; Ghosh, M.; Sahoo, S.; Padhi, S.; Misra, N.; Raina, V.; Suar, M.; Son, Y.-O. Next-Generation Bioinformatics Approaches and Resources for Coronavirus Vaccine Discovery and Development—A Perspective Review. Vaccines 2021, 9, 812. https://doi.org/10.3390/vaccines9080812
Chatterjee R, Ghosh M, Sahoo S, Padhi S, Misra N, Raina V, Suar M, Son Y-O. Next-Generation Bioinformatics Approaches and Resources for Coronavirus Vaccine Discovery and Development—A Perspective Review. Vaccines. 2021; 9(8):812. https://doi.org/10.3390/vaccines9080812
Chicago/Turabian StyleChatterjee, Rahul, Mrinmoy Ghosh, Susrita Sahoo, Santwana Padhi, Namrata Misra, Visakha Raina, Mrutyunjay Suar, and Young-Ok Son. 2021. "Next-Generation Bioinformatics Approaches and Resources for Coronavirus Vaccine Discovery and Development—A Perspective Review" Vaccines 9, no. 8: 812. https://doi.org/10.3390/vaccines9080812
APA StyleChatterjee, R., Ghosh, M., Sahoo, S., Padhi, S., Misra, N., Raina, V., Suar, M., & Son, Y.-O. (2021). Next-Generation Bioinformatics Approaches and Resources for Coronavirus Vaccine Discovery and Development—A Perspective Review. Vaccines, 9(8), 812. https://doi.org/10.3390/vaccines9080812







