Overview of Neutralizing Antibodies and Their Potential in COVID-19
Abstract
1. Introduction
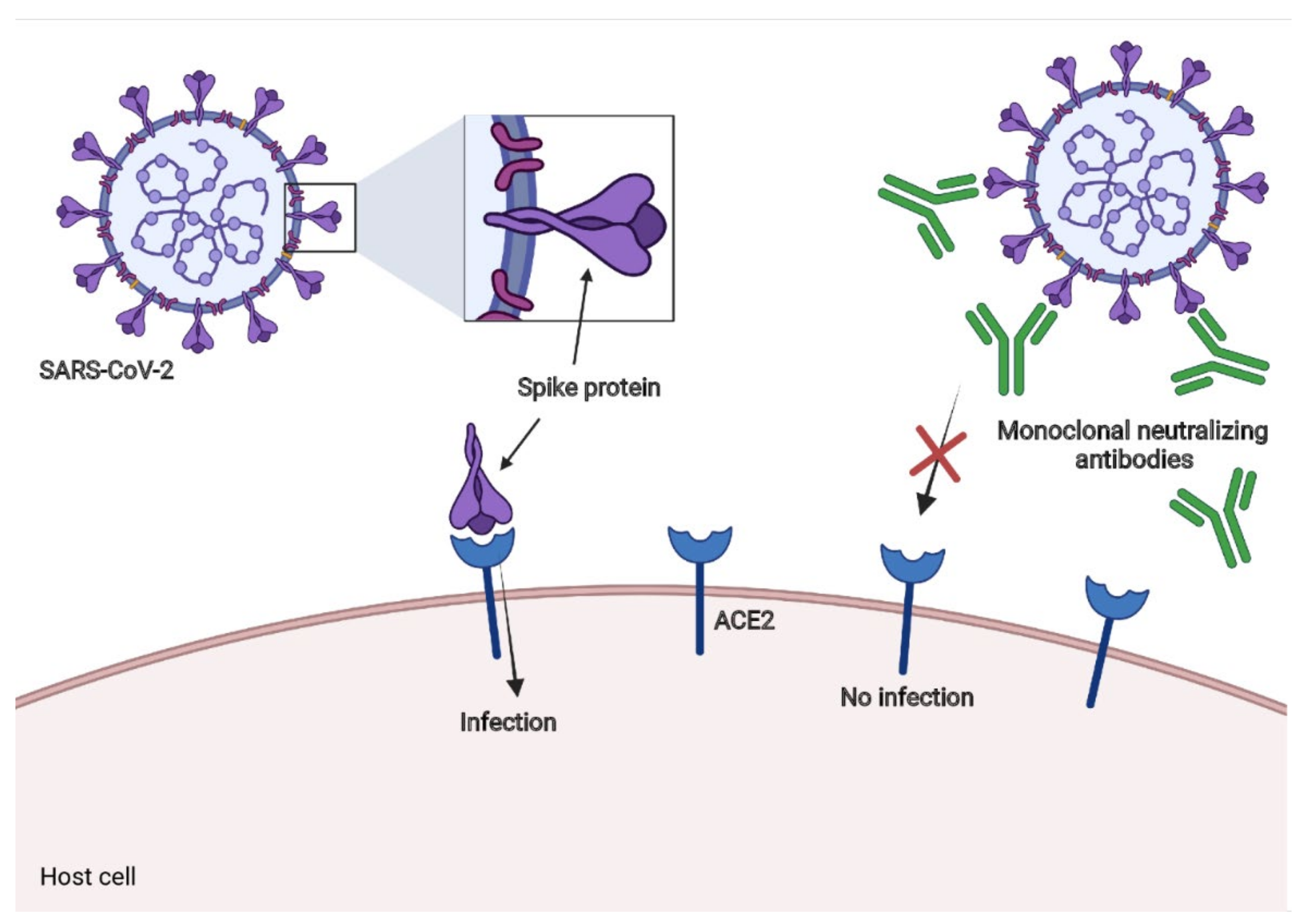
2. Definition of Neutralizing Antibody
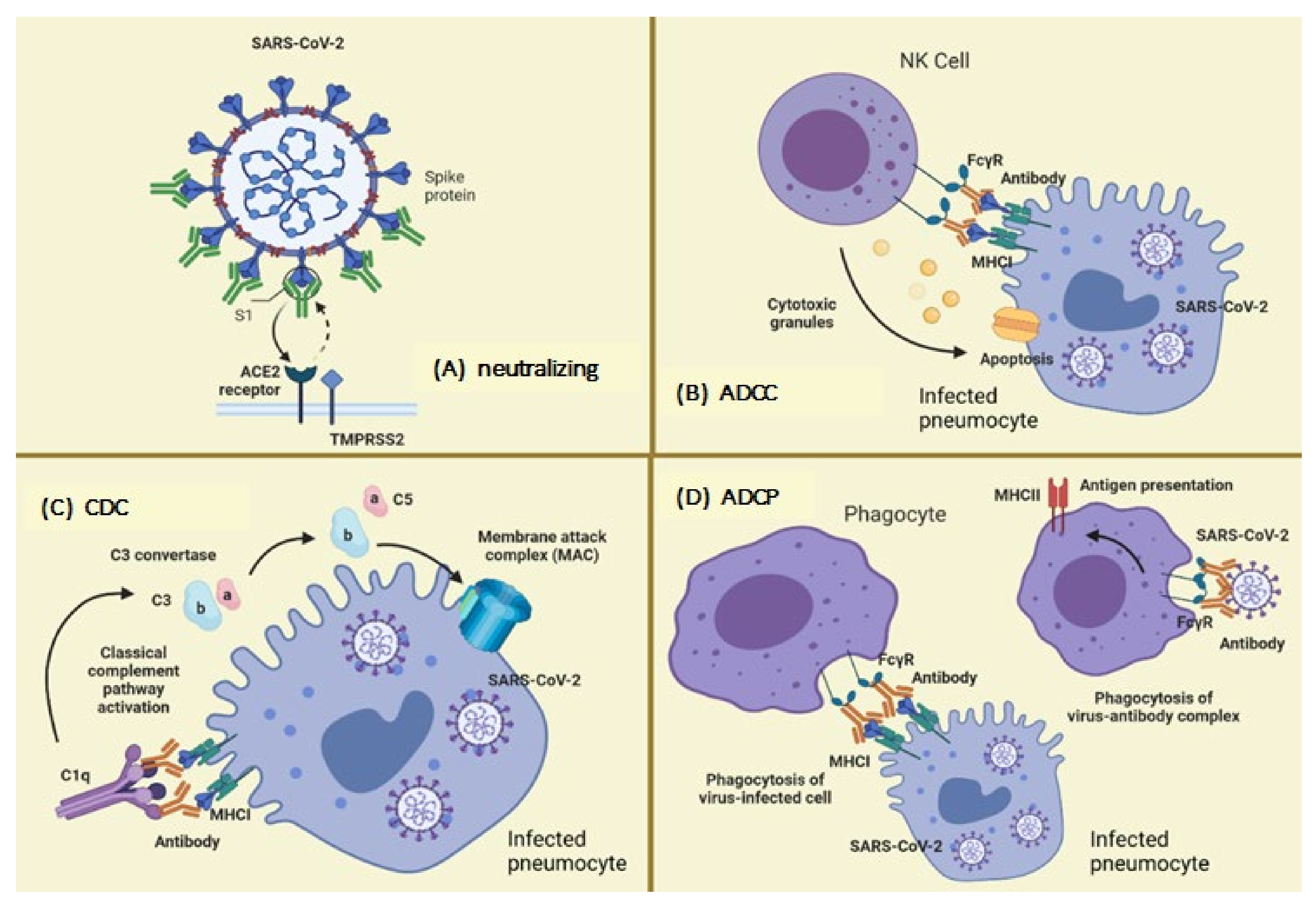
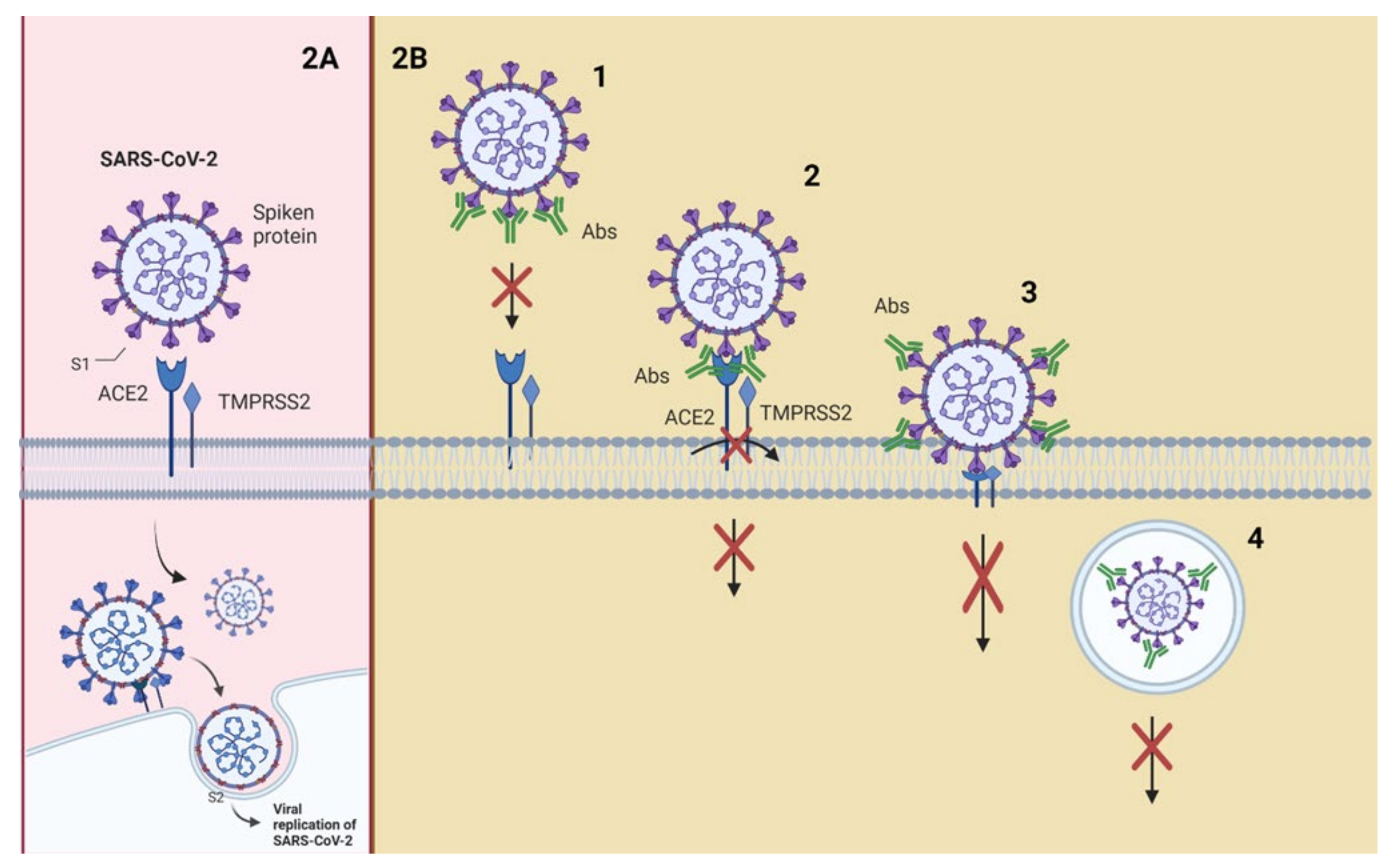
3. Mechanisms of Action of Neutralizing Antibodies
4. Generation and Characteristics of a Neutralizing Antibody
5. Neutralizing Antibodies and SARS-CoV-2
5.1. Challenges in Comparing Antibody Titers
5.2. NAbs Induced by Natural Infection and Their Protective Role
5.3. NAbs Induced by Vaccination
5.4. Factors Affecting NAb Production
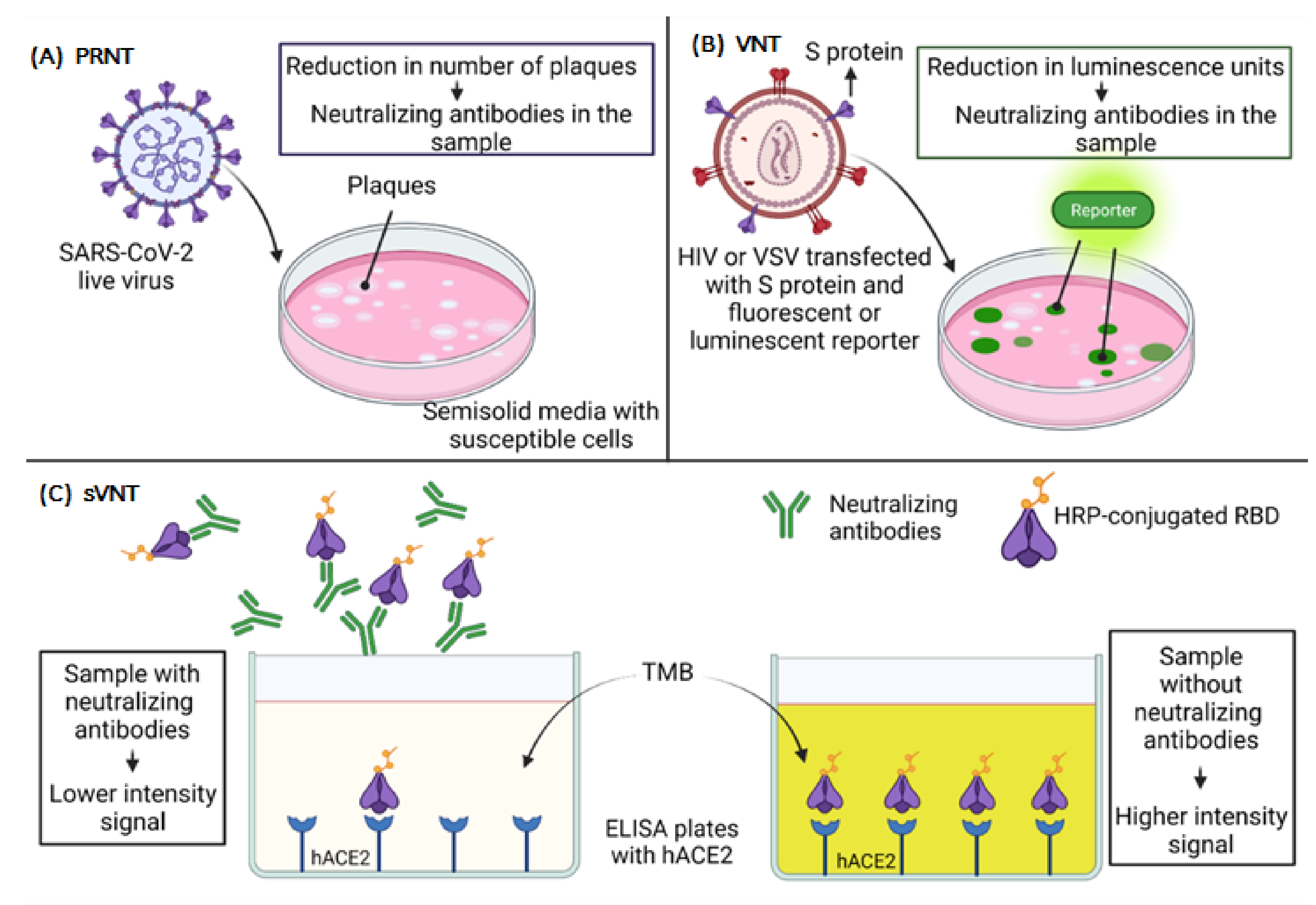
6. Neutralization Assays
7. Therapeutic Applications of Neutralizing Monoclonal Antibodies: Can We Take Advantage of This for COVID-19?
7.1. LY-CoV555
7.2. VIR-7831 and VIR-7832
7.3. BGB-DXP593
7.4. REGN-COV2
7.5. CT-P59
7.6. Nanobodies
8. Disadvantages of Neutralizing Antibodies: Antibody-Dependent Enhancement (ADE)
9. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Coronaviridae Study Group of the International Committee on Taxonomy of Viruses. The Species Severe Acute Respiratory Syndrome-Related Coronavirus: Classifying 2019-Ncov and Naming it SARS-CoV-2. Nat. Microbiol. 2020, 5, 536–544. [Google Scholar] [CrossRef] [PubMed]
- Johns Hopkins University. Coronavirus COVID-19 Dashboard by the Center for Systems Science and Engineering (Csse). Johns Hopkins University-Baltimore. Available online: https://coronavirus.jhu.edu/map.html (accessed on 8 July 2021).
- Who Coronavirus Disease (COVID-19) Dashboard. Available online: https://covid19.who.int (accessed on 26 July 2021).
- Katz, L.M. (A Little) Clarity on Convalescent Plasma for COVID-19. N. Engl. J. Med. 2021, 384, 666–668. [Google Scholar] [CrossRef]
- Li, G.; De Clercq, E. Therapeutic Options for the 2019 Novel Coronavirus (2019-Ncov). Nat. Rev. Drug Discov. 2020, 19, 149–150. [Google Scholar] [CrossRef]
- Zhou, G.; Zhao, Q. Perspectives on Therapeutic Neutralizing Antibodies against the Novel Coronavirus SARS-CoV-2. Int. J. Biol. Sci. 2020, 16, 1718–1723. [Google Scholar] [CrossRef]
- Pallikkuth, S.; De Armas, L.; Rinaldi, S.; Pahwa, S. T Follicular Helper Cells and B Cell Dysfunction in a ging and Hiv-1 Infection. Front. Immunol. 2017, 8, 1380. [Google Scholar] [CrossRef]
- Corti, D.; Lanzavecchia, A. Broadly Neutralizing Antiviral Antibodies. Annu. Rev. Immunol. 2013, 31, 705–742. [Google Scholar] [CrossRef] [PubMed]
- Jiang, S.; Du, L. Effect of Low-Pathogenic Human Coronavirus-Specific Antibodies on SARS-CoV-2. Trends Immunol. 2020, 41, 853–854. [Google Scholar] [CrossRef]
- Padilla-Quirarte, H.O.; Lopez-Guerrero, D.V.; Gutierrez-Xicotencatl, L.; Esquivel-Guadarrama, F. Protective Antibodies Against Influenza Proteins. Front. Immunol. 2019, 10, 1677. [Google Scholar] [CrossRef]
- Arvin, A.M.; Fink, K.; Schmid, M.A.; Cathcart, A.; Spreafico, R.; Havenar-Daughton, C.; Lanzavecchia, A.; Corti, D.; Virgin, H.W. A Perspective on Potential Antibody-Dependent Enhancement of SARS-CoV-2. Nature 2020, 584, 353–363. [Google Scholar] [CrossRef] [PubMed]
- Raybould, M.I.J.; Kovaltsuk, A.; Marks, C.; Deane, C.M. Cov-Abdab: The Coronavirus Antibody Database. Bioinformatics 2020, 37, 734–735. [Google Scholar] [CrossRef]
- Dörner, T.; Radbruch, A. Antibodies and B Cell Memory in Viral Immunity. Immunity 2007, 27, 384–392. [Google Scholar] [CrossRef]
- Hua, C.K.; Ackerman, M.E. Increasing the Clinical Potential and Applications of Anti-Hiv Antibodies. Front. Immunol. 2017, 8, 1655. [Google Scholar] [CrossRef]
- Murin, C.D.; Wilson, I.A.; Ward, A.B.; Biology, C.; Jolla, L.; Discovery, I.; Initiative, V.; Antibody, N.; Jolla, L.; Jolla, L. Across Three Different Enveloped Viruses. Nat. Microbiol. 2019, 4, 734–747. [Google Scholar] [CrossRef] [PubMed]
- Van Erp, E.A.; Luytjes, W.; Ferwerda, G.; Van Kasteren, P.B. Fc-Mediated Antibody Effector Functions during Respiratory Syncytial Virus Infection and Disease. Front. Immunol. 2019, 10, 548. [Google Scholar] [CrossRef]
- Lu, L.L.; Suscovich, T.J.; Fortune, S.M.; Alter, G. Beyond Binding: Antibody Effector Functions in Infectious Diseases. Nat. Rev. Immunol. 2017, 18, 46–61. [Google Scholar] [CrossRef] [PubMed]
- Overbaugh, J.; Morris, L. The Antibody Response against Hiv-1. Cold Spring Harb. Perspect. Med. 2011, 2, A007039. [Google Scholar] [CrossRef]
- Neurath, A.R. Immune Response to Viruses: Antibody-Mediated Immunity. In Encyclopedia Of Virology, 3rd ed.; Mahy, B.W.J., Van Regenmortel, M.H.V., Eds.; Academic Press: Oxford, UK, 2008; pp. 56–70. [Google Scholar] [CrossRef]
- Klasse, P.J. Neutralization of Virus Infectivity by Antibodies: Old Problems in New Perspectives. Adv. Biol. 2014, 2014, 1–24. [Google Scholar] [CrossRef]
- Klasse, P.J.; Sattentau, Q.J. Occupancy and Mechanism in Antibody-Mediated Neutralization of Animal Viruses. J. Gen. Virol. 2002, 83, 2091–2108. [Google Scholar] [CrossRef]
- Bachmann, M.F.; Mohsen, M.O.; Zha, L.; Vogel, M.; Speiser, D.E. SARS-CoV-2 Structural Features May Explain Limited Neutralizing-Antibody Responses. NPJ Vaccines 2021, 6, 2. [Google Scholar] [CrossRef] [PubMed]
- Sedova, E.; Scherbinin, D.N.; Lysenko, A.; Alekseeva, S.V.; Artemova, E.A.; Shmarov, M. Non-Neutralizing Antibodies Directed at Conservative Influenza Antigens. Acta Nat. 2019, 11, 22–32. [Google Scholar] [CrossRef]
- Vangelista, L.; Secchi, M. Prepare for the Future: Dissecting the Spike to Seek Broadly Neutralizing Antibodies and Universal Vaccine for Pandemic Coronaviruses. Front. Mol. Biosci. 2020, 7, 226. [Google Scholar] [CrossRef] [PubMed]
- Subbaraman, H.; Schanz, M.; Trkola, A. Broadly Neutralizing Antibodies: What is Needed to Move from a Rare Event in Hiv-1 Infection to Vaccine Efficacy? Retrovirology 2018, 15, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Burton, D.R.; Hangartner, L. Broadly Neutralizing Antibodies to Hiv and Their Role in Vaccine Design. Annu. Rev. Immunol. 2016, 34, 635–659. [Google Scholar] [CrossRef]
- Ronen, K.; Dingens, A.S.; Graham, S.M.; Jaoko, W.; Mandaliya, K.; Mcclelland, R.S.; Overbaugh, J. Comprehensive Characterization of Humoral Correlates of Human Immunodeficiency Virus 1 Superinfection Acquisition in High-Risk Kenyan Women. Ebiomedicine 2017, 18, 216–224. [Google Scholar] [CrossRef][Green Version]
- Gray, E.; Madiga, M.C.; Hermanus, T.; Moore, P.; Wibmer, C.K.; Tumba, N.L.; Werner, L.; Mlisana, K.; Sibeko, S.; Williamson, C.; et al. The Neutralization Breadth of Hiv-1 Develops Incrementally over Four Years and is Associated with Cd4+ T Cell Decline and High Viral Load During Acute Infection. J. Virol. 2011, 85, 4828–4840. [Google Scholar] [CrossRef] [PubMed]
- Derdeyn, C.A.; Moore, P.L.; Morris, L. Development of Broadly Neutralizing Antibodies from Autologous Neutralizing Antibody Responses in Hiv Infection. Curr. Opin. HIV Aids 2014, 9, 210–216. [Google Scholar] [CrossRef]
- Gupta, A.; Madhavan, M.V.; Sehgal, K.; Nair, N.; Mahajan, S.; Sehrawat, T.S.; Bikdeli, B.; Ahluwalia, N.; Ausiello, J.C.; Wan, E.Y.; et al. Extrapulmonary Manifestations of COVID-19. Nat. Med. 2020, 26, 1017–1032. [Google Scholar] [CrossRef]
- Yuki, K.; Fujiogi, M.; Koutsogiannaki, S. COVID-19 Pathophysiology: A Review. Clin. Immunol. 2020, 215, 108427. [Google Scholar] [CrossRef]
- Zhao, M.-M.; Yang, W.-L.; Yang, F.-Y.; Zhang, L.; Huang, W.-J.; Hou, W.; Fan, C.-F.; Jin, R.-H.; Feng, Y.-M.; Wang, Y.-C.; et al. Cathepsin L Plays a Key Role in SARS-CoV-2 Infection in Humans and Humanized Mice and is a Promising Target for New Drug Development. Signal Transduct. Target. Ther. 2021, 6, 1–12. [Google Scholar] [CrossRef]
- Duan, L.; Zheng, Q.; Zhang, H.; Niu, Y.; Lou, Y.; Wang, H. The SARS-CoV-2 Spike Glycoprotein Biosynthesis, Structure, Function, and Antigenicity: Implications for the Design of Spike-Based Vaccine Immunogens. Front. Immunol. 2020, 11, 576622. [Google Scholar] [CrossRef]
- Ali, A.; Vijayan, R. Dynamics of the Ace2–SARS-CoV-2/SARS-CoV Spike Protein Interface Reveal Unique Mechanisms. Sci. Rep. 2020, 10, 1–12. [Google Scholar] [CrossRef]
- Ju, B.; Zhang, Q.; Ge, J.; Wang, R.; Sun, J.; Ge, X.; Yu, J.; Shan, S.; Zhou, B.; Song, S.; et al. Human Neutralizing Antibodies Elicited By SARS-CoV-2 Infection. Nature 2020, 584, 115–119. [Google Scholar] [CrossRef] [PubMed]
- Cheedarla, N.; Hanna, L.E. Chapter 7-Functional and Protective Role of Neutralizing Antibodies (NAbs) Against Viral Infections. In Recent Developments in Applied Microbiology and Biochemistry; Buddolla, V., Ed.; Academic Press: Cambridge, MA, USA, 2019; pp. 83–93. [Google Scholar]
- Ali, M.G.; Zhang, Z.; Gao, Q.; Pan, M.; Rowan, E.G.; Zhang, J. Recent Advances in Therapeutic Applications of Neutralizing Antibodies for Virus Infections: An Overview. Immunol. Res. 2020, 68, 325–339. [Google Scholar] [CrossRef]
- Dimitrov, D.S. Virus Entry: Molecular Mechanisms and Biomedical Applications. Nat. Rev. Genet. 2004, 2, 109–122. [Google Scholar] [CrossRef]
- Thorley, J.A.; Mckeating, J.A.; Rappoport, J.Z. Mechanisms of Viral Entry: Sneaking in the Front Door. Protoplasma 2010, 244, 15–24. [Google Scholar] [CrossRef]
- Baum, L.L.; Mathieson, B.J.; Connick, E. Immunity to Hiv. Encycl. Immunobiol. 2016, 4, 342–354. [Google Scholar] [CrossRef]
- Zheng, Q.; Jiang, J.; He, M.; Zheng, Z.; Yu, H.; Li, T.; Xue, W.; Tang, Z.; Ying, D.; Li, Z.; et al. Viral Neutralization by Antibody-Imposed Physical Disruption. Proc. Natl. Acad. Sci. USA 2019, 116, 26933–26940. [Google Scholar] [CrossRef]
- Klasse, P.J.; Bron, R.; Marsh, M. Mechanisms of Enveloped Virus Entry into Animal Cells. Adv. Drug Deliv. Rev. 1998, 34, 65–91. [Google Scholar] [CrossRef]
- Reading, S.A.; Dimmock, N.J. Neutralization of Animal Virus Infectivity by Antibody. Arch. Virol. 2007, 152, 1047–1059. [Google Scholar] [CrossRef] [PubMed]
- Nelson, C.D.; Palermo, L.M.; Hafenstein, S.L.; Parrish, C.R. Different Mechanisms of Antibody-Mediated Neutralization of Parvoviruses Revealed using the Fab Fragments of Monoclonal Antibodies. Virology 2007, 361, 283–293. [Google Scholar] [CrossRef] [PubMed]
- Xiao, Y.; Dong, X.; Chen, Y.-H. Neutralizing Antibodies Mechanism of Neutralization and Protective Activity against Hiv-1. Immunol. Res. 2002, 25, 193–200. [Google Scholar] [CrossRef]
- Marsh, M.; Helenius, A. Virus Entry: Open Sesame. Cell 2006, 124, 729–740. [Google Scholar] [CrossRef]
- Florese, R.H.; Robert-Guroff, M. Non-Neutralizing Antibodies and Vaccine-Induced Protection Against Hiv. Futur. HIV Ther. 2007, 1, 259–271. [Google Scholar] [CrossRef]
- Mayr, L.M.; Su, B.; Moog, C. Non-Neutralizing Antibodies Directed against Hiv and their Functions. Front. Immunol. 2017, 8, 1590. [Google Scholar] [CrossRef] [PubMed]
- Tas, J.M.J.; Mesin, L.; Pasqual, G.; Targ, S.; Jacobsen, J.T.; Mano, Y.M.; Chen, C.S.; Weill, J.-C.; Reynaud, C.-A.; Browne, E.P.; et al. Visualizing Antibody Affinity Maturation in Germinal Centers. Science 2016, 351, 1048–1054. [Google Scholar] [CrossRef]
- Pilzecker, B.; Jacobs, H. Mutating for Good: DNA Damage Responses During Somatic Hypermutation. Front. Immunol. 2019, 10, 438. [Google Scholar] [CrossRef] [PubMed]
- Teng, G.; Papavasiliou, F.N. Immunoglobulin Somatic Hypermutation. Annu. Rev. Genet. 2007, 41, 107–120. [Google Scholar] [CrossRef]
- Stebegg, M.; Kumar, S.; Silva-Cayetano, A.; Fonseca, V.R.; Linterman, M.A.; Graca, L. Regulation of the Germinal Center Response. Front. Immunol. 2018, 9, 2469. [Google Scholar] [CrossRef]
- Oropallo, M.A.; Cerutti, A. Germinal Center Reaction: Antigen Affinity and Presentation Explain it All. Trends Immunol. 2014, 35, 287–289. [Google Scholar] [CrossRef]
- Nouri, N.; Kleinstein, S.H. Somatic Hypermutation Analysis for Improved Identification of B Cell Clonal Families from Next-Generation Sequencing Data. PLoS Comput. Biol. 2020, 16, E1007977. [Google Scholar] [CrossRef]
- Mesin, L.; Ersching, J.; Victora, G.D. Germinal Center B Cell Dynamics. Immunity 2016, 45, 471–482. [Google Scholar] [CrossRef]
- Stavnezer, J.; Guikema, J.E.; Schrader, C.E. Mechanism and Regulation of Class Switch Recombination. Annu. Rev. Immunol. 2008, 26, 261–292. [Google Scholar] [CrossRef]
- Liechti, T. Cd40 Signaling—Germinal Center B Cells Go at their Own Pace. Cytom. Part A 2018, 95, 419–421. [Google Scholar] [CrossRef]
- Zhang, M.; Srivastava, G.; Lu, L. The Pre-B Cell Receptor and its Function During B Cell Development. Cell. Mol. Immunol. 2004, 1, 89–94. [Google Scholar] [PubMed]
- Papa, I.; Vinuesa, C.G. Synaptic Interactions in Germinal Centers. Front. Immunol. 2018, 9, 1858. [Google Scholar] [CrossRef] [PubMed]
- Chen, Z.; Wang, J.H. Signaling Control of Antibody Isotype Switching. Adv. Immunol. 2019, 141, 105–164. [Google Scholar] [CrossRef] [PubMed]
- Jhamnani, R.D.; Nunes-Santos, C.J.; Bergerson, J.; Rosenzweig, S.D. Class-Switch Recombination (Csr)/Hyper-Igm (Higm) Syndromes and Phosphoinositide 3-Kinase (Pi3k) Defects. Front. Immunol. 2018, 9, 2172. [Google Scholar] [CrossRef] [PubMed]
- Radtke, D.; Bannard, O. Expression of the Plasma Cell Transcriptional Regulator Blimp-1 By Dark Zone Germinal Center B Cells During Periods of Proliferation. Front. Immunol. 2019, 9, 3106. [Google Scholar] [CrossRef]
- Recaldin, T.; Fear, D.J. Transcription Factors Regulating B Cell Fate in the Germinal Centre. Clin. Exp. Immunol. 2015, 183, 65–75. [Google Scholar] [CrossRef] [PubMed]
- Huang, C.; Geng, H.; Boss, I.; Wang, L.; Melnick, A. Cooperative Transcriptional Repression by Bcl6 And Bach2 in Germinal Center B-Cell Differentiation. Blood 2014, 123, 1012–1020. [Google Scholar] [CrossRef] [PubMed]
- Palm, A.-K.E.; Henry, C. Remembrance of Things Past: Long-Term B Cell Memory after Infection and Vaccination. Front. Immunol. 2019, 10, 1787. [Google Scholar] [CrossRef]
- Brynjolfsson, S.; Berg, L.P.; Ekerhult, T.O.; Rimkute, I.; Wick, M.-J.; Mårtensson, I.-L.; Grimsholm, O. Long-Lived Plasma Cells In Mice and Men. Front. Immunol. 2018, 9, 2673. [Google Scholar] [CrossRef]
- Lightman, S.M.; Utley, A.; Lee, K.P. Survival of Long-Lived Plasma Cells (Llpc): Piecing Together the Puzzle. Front. Immunol. 2019, 10, 965. [Google Scholar] [CrossRef]
- Slifka, M.K.; Antia, R.; Whitmire, J.; Ahmed, R. Humoral Immunity Due to Long-Lived Plasma Cells. Immunity 1998, 8, 363–372. [Google Scholar] [CrossRef]
- Akkaya, M.; Kwak, K.; Pierce, S.K. B Cell Memory: Building Two Walls of Protection against Pathogens. Nat. Rev. Immunol. 2019, 20, 229–238. [Google Scholar] [CrossRef] [PubMed]
- Weisel, F.; Shlomchik, M. Memory B Cells of Mice and Humans. Annu. Rev. Immunol. 2017, 35, 255–284. [Google Scholar] [CrossRef] [PubMed]
- Wong, R.; Bhattacharya, D. Basics of Memory B-Cell Responses: Lessons From and for the Real World. Immunology 2018, 156, 120–129. [Google Scholar] [CrossRef] [PubMed]
- Purtha, W.E.; Tedder, T.F.; Johnson, S.; Bhattacharya, D.; Diamond, M.S. Memory B Cells, but Not Long-Lived Plasma Cells, Possess Antigen Specificities For Viral Escape Mutants. J. Exp. Med. 2011, 208, 2599–2606. [Google Scholar] [CrossRef]
- Wang, Z.; Muecksch, F.; Schaefer-Babajew, D.; Finkin, S.; Viant, C.; Gaebler, C.; Hoffmann, H.-H.; Barnes, C.O.; Cipolla, M.; Ramos, V.; et al. Naturally Enhanced Neutralizing Breadth against SARS-CoV-2 One Year after Infection. Nature 2021, 595, 426–431. [Google Scholar] [CrossRef] [PubMed]
- Vanblargan, L.A.; Goo, L.; Pierson, T.C. Deconstructing the Antiviral Neutralizing-Antibody Response: Implications for Vaccine Development and Immunity. Microbiol. Mol. Biol. Rev. 2016, 80, 989–1010. [Google Scholar] [CrossRef]
- Doria-Rose, N.A.; Joyce, M.G. Strategies to Guide the Antibody Affinity Maturation Process. Curr. Opin. Virol. 2015, 11, 137–147. [Google Scholar] [CrossRef]
- West, A.P.; Diskin, R.; Nussenzweig, M.C.; Bjorkman, P.J. Structural Basis For Germ-Line Gene Usage of a Potent Class of Antibodies Targeting the Cd4-Binding Site of Hiv-1 Gp120. Proc. Natl. Acad. Sci. USA 2012, 109, E2083–E2090. [Google Scholar] [CrossRef]
- Doria-Rose, N.A.; Sequencing, N.C.; Schramm, C.A.; Gorman, J.; Moore, P.L.; Bhiman, J.N.; Dekosky, B.J.; Ernandes, M.J.; Georgiev, I.S.; Kim, H.J.; et al. Developmental Pathway for Potent V1v2-Directed Hiv-Neutralizing Antibodies. Nature 2014, 509, 55–62. [Google Scholar] [CrossRef] [PubMed]
- Wrammert, J.; Smith, K.; Miller, J.; Langley, W.A.; Kokko, K.; Larsen, C.; Zheng, N.-Y.; Mays, I.; Garman, L.; Helms, C.; et al. Rapid Cloning of High-Affinity Human Monoclonal Antibodies Against Influenza Virus. Nature 2008, 453, 667–671. [Google Scholar] [CrossRef] [PubMed]
- Mclellan, J.S.; Pancera, M.; Carrico, C.; Gorman, J.; Julien, J.-P.; Khayat, R.; Louder, R.; Pejchal, R.; Sastry, M.; Dai, K.; et al. Structure of Hiv-1 Gp120 V1/V2 Domain with Broadly Neutralizing Antibody Pg9. Nature 2011, 480, 336–343. [Google Scholar] [CrossRef]
- Graham, C.; Seow, J.; Huettner, I.; Khan, H.; Kouphou, N.; Acors, S.; Winstone, H.; Pickering, S.; Galao, R.P.; Dupont, L.; et al. Neutralization Potency of Monoclonal Antibodies Recognizing Dominant and Subdominant Epitopes on SARS-CoV-2 Spike is Impacted by the B.1.1.7 Variant. Immunity 2021, 54, 1276–1289.e6. [Google Scholar] [CrossRef]
- Astronomo, R.D.; Santra, S.; Ballweber-Fleming, L.; Westerberg, K.G.; Mach, L.; Hensley-Mcbain, T.; Sutherland, L.; Mildenberg, B.; Morton, G.; Yates, N.L.; et al. Neutralization Takes Precedence over Igg or Iga Isotype-Related Functions in Mucosal Hiv-1 Antibody-Mediated Protection. Ebiomedicine 2016, 14, 97–111. [Google Scholar] [CrossRef] [PubMed]
- Vukovic, N.; Van Elsas, A.; Verbeek, J.S.; Zaiss, D.M. Isotype Selection for Antibody-Based Cancer Therapy. Clin. Exp. Immunol. 2020, 203, 351–365. [Google Scholar] [CrossRef]
- Davies, A.M.; Sutton, B.J. Human Igg4: A Structural Perspective. Immunol. Rev. 2015, 268, 139–159. [Google Scholar] [CrossRef]
- Demonbreun, A.R.; Sancilio, A.; Velez, M.E.; Ryan, D.T.; Saber, R.; Vaught, L.A.; McDade, T.W. Comparison of Igg And Neutralizing Antibody Responses after one or two doses of COVID-19 Mrna Vaccine in Previously Infected and Uninfected Persons. medRxiv 2021. [Google Scholar] [CrossRef]
- Kallolimath, S.; Sun, L.; Palt, R.; Stiasny, K.; Mayrhofer, P.; Gruber, C.; Kogelmann, B.; Chen, Q.; Steinkellner, H. Highly Active Engineered Igg3 Antibodies against SARS-CoV-2. Proc. Natl. Acad. Sci. USA 2021, 42, e2107249118. [Google Scholar] [CrossRef]
- Corthésy, B. Multi-Faceted Functions of Secretory Iga at Mucosal Surfaces. Front. Immunol. 2013, 4, 185. [Google Scholar] [CrossRef] [PubMed]
- Woof, J.M.; Russell, M. Structure and Function Relationships in Iga. Mucosal Immunol. 2011, 4, 590–597. [Google Scholar] [CrossRef]
- Sterlin, D.; Mathian, A.; Miyara, M.; Mohr, A.; Anna, F.; Claër, L.; Quentric, P.; Fadlallah, J.; Devilliers, H.; Ghillani, P.; et al. Iga Dominates the Early Neutralizing Antibody Response to SARS-CoV-2. Sci. Transl. Med. 2021, 13. [Google Scholar] [CrossRef] [PubMed]
- Sharp, T.H.; Boyle, A.L.; Diebolder, C.A.; Kros, A.; Koster, A.J.; Gros, P. Insights into Igm-Mediated Complement Activation Based on in situ Structures of Igm-C1-C4b. Proc. Natl. Acad. Sci. USA 2019, 116, 11900–11905. [Google Scholar] [CrossRef] [PubMed]
- Daha, N.A.; Banda, N.K.; Roos, A.; Beurskens, F.J.; Bakker, J.M.; Daha, M.R.; Trouw, L.A. Complement Activation by (Auto-) Antibodies. Mol. Immunol. 2011, 48, 1656–1665. [Google Scholar] [CrossRef]
- Liu, J.; Wang, Y.; Xiong, E.; Hong, R.; Lu, Q.; Ohno, H.; Wang, J.-Y. Role of the Igm Fc Receptor in Immunity and Tolerance. Front. Immunol. 2019, 10, 529. [Google Scholar] [CrossRef]
- Jiang, S.; Hillyer, C.; Du, L. Neutralizing Antibodies against SARS-CoV-2 and Other Human Coronaviruses. Trends Immunol. 2020, 41, 355–359. [Google Scholar] [CrossRef]
- Putcharoen, O.; Wacharapluesadee, S.; Ni Chia, W.; Paitoonpong, L.; Tan, C.W.; Suwanpimolkul, G.; Jantarabenjakul, W.; Ruchisrisarod, C.; Wanthong, P.; Sophonphan, J.; et al. Early Detection of Neutralizing Antibodies against SARS-CoV-2 in COVID-19 Patients in Thailand. PLoS ONE 2021, 16, E0246864. [Google Scholar] [CrossRef]
- Krammer, F. A Correlate of Protection for SARS-CoV-2 Vaccines is Urgently Needed. Nat. Med. 2021, 27, 1147–1148. [Google Scholar] [CrossRef]
- Kristiansen, P.A.; Page, M.; Bernasconi, V.; Mattiuzzo, G.; Dull, P.; Makar, K.; Plotkin, S.; Knezevic, I. Who International Standard for Anti-SARS-CoV-2 Immunoglobulin. Lancet 2021, 397, 1347–1348. [Google Scholar] [CrossRef]
- Lau, E.H.Y.; Tsang, O.T.Y.; Hui, D.S.C.; Kwan, M.Y.W.; Chan, W.-H.; Chiu, S.S.; Ko, R.L.W.; Chan, K.H.; Cheng, S.M.S.; Perera, R.A.P.M.; et al. Neutralizing Antibody Titres in SARS-CoV-2 Infections. Nat. Commun. 2021, 12, 63. [Google Scholar] [CrossRef] [PubMed]
- Altawalah, H. Antibody Responses to Natural SARS-CoV-2 Infection or after COVID-19 Vaccination. Vaccines 2021, 9, 910. [Google Scholar] [CrossRef] [PubMed]
- Dan, J.M.; Mateus, J.; Kato, Y.; Hastie, K.M.; Yu, E.D.; Faliti, C.E.; Grifoni, A.; Ramirez, S.I.; Haupt, S.; Frazier, A.; et al. Immunological Memory to SARS-CoV-2 Assessed for Up To 8 Months after Infection. Science 2021, 371, Eabf4063. [Google Scholar] [CrossRef]
- Marot, S.; Malet, I.; Leducq, V.; Zafilaza, K.; Sterlin, D.; Planas, D.; Gothland, A.; Jary, A.; Dorgham, K.; Bruel, T.; et al. Rapid Decline of Neutralizing Antibodies against SARS-CoV-2 among Infected Healthcare Workers. Nat. Commun. 2021, 12, 844. [Google Scholar] [CrossRef] [PubMed]
- Seow, J.; Graham, C.; Merrick, B.; Acors, S.; Pickering, S.; Steel, K.J.A.; Hemmings, O.; O’byrne, A.; Kouphou, N.; Galao, R.P.; et al. Longitudinal Observation and Decline of Neutralizing Antibody Responses in the Three Months Following SARS-CoV-2 Infection in Humans. Nat. Microbiol. 2020, 5, 1598–1607. [Google Scholar] [CrossRef] [PubMed]
- Garcia-Beltran, W.F.; Lam, E.C.; Astudillo, M.G.; Yang, D.; Miller, T.E.; Feldman, J.; Hauser, B.M.; Caradonna, T.M.; Clayton, K.L.; Nitido, A.D.; et al. COVID-19-Neutralizing Antibodies Predict Disease Severity and Survival. Cell 2020, 184, 476–488.e11. [Google Scholar] [CrossRef]
- Klein, S.; Cortese, M.; Winter, S.L.; Wachsmuth-Melm, M.; Neufeldt, C.J.; Cerikan, B.; Stanifer, M.L.; Boulant, S.; Bartenschlager, R.; Chlanda, P. SARS-CoV-2 Structure and Replication Characterized by in situ Cryo-Electron Tomography. Nat. Commun. 2020, 11, 5885. [Google Scholar] [CrossRef] [PubMed]
- Liu, L.; To, K.K.-W.; Chan, K.-H.; Wong, Y.-C.; Zhou, R.; Kwan, K.-Y.; Fong, C.H.-Y.; Chen, L.-L.; Choi, C.Y.-K.; Lu, L.; et al. High Neutralizing Antibody Titer in Intensive Care Unit Patients with COVID-19. Emerg. Microbes Infect. 2020, 9, 1664–1670. [Google Scholar] [CrossRef]
- Schreiber, G. The Role of Type I Interferons in the Pathogenesis and Treatment of COVID-19. Front. Immunol. 2020, 11, 595739. [Google Scholar] [CrossRef]
- Röltgen, K.; Powell, A.E.; Wirz, O.F.; Stevens, B.A.; Hogan, C.A.; Najeeb, J.; Hunter, M.; Wang, H.; Sahoo, M.K.; Huang, C.; et al. Defining the Features and Duration of Antibody Responses to SARS-CoV-2 Infection Associated with Disease Severity and Outcome. Sci. Immunol. 2020, 5, eabe0240. [Google Scholar] [CrossRef]
- Dugan, H.L.; Stamper, C.T.; Li, L.; Changrob, S.; Asby, N.W.; Halfmann, P.J.; Zheng, N.-Y.; Huang, M.; Shaw, D.G.; Cobb, M.S.; et al. Profiling B Cell Immunodominance after SARS-CoV-2 Infection Reveals Antibody Evolution to Non-Neutralizing Viral Targets. Immunity 2021, 54, 1290–1303.e7. [Google Scholar] [CrossRef] [PubMed]
- Rodda, L.B.; Netland, J.; Shehata, L.; Pruner, K.B.; Morawski, P.A.; Thouvenel, C.D.; Takehara, K.K.; Eggenberger, J.; Hemann, E.A.; Waterman, H.R.; et al. Functional SARS-CoV-2-Specific Immune Memory Persists after Mild COVID-19. Cell 2021, 184, 169–183.e17. [Google Scholar] [CrossRef] [PubMed]
- Dolgin, E. COVID Vaccine Immunity is Waning—How Much does that Matter? Nature 2021, 597, 606–607. [Google Scholar] [CrossRef] [PubMed]
- Levin, E.G.; Lustig, Y.; Cohen, C.; Fluss, R.; Indenbaum, V.; Amit, S.; Doolman, R.; Asraf, K.; Mendelson, E.; Ziv, A.; et al. Waning Immune Humoral Response to Bnt162b2 COVID-19 Vaccine over 6 Months. N. Engl. J. Med. 2021. [Google Scholar] [CrossRef]
- Gobbi, F.; Buonfrate, D.; Moro, L.; Rodari, P.; Piubelli, C.; Caldrer, S.; Riccetti, S.; Sinigaglia, A.; Barzon, L. Antibody Response to the Bnt162b2 Mrna COVID-19 Vaccine in Subjects with Prior SARS-CoV-2 Infection. Viruses 2021, 13, 422. [Google Scholar] [CrossRef]
- Ciabattini, A.; Pastore, G.; Fiorino, F.; Polvere, J.; Lucchesi, S.; Pettini, E.; Auddino, S.; Rancan, I.; Durante, M.; Miscia, M.; et al. Evidence of SARS-CoV-2-Specific Memory B Cells Six Months after Vaccination with the Bnt162b2 Mrna Vaccine. Front. Immunol. 2021, 12, 708. [Google Scholar] [CrossRef]
- Khoury, D.S.; Cromer, D.; Reynaldi, A.; Schlub, T.E.; Wheatley, A.K.; Juno, J.A.; Davenport, M.P. What Level of Neutralising Antibody Protects from COVID-19? MedRxiv 2021. [Google Scholar] [CrossRef]
- Khoury, D.S.; Cromer, D.; Reynaldi, A.; Schlub, T.E.; Wheatley, A.K.; Juno, J.A.; Subbarao, K.; Kent, S.J.; Triccas, J.A.; Davenport, M.P. Neutralizing Antibody Levels are Highly Predictive of Immune Protection From Symptomatic SARS-CoV-2 Infection. Nat. Med. 2021, 27, 1205–1211. [Google Scholar] [CrossRef]
- Tracking SARS-CoV-2 Variants. Available online: https://www.who.int/activities/tracking-sars-cov-2-variants (accessed on 21 July 2021).
- Tada, T.; Zhou, H.; Samanovic, M.I.; Dcosta, B.M.; Cornelius, A.; Mulligan, M.J.; Landau, N.R. Comparison of Neutralizing Antibody Titers Elicited by Mrna and Adenoviral vector Vaccine against SARS-CoV-2 Variants. Biorxiv 2021. [Google Scholar] [CrossRef]
- Schmitz, A.J.; Turner, J.S.; Liu, Z.; Zhou, J.Q.; Aziati, I.D.; Chen, R.E.; Joshi, A.; Bricker, T.L.; Darling, T.L.; Adelsberg, D.C.; et al. A Vaccine-Induced Public Antibody Protects Against SARS-CoV-2 and Emerging Variants. Immunity 2021, 54, 2159–2166.e6. [Google Scholar] [CrossRef] [PubMed]
- Mishra, N.; Kumar, S.; Singh, S.; Bansal, T.; Jain, N.; Saluja, S.; Kumar, R.; Bhattacharyya, S.; Palanichamy, J.K.; Mir, R.A.; et al. Cross-Neutralization of SARS-CoV-2 by Hiv-1 Specific Broadly Neutralizing Antibodies and Polyclonal Plasma. PLoS Pathog. 2021, 17, E1009958. [Google Scholar] [CrossRef] [PubMed]
- Song, G.; He, W.-T.; Callaghan, S.; Anzanello, F.; Huang, D.; Ricketts, J.; Torres, J.L.; Beutler, N.; Peng, L.; Vargas, S.; et al. Cross-Reactive Serum and Memory B-Cell Responses to Spike Protein in SARS-CoV-2 and Endemic Coronavirus Infection. Nat. Commun. 2021, 12, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Kageyama, T.; Ikeda, K.; Tanaka, S.; Taniguchi, T.; Igari, H.; Onouchi, Y.; Kaneda, A.; Matsushita, K.; Hanaoka, H.; Nakada, T.-A.; et al. Antibody Responses to Bnt162b2 Mrna COVID-19 Vaccine and Their Predictors among Healthcare Workers in A Tertiary Referral Hospital in Japan. Clin. Microbiol. Infect. 2021, 21, 00437-7. [Google Scholar] [CrossRef] [PubMed]
- Ebinger, J.E.; Fert-Bober, J.; Printsev, I.; Wu, M.; Sun, N.; Prostko, J.C.; Frias, E.C.; Stewart, J.L.; Van Eyk, J.E.; Braun, J.G.; et al. Antibody Responses to the Bnt162b2 Mrna Vaccine in Individuals Previously Infected with SARS-CoV-2. Nat. Med. 2021, 27, 981–984. [Google Scholar] [CrossRef]
- Morales-Núñez, J.; Muñoz-Valle, J.; Meza-López, C.; Wang, L.-F.; Sulbarán, A.M.; Torres-Hernández, P.; Bedolla-Barajas, M.; De La O-Gómez, B.; Balcázar-Félix, P.; Hernández-Bello, J. Neutralizing Antibodies Titers and Side Effects in Response to Bnt162b2 Vaccine in Healthcare Workers with and Without Prior SARS-CoV-2 Infection. Vaccines 2021, 9, 742. [Google Scholar] [CrossRef]
- Zimmermann, P.; Curtis, N. Factors that Influence the Immune Response to Vaccination. Clin. Microbiol. Rev. 2019, 32, e00084-18. [Google Scholar] [CrossRef]
- Pang, N.Y.-L.; Pang, A.S.-R.; Chow, V.T.; Wang, D.-Y. Understanding Neutralising Antibodies against SARS-CoV-2 and Their Implications in Clinical Practice. Mil. Med. Res. 2021, 8, 1–17. [Google Scholar] [CrossRef]
- Gozalbo-Rovira, R.; Gimenez, E.; Latorre, V.; Francés-Gómez, C.; Albert, E.; Buesa, J.; Marina, A.; Blasco, M.L.; Signes-Costa, J.; Rodríguez-Díaz, J.; et al. SARS-CoV-2 Antibodies, Serum Inflammatory Biomarkers and Clinical Severity of Hospitalized COVID-19 Patients. J. Clin. Virol. 2020, 131, 104611. [Google Scholar] [CrossRef]
- Eroshenko, N.; Gill, T.; Keaveney, M.K.; Church, G.M.; Trevejo, J.M.; Rajaniemi, H. Implications of Antibody-Dependent Enhancement of Infection for SARS-CoV-2 Countermeasures. Nat. Biotechnol. 2020, 38, 789–791. [Google Scholar] [CrossRef]
- Chen, W.; Zhang, J.; Qin, X.; Wang, W.; Xu, M.; Wang, L.-F.; Xu, C.; Tang, S.; Liu, P.; Zhang, L.; et al. SARS-CoV-2 Neutralizing Antibody Levels are Correlated with Severity of COVID-19 Pneumonia. Biomed. Pharmacother. 2020, 130, 110629. [Google Scholar] [CrossRef]
- Mendoza, E.J.; Manguiat, K.; Wood, H.; Drebot, M. Two Detailed Plaque Assay Protocols for the Quantification of Infectious SARS-CoV-2. Curr. Protoc. Microbiol. 2020, 57, Ecpmc105. [Google Scholar] [CrossRef] [PubMed]
- Roehrig, J.T.; Hombach, J.; Barrett, A.D. Guidelines for Plaque Reduction Neutralization Testing of Human Antibodies to Dengue Viruses. Viral Immunol. 2007, 21, 123–132. [Google Scholar] [CrossRef]
- Abe, K.T.; Li, Z.; Samson, R.; Samavarchi-Tehrani, P.; Valcourt, E.J.; Wood, H.; Dupuis, A.P. A Simple Protein-based Surrogate Neutralization Assay for SARS-CoV-2. JCI insiggt. 2020, 5, e142362. [Google Scholar] [CrossRef] [PubMed]
- Perera, R.A.; Mok, C.K.; Tsang, O.T.; Lv, H.; Ko, R.L.; Wu, N.C.; Yuan, M.; Leung, W.S.; Mc Chan, J.; Chik, T.S.; et al. Serological Assays for Severe Acute Respiratory Syndrome Coronavirus 2 (SARS-CoV-2). Eurosurveillance 2020, 25, 2000421. [Google Scholar] [CrossRef]
- Okba, N.M.A.; Müller, M.A.; Li, W.; Wang, C.; Geurtsvankessel, C.H.; Corman, V.M.; Lamers, M.M.; Sikkema, R.S.; De Bruin, E.; Chandler, F.D.; et al. Severe Acute Respiratory Syndrome Coronavirus 2−Specific Antibody Responses in Coronavirus Disease Patients. Emerg. Infect. Dis. 2020, 26, 1478–1488. [Google Scholar] [CrossRef]
- Crawford, K.H.; Eguia, R.; Dingens, A.S.; Loes, A.N.; Malone, K.D.; Wolf, C.R.; Bloom, J.D. Protocol and Reagents for Pseudotyping Lentiviral Particles with SARS-CoV-2 Spike Protein for Neutralization Assays. Biorxiv 2020, 12, 513. [Google Scholar] [CrossRef] [PubMed]
- Ferrara, F.; Temperton, N. Pseudotype Neutralization Assays: From Laboratory Bench to Data Analysis. Methods Protoc. 2018, 1, 8. [Google Scholar] [CrossRef]
- Ou, X.; Liu, Y.; Lei, X.; Li, P.; Mi, D.; Ren, L.; Guo, L.; Guo, R.; Chen, T.; Hu, J.; et al. Characterization OF Spike Glycoprotein of SARS-CoV-2 on Virus Entry and its Immune Cross-Reactivity with SARS-CoV. Nat. Commun. 2020, 11, 1620. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Li, R.; Pan, Z.; Qian, C.; Yang, Y.; You, R.; Zhao, J.; Liu, P.; Gao, L.; Li, Z.; et al. Human Monoclonal Antibodies Block the Binding of SARS-CoV-2 Spike Protein to Angiotensin Converting Enzyme 2 Receptor. Cell. Mol. Immunol. 2020, 17, 647–649. [Google Scholar] [CrossRef]
- Xiong, H.-L.; Wu, Y.-T.; Cao, J.-L.; Yang, R.; Liu, Y.-X.; Ma, J.; Qiao, X.-Y.; Yao, X.-Y.; Zhang, B.-H.; Zhang, Y.-L.; et al. Robust Neutralization Assay based on SARS-CoV-2 S-Protein-Bearing Vesicular Stomatitis Virus (Vsv) Pseudovirus and Ace2-Overexpressing Bhk21 Cells. Emerg. Microbes Infect. 2020, 9, 2105–2113. [Google Scholar] [CrossRef]
- Nie, J.; Li, Q.; Wu, J.; Zhao, C.; Hao, H.; Liu, H.; Zhang, L.; Nie, L.; Qin, H.; Wang, M.; et al. Establishment and Validation of A Pseudovirus Neutralization Assay for SARS-CoV-2. Emerg. Microbes Infect. 2020, 9, 680–686. [Google Scholar] [CrossRef]
- Hoffmann, M.; Kleine-Weber, H.; Schroeder, S.; Krüger, N.; Herrler, T.; Erichsen, S.; Schiergens, T.S.; Herrler, G.; Wu, N.-H.; Nitsche, A.; et al. SARS-CoV-2 Cell Entry Depends on Ace2 and Tmprss2 and is Blocked by a Clinically Proven Protease Inhibitor. Cell 2020, 181, 271–280.e8. [Google Scholar] [CrossRef] [PubMed]
- Chen, Q.; Nie, J.; Huang, W.; Jiao, Y.; Li, L.; Zhang, T.; Zhao, J.; Wu, H.; Wang, Y. Development and Optimization of a Sensitive Pseudovirus-based Assay for Hiv-1 Neutralizing Antibodies Detection Using A3r5 Cells. Hum. Vaccines Immunother. 2017, 14, 199–208. [Google Scholar] [CrossRef]
- Case, J.B.; Rothlauf, P.W.; Chen, R.E.; Liu, Z.; Zhao, H.; Kim, A.S.; Bloyet, L.-M.; Zeng, Q.; Tahan, S.; Droit, L.; et al. Neutralizing Antibody and Soluble Ace2 Inhibition of a Replication-Competent Vsv-SARS-CoV-2 and a Clinical Isolate of SARS-CoV-2. Cell Host Microbe 2020, 28, 475–485.e5. [Google Scholar] [CrossRef] [PubMed]
- Dieterle, M.E.; Haslwanter, D.; Bortz, R.H.; Wirchnianski, A.S.; Lasso, G.; Vergnolle, O.; Abbasi, S.A.; Fels, J.M.; Laudermilch, E.; Florez, C.; et al. A Replication-Competent Vesicular Stomatitis Virus for Studies of SARS-CoV-2 Spike-Mediated Cell Entry and its Inhibition. Cell Host Microbe 2020, 28, 486–496.e6. [Google Scholar] [CrossRef]
- Vandergaast, R.; Carey, T.; Reiter, S.; Lech, P.; Gnanadurai, C.; Tesfay, M.; Russell, S.J. Development and Validation of Immuno-CovTM: A High-Throughput Clinical Assay for Detecting Antibodies that Neutralize SARS-CoV-2. Biorxiv 2020, 1–32. [Google Scholar] [CrossRef]
- Nakane, S.; Matsuda, Z. Dual Split Protein (Dsp) Assay to Monitor Cell–Cell Membrane Fusion. Methods Mol. Biol. 2015, 1313, 229–236. [Google Scholar] [CrossRef]
- Meyer, B.; Reimerink, J.; Torriani, G.; Brouwer, F.; Godeke, G.J.; Yerly, S.; Reusken, C. Validation and Clinical Evaluation of a SARS-CoV-2 Surrogate Virus Neutralization Test (Svnt). Medrxiv 2020, 9, 2394–2403. [Google Scholar] [CrossRef]
- Andabaka, T.; Nickerson, J.W.; Rojas-Reyes, M.X.; Rueda, J.D.; Vrca, V.B.; Barsic, B. Monoclonal Antibody for Reducing the Risk of Respiratory Syncytial Virus Infection in Children. Cochrane Database Syst. Rev. 2013, 8, 2243–2376. [Google Scholar] [CrossRef]
- Mulangu, S.; Dodd, L.E.; Davey, R.T.; Tshiani Mbaya, O.; Proschan, M.; Mukadi, D.; Lusakibanza Manzo, M.; Nzolo, D.; Tshomba Oloma, A.; Ibanda, A.; et al. A Randomized, Controlled Trial of Ebola Virus Disease Therapeutics. N. Engl. J. Med. 2019, 381, 2293–2303. [Google Scholar] [CrossRef]
- Levine, M.M. Monoclonal Antibody Therapy for Ebola Virus Disease. N. Engl. J. Med. 2019, 381, 2365–2366. [Google Scholar] [CrossRef]
- Tuccori, M.; Ferraro, S.; Convertino, I.; Cappello, E.; Valdiserra, G.; Blandizzi, C.; Maggi, F.; Focosi, D. Anti-SARS-CoV-2 Neutralizing Monoclonal Antibodies: Clinical Pipeline. Mabs 2020, 12, 1854149. [Google Scholar] [CrossRef]
- Renn, A.; Fu, Y.; Hu, X.; Hall, M.D.; Simeonov, A. Fruitful Neutralizing Antibody Pipeline Brings Hope to Defeat SARS-CoV-2. Trends Pharmacol. Sci. 2020, 41, 815–829. [Google Scholar] [CrossRef]
- Jones, B.E.; Brown-Augsburger, P.L.; Corbett, K.S.; Westendorf, K.; Davies, J.; Cujec, T.P.; Wiethoff, C.M.; Blackbourne, J.L.; Heinz, B.A.; Foster, D.; et al. The Neutralizing Antibody, Ly-Cov555, Protects against SARS-CoV-2 Infection in Nonhuman Primates. Sci. Transl. Med. 2021, 13, eabf1906. [Google Scholar] [CrossRef] [PubMed]
- Chen, P.; Nirula, A.; Heller, B.; Gottlieb, R.L.; Boscia, J.; Morris, J.; Huhn, G.; Cardona, J.; Mocherla, B.; Stosor, V.; et al. SARS-CoV-2 Neutralizing Antibody Ly-Cov555 in Outpatients with COVID-19. N. Engl. J. Med. 2021, 384, 229–237. [Google Scholar] [CrossRef]
- Cathcart, A.L.; Havenar-Daughton, C.; Lempp, F.A.; Ma, D.; Schmid, M.; Agostini, M.L.; Hebner, C.M. The Dual Function Monoclonal Antibodies Vir-7831 and Vir-7832 Demonstrate Potent In Vitro And In Vivo Activity against SARS-CoV-2. Biorxiv 2021, 6. [Google Scholar] [CrossRef]
- Cao, Y.; Su, B.; Guo, X.; Sun, W.; Deng, Y.; Bao, L.; Zhu, Q.; Zhang, X.; Zheng, Y.; Geng, C.; et al. potent Neutralizing Antibodies against SARS-CoV-2 Identified by High-Throughput Single-Cell Sequencing of Convalescent Patients’ B Cells. Cell 2020, 182, 73–84.e16. [Google Scholar] [CrossRef] [PubMed]
- Baum, A.; Fulton, B.O.; Wloga, E.; Copin, R.; Pascal, K.E.; Russo, V.; Giordano, S.; Lanza, K.; Negron, N.; Ni, M.; et al. Antibody Cocktail to SARS-CoV-2 Spike Protein Prevents Rapid Mutational Escape Seen with Individual Antibodies. Science 2020, 369, 1014–1018. [Google Scholar] [CrossRef]
- Hansen, J.; Baum, A.; Pascal, K.E.; Russo, V.; Giordano, S.; Wloga, E.; Fulton, B.O.; Yan, Y.; Koon, K.; Patel, K.; et al. Studies in Humanized Mice and Convalescent Humans Yield a SARS-CoV-2 Antibody Cocktail. Science 2020, 369, 1010–1014. [Google Scholar] [CrossRef] [PubMed]
- Weinreich, D.M.; Sivapalasingam, S.; Norton, T.; Ali, S.; Gao, H.; Bhore, R.; Musser, B.J.; Soo, Y.; Rofail, D.; Im, J.; et al. Regn-Cov2, a Neutralizing Antibody Cocktail, in Outpatients with COVID-19. N. Engl. J. Med. 2021, 384, 238–251. [Google Scholar] [CrossRef]
- Kim, C.; Ryu, D.-K.; Lee, J.; Kim, Y.-I.; Seo, J.-M.; Kim, Y.-G.; Jeong, J.-H.; Kim, M.; Kim, J.-I.; Kim, P.; et al. A Therapeutic Neutralizing Antibody Targeting Receptor Binding Domain of SARS-CoV-2 Spike Protein. Nat. Commun. 2021, 12, 1–10. [Google Scholar] [CrossRef]
- Muyldermans, S. Nanobodies: Natural Single-Domain Antibodies. Annu. Rev. Biochem. 2013, 82, 775–797. [Google Scholar] [CrossRef]
- Sasisekharan, R.; Ph, Y.D. Preparing for The Future—Nanobodies for COVID-19? N. Eng. J. Med. 2021, 384, 1–4. [Google Scholar] [CrossRef] [PubMed]
- Vanlandschoot, P.; Stortelers, C.; Beirnaert, E.; Ibañez, L.I.; Schepens, B.; Depla, E.; Saelens, X. Nanobodies®: New Ammunition To Battle Viruses. Antivir. Res. 2011, 92, 389–407. [Google Scholar] [CrossRef] [PubMed]
- Xiang, Y.; Nambulli, S.; Xiao, Z.; Liu, H.; Sang, Z.; Duprex, W.P.; Shi, Y. Versatile, Multivalent Nanobody Cocktails Efficiently Neutralize SARS-CoV-2. Biorxiv 2020, 1484, 1479–1484. [Google Scholar] [CrossRef]
- Koenig, P.-A.; Das, H.; Liu, H.; Kümmerer, B.M.; Gohr, F.N.; Jenster, L.-M.; Schiffelers, L.D.J.; Tesfamariam, Y.M.; Uchima, M.; Wuerth, J.D.; et al. Structure-Guided Multivalent Nanobodies Block SARS-CoV-2 Infection and Suppress Mutational Escape. Science 2021, 371, eabe6230. [Google Scholar] [CrossRef]
- Tirado, S.M.C.; Yoon, K.-J. Antibody-Dependent Enhancement of Virus Infection and Disease. Viral Immunol. 2003, 16, 69–86. [Google Scholar] [CrossRef]
- Horwitz, J.A.; Bar-On, Y.; Lu, C.-L.; Fera, D.; Lockhart, A.; Lorenzi, J.C.C.; Nogueira, L.; Golijanin, J.; Scheid, J.F.; Seaman, M.S.; et al. Non-Neutralizing Antibodies Alter the Course of Hiv-1 Infection In Vivo. Cell 2017, 170, 637–648.e10. [Google Scholar] [CrossRef] [PubMed]
- Willey, S.; Aasa-Chapman, M.M.; O’farrell, S.; Pellegrino, P.; Williams, I.; Weiss, R.A.; Neil, S.J. Extensive Complement-Dependent Enhancement of Hiv-1 By Autologous Non-Neutralising Antibodies at Early Stages of Infection. Retrovirology 2011, 8, 16. [Google Scholar] [CrossRef]
- Wen, J.; Cheng, Y.; Ling, R.; Dai, Y.; Huang, B.; Huang, W.; Zhang, S.; Jiang, Y. Antibody-Dependent Enhancement of Coronavirus. Int. J. Infect. Dis. 2020, 100, 483–489. [Google Scholar] [CrossRef]
- Huang, X.; Yue, Y.; Li, D.; Zhao, Y.; Qiu, L.; Chen, J.; Pan, Y.; Xi, J.; Wang, X.; Sun, Q.; et al. Antibody-Dependent Enhancement of Dengue Virus Infection Inhibits Rlr-Mediated Type-I Ifn-Independent Signalling through Upregulation of Cellular Autophagy. Sci. Rep. 2016, 6, 22303. [Google Scholar] [CrossRef]
- Nimmerjahn, F.; Ravetch, J.V. Fc-Receptors as Regulators of Immunity. Adv. Immunol. 2007, 96, 179–204. [Google Scholar] [CrossRef] [PubMed]
- Morrone, S.R.; Lok, S.-M. Structural Perspectives of Antibody-Dependent Enhancement of Infection of Dengue Virus. Curr. Opin. Virol. 2019, 36, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Sánchez-Zuno, G.A.; Matuz-Flores, M.G.; González-Estevez, G.; Nicoletti, F.; Turrubiates-Hernández, F.J.; Mangano, K.; Muñoz-Valle, J.F. A Review: Antibody-Dependent Enhancement In COVID-19: The not so Friendly Side of Antibodies. Int. J. Immunopathol. Pharmacol. 2021, 35, 20587384211050199. [Google Scholar] [CrossRef] [PubMed]
- Lee, W.S.; Wheatley, A.K.; Kent, S.J.; Dekosky, B.J. Antibody-Dependent Enhancement and SARS-CoV-2 Vaccines and Therapies. Nat. Microbiol. 2020, 5, 1185–1191. [Google Scholar] [CrossRef]
- Ricke, D.O. Two Different Antibody-Dependent Enhancement (Ade) Risks for SARS-CoV-2 Antibodies. Front. Immunol. 2021, 12, 640093. [Google Scholar] [CrossRef]
- Halstead, S.B.; Katzelnick, L. COVID-19 Vaccines: Should We Fear Ade? J. Infect. Dis. 2020, 222, 1946–1950. [Google Scholar] [CrossRef]
- Munar, B.A.C.; Parra, J.E.C.; Panduro, M.H.R. Amplificación de la Infección Dependiente de Anticuerpos en la Inmunopatogénesis del Dengue Grave, Implicaciones Para el Desarrollo y uso de las Vacunas. Acta Biol. Colomb. 2019, 24, 439–451. [Google Scholar] [CrossRef]
- Zhou, Y.; Liu, Z.; Li, S.; Xu, W.; Zhang, Q.; Silva, I.T.; Li, C.; Wu, Y.; Jiang, Q.; Liu, Z.; et al. Enhancement Versus Neutralization by SARS-CoV-2 Antibodies from a Convalescent Donor Associates with Distinct Epitopes on the Rbd. Cell Rep. 2021, 34, 108699. [Google Scholar] [CrossRef]

Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Morales-Núñez, J.J.; Muñoz-Valle, J.F.; Torres-Hernández, P.C.; Hernández-Bello, J. Overview of Neutralizing Antibodies and Their Potential in COVID-19. Vaccines 2021, 9, 1376. https://doi.org/10.3390/vaccines9121376
Morales-Núñez JJ, Muñoz-Valle JF, Torres-Hernández PC, Hernández-Bello J. Overview of Neutralizing Antibodies and Their Potential in COVID-19. Vaccines. 2021; 9(12):1376. https://doi.org/10.3390/vaccines9121376
Chicago/Turabian StyleMorales-Núñez, José Javier, José Francisco Muñoz-Valle, Paola Carolina Torres-Hernández, and Jorge Hernández-Bello. 2021. "Overview of Neutralizing Antibodies and Their Potential in COVID-19" Vaccines 9, no. 12: 1376. https://doi.org/10.3390/vaccines9121376
APA StyleMorales-Núñez, J. J., Muñoz-Valle, J. F., Torres-Hernández, P. C., & Hernández-Bello, J. (2021). Overview of Neutralizing Antibodies and Their Potential in COVID-19. Vaccines, 9(12), 1376. https://doi.org/10.3390/vaccines9121376






