Maternal COVID-19 Vaccination and Its Potential Impact on Fetal and Neonatal Development
Abstract
1. Introduction
2. Lipid Nanoparticles (LNPs) in the COVID-19 mRNA Vaccines
3. Viral Vector COVID-19 Vaccines
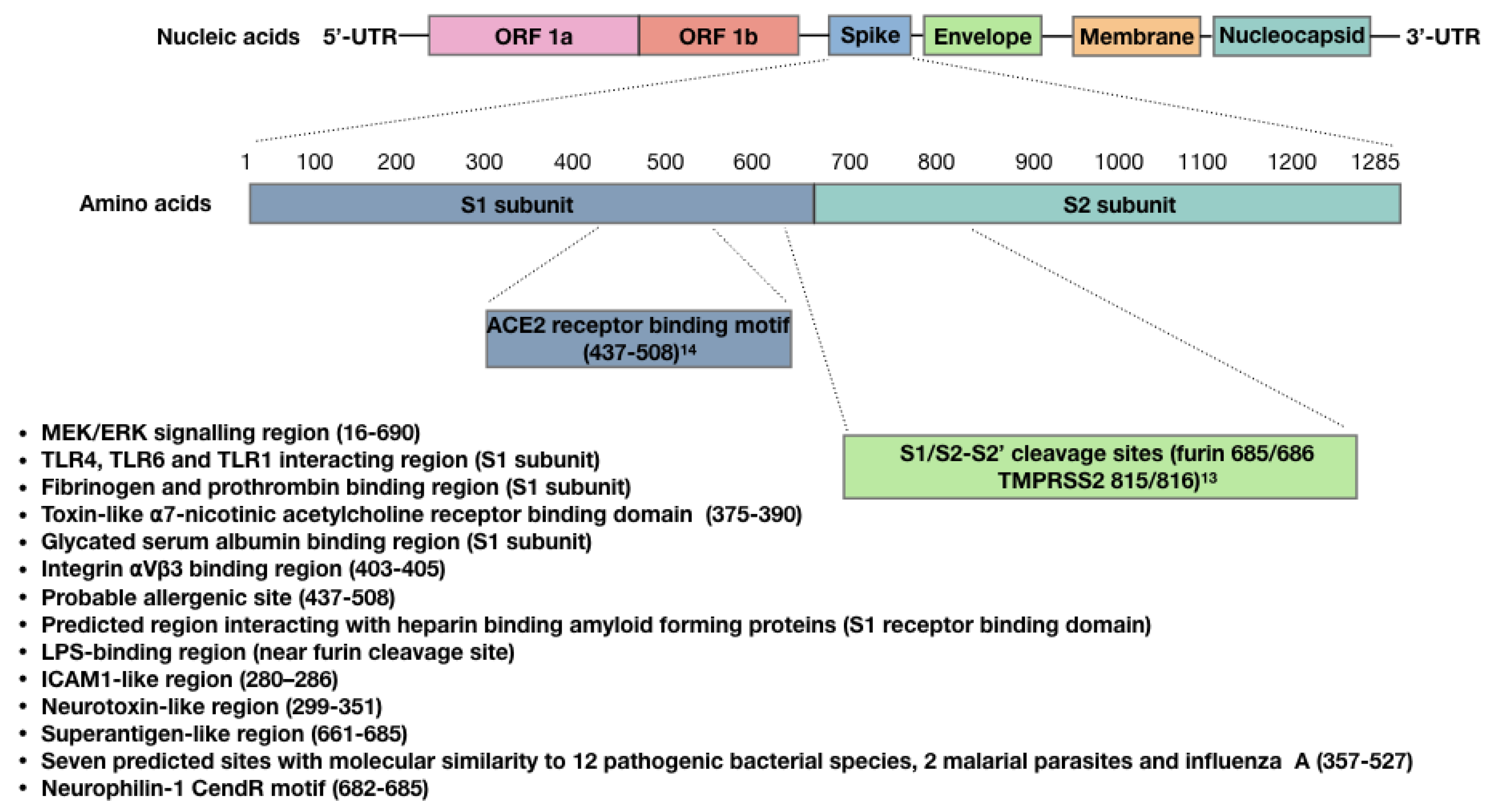
4. Bioactivity of the SARS-CoV-2 Spike Protein
5. The SARS-CoV-2 Spike Protein Triggers Autoimmune Responses
6. The SARS-CoV-2 Spike Protein and Antibody-Dependent Enhancement
7. Using an Ovine Model to Study Long-Term Outcomes of Maternal COVID-19 Vaccination
8. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Conflicts of Interest
References
- Canada PHA of Demographics: COVID-19 Vaccination Coverage in Canada—Canada.ca [Internet]. aem. 2021. Available online: https://health-infobase.canada.ca/covid-19/vaccination-coverage/ (accessed on 13 September 2021).
- Reuters. WHO Warns Individuals against Mixing and Matching COVID Vaccines. Reuters. 14 July 2021. Available online: https://www.reuters.com/business/healthcare-pharmaceuticals/who-warns-against-mixing-matching-covid-vaccines-2021-07-12/ (accessed on 13 September 2021).
- Ciarambino, T.; Para, O.; Giordano, M. Immune system and COVID-19 by sex differences and age. Womens Health 2021, 17, 17455065211022262. [Google Scholar] [CrossRef]
- Number of births in Canada 2020 [Internet]. Statista. Available online: https://www.statista.com/statistics/443051/number-of-births-in-canada/ (accessed on 13 September 2021).
- Maykin, M.M.; Heuser, C.; Feltovich, H. With the Society for Maternal-Fetal Medicine Health Policy Advocacy Committee. Pregnant people deserve the protection offered by SARS-CoV-2 vaccines. Vaccine 2021, 39, 171–172. [Google Scholar] [CrossRef]
- Klein, S.L.; Creisher, P.S.; Burd, I. COVID-19 vaccine testing in pregnant females is necessary. J. Clin. Investig. 2021, 131, 147553. [Google Scholar] [CrossRef]
- Pineles, B.L.; Goodman, K.E.; Pineles, L.; O’Hara, L.M.; Nadimpalli, G.; Magder, L.S.; Baghdadi, J.D.; Parchem, J.G.; Harris, A.D. In-Hospital Mortality in a Cohort of Hospitalized Pregnant and Nonpregnant Patients With COVID-19. Ann Intern Med. 2021, 174, 1186–1188. [Google Scholar] [CrossRef]
- CDC. COVID-19 Vaccination [Internet]. Centers for Disease Control and Prevention. 2020. Available online: https://www.cdc.gov/coronavirus/2019-ncov/vaccines/safety/vsafepregnancyregistry.html (accessed on 13 September 2021).
- Shimabukuro, T.T.; Kim, S.Y.; Myers, T.R.; Moro, P.L.; Oduyebo, T.; Panagiotakopoulos, L.; Marquez, P.L.; Olson, C.K.; Liu, R.; Cheng, R.T.; et al. Preliminary Findings of mRNA Covid-19 Vaccine Safety in Pregnant Persons. N. Engl. J. Med. 2021, 384, 2273–2282. [Google Scholar] [CrossRef]
- Preliminary Findings of mRNA Covid-19 Vaccine Safety in Pregnant Persons. New England Journal of Medicine [Internet]. 8 September 2021. Available online: https://www.nejm.org/doi/10.1056/NEJMx210016 (accessed on 4 November 2021).
- Lancet Commission on COVID-19 Vaccines and Therapeutics Task Force Members. Operation Warp Speed: implications for global vaccine security. Lancet Glob Health 2021, 9, e1017–e1021. [Google Scholar]
- Sun, H. On Preliminary Findings of mRNA Covid-19 Vaccine Safety in Pregnant Persons. N. Engl. J. Med. 2021, 385, 1535–1536. [Google Scholar] [CrossRef]
- Meaney-Delman, D.M.; Ellington, S.R.; Shimabukuro, T.T. On Preliminary Findings of mRNA Covid-19 Vaccine Safety in Pregnant Persons. Reply. N. Engl. J. Med. 2021, 385, 1536. [Google Scholar] [CrossRef]
- Sprent, J.; King, C. COVID-19 vaccine side effects: The positives about feeling bad. Sci. Immunol. 2021, 6, eabj9256. [Google Scholar] [CrossRef]
- Casazza, R.L.; Lazear, H.M.; Miner, J.J. Protective and Pathogenic Effects of Interferon Signaling During Pregnancy. Viral Immunol. 2020, 33, 3–11. [Google Scholar] [CrossRef]
- Gómez-Chávez, F.; Correa, D.; Navarrete-Meneses, P.; Cancino-Diaz, J.C.; Cancino-Diaz, M.E.; Rodríguez-Martínez, S. NF-κB and Its Regulators During Pregnancy. Front. Immunol. 2021, 12, 679106. [Google Scholar] [CrossRef]
- Seneff, S.; Nigh, G. Worse Than the Disease? Reviewing Some Possible Unintended Consequences of the mRNA Vaccines Against COVID-19. Int. J. Vaccine Theory Pract. Res. 2021, 2, 38–79. [Google Scholar]
- Schoenmaker, L.; Witzigmann, D.; Kulkarni, J.A.; Verbeke, R.; Kersten, G.; Jiskoot, W.; Crommelin, D.J.A. mRNA-lipid nanoparticle COVID-19 vaccines: Structure and stability. Int. J. Pharm. 2021, 601, 120586. [Google Scholar] [CrossRef] [PubMed]
- Schädlich, A.; Hoffmann, S.; Mueller, T.; Caysa, H.; Rose, C.; Göpferich, A.; Li, J.; Kuntsche, J.; Mäder, K. Accumulation of nanocarriers in the ovary: A neglected toxicity risk? J. Control Release 2012, 160, 105–112. [Google Scholar] [CrossRef]
- Doshi, P. Covid-19 vaccines: In the rush for regulatory approval, do we need more data? BMJ 2021, 373, n1244. [Google Scholar] [CrossRef] [PubMed]
- Lyzogubov, V.V.; Tytarenko, R.G.; Liu, J.; Bora, N.S.; Bora, P.S. Polyethylene glycol (PEG)-induced mouse model of choroidal neovascularization. J. Biol. Chem. 2011, 286, 16229–16237. [Google Scholar] [CrossRef]
- Patel, S.; Barnett, J.M.; Kim, S.J. Retinal Toxicity of Intravitreal Polyethylene Glycol 400. J. Ocul. Pharmacol. Ther. 2016, 32, 97–101. [Google Scholar] [CrossRef]
- Subramony, R.; Lin, L.C.; Knight, D.K.; Aminlari, A.; Belovarski, I. Bilateral Retinal Detachments in a Healthy 22-year-old Female Following Moderna SARS-CoV-2 Vaccination. J. Emerg. Med. 2021. Available online: https://www.sciencedirect.com/science/article/pii/S0736467921006119 (accessed on 10 September 2021). [CrossRef]
- Fowler, N.; Martinez, N.R.M.; Pallares, B.V.; Maldonado, R.S. Acute-onset central serous retinopathy after immunization with COVID-19 mRNA vaccine. Am. J. Ophthalmol. Case Rep. 2021, 23, 101136. [Google Scholar] [CrossRef]
- Samaridou, E.; Heyes, J.; Lutwyche, P. Lipid nanoparticles for nucleic acid delivery: Current perspectives. Adv. Drug Deliv. Rev. 2020, 154–155, 37–63. [Google Scholar] [CrossRef]
- Ndeupen, S.; Qin, Z.; Jacobsen, S.; Estanbouli, H.; Bouteau, A.; Igyártó, B.Z. The mRNA-LNP platform’s lipid nanoparticle component used in preclinical vaccine studies is highly inflammatory. bioRxiv 2021. [Google Scholar] [CrossRef]
- De Oliveira Viana, I.M.; Roussel, S.; Defrêne, J.; Lima, E.M.; Barabé, F.; Bertrand, N. Innate and adaptive immune responses toward nanomedicines. Acta Pharm. Sin. B 2021, 11, 852–870. [Google Scholar] [CrossRef]
- Sokolowska, M.; Eiwegger, T.; Ollert, M.; Torres, M.J.; Barber, D.; Del Giacco, S.; Jutel, M.; Nadeau, K.C.; Palomares, O.; Rabin, R.L.; et al. EAACI statement on the diagnosis, management and prevention of severe allergic reactions to COVID-19 vaccines. Allergy 2021, 76, 1629–1639. [Google Scholar] [CrossRef] [PubMed]
- Castells, M.C.; Phillips, E.J. Maintaining Safety with SARS-CoV-2 Vaccines. N. Engl. J. Med. 2021, 384, 643–649. [Google Scholar] [CrossRef] [PubMed]
- Selvaraj, G.; Kaliamurthi, S.; Peslherbe, G.H.; Wei, D.-Q. Are the Allergic Reactions of COVID-19 Vaccines Caused by mRNA Constructs or Nanocarriers? Immunological Insights. Interdiscip Sci. 2021, 13, 344–347. [Google Scholar] [CrossRef] [PubMed]
- McCall, S.J.; Bonnet, M.-P.; Äyräs, O.; Vandenberghe, G.; Gissler, M.; Zhang, W.-H.; Van Leeuw, V.; Deneux-Tharaux, C.; Kurinczuk, J.J.; Knight, M. Anaphylaxis in pregnancy: A population-based multinational European study. Anaesthesia 2020, 75, 1469–1475. [Google Scholar] [CrossRef]
- Othman, M.; Labelle, A.; Mazzetti, I.; Elbatarny, H.S.; Lillicrap, D. Adenovirus-induced thrombocytopenia: The role of von Willebrand factor and P-selectin in mediating accelerated platelet clearance. Blood 2007, 109, 2832–2839. [Google Scholar] [CrossRef]
- Tsilingiris, D.; Vallianou, N.G.; Karampela, I.; Dalamaga, M. Vaccine induced thrombotic thrombocytopenia: The shady chapter of a success story. Metabol. Open 2021, 11, 100101. [Google Scholar] [CrossRef]
- Rzymski, P.; Perek, B.; Flisiak, R. Thrombotic Thrombocytopenia after COVID-19 Vaccination: In Search of the Underlying Mechanism. Vaccines 2021, 9, 559. [Google Scholar] [CrossRef]
- Tang, M.W.; Nur, E.; Biemond, B.J. Immune thrombocytopenia due to COVID-19 during pregnancy. Am. J. Hematol. 2020, 95, E191–E192. [Google Scholar] [CrossRef]
- Aminimoghaddam, S.; Afrooz, N.; Nasiri, S.; Motaghi Nejad, O.; Mahmoudzadeh, F. A COVID-19 pregnant patient with thrombotic thrombocytopenic purpura: A case report. J. Med. Case Rep. 2021, 15, 104. [Google Scholar] [CrossRef]
- Serrano, C.; Español, I.; Cascales, A.; Moraleda, J.M. Frequently Relapsing Post-COVID-19 Immune Thrombocytopenia. SN Compr Clin Med [Internet]. 20 July 2021. Available online: https://doi.org/10.1007/s42399-021-01019-7 (accessed on 25 August 2021).
- Jawed, M.; Khalid, A.; Rubin, M.; Shafiq, R.; Cemalovic, N. Acute Immune Thrombocytopenia (ITP) Following COVID-19 Vaccination in a Patient With Previously Stable ITP. Open Forum Infect Dis. 2021, 8, ofab343. [Google Scholar] [CrossRef]
- Virani, A.; Rabold, E.; Hanson, T.; Haag, A.; Elrufay, R.; Cheema, T.; Balaan, M.; Bhanot, N. Guillain-Barré Syndrome associated with SARS-CoV-2 infection. IDCases 2020, 20, e00771. [Google Scholar] [CrossRef]
- Zhao, H.; Shen, D.; Zhou, H.; Liu, J.; Chen, S. Guillain-Barré syndrome associated with SARS-CoV-2 infection: Causality or coincidence? Lancet Neurol. 2020, 19, 383–384. [Google Scholar] [CrossRef]
- Sedaghat, Z.; Karimi, N. Guillain Barre syndrome associated with COVID-19 infection: A case report. J. Clin. Neurosci. 2020, 76, 233–235. [Google Scholar] [CrossRef] [PubMed]
- Toscano, G.; Palmerini, F.; Ravaglia, S.; Ruiz, L.; Invernizzi, P.; Cuzzoni, M.G.; Franciotta, D.; Baldanti, F.; Daturi, R.; Postorino, P.; et al. Guillain-Barré Syndrome Associated with SARS-CoV-2. N. Engl. J. Med. 2020, 382, 2574–2576. [Google Scholar] [CrossRef]
- Patel, S.U.; Khurram, R.; Lakhani, A.; Quirk, B. Guillain-Barre syndrome following the first dose of the chimpanzee adenovirus-vectored COVID-19 vaccine, ChAdOx1. BMJ Case Rep. 2021, 14, e242956. [Google Scholar] [CrossRef] [PubMed]
- Donofrio, P.D. Guillain-Barré Syndrome. CONTINUUM Lifelong Learn. Neurol. 2017, 23, 1295–1309. [Google Scholar] [CrossRef]
- Sharma, S.R.; Sharma, N.; Masaraf, H.; Singh, S.A. Guillain-Barré syndrome complicating pregnancy and correlation with maternal and fetal outcome in North Eastern India: A retrospective study. Ann. Indian Acad. Neurol. 2015, 18, 215–218. [Google Scholar] [CrossRef] [PubMed]
- Mohkhedkar, M.; Venigalla, S.S.K.; Janakiraman, V. Autoantigens That May Explain Postinfection Autoimmune Manifestations in Patients With Coronavirus Disease 2019 Displaying Neurological Conditions. J. Infect. Dis. 2021, 223, 536–537. [Google Scholar] [CrossRef]
- Schwartzberg, L.S.; Navari, R.M. Safety of Polysorbate 80 in the Oncology Setting. Adv. Ther. 2018, 35, 754–767. [Google Scholar] [CrossRef]
- Stone, C.A.; Liu, Y.; Relling, M.V.; Krantz, M.S.; Pratt, A.L.; Abreo, A.; Hemler, J.A.; Phillips, E.J. Immediate Hypersensitivity to Polyethylene Glycols and Polysorbates: More Common Than We Have Recognized. J. Allergy Clin. Immunol. Pract. 2019, 7, 1533–1540.e8. [Google Scholar] [CrossRef]
- Zhang, L.; Richards, A.; Barrasa, M.I.; Hughes, S.H.; Young, R.A.; Jaenisch, R. Reverse-Transcribed SARS-CoV-2 RNA Can Integrate into the Genome of Cultured Human Cells and can be Expressed in Patient-Derived Tissues. PNAS 2021, 118. Available online: https://www.pnas.org/content/118/21/e2105968118 (accessed on 25 August 2021). [CrossRef]
- Doerfler, W. Adenoviral Vector DNA- and SARS-CoV-2 mRNA-Based Covid-19 Vaccines: Possible Integration into the Human Genome—Are Adenoviral Genes Expressed in Vector-based Vaccines? Virus Res. 2021, 302, 198466. [Google Scholar] [CrossRef] [PubMed]
- Al-Benna, S. Association of high level gene expression of ACE2 in adipose tissue with mortality of COVID-19 infection in obese patients. Obes. Med. 2020, 19, 100283. [Google Scholar] [CrossRef] [PubMed]
- Ashraf, U.M.; Abokor, A.A.; Edwards, J.M.; Waigi, E.W.; Royfman, R.S.; Hasan, S.A.-M.; Smedlund, K.B.; Hardy, A.M.G.; Chakravarti, R.; Koch, L.G. SARS-CoV-2, ACE2 expression, and systemic organ invasion. Physiol. Genom. 2021, 53, 51–60. [Google Scholar] [CrossRef] [PubMed]
- Gengler, C.; Dubruc, E.; Favre, G.; Greub, G.; de Leval, L.; Baud, D. SARS-CoV-2 ACE-receptor detection in the placenta throughout pregnancy. Clin. Microbiol. Infect. 2021, 27, 489–490. [Google Scholar] [CrossRef]
- Dhaundiyal, A.; Kumari, P.; Jawalekar, S.S.; Chauhan, G.; Kalra, S.; Navik, U. Is highly expressed ACE 2 in pregnant women “a curse” in times of COVID-19 pandemic? Life Sci. 2021, 264, 118676. [Google Scholar] [CrossRef]
- Argueta, L.B.; Lacko, L.A.; Bram, Y.; Tada, T.; Carrau, L.; Zhang, T.; Uhl, S.; Lubor, B.C.; Chandar, V.; Gil, C.; et al. SARS-CoV-2 Infects Syncytiotrophoblast and Activates Inflammatory Responses in the Placenta. bioRxiv 2021, 17, 2021. [Google Scholar] [CrossRef]
- Flores-Pliego, A.; Miranda, J.; Vega-Torreblanca, S.; Valdespino-Vázquez, Y.; Helguera-Repetto, C.; Espejel-Nuñez, A.; Borboa-Olivares, H.; Espino Y Sosa, S.; Mateu-Rogell, P.; León-Juárez, M.; et al. Molecular Insights into the Thrombotic and Microvascular Injury in Placental Endothelium of Women with Mild or Severe COVID-19. Cells 2021, 10, 364. [Google Scholar] [CrossRef]
- Ogata, A.F.; Cheng, C.-A.; Desjardins, M.; Senussi, Y.; Sherman, A.C.; Powell, M.; Novack, L.; Von, S.; Li, X.; Baden, L.R.; et al. Circulating SARS-CoV-2 Vaccine Antigen Detected in the Plasma of mRNA-1273 Vaccine Recipients. Clin Infect Dis. 2021, 20, ciab465. [Google Scholar] [CrossRef]
- Lei, Y.; Zhang, J.; Schiavon, C.R.; He, M.; Chen, L.; Shen, H.; Zhang, Y.; Yin, Q.; Cho, Y.; Andrade, L.; et al. SARS-CoV-2 Spike Protein Impairs Endothelial Function via Downregulation of ACE2. bioRxiv 2021, 128, 1323–1326. [Google Scholar] [CrossRef]
- Zhou, Y.; Wang, M.; Li, Y.; Wang, P.; Zhao, P.; Yang, Z.; Wang, S.; Zhang, L.; Li, Z.; Jia, K.; et al. SARS-CoV-2 Spike protein enhances ACE2 expression via facilitating Interferon effects in bronchial epithelium. Immunol. Lett. 2021, 237, 33–41. [Google Scholar] [CrossRef]
- Ratajczak, M.Z.; Bujko, K.; Ciechanowicz, A.; Sielatycka, K.; Cymer, M.; Marlicz, W.; Kucia, M. SARS-CoV-2 Entry Receptor ACE2 Is Expressed on Very Small CD45- Precursors of Hematopoietic and Endothelial Cells and in Response to Virus Spike Protein Activates the Nlrp3 Inflammasome. Stem Cell Rev. Rep. 2021, 17, 266–277. [Google Scholar] [CrossRef] [PubMed]
- Ropa, J.; Cooper, S.; Capitano, M.L.; Van’t Hof, W.; Broxmeyer, H.E. Human Hematopoietic Stem, Progenitor, and Immune Cells Respond Ex Vivo to SARS-CoV-2 Spike Protein. Stem Cell Rev. Rep. 2021, 17, 253–265. [Google Scholar] [CrossRef]
- Colunga Biancatelli, R.M.L.; Solopov, P.A.; Sharlow, E.R.; Lazo, J.S.; Marik, P.E.; Catravas, J.D. The SARS-CoV-2 spike protein subunit S1 induces COVID-19-like acute lung injury in Κ18-hACE2 transgenic mice and barrier dysfunction in human endothelial cells. Am. J. Physiol. Lung Cell Mol. Physiol. 2021, 321, L477–L484. [Google Scholar] [CrossRef]
- Suzuki, Y.J.; Nikolaienko, S.I.; Dibrova, V.A.; Dibrova, Y.V.; Vasylyk, V.M.; Novikov, M.Y.; Shults, N.V.; Gychka, S.G. SARS-CoV-2 spike protein-mediated cell signaling in lung vascular cells. Vascul Pharmacol. 2021, 137, 106823. [Google Scholar] [CrossRef]
- Chen, I.-Y.; Chang, S.C.; Wu, H.-Y.; Yu, T.-C.; Wei, W.-C.; Lin, S.; Chien, C.L.; Chang, M.F. Upregulation of the chemokine (C-C motif) ligand 2 via a severe acute respiratory syndrome coronavirus spike-ACE2 signaling pathway. J. Virol. 2010, 84, 7703–7712. [Google Scholar] [CrossRef]
- Nader, D.; Fletcher, N.; Curley, G.F.; Kerrigan, S.W. SARS-CoV-2 uses major endothelial integrin αvβ3 to cause vascular dysregulation in-vitro during COVID-19. PLoS ONE 2021, 16, e0253347. [Google Scholar] [CrossRef] [PubMed]
- Gautam, I.; Storad, Z.; Filipiak, L.; Huss, C.; Meikle, C.K.; Worth, R.G.; Wuescher, L.M. From Classical to Unconventional: The Immune Receptors Facilitating Platelet Responses to Infection and Inflammation. Biology 2020, 9, 343. [Google Scholar] [CrossRef]
- Shen, S.; Zhang, J.; Fang, Y.; Lu, S.; Wu, J.; Zheng, X.; Deng, F. SARS-CoV-2 interacts with platelets and megakaryocytes via ACE2-independent mechanism. J. Hematol. Oncol. 2021, 14, 72. [Google Scholar] [CrossRef] [PubMed]
- Campbell, R.A.; Boilard, E.; Rondina, M.T. Is there a role for the ACE2 receptor in SARS-CoV-2 interactions with platelets? J. Thromb. Haemost. 2021, 19, 46–50. [Google Scholar] [CrossRef]
- Choudhury, A.; Mukherjee, S. In silico studies on the comparative characterization of the interactions of SARS-CoV-2 spike glycoprotein with ACE-2 receptor homologs and human TLRs. J. Med. Virol. 2020, 92, 2105–2113. [Google Scholar] [CrossRef] [PubMed]
- Shirato, K.; Kizaki, T. SARS-CoV-2 spike protein S1 subunit induces pro-inflammatory responses via toll-like receptor 4 signaling in murine and human macrophages. Heliyon 2021, 7, e06187. [Google Scholar] [CrossRef] [PubMed]
- Andonegui, G.; Kerfoot, S.M.; McNagny, K.; Ebbert, K.V.J.; Patel, K.D.; Kubes, P. Platelets express functional Toll-like receptor-4. Blood 2005, 106, 2417–2423. [Google Scholar] [CrossRef] [PubMed]
- Ouyang, W.; Xie, T.; Fang, H.; Gao, C.; Stantchev, T.; Clouse, K.A.; Yuan, K.; Ju, T.; Frucht, D.M. Variable Induction of Pro-Inflammatory Cytokines by Commercial SARS CoV-2 Spike Protein Reagents: Potential Impacts of LPS on In Vitro Modeling and Pathogenic Mechanisms In Vivo. Int. J. Mol. Sci. 2021, 22, 7540. [Google Scholar] [CrossRef]
- Petruk, G.; Puthia, M.; Petrlova, J.; Samsudin, F.; Strömdahl, A.-C.; Cerps, S.; Uller, L.; Kjellström, S.; Bond, P.J.; Schmidtchen, A.A. SARS-CoV-2 spike protein binds to bacterial lipopolysaccharide and boosts proinflammatory activity. J. Mol. Cell Biol. 2020, 12, 916–932. [Google Scholar] [CrossRef] [PubMed]
- Grobbelaar, L.M.; Venter, C.; Vlok, M.; Ngoepe, M.; Laubscher, G.J.; Lourens, P.J.; Steenkamp, J.; Kell, D.B.; Pretorius, E. SARS-CoV-2 spike protein S1 induces fibrin(ogen) resistant to fibrinolysis: Implications for microclot formation in COVID-19. Biosci Rep. 2021, 41, BSR20210611. [Google Scholar] [CrossRef]
- IIes, J.K.; Zmuidinaite, R.; Sadee, C.; Gardiner, A.; Lacey, J.; Harding, S.; Ule, J.; Roblett, D.; Heeney, J.L.; Baxendale, H.E.; et al. SARS-CoV-2 Spike Protein Binding of Glycated Serum Albumin—Its Potential Role in the Pathogenesis of the COVID-19 Clinical Syndromes and Bias towards Individuals with Pre-Diabetes/Type 2 Diabetes & Metabolic Diseases [Internet]. 2021, p. 21258871. Available online: https://www.medrxiv.org/content/10.1101/2021.06.14.21258871v3 (accessed on 28 August 2021).
- Lagoumintzis, G.; Chasapis, C.T.; Alexandris, N.; Kouretas, D.; Tzartos, S.; Eliopoulos, E.; Farsalinos, K.; Poulas, K. Nicotinic cholinergic system and COVID-19: In silico identification of interactions between α7 nicotinic acetylcholine receptor and the cryptic epitopes of SARS-Co-V and SARS-CoV-2 Spike glycoproteins. Food Chem. Toxicol. 2021, 149, 112009. [Google Scholar] [CrossRef]
- Cantuti-Castelvetri, L.; Ojha, R.; Pedro, L.D.; Djannatian, M.; Franz, J.; Kuivanen, S.; van der Meer, F.; Kallio, K.; Kaya, T.; Anastasina, M.; et al. Neuropilin-1 facilitates SARS-CoV-2 cell entry and infectivity. Science 2020, 370, 856–860. [Google Scholar] [CrossRef]
- Shiers, S.; Ray, P.R.; Wangzhou, A.; Tatsui, C.E.; Rhines, L.; Li, Y.; Uhelski, M.L.; Dougherty, P.M.; Price, T.J. ACE2 expression in human dorsal root ganglion sensory neurons: Implications for SARS-CoV-2 virus-induced neurological effects [Internet]. May 2020, p. 2020.05.28.122374. Available online: https://www.biorxiv.org/content/10.1101/2020.05.28.122374v1 (accessed on 25 August 2021).
- Mao, L.; Jin, H.; Wang, M.; Hu, Y.; Chen, S.; He, Q.; Chang, J.; Hong, C.; Zhou, Y.; Wang, D.; et al. Neurologic Manifestations of Hospitalized Patients With Coronavirus Disease 2019 in Wuhan, China. JAMA Neurol. 2020, 77, 683–690. [Google Scholar] [CrossRef]
- Daly, J.L.; Simonetti, B.; Klein, K.; Chen, K.-E.; Williamson, M.K.; Antón-Plágaro, C.; Shoemark, D.K.; Simón-Gracia, L.; Bauer, M.; Hollandi, R.; et al. Neuropilin-1 is a host factor for SARS-CoV-2 infection. Science 2020, 370, 861–865. [Google Scholar] [CrossRef]
- Moutal, A.; Martin, L.F.; Boinon, L.; Gomez, K.; Ran, D.; Zhou, Y.; Stratton, H.J.; Cai, S.; Luo, S.; Gonzalez, K.B.; et al. SARS-CoV-2 spike protein co-opts VEGF-A/neuropilin-1 receptor signaling to induce analgesia. Pain 2021, 162, 243–252. [Google Scholar] [CrossRef] [PubMed]
- Idrees, D.; Kumar, V. SARS-CoV-2 spike protein interactions with amyloidogenic proteins: Potential clues to neurodegeneration. Biochem. Biophys. Res. Commun. 2021, 554, 94–98. [Google Scholar] [CrossRef] [PubMed]
- Classen, J.B. COVID-19 RNA Based Vaccines and the Risk of Prion Disease. Microbiol. Infect Dis. 2021, 5. Available online: https://scivisionpub.com/pdfs/covid19-rna-based-vaccines-and-the-risk-of-prion-disease-1503.pdf (accessed on 25 August 2021). [CrossRef]
- Dakal, T.C. Antigenic sites in SARS-CoV-2 spike RBD show molecular similarity with pathogenic antigenic determinants and harbors peptides for vaccine development. Immunobiology 2021, 226, 152091. [Google Scholar] [CrossRef] [PubMed]
- Noval Rivas, M.; Porritt, R.A.; Cheng, M.H.; Bahar, I.; Arditi, M. COVID-19-associated multisystem inflammatory syndrome in children (MIS-C): A novel disease that mimics toxic shock syndrome-the superantigen hypothesis. J. Allergy Clin. Immunol. 2021, 147, 57–59. [Google Scholar] [CrossRef]
- Cheng, M.H.; Zhang, S.; Porritt, R.A.; Noval Rivas, M.; Paschold, L.; Willscher, E.; Binder, M.; Arditi, M.; Bahar, I. Superantigenic character of an insert unique to SARS-CoV-2 spike supported by skewed TCR repertoire in patients with hyperinflammation. Proc. Natl. Acad. Sci. USA 2020, 117, 25254–25262. [Google Scholar] [CrossRef]
- Cheng, M.H.; Porritt, R.A.; Rivas, M.N.; Krieger, J.M.; Ozdemir, A.B.; Garcia, G.; Arumugaswami, V.; Fries, B.C.; Arditi, M.; Bahar, I. A monoclonal antibody against staphylococcal enterotoxin B superantigen inhibits SARS-CoV-2 entry in vitro. Structure 2021, 29, 951–962. [Google Scholar] [CrossRef]
- Arashkia, A.; Jalilvand, S.; Mohajel, N.; Afchangi, A.; Azadmanesh, K.; Salehi-Vaziri, M.; Fazlalipour, M.; Pouriayevali, M.H.; Jalali, T.; Mousavi Nasab, S.D.; et al. Severe acute respiratory syndrome-coronavirus-2 spike (S) protein based vaccine candidates: State of the art and future prospects. Rev. Med. Virol. 2021, 31, e2183. [Google Scholar] [CrossRef]
- Verbeke, R.; Lentacker, I.; De Smedt, S.C.; Dewitte, H. The dawn of mRNA vaccines: The COVID-19 case. J. Control Release 2021, 333, 511–520. [Google Scholar] [CrossRef]
- Patterson, B.K.; Francisco, E.B.; Yogendra, R.; Long, E.; Pise, A.; Rodrigues, H. Persistence of SARS CoV-2 S1 Protein in CD16+ Monocytes in Post-Acute Sequelae of COVID-19 (PASC) Up to 15 Months Post-Infection [Internet]. 2021, p. 449905. Available online: https://www.biorxiv.org/content/10.1101/2021.06.25.449905v3 (accessed on 25 August 2021).
- Golan, Y.; Prahl, M.; Cassidy, A.; Lin, C.Y.; Ahituv, N.; Flaherman, V.J.; Gaw, S.L. Evaluation of Messenger RNA From COVID-19 BTN162b2 and mRNA-1273 Vaccines in Human Milk. JAMA Pediatr. 2021, 175, 1069–1071. [Google Scholar] [CrossRef] [PubMed]
- Bansal, S.; Perincheri, S.; Fleming, T.; Poulson, C.; Tiffany, B.; Bremner, R.M. Cutting Edge: Circulating Exosomes with COVID Spike Protein Are Induced by BNT162b2 (Pfizer-BioNTech) Vaccination prior to Development of Antibodies: A Novel Mechanism for Immune Activation by mRNA Vaccines. J. Immunol. 2021, 15, ji2100637. [Google Scholar] [CrossRef]
- Ross, M.; Atalla, H.; Karrow, N.; Mallard, B.A. The bioactivity of colostrum and milk exosomes of high, average, and low immune responder cows on human intestinal epithelial cells. J. Dairy Sci. 2021, 104, 2499–2510. [Google Scholar] [CrossRef]
- He, Y.; He, Z.; Leone, S.; Liu, S. Milk Exosomes Transfer Oligosaccharides into Macrophages to Modulate Immunity and Attenuate Adherent-Invasive E. coli (AIEC) Infection. Nutrients 2021, 13, 3198. [Google Scholar] [CrossRef]
- Cappello, F.; Marino Gammazza, A.; Dieli, F.; Conway de Macario, E.; Macario, A.J. Does SARS-CoV-2 Trigger Stress-Induced Autoimmunity by Molecular Mimicry? A Hypothesis. J. Clin. Med. 2020, 9, 2038. [Google Scholar] [CrossRef]
- Liu, Y.; Sawalha, A.H.; Lu, Q. COVID-19 and autoimmune diseases. Curr. Opin. Rheumatol. 2021, 33, 155–162. [Google Scholar] [CrossRef] [PubMed]
- Sacchi, M.C.; Tamiazzo, S.; Stobbione, P.; Agatea, L.; De Gaspari, P.; Stecca, A.; Lauritano, E.C.; Roveta, A.; Tozzoli, R.; Guaschino, R.; et al. SARS-CoV-2 infection as a trigger of autoimmune response. Clin. Transl. Sci. 2021, 14, 898–907. [Google Scholar] [CrossRef]
- Gupta, M.; Weaver, D.F. COVID-19 as a Trigger of Brain Autoimmunity. ACS Chem. Neurosci. 2021, 12, 2558–2561. [Google Scholar] [CrossRef]
- Jacobs, J.J.L. Persistent SARS-2 infections contribute to long COVID-19. Med. Hypotheses 2021, 149, 110538. [Google Scholar] [CrossRef] [PubMed]
- Zuo, Y.; Yalavarthi, S.; Navaz, S.A.; Hoy, C.K.; Harbaugh, A.; Gockman, K.; Zuo, M.; Madison, J.A.; Kanthi, Y.; Knight, J.S. Autoantibodies stabilize neutrophil extracellular traps in COVID-19. JCI Insight 2021, 6, 150111. [Google Scholar] [CrossRef] [PubMed]
- Holm, S.; Kared, H.; Michelsen, A.E.; Kong, X.Y.; Dahl, T.B.; Schultz, N.H.; Nyman, T.A.; Fladeby, C.; Seljeflot, I.; Ueland, T.; et al. Immune complexes, innate immunity, and NETosis in ChAdOx1 vaccine-induced thrombocytopenia. Eur. Heart J. 2021, 18, ehab506. [Google Scholar] [CrossRef] [PubMed]
- Zheng, Y.; Qin, C.; Zhang, X.; Zhu, Y.; Li, A.; Wang, M.; Tang, Y.; Kreiswirth, B.N.; Chen, L.; Zhang, H.; et al. The tst gene associated Staphylococcus aureus pathogenicity island facilitates its pathogenesis by promoting the secretion of inflammatory cytokines and inducing immune suppression. Microb. Pathog. 2020, 138, 103797. [Google Scholar] [CrossRef]
- Pakbaz, Z.; Sahraian, M.A.; Noorbakhsh, F.; Salami, S.A.; Pourmand, M.R. Staphylococcal enterotoxin B increased severity of experimental model of multiple sclerosis. Microb. Pathog. 2020, 142, 104064. [Google Scholar] [CrossRef]
- Gao, S.-X.; Sun, J.; Li, H.-H.; Chen, J.; Kashif, M.R.; Zhou, P.; Wei, L.; Zheng, Q.W.; Wu, L.G.; Guan, J.C. Prenatal exposure of staphylococcal enterotoxin B attenuates the development and function of blood regulatory T cells to repeated staphylococcal enterotoxin B exposure in adult offspring rats. J. Med. Microbiol. 2020, 69, 591–599. [Google Scholar] [CrossRef] [PubMed]
- Glass, R.; Norton, S.; Fox, N.; Kusnecov, A.W. Maternal immune activation with staphylococcal enterotoxin A produces unique behavioral changes in C57BL/6 mouse offspring. Brain Behav. Immun. 2019, 75, 12–25. [Google Scholar] [CrossRef]
- Karagöz, I.K.; Munk, M.R.; Kaya, M.; Rückert, R.; Yıldırım, M.; Karabaş, L. Using bioinformatic protein sequence similarity to investigate if SARS CoV-2 infection could cause an ocular autoimmune inflammatory reactions? Exp. Eye Res. 2021, 203, 108433. [Google Scholar] [CrossRef]
- Adiguzel, Y. Molecular mimicry between SARS-CoV-2 and human proteins. Autoimmun. Rev. 2021, 20, 102791. [Google Scholar] [CrossRef]
- Kanduc, D.; Shoenfeld, Y. Molecular mimicry between SARS-CoV-2 spike glycoprotein and mammalian proteomes: Implications for the vaccine. Immunol. Res. 2020, 68, 310–313. [Google Scholar] [CrossRef] [PubMed]
- Martínez, Y.A.; Guo, X.; Portales-Pérez, D.P.; Rivera, G.; Castañeda-Delgado, J.E.; García-Pérez, C.A.; Enciso-Moreno, J.A.; Lara-Ramírez, E.E. The analysis on the human protein domain targets and host-like interacting motifs for the MERS-CoV and SARS-CoV/CoV-2 infers the molecular mimicry of coronavirus. PLoS ONE 2021, 16, e0246901. [Google Scholar] [CrossRef]
- Morsy, S.; Morsy, A. Epitope mimicry analysis of SARS-COV-2 surface proteins and human lung proteins. J. Mol. Graph Model. 2021, 105, 107836. [Google Scholar] [CrossRef]
- Dotan, A.; Kanduc, D.; Muller, S.; Makatsariya, A.; Shoenfeld, Y. Molecular mimicry between SARS-CoV-2 and the female reproductive system. Am. J. Reprod. Immunol. 2021. [Google Scholar] [CrossRef]
- Vojdani, A.; Kharrazian, D. Potential antigenic cross-reactivity between SARS-CoV-2 and human tissue with a possible link to an increase in autoimmune diseases. Clin. Immunol. 2020, 217, 108480. [Google Scholar] [CrossRef]
- Bozkurt, B.; Kamat, I.; Hotez, P.J. Myocarditis With COVID-19 mRNA Vaccines. Circulation 2021, 144, 471–484. [Google Scholar] [CrossRef]
- Huynh, A.; Kelton, J.G.; Arnold, D.M.; Daka, M.; Nazy, I. Antibody epitopes in vaccine-induced immune thrombotic thrombocytopaenia. Nature 2021, 596, 565–569. [Google Scholar] [CrossRef]
- Greinacher, A.; Selleng, K.; Mayerle, J.; Palankar, R.; Wesche, J.; Reiche, S.; Aebischer, A.; Warkentin, T.E.; Muenchhoff, M.; Hellmuth, J.C.; et al. Anti-Platelet Factor 4 Antibodies Causing VITT do not Cross-React with SARS-CoV-2 Spike Protein. Blood 2021, 138, 1269–1277. [Google Scholar] [CrossRef]
- Liu, L.; Sun, D. Pregnancy outcomes in patients with primary antiphospholipid syndrome: A systematic review and meta-analysis. Medicine 2019, 98, e15733. [Google Scholar] [CrossRef]
- Talotta, R.; Robertson, E.S. Antiphospholipid antibodies and risk of post-COVID-19 vaccination thrombophilia: The straw that breaks the camel’s back? Cytokine Growth Factor Rev. 2021, 60, 52–60. [Google Scholar] [CrossRef] [PubMed]
- Harel, M.; Aron-Maor, A.; Sherer, Y.; Blank, M.; Shoenfeld, Y. The infectious etiology of the antiphospholipid syndrome: Links between infection and autoimmunity. Immunobiology 2005, 210, 743–747. [Google Scholar] [CrossRef]
- Zussman, R.; Xu, L.Y.; Damani, T.; Groom, K.M.; Chen, Q.; Seers, B.; Viall, C.A.; Chamley, L.W.; Hickey, A. Antiphospholipid antibodies can specifically target placental mitochondria and induce ROS production. J. Autoimmun. 2020, 111, 102437. [Google Scholar] [CrossRef] [PubMed]
- Martirosyan, A.; Aminov, R.; Manukyan, G. Environmental Triggers of Autoreactive Responses: Induction of Antiphospholipid Antibody Formation. Front. Immunol. 2019, 10, 1609. [Google Scholar] [CrossRef] [PubMed]
- Cimolai, N. Untangling the Intricacies of Infection, Thrombosis, Vaccination, and Antiphospholipid Antibodies for COVID-19. SN Compr. Clin. Med. 2021, 22, 1–16. [Google Scholar] [CrossRef] [PubMed]
- Arthur, J.M.; Forrest, J.C.; Boehme, K.W.; Kennedy, J.L.; Owens, S.; Herzog, C.; Liu, J.; Harville, T.O. Development of ACE2 autoantibodies after SARS-CoV-2 infection. PLoS ONE 2021, 16, e0257016. [Google Scholar] [CrossRef]
- Wen, J.; Cheng, Y.; Ling, R.; Dai, Y.; Huang, B.; Huang, W.; Zhang, S.; Jiang, Y. Antibody-dependent enhancement of coronavirus. Int. J. Infect. Dis. 2020, 100, 483–489. [Google Scholar] [CrossRef]
- Ricke, D.O. Two Different Antibody-Dependent Enhancement (ADE) Risks for SARS-CoV-2 Antibodies. Front. Immunol. 2021, 12, 640093. [Google Scholar] [CrossRef] [PubMed]
- Rothan, H.A.; Byrareddy, S.N. The potential threat of multisystem inflammatory syndrome in children during the COVID-19 pandemic. Pediatr. Allergy Immunol. 2021, 32, 17–22. [Google Scholar] [CrossRef]
- Liu, Y.; Soh, W.T.; Kishikawa, J.-I.; Hirose, M.; Nakayama, E.E.; Li, S.; Sasai, M.; Suzuki, T.; Tada, A.; Arakawa, A. An infectivity-enhancing site on the SARS-CoV-2 spike protein targeted by antibodies. Cell 2021, 184, 3452–3466.e18. [Google Scholar] [CrossRef] [PubMed]
- Yahi, N.; Chahinian, H.; Fantini, J. Infection-enhancing anti-SARS-CoV-2 antibodies recognize both the original Wuhan/D614G strain and Delta variants. A potential risk for mass vaccination? J. Infect. 2021. [Google Scholar] [CrossRef] [PubMed]
- Martínez Gómez, J.M.; Ong, L.C.; Lam, J.H.; Binte Aman, S.A.; Libau, E.A.; Lee, P.X.; St. John, A.L.; Alonso, S. Maternal Antibody-Mediated Disease Enhancement in Type I Interferon-Deficient Mice Leads to Lethal Disease Associated with Liver Damage. PLoS Negl. Trop Dis. 2016, 10, e0004536. [Google Scholar] [CrossRef]
- Lee, P.X.; Ong, L.C.; Libau, E.A.; Alonso, S. Relative Contribution of Dengue IgG Antibodies Acquired during Gestation or Breastfeeding in Mediating Dengue Disease Enhancement and Protection in Type I Interferon Receptor-Deficient Mice. PLoS Negl. Trop Dis. 2016, 10, e0004805. [Google Scholar] [CrossRef]
- Pace, R.M.; Williams, J.E.; Järvinen, K.M.; Belfort, M.B.; Pace, C.D.W.; Lackey, K.A.; Gogel, A.C.; Nguyen-Contant, P.; Kanagaiah, P.; Fitzgerald, T.; et al. Characterization of SARS-CoV-2 RNA, Antibodies, and Neutralizing Capacity in Milk Produced by Women with COVID-19. mBio 2021, 12, e03192-20. [Google Scholar] [CrossRef]
- Baird, J.K.; Jensen, S.M.; Urba, W.J.; Fox, B.A.; Baird, J.R. SARS-CoV-2 Antibodies Detected in Mother’s Milk Post-Vaccination. J. Hum. Lact. 2021. [Google Scholar] [CrossRef]
- Hsu, C.-N.; Tain, Y.-L. Animal Models for DOHaD Research: Focus on Hypertension of Developmental Origins. Biomedicines 2021, 9, 623. [Google Scholar] [CrossRef]
- Beena, V. Applications of livestock in biomedical research. J. Dairy Vet. Anim. Res. 2021, 8, 107–109. [Google Scholar]
- Gatford, K.L.; Wooldridge, A.L.; Kind, K.L.; Bischof, R.; Clifton, V.L. Pre-birth origins of allergy and asthma. J. Reprod. Immunol. 2017, 123, 88–93. [Google Scholar] [CrossRef]
- Wooldridge, A.L.; Clifton, V.L.; Moss, T.J.M.; Lu, H.; Jamali, M.; Agostino, S.; Muhlhausler, B.S.; Morrison, J.L.; De Matteo, R.; Wallace, M.J.; et al. Maternal allergic asthma during pregnancy alters fetal lung and immune development in sheep: Potential mechanisms for programming asthma and allergy. J. Physiol. 2019, 597, 4251–4262. [Google Scholar] [CrossRef] [PubMed]
- Alvites, R.D.; Branquinho, M.V.; Sousa, A.C.; Lopes, B.; Sousa, P.; Mendonça, C.; Atayde, L.M.; Maurício, A.C. Small Ruminants and Its Use in Regenerative Medicine: Recent Works and Future Perspectives. Biology 2021, 10, 249. [Google Scholar] [CrossRef] [PubMed]
- Peden, A.H.; Suleiman, S.; Barria, M.A. Understanding Intra-Species and Inter-Species Prion Conversion and Zoonotic Potential Using Protein Misfolding Cyclic Amplification. Front. Aging Neurosci. 2021, 13, 716452. [Google Scholar] [CrossRef] [PubMed]
- Salamat, M.K.F.; Gossner, A.; Bradford, B.; Hunter, N.; Hopkins, J.; Houston, F. Scrapie infection and endogenous retroviral expression in sheep lymphoid tissues. Vet Immunol. Immunopathol. 2021, 233, 110194. [Google Scholar] [CrossRef]
- Entrican, G.; Wattegedera, S.R.; Griffiths, D.J. Exploiting ovine immunology to improve the relevance of biomedical models. Mol. Immunol. 2015, 66, 68–77. [Google Scholar] [CrossRef]
- Scheerlinck, J.-P.Y.; Snibson, K.J.; Bowles, V.M.; Sutton, P. Biomedical applications of sheep models: From asthma to vaccines. Trends Biotechnol. 2008, 26, 259–266. [Google Scholar] [CrossRef]
- Luján, L.; Pérez, M.; Salazar, E.; Álvarez, N.; Gimeno, M.; Pinczowski, P.; Irusta, S.; Santamaría, J.; Insausti, N.; Cortés, Y.; et al. Autoimmune/autoinflammatory syndrome induced by adjuvants (ASIA syndrome) in commercial sheep. Immunol. Res. 2013, 56, 317–324. [Google Scholar] [CrossRef] [PubMed]
- Goldstein, J.A.; Norris, S.A.; Aronoff, D.M. DOHaD at the intersection of maternal immune activation and maternal metabolic stress: A scoping review. J. Dev. Orig. Health Dis. 2017, 8, 273–283. [Google Scholar] [CrossRef] [PubMed]
- Fisher-Heffernan, R.E.; Or’Rashid, M.M.; AlZahal, O.; Quinton, M.; Boermans, H.J.; McBride, B.W.; Regnault, T.R.; Karrow, N.A. Fishmeal supplementation during ovine pregnancy and lactation protects against maternal stress-induced programming of the offspring immune system. BMC Vet. Res. 2015, 11, 266. [Google Scholar] [CrossRef] [PubMed]
- Fisher, R.E.; Or’Rashid, M.; Quinton, M.; AlZahal, O.; Boermans, H.J.; McBride, B.W.; Karrow, N.A. Maternal supplementation with fishmeal protects against late gestation endotoxin-induced fetal programming of the ovine hypothalamic-pituitary-adrenal axis. J. Dev. Orig. Health Dis. 2014, 5, 206–213. [Google Scholar] [CrossRef]
- Karrow, N.A.; Lee, A.V.; Quinton, M.; McBride, B.W.; Fisher-Heffernan, R.E. Ovine hippocampal mRNA expression in offspring from dams supplemented with fishmeal and stress challenged in late pregnancy with endotoxin. Anim. Nutr. 2017, 3, 39–45. [Google Scholar] [CrossRef] [PubMed]
- Naylor, D.; Sharma, A.; Li, Z.; Monteith, G.; Sullivan, T.; Canovas, A.; Mallard, B.A.; Baes, C.; Karrow, N.A. Short communication: Characterizing ovine serum stress biomarkers during endotoxemia. J. Dairy Sci. 2020, 103, 5501–5508. [Google Scholar] [CrossRef]
- Sharma, A.; Shandilya, U.K.; Sullivan, T.; Naylor, D.; Canovas, A.; Mallard, B.A.; Karrow, N.A. Identification of Ovine Serum miRNAs Following Bacterial Lipopolysaccharide Challenge. Int. J. Mol. Sci. 2020, 21, E7920. [Google Scholar] [CrossRef]
- Naylor, D.; Sharma, A.; Li, Z.; Monteith, G.; Mallard, B.A.; Bergeron, R.; Baes, C.; Karrow, N.A. Endotoxin-induced cytokine, chemokine and white blood cell profiles of variable stress-responding sheep. Stress 2021, 28, 1–10. [Google Scholar] [CrossRef]
- Pant, S.D.; You, Q.; Schenkel, L.C.; Voort, G.V.; Schenkel, F.S.; Wilton, J.; Cain, L.; Karrow, N.A. A genome-wide association study to identify chromosomal regions influencing ovine cortisol response. Livest. Sci. 2016, 187, 40–47. [Google Scholar] [CrossRef]
- Forestieri, S.; Pintus, R.; Marcialis, M.A.; Pintus, M.C.; Fanos, V. COVID-19 and developmental origins of health and disease. Early Hum. Dev. 2021, 155, 105322. [Google Scholar] [CrossRef]
- Hernández, A.F.; Calina, D.; Poulas, K.; Docea, A.O.; Tsatsakis, A.M. Safety of COVID-19 vaccines administered in the EU: Should we be concerned? Toxicol. Rep. 2021, 8, 871–879. [Google Scholar] [CrossRef] [PubMed]
- Fowlkes, A.; Gaglani, M.; Groover, K.; Thiese, M.S.; Tyner, H.; Ellingson, K.; Cohorts, H.R. Effectiveness of COVID-19 Vaccines in Preventing SARS-CoV-2 Infection Among Frontline Workers Before and During B.1.617.2 (Delta) Variant Predominance—Eight U.S. Locations, December 2020-August 2021. MMWR Morb Mortal Wkly Rep. 2021, 70, 1167–1169. [Google Scholar] [CrossRef]
- Iacobucci, G. Covid-19: Protection from two doses of vaccine wanes within six months, data suggest. BMJ 2021, 374, n2113. [Google Scholar] [CrossRef] [PubMed]
- Bianco-Miotto, T.; Craig, J.M.; Gasser, Y.P.; van Dijk, S.J.; Ozanne, S.E. Epigenetics and DOHaD: From basics to birth and beyond. J. Dev. Orig. Health Dis. 2017, 8, 513–519. [Google Scholar] [CrossRef]
- Fang, S.; Zheng, R.; Lei, C.; Wang, J.; Zheng, R.; Li, M. Key residues influencing binding affinities of 2019-nCoV with ACE2 in different species. Brief Bioinform. 2021, 22, 963–975. [Google Scholar] [CrossRef] [PubMed]
- Di Teodoro, G.; Valleriani, F.; Puglia, I.; Monaco, F.; Di Pancrazio, C.; Luciani, M.; Krasteva, I.; Petrini, A.; Marcacci, M.; D’Alterio, N.; et al. SARS-CoV-2 replicates in respiratory ex vivo organ cultures of domestic ruminant species. Vet Microbiol. 2021, 252, 108933. [Google Scholar] [CrossRef]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Karrow, N.A.; Shandilya, U.K.; Pelech, S.; Wagter-Lesperance, L.; McLeod, D.; Bridle, B.; Mallard, B.A. Maternal COVID-19 Vaccination and Its Potential Impact on Fetal and Neonatal Development. Vaccines 2021, 9, 1351. https://doi.org/10.3390/vaccines9111351
Karrow NA, Shandilya UK, Pelech S, Wagter-Lesperance L, McLeod D, Bridle B, Mallard BA. Maternal COVID-19 Vaccination and Its Potential Impact on Fetal and Neonatal Development. Vaccines. 2021; 9(11):1351. https://doi.org/10.3390/vaccines9111351
Chicago/Turabian StyleKarrow, Niel A., Umesh K. Shandilya, Steven Pelech, Lauraine Wagter-Lesperance, Deanna McLeod, Byram Bridle, and Bonnie A. Mallard. 2021. "Maternal COVID-19 Vaccination and Its Potential Impact on Fetal and Neonatal Development" Vaccines 9, no. 11: 1351. https://doi.org/10.3390/vaccines9111351
APA StyleKarrow, N. A., Shandilya, U. K., Pelech, S., Wagter-Lesperance, L., McLeod, D., Bridle, B., & Mallard, B. A. (2021). Maternal COVID-19 Vaccination and Its Potential Impact on Fetal and Neonatal Development. Vaccines, 9(11), 1351. https://doi.org/10.3390/vaccines9111351





