A Comprehensive Review of the Immunological Response against Foot-and-Mouth Disease Virus Infection and Its Evasion Mechanisms
Abstract
1. Introduction
2. Foot-and-Mouth Disease Virus Pathogenesis
- Entry to a susceptible host;
- Replication for increasing viral load;
- Dissemination from the entry site to the tissues and/or target organs, developing the infection, and damage to cells and organs;
- Elimination, contamination, and dissemination to the environment;
- Persistence in the environment; and
- Transmission to new hosts and beginning a new cycle [5].
2.1. Foot-and-Mouth Disease Virus Entry to a Host
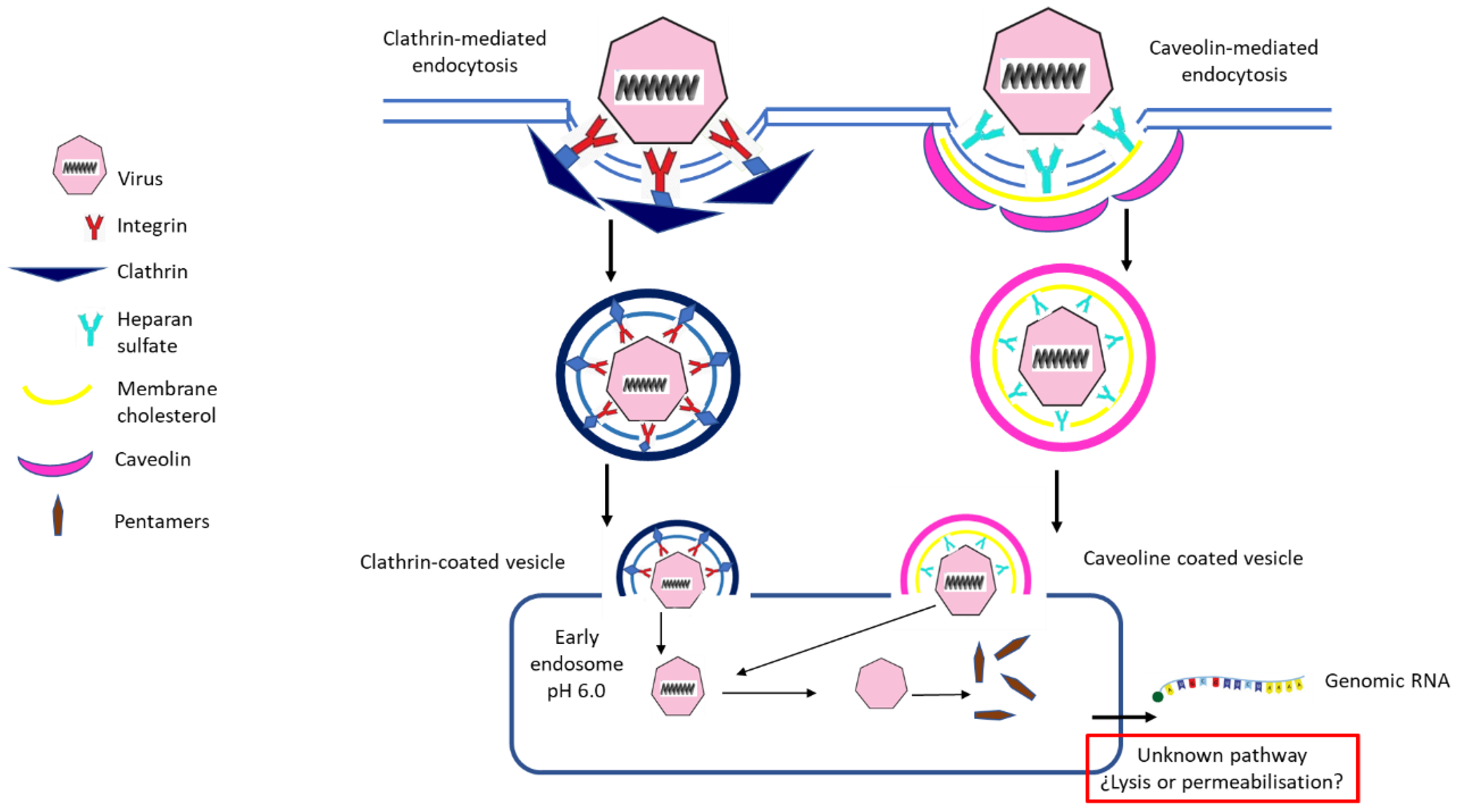
2.2. Foot-and-Mouth Disease Virus Entry to Cells
2.3. Foot-and-Mouth Disease Virus Replication, Dissemination, and Effect on Target Organs
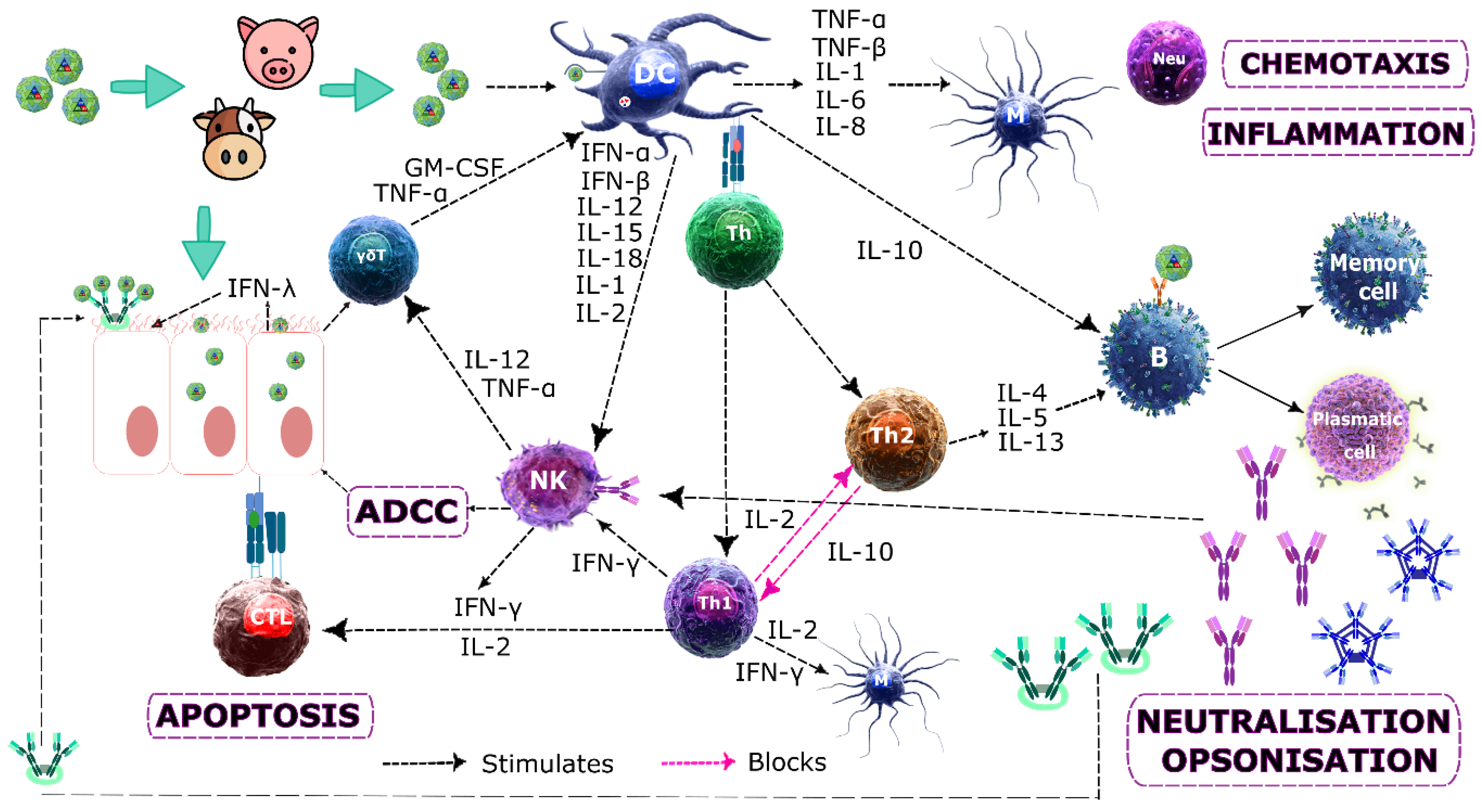
2.4. Immune Response
2.4.1. Innate Response
2.4.2. Acquired Response
Cellular Response
Humoral Response
2.5. Evasion Mechanisms
3. Conclusions and Perspectives
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Morante Yamir, L.; Guibarra Escobar, V.H. Barreras anatómicas del sistema inmunitario. Rev. Actual. Clín. 2011, 13, 634–638. [Google Scholar]
- Campos-Granados, C. El sistema inmune en los mamíferos: Las defensas del cuerpo. Nutr. Anim. Trop. 2014, 8, 80–93. [Google Scholar]
- Doménech, A.; Gibello, A.; Collado, V.M.; Porras, R.; Blanco, M. The innate immune system II: First response against infection. Rev. Complut. Cienc. Vet. 2008, 2, 17–30. [Google Scholar]
- Klossek, J.M.; Fontanel, D.J.P. Fisiología de la mucosa respiratoria nasal y trastornos funcionales. EMC Otorrinolaringol. 2000, 38, 1–11. [Google Scholar] [CrossRef]
- Finlay, B.B.; Mcfadden, G. Review Anti-Immunology: Evasion of the Host Immune System by Bacterial and Viral Pathogens. Cell 2006, 124, 767–782. [Google Scholar] [CrossRef]
- Golde, W.T.; Nfon, C.K.; Toka, F.N. Immune evasion during foot-and-mouth disease virus infection of swine. Immunol. Rev. 2008, 225, 85–95. [Google Scholar] [CrossRef]
- OIE. Foot and Mouth Disease Virus. 2013. Available online: https://www.oie.int/fileadmin/Home/eng/Animal_Health_in_the_World/docs/pdf/Disease_cards/FOOT_AND_MOUTH_DISEASE.pdf (accessed on 25 June 2020).
- Brito, B.P.; Jori, F.; Dwarka, R.; Maree, F.F.; Heath, L.; Perez, A.M. Transmission of foot-and-mouth disease SAT2 viruses at the wildlife-livestock interface of two major transfrontier conservation areas in Southern Africa. Front. Microbiol. 2016, 7, 1–10. [Google Scholar] [CrossRef]
- Sebhatu, T.T. Foot-and-Mouth Disease Sero-Surveillance in Africa and Vaccine Matching; Utrecht University: Utrecht, The Netherlands, 2014. [Google Scholar]
- Kitching, P.; Hammond, J.; Jeggo, M.; Charleston, B.; Paton, D.; Rodriguez, L.; Heckert, R. Global FMD control-Is it an option? Vaccine 2007, 25, 5660–5664. [Google Scholar] [CrossRef]
- Kahrs, R.F. Fiebre Aftosa. In Enfermedades Víricas del Ganado Vacuno, 1st ed.; Editorial Acribia, S.A.: Zaragoza, España, 1985; pp. 319–328. [Google Scholar]
- Briones, V. Algunas características del virus de la fiebre aftosa. Mundo Vet. 2001, 133, 72–74. [Google Scholar]
- Reeve, R.; Borley, D.W.; Maree, F.F.; Upadhyaya, S.; Lukhwareni, A.; Esterhuysen, J.J.; Harvey, W.T.; Blignaut, B.; Fry, E.E.; Parida, S.; et al. Tracking the antigenic evolution of foot-and-mouth disease virus. PLoS ONE 2016, 11, e159360. [Google Scholar] [CrossRef]
- Kamel, M.; El, A.; Hugo, S.; Vazquez, C. Foot-and-mouth disease vaccines: Recent updates and future perspectives. Arch. Virol. 2019, 164, 1501–1513. [Google Scholar] [CrossRef] [PubMed]
- Ferrari, G.; Paton, D.; Duffy, S.; Bartels, C.; Knight-jones, T. Foot and mouth disease vaccination monitoring Guidelines. Chapter 2 Vaccine Programme, Delivery, Schedule and Coverage. In Foot and Mouth Disease Vaccination and Post-Vaccination Monitoring Guidelines; Metwally Samia, M.S., Ed.; FAO: Rome, Italy, 2 December 2016; pp. 15–20. [Google Scholar]
- Domingo, E. RNA Virus evolution and the control of viral disease. Prog. Drug Res. 1989, 33, 93–133. [Google Scholar] [PubMed]
- Pulido, M.R. Estructura y Función de la Región 3’ no Codificante del Virus de la Fiebre Aftosa. Aplicación a Nuevas Estrategias Vacunales Basadas en RNA. Ph.D. Thesis, Universidad Autónoma de Madrid, Madrid, Spain, 2009. [Google Scholar]
- Barteling, S.J.; Woortmeyer, R. Multiple variants in foot-and-mouse disease virus (FMDV) populations: The Achilles heel for peptide and rec. DNA vaccines? Dev. Biol. Stand. 1987, 66, 511–521. [Google Scholar] [PubMed]
- Grubman, M.J.; Moraes, M.P.; Diaz-San Segundo, F.; Pena, L.; De Los Santos, T. Evading the host immune response: How foot-and-mouth disease virus has become an effective pathogen. FEMS Immunol. Med. Microbiol. 2008, 53, 8–17. [Google Scholar] [CrossRef] [PubMed]
- Barlow, D.F. The Aerosol Stability of a Strain of Foot-and-Mouth Disease Virus and the Effects on Stability of Precipitation with Ammonium Sulphate, Methanol or Polyethylene Glycol. J. Gen. Virol. 2016, 15, 17–24. [Google Scholar] [CrossRef]
- Madero, M.M.J. El Sistema Inmunológico Cutáneo 2007. Available online: http://www.medicosecuador.com/librodermatologia/capitulos/capitulo_3.htm (accessed on 14 May 2020).
- Ramos, A.P.; Desgarennes, C.P. La función inmunológica de la piel. Dermatol. Rev. Mex. 2008, 52, 211–224. [Google Scholar]
- Alexandersen, S.; Zhang, Z.; Donaldson, A.I.; Garland, A.J.M. The Pathogenesis and Diagnosis of Foot-and-Mouth Disease. J. Comp. Pathol. 2003, 129, 1–36. [Google Scholar] [CrossRef]
- Alexandersen, S.; Brotherhood, I.; Donaldson, A.I. Natural aerosol transmission of foot-and-mouth disease virus to pigs: Minimal infectious dose for strain O 1 Lausanne. Epidemiol. Infect. 2002, 128, 301–312. [Google Scholar] [CrossRef]
- Pharo, H. Foot-and-mouth disease: An assessment of the risks facing New Zealand Although New Zealand has never had a case of foot-and-mouth The emergence of the type-O PanAsia strain. N. Z. Vet. J. 2002, 50, 46–55. [Google Scholar] [CrossRef]
- Carroll, K.; Butel, J.; Morse, S.; Mietzer, T. Medical Microbiology, 27th ed.; McGraw-Hill Education: New York, NY, USA, 2015. [Google Scholar]
- Segundo, F.D.; Medina, G.N.; Grubman, M.J.; Santos, T.D.L. Animal Health: Foot-and-Mouth Disease; Elsevier Ltd.: Amsterdam, The Netherlands, 2003; Volume 1, pp. 327–345. [Google Scholar] [CrossRef]
- Ruiz-Saenz, J.; Goez, Y.; Tabares, W. Cellular Receptors for Foot and Mouth Disease Virus. Intervirology 2009, 52, 201–212. [Google Scholar] [CrossRef]
- Wang, G.; Wang, Y.; Shang, Y.; Zhang, Z.; Liu, X. How foot-and-mouth disease virus receptor mediates foot-and-mouth disease virus infection. Virol. J. 2015, 12, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Ruiz-Sáenz, J.; Jaime, J.; Vera, V. Foot and Mouth Disease virus: An Approach to the state of the art. Rev. Colomb. Cienc. Pecu. 2009, 22, 209–220. [Google Scholar]
- O’Donnell, V.; Pacheco, J.M.; Gregg, D.; Baxt, B. Analysis of Foot-and-Mouth Disease Virus Integrin Receptor Expression in Tissues from Naïve and Infected Cattle. J. Comp. Pathol. 2009, 141, 98–112. [Google Scholar] [CrossRef] [PubMed]
- Jackson, T.; Ellard, F.M.; A Ghazaleh, R.; Brookes, S.M.; E Blakemore, W.; Corteyn, A.H.; I Stuart, D.; Newman, J.W.; King, A.M. Efficient Infection of Cells in Culture by Type O Foot-and-Mouth Disease Virus Requires Binding to Cell Surface Heparan Sulfate. J. Virol. 1996, 70, 5282–5287. [Google Scholar] [CrossRef]
- Fry, E.E.; Lea, S.M.; Jackson, T.; Newman, J.W.; Ellard, F.M.; Blakemore, W.E.; Abu-Ghazaleh, R.; Samuel, A.; King, A.M.; Stuart, D.I. The structure and function of a foot-and-mouth disease virus–oligosaccharide receptor complex. EMBO J. 1999, 18, 543–554. [Google Scholar] [CrossRef]
- Baranowski, E.; Ruiz-jarabo, C.M.; Sevilla, N.; Andreu, D.; Beck, E.; Domingo, E. Cell Recognition by Foot-and-Mouth Disease Virus That Lacks the RGD Integrin-Binding Motif: Flexibility in Aphthovirus Receptor Usage. J. Virol. 2000, 74, 1641–1647. [Google Scholar] [CrossRef]
- Baxt, B.; Bachrach, H.L. Early Interactions of Foot-and-Mouth Disease Virus with Cultured Cells. Virology 1980, 104, 42–55. [Google Scholar] [CrossRef]
- Han, S.-C.; Guo, H.-C.; Sun, S. Three-dimensional structure of foot-and-mouth disease virus and its biological functions. Arch. Virol. 2014, 160, 1–16. [Google Scholar] [CrossRef]
- Bayry, J.; Tough, D.F. Interaction of foot-and-mouth disease virus with dendritic cells. Trends Microbiol. 2006, 14, 346–347. [Google Scholar] [CrossRef]
- Baxt, B.; Mason, P.W. FMDV undergoes resticted replicaton in macrophage cells. Virology 1995, 207, 503–509. [Google Scholar] [CrossRef]
- Gutiérrez, M.; López, S. Mechanisms of virus entry: A way to learn about the host cell. Rev. Espec. Cienc. Quím. 2010, 13, 26–34. [Google Scholar]
- Agudo Torres, R. Caracterización de las Proteínas del Virus de la Fiebre Aftosa Implicadas en Respuesta a Mutagénesis Letal por Análogos de Nucleótido. Ph.D. Thesis, Universidad Autónoma de Madrid, Madrid, Spain, 2009. [Google Scholar]
- Smitsaart, E.N.; Saiz, J.C.; Yedloutschnig, R.J.; Morgan, D.O. Detection of foot-and-mouth disease virus by competitive ELISA using a monoclonal antibody specific for the 12S protein subunit from six of the seven serotypes. Vet. Immunol. Immunopathol. 1990, 26, 251–265. [Google Scholar] [CrossRef]
- Gao, Y.; Sun, S.-Q.; Guo, H.-C. Biological function of Foot-and-mouth disease virus non-structural proteins and non-coding elements. Virol. J. 2016, 13, 107. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Zhu, Z.; Zhang, M.; Zheng, H. Multifunctional roles of leader protein of foot - and - mouth disease viruses in suppressing host antiviral responses. Vet. Res. 2015, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Taboga, O.A. Expresión de Antígenos Derivados del Virus de la Fiebre Aftosa en Diferentes Sistemas Eucarióticos. Ph.D. Thesis, Universidad de Buenos Aires, Buenos Aires, Argentina, 2000. [Google Scholar]
- Gullberg, M.; Muszynski, B.; Organtini, L.J.; Ashley, R.E.; Hafenstein, S.L.; Belsham, G.J.; Polacek, C. Assembly and characterization of foot-and-mouth disease virus empty capsid particles expressed within mammalian cells. J. Gen. Virol. 2017, 1769–1779. [Google Scholar] [CrossRef]
- Xie, Y.; Gao, P.; Li, Z. A Recombinant Adenovirus Expressing P12A and 3C Protein of the Type O Foot-and-Mouth Disease Virus Stimulates Systemic and Mucosal Immune Responses in Mice. BioMed Res. Int. 2016, 2016, 7849203. [Google Scholar] [CrossRef]
- Segundo, F.D.; Medina, G.N.; Stenfeldt, C.; Arzt, J.; Santos, T.D.L. Foot-and-mouth disease vaccines. Vet. Microbiol. 2016. [Google Scholar] [CrossRef]
- Patch, J.R.; Dar, P.A.; Waters, R.; Toka, F.N.; Barrera, J.; Schutta, C.; Kondabattula, G.; Golde, W.T. Infection with foot-and-mouth disease virus (FMDV) induces a natural killer (NK) cell response in cattle that is lacking following vaccination. Comp. Immunol. Microbiol. Infect. Dis. 2014, 37, 249–257. [Google Scholar] [CrossRef]
- Arzt, J.; Juleff, N.; Zhang, Z.; Rodriguez, L.L. The Pathogenesis of Foot-and-Mouth Disease I: Viral Pathways in Cattle. Transbound. Emerg. Dis. 2011, 58, 291–304. [Google Scholar] [CrossRef]
- Arzt, J.; Pacheco, J.M.; Smoliga, G.; Tucker, M.T.; Bishop, E.; Pauszek, S.; Hartwig, E.; Santos, T.D.L.; Rodriguez, L.L. Foot-and-mouth disease virus virulence in cattle is co-determined by viral replication dynamics and route of infection. Virology 2014, 452–453, 12–22. [Google Scholar] [CrossRef]
- Robinson, L.; Charleston, B.; Rodriguez, L.L.; Gay, C.G.; Sumption, K.J. Global Foot-and-Mouth Disease Research Update and Gap Analysis: 7–Pathogenesis and Molecular Biology. Transbound. Emerg. Dis. 2016, 63, 63–71. [Google Scholar] [CrossRef] [PubMed]
- Martínez, R.-C.; Mayayo, A.I.; Urraca Pinilla, T.M. Fisiología de la Nariz y de los Senos Paranasales. Mecanismos de la Olfacción. Libro Virtual de Formación en Otorrinolaringología. Edited by Sociedad Española de Otorrinolaringología y Patología Cervico-Facial SEORL-PCF. 2014. Available online: https://seorl.net/PDF/Nariz%20y%20senos%20paranasales/042%20-%20FISIOLOG%C3%8DA%20DE%20LA%20NARIZ%20Y%20DE%20LOS%20SENOS%20PARANASALES.%20MECANISMOS%20DE%20LA%20OLFACCI%C3%93N.pdf (accessed on 14 May 2020).
- Moran, P. Mecanismos de infección viral y diseminación de los virus. Igarss 2011, 3, 1–12. [Google Scholar] [CrossRef]
- Dunham, S.; MacLachlan, N.J.; Dubovi, E.J. Fenner’s Veterinary Virology; Academic Press: New York, NY, USA, 2011. [Google Scholar]
- García-Bellmunt, L.; Sibila, O. Mecanismos de defensa pulmonar. Med. Respir. 2013, 6, 15–24. [Google Scholar]
- Kawai, T.; Akira, S. TLR signaling. Semin. Immunol. 2007, 19, 24–32. [Google Scholar] [CrossRef]
- Kawai, T.; Akira, S. The role of pattern-recognition receptors in innate immunity: Update on toll-like receptors. Nat. Immunol. 2010, 11, 373–384. [Google Scholar] [CrossRef]
- Zhu, J.J.; Arzt, J.; Puckette, M.C.; Smoliga, G.R.; Pacheco, J.M.; Rodriguez, L.L. Mechanisms of Foot-and-Mouth Disease Virus Tropism Inferred from Differential Tissue Gene Expression. PLoS ONE 2013, 8, e64119. [Google Scholar] [CrossRef]
- Parker, D.; Prince, A. Innate Immunity in the Respiratory Epithelium. Am. J. Respir. Cell Mol. Biol. 2011, 45. [Google Scholar] [CrossRef]
- Nicod, L.P. Pulmonary Defence Mechanisms. Respiration 1999, 14, 2–11. [Google Scholar] [CrossRef]
- Schulz, O.; Diebold, S.S.; Chen, M.; Näslund, T.I.; Nolte, M.A.; Alexopoulou, L.; Azuma, Y.-T.; Flavell, R.A.; Liljeström, P.; E Sousa, C.R. Toll-like receptor 3 promotes cross-priming to virus-infected cells. Nature 2005, 433, 887–892. [Google Scholar] [CrossRef]
- Ranjan, R.; Biswal, J.K.; Sharma, A.K.; Misri, J.; Pattnaik, B. Profiling of bovine toll like receptors (TLRs) in foot and mouth disease vaccinated cattle. Indian J. Anim. Sci. 2016, 86, 367–371. [Google Scholar]
- Cheng, G.; Zhao, X.; Yan, W.; Wang, W.; Zuo, X.; Huang, K.; Liu, Y.; Chen, J.; Wang, J.; Cong, W.; et al. Alpha interferon is a powerful adjuvant for a recombinant protein vaccine against foot-and-mouth disease virus in swine, and an effective stimulus of in vivo immune response. Vaccine 2007, 25, 5199–5208. [Google Scholar] [CrossRef] [PubMed]
- Toka, F.N.; Golde, W.T. Cell mediated innate responses of cattle and swine are diverse during foot-and-mouth disease virus (FMDV) infection: A unique landscape of innate immunity. Immunol. Lett. 2013, 152, 135–143. [Google Scholar] [CrossRef] [PubMed]
- Banchereau, J.; Steinman, R.M. Dendritic cells and the control of immunity. Nature 2016, 245–252. [Google Scholar] [CrossRef] [PubMed]
- Reid, E.; Juleff, N.; Gubbins, S.; Prentice, H.; Seago, J.; Charleston, B. Bovine plasmacytoid dendritic cells are the major source of type I interferon in response to foot-and-mouth disease virus in vitro and in vivo. J. Virol. 2011, 85, 4297–4308. [Google Scholar] [CrossRef] [PubMed]
- Guzylack-Piriou, L.; Bergamin, F.; Gerber, M.; McCullough, K.C.; Summerfield, A. Plasmacytoid dendritic cell activation by foot-and-mouth disease virus requires immune complexes. Eur. J. Immunol. 2006, 36, 1674–1683. [Google Scholar] [CrossRef]
- Bautista, E.M.; Ferman, G.S.; Gregg, D.R.; Brum, M.C.S.; Grubman, M.J.; Golde, W.T. Constitutive expression of alpha interferon by skin dendritic cells confers resistance to infection by Foot-and-mouth disease virus. J. Virol. 2005, 79, 4838–4847. [Google Scholar] [CrossRef]
- Sei, J.J.; Waters, R.A.; Kenney, M.; Barlow, J.W.; Golde, W.T. Effect of Foot-and-Mouth Disease Virus Infection on the Frequency, Phenotype and Function of Circulating Dendritic Cells in Cattle. PLoS ONE 2016, 11, e0152192. [Google Scholar] [CrossRef]
- Lannes, N.; Python, S.; Summerfield, A. Interplay of foot-and-mouth disease virus, antibodies and plasmacytoid dendritic cells: Virus opsonization under non-neutralizing conditions results in enhanced interferon-alpha responses. Vet. Res. 2012, 43, 1. [Google Scholar] [CrossRef]
- Russo, R.C.; Garcia, C.C.; Teixeira, M.M.; Amaral, F.A. The CXCL8/IL-8 chemokine family and its receptors in inflammatory diseases. Expert Rev. Clin. Immunol. 2014, 10, 593–619. [Google Scholar] [CrossRef]
- Zlotnik, A.; Yoshie, O. Chemokines: A new classification system and their role in immunity. J. Cult. Herit. 2000, 1, 121–127. [Google Scholar] [CrossRef]
- Segundo, F.D.-S.; Weiss, M.; Pérez-Martín, E.; Koster, M.J.; Zhu, J.; Grubman, M.J.; Santos, T.D.L. Antiviral activity of bovine type III interferon against foot-and-mouth disease virus. Virology 2011, 413, 283–292. [Google Scholar] [CrossRef] [PubMed]
- Chinsangaram, J.; Moraes, M.P.; Koster, M.; Grubman, M.J. Novel Viral Disease Control Strategy: Adenovirus Expressing Alpha Interferon Rapidly Protects Swine from Foot-and-Mouth Disease. J. Virol. 2003, 77, 1621–1625. [Google Scholar] [CrossRef] [PubMed]
- Mordstein, M.; Neugebauer, E.; Ditt, V.; Jessen, B.; Rieger, T.; Falcone, V.; Sorgeloos, F.; Ehl, S.; Mayer, D.; Kochs, G.; et al. Lambda Interferon Renders Epithelial Cells of the Respiratory and Gastrointestinal Tracts Resistant to Viral Infections. J. Virol. 2010, 84, 5670–5677. [Google Scholar] [CrossRef] [PubMed]
- Perez-Martin, E.; Weiss, M.; Segundo, F.D.-S.; Pacheco, J.M.; Arzt, J.; Grubman, M.J.; Santos, T.D.L. Bovine Type III Interferon Significantly Delays and Reduces the Severity of Foot-and-Mouth Disease in Cattle. J. Virol. 2012, 86, 4477–4487. [Google Scholar] [CrossRef]
- Golde, W.T.; Santos, T.D.L.; Robinson, L.; Grubman, M.J.; Sevilla, N.; Summerfield, A.; Charleston, B. Evidence of Activation and Suppression during the Early Immune Response to Foot-and-Mouth Disease Virus. Transbound. Emerg. Dis. 2011, 58, 283–290. [Google Scholar] [CrossRef]
- Herzig, C.T.A.; Lefranc, M.; Baldwin, C.L. Annotation and classification of the bovine T cell receptor delta genes. BMC Genom. 2010, 11, 100. [Google Scholar] [CrossRef]
- Bautista, C. Importance of γδT lymphocytes in the bovine immune response. Vet. Méx. 2011, 42, 65–75. [Google Scholar]
- Abubakar, M.; Ahmed, A. Interplay of foot and mouth disease virus with cell-mediated and humoral immunity of host. Rev. Med. Virol. 2017, 1–8. [Google Scholar] [CrossRef]
- Toka, F.N.; Kenney, M.A.; Golde, W.T. Rapid and Transient Activation of γ δ T Cells to IFN- γ Production, NK Cell-Like Killing, and Antigen Processing during Acute Virus Infection. J. Immunol. 2019, 186, 4853–4861. [Google Scholar] [CrossRef]
- Vivier, E.; Tomasello, E.; Baratin, M.; Walzer, T.; Ugolini, S. Functions of natural killer cells. Nat. Immunol. 2008, 9, 503–510. [Google Scholar] [CrossRef]
- Sun, J.C.; Lanier, L.L. Cell development NK: Homeostasis and functions: Parallels with CD8+ T cells. Nat. Rev. Immunol. 2015, 11, 645–657. [Google Scholar] [CrossRef] [PubMed]
- Shekhar, S.; Yang, X. Natural killer cells in host defense against veterinary pathogens. Vet. Immunol. Immunopathol. 2015, 168, 30–34. [Google Scholar] [CrossRef] [PubMed]
- Childerstone, A.J.; Cedillo-baron, L.; Foster-cuevas, M.; Parkhouse, R.M.E. Demonstration of bovine CD8 M T-cell responses to foot-and-mouth disease virus. J. Gen. Virol. 1999, 80, 663–669. [Google Scholar] [CrossRef] [PubMed]
- Doel, T.R. FMD vaccines. Virus Res. 2003, 91, 81–99. [Google Scholar] [CrossRef]
- Windsor, M.A.; Carr, B.V.; Bankowski, B.; Gibson, D.; Reid, E.; Hamblin, P.; Gubbins, S.; Juleff, N.; Charleston, B. Cattle remain immunocompetent during the acute phase of foot-and-mouth disease virus infection. Vet. Res. 2011, 42, 108. [Google Scholar] [CrossRef] [PubMed]
- Robinson, L.; Charleston, B.; Rodriguez, L.L.; Gay, C.G.; Sumption, K.J. Global Foot-and-Mouth Disease Research Update and Gap Analysis: 6–Immunology. Transbound. Emerg. Dis. 2016, 63, 56–62. [Google Scholar] [CrossRef]
- Gowane, G.R.; Sharma, A.; Sankar, M.; Narayanan, K.; Das, B.; Subramaniam, S.; Pattnaik, B. Research in Veterinary Science Association of BoLA DRB3 alleles with variability in immune response among the crossbred cattle vaccinated for foot-and-mouth disease (FMD). Res. Vet. Sci. 2013, 95, 156–163. [Google Scholar] [CrossRef]
- López-Martínez, A.; Chávez-muñoz, C.; Granados, J. Histocompability main complex biological function. Rev. Investig. Clin. 2005, 57, 132–141. [Google Scholar]
- Baxter, R.; Craigmile, S.C.; Haley, C.; Douglas, A.J.; Williams, J.L.; Glass, E.J. BoLA-DR peptide binding pockets are fundamental for foot-and-mouth disease virus vaccine design in cattle. Vaccine 2010, 28, 28–37. [Google Scholar] [CrossRef]
- De Groot, A.S.; Nene, V.; Hegde, N.R. T cell epitope identification for bovine vaccines: An epitope mapping method for BoLA A-11. Int. J. Parasitol. 2003, 7519. [Google Scholar] [CrossRef]
- Longjam, N.; Deb, R.; Sarmah, A.K.; Tayo, T.; Awachat, V.B.; Saxena, V.K. A Brief Review on Diagnosis of Foot-and-Mouth Disease of Livestock: Conventional to Molecular Tools. Vet. Med. Int. 2011. [Google Scholar] [CrossRef] [PubMed]
- Langellotti, C.A. Interacción del Virus de la Fiebre Aftosa con las Células Dendríticas: Génesis de la Respuesta Inmune Adaptativa. Ph.D. Thesis, Universidad de Buenos Aires, Buenos Aires, Argentina, 2011. [Google Scholar]
- Schutta, C.; Barrera, J.; Pisano, M.; Zsak, L.; Grubman, M.J.; Mayr, G.A.; Moraes, M.P.; Kamicker, B.J.; Brake, D.A.; Ettyreddy, D.; et al. Multiple efficacy studies of an adenovirus-vectored foot-and-mouth disease virus serotype A24 subunit vaccine in cattle using homologous challenge. Vaccine 2017, 34, 3214–3220. [Google Scholar] [CrossRef] [PubMed]
- Romanutti, C. Evaluación de la Respuesta Inmune Inducida por Distintos Inmunógenos Recombinantes Dirigidos contra el Virus de la Fiebre Aftosa. Ph.D. Thesis, Universidad Nacional de Buenos Aires, Buenos Aires, Argentina, 2012. [Google Scholar]
- Mulcahy, G.; Gale, C.; Robertson, P.; Iyisan, S.; DiMarchi, R.D.; Doel, T. Isotype responses of infected, virus-vaccinated and peptide-vaccinated cattle to foot-and-mouth disease virus. Vaccine 1990, 8, 249–256. [Google Scholar] [CrossRef]
- Pega, J.; Di Giacomo, S.; Bucafusco, D.; Schammas, J.M.; Malacari, D.; Barrionuevo, F.; Capozzo, A.V.; Rodriguez, L.L.; Borca, M.V.; Pérez-Filgueira, M. Systemic Foot-and-Mouth Disease Vaccination in Cattle Promotes Specific Antibody-Secreting Cells at the Respiratory Tract and Triggers Local Anamnestic Responses upon Aerosol Infection. J. Virol. 2015, 89, 9581–9590. [Google Scholar] [CrossRef] [PubMed]
- Grubman, M.J. Development of novel strategies to control foot-and-mouth disease: Marker vaccines and antivirals IRES. Biologicals 2005, 33, 227–234. [Google Scholar] [CrossRef]
- Bautista, E.M.; Ferman, G.S.; Golde, W.T. Induction of lymphopenia and inhibition of T cell function during acute infection of swine with foot and mouth disease virus (FMDV). Vet. Immunol. Immunopathol. 2003, 92, 4–7. [Google Scholar] [CrossRef]
- Rodríguez Pulido, M.; Sáiz, M. Molecular Mechanisms of Foot-and-Mouth Disease Virus Targeting the Host Antiviral Response. Front. Cell Infect. Microbiol. 2017, 7, 1–9. [Google Scholar] [CrossRef]
- Chinsangaram, J.; Piccone, M.E.; Grubman, M.J. Ability of Foot-and-Mouth Disease Virus To Form Plaques in Cell Culture Is Associated with Suppression of Alpha/Beta Interferon. J. Virol. 1999, 73, 9891–9898. [Google Scholar] [CrossRef]
- Capozzo, V. Evaluación de la Respuesta Inmune Humoral Contra el Virus de la Fiebre Aftosa Inducida por Inmunógenos Tradicionales y Recombinantes: Desarrollo de métodos de Vacunación Alternativa Utilizando Proteínas Quiméricas y ADN como Inmunógenos. Ph.D. Thesis, Universidad de Buenos Aires, Buenos Aires, Argentina, 2002. [Google Scholar]
- García-briones, M.; Rosas, M.F.; González-magaldi, M. Differential distribution of non-structural proteins of foot-and-mouth disease virus in BHK-21 cells. Virology 2006, 349, 409–421. [Google Scholar] [CrossRef]
- Ho, M.Y.; Hung, S.W.; Liang, C.M.; Liang, S.M. Recombinant viral capsid protein VP1 suppresses lung cancer metastasis by inhibiting COX-2/PGE2 and MIG-7. Oncotarget 2014, 5, 3931–3943. [Google Scholar] [CrossRef][Green Version]
- Arzt, J.; Pacheco, J.M.; Stenfeldt, C.; Rodriguez, L.L. Pathogenesis of virulent and attenuated foot-and-mouth disease virus in cattle. Virol. J. 2017, 14, 1–15. [Google Scholar] [CrossRef] [PubMed]
- Belsham, G.J. Towards improvements in foot-and-mouth disease vaccine performance. Acta Vet. Scand. 2020, 62, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Singh, R.K.; Sharma, G.K.; Mahajan, S.; Dhama, K.; Basagoudanavar, S.H.; Hosamani, M.; Sreenivasa, B.P.; Chaicumpa, W.; Gupta, V.; Sanyal, A. Foot-and-mouth disease virus: Immunobiology advances in vaccines and vaccination strategies addressing vaccine failures—An Indian perspective. Vaccines 2019, 7, 90. [Google Scholar] [CrossRef] [PubMed]

Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Rodríguez-Habibe, I.; Celis-Giraldo, C.; Patarroyo, M.E.; Avendaño, C.; Patarroyo, M.A. A Comprehensive Review of the Immunological Response against Foot-and-Mouth Disease Virus Infection and Its Evasion Mechanisms. Vaccines 2020, 8, 764. https://doi.org/10.3390/vaccines8040764
Rodríguez-Habibe I, Celis-Giraldo C, Patarroyo ME, Avendaño C, Patarroyo MA. A Comprehensive Review of the Immunological Response against Foot-and-Mouth Disease Virus Infection and Its Evasion Mechanisms. Vaccines. 2020; 8(4):764. https://doi.org/10.3390/vaccines8040764
Chicago/Turabian StyleRodríguez-Habibe, Ibett, Carmen Celis-Giraldo, Manuel Elkin Patarroyo, Catalina Avendaño, and Manuel Alfonso Patarroyo. 2020. "A Comprehensive Review of the Immunological Response against Foot-and-Mouth Disease Virus Infection and Its Evasion Mechanisms" Vaccines 8, no. 4: 764. https://doi.org/10.3390/vaccines8040764
APA StyleRodríguez-Habibe, I., Celis-Giraldo, C., Patarroyo, M. E., Avendaño, C., & Patarroyo, M. A. (2020). A Comprehensive Review of the Immunological Response against Foot-and-Mouth Disease Virus Infection and Its Evasion Mechanisms. Vaccines, 8(4), 764. https://doi.org/10.3390/vaccines8040764




