Schistosoma haematobium Extracellular Vesicle Proteins Confer Protection in a Heterologous Model of Schistosomiasis
Abstract
1. Introduction
2. Materials and Methods
2.1. Ethics Statement
2.2. Parasite Material and Experimental Animals
2.3. Adult Worm Culture, ES Collection, and EV Purification
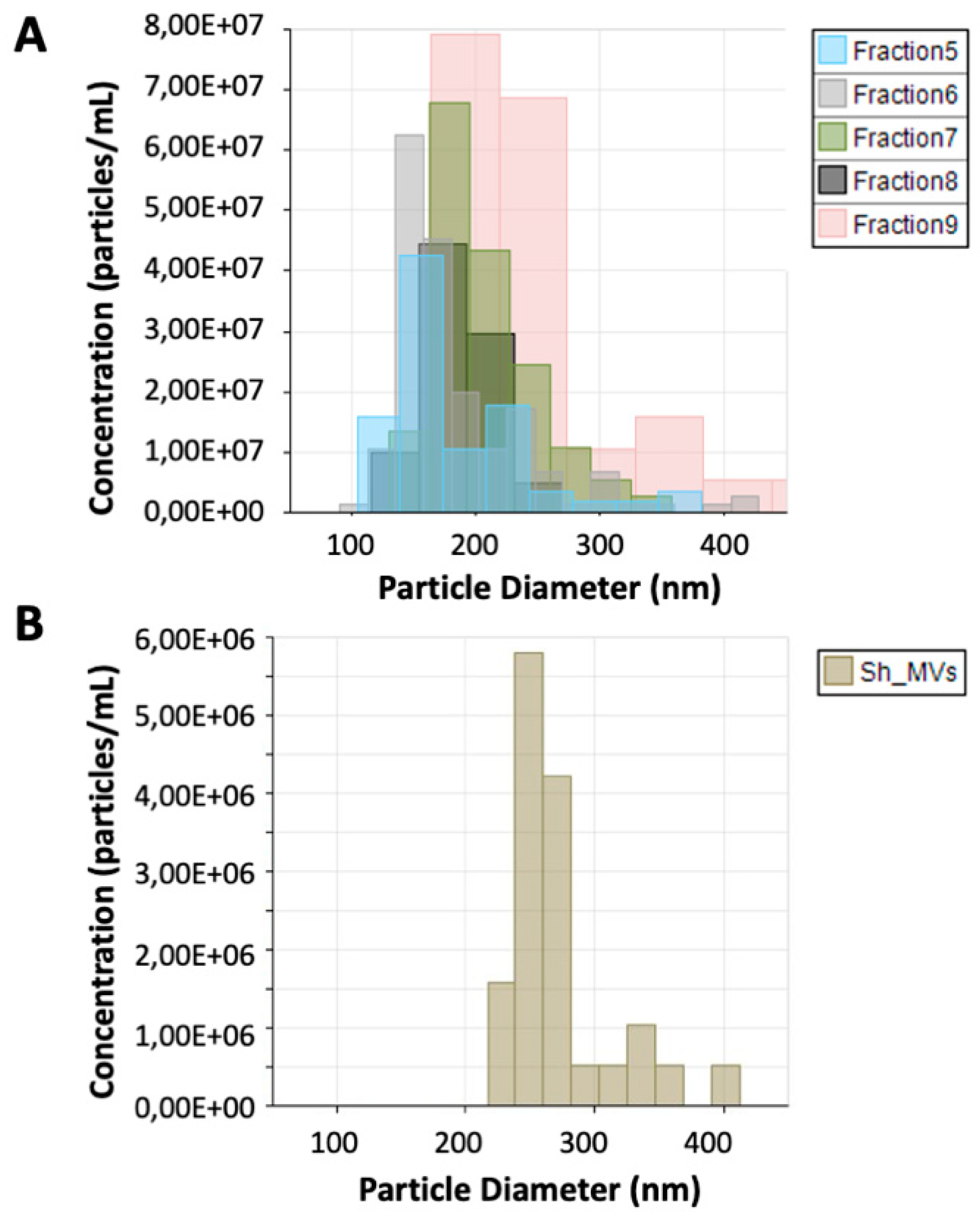
2.4. Determination of the Size and Concentration of EVs
2.5. In-Gel Trypsin Digestion of EVs
2.6. LC-MS/MS Analysis, Database Search, and Bioinformatic Analysis
2.7. Cloning and Expression of Sh-TSP2 and MS3_09198 in Pichia Pastoris
2.8. Cloning and Expression of MS3_01370 in Escherichia Coli
2.9. Vaccine Formulation and Immunization Schedule
2.10. Necropsy and Estimation of Parasite Burden
2.11. Statistics
3. Results
3.1. Density, Protein Concentration, Particle Concentration, and Purity of 120 k and 15 k Pellet Vesicles from Schistosoma haematobium
3.2. Proteomic Analysis of Schistosoma haematobium 120 k and 15 k Pellet Vesicles
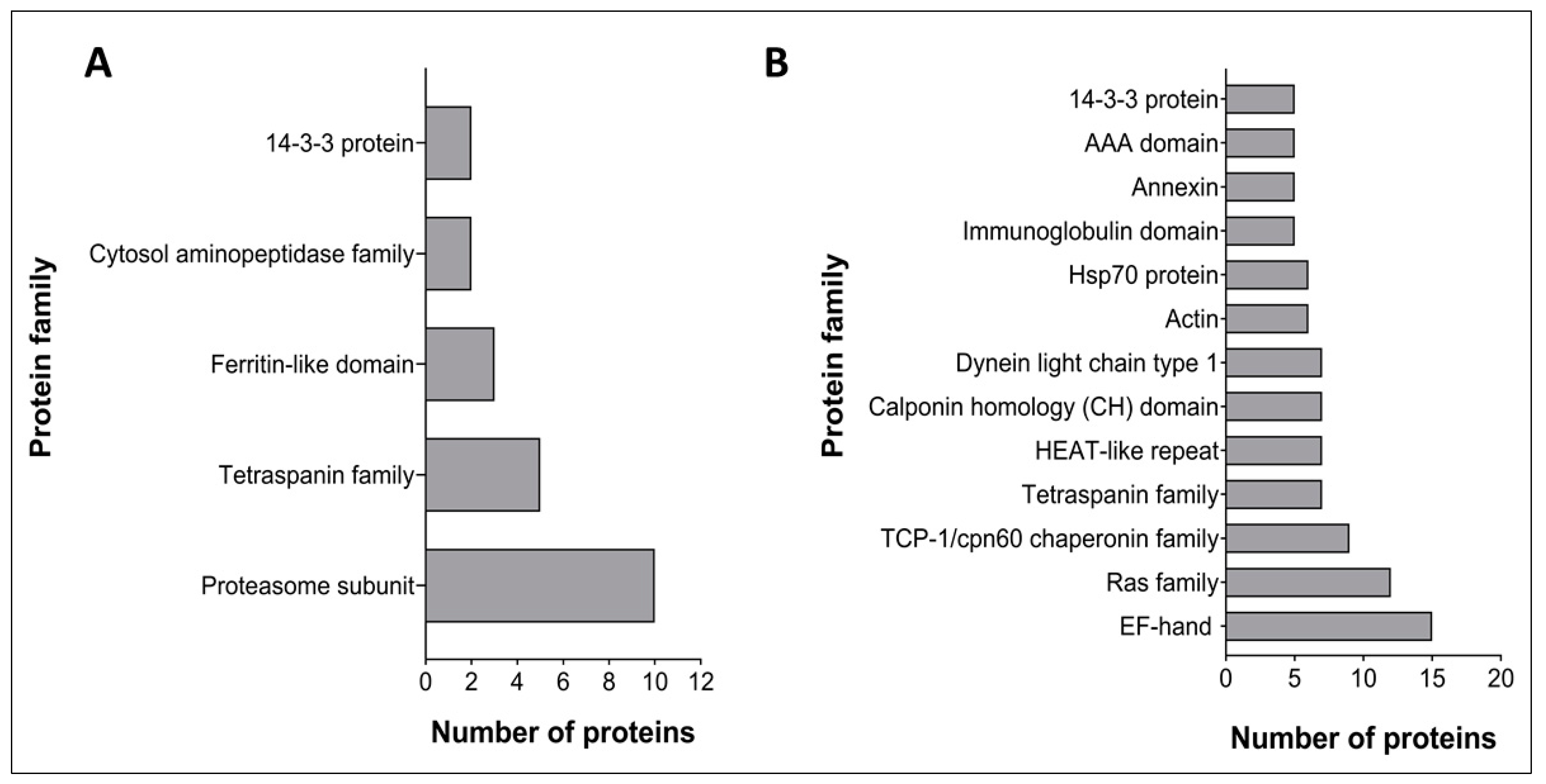
3.3. Protein Families Present in Schistosoma haematobium 120 k and 15 k Pellet Vesicles
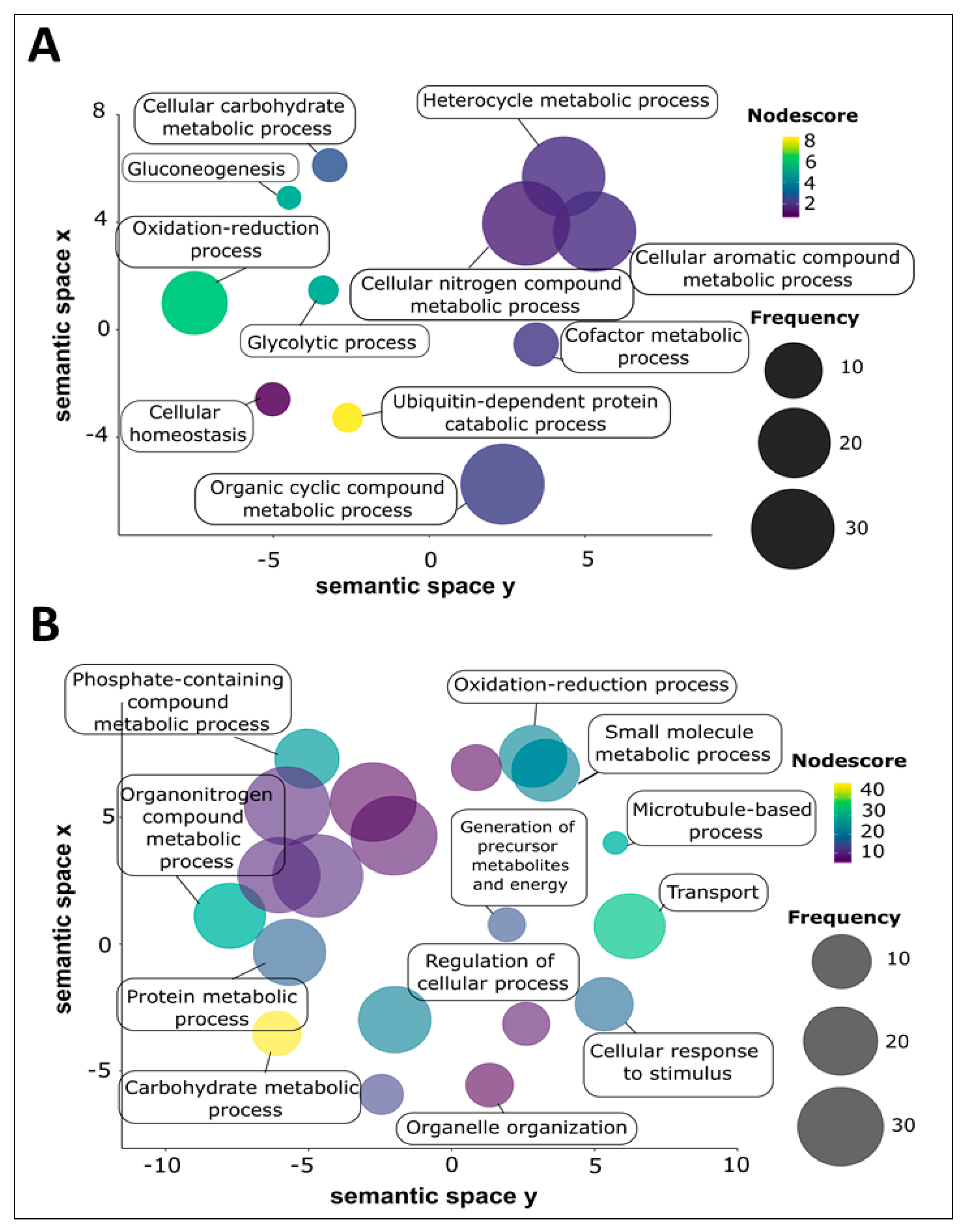
3.4. Gene Ontology of Proteins Identified from Schistosoma haematobium 120 k and 15 k Pellet Vesicles
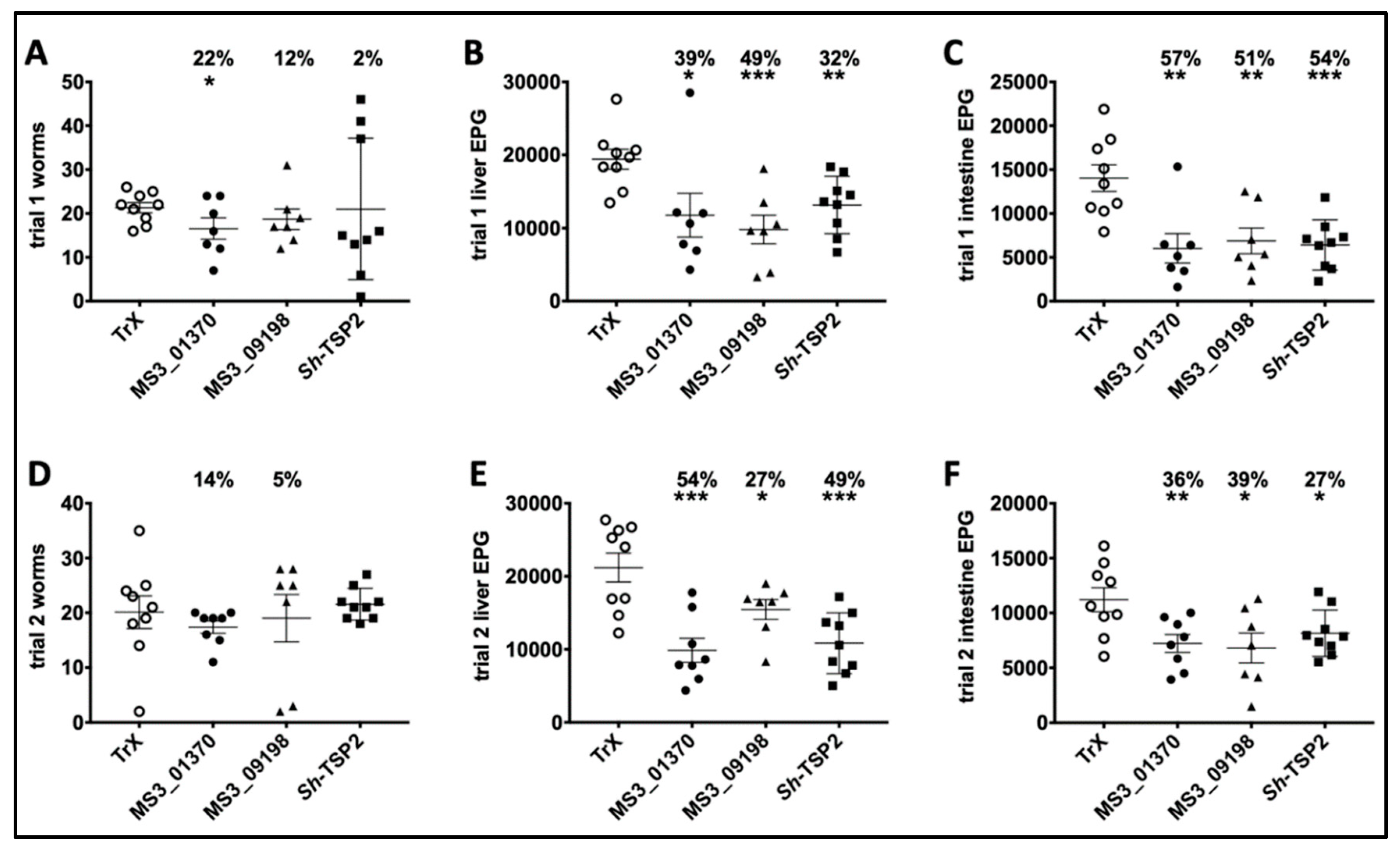
3.5. Parasite Burdens in Vaccinated and Control Mice
4. Discussion
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- McManus, D.P.; Dunne, D.W.; Sacko, M.; Utzinger, J.; Vennervald, B.; Zhou, X.N. Schistosomiasis. Nat. Rev. Dis. Prim. 2018, 4, 13. [Google Scholar] [CrossRef]
- Chitsulo, L.; Engels, D.; Montresor, A.; Savioli, L. The global status of schistosomiasis and its control. Acta Trop. 2000, 77, 41–51. [Google Scholar] [CrossRef]
- Boissier, J.; Grech-Angelini, S.; Webster, B.L.; Allienne, J.-F.; Huyse, T.; Mas-Coma, S.; Toulza, E.; Barré-Cardi, H.; Rollinson, D.; Kincaid-Smith, J.; et al. Outbreak of urogenital schistosomiasis in Corsica (France): An epidemiological case study. Lancet Infect. Dis. 2016, 16, 971–979. [Google Scholar] [CrossRef]
- Berger, S. Schistosoma Haematobium, 2017 ed.; GIDEON Informatics Inc.: Los Angeles, CA, USA, 2017. [Google Scholar]
- World Health Organization. Schistosomiasis: Number of People Treated Worldwide in 2014. Weekly Epidemiol. Record 2016, 91, 53–60. [Google Scholar]
- Berry, A.; Moné, H.; Iriart, X.; Mouahid, G.; Aboo, O.; Boissier, J.; Fillaux, J.; Cassaing, S.; Debuisson, C.; Valentin, A.; et al. schistosomiasis haematobium, Corsica, France. Emerg. Infect. Dis. 2014, 20, 1595–1597. [Google Scholar] [CrossRef] [PubMed]
- Wang, W.; Wang, L.; Liang, Y.S. Susceptibility or resistance of praziquantel in human schistosomiasis: A review. Parasitol. Res. 2012, 111, 1871–1877. [Google Scholar] [CrossRef] [PubMed]
- Vale, N.; Gouveia, M.J.; Rinaldi, G.; Brindley, P.J.; Gartner, F.; da Correia Costa, J.M. Praziquantel for Schistosomiasis: Single-Drug Metabolism Revisited, Mode of Action, and Resistance. Antimicrob. Agents Chemother. 2017, 61. [Google Scholar] [CrossRef] [PubMed]
- Gönnert, R.; Andrews, P. Praziquantel, a new broad-spectrum antischistosomal agent. Z. Parasitenkunde 1977, 52, 129–150. [Google Scholar] [CrossRef]
- McManus, D.P.; Bergquist, R.; Cai, P.; Ranasinghe, S.; Tebeje, B.M.; You, H. Schistosomiasis—From immunopathology to vaccines. Semin. Immunopathol. 2020, 42, 355–371. [Google Scholar] [CrossRef]
- Wilson, R.; Coulson, P. Why Don’t We Have a Schistosomiasis Vaccine? Parasitol. Today 1998, 14, 97–99. [Google Scholar] [CrossRef]
- Tebeje, B.M.; Harvie, M.; You, H.; Loukas, A.; McManus, D.P. Schistosomiasis vaccines: Where do we stand? Parasites Vectors 2016, 9, 1–5. [Google Scholar] [CrossRef] [PubMed]
- Riveau, G.; Schacht, A.-M.; Dompnier, J.-P.; Deplanque, D.; Seck, M.; Waucquier, N.; Senghor, S.; Delcroix-Genete, D.; Hermann, E.; Idris-Khodja, N.; et al. Safety and efficacy of the rSh28GST urinary schistosomiasis vaccine: A phase 3 randomized, controlled trial in Senegalese children. PLoS Negl. Trop. Dis. 2018, 12, e0006968. [Google Scholar] [CrossRef] [PubMed]
- Crowe, J.; Lumb, F.E.; Harnett, M.M.; Harnett, W. Parasite excretory-secretory products and their effects on metabolic syndrome. Parasite Immunol. 2017, 39, e12410. [Google Scholar] [CrossRef] [PubMed]
- Delcroix, M.; Medzihradsky, K.; Caffrey, C.R.; Fetter, R.D.; McKerrow, J.H. Proteomic analysis of adult S. mansoni gut contents. Mol. Biochem. Parasitol. 2007, 154, 95–97. [Google Scholar] [CrossRef]
- Hall, S.L.; Braschi, S.; Truscott, M.; Mathieson, W.; Cesari, I.M.; Wilson, R.A. Insights into blood feeding by schistosomes from a proteomic analysis of worm vomitus. Mol. Biochem. Parasitol. 2011, 179, 18–29. [Google Scholar] [CrossRef] [PubMed]
- Liu, F.; Cui, S.-J.; Hu, W.; Feng, Z.; Wang, Z.-Q.; Han, Z.-G. Excretory/Secretory Proteome of the Adult Developmental Stage of Human Blood Fluke, Schistosoma japonicum. Mol. Cell. Proteom. 2009, 8, 1236–1251. [Google Scholar] [CrossRef]
- Sotillo, J.; Pearson, M.S.; Becker, L.; Mekonnen, G.G.; Amoah, A.S.; Van Dam, G.J.; Corstjens, P.L.A.M.; Murray, J.; Mduluza, T.; Mutapi, F.; et al. In-depth proteomic characterization of Schistosoma haematobium: Towards the development of new tools for elimination. PLoS Negl. Trop. Dis. 2019, 13, e0007362. [Google Scholar] [CrossRef]
- Eichenberger, R.M.; Sotillo, J.; Loukas, A. Immunobiology of parasitic worm extracellular vesicles. Immunol. Cell Biol. 2018, 96, 704–713. [Google Scholar] [CrossRef]
- Kifle, D.W.; Pearson, M.S.; Becker, L.; Pickering, D.; Loukas, A.; Sotillo, J. Proteomic analysis of two populations of Schistosoma mansoni-derived extracellular vesicles: 15 k pellet and 120 k pellet vesicles. Mol. Biochem. Parasitol. 2020, 236, 111264. [Google Scholar] [CrossRef]
- Mekonnen, G.G.; Pearson, M.S.; Loukas, A.; Sotillo, J. Extracellular vesicles from parasitic helminths and their potential utility as vaccines. Expert Rev. Vaccines 2018, 17, 1–9. [Google Scholar] [CrossRef]
- Kifle, D.W.; Sotillo, J.; Pearson, M.S.; Loukas, A. Extracellular vesicles as a target for the development of anti-helminth vaccines. Emerg. Top. Life Sci. 2017, 1, 659–665. [Google Scholar] [CrossRef]
- Kim, K.M.; Abdelmohsen, K.; Mustapic, M.; Kapogiannis, D.; Gorospe, M. RNA in extracellular vesicles. Wiley Interdiscip. Rev. RNA 2017, 8, e1413. [Google Scholar] [CrossRef] [PubMed]
- Pathan, M.; Fonseka, P.; Chitti, S.V.; Kang, T.; Sanwlani, R.; Van Deun, J.; Hendrix, A.; Mathivanan, S. Vesiclepedia 2019: A compendium of RNA, proteins, lipids and metabolites in extracellular vesicles. Nucleic Acids Res. 2019, 47, D516–D519. [Google Scholar] [CrossRef] [PubMed]
- Coakley, G.; McCaskill, J.L.; Borger, J.G.; Simbari, F.; Robertson, E.; Millar, M.; Harcus, Y.; McSorley, H.J.; Maizels, R.M.; Buck, A.H. Extracellular vesicles from a helminth parasite suppress macrophage activation and constitute an effective vaccine for protective immunity. Cell Rep. 2017, 19, 1545–1557. [Google Scholar] [CrossRef]
- Tritten, L.; Geary, T.G. Helminth extracellular vesicles in host-parasite interactions. Curr. Opin. Microbiol. 2018, 46, 73–79. [Google Scholar] [CrossRef]
- Sotillo, J.; Pearson, M.S.; Potriquet, J.; Becker, L.; Pickering, D.; Mulvenna, J.; Loukas, A. Extracellular vesicles secreted by Schistosoma mansoni contain protein vaccine candidates. Int. J. Parasitol. 2016, 46, 1–5. [Google Scholar] [CrossRef] [PubMed]
- Nowacki, F.C.; Swain, M.T.; Klychnikov, O.I.; Niazi, U.; Ivens, A.; Quintana, J.F.; Hensbergen, P.J.; Hokke, C.H.; Buck, A.H.; Hoffmann, K.F. Protein and small non-coding RNA-enriched extracellular vesicles are released by the pathogenic blood fluke Schistosoma mansoni. J. Extracell. Vesicles 2015, 4, 28665. [Google Scholar] [CrossRef]
- Chaiyadet, S.; Sotillo, J.; Smout, M.J.; Cantacessi, C.; Jones, M.K.; Johnson, M.S.; Turnbull, L.; Whitchurch, C.B.; Potriquet, J.; Laohaviroj, M.; et al. Carcinogenic Liver Fluke Secretes Extracellular Vesicles That Promote Cholangiocytes to Adopt a Tumorigenic Phenotype. J. Infect. Dis. 2015, 212, 1636–1645. [Google Scholar] [CrossRef]
- Cwiklinski, K.; De La Torre-Escudero, E.; Trelis, M.; Bernal, D.; Dufresne, P.J.; Brennan, G.P.; O’Neill, S.; Tort, J.F.; Paterson, S.; Marcilla, A.; et al. The Extracellular Vesicles of the Helminth Pathogen, Fasciola hepatica: Biogenesis Pathways and Cargo Molecules Involved in Parasite Pathogenesis. Mol. Cell. Proteom. 2015, 14, 3258–3273. [Google Scholar] [CrossRef]
- Zhu, L.; Liu, J.; Dao, J.; Lu, K.; Li, H.; Gu, H.; Liu, J.; Feng, X.; Cheng, G. Molecular characterization of S. japonicum exosome-like vesicles reveals their regulatory roles in parasite-host interactions. Sci. Rep. 2016, 6, 25885. [Google Scholar] [CrossRef]
- Zheng, Y.; Guo, X.; Su, M.; Guo, A.; Ding, J.; Yang, J.; Xiang, H.; Cao, X.; Zhang, S.; Ayaz, M.; et al. Regulatory effects of Echinococcus multilocularis extracellular vesicles on RAW264.7 macrophages. Veter. Parasitol. 2017, 235, 29–36. [Google Scholar] [CrossRef]
- Nicolao, M.C.; Rodrigues, C.R.; Cumino, A.C. Extracellular vesicles from Echinococcus granulosus larval stage: Isolation, characterization and uptake by dendritic cells. PLoS Negl. Trop. Dis. 2019, 13, e0007032. [Google Scholar] [CrossRef] [PubMed]
- Eichenberger, R.M.; Ryan, S.; Jones, L.; Buitrago, G.; Polster, R.; De Oca, M.M.; Zuvelek, J.; Giacomin, P.R.; Dent, L.A.; Engwerda, C.R.; et al. Hookworm Secreted Extracellular Vesicles Interact With Host Cells and Prevent Inducible Colitis in Mice. Front. Immunol. 2018, 9, 850. [Google Scholar] [CrossRef]
- Eichenberger, R.M.; Talukder, H.; Field, M.A.; Wangchuk, P.; Giacomin, P.R.; Loukas, A.; Sotillo, J. Characterization of Trichuris muris secreted proteins and extracellular vesicles provides new insights into host–parasite communication. J. Extracell. Vesicles 2018, 7, 1428004–1428016. [Google Scholar] [CrossRef] [PubMed]
- Bernal, L.; Trelis, M.; Montaner, S.; Cantalapiedra, F.; Galiano, A.; Hackenberg, M.; Gomis, A.M.; Tarbes, S.M. Surface analysis of Dicrocoelium dendriticum. The molecular characterization of exosomes reveals the presence of miRNAs. J. Proteom. 2014, 105, 232. [Google Scholar] [CrossRef] [PubMed]
- Buck, A.H.; Coakley, G.; Simbari, F.; McSorley, H.J.; Quintana, J.F.; Le Bihan, T.; Kumar, S.; Abreu-Goodger, C.; Lear, M.; Harcus, Y.; et al. Exosomes secreted by nematode parasites transfer small RNAs to mammalian cells and modulate innate immunity. Nat. Commun. 2014, 5, 5488. [Google Scholar] [CrossRef]
- Hansen, E.P.; Fromm, B.; Andersen, S.D.; Marcilla, A.; Andersen, K.L.; Borup, A.; Williams, A.R.; Jex, A.R.; Gasser, R.B.; Young, N.D.; et al. Exploration of extracellular vesicles from Ascaris suum provides evidence of parasite–host cross talk. J. Extracell. Vesicles 2019, 8, 1578116. [Google Scholar] [CrossRef]
- Wang, L.; Li, Z.; Shen, J.; Liu, Z.; Liang, J.; Wu, X.; Sun, X.; Wu, Z. Exosome-like vesicles derived by Schistosoma japonicum adult worms mediates M1 type immune-activity of macrophage. Parasitol. Res. 2015, 114, 1865–1873. [Google Scholar] [CrossRef] [PubMed]
- Zamanian, M.; Fraser, L.M.; Agbedanu, P.N.; Harischandra, H.; Moorhead, A.R.; Day, T.; Bartholomay, L.C.; Kimber, M.J. Release of Small RNA-containing Exosome-like Vesicles from the Human Filarial Parasite Brugia malayi. PLoS Negl. Trop. Dis. 2015, 9, e0004069. [Google Scholar] [CrossRef] [PubMed]
- Shears, R.K.; Bancroft, A.J.; Hughes, G.W.; Grencis, R.K.; Thornton, D.J. Extracellular vesicles induce protective immunity against Trichuris muris. Parasite Immunol. 2018, 40, e12536. [Google Scholar] [CrossRef]
- Chaiyadet, S.; Sotillo, J.; Krueajampa, W.; Thongsen, S.; Brindley, P.J.; Sripa, B.; Loukas, A.; Laha, T. Vaccination of hamsters with Opisthorchis viverrini extracellular vesicles and vesicle-derived recombinant tetraspanins induces antibodies that block vesicle uptake by cholangiocytes and reduce parasite burden after challenge infection. PLoS Negl. Trop. Dis. 2019, 13, e0007450. [Google Scholar] [CrossRef] [PubMed]
- Trelis, M.; Galiano, A.; Bolado, A.; Toledo, R.; Gomis, A.M.; Bernal, D. Subcutaneous injection of exosomes reduces symptom severity and mortality induced by Echinostoma caproni infection in BALB/c mice. Int. J. Parasitol. 2016, 46, 799–808. [Google Scholar] [CrossRef] [PubMed]
- Coakley, G.; Maizels, R.M.; Buck, A.H. Exosomes and Other Extracellular Vesicles: The New Communicators in Parasite Infections. Trends Parasitol. 2015, 31, 477–489. [Google Scholar] [CrossRef] [PubMed]
- Da’Dara, A.; Skelly, P.J.; Wang, M.; Harn, D.A. Immunization with plasmid DNA encoding the integral membrane protein, Sm23, elicits a protective immune response against schistosome infection in mice. Vaccine 2001, 20, 359–369. [Google Scholar] [CrossRef]
- Pearson, M.S.; Pickering, D.A.; McSorley, H.J.; Bethony, J.M.; Tribolet, L.; Dougall, A.M.; Hotez, P.J.; Loukas, A. Enhanced Protective Efficacy of a Chimeric Form of the Schistosomiasis Vaccine Antigen Sm-TSP-2. PLoS Negl. Trop. Dis. 2012, 6, e1564. [Google Scholar] [CrossRef]
- Tran, M.H.; Pearson, M.S.; Bethony, J.; Smyth, D.J.; Jones, M.K.; Duke, M.; Don, A.T.; McManus, D.P.; Correa-Oliveira, R.; Loukas, A. Tetraspanins on the surface of Schistosoma mansoni are protective antigens against schistosomiasis. Nat. Med. 2006, 12, 835–840. [Google Scholar] [CrossRef]
- Keitel, W.; Potter, G.; Diemert, D.; Bethony, J.; El Sahly, H.; Kennedy, J.; Patel, S.; Plieskatt, J.; Jones, W.; Deye, G.; et al. A phase 1 study of the safety, reactogenicity, and immunogenicity of a Schistosoma mansoni vaccine with or without glucopyranosyl lipid A aqueous formulation (GLA-AF) in healthy adults from a non-endemic area. Vaccine 2019, 37, 6500–6509. [Google Scholar] [CrossRef]
- Yuan, C.; Fu, Y.-J.; Li, J.; Yue, Y.-F.; Cai, L.-L.; Xiao, W.-J.; Chen, J.-P.; Yang, L. Schistosoma japonicum: Efficient and rapid purification of the tetraspanin extracellular loop 2, a potential protective antigen against schistosomiasis in mammalian. Exp. Parasitol. 2010, 126, 456–461. [Google Scholar] [CrossRef]
- Tucker, M.S.; Karunaratne, L.B.; Lewis, F.A.; Frietas, T.C.; Liang, Y.-S. Schistosomiasis. In Current Protocols in Immunology; Coico, R., Ed.; John Wiley and Sons, Inc.: Hoboken, NJ, USA, 2013; pp. 19.11.11–19.11.57. [Google Scholar] [CrossRef]
- Stroehlein, A.J.; Korhonen, P.K.; Chong, T.M.; Lim, Y.L.; Chan, K.-G.; Webster, L.B.; Rollinson, D.; Brindley, P.J.; Gasser, R.; Young, N.D. High-quality Schistosoma haematobium genome achieved by single-molecule and long-range sequencing. GigaScience 2019, 8. [Google Scholar] [CrossRef]
- Young, N.D.; Jex, A.R.; Li, B.; Liu, S.; Yang, L.; Xiong, Z.; Li, Y.; Cantacessi, C.; Hall, R.S.; Xu, X.; et al. Whole-genome sequence of Schistosoma haematobium. Nat. Genet. 2012, 44, 221–225. [Google Scholar] [CrossRef]
- Craig, R.; Cortens, A.J.P.; Beavis, R.C. Open Source System for Analyzing, Validating, and Storing Protein Identification Data. J. Proteome Res. 2004, 3, 1234–1242. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.; Pevzner, P.A. MS-GF+ makes progress towards a universal database search tool for proteomics. Nat. Commun. 2014, 5, 5277. [Google Scholar] [CrossRef] [PubMed]
- Diament, B.J.; Noble, W.S. Faster SEQUEST Searching for Peptide Identification from Tandem Mass Spectra. J. Proteome Res. 2011, 10, 3871–3879. [Google Scholar] [CrossRef] [PubMed]
- Vaudel, M.; Barsnes, H.; Berven, F.S.; Sickmann, A.; Martens, L. SearchGUI: An open? Source graphical user interface for simultaneous OMSSA and X!Tandem searches. Proteomics 2011, 11, 996–999. [Google Scholar] [CrossRef] [PubMed]
- Finn, R.D.; Clements, J.; Eddy, S.R. HMMER web server: Interactive sequence similarity searching. Nucleic Acids Res. 2011, 39 (Suppl. 2), W29–W37. [Google Scholar] [CrossRef] [PubMed]
- Conesa, A.; Goetz, S.; García-Gómez, J.M.; Terol, J.; Talon, M.; Robles, M. Blast2GO: A universal tool for annotation, visualization and analysis in functional genomics research. Bioinformation 2005, 21, 3674–3676. [Google Scholar] [CrossRef]
- Supek, F.; Bosnjak, M.; Skunca, N.; Šmuc, T. REVIGO Summarizes and Visualizes Long Lists of Gene Ontology Terms. PLoS ONE 2011, 6, e21800. [Google Scholar] [CrossRef]
- Sonnhammer, E.L.; von Heijne, G.; Krogh, A. A hidden Markov model for predicting transmembrane helices in protein sequences. Proc. Int. Conf. Intell. Syst. Mol. Biol. 1998, 6, 175–182. [Google Scholar]
- Petersen, T.N.; Brunak, S.; Heijne, G.; Nielsen, H. SignalP 4.0: Discriminating signal peptides from transmembrane regions. Nat. Methods 2011, 8, 785–786. [Google Scholar] [CrossRef]
- Tedla, B.A.; Pickering, D.; Becker, L.; Loukas, A.; Pearson, M.S. Vaccination with Schistosoma mansoni Cholinesterases Reduces the Parasite Burden and Egg Viability in a Mouse Model of Schistosomiasis. Vaccines 2020, 8, 162. [Google Scholar] [CrossRef]
- WHO. Fact Sheet on Neglected Tropical Diseases. Available online: https://www.who.int/en/news-room/fact-sheets/detail/schistosomiasis (accessed on 20 January 2019).
- Bergquist, R.; Utzinger, J.; Keiser, J. Controlling schistosomiasis with praziquantel: How much longer without a viable alternative? Infect. Dis. Poverty 2017, 6, 74. [Google Scholar] [CrossRef] [PubMed]
- Cioli, D. Praziquantel: Is there real resistance and are there alternatives? Curr. Opin. Infect. Dis. 2000, 13, 659–663. [Google Scholar] [CrossRef] [PubMed]
- Tucher, C.; Bode, K.; Schiller, P.; Classen, L.; Birr, C.; Souto-Carneiro, M.M.; Blank, N.; Lorenz, H.-M.; Schiller, M. Extracellular Vesicle Subtypes Released From Activated or Apoptotic T-Lymphocytes Carry a Specific and Stimulus-Dependent Protein Cargo. Front. Immunol. 2018, 9, 534. [Google Scholar] [CrossRef] [PubMed]
- Bhattacharyya, S.; Yu, H.; Mim, C.; Matouschek, A. Regulated protein turnover: Snapshots of the proteasome in action. Nat. Rev. Mol. Cell Biol. 2014, 15, 122–133. [Google Scholar] [CrossRef]
- De Paula, R.G.; Ornelas, A.M.D.M.; Morais, E.R.; Gomes, M.D.S.; Aguiar, D.D.P.; Magalhães, L.G.; Rodrigues, V. Proteasome stress responses in Schistosoma mansoni. Parasitol. Res. 2015, 114, 1747–1760. [Google Scholar] [CrossRef]
- Guerra-Sá, R.; Castro-Borges, W.; Evangelista, E.; Kettelhut, I.C.; Rodrigues, V. Schistosoma mansoni: Functional proteasomes are required for development in the vertebrate host. Exp. Parasitol. 2005, 109, 228–236. [Google Scholar] [CrossRef]
- Nabhan, J.F.; El-Shehabi, F.; Patocka, N.; Ribeiro, P. The 26S proteasome in Schistosoma mansoni: Bioinformatics analysis, developmental expression, and RNA interference (RNAi) studies. Exp. Parasitol. 2007, 117, 337–347. [Google Scholar] [CrossRef]
- Arosio, P.; Elia, L.; Poli, M. Ferritin, cellular iron storage and regulation. IUBMB Life 2017, 69, 414–422. [Google Scholar] [CrossRef]
- Orino, K.; Lehman, L.; Tsuji, Y.; Ayaki, H.; Torti, S.V.; Torti, F.M. Ferritin and the response to oxidative stress. Biochem. J. 2001, 357, 241–247. [Google Scholar] [CrossRef]
- Jones, M.K.; McManus, D.P.; Sivadorai, P.; Glanfield, A.; Moertel, L.; Belli, S.I.; Gobert, G.N. Tracking the fate of iron in early development of human blood flukes. Int. J. Biochem. Cell Biol. 2007, 39, 1646–1658. [Google Scholar] [CrossRef][Green Version]
- Chen, L.-Y.; Yi, X.-Y.; Zeng, X.-F.; Zhang, S.-K.; McReynolds, L. Mucosal immunization of recombinant Schistosoma japonicum ferritin. Zhongguo Ji Sheng Chong Xue Yu Ji Sheng Chong Bing Za Zhi 2004, 22, 129–132. [Google Scholar] [PubMed]
- Nagamune, K.; Moreno, S.N.; Chini, E.N.; Sibley, L.D. Calcium regulation and signaling in apicomplexan parasites. Subcell. Biochem. 2008, 47, 70–81. [Google Scholar] [CrossRef] [PubMed]
- Bucki, R.; Bachelot-Loza, C.; Zachowski, A.; Giraud, F.; Sulpice, J.-C. Calcium Induces Phospholipid Redistribution and Microvesicle Release in Human Erythrocyte Membranes by Independent Pathways†. Biochemistry 1998, 37, 15383–15391. [Google Scholar] [CrossRef] [PubMed]
- Sotillo, J.; Robinson, M.W.; Kimber, M.J.; Cucher, M.; Ancarola, M.E.; Nejsum, P.; Marcilla, A.; Eichenberger, R.M.; Tritten, L. The protein and microRNA cargo of extracellular vesicles from parasitic helminths—Current status and research priorities. Int. J. Parasitol. 2020. [Google Scholar] [CrossRef] [PubMed]
- Repasky, G.; Chenette, E.J.; Der, C.J. Renewing the conspiracy theory debate: Does Raf function alone to mediate Ras oncogenesis? Trends Cell Biol. 2004, 14, 639–647. [Google Scholar] [CrossRef] [PubMed]
- Abels, E.R.; Breakefield, X.O. Introduction to Extracellular Vesicles: Biogenesis, RNA Cargo Selection, Content, Release, and Uptake. Cell. Mol. Neurobiol. 2016, 36, 301–312. [Google Scholar] [CrossRef] [PubMed]
- Schüßler, P.; Grevelding, C.G.; Kunz, W. Identification of Ras, MAP kinases, and a GAP protein in Schistosoma mansoni by immunoblotting and their putative involvement in male–female interaction. Parasitology 1997, 115 Pt 6, 629–634. [Google Scholar] [CrossRef]
- Gómez-Puertas, P.; Martín-Benito, J.; Carrascosa, J.L.; Willison, K.R.; Valpuesta, J.M. The substrate recognition mechanisms in chaperonins. J. Mol. Recognit. 2004, 17, 85–94. [Google Scholar] [CrossRef]
- Frydman, J.; Nimmesgern, E.; Erdjument-Bromage, H.; Wall, J.; Tempst, P.; Hartl, F. Function in protein folding of TRiC, a cytosolic ring complex containing TCP-1 and structurally related subunits. EMBO J. 1992, 11, 4767–4778. [Google Scholar] [CrossRef]
- Andreu, Z.; Yáñez-Mó, M. Tetraspanins in Extracellular Vesicle Formation and Function. Front. Immunol. 2014, 5, 442. [Google Scholar] [CrossRef]
- Chaiyadet, S.; Krueajampa, W.; Hipkaeo, W.; Plosan, Y.; Piratae, S.; Sotillo, J.; Smout, M.J.; Sripa, B.; Brindley, P.J.; Loukas, A.; et al. Suppression of mRNAs encoding CD63 family tetraspanins from the carcinogenic liver fluke Opisthorchis viverrini results in distinct tegument phenotypes. Sci. Rep. 2017, 7, 14342. [Google Scholar] [CrossRef] [PubMed]
- Piratae, S.; Tesana, S.; Jones, M.K.; Brindley, P.J.; Loukas, A.; Lovas, E.; Eursitthichai, V.; Sripa, B.; Thanasuwan, S.; Laha, T. Molecular Characterization of a Tetraspanin from the Human Liver Fluke, Opisthorchis viverrini. PLoS Negl. Trop. Dis. 2012, 6, e1939. [Google Scholar] [CrossRef] [PubMed]
- Tran, M.H.; Freitas, T.C.; Cooper, L.; Gaze, S.; Gatton, M.; Jones, M.K.; Lovas, E.; Pearce, E.J.; Loukas, A. Suppression of mRNAs Encoding Tegument Tetraspanins from Schistosoma mansoni Results in Impaired Tegument Turnover. PLoS Pathog. 2010, 6, e1000840. [Google Scholar] [CrossRef] [PubMed]
- Zhang, W.; Li, J.; Duke, M.; Jones, M.K.; Kuang, L.; Zhang, J.; Blair, D.; Li, Y.; McManus, D.P. Inconsistent Protective Efficacy and Marked Polymorphism Limits the Value of Schistosoma japonicum Tetraspanin-2 as a Vaccine Target. PLoS Negl. Trop. Dis. 2011, 5, e1166. [Google Scholar] [CrossRef] [PubMed]
- Kifle, D.W.; Chaiyadet, S.; Waardenberg, A.J.; Wise, I.; Cooper, M.; Becker, L.; Doolan, D.L.; Laha, T.; Sotillo, J.; Pearson, M.S.; et al. Uptake of Schistosoma mansoni extracellular vesicles by human endothelial and monocytic cell lines and impact on vascular endothelial cell gene expression. Int. J. Parasitol. 2020. [Google Scholar] [CrossRef] [PubMed]
- Burke, M.L.; Jones, M.K.; Gobert, G.N.; Li, Y.S.; Ellis, M.K.; McManus, D.P. Immunopathogenesis of human schistosomiasis. Parasite Immunol. 2009, 31, 163–176. [Google Scholar] [CrossRef]
- Beaumier, C.M.; Gillespie, P.M.; Hotez, P.J.; Bottazzi, M.E. New vaccines for neglected parasitic diseases and dengue. Transl. Res. 2013, 162, 144–155. [Google Scholar] [CrossRef]
- Bushara, H.O.; Bashir, M.E.N.; Malik, K.H.E.; Mukhtar, M.M.; Trottein, F.; Capron, A.; Taylor, M.G. Suppression of Schistosoma bovis egg production in cattle by vaccination with either glutathione S-transferase or keyhole limpet haemocyanin. Parasite Immunol. 1993, 15, 383–390. [Google Scholar] [CrossRef]
- Mbanefo, E.; Kumagai, T.; Kodama, Y.; Kurosaki, T.; Furushima-Shimogawara, R.; Cherif, M.S.; Mizukami, S.; Kikuchi, M.; Huy, N.T.; Ohta, N.; et al. Immunogenicity and anti-fecundity effect of nanoparticle coated glutathione S-transferase (SjGST) DNA vaccine against murine Schistosoma japonicum infection. Parasitol. Int. 2015, 64, 24–31. [Google Scholar] [CrossRef]
- Bushara, H.O.; Majid, A.A.; Saad, A.M.; Hussein, M.F.; Taylor, M.G.; Dargie, J.D.; Marshall, T.F.; Nelson, G.S. Observations on cattle schistosomiasis in the Sudan, a study in comparative medicine. II. Experimental demonstration of naturally acquired resistance to Schistosoma bovis. Am. J. Trop. Med. Hyg. 1980, 29, 442–451. [Google Scholar] [CrossRef]
- Saad, A.; Hussein, M.; Dargie, J.; Taylor, M.; Nelson, G. Schistosoma bovis in calves: The development and clinical pathology of primary infections. Res. Veter. Sci. 1980, 28, 105–111. [Google Scholar] [CrossRef]
- Massoud, J. Parasitological and Pathological Observations on Schistosoma bovis Sonsion, 1876, in Calves, Sheep and Goats in Iran. J. Helminthol. 1973, 47, 155–164. [Google Scholar] [CrossRef] [PubMed]
- Cheever, A.W.; Powers, K.G. Schistosoma mansoni infection in rhesus monkeys: Changes in egg production and egg distribution in prolonged infections in intact and splenectomized monkeys. Ann. Trop. Med. Parasitol. 1969, 63, 83–93. [Google Scholar] [CrossRef] [PubMed]
- Webbe, G.; James, C.; Nelson, G.S.; Smithers, S.R.; Terry, R.J. Acquired resistance to Schistosoma haematobium in the baboon (Papio anubis) after cercarial exposure and adult worm transplantation. Ann. Trop. Med. Parasitol. 1976, 70, 411–424. [Google Scholar] [CrossRef] [PubMed]
- WHO. Ending the Neglect to Attain the Sustainable Development Goals: A Roadmap for Neglected Tropical Diseases 2021–2030. 2020. Available online: https://www.who.int/neglected_diseases/Ending-the-neglect-to-attain-the-SDGs--NTD-Roadmap.pdf?ua=1 (accessed on 2 July 2020).
- Kura, K.; Truscott, J.E.; Toor, J.; Anderson, R.M. Modelling the impact of a Schistosoma mansoni vaccine and mass drug administration to achieve morbidity control and transmission elimination. PLoS Negl. Trop. Dis. 2019, 13, e0007349. [Google Scholar] [CrossRef] [PubMed]
| Protein Category | Protein Accession Numbers |
|---|---|
| 120 k vesicles | |
| Proteasome subunit | MS3_10249.1, MS3_05734.1, MS3_01483.1, MS3_06009.1, MS3_04526.1, MS3_08808.1, MS3_07240.1, MS3_02807.1, MS3_09236.1, MS3_03070.1 |
| GAPDH | MS3_10141.1 |
| Papain family cysteine protease | MS3_08498.1 |
| C-terminal domain of 1-Cys peroxiredoxin | MS3_08460.1 |
| Ferritin-like domain | MS3_08059.1 |
| S-adenosyl-L-homocysteine hydrolase | MS3_04449.1 |
| Cytosol amino peptidase | MS3_01749.1 |
| Trefoil (P-type) domain-containing protein | MS3_00004.1 |
| 15k vesicles | |
| EF hand | MS3_05735.1, MS3_00180.1, MS3_09846.1, MS3_05877.1, MS3_05317.1, MS3_04536.1, MS3_10043.1, MS3_05959.1, MS3_05150.1, MS3_04275.1, MS3_05958.1 MS3_05952.1, MS3_00361.1, MS3_02003.1 |
| Ras family | MS3_10193.1, MS3_05953.1, MS3_05910.1, MS3_05976.1, MS3_07854.1, MS3_11139.1, MS3_02375.1, MS3_01653.1, MS3_04355.1, MS3_09110.1, MS3_09593.1, MS3_03443.1 |
| TCP-1/cpn60 chaperonin family | MS3_03054.1, MS3_06928.1, MS3_01627.1, MS3_10572.1, MS3_06669.1, MS3_07556.1, MS3_08399.1, MS3_00785.1, MS3_08926.1 |
| Tetraspanins | MS3_01905.1, MS3_01370 |
| Heat-like repeat | MS3_08696.1, MS3_01642.1, MS3_09658.1, MS3_10590.1, MS3_05814.1, MS3_02928.1, MS3_06293.1 |
| Calponin homology (CH) domain | MS3_07481.1, MS3_05505.1, MS3_01744.1, MS3_00852.1, MS3_00361.1, MS3_03766.1, MS3_10701.1 |
| Dynein light chain type 1 | MS3_05351.1, MS3_08569.1, MS3_05345.1, MS3_01173.1, MS3_05342.1, MS3_04412.1, MS3_05960.1 |
| Actin | MS3_07374.1, MS3_04014.1, MS3_00351.1, MS3_02465.1, MS3_04907.1, MS3_01922.1 |
| HSP-70 protein | MS3_10713.1, MS3_11293.1, MS3_11411.1, MS3_10049.1, MS3_02688.1, MS3_02787.1 |
| Immunoglobulin domain | MS3_03027.1, MS3_01271.1, MS3_03208.1, MS3_07594.1, MS3_01223.1 |
| Annexin | MS3_08725.1, MS3_08723.1, MS3_04598.1, MS3_01964.1, MS3_01952.1 |
| AAA domain | MS3_03802.1, MS3_02581.1, MS3_01139.1, MS3_01650.1, MS3_07031.1 |
| 14-3-3 protein | MS3_03977.1, MS3_05219.1, MS3_00047.1, MS3_01871.1, MS3_03976.1 |
| 120 k and 15 k vesicles | |
| Tetraspanins | MS3_09198, Sh-TSP-2, MS3_05226, MS3_05289, MS3_01153 |
| Ferritin-like domain | MS3_07972.1, MS3_07178.1 |
| 14-3-3 protein | MS3_03977.1, MS3_00047.1 |
| Elongation factor Tu C-terminal domain | MS3_08479.1 |
| EF hand | MS3_08446.1 |
| Actin | MS3_07374.1 |
| GST, N-terminal domain | MS3_06482.1 |
| Cytosol aminopeptidase family, catalytic domain | MS3_08450.1 |
| Lipocalin/cytosolic fatty-acid binding protein family | MS3_04307.1 |
| Immunoglobulin domain | MS3_03208.1 |
| Saposin-like type B, region 2 | MS3_02805.1 |
| Enolase, N-terminal domain | MS3_02425.1 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Mekonnen, G.G.; Tedla, B.A.; Pickering, D.; Becker, L.; Wang, L.; Zhan, B.; Bottazzi, M.E.; Loukas, A.; Sotillo, J.; Pearson, M.S. Schistosoma haematobium Extracellular Vesicle Proteins Confer Protection in a Heterologous Model of Schistosomiasis. Vaccines 2020, 8, 416. https://doi.org/10.3390/vaccines8030416
Mekonnen GG, Tedla BA, Pickering D, Becker L, Wang L, Zhan B, Bottazzi ME, Loukas A, Sotillo J, Pearson MS. Schistosoma haematobium Extracellular Vesicle Proteins Confer Protection in a Heterologous Model of Schistosomiasis. Vaccines. 2020; 8(3):416. https://doi.org/10.3390/vaccines8030416
Chicago/Turabian StyleMekonnen, Gebeyaw G., Bemnet A. Tedla, Darren Pickering, Luke Becker, Lei Wang, Bin Zhan, Maria Elena Bottazzi, Alex Loukas, Javier Sotillo, and Mark S. Pearson. 2020. "Schistosoma haematobium Extracellular Vesicle Proteins Confer Protection in a Heterologous Model of Schistosomiasis" Vaccines 8, no. 3: 416. https://doi.org/10.3390/vaccines8030416
APA StyleMekonnen, G. G., Tedla, B. A., Pickering, D., Becker, L., Wang, L., Zhan, B., Bottazzi, M. E., Loukas, A., Sotillo, J., & Pearson, M. S. (2020). Schistosoma haematobium Extracellular Vesicle Proteins Confer Protection in a Heterologous Model of Schistosomiasis. Vaccines, 8(3), 416. https://doi.org/10.3390/vaccines8030416







