Abstract
Myxoma virus (MYXV) is the prototypic member of the Leporipoxvirus genus of the Poxviridae family of viruses. In nature, MYXV is highly restricted to leporids and causes a lethal disease called myxomatosis only in European rabbits (Oryctologous cuniculus). However, MYXV has been shown to also productively infect various types of nonrabbit transformed and cancer cells in vitro and in vivo, whereas their normal somatic cell counterparts undergo abortive infections. This selective tropism of MYXV for cancer cells outside the rabbit host has facilitated its development as an oncolytic virus for the treatment of different types of cancers. Like other poxviruses, MYXV possesses a large dsDNA genome which encodes an array of dozens of immunomodulatory proteins that are important for host and cellular tropism and modulation of host antiviral innate immune responses, some of which are rabbit-specific and others can function in nonrabbit cells as well. This review summarizes the functions of one such MYXV host range protein, M029, an ortholog of the larger superfamily of poxvirus encoded E3-like dsRNA binding proteins. M029 has been identified as a multifunctional protein involved in MYXV cellular and host tropism, antiviral responses, and pathogenicity in rabbits.
1. Introduction
Myxoma virus (MYXV) is the prototypic member of the Leporipoxvirus genus of the Poxviridae family of viruses. Other members of Leporipoxviruses are Shope (Rabbit) fibroma virus (SFV), hare fibroma virus, and Squirrel fibroma virus. The natural hosts for these viruses are Lagomorphs and squirrels. The natural evolutionary host of MYXV is the South American leporid Sylvilagus brasiliensis, also called tapeti or brush rabbit. In these animals, MYXV causes a benign cutaneous fibroma, which persists for a few weeks before clearance. The virus in these natural habitats of South America is transmitted by mosquitoes or fleas from one host to another without virus replication in the vector. However, infection of the European rabbit (Oryctolagus cuniculus) causes a rapid systemic and lethal disease called myxomatosis [1,2,3,4,5]. The disease myxomatosis is characterized by extensive internal and external lesions and severe immunosuppression accompanied by Gram-negative bacterial infections of the respiratory tract [6]. The natural host of SFV is the eastern cottontail rabbit Sylvilagus floridanus, where it causes fibrosarcoma-like tumors [7,8]. Unlike MYXV, SFV causes only benign localized fibromas in European rabbits [3]. These apparent differences between the two viruses have been partially attributed to the fact that MYXV, unlike SFV, can inhibit an apoptotic response in rabbit T lymphocytes and thus is able to replicate and spread via infected leukocytes through the lymphatic system [9]. However, other immunomodulatory proteins in MYXV as well likely contribute to the unique lethality in European rabbits. Due to the extreme lethality of MYXV in European rabbits and the ability of the virus to act as a highly transmissible host-restricted pathogen, MYXV was released in Australia and Europe to control the feral European rabbit populations [10]. Since then, the virus and the host have undergone continuous co-evolution in real-time and is one of the most well-documented natural host-microbial pathogen experiments in evolutionary biology [11]. This virus-host interaction also provides new insights into virus adaptation to new hosts and the evolution of virulence. Since the time MYXV was released in Australia in the 1950s, many virus isolates were sequenced in different phases to track the phenotypic evolution of virulence in both Australia and Europe [10,12,13,14,15]. On the other hand, pathogen pressure also caused changes in the antiviral genes in the host European rabbits across two continents [16]. Interestingly, recent reports describe a new natural recombinant version of MYXV from Spain and Portugal that has leaped from rabbits into hares and caused a novel myxomatosis-like disease in this new host [17,18,19]. It will be of interest to deduce the viral genes in this new virus (designated MYXV-Toledo) that is/are responsible for mediating this host species leap into hares.
The genomes of both MYXV and SFV have been sequenced [20,21]. The double-stranded DNA genome (161.8 kb) of MYXV encodes about 171 functional Open Reading Frames (ORFs), many of which are unique to the leporipoxviruses. The central region of the genome harbors about 100 genes that are conserved among poxviruses, encoding proteins involved in mostly structural and housekeeping functions. The rest of the genes located in the terminal-inverted repeats (TIRs) and the near-terminal unique regions are mostly immunomodulatory genes involved in subverting the host immune system. These genes are predicted to be involved in host-specific functions. Many of these gene functions are studied in MYXV [22,23,24]. In SFV, five MYXV gene orthologs are deleted, which may have caused SFV to become less pathogenic in European rabbits [21]. The immunomodulatory proteins encoded by poxviruses have been classified as viroceptors (virus-encoded mimics of immune receptors), virokines (virus-encoded mimics of immune proteins like cytokines), and intracellular signaling modulators [22,25,26]. This review focuses on one of the vital MYXV-encoded intracellular modulators, and considered a major host range factor, called M029.
3. Role in Poxvirus Replication
Poxvirus encoded E3-like proteins play an essential role in virus replication in host cell lines. In the case of VACV, the deletion of the E3L gene compromised the ability of the virus to replicate in many diverse cell lines [40,41,42,43,44,45]. However, VACV E3 mutant virus was successfully generated and grown in RK13 or BHK21 cell lines derived from rabbit and hamster, respectively [40,46]. Unlike VACV, generation of MYXV and Ectromelia virus (ECTV) E3 orthologs mutant viruses by homologous recombination using flanking sequences and insertion of a reporter gene was not successful in cell lines where these viruses naturally can replicate. In both cases, engineered complementing cell lines were required for constructing and isolating the E3/M029 mutant viruses. In the case of MYXV, the rabbit RK13 cell line engineered to express stably the E3 protein of VACV was used to create the vMyx-M029KO virus [47]. For ECTV, engineered BS-C-1 cells that stably express the E3 protein of VACV were used to generate the ECTVΔE3L virus [48]. Both knockout viruses were only able to be propagated in these complementing cell lines. The purified deletion viruses, when tested for infection and replication in cultured cell lines originated from different species; the viruses were invariably highly defective in replication. For example, vMyx-M029KO virus was not able to replicate at all in tested cell lines originating from human, mouse, or nonhuman primates [47]. The M029-minus mutant virus was also compromised in the viral replication cycle (single step or multistep) in rabbit cell lines like RK13 and RL5 [47]. Similar results were reported in the case of the ECTVΔE3L virus, which was unable to replicate in many mammalian cell lines originating from human, monkey, and mouse sources [48]. This defect in MYXV replication in the absence of M029 and ECTV in the absence of E3 was due to defects that manifest in late gene expression. In the infected cells, the early gene expression and DNA replication allowed the formation of a small viral factory, which indicated the abortive replication of the knockout viruses [48]. The importance of E3 in the life cycle of Modified vaccinia Ankara (MVA), a highly attenuated strain of VACV developed as a live but nonreplicating vaccine, has also been studied [44]. In the absence of E3L, MVA was defective in DNA replication, late transcription, and protein synthesis in HeLa cells [44]. These studies thus highlight the importance of E3-like proteins in the viral replication cycle.
4. Role in Pathogenesis
Poxvirus-encoded E3-like proteins are universally essential for the pathogenesis in their native susceptible hosts. Most of the in vivo functions of these proteins are known from the studies on VACV E3. For E3, both the N and C-terminal domains are required for the in vivo pathogenesis of VACV in mice [32,46,49,50]. Similar results were reported with E3 from ECTV. Infection of susceptible BALB/c mice with ECTVΔE3L resulted in the survival of all the mice, whereas infection with wildtype (WT) ECTV and an ECTVΔE3L-revertant virus caused 100% mortality [48]. These results clearly demonstrated that E3 proteins encoded by orthopoxviruses are essential for the virulence in their pathogenic host. Since MYXV causes the lethal disease myxomatosis only in European rabbits, the role of M029 was studied in lab rabbits. In order to understand the role of M029 in MYXV pathogenesis, the vMyx-M029KO virus was intradermally injected in the flank of lab rabbits [47]. Using the same inoculation route, the WT-MYXV- and a M029KO-revertant virus caused 100% mortality of rabbits, whereas the vMyx-M029KO caused only very mild redness and transient swelling at the inoculation site. Unlike the WT-MYXV, no secondary lesion was observed with vMyx-M029KO-infected rabbits [47]. These results suggested that the vMyx-M029KO was highly defective in replication and spread in vivo. Studies with intratracheal injection of VACVΔE3L in C57BL/6 mice exhibited immune cell infiltration and mild edema around the infected bronchi of the lung and very little damage to the lung compared to the WT-VACV infection [46]. Similar results were observed from vMyx-M029KO infection in rabbits, where in the absence of M029, the mutant virus was rapidly cleared by the immune system. In order to test whether the E3-like proteins can complement the in vivo function of VACV E3, several in vivo studies were done in mice [39]. Surprisingly, in all the studies, the E3 protein orthologs were not able to rescue the full pathogenesis of VACV in the host. For example, in the absence of VACV E3, the orthologous proteins SPPV34, YMTV34, SPV032 and M029 were not able to restore the in vivo pathogenesis of VACV in the BALB/c mice when viruses were delivered intranasally [39]. However, as mentioned before, in vitro, both M029 and SPV032 restored the replication of the VACVΔE3L virus. Similar results were reported when the orf E3 protein was expressed in VACV lacking E3L [39]. The orf E3 expressing VACV was more than 1000-fold less pathogenic than WT VACV [38]. These results suggest that E3 orthologous proteins encoded by different poxviruses have evolved to be adapted for more host-specific functions for pathogenesis in specific host species, which are yet to be identified.
6. Myxoma Virus Interference of the Type I IFN Pathway and the Role of M029
Type I IFNs are produced and released from cells in response to virus infection or other stimulation. They play a major role in activating host antiviral innate and adaptive immune responses. Type I IFNs bind to specific IFN-receptors on cells to trigger signaling pathways that result in the expression of genes called IFN-stimulated genes (ISGs). The expression of ISGs render the cells resistant to subsequent virus infection. Poxviruses have acquired different mechanisms to inhibit production of IFNs, IFN signaling and function of ISGs [25,26,84]. However, among the members of poxviruses, this ability to interfere with type I IFN signaling and responses varies widely. For example, VACV can inhibit almost every aspect of type I IFN signaling in cell lines originating from different species [80,84]. MYXV, on the other hand, is very species-specific in terms of inhibition of type I IFN and may explain why MYXV is restricted to only lagomorphs. This was first reported in a study using primary mouse embryonic fibroblasts (pMEFs) [85]. MYXV infection of pMEFs activated ERK signaling which activated IRF3 and eventually type I IFN induction. This early induction of type I IFN prevented virus gene expression and replication in pMEFs [85]. The role of ERK was further confirmed by inhibition of ERK signaling or disruption of STAT1-mediated IFN signaling, both of which completely rescued MYXV replication in the pMEFs. This was further confirmed in vivo in mice where MYXV did not cause any disease. However, STAT1 deficiency made the mice highly susceptible to lethal MYXV infection [85]. Thus, the ERK-IFN-STAT1 signaling cascade is the key to protect mice against MYXV infection. Although this signaling cascade is highly conserved across the species, MYXV can completely inhibit this cascade and cause lethal disease in rabbits. So, this suggests that MYXV-encoded proteins that target this pathway are highly host- and species-specific.
The IFN pathway is also a vital restriction determinant of MYXV replication in primary human fibroblasts [86]. In contrast to pMEFs, when they were spontaneously immortalized (iMEFs) by multiple passages, these immortalized cells now fully supported productive MYXV infection [87]. In iMEFS, the IFN signaling pathway was intact; however, the MYXV infection-induced type I IFN induction was compromised, possibly at the level of the initial sensing of the virus infection [87]. These findings were further confirmed by the observation that unlike pMEFs, the phosphorylation of ERK1/2 was not detectable, which was further confirmed that there was little or no induction of type I IFN production in the iMEFs [87]. These results indicated that pathogen-associated molecular patterns (PAMPs) sensing of MYXV in pMEFs is likely a key step in induction of type I IFN that is completely lost during the immortalization process. These studies thus confirmed that in immortalized or transformed nonrabbit cells (such as NIH3T3 or iMEFs), the type I IFN-induced antiviral state can restrict MYXV replication. In a later study, this observation was further extended by comparing cells derived from rabbits, humans and mice [88]. Representative cell lines from all these species were able to induce an antiviral state when treated with type I IFN. But surprisingly, when they were infected with MYXV, the viral ability to infect the various type I IFN-treated cells was dramatically different. In rabbit cells, despite the induction of antiviral state, the virus was able to replicate completely even in the presence of the fully-formed antiviral state. In human cells, on the other hand, the virus was able to partially replicate, whereas in the mouse cell line the virus was not able to replicate to any measurable extent [88]. This suggests that MYXV’s ability to counteract the effector functions of the type I IFN induced antiviral state is highly species-specific.
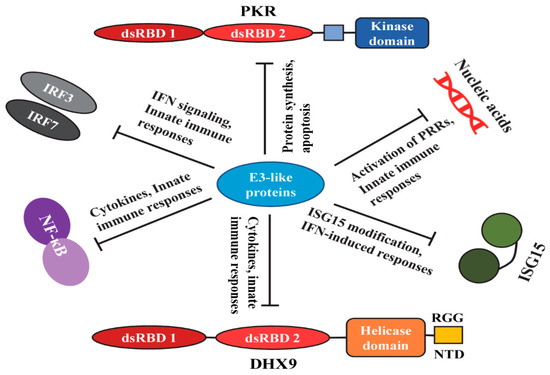
This species-specific ability to inhibit the operational IFN-induced antiviral state was completely dependent on the presence of the M029 protein [88]. In the absence of M029, the vMyxM029KO virus was not able to inhibit the type I antiviral state in rabbit cells. Even in the human cells, the partial inhibition of the antiviral state was also completely lost in the absence of M029 [88]. These results indicated that unlike VACV E3, which is relatively promiscuous in terms of host species specificity, the MYXV M029 retained rabbit and to some extent human-specific inhibitory activity against a type I IFN-induced antiviral state. Furthermore, it was confirmed that this antiviral state was independent of the host cell PKR [88]. In the absence of PKR expression, the vMyxM029KO was inhibited by the antiviral state in rabbit and human cells. This suggests that ISG(s), in addition to PKR, remain to be identified as likely targets of M029 for the inhibition of the type I IFN antiviral state in rabbit and human cells. Based on the studies from VACV, the candidate ISGs could be ISG15, OAS, or IFITs [89,90,91]. Among these ISGs, ISG15 is directly targeted by VACV E3 (Figure 2). In addition, MYXV lacked a homolog of VACV B18, a soluble inhibitor of type I IFNs that directly binds to type I IFN and inhibits type I IFN-induced signaling. B18 showed broad species specificity and can inhibit type I IFNs from humans, rabbits, mice, rats, and bovine animals [92,93,94]. The absence of such an inhibitor also restricts MYXV capacity to infect species other than the leporids.
7. Other Cellular Targets of E3 and M029
E3 orthologs are one of the critical proteins in host immune modulation by targeting different cellular pathways. However, many cellular- and host-specific protein targets are yet to be identified. Expression of tagged E3-like proteins in the infected cells and mass-spectrometry followed by co-immunoprecipitation (co-IP) will allow identification of unknown interacting host proteins. To identify the cellular interactions of M029, a recombinant MYXV expressing a V5-tagged M029 protein under the native promoter was constructed [47]. Co-IP using the anti-V5 antibody allowed the pull-down of proteins that interacted with M029 during MYXV infection. Using this approach, cellular DHX9 was identified as an additional cellular interaction of M029 in human cancer cells [47]. However, unlike PKR knockdown, the DHX9 knockdown did not rescue the replication of MYXV in the absence of M029. In another study, DHX9 was identified as a host antiviral RNA helicase in human cancer cell lines where MYXV replicated poorly, for example, pancreatic cancer cell line PANC-1 and renal cancer cell line 786-0 [95]. In these cell lines, knockdown of DHX9 significantly enhanced MYXV replication. In addition, DHX9 has proviral functions based on its requirement for the replication of multiple RNA viruses [96]. DHX9 also functions as a sensor for induction of innate immune responses against PAMPs and viruses in immune cells [97,98,99]. Thus, both RNA and DNA viruses exploit DHX9 to optimize their replication. Like MYXV M029, the VACV E3 protein has been shown to interact with DHX9 in human monocytes, such as THP1 cells. Unlike dendritic cells, in this cell line, the nucleic acid-sensing role of DHX9 was not found, suggesting the cell-type-specific function of DHX9 as a viral PAMP sensor. However, in these cells, DHX9 played a role in IL-6 induction, which was directly antagonized by VACVC E3 [100] (Figure 2). These results suggest that by direct interaction with DHX9, E3 might suppress innate immunity by reducing the synthesis of selected innate defense cytokines.
8. Vaccine Vector and Gene Delivery
Several members of the poxvirus family have been developed as a vaccine platform because of their host restriction. For example, VACV strain MVA, fowlpox, and canarypox (members of the genus avipoxvirus), have been developed as vaccine vectors for different host species [101,102,103,104,105]. Since MYXV only infects leporids but still can bind to most mammalian cells of any species, and causes abortive infections in primary nonrabbit cells, it has the potential to be developed as safe vaccine platform for hosts outside of rabbits. As a vaccine vector platform outside of leporids, the safety and efficacy of MYXV has been tested by vaccination of cats against feline calicivirus [106,107] and in sheep against bluetongue virus [108,109]. Also as a rabbit-specific vaccine platform, recombinant attenuated strains of MYXV that do not cause myxomatosis have been used for rabbit vaccination against myxomatosis and rabbit hemorrhagic disease [110]. The M029-deleted MYXV could be a safe alternative for the development of the attenuated vaccine platform in rabbits, and possibly other species as well. VACV E3L mutants are investigated as a vaccine platform due to their greater safety profile in comparison to WT-VACV [111,112]. The New York City Board of Health (NYCBH) strain of VACV was compared to NYCBH deleted for E3L (NYCBHΔE3L) in a rabbitpox virus (RPXV) challenge study in the rabbit model [113]. Both the WT-NYCBH and NYCBHΔE3L vaccines completely protected rabbits against a lethal challenge of RPXV. However, unlike a single dose of WT-NYCBH, two doses of NYCBHΔE3L were required to prevent weight loss, fever, and clinical symptoms following challenge with RPV [113]. The attenuated NYCBHΔE3L was also tested in cynomolgus macaques, which were challenged with heterologous MPXV. However, in this model, NYCBHΔE3L provided partial protection against the MPXV challenge [114]. Unlike different VACV strains, MYXV has not been widely tested as a vaccine candidate, other than the examples cited above. However, the inability to cause disease outside the leporids and the capacity to generate robust immune responses in murine or human dendritic cells suggests that MYXV could readily be adapted as a vaccine platform in humans [115,116]. In this context, the M029-deleted MYXV could be a candidate vaccine platform for this purpose.
9. Conclusions and Future Directions
Among the many members of poxviruses with restricted host tropism, MYXV is by far the most well-studied model to understand the regulation of viral pathogen-host species interactions. Since MYXV was released in Australia and Europe in the 1950s to control feral rabbit populations, it also serves as a real-time model to study co-evolution of virus and host in the wild. MYXV encodes many unique proteins which have, or are predicted to have, host-restricted immunomodulatory functions. Studies of MYXV-encoded genes by genetic manipulation have allowed greater understanding of how poxviruses like MYXV can interact with, and subvert, the host immune system in a species-specific manner. Due to this natural host restriction, coupled with the acquired ability to infect transformed or cancerous cells originating from different species outside of the rabbit, MYXV is currently being developed as an oncolytic virus for the treatment of different types of human cancers. To mediate the tropism of MYXV for cancer cells, the virus-encoded immunomodulatory proteins clearly play an important role. However, in an intact nonrabbit host with a functional immune system, the operational activities of these immunomodulatory proteins can have a much different outcome than in rabbits. In this context, for the clinical development of MYXV, it is important to understand the function of these proteins in both lagomorphs and non-lagomorphs. M029 is an essential immune regulatory protein encoded by MYXV that mediates the virus tropism in both primary rabbit cells and in nonrabbit cancer cells. As a member of the E3 family of poxvirus host range proteins, M029 retains vital features important for MYXV tropism in all rabbit cells and many nonrabbit cancer cells by the regulation of the early innate immune responses in the infected cells. To further understand the function(s) of M029 and other E3-like proteins, the identification of additional cellular interactions will be crucial.
Author Contributions
M.M.R. and G.M. wrote the manuscript. All authors have read and agreed to the published version of the manuscript.
Funding
G.M. and M.M.R.’s research is supported by NIH grant R01 AI080607 and a start-up fund from Arizona State University.
Conflicts of Interest
The authors declare no conflict of interest. G.M. is co-founder and M.M.R. is consultant of a biotech company, OncoMyx Therapeutics, devoted to development of multi-armed MYXV for treatment of cancer.
References
- Kerr, P.; Best, S. Myxoma virus in rabbits. Revue Scientifique Technique 1998, 17, 256–268. [Google Scholar] [CrossRef] [PubMed]
- Bertagnoli, S.; Marchandeau, S. Myxomatosis. Revue Scientifique Technique 2015, 34, 549–556. [Google Scholar]
- Kerr, P.J.; Donnelly, T.M. Viral Infections of Rabbits. Vet. Clin. N. Am. Exot. Anim. Pract. 2013, 16, 437–468. [Google Scholar] [CrossRef] [PubMed]
- Fenner, F.; Woodroofe, G.M. The Pathogenesis of Infectious Myxomatosis: The Mechanism of Infection and the Immunological Response in the European Rabbit (Oryctolagus cuniculus). Br. J. Exp. Pathol. 1953, 34, 400–411. [Google Scholar]
- Fenner, F.; Day, M.F.; Woodroofe, G.M. Epidemiological consequences of the mechanical transmission of myxomatosis by mosquitoes. J. Hyg. 1956, 54, 284. [Google Scholar] [CrossRef]
- Fenner, F. Myxomatosis; University Press: Cambridge, MA, USA, 1965. [Google Scholar]
- Cikanek, S.J.; Carpenter, J.W.; Lindemann, D.M.; Hallman, R.; Eshar, D.; Kim, I.J.; Almes, K.M. Shope Fibroma in the External Ear Canal of a Domestic Rabbit. Comp. Med. 2017, 67, 51–55. [Google Scholar]
- Shope, R.E. A Transmissible Tumor-Like Condition in Rabbits. J. Exp. Med. 1932, 56, 793–802. [Google Scholar] [CrossRef]
- Macen, J.L.; Graham, K.A.; Lee, S.F.; Schreiber, M.; Boshkov, L.K.; McFadden, G. Expression of the Myxoma Virus Tumor Necrosis Factor Receptor Homologue and M11L Genes Is Required to Prevent Virus-Induced Apoptosis in Infected Rabbit T Lymphocytes. Virology 1996, 218, 232–237. [Google Scholar] [CrossRef]
- Kerr, P. Myxomatosis in Australia and Europe: A model for emerging infectious diseases. Antivir. Res. 2012, 93, 387–415. [Google Scholar] [CrossRef]
- Fenner, F. Evolutionary aspects of Myxomatosis in Australia. Memórias Instituto Oswaldo Cruz 1956, 54, 269–278. [Google Scholar] [CrossRef]
- Kerr, P.; Liu, J.; Cattadori, I.; Ghedin, E.; Read, A.; Holmes, E.C. Myxoma Virus and the Leporipoxviruses: An Evolutionary Paradigm. Viruses 2015, 7, 1020–1061. [Google Scholar] [CrossRef]
- Kerr, P.J.; Ghedin, E.; DePasse, J.V.; Fitch, A.; Cattadori, I.; Hudson, P.J.; Tscharke, D.C.; Read, A.F.; Holmes, E.C. Evolutionary History and Attenuation of Myxoma Virus on Two Continents. PLoS Pathog. 2012, 8, e1002950. [Google Scholar] [CrossRef] [PubMed]
- Kerr, P.J.; Rogers, M.B.; Fitch, A.; DePasse, J.V.; Cattadori, I.; Twaddle, A.; Hudson, P.J.; Tscharke, D.; Read, A.F.; Holmes, E.C.; et al. Genome Scale Evolution of Myxoma Virus Reveals Host-Pathogen Adaptation and Rapid Geographic Spread. J. Virol. 2013, 87, 12900–12915. [Google Scholar] [CrossRef] [PubMed]
- Kerr, P.J.; Cattadori, I.; Liu, J.; Sim, D.G.; Dodds, J.W.; Brooks, J.W.; Kennett, M.J.; Holmes, E.C.; Read, A.F. Next step in the ongoing arms race between myxoma virus and wild rabbits in Australia is a novel disease phenotype. Proc. Natl. Acad. Sci. USA 2017, 114, 9397–9402. [Google Scholar] [CrossRef] [PubMed]
- Alves, J.M.; Carneiro, M.; Cheng, J.Y.; De Matos, A.L.; Rahman, M.M.; Loog, L.; Campos, P.; Wales, N.; Eriksson, A.; Manica, A.; et al. Parallel adaptation of rabbit populations to myxoma virus. Science (N.Y.) 2019, 363, 1319–1326. [Google Scholar] [CrossRef] [PubMed]
- Águeda-Pinto, A.; De Matos, A.L.; Abrantes, M.; Kraberger, S.; Risalde, M.A.; Risalde, M.A.; McFadden, G.; Varsani, A.; Esteves, P. Genetic Characterization of a Recombinant Myxoma Virus in the Iberian Hare (Lepus granatensis). Viruses 2019, 11, 530. [Google Scholar] [CrossRef] [PubMed]
- Dalton, K.P.; Martín, J.M.; Nicieza, I.; Podadera, A.; De Llano, D.; Casais, R.; Gimenez, S.; Badiola, I.; Agüero, M.; Duran, M.; et al. Myxoma virus jumps species to the Iberian hare. Transbound. Emerg. Dis. 2019, 66, 2218–2226. [Google Scholar] [CrossRef]
- García-Bocanegra, I.; Camacho-Sillero, L.; Risalde, M.A.; Dalton, K.P.; Caballero-Gómez, J.; Agüero, M.; Zorrilla, I.; Gómez-Guillamón, F. First outbreak of myxomatosis in Iberian hares (Lepus granatensis). Transbound. Emerg. Dis. 2019, 66, 2204–2208. [Google Scholar] [CrossRef]
- Cameron, C.; Hota-Mitchell, S.; Chen, L.; Barrett, J.; Cao, J.-X.; Macaulay, C.; Willer, D.; Evans, D.; McFadden, G. The Complete DNA Sequence of Myxoma Virus. Virology 1999, 264, 298–318. [Google Scholar] [CrossRef]
- Willer, D.O.; McFadden, G.; Evans, D.H. The Complete Genome Sequence of Shope (Rabbit) Fibroma Virus. Virology 1999, 264, 319–343. [Google Scholar] [CrossRef]
- Liu, J.; Wennier, S.; McFadden, G. The immunoregulatory properties of oncolytic myxoma virus and their implications in therapeutics. Microbes Infect. 2010, 12, 1144–1152. [Google Scholar] [CrossRef] [PubMed]
- Stanford, M.M.; Werden, S.J.; McFadden, G. Myxoma virus in the European rabbit: Interactions between the virus and its susceptible host. Vet. Res. 2007, 38, 299–318. [Google Scholar] [CrossRef] [PubMed]
- Rahman, M.M.; McFadden, G. Oncolytic Virotherapy with Myxoma Virus. J. Clin. Med. 2020, 9, 171. [Google Scholar] [CrossRef] [PubMed]
- Seet, B.T.; Johnston, J.B.; Brunetti, C.R.; Barrett, J.W.; Everett, H.; Cameron, C.; Sypula, J.; Nazarian, S.H.; Lucas, A.; McFadden, G. Poxviruses and immune evasion. Ann. Rev. Immunol. 2003, 21, 377–423. [Google Scholar] [CrossRef]
- Johnston, J.B.; McFadden, G. Poxvirus Immunomodulatory Strategies: Current Perspectives. J. Virol. 2003, 77, 6093–6100. [Google Scholar] [CrossRef]
- Chang, H.W.; Watson, J.C.; Jacobs, B.L. The E3L gene of vaccinia virus encodes an inhibitor of the interferon-induced, double-stranded RNA-dependent protein kinase. Proc. Natl. Acad. Sci. USA 1992, 89, 4825–4829. [Google Scholar] [CrossRef]
- Deng, L.; Dai, P.; Parikh, T.; Cao, H.; Bhoj, V.; Sun, Q.; Chen, Z.; Merghoub, T.; Houghton, A.; Shuman, S. Vaccinia Virus Subverts a Mitochondrial Antiviral Signaling Protein-Dependent Innate Immune Response in Keratinocytes through Its Double-Stranded RNA Binding Protein, E3. J. Virol. 2008, 82, 10735–10746. [Google Scholar] [CrossRef]
- Langland, J.O.; Kash, J.C.; Carter, V.; Thomas, M.J.; Katze, M.G.; Jacobs, B.L. Suppression of Proinflammatory Signal Transduction and Gene Expression by the DualNucleic Acid Binding Domains of the Vaccinia Virus E3L Proteins. J. Virol. 2006, 80, 10083–10095. [Google Scholar] [CrossRef]
- Myskiw, C.; Arsenio, J.; Van Bruggen, R.; Deschambault, Y.; Cao, J. Vaccinia Virus E3 Suppresses Expression of Diverse Cytokines through Inhibition of the PKR, NF-κB, and IRF3 Pathways. J. Virol. 2009, 83, 6757–6768. [Google Scholar] [CrossRef]
- Kim, Y.G.; Lowenhaupt, K.; Oh, D.B.; Kim, K.K.; Rich, A. Evidence that vaccinia virulence factor E3L binds to Z-DNA in vivo: Implications for development of a therapy for poxvirus infection. Proc. Natl. Acad. Sci. USA 2004, 101, 1514–1518. [Google Scholar] [CrossRef]
- Kim, Y.-G.; Muralinath, M.; Brandt, T.; Pearcy, M.; Hauns, K.; Lowenhaupt, K.; Jacobs, B.L.; Rich, A. A role for Z-DNA binding in vaccinia virus pathogenesis. Proc. Natl. Acad. Sci. USA 2003, 100, 6974–6979. [Google Scholar] [CrossRef] [PubMed]
- Koehler, H.; Cotsmire, S.; Langland, J.; Kibler, K.V.; Kalman, D.; Upton, J.W.; Mocarski, E.S.; Jacobs, B.L. Inhibition of DAI-dependent necroptosis by the Z-DNA binding domain of the vaccinia virus innate immune evasion protein, E3. Proc. Natl. Acad. Sci. USA 2017, 114, 11506–11511. [Google Scholar] [CrossRef] [PubMed]
- Arndt, W.D.; Cotsmire, S.; Trainor, K.; Harrington, H.; Hauns, K.; Kibler, K.V.; Huynh, T.P.; Jacobs, B.L. Evasion of the Innate Immune Type I Interferon System by Monkeypox Virus. J. Virol. 2015, 89, 10489–10499. [Google Scholar] [CrossRef] [PubMed]
- Gil, J.; Rullas, J.; Alcami, J.; Esteban, M. MC159L protein from the poxvirus molluscum contagiosum virus inhibits NF-kappaB activation and apoptosis induced by PKR. J. Gen. Virol. 2001, 82 Pt 12, 3027–3034. [Google Scholar] [CrossRef]
- Shors, T.; Jacobs, B.L. Complementation of Deletion of the Vaccinia Virus E3L Gene by theEscherichia coliRNase III Gene. Virology 1997, 227, 77–87. [Google Scholar] [CrossRef]
- Guerra, S.; Abaitua, F.; Martinez-Sobrido, L.; Esteban, M.; García-Sastre, A.; Rodríguez, D. Host-Range Restriction of Vaccinia Virus E3L Deletion Mutant Can Be Overcome In Vitro, but Not In Vivo, by Expression of the Influenza Virus NS1 Protein. PLoS ONE 2011, 6, e28677. [Google Scholar] [CrossRef]
- Vijaysri, S.; Talasela, L.; Mercer, A.; McInnes, C.J.; Jacobs, B.L.; Langland, J.O. The Orf virus E3L homologue is able to complement deletion of the vaccinia virus E3L gene in vitro but not in vivo. Virology 2003, 314, 305–314. [Google Scholar] [CrossRef]
- Myskiw, C.; Arsenio, J.; Hammett, C.; Van Bruggen, R.; Deschambault, Y.; Beausoleil, N.; Babiuk, S.; Cao, J. Comparative Analysis of Poxvirus Orthologues of the Vaccinia Virus E3 Protein: Modulation of Protein Kinase R Activity, Cytokine Responses, and Virus Pathogenicity. J. Virol. 2011, 85, 12280–12291. [Google Scholar] [CrossRef]
- Shors, T.; Kibler, K.; Perkins, K.B.; Seidler-Wulff, R.; Banaszak, M.P.; Jacobs, B.L. Complementation of Vaccinia Virus Deleted of the E3L Gene by Mutants of E3L. Virology 1997, 239, 269–276. [Google Scholar] [CrossRef]
- Chang, H.W.; Uribe, L.H.; Jacobs, B.L. Rescue of vaccinia virus lacking the E3L gene by mutants of E3L. J. Virol. 1995, 69, 6605–6608. [Google Scholar] [CrossRef]
- Beattie, E.; Kauffman, E.B.; Martínez, H.; Perkus, M.E.; Jacobs, B.L.; Paoletti, E.; Tartaglia, J. Host-range restriction of vaccinia virus E3L-specific deletion mutants. Virus Genes 1996, 12, 89–94. [Google Scholar] [CrossRef] [PubMed]
- Langland, J.O.; Jacobs, B.L. The Role of the PKR-Inhibitory Genes, E3L and K3L, in Determining Vaccinia Virus Host Range. Virology 2002, 299, 133–141. [Google Scholar] [CrossRef] [PubMed]
- Ludwig, H.; Mages, J.; Staib, C.; Lehmann, M.; Lang, R.; Sutter, G. Role of Viral Factor E3L in Modified Vaccinia Virus Ankara Infection of Human HeLa Cells: Regulation of the Virus Life Cycle and Identification of Differentially Expressed Host Genes. J. Virol. 2005, 79, 2584–2596. [Google Scholar] [CrossRef] [PubMed]
- Hornemann, S.; Harlin, O.; Staib, C.; Kisling, S.; Erfle, V.; Kaspers, B.; Hacker, G.; Sutter, G. Replication of Modified Vaccinia Virus Ankara in Primary Chicken Embryo Fibroblasts Requires Expression of the Interferon Resistance Gene E3L. J. Virol. 2003, 77, 8394–8407. [Google Scholar] [CrossRef]
- Rice, A.; Turner, P.C.; Embury, J.E.; Moldawer, L.L.; Baker, H.V.; Moyer, R.W. Roles of Vaccinia Virus Genes E3L and K3L and Host Genes PKR and RNase L during Intratracheal Infection of C57BL/6 Mice. J. Virol. 2010, 85, 550–567. [Google Scholar] [CrossRef]
- Rahman, M.M.; Liu, J.; Chan, W.M.; Rothenburg, S.; McFadden, G. Myxoma Virus Protein M029 Is a Dual Function Immunomodulator that Inhibits PKR and Also Conscripts RHA/DHX9 to Promote Expanded Host Tropism and Viral Replication. PLoS Pathog. 2013, 9, e1003465. [Google Scholar] [CrossRef]
- Frey, T.R.; Forsyth, K.; Sheehan, M.M.; De Haven, B.C.; Pevarnik, J.G.; Hand, E.S.; Pizzorno, M.C.; Eisenlohr, L.C.; Hersperger, A.R. Ectromelia virus lacking the E3L ortholog is replication-defective and nonpathogenic but does induce protective immunity in a mouse strain susceptible to lethal mousepox. Virology 2018, 518, 335–348. [Google Scholar] [CrossRef]
- Brandt, T.A.; Jacobs, B.L. Both Carboxy- and Amino-Terminal Domains of the Vaccinia Virus Interferon Resistance Gene, E3L, Are Required for Pathogenesis in a Mouse Model. J. Virol. 2001, 75, 850–856. [Google Scholar] [CrossRef]
- Brandt, T.; Heck, M.C.; Vijaysri, S.; Jentarra, G.M.; Cameron, J.M.; Jacobs, B.L. The N-terminal domain of the vaccinia virus E3L-protein is required for neurovirulence, but not induction of a protective immune response. Virology 2005, 333, 263–270. [Google Scholar] [CrossRef]
- Colby, C.; Duesberg, P.H.; Colby, P.H.D.C. Double-stranded RNA in Vaccinia Virus Infected Cells. Nature 1969, 222, 940–944. [Google Scholar] [CrossRef]
- Colby, C.; Jurale, C.; Kates, J.R. Mechanism of Synthesis of Vaccinia Virus Double-Stranded Ribonucleic Acid In Vivo and In Vitro. J. Virol. 1971, 7, 71–76. [Google Scholar] [CrossRef] [PubMed]
- Duesberg, P.H.; Colby, C. On the biosynthesis and structure of double-stranded RNA in vaccinia virus-infected cells. Proc. Natl. Acad. Sci. USA 1969, 64, 396–403. [Google Scholar] [CrossRef] [PubMed]
- Broyles, S.S. Vaccinia virus transcription. J. Gen. Virol. 2003, 84, 2293–2303. [Google Scholar] [CrossRef] [PubMed]
- Gantier, M.P. Processing of Double-Stranded RNA in Mammalian Cells: A Direct Antiviral Role? J. Interf. Cytokine Res. 2014, 34, 469–477. [Google Scholar] [CrossRef]
- Brennan, K.; Bowie, A.G. Activation of host pattern recognition receptors by viruses. Curr. Opin. Microbiol. 2010, 13, 503–507. [Google Scholar] [CrossRef]
- Gürtler, C.; Bowie, A.G. Innate immune detection of microbial nucleic acids. Trends Microbiol. 2013, 21, 413–420. [Google Scholar] [CrossRef]
- Arndt, W.D.; White, S.D.; Johnson, B.P.; Huynh, T.; Liao, J.; Harrington, H.; Cotsmire, S.; Kibler, K.; Langland, J.; Jacobs, B.L. Monkeypox virus induces the synthesis of less dsRNA than vaccinia virus, and is more resistant to the anti-poxvirus drug, IBT, than vaccinia virus. Virology 2016, 497, 125–135. [Google Scholar] [CrossRef]
- Frey, T.R.; Lehmann, M.; Ryan, C.M.; Pizzorno, M.; Sutter, G.; Hersperger, A.R. Ectromelia virus accumulates less double-stranded RNA compared to vaccinia virus in BS-C-1 cells. Virology 2017, 509, 98–111. [Google Scholar] [CrossRef]
- Paez, E.; Esteban, M. Resistance of vaccinia virus to interferon is related to an interference phenomenon between the virus and the interferon system. Virology 1984, 134, 12–28. [Google Scholar] [CrossRef]
- Rice, A.P.; Kerr, I.M. Interferon-mediated, double-stranded RNA-dependent protein kinase is inhibited in extracts from vaccinia virus-infected cells. J. Virol. 1984, 50, 229–236. [Google Scholar] [CrossRef]
- Whitaker-Dowling, P.; Youngner, J.S. Characterization of a specific kinase inhibitory factor produced by vaccinia virus which inhibits the interferon-induced protein kinase. Virology 1984, 137, 171–181. [Google Scholar] [CrossRef]
- Romano, P.R.; Zhang, F.; Tan, S.-L.; Garcia-Barrio, M.T.; Katze, M.G.; Dever, T.E.; Hinnebusch, A.G. Inhibition of Double-Stranded RNA-Dependent Protein Kinase PKR by Vaccinia Virus E3: Role of Complex Formation and the E3 N-Terminal Domain. Mol. Cell. Biol. 1998, 18, 7304–7316. [Google Scholar] [CrossRef] [PubMed]
- Dueck, K.J.; Hu, Y.S.; Chen, P.; Deschambault, Y.; Lee, J.; Varga, J.; Cao, J. Mutational Analysis of Vaccinia Virus E3 Protein: The Biological Functions Do Not Correlate with Its Biochemical Capacity To Bind Double-Stranded RNA. J. Virol. 2015, 89, 5382–5394. [Google Scholar] [CrossRef] [PubMed]
- Meurs, E.; Chong, K.; Galabru, J.; Thomas, N.B.; Kerr, I.M.; Williams, B.R.; Hovanessian, A.G. Molecular cloning and characterization of the human double-stranded RNA-activated protein kinase induced by interferon. Cell 1990, 62, 379–390. [Google Scholar] [CrossRef]
- Metz, D.H.; Esteban, M. Interferon inhibits Viral Protein Synthesis in L Cells infected with Vaccinia Virus. Nature 1972, 238, 385–388. [Google Scholar] [CrossRef]
- Friedman, R.M.; Metz, D.H.; Esteban, R.M.; Tovell, D.R.; Ball, L.A.; Kerr, I.M. Mechanism of Interferon Action: Inhibition of Viral Messenger Ribonucleic Acid Translation in L-Cell Extracts. J. Virol. 1972, 10, 1184–1198. [Google Scholar] [CrossRef]
- Davies, M.V.; Chang, H.W.; Jacobs, B.L.; Kaufman, R.J. The E3L and K3L vaccinia virus gene products stimulate translation through inhibition of the double-stranded RNA-dependent protein kinase by different mechanisms. J. Virol. 1993, 67, 1688–1692. [Google Scholar] [CrossRef]
- Sharp, T.; Moonan, F.; Romashko, A.; Joshi, B.; Barber, G.; Jagus, R. The Vaccinia Virus E3L Gene Product Interacts with both the Regulatory and the Substrate Binding Regions of PKR: Implications for PKR Autoregulation. Virology 1998, 250, 302–315. [Google Scholar] [CrossRef]
- Wu, S.; Kaufman, R.J. A Model for the Double-stranded RNA (dsRNA)-dependent Dimerization and Activation of the dsRNA-activated Protein Kinase PKR. J. Biol. Chem. 1997, 272, 1291–1296. [Google Scholar] [CrossRef]
- Bou-Nader, C.; Gordon, J.M.; Henderson, F.E.; Zhang, J. The search for a PKR code-differential regulation of protein kinase R activity by diverse RNA and protein regulators. RNA 2019, 25, 539–556. [Google Scholar] [CrossRef]
- Gil, J.; Alcamí, J.; Esteban, M. Induction of Apoptosis by Double-Stranded-RNA-Dependent Protein Kinase (PKR) Involves the α Subunit of Eukaryotic Translation Initiation Factor 2 and NF-κB. Mol. Cell. Biol. 1999, 19, 4653–4663. [Google Scholar] [CrossRef] [PubMed]
- Gil, J.; Alcamí, J.; Esteban, M. Activation of NF-κB by the dsRNA-dependent protein kinase, PKR involves the IκB kinase complex. Oncogene 2000, 19, 1369–1378. [Google Scholar] [CrossRef] [PubMed]
- Gil, J.; Rullas, J.; Garcia, M.A.; Alcami, J.; Esteban, M. The catalytic activity of dsRNA-dependent protein kinase, PKR, is required for NF-κB activation. Oncogene 2001, 20, 385–394. [Google Scholar] [CrossRef] [PubMed]
- Dzananovic, E.; McKenna, S.; Patel, T.R. Viral proteins targeting host protein kinase R to evade an innate immune response: A mini review. Biotechnol. Genet. Eng. Rev. 2018, 34, 33–59. [Google Scholar] [CrossRef]
- Garcia, M.A.; Meurs, E.F.; Esteban, M. The dsRNA protein kinase PKR: Virus and cell control. Biochimie 2007, 89, 799–811. [Google Scholar] [CrossRef]
- Langland, J.O.; Cameron, J.M.; Heck, M.C.; Jancovich, J.K.; Jacobs, B.L. Inhibition of PKR by RNA and DNA viruses. Virus Res. 2006, 119, 100–110. [Google Scholar] [CrossRef]
- Zhang, P.; Jacobs, B.L.; Samuel, C.E. Loss of Protein Kinase PKR Expression in Human HeLa Cells Complements the Vaccinia Virus E3L Deletion Mutant Phenotype by Restoration of Viral Protein Synthesis. J. Virol. 2007, 82, 840–848. [Google Scholar] [CrossRef]
- Liem, J.; Liu, J. Stress Beyond Translation: Poxviruses and More. Viruses 2016, 8, 169. [Google Scholar] [CrossRef]
- Perdiguero, B.; Esteban, M. The Interferon System and Vaccinia Virus Evasion Mechanisms. J. Interf. Cytokine Res. 2009, 29, 581–598. [Google Scholar] [CrossRef]
- Pham, A.M.; Maria, F.G.S.; Lahiri, T.; Friedman, E.; Marie, I.; Levy, D.E. PKR Transduces MDA5-Dependent Signals for Type I IFN Induction. PLoS Pathog. 2016, 12, e1005489. [Google Scholar] [CrossRef]
- Liu, R.; Moss, B. Opposing Roles of Double-Stranded RNA Effector Pathways and Viral Defense Proteins Revealed with CRISPR-Cas9 Knockout Cell Lines and Vaccinia Virus Mutants. J. Virol. 2016, 90, 7864–7879. [Google Scholar] [CrossRef] [PubMed]
- Peng, C.; Haller, S.L.; Rahman, M.M.; McFadden, G.; Rothenburg, S. Myxoma virus M156 is a specific inhibitor of rabbit PKR but contains a loss-of-function mutation in Australian virus isolates. Proc. Natl. Acad. Sci. USA 2016, 113, 3855–3860. [Google Scholar] [CrossRef] [PubMed]
- Smith, G.L.; Talbot-Cooper, C.; Lu, Y. How Does Vaccinia Virus Interfere With Interferon? Adv. Virus Res. 2018, 100, 355–378. [Google Scholar] [PubMed]
- Wang, F.; Ma, Y.; Barrett, J.W.; Gao, X.; Loh, J.; Barton, E.; Virgin, H.W.; McFadden, G.; Iv, H.W.V. Disruption of Erk-dependent type I interferon induction breaks the myxoma virus species barrier. Nat. Immunol. 2004, 5, 1266–1274. [Google Scholar] [CrossRef] [PubMed]
- Wang, F.; Gao, X.; Barrett, J.W.; Shao, Q.; Bartee, E.; Mohamed, M.R.; Rahman, M.; Werden, S.; Irvine, T.; Cao, J.; et al. RIG-I Mediates the Co-Induction of Tumor Necrosis Factor and Type I Interferon Elicited by Myxoma Virus in Primary Human Macrophages. PLoS Pathog. 2008, 4, e1000099. [Google Scholar] [CrossRef]
- Wang, F.; Barrett, J.W.; Ma, Y.; Dekaban, G.A.; McFadden, G. Induction of alpha/beta interferon by myxoma virus is selectively abrogated when primary mouse embryo fibroblasts become immortalized. J. Virol. 2009, 83, 5928–5932. [Google Scholar] [CrossRef]
- Rahman, M.M.; McFadden, G. Myxoma Virus dsRNA Binding Protein M029 Inhibits the Type I IFN-Induced Antiviral State in a Highly Species-Specific Fashion. Viruses 2017, 9, 27. [Google Scholar] [CrossRef]
- Liu, R.; Olano, L.R.; Mirzakhanyan, Y.; Gershon, P.D.; Moss, B. Vaccinia Virus Ankyrin-Repeat/F-Box Protein Targets Interferon-Induced IFITs for Proteasomal Degradation. Cell Rep. 2019, 29, 816–828.e6. [Google Scholar] [CrossRef]
- Guerra, S.; Cáceres, A.; Knobeloch, K.-P.; Horak, I.; Esteban, M. Vaccinia Virus E3 Protein Prevents the Antiviral Action of ISG. PLoS Pathog. 2008, 4, e1000096. [Google Scholar] [CrossRef]
- Eduardo-Correia, B.; Martinez-Romero, C.; García-Sastre, A.; Guerra, S. ISG15 Is Counteracted by Vaccinia Virus E3 Protein and Controls the Proinflammatory Response against Viral Infection. J. Virol. 2013, 88, 2312–2318. [Google Scholar] [CrossRef]
- Symons, J.A.; Alcamí, A.; Smith, G.L. Vaccinia virus encodes a soluble type I interferon receptor of novel structure and broad species specificity. Cell 1995, 81, 551–560. [Google Scholar] [CrossRef]
- Colamonici, O.R.; Domanski, P.; Sweitzer, S.M.; Larner, A.; Buller, R.M.L. Vaccinia Virus B18R Gene Encodes a Type I Interferon-binding Protein That Blocks Interferon α Transmembrane Signaling. J. Biol. Chem. 1995, 270, 15974–15978. [Google Scholar] [CrossRef] [PubMed]
- Alcami, A.; Symons, J.A.; Smith, G.L. The vaccinia virus soluble alpha/beta interferon (IFN) receptor binds to the cell surface and protects cells from the antiviral effects of IFN. J. Virol. 2000, 74, 11230–11239. [Google Scholar] [CrossRef]
- Rahman, M.M.; Bagdassarian, E.; Ali, M.A.M.; McFadden, G. Identification of host DEAD-box RNA helicases that regulate cellular tropism of oncolytic Myxoma virus in human cancer cells. Sci. Rep. 2017, 7, 15710. [Google Scholar] [CrossRef] [PubMed]
- Fullam, A.; Schröder, M. DExD/H-box RNA helicases as mediators of anti-viral innate immunity and essential host factors for viral replication. Biochim. Biophys. Acta (BBA) Bioenerg. 2013, 1829, 854–865. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.; Yuan, B.; Lu, N.; Facchinetti, V.; Liu, Y.-J. DHX9 pairs with IPS-1 to sense double-stranded RNA in myeloid dendritic cells. J. Immunol. 2011, 187, 4501–4508. [Google Scholar] [CrossRef] [PubMed]
- Ng, Y.C.; Chung, W.C.; Kang, H.R.; Cho, H.J.; Park, E.B.; Kang, S.J.; Song, M.J. A DNA-sensing-independent role of a nuclear RNA helicase, DHX9, in stimulation of NF-kappaB-mediated innate immunity against DNA virus infection. Nucleic Acids Res. 2018, 46, 9011–9026. [Google Scholar] [CrossRef] [PubMed]
- Kim, T.; Pazhoor, S.; Bao, M.; Zhang, Z.; Hanabuchi, S.; Facchinetti, V.; Bover, L.; Plumas, J.; Chaperot, L.; Qin, J.; et al. Aspartate-glutamate-alanine-histidine box motif (DEAH)/RNA helicase A helicases sense microbial DNA in human plasmacytoid dendritic cells. Proc. Natl. Acad. Sci. USA 2010, 107, 15181–15186. [Google Scholar] [CrossRef]
- Dempsey, A.; Keating, S.E.; Carty, M.; Bowie, A.G. Poxviral protein E3–altered cytokine production reveals that DExD/H-box helicase 9 controls Toll-like receptor–stimulated immune responses. J. Biol. Chem. 2018, 293, 14989–15001. [Google Scholar] [CrossRef]
- Nagata, L.P.; Irwin, C.R.; Hu, W.-G.; Evans, D.H. Vaccinia-based vaccines to biothreat and emerging viruses. Biotechnol. Genet. Eng. Rev. 2018, 34, 107–121. [Google Scholar] [CrossRef]
- Altenburg, A.; Van De Sandt, C.E.; Li, B.W.S.; MacLoughlin, R.; Fouchier, R.A.M.; Van Amerongen, G.; Volz, A.; Hendriks, R.W.; De Swart, R.L.; Sutter, G.; et al. Modified Vaccinia Virus Ankara Preferentially Targets Antigen Presenting Cells In Vitro, Ex Vivo and In Vivo. Sci. Rep. 2017, 7, 8580. [Google Scholar] [CrossRef] [PubMed]
- García-Arriaza, J.; Esteban, M. Enhancing poxvirus vectors vaccine immunogenicity. Hum. Vaccines Immunother. 2014, 10, 2235–2244. [Google Scholar] [CrossRef] [PubMed]
- Moss, B. Reflections on the early development of poxvirus vectors. Vaccine 2013, 31, 4220–4222. [Google Scholar] [CrossRef] [PubMed]
- Sánchez-Sampedro, L.; Perdiguero, B.; Mejías-Pérez, E.; García-Arriaza, J.; Di Pilato, M.; Esteban, M. The Evolution of Poxvirus Vaccines. Viruses 2015, 7, 1726–1803. [Google Scholar] [CrossRef]
- McCabe, V.J.; Spibey, N. Potential for broad-spectrum protection against feline calicivirus using an attenuated myxoma virus expressing a chimeric FCV capsid protein. Vaccine 2005, 23, 5380–5388. [Google Scholar] [CrossRef]
- McCabe, V.J.; Tarpey, I.; Spibey, N. Vaccination of cats with an attenuated recombinant myxoma virus expressing feline calicivirus capsid protein. Vaccine 2002, 20, 2454–2462. [Google Scholar] [CrossRef]
- Top, S.; Foucras, G.; Deplanche, M.; Rives, G.; Calvalido, J.; Comtet, L.; Bertagnoli, S.; Meyer, G. Myxomavirus as a vector for the immunisation of sheep: Protection study against challenge with bluetongue virus. Vaccine 2012, 30, 1609–1616. [Google Scholar] [CrossRef]
- Pignolet, B.; Boullier, S.; Gelfi, J.; Bozzetti, M.; Russo, P.; Foulon, E.; Meyer, G.; Delverdier, M.; Foucras, G.; Bertagnoli, S. Safety and immunogenicity of myxoma virus as a new viral vector for small ruminants. J. Gen. Virol. 2008, 89, 1371–1379. [Google Scholar] [CrossRef]
- Bárcena, J.; Morales, M.; Vázquez, B.; Boga, J.A.; Parra, F.; Lucientes, J.; Pagès-Manté, A.; Sánchez-Vizcaíno, J.; Blasco, R.; Torres, J.M. Horizontal Transmissible Protection against Myxomatosis and Rabbit Hemorrhagic Disease by Using a Recombinant Myxoma Virus. J. Virol. 2000, 74, 1114–1123. [Google Scholar] [CrossRef]
- Jentarra, G.M.; Heck, M.C.; Youn, J.W.; Kibler, K.; Langland, J.O.; Baskin, C.R.; Ananieva, O.; Chang, Y.; Jacobs, B.L. Vaccinia viruses with mutations in the E3L gene as potential replication-competent, attenuated vaccines: Scarification vaccination. Vaccine 2008, 26, 2860–2872. [Google Scholar] [CrossRef]
- Vijaysri, S.; Jentarra, G.; Heck, M.C.; Mercer, A.; McInnes, C.J.; Jacobs, B.L. Vaccinia viruses with mutations in the E3L gene as potential replication-competent, attenuated vaccines: Intra-nasal vaccination. Vaccine 2008, 26, 664–676. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Denzler, K.L.; Rice, A.; MacNeill, A.L.; Fukushima, N.; Lindsey, S.F.; Wallace, G.; Burrage, A.M.; Smith, A.J.; Manning, B.R.; Swetnam, D.M.; et al. The NYCBH vaccinia virus deleted for the innate immune evasion gene, E3L, protects rabbits against lethal challenge by rabbitpox virus. Vaccine 2011, 29, 7659–7669. [Google Scholar] [CrossRef] [PubMed]
- Denzler, K.L.; Babas, T.; Rippeon, A.; Huynh, T.; Fukushima, N.; Rhodes, L.; Silvera, P.M.; Jacobs, B.L. Attenuated NYCBH vaccinia virus deleted for the E3L gene confers partial protection against lethal monkeypox virus disease in cynomolgus macaques. Vaccine 2011, 29, 9684–9690. [Google Scholar] [CrossRef] [PubMed]
- Dai, P.; Cao, H.; Merghoub, T.; Avogadri, F.; Wang, W.; Parikh, T.; Fang, C.M.; Pitha, P.M.; Fitzgerald, K.A.; Rahman, M.M.; et al. Myxoma virus induces type I interferon production in murine plasmacytoid dendritic cells via a TLR9/MyD88-, IRF5/IRF7-, and IFNAR-dependent pathway. J. Virol. 2011, 85, 10814–10825. [Google Scholar] [CrossRef]
- Cao, H.; Dai, P.; Wang, W.; Li, H.; Yuan, J.; Wang, F.; Fang, C.-M.; Pitha, P.M.; Liu, J.; Condit, R.C.; et al. Innate Immune Response of Human Plasmacytoid Dendritic Cells to Poxvirus Infection Is Subverted by Vaccinia E3 via Its Z-DNA/RNA Binding Domain. PLoS ONE 2012, 7, e36823. [Google Scholar] [CrossRef]
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).