Hepatitis C Virus Vaccine: Challenges and Prospects
Abstract
1. Introduction
2. Host Immune Responses to Hepatitis C Virus
2.1. Innate Immune Reponses
2.2. Ceullar Immune Responses
2.3. Humoral Immune Responses
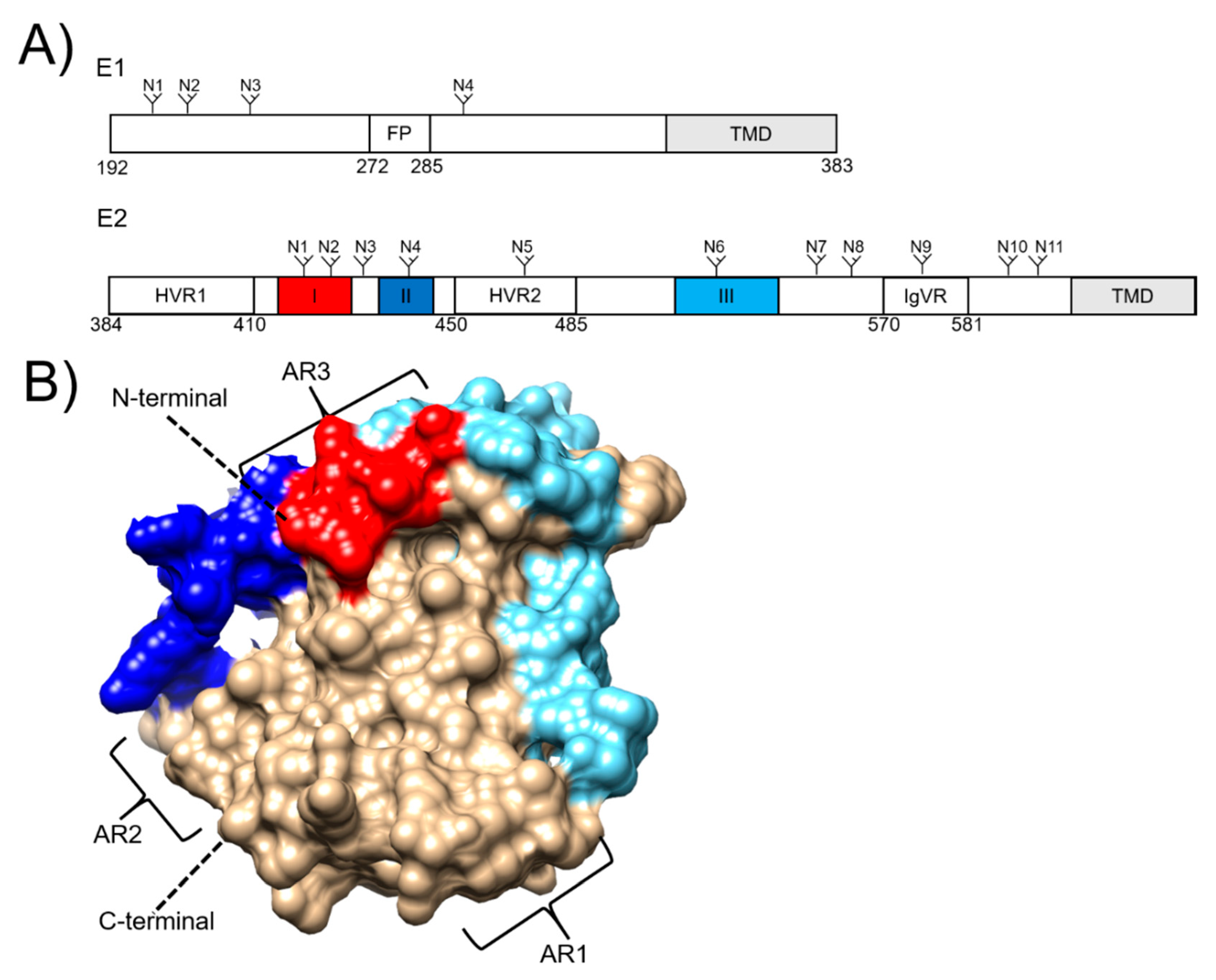
3. Hepatitis C Virus Envelope Proteins as Vaccine Targets
4. Challenges to Hepatitis C Virus Vaccine Design
4.1. Genetic Diversity
4.2. Evading the Host Adaptive Immune Response
4.3. Models for Hepatitis C Virus Infection
4.3.1. In Vitro Models
4.3.2. In Vivo Models
4.4. Rational Design of Immunogens to Elicit Cross-Reactive Neutralising Antibodies
5. Vaccine Prospects
5.1. Recombinant Subunit Vaccines
5.2. Virus-Like Particles
5.3. Viral Vector Vaccines
6. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Choo, Q.L.; Kuo, G.; Weiner, A.J.; Overby, L.R.; Bradley, D.W.; Houghton, M. Isolation of a cDNA clone derived from a blood-borne non-A, non-B viral hepatitis genome. Science 1989, 244, 359–362. [Google Scholar] [CrossRef]
- Collaborators, H. Global prevalence and genotype distribution of hepatitis C virus infection in 2015: A modelling study. Lancet Gastroenterol. Hepatol. 2017, 2, 161–176. [Google Scholar] [CrossRef]
- Ly, K.N.; Xing, J.; Klevens, R.M.; Jiles, R.B.; Holmberg, S.D. Causes of death and characteristics of decedents with viral hepatitis, United States, 2010. Clin. Infect. Dis. 2014, 58, 40–49. [Google Scholar] [CrossRef]
- Joshi, D.; Pinzani, M.; Carey, I.; Agarwal, K. Recurrent HCV after liver transplantation-mechanisms, assessment and therapy. Nat. Rev. Gastroenterol. Hepatol. 2014, 11, 710–721. [Google Scholar] [CrossRef] [PubMed]
- Mitchell, O.; Gurakar, A. Management of Hepatitis C Post-liver Transplantation: A Comprehensive Review. J. Clin. Transl. Hepatol. 2015, 3, 140–148. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Liang, T.J.; Ghany, M.G. Current and future therapies for hepatitis C virus infection. N. Engl. J. Med. 2013, 368, 1907–1917. [Google Scholar] [CrossRef] [PubMed]
- Falade-Nwulia, O.; Suarez-Cuervo, C.; Nelson, D.R.; Fried, M.W.; Segal, J.B.; Sulkowski, M.S. Oral Direct-Acting Agent Therapy for Hepatitis C Virus Infection: A Systematic Review. Ann. Intern. Med. 2017, 166, 637–648. [Google Scholar] [CrossRef] [PubMed]
- Chen, Q.; Ayer, T.; Bethea, E.; Kanwal, F.; Wang, X.; Roberts, M.; Zhuo, Y.; Fagiuoli, S.; Petersen, J.; Chhatwal, J. Changes in hepatitis C burden and treatment trends in Europe during the era of direct-acting antivirals: A modelling study. BMJ Open 2019, 9, e026726. [Google Scholar] [CrossRef]
- Rong, L.; Dahari, H.; Ribeiro, R.M.; Perelson, A.S. Rapid emergence of protease inhibitor resistance in hepatitis C virus. Sci. Transl. Med. 2010, 2, 30–32. [Google Scholar] [CrossRef]
- Bagaglio, S.; Uberti-Foppa, C.; Morsica, G. Resistance Mechanisms in Hepatitis C Virus: Implications for Direct-Acting Antiviral Use. Drugs 2017, 77, 1043–1055. [Google Scholar] [CrossRef]
- Raj, V.S.; Hundie, G.B.; Schurch, A.C.; Smits, S.L.; Pas, S.D.; Le Pogam, S.; Janssen, H.L.A.; de Knegt, R.J.; Osterhaus, A.; Najera, I.; et al. Identification of HCV Resistant Variants against Direct Acting Antivirals in Plasma and Liver of Treatment Naive Patients. Sci. Rep. 2017, 7, 4688. [Google Scholar] [CrossRef]
- Sarrazin, C.; Zeuzem, S. Resistance to direct antiviral agents in patients with hepatitis C virus infection. Gastroenterology 2010, 138, 447–462. [Google Scholar] [CrossRef]
- McCown, M.F.; Rajyaguru, S.; Kular, S.; Cammack, N.; Najera, I. GT-1a or GT-1b subtype-specific resistance profiles for hepatitis C virus inhibitors telaprevir and HCV-796. Antimicrob. Agents Chemother. 2009, 53, 2129–2132. [Google Scholar] [CrossRef] [PubMed]
- Svarovskaia, E.S.; Dvory-Sobol, H.; Parkin, N.; Hebner, C.; Gontcharova, V.; Martin, R.; Ouyang, W.; Han, B.; Xu, S.; Ku, K.; et al. Infrequent development of resistance in genotype 1-6 hepatitis C virus-infected subjects treated with sofosbuvir in phase 2 and 3 clinical trials. Clin. Infect. Dis. 2014, 59, 1666–1674. [Google Scholar] [CrossRef] [PubMed]
- Vranjkovic, A.; Deonarine, F.; Kaka, S.; Angel, J.B.; Cooper, C.L.; Crawley, A.M. Direct-Acting Antiviral Treatment of HCV Infection Does Not Resolve the Dysfunction of Circulating CD8(+) T-Cells in Advanced Liver Disease. Front. Immunol. 2019, 10, 1926. [Google Scholar] [CrossRef]
- Conti, F.; Buonfiglioli, F.; Scuteri, A.; Crespi, C.; Bolondi, L.; Caraceni, P.; Foschi, F.G.; Lenzi, M.; Mazzella, G.; Verucchi, G.; et al. Early occurrence and recurrence of hepatocellular carcinoma in HCV-related cirrhosis treated with direct-acting antivirals. J. Hepatol. 2016, 65, 727–733. [Google Scholar] [CrossRef]
- Kanwal, F.; Kramer, J.; Asch, S.M.; Chayanupatkul, M.; Cao, Y.; El-Serag, H.B. Risk of Hepatocellular Cancer in HCV Patients Treated With Direct-Acting Antiviral Agents. Gastroenterology 2017, 153, 996–1005.e1001. [Google Scholar] [CrossRef]
- Major, M.; Gutfraind, A.; Shekhtman, L.; Cui, Q.; Kachko, A.; Cotler, S.J.; Hajarizadeh, B.; Sacks-Davis, R.; Page, K.; Boodram, B.; et al. Modeling of patient virus titers suggests that availability of a vaccine could reduce hepatitis C virus transmission among injecting drug users. Sci. Transl. Med. 2018, 10. [Google Scholar] [CrossRef]
- Micallef, J.M.; Kaldor, J.M.; Dore, G.J. Spontaneous viral clearance following acute hepatitis C infection: A systematic review of longitudinal studies. J. Viral. Hepat. 2006, 13, 34–41. [Google Scholar] [CrossRef]
- Poustchi, H.; Esmaili, S.; Mohamadkhani, A.; Nikmahzar, A.; Pourshams, A.; Sepanlou, S.G.; Merat, S.; Malekzadeh, R. The impact of illicit drug use on spontaneous hepatitis C clearance: Experience from a large cohort population study. PLoS ONE 2011, 6, e23830. [Google Scholar] [CrossRef]
- Mosley, J.W.; Operskalski, E.A.; Tobler, L.H.; Buskell, Z.J.; Andrews, W.W.; Phelps, B.; Dockter, J.; Giachetti, C.; Seeff, L.B.; Busch, M.P. The course of hepatitis C viraemia in transfusion recipients prior to availability of antiviral therapy. J. Viral. Hepat. 2008, 15, 120–128. [Google Scholar] [CrossRef] [PubMed]
- Page, K.; Hahn, J.A.; Evans, J.; Shiboski, S.; Lum, P.; Delwart, E.; Tobler, L.; Andrews, W.; Avanesyan, L.; Cooper, S.; et al. Acute hepatitis C virus infection in young adult injection drug users: A prospective study of incident infection, resolution, and reinfection. J. Infect. Dis. 2009, 200, 1216–1226. [Google Scholar] [CrossRef] [PubMed]
- Osburn, W.O.; Fisher, B.E.; Dowd, K.A.; Urban, G.; Liu, L.; Ray, S.C.; Thomas, D.L.; Cox, A.L. Spontaneous control of primary hepatitis C virus infection and immunity against persistent reinfection. Gastroenterology 2010, 138, 315–324. [Google Scholar] [CrossRef]
- Sacks-Davis, R.; Aitken, C.K.; Higgs, P.; Spelman, T.; Pedrana, A.E.; Bowden, S.; Bharadwaj, M.; Nivarthi, U.K.; Suppiah, V.; George, J.; et al. High rates of hepatitis C virus reinfection and spontaneous clearance of reinfection in people who inject drugs: A prospective cohort study. PLoS ONE 2013, 8, e80216. [Google Scholar] [CrossRef] [PubMed]
- Brown, K.S.; Keogh, M.J.; Owsianka, A.M.; Adair, R.; Patel, A.H.; Arnold, J.N.; Ball, J.K.; Sim, R.B.; Tarr, A.W.; Hickling, T.P. Specific interaction of hepatitis C virus glycoproteins with mannan binding lectin inhibits virus entry. Protein Cell 2010, 1, 664–674. [Google Scholar] [CrossRef] [PubMed]
- Hamed, M.R.; Brown, R.J.; Zothner, C.; Urbanowicz, R.A.; Mason, C.P.; Krarup, A.; McClure, C.P.; Irving, W.L.; Ball, J.K.; Harris, M.; et al. Recombinant human L-ficolin directly neutralizes hepatitis C virus entry. J. Innate Immun. 2014, 6, 676–684. [Google Scholar] [CrossRef] [PubMed]
- Xu, Y.; Zhong, J. Innate immunity against hepatitis C virus. Curr. Opin. Immunol. 2016, 42, 98–104. [Google Scholar] [CrossRef]
- Chigbu, D.I.; Loonawat, R.; Sehgal, M.; Patel, D.; Jain, P. Hepatitis C Virus Infection: Host(-)Virus Interaction and Mechanisms of Viral Persistence. Cells 2019, 8, 376. [Google Scholar] [CrossRef]
- Wong, M.T.; Chen, S.S. Emerging roles of interferon-stimulated genes in the innate immune response to hepatitis C virus infection. Cell Mol. Immunol. 2016, 13, 11–35. [Google Scholar] [CrossRef] [PubMed]
- Schoggins, J.W.; Wilson, S.J.; Panis, M.; Murphy, M.Y.; Jones, C.T.; Bieniasz, P.; Rice, C.M. A diverse range of gene products are effectors of the type I interferon antiviral response. Nature 2011, 472, 481–485. [Google Scholar] [CrossRef]
- Patra, T.; Ray, R.B.; Ray, R. Strategies to Circumvent Host Innate Immune Response by Hepatitis C Virus. Cells 2019, 8, 274. [Google Scholar] [CrossRef] [PubMed]
- Heim, M.H. Innate immunity and HCV. J. Hepatol. 2013, 58, 564–574. [Google Scholar] [CrossRef] [PubMed]
- Thomas, D.L.; Thio, C.L.; Martin, M.P.; Qi, Y.; Ge, D.; O’Huigin, C.; Kidd, J.; Kidd, K.; Khakoo, S.I.; Alexander, G.; et al. Genetic variation in IL28B and spontaneous clearance of hepatitis C virus. Nature 2009, 461, 798–801. [Google Scholar] [CrossRef] [PubMed]
- Ge, D.; Fellay, J.; Thompson, A.J.; Simon, J.S.; Shianna, K.V.; Urban, T.J.; Heinzen, E.L.; Qiu, P.; Bertelsen, A.H.; Muir, A.J.; et al. Genetic variation in IL28B predicts hepatitis C treatment-induced viral clearance. Nature 2009, 461, 399–401. [Google Scholar] [CrossRef] [PubMed]
- Khakoo, S.I.; Thio, C.L.; Martin, M.P.; Brooks, C.R.; Gao, X.; Astemborski, J.; Cheng, J.; Goedert, J.J.; Vlahov, D.; Hilgartner, M.; et al. HLA and NK cell inhibitory receptor genes in resolving hepatitis C virus infection. Science 2004, 305, 872–874. [Google Scholar] [CrossRef]
- Amadei, B.; Urbani, S.; Cazaly, A.; Fisicaro, P.; Zerbini, A.; Ahmed, P.; Missale, G.; Ferrari, C.; Khakoo, S.I. Activation of natural killer cells during acute infection with hepatitis C virus. Gastroenterology 2010, 138, 1536–1545. [Google Scholar] [CrossRef]
- Yoon, J.C.; Yang, C.M.; Song, Y.; Lee, J.M. Natural killer cells in hepatitis C: Current progress. World J. Gastroenterol. 2016, 22, 1449–1460. [Google Scholar] [CrossRef]
- Missale, G.; Bertoni, R.; Lamonaca, V.; Valli, A.; Massari, M.; Mori, C.; Rumi, M.G.; Houghton, M.; Fiaccadori, F.; Ferrari, C. Different clinical behaviors of acute hepatitis C virus infection are associated with different vigor of the anti-viral cell-mediated immune response. J. Clin. Investig. 1996, 98, 706–714. [Google Scholar] [CrossRef]
- Mittrucker, H.W.; Visekruna, A.; Huber, M. Heterogeneity in the differentiation and function of CD8(+) T cells. Arch. Immunol. Ther. Exp. (Warsz) 2014, 62, 449–458. [Google Scholar] [CrossRef]
- Luckheeram, R.V.; Zhou, R.; Verma, A.D.; Xia, B. CD4(+)T cells: Differentiation and functions. Clin. Dev. Immunol. 2012, 2012, 925135. [Google Scholar] [CrossRef]
- Laidlaw, B.J.; Craft, J.E.; Kaech, S.M. The multifaceted role of CD4(+) T cells in CD8(+) T cell memory. Nat. Rev. Immunol. 2016, 16, 102–111. [Google Scholar] [CrossRef] [PubMed]
- Swain, S.L.; McKinstry, K.K.; Strutt, T.M. Expanding roles for CD4(+) T cells in immunity to viruses. Nat. Rev. Immunol. 2012, 12, 136–148. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Liu, W.; Wen, B.; Xie, T.; Tang, P.; Hu, Y.; Huang, L.; Jin, K.; Zhang, P.; Liu, Z.; et al. Circulating CXCR3(+) Tfh cells positively correlate with neutralizing antibody responses in HCV-infected patients. Sci. Rep. 2019, 9, 10090. [Google Scholar] [CrossRef] [PubMed]
- Semmo, N.; Klenerman, P. CD4+ T cell responses in hepatitis C virus infection. World J. Gastroenterol. 2007, 13, 4831–4838. [Google Scholar] [CrossRef]
- Schulze Zur Wiesch, J.; Ciuffreda, D.; Lewis-Ximenez, L.; Kasprowicz, V.; Nolan, B.E.; Streeck, H.; Aneja, J.; Reyor, L.L.; Allen, T.M.; Lohse, A.W.; et al. Broadly directed virus-specific CD4+ T cell responses are primed during acute hepatitis C infection, but rapidly disappear from human blood with viral persistence. J. Exp. Med. 2012, 209, 61–75. [Google Scholar] [CrossRef]
- Ulsenheimer, A.; Gerlach, J.T.; Gruener, N.H.; Jung, M.C.; Schirren, C.A.; Schraut, W.; Zachoval, R.; Pape, G.R.; Diepolder, H.M. Detection of functionally altered hepatitis C virus-specific CD4 T cells in acute and chronic hepatitis C. Hepatology 2003, 37, 1189–1198. [Google Scholar] [CrossRef]
- Dustin, L.B. Innate and Adaptive Immune Responses in Chronic HCV Infection. Curr. Drug Targets 2017, 18, 826–843. [Google Scholar] [CrossRef]
- Grakoui, A.; Shoukry, N.H.; Woollard, D.J.; Han, J.H.; Hanson, H.L.; Ghrayeb, J.; Murthy, K.K.; Rice, C.M.; Walker, C.M. HCV persistence and immune evasion in the absence of memory T cell help. Science 2003, 302, 659–662. [Google Scholar] [CrossRef]
- Shoukry, N.H.; Grakoui, A.; Houghton, M.; Chien, D.Y.; Ghrayeb, J.; Reimann, K.A.; Walker, C.M. Memory CD8+ T cells are required for protection from persistent hepatitis C virus infection. J. Exp. Med. 2003, 197, 1645–1655. [Google Scholar] [CrossRef]
- Farci, P.; Alter, H.J.; Wong, D.C.; Miller, R.H.; Govindarajan, S.; Engle, R.; Shapiro, M.; Purcell, R.H. Prevention of hepatitis C virus infection in chimpanzees after antibody-mediated in vitro neutralization. Proc. Natl. Acad. Sci. USA 1994, 91, 7792–7796. [Google Scholar] [CrossRef]
- Grebely, J.; Prins, M.; Hellard, M.; Cox, A.L.; Osburn, W.O.; Lauer, G.; Page, K.; Lloyd, A.R.; Dore, G.J. Hepatitis C virus clearance, reinfection, and persistence, with insights from studies of injecting drug users: Towards a vaccine. Lancet Infect. Dis. 2012, 12, 408–414. [Google Scholar] [CrossRef]
- Zeremski, M.; Shu, M.A.; Brown, Q.; Wu, Y.; Des Jarlais, D.C.; Busch, M.P.; Talal, A.H.; Edlin, B.R. Hepatitis C virus-specific T-cell immune responses in seronegative injection drug users. J. Viral. Hepat. 2009, 16, 10–20. [Google Scholar] [CrossRef] [PubMed]
- Post, J.J.; Pan, Y.; Freeman, A.J.; Harvey, C.E.; White, P.A.; Palladinetti, P.; Haber, P.S.; Marinos, G.; Levy, M.H.; Kaldor, J.M.; et al. Clearance of hepatitis C viremia associated with cellular immunity in the absence of seroconversion in the hepatitis C incidence and transmission in prisons study cohort. J. Infect. Dis. 2004, 189, 1846–1855. [Google Scholar] [CrossRef]
- Strasak, A.M.; Kim, A.Y.; Lauer, G.M.; de Sousa, P.S.; Ginuino, C.F.; Fernandes, C.A.; Velloso, C.E.; de Almeida, A.J.; de Oliveira, J.M.; Yoshida, C.F.; et al. Antibody dynamics and spontaneous viral clearance in patients with acute hepatitis C infection in Rio de Janeiro, Brazil. BMC Infect. Dis. 2011, 11, 15. [Google Scholar] [CrossRef] [PubMed]
- Pestka, J.M.; Zeisel, M.B.; Bläser, E.; Schürmann, P.; Bartosch, B.; Cosset, F.L.; Patel, A.H.; Meisel, H.; Baumert, J.; Viazov, S.; et al. Rapid induction of virus-neutralizing antibodies and viral clearance in a single-source outbreak of hepatitis C. Proc. Natl. Acad. Sci. USA 2007, 104, 6025–6030. [Google Scholar] [CrossRef] [PubMed]
- Dowd, K.A.; Netski, D.M.; Wang, X.H.; Cox, A.L.; Ray, S.C. Selection pressure from neutralizing antibodies drives sequence evolution during acute infection with hepatitis C virus. Gastroenterology 2009, 136, 2377–2386. [Google Scholar] [CrossRef]
- Osburn, W.O.; Snider, A.E.; Wells, B.L.; Latanich, R.; Bailey, J.R.; Thomas, D.L.; Cox, A.L.; Ray, S.C. Clearance of hepatitis C infection is associated with the early appearance of broad neutralizing antibody responses. Hepatology 2014, 59, 2140–2151. [Google Scholar] [CrossRef]
- Walker, M.R.; Leung, P.; Eltahla, A.A.; Underwood, A.; Abayasingam, A.; Brasher, N.A.; Li, H.; Wu, B.R.; Maher, L.; Luciani, F.; et al. Clearance of hepatitis C virus is associated with early and potent but narrowly-directed, Envelope-specific antibodies. Sci. Rep. 2019, 9, 13300. [Google Scholar] [CrossRef]
- Kinchen, V.J.; Zahid, M.N.; Flyak, A.I.; Soliman, M.G.; Learn, G.H.; Wang, S.; Davidson, E.; Doranz, B.J.; Ray, S.C.; Cox, A.L.; et al. Broadly Neutralizing Antibody Mediated Clearance of Human Hepatitis C Virus Infection. Cell Host Microbe 2018, 24, 717–730.e715. [Google Scholar] [CrossRef]
- Gu, J.; Hardy, J.; Boo, I.; Vietheer, P.; McCaffrey, K.; Alhammad, Y.; Chopra, A.; Gaudieri, S.; Poumbourios, P.; Coulibaly, F.; et al. Escape of Hepatitis C Virus from Epitope I Neutralization Increases Sensitivity of Other Neutralization Epitopes. J. Virol. 2018, 92. [Google Scholar] [CrossRef]
- Hadlock, K.G.; Lanford, R.E.; Perkins, S.; Rowe, J.; Yang, Q.; Levy, S.; Pileri, P.; Abrignani, S.; Foung, S.K. Human monoclonal antibodies that inhibit binding of hepatitis C virus E2 protein to CD81 and recognize conserved conformational epitopes. J. Virol. 2000, 74, 10407–10416. [Google Scholar] [CrossRef] [PubMed]
- Allander, T.; Drakenberg, K.; Beyene, A.; Rosa, D.; Abrignani, S.; Houghton, M.; Widell, A.; Grillner, L.; Persson, M.A. Recombinant human monoclonal antibodies against different conformational epitopes of the E2 envelope glycoprotein of hepatitis C virus that inhibit its interaction with CD81. J. Gen. Virol. 2000, 81, 2451–2459. [Google Scholar] [CrossRef] [PubMed]
- Johansson, D.X.; Voisset, C.; Tarr, A.W.; Aung, M.; Ball, J.K.; Dubuisson, J.; Persson, M.A. Human combinatorial libraries yield rare antibodies that broadly neutralize hepatitis C virus. Proc. Natl. Acad. Sci. USA 2007, 104, 16269–16274. [Google Scholar] [CrossRef] [PubMed]
- Law, M.; Maruyama, T.; Lewis, J.; Giang, E.; Tarr, A.W.; Stamataki, Z.; Gastaminza, P.; Chisari, F.V.; Jones, I.M.; Fox, R.I.; et al. Broadly neutralizing antibodies protect against hepatitis C virus quasispecies challenge. Nat. Med. 2008, 14, 25–27. [Google Scholar] [CrossRef] [PubMed]
- Swann, R.E.; Cowton, V.M.; Robinson, M.W.; Cole, S.J.; Barclay, S.T.; Mills, P.R.; Thomson, E.C.; McLauchlan, J.; Patel, A.H. Broad Anti-Hepatitis C Virus (HCV) Antibody Responses Are Associated with Improved Clinical Disease Parameters in Chronic HCV Infection. J. Virol. 2016, 90, 4530–4543. [Google Scholar] [CrossRef]
- Bjoro, K.; Froland, S.S.; Yun, Z.; Samdal, H.H.; Haaland, T. Hepatitis C infection in patients with primary hypogammaglobulinemia after treatment with contaminated immune globulin. N. Engl. J. Med. 1994, 331, 1607–1611. [Google Scholar] [CrossRef]
- Bailey, J.R.; Flyak, A.I.; Cohen, V.J.; Li, H.; Wasilewski, L.N.; Snider, A.E.; Wang, S.; Learn, G.H.; Kose, N.; Loerinc, L.; et al. Broadly neutralizing antibodies with few somatic mutations and hepatitis C virus clearance. JCI Insight 2017, 2. [Google Scholar] [CrossRef]
- Chan, C.H.; Hadlock, K.G.; Foung, S.K.; Levy, S. V(H)1-69 gene is preferentially used by hepatitis C virus-associated B cell lymphomas and by normal B cells responding to the E2 viral antigen. Blood 2001, 97, 1023–1026. [Google Scholar] [CrossRef]
- Giang, E.; Dorner, M.; Prentoe, J.C.; Dreux, M.; Evans, M.J.; Bukh, J.; Rice, C.M.; Ploss, A.; Burton, D.R.; Law, M. Human broadly neutralizing antibodies to the envelope glycoprotein complex of hepatitis C virus. Proc. Natl. Acad. Sci. USA 2012, 109, 6205–6210. [Google Scholar] [CrossRef]
- Chen, F.; Tzarum, N.; Wilson, I.A.; Law, M. VH1-69 antiviral broadly neutralizing antibodies: Genetics, structures, and relevance to rational vaccine design. Curr. Opin. Virol. 2019, 34, 149–159. [Google Scholar] [CrossRef]
- Merat, S.J.; Bru, C.; van de Berg, D.; Molenkamp, R.; Tarr, A.W.; Koekkoek, S.; Kootstra, N.A.; Prins, M.; Ball, J.K.; Bakker, A.Q.; et al. Cross-genotype AR3-specific neutralizing antibodies confer long-term protection in injecting drug users after HCV clearance. J. Hepatol. 2019, 71, 14–24. [Google Scholar] [CrossRef] [PubMed]
- Flyak, A.I.; Ruiz, S.; Colbert, M.D.; Luong, T.; Crowe, J.E., Jr.; Bailey, J.R.; Bjorkman, P.J. HCV Broadly Neutralizing Antibodies Use a CDRH3 Disulfide Motif to Recognize an E2 Glycoprotein Site that Can Be Targeted for Vaccine Design. Cell Host Microbe 2018, 24, 703–716.e703. [Google Scholar] [CrossRef] [PubMed]
- Tzarum, N.; Giang, E.; Kong, L.; He, L.; Prentoe, J.; Augestad, E.; Hua, Y.; Castillo, S.; Lauer, G.M.; Bukh, J.; et al. Genetic and structural insights into broad neutralization of hepatitis C virus by human VH1-69 antibodies. Sci. Adv. 2019, 5, eaav1882. [Google Scholar] [CrossRef] [PubMed]
- Ball, J.K.; Tarr, A.W.; McKeating, J.A. The past, present and future of neutralizing antibodies for hepatitis C virus. Antiviral Res. 2014, 105, 100–111. [Google Scholar] [CrossRef]
- Vieyres, G.; Thomas, X.; Descamps, V.; Duverlie, G.; Patel, A.H.; Dubuisson, J. Characterization of the envelope glycoproteins associated with infectious hepatitis C virus. J. Virol. 2010, 84, 10159–10168. [Google Scholar] [CrossRef]
- Vieyres, G.; Dubuisson, J.; Pietschmann, T. Incorporation of hepatitis C virus E1 and E2 glycoproteins: The keystones on a peculiar virion. Viruses 2014, 6, 1149–1187. [Google Scholar] [CrossRef]
- Kong, L.; Giang, E.; Nieusma, T.; Kadam, R.U.; Cogburn, K.E.; Hua, Y.; Dai, X.; Stanfield, R.L.; Burton, D.R.; Ward, A.B.; et al. Hepatitis C virus E2 envelope glycoprotein core structure. Science 2013, 342, 1090–1094. [Google Scholar] [CrossRef]
- Khan, A.G.; Whidby, J.; Miller, M.T.; Scarborough, H.; Zatorski, A.V.; Cygan, A.; Price, A.A.; Yost, S.A.; Bohannon, C.D.; Jacob, J.; et al. Structure of the core ectodomain of the hepatitis C virus envelope glycoprotein 2. Nature 2014, 509, 381–384. [Google Scholar] [CrossRef]
- El Omari, K.; Iourin, O.; Kadlec, J.; Sutton, G.; Harlos, K.; Grimes, J.M.; Stuart, D.I. Unexpected structure for the N-terminal domain of hepatitis C virus envelope glycoprotein E1. Nat. Commun. 2014, 5, 4874. [Google Scholar] [CrossRef]
- Kong, L.; Kadam, R.U.; Giang, E.; Ruwona, T.B.; Nieusma, T.; Culhane, J.C.; Stanfield, R.L.; Dawson, P.E.; Wilson, I.A.; Law, M. Structure of Hepatitis C Virus Envelope Glycoprotein E1 Antigenic Site 314–324 in Complex with Antibody IGH526. J. Mol. Biol. 2015, 427, 2617–2628. [Google Scholar] [CrossRef]
- Bartosch, B.; Vitelli, A.; Granier, C.; Goujon, C.; Dubuisson, J.; Pascale, S.; Scarselli, E.; Cortese, R.; Nicosia, A.; Cosset, F.L. Cell entry of hepatitis C virus requires a set of co-receptors that include the CD81 tetraspanin and the SR-B1 scavenger receptor. J. Biol. Chem. 2003, 278, 41624–41630. [Google Scholar] [CrossRef]
- Pileri, P.; Uematsu, Y.; Campagnoli, S.; Galli, G.; Falugi, F.; Petracca, R.; Weiner, A.J.; Houghton, M.; Rosa, D.; Grandi, G.; et al. Binding of hepatitis C virus to CD81. Science 1998, 282, 938–941. [Google Scholar] [CrossRef] [PubMed]
- Scarselli, E.; Ansuini, H.; Cerino, R.; Roccasecca, R.M.; Acali, S.; Filocamo, G.; Traboni, C.; Nicosia, A.; Cortese, R.; Vitelli, A. The human scavenger receptor class B type I is a novel candidate receptor for the hepatitis C virus. Embo J. 2002, 21, 5017–5025. [Google Scholar] [CrossRef] [PubMed]
- Evans, M.J.; von Hahn, T.; Tscherne, D.M.; Syder, A.J.; Panis, M.; Wolk, B.; Hatziioannou, T.; McKeating, J.A.; Bieniasz, P.D.; Rice, C.M. Claudin-1 is a hepatitis C virus co-receptor required for a late step in entry. Nature 2007, 446, 801–805. [Google Scholar] [CrossRef] [PubMed]
- Ploss, A.; Evans, M.J.; Gaysinskaya, V.A.; Panis, M.; You, H.; de Jong, Y.P.; Rice, C.M. Human occludin is a hepatitis C virus entry factor required for infection of mouse cells. Nature 2009, 457, 882–886. [Google Scholar] [CrossRef] [PubMed]
- Liu, S.; Yang, W.; Shen, L.; Turner, J.R.; Coyne, C.B.; Wang, T. Tight junction proteins claudin-1 and occludin control hepatitis C virus entry and are downregulated during infection to prevent superinfection. J. Virol. 2009, 83, 2011–2014. [Google Scholar] [CrossRef]
- Dubuisson, J.; Cosset, F.L. Virology and cell biology of the hepatitis C virus life cycle: An update. J. Hepatol. 2014, 61, S3–S13. [Google Scholar] [CrossRef]
- Tscherne, D.M.; Jones, C.T.; Evans, M.J.; Lindenbach, B.D.; McKeating, J.A.; Rice, C.M. Time- and temperature-dependent activation of hepatitis C virus for low-pH-triggered entry. J. Virol. 2006, 80, 1734–1741. [Google Scholar] [CrossRef]
- Tong, Y.; Lavillette, D.; Li, Q.; Zhong, J. Role of Hepatitis C Virus Envelope Glycoprotein E1 in Virus Entry and Assembly. Front. Immunol. 2018, 9, 1411. [Google Scholar] [CrossRef]
- Drummer, H.E. Challenges to the development of vaccines to hepatitis C virus that elicit neutralizing antibodies. Front. Microbiol. 2014, 5, 329. [Google Scholar] [CrossRef]
- Keck, Z.; Wang, W.; Wang, Y.; Lau, P.; Carlsen, T.H.; Prentoe, J.; Xia, J.; Patel, A.H.; Bukh, J.; Foung, S.K. Cooperativity in virus neutralization by human monoclonal antibodies to two adjacent regions located at the amino terminus of hepatitis C virus E2 glycoprotein. J. Virol. 2013, 87, 37–51. [Google Scholar] [CrossRef] [PubMed]
- Tzarum, N.; Wilson, I.A.; Law, M. The Neutralizing Face of Hepatitis C Virus E2 Envelope Glycoprotein. Front. Immunol. 2018, 9, 1315. [Google Scholar] [CrossRef] [PubMed]
- Cuevas, J.M.; Gonzalez-Candelas, F.; Moya, A.; Sanjuan, R. Effect of ribavirin on the mutation rate and spectrum of hepatitis C virus in vivo. J. Virol. 2009, 83, 5760–5764. [Google Scholar] [CrossRef] [PubMed]
- Echeverria, N.; Moratorio, G.; Cristina, J.; Moreno, P. Hepatitis C virus genetic variability and evolution. World J. Hepatol. 2015, 7, 831–845. [Google Scholar] [CrossRef]
- Smith, D.B.; Bukh, J.; Kuiken, C.; Muerhoff, A.S.; Rice, C.M.; Stapleton, J.T.; Simmonds, P. Expanded classification of hepatitis C virus into 7 genotypes and 67 subtypes: Updated criteria and genotype assignment web resource. Hepatology 2014, 59, 318–327. [Google Scholar] [CrossRef]
- Petruzziello, A.; Marigliano, S.; Loquercio, G.; Cozzolino, A.; Cacciapuoti, C. Global epidemiology of hepatitis C virus infection: An up-date of the distribution and circulation of hepatitis C virus genotypes. World J. Gastroenterol. 2016, 22, 7824–7840. [Google Scholar] [CrossRef]
- Borgia, S.M.; Hedskog, C.; Parhy, B.; Hyland, R.H.; Stamm, L.M.; Brainard, D.M.; Subramanian, M.G.; McHutchison, J.G.; Mo, H.; Svarovskaia, E.; et al. Identification of a Novel Hepatitis C Virus Genotype From Punjab, India: Expanding Classification of Hepatitis C Virus Into 8 Genotypes. J. Infect. Dis. 2018, 218, 1722–1729. [Google Scholar] [CrossRef]
- Forns, X.; Purcell, R.H.; Bukh, J. Quasispecies in viral persistence and pathogenesis of hepatitis C virus. Trends Microbiol. 1999, 7, 402–410. [Google Scholar] [CrossRef]
- Farci, P.; Shimoda, A.; Coiana, A.; Diaz, G.; Peddis, G.; Melpolder, J.C.; Strazzera, A.; Chien, D.Y.; Munoz, S.J.; Balestrieri, A.; et al. The outcome of acute hepatitis C predicted by the evolution of the viral quasispecies. Science 2000, 288, 339–344. [Google Scholar] [CrossRef]
- Liu, L.; Fisher, B.E.; Dowd, K.A.; Astemborski, J.; Cox, A.L.; Ray, S.C. Acceleration of hepatitis C virus envelope evolution in humans is consistent with progressive humoral immune selection during the transition from acute to chronic infection. J. Virol. 2010, 84, 5067–5077. [Google Scholar] [CrossRef]
- Islam, N.; Krajden, M.; Shoveller, J.; Gustafson, P.; Gilbert, M.; Wong, J.; Tyndall, M.W.; Janjua, N.Z. Hepatitis C cross-genotype immunity and implications for vaccine development. Sci. Rep. 2017, 7, 12326. [Google Scholar] [CrossRef] [PubMed]
- Tarr, A.W.; Urbanowicz, R.A.; Hamed, M.R.; Albecka, A.; McClure, C.P.; Brown, R.J.; Irving, W.L.; Dubuisson, J.; Ball, J.K. Hepatitis C patient-derived glycoproteins exhibit marked differences in susceptibility to serum neutralizing antibodies: Genetic subtype defines antigenic but not neutralization serotype. J. Virol. 2011, 85, 4246–4257. [Google Scholar] [CrossRef] [PubMed]
- Luxenburger, H.; Grass, F.; Baermann, J.; Boettler, T.; Marget, M.; Emmerich, F.; Panning, M.; Thimme, R.; Nitschke, K.; Neumann-Haefelin, C. Differential virus-specific CD8(+) T-cell epitope repertoire in hepatitis C virus genotype 1 versus 4. J. Viral. Hepat. 2018, 25, 779–790. [Google Scholar] [CrossRef] [PubMed]
- Von Delft, A.; Humphreys, I.S.; Brown, A.; Pfafferott, K.; Lucas, M.; Klenerman, P.; Lauer, G.M.; Cox, A.L.; Gaudieri, S.; Barnes, E. The broad assessment of HCV genotypes 1 and 3 antigenic targets reveals limited cross-reactivity with implications for vaccine design. Gut 2016, 65, 112–123. [Google Scholar] [CrossRef]
- Bankwitz, D.; Steinmann, E.; Bitzegeio, J.; Ciesek, S.; Friesland, M.; Herrmann, E.; Zeisel, M.B.; Baumert, T.F.; Keck, Z.Y.; Foung, S.K.; et al. Hepatitis C virus hypervariable region 1 modulates receptor interactions, conceals the CD81 binding site, and protects conserved neutralizing epitopes. J. Virol. 2010, 84, 5751–5763. [Google Scholar] [CrossRef]
- Prentoe, J.; Jensen, T.B.; Meuleman, P.; Serre, S.B.; Scheel, T.K.; Leroux-Roels, G.; Gottwein, J.M.; Bukh, J. Hypervariable region 1 differentially impacts viability of hepatitis C virus strains of genotypes 1 to 6 and impairs virus neutralization. J. Virol. 2011, 85, 2224–2234. [Google Scholar] [CrossRef]
- Bazzill, J.D.; Ochyl, L.J.; Giang, E.; Castillo, S.; Law, M.; Moon, J.J. Interrogation of Antigen Display on Individual Vaccine Nanoparticles for Achieving Neutralizing Antibody Responses against Hepatitis C Virus. Nano. Lett. 2018, 18, 7832–7838. [Google Scholar] [CrossRef]
- Lavie, M.; Hanoulle, X.; Dubuisson, J. Glycan Shielding and Modulation of Hepatitis C Virus Neutralizing Antibodies. Front. Immunol. 2018, 9, 910. [Google Scholar] [CrossRef]
- Helle, F.; Goffard, A.; Morel, V.; Duverlie, G.; McKeating, J.; Keck, Z.Y.; Foung, S.; Penin, F.; Dubuisson, J.; Voisset, C. The neutralizing activity of anti-hepatitis C virus antibodies is modulated by specific glycans on the E2 envelope protein. J. Virol. 2007, 81, 8101–8111. [Google Scholar] [CrossRef]
- Helle, F.; Vieyres, G.; Elkrief, L.; Popescu, C.I.; Wychowski, C.; Descamps, V.; Castelain, S.; Roingeard, P.; Duverlie, G.; Dubuisson, J. Role of N-linked glycans in the functions of hepatitis C virus envelope proteins incorporated into infectious virions. J. Virol. 2010, 84, 11905–11915. [Google Scholar] [CrossRef]
- Li, D.; von Schaewen, M.; Wang, X.; Tao, W.; Zhang, Y.; Li, L.; Heller, B.; Hrebikova, G.; Deng, Q.; Ploss, A.; et al. Altered Glycosylation Patterns Increase Immunogenicity of a Subunit Hepatitis C Virus Vaccine, Inducing Neutralizing Antibodies Which Confer Protection in Mice. J. Virol. 2016, 90, 10486–10498. [Google Scholar] [CrossRef] [PubMed]
- Ren, Y.; Min, Y.Q.; Liu, M.; Chi, L.; Zhao, P.; Zhang, X.L. N-glycosylation-mutated HCV envelope glycoprotein complex enhances antigen-presenting activity and cellular and neutralizing antibody responses. Biochim. Biophys. Acta 2016, 1860, 1764–1775. [Google Scholar] [CrossRef] [PubMed]
- Urbanowicz, R.A.; Wang, R.; Schiel, J.E.; Keck, Z.Y.; Kerzic, M.C.; Lau, P.; Rangarajan, S.; Garagusi, K.J.; Tan, L.; Guest, J.D.; et al. Antigenicity and Immunogenicity of Differentially Glycosylated Hepatitis C Virus E2 Envelope Proteins Expressed in Mammalian and Insect Cells. J. Virol. 2019, 93. [Google Scholar] [CrossRef] [PubMed]
- Pantua, H.; Diao, J.; Ultsch, M.; Hazen, M.; Mathieu, M.; McCutcheon, K.; Takeda, K.; Date, S.; Cheung, T.K.; Phung, Q.; et al. Glycan shifting on hepatitis C virus (HCV) E2 glycoprotein is a mechanism for escape from broadly neutralizing antibodies. J. Mol. Biol. 2013, 425, 1899–1914. [Google Scholar] [CrossRef]
- Owsianka, A.; Tarr, A.W.; Juttla, V.S.; Lavillette, D.; Bartosch, B.; Cosset, F.L.; Ball, J.K.; Patel, A.H. Monoclonal antibody AP33 defines a broadly neutralizing epitope on the hepatitis C virus E2 envelope glycoprotein. J. Virol. 2005, 79, 11095–11104. [Google Scholar] [CrossRef]
- Voisset, C.; Op de Beeck, A.; Horellou, P.; Dreux, M.; Gustot, T.; Duverlie, G.; Cosset, F.L.; Vu-Dac, N.; Dubuisson, J. High-density lipoproteins reduce the neutralizing effect of hepatitis C virus (HCV)-infected patient antibodies by promoting HCV entry. J. Gen. Virol. 2006, 87, 2577–2581. [Google Scholar] [CrossRef]
- Dreux, M.; Pietschmann, T.; Granier, C.; Voisset, C.; Ricard-Blum, S.; Mangeot, P.E.; Keck, Z.; Foung, S.; Vu-Dac, N.; Dubuisson, J.; et al. High density lipoprotein inhibits hepatitis C virus-neutralizing antibodies by stimulating cell entry via activation of the scavenger receptor BI. J. Biol. Chem. 2006, 281, 18285–18295. [Google Scholar] [CrossRef]
- Fauvelle, C.; Felmlee, D.J.; Crouchet, E.; Lee, J.; Heydmann, L.; Lefevre, M.; Magri, A.; Hiet, M.S.; Fofana, I.; Habersetzer, F.; et al. Apolipoprotein E Mediates Evasion From Hepatitis C Virus Neutralizing Antibodies. Gastroenterology 2016, 150, 206–217.e204. [Google Scholar] [CrossRef]
- Deng, L.; Jiang, W.; Wang, X.; Merz, A.; Hiet, M.S.; Chen, Y.; Pan, X.; Jiu, Y.; Yang, Y.; Yu, B.; et al. Syntenin regulates hepatitis C virus sensitivity to neutralizing antibody by promoting E2 secretion through exosomes. J. Hepatol. 2019, 71, 52–61. [Google Scholar] [CrossRef]
- Wrensch, F.; Ligat, G.; Heydmann, L.; Schuster, C.; Zeisel, M.B.; Pessaux, P.; Habersetzer, F.; King, B.J.; Tarr, A.W.; Ball, J.K.; et al. Interferon-Induced Transmembrane Proteins Mediate Viral Evasion in Acute and Chronic Hepatitis C Virus Infection. Hepatology 2019, 70, 1506–1520. [Google Scholar] [CrossRef]
- Lohmann, V.; Bartenschlager, R. On the history of hepatitis C virus cell culture systems. J. Med. Chem. 2014, 57, 1627–1642. [Google Scholar] [CrossRef] [PubMed]
- Pietschmann, T.; Kaul, A.; Koutsoudakis, G.; Shavinskaya, A.; Kallis, S.; Steinmann, E.; Abid, K.; Negro, F.; Dreux, M.; Cosset, F.L.; et al. Construction and characterization of infectious intragenotypic and intergenotypic hepatitis C virus chimeras. Proc. Natl. Acad. Sci. USA 2006, 103, 7408–7413. [Google Scholar] [CrossRef] [PubMed]
- Gottwein, J.M.; Scheel, T.K.; Jensen, T.B.; Lademann, J.B.; Prentoe, J.C.; Knudsen, M.L.; Hoegh, A.M.; Bukh, J. Development and characterization of hepatitis C virus genotype 1-7 cell culture systems: Role of CD81 and scavenger receptor class B type I and effect of antiviral drugs. Hepatology 2009, 49, 364–377. [Google Scholar] [CrossRef] [PubMed]
- Carlsen, T.H.; Pedersen, J.; Prentoe, J.C.; Giang, E.; Keck, Z.Y.; Mikkelsen, L.S.; Law, M.; Foung, S.K.; Bukh, J. Breadth of neutralization and synergy of clinically relevant human monoclonal antibodies against HCV genotypes 1a, 1b, 2a, 2b, 2c, and 3a. Hepatology 2014, 60, 1551–1562. [Google Scholar] [CrossRef] [PubMed]
- McClure, C.P.; Urbanowicz, R.A.; King, B.J.; Cano-Crespo, S.; Tarr, A.W.; Ball, J.K. Flexible and rapid construction of viral chimeras applied to hepatitis C virus. J. Gen. Virol. 2016, 97, 2187–2193. [Google Scholar] [CrossRef]
- King, B.; Urbanowicz, R.; Tarr, A.W.; Ball, J.K.; McClure, C.P. InFusion Cloning for the Generation of Biologically Relevant HCV Chimeric Molecular Clones. Methods Mol. Biol. (Clifton N.J.) 2019, 1911, 93–104. [Google Scholar]
- Bartosch, B.; Dubuisson, J.; Cosset, F.L. Infectious hepatitis C virus pseudo-particles containing functional E1-E2 envelope protein complexes. J. Exp. Med. 2003, 197, 633–642. [Google Scholar] [CrossRef]
- Flint, M.; Logvinoff, C.; Rice, C.M.; McKeating, J.A. Characterization of infectious retroviral pseudotype particles bearing hepatitis C virus glycoproteins. J. Virol. 2004, 78, 6875–6882. [Google Scholar] [CrossRef]
- Urbanowicz, R.A.; McClure, C.P.; Brown, R.J.P.; Tsoleridis, T.; Persson, M.A.A.; Krey, T.; Irving, W.L.; Ball, J.K.; Tarr, A.W. A Diverse Panel of Hepatitis C Virus Glycoproteins for Use in Vaccine Research Reveals Extremes of Monoclonal Antibody Neutralization Resistance. J. Virol. 2016, 90, 3288–3301. [Google Scholar] [CrossRef]
- Urbanowicz, R.A.; McClure, C.P.; King, B.; Mason, C.P.; Ball, J.K.; Tarr, A.W. Novel functional hepatitis C virus glycoprotein isolates identified using an optimized viral pseudotype entry assay. J. Gen. Virol. 2016, 97, 2265–2279. [Google Scholar] [CrossRef]
- Bailey, J.R.; Wasilewski, L.N.; Snider, A.E.; El-Diwany, R.; Osburn, W.O.; Keck, Z.; Foung, S.K.; Ray, S.C. Naturally selected hepatitis C virus polymorphisms confer broad neutralizing antibody resistance. J. Clin. Investig. 2015, 125, 437–447. [Google Scholar] [CrossRef] [PubMed]
- Kolykhalov, A.A.; Agapov, E.V.; Blight, K.J.; Mihalik, K.; Feinstone, S.M.; Rice, C.M. Transmission of hepatitis C by intrahepatic inoculation with transcribed RNA. Science 1997, 277, 570–574. [Google Scholar] [CrossRef]
- Feinstone, S.M.; Alter, H.J.; Dienes, H.P.; Shimizu, Y.; Popper, H.; Blackmore, D.; Sly, D.; London, W.T.; Purcell, R.H. Non-A, non-B hepatitis in chimpanzees and marmosets. J. Infect. Dis. 1981, 144, 588–598. [Google Scholar] [CrossRef] [PubMed]
- Yanagi, M.; Purcell, R.H.; Emerson, S.U.; Bukh, J. Transcripts from a single full-length cDNA clone of hepatitis C virus are infectious when directly transfected into the liver of a chimpanzee. Proc. Natl. Acad. Sci. USA 1997, 94, 8738–8743. [Google Scholar] [CrossRef]
- Seaman, M.S.; Janes, H.; Hawkins, N.; Grandpre, L.E.; Devoy, C.; Giri, A.; Coffey, R.T.; Harris, L.; Wood, B.; Daniels, M.G.; et al. Tiered categorization of a diverse panel of HIV-1 Env pseudoviruses for assessment of neutralizing antibodies. J. Virol. 2010, 84, 1439–1452. [Google Scholar] [CrossRef]
- Mercer, D.F.; Schiller, D.E.; Elliott, J.F.; Douglas, D.N.; Hao, C.; Rinfret, A.; Addison, W.R.; Fischer, K.P.; Churchill, T.A.; Lakey, J.R.; et al. Hepatitis C virus replication in mice with chimeric human livers. Nat. Med. 2001, 7, 927–933. [Google Scholar] [CrossRef] [PubMed]
- Meuleman, P.; Libbrecht, L.; De Vos, R.; de Hemptinne, B.; Gevaert, K.; Vandekerckhove, J.; Roskams, T.; Leroux-Roels, G. Morphological and biochemical characterization of a human liver in a uPA-SCID mouse chimera. Hepatology 2005, 41, 847–856. [Google Scholar] [CrossRef]
- Chen, J.; Zhao, Y.; Zhang, C.; Chen, H.; Feng, J.; Chi, X.; Pan, Y.; Du, J.; Guo, M.; Cao, H.; et al. Persistent hepatitis C virus infections and hepatopathological manifestations in immune-competent humanized mice. Cell Res. 2014, 24, 1050–1066. [Google Scholar] [CrossRef]
- Ding, Q.; von Schaewen, M.; Hrebikova, G.; Heller, B.; Sandmann, L.; Plaas, M.; Ploss, A. Mice Expressing Minimally Humanized CD81 and Occludin Genes Support Hepatitis C Virus Uptake In Vivo. J. Virol. 2017, 91. [Google Scholar] [CrossRef]
- Yong, K.S.M.; Her, Z.; Chen, Q. Humanized Mouse Models for the Study of Hepatitis C and Host Interactions. Cells 2019, 8, 604. [Google Scholar] [CrossRef]
- Firth, C.; Bhat, M.; Firth, M.A.; Williams, S.H.; Frye, M.J.; Simmonds, P.; Conte, J.M.; Ng, J.; Garcia, J.; Bhuva, N.P.; et al. Detection of zoonotic pathogens and characterization of novel viruses carried by commensal Rattus norvegicus in New York City. Mbio 2014, 5, e01933-14. [Google Scholar] [CrossRef]
- Trivedi, S.; Murthy, S.; Sharma, H.; Hartlage, A.S.; Kumar, A.; Gadi, S.V.; Simmonds, P.; Chauhan, L.V.; Scheel, T.K.H.; Billerbeck, E.; et al. Viral persistence, liver disease, and host response in a hepatitis C-like virus rat model. Hepatology 2018, 68, 435–448. [Google Scholar] [CrossRef] [PubMed]
- Atcheson, E.; Li, W.; Bliss, C.M.; Chinnakannan, S.; Heim, K.; Sharpe, H.; Hutchings, C.; Dietrich, I.; Nguyen, D.; Kapoor, A.; et al. Use of an outbred rat hepacivirus challenge model for design and evaluation of efficacy of different immunisation strategies for HCV. Hepatology 2019. [Google Scholar] [CrossRef] [PubMed]
- Kapoor, A.; Simmonds, P.; Gerold, G.; Qaisar, N.; Jain, K.; Henriquez, J.A.; Firth, C.; Hirschberg, D.L.; Rice, C.M.; Shields, S.; et al. Characterization of a canine homolog of hepatitis C virus. Proc. Natl. Acad. Sci. USA 2011, 108, 11608–11613. [Google Scholar] [CrossRef] [PubMed]
- Burbelo, P.D.; Dubovi, E.J.; Simmonds, P.; Medina, J.L.; Henriquez, J.A.; Mishra, N.; Wagner, J.; Tokarz, R.; Cullen, J.M.; Iadarola, M.J.; et al. Serology-enabled discovery of genetically diverse hepaciviruses in a new host. J. Virol. 2012, 86, 6171–6178. [Google Scholar] [CrossRef]
- Pfaender, S.; Brown, R.J.; Pietschmann, T.; Steinmann, E. Natural reservoirs for homologs of hepatitis C virus. Emerg. Microbes Infect. 2014, 3, e21. [Google Scholar] [CrossRef]
- Pfaender, S.; Cavalleri, J.M.; Walter, S.; Doerrbecker, J.; Campana, B.; Brown, R.J.; Burbelo, P.D.; Postel, A.; Hahn, K.; Anggakusuma; et al. Clinical course of infection and viral tissue tropism of hepatitis C virus-like nonprimate hepaciviruses in horses. Hepatology 2015, 61, 447–459. [Google Scholar] [CrossRef]
- Tegtmeyer, B.; Echelmeyer, J.; Pfankuche, V.M.; Puff, C.; Todt, D.; Fischer, N.; Durham, A.; Feige, K.; Baumgartner, W.; Steinmann, E.; et al. Chronic equine hepacivirus infection in an adult gelding with severe hepatopathy. Vet. Med. Sci. 2019, 5, 372–378. [Google Scholar] [CrossRef]
- Shi, B.; Ma, L.; He, X.; Wang, X.; Wang, P.; Zhou, L.; Yao, X. Comparative analysis of human and mouse immunoglobulin variable heavy regions from IMGT/LIGM-DB with IMGT/HighV-QUEST. Theor. Biol. Med. Model. 2014, 11, 30. [Google Scholar] [CrossRef]
- Manso, T.C.; Groenner-Penna, M.; Minozzo, J.C.; Antunes, B.C.; Ippolito, G.C.; Molina, F.; Felicori, L.F. Next-generation sequencing reveals new insights about gene usage and CDR-H3 composition in the horse antibody repertoire. Mol. Immunol. 2019, 105, 251–259. [Google Scholar] [CrossRef]
- Li, Y.; Pierce, B.G.; Wang, Q.; Keck, Z.Y.; Fuerst, T.R.; Foung, S.K.; Mariuzza, R.A. Structural basis for penetration of the glycan shield of hepatitis C virus E2 glycoprotein by a broadly neutralizing human antibody. J. Biol. Chem. 2015, 290, 10117–10125. [Google Scholar] [CrossRef] [PubMed]
- Keck, Z.Y.; Angus, A.G.; Wang, W.; Lau, P.; Wang, Y.; Gatherer, D.; Patel, A.H.; Foung, S.K. Non-random escape pathways from a broadly neutralizing human monoclonal antibody map to a highly conserved region on the hepatitis C virus E2 glycoprotein encompassing amino acids 412–423. PLoS Pathog. 2014, 10, e1004297. [Google Scholar] [CrossRef] [PubMed]
- Pierce, B.G.; Boucher, E.N.; Piepenbrink, K.H.; Ejemel, M.; Rapp, C.A.; Thomas, W.D., Jr.; Sundberg, E.J.; Weng, Z.; Wang, Y. Structure-Based Design of Hepatitis C Virus Vaccines That Elicit Neutralizing Antibody Responses to a Conserved Epitope. J. Virol. 2017, 91. [Google Scholar] [CrossRef] [PubMed]
- Filskov, J.; Andersen, P.; Agger, E.M.; Bukh, J. HCV p7 as a novel vaccine-target inducing multifunctional CD4(+) and CD8(+) T-cells targeting liver cells expressing the viral antigen. Sci. Rep. 2019, 9, 14085. [Google Scholar] [CrossRef] [PubMed]
- Dawood, R.M.; Moustafa, R.I.; Abdelhafez, T.H.; El-Shenawy, R.; El-Abd, Y.; Bader El Din, N.G.; Dubuisson, J.; El Awady, M.K. A multiepitope peptide vaccine against HCV stimulates neutralizing humoral and persistent cellular responses in mice. BMC Infect. Dis. 2019, 19, 932. [Google Scholar] [CrossRef] [PubMed]
- Marin, M.Q.; Perez, P.; Ljungberg, K.; Sorzano, C.O.S.; Gomez, C.E.; Liljestrom, P.; Esteban, M.; Garcia-Arriaza, J. Potent Anti-hepatitis C Virus (HCV) T Cell Immune Responses Induced in Mice Vaccinated with DNA-Launched RNA Replicons and Modified Vaccinia Virus Ankara-HCV. J. Virol. 2019, 93. [Google Scholar] [CrossRef]
- Wijesundara, D.K.; Gummow, J.; Li, Y.; Yu, W.; Quah, B.J.; Ranasinghe, C.; Torresi, J.; Gowans, E.J.; Grubor-Bauk, B. Induction of Genotype Cross-Reactive, Hepatitis C Virus-Specific, Cell-Mediated Immunity in DNA-Vaccinated Mice. J. Virol. 2018, 92. [Google Scholar] [CrossRef]
- Masavuli, M.G.; Wijesundara, D.K.; Underwood, A.; Christiansen, D.; Earnest-Silveira, L.; Bull, R.; Torresi, J.; Gowans, E.J.; Grubor-Bauk, B. A Hepatitis C Virus DNA Vaccine Encoding a Secreted, Oligomerized Form of Envelope Proteins Is Highly Immunogenic and Elicits Neutralizing Antibodies in Vaccinated Mice. Front. Immunol. 2019, 10, 1145. [Google Scholar] [CrossRef]
- Plotkin, S.A. Vaccines: Past, present and future. Nat. Med. 2005, 11, S5–S11. [Google Scholar] [CrossRef]
- Choo, Q.L.; Kuo, G.; Ralston, R.; Weiner, A.; Chien, D.; Van Nest, G.; Han, J.; Berger, K.; Thudium, K.; Kuo, C.; et al. Vaccination of chimpanzees against infection by the hepatitis C virus. Proc. Natl. Acad. Sci. USA 1994, 91, 1294–1298. [Google Scholar] [CrossRef]
- Houghton, M. Prospects for prophylactic and therapeutic vaccines against the hepatitis C viruses. Immunol. Rev. 2011, 239, 99–108. [Google Scholar] [CrossRef] [PubMed]
- Frey, S.E.; Houghton, M.; Coates, S.; Abrignani, S.; Chien, D.; Rosa, D.; Pileri, P.; Ray, R.; Di Bisceglie, A.M.; Rinella, P.; et al. Safety and immunogenicity of HCV E1E2 vaccine adjuvanted with MF59 administered to healthy adults. Vaccine 2010, 28, 6367–6373. [Google Scholar] [CrossRef] [PubMed]
- Ray, R.; Meyer, K.; Banerjee, A.; Basu, A.; Coates, S.; Abrignani, S.; Houghton, M.; Frey, S.E.; Belshe, R.B. Characterization of antibodies induced by vaccination with hepatitis C virus envelope glycoproteins. J. Infect. Dis. 2010, 202, 862–866. [Google Scholar] [CrossRef] [PubMed]
- Law, J.L.; Chen, C.; Wong, J.; Hockman, D.; Santer, D.M.; Frey, S.E.; Belshe, R.B.; Wakita, T.; Bukh, J.; Jones, C.T.; et al. A hepatitis C virus (HCV) vaccine comprising envelope glycoproteins gpE1/gpE2 derived from a single isolate elicits broad cross-genotype neutralizing antibodies in humans. PLoS ONE 2013, 8, e59776. [Google Scholar] [CrossRef] [PubMed]
- Wong, J.A.; Bhat, R.; Hockman, D.; Logan, M.; Chen, C.; Levin, A.; Frey, S.E.; Belshe, R.B.; Tyrrell, D.L.; Law, J.L.; et al. Recombinant hepatitis C virus envelope glycoprotein vaccine elicits antibodies targeting multiple epitopes on the envelope glycoproteins associated with broad cross-neutralization. J. Virol. 2014, 88, 14278–14288. [Google Scholar] [CrossRef] [PubMed]
- Logan, M.; Law, J.; Wong, J.A.J.; Hockman, D.; Landi, A.; Chen, C.; Crawford, K.; Kundu, J.; Baldwin, L.; Johnson, J.; et al. Native Folding of a Recombinant gpE1/gpE2 Heterodimer Vaccine Antigen from a Precursor Protein Fused with Fc IgG. J. Virol. 2017, 91. [Google Scholar] [CrossRef]
- Ruwona, T.B.; Giang, E.; Nieusma, T.; Law, M. Fine mapping of murine antibody responses to immunization with a novel soluble form of hepatitis C virus envelope glycoprotein complex. J. Virol. 2014, 88, 10459–10471. [Google Scholar] [CrossRef]
- Krapchev, V.B.; Rychlowska, M.; Chmielewska, A.; Zimmer, K.; Patel, A.H.; Bienkowska-Szewczyk, K. Recombinant Flag-tagged E1E2 glycoproteins from three hepatitis C virus genotypes are biologically functional and elicit cross-reactive neutralizing antibodies in mice. Virology 2018, 519, 33–41. [Google Scholar] [CrossRef]
- McCaffrey, K.; Boo, I.; Poumbourios, P.; Drummer, H.E. Expression and characterization of a minimal hepatitis C virus glycoprotein E2 core domain that retains CD81 binding. J. Virol. 2007, 81, 9584–9590. [Google Scholar] [CrossRef]
- Vietheer, P.T.; Boo, I.; Gu, J.; McCaffrey, K.; Edwards, S.; Owczarek, C.; Hardy, M.P.; Fabri, L.; Center, R.J.; Poumbourios, P.; et al. The core domain of hepatitis C virus glycoprotein E2 generates potent cross-neutralizing antibodies in guinea pigs. Hepatology 2017, 65, 1117–1131. [Google Scholar] [CrossRef]
- McCaffrey, K.; Boo, I.; Owczarek, C.M.; Hardy, M.P.; Perugini, M.A.; Fabri, L.; Scotney, P.; Poumbourios, P.; Drummer, H.E. An Optimized Hepatitis C Virus E2 Glycoprotein Core Adopts a Functional Homodimer That Efficiently Blocks Virus Entry. J. Virol. 2017, 91. [Google Scholar] [CrossRef] [PubMed]
- Law, J.L.M.; Logan, M.; Wong, J.; Kundu, J.; Hockman, D.; Landi, A.; Chen, C.; Crawford, K.; Wininger, M.; Johnson, J.; et al. Role of the E2 Hypervariable Region (HVR1) in the Immunogenicity of a Recombinant Hepatitis C Virus Vaccine. J. Virol. 2018, 92. [Google Scholar] [CrossRef]
- Torresi, J. The Rationale for a Preventative HCV Virus-Like Particle (VLP) Vaccine. Front. Microbiol. 2017, 8, 2163. [Google Scholar] [CrossRef] [PubMed]
- Bachmann, M.F.; Jennings, G.T. Vaccine delivery: A matter of size, geometry, kinetics and molecular patterns. Nat. Rev. Immunol. 2010, 10, 787–796. [Google Scholar] [CrossRef] [PubMed]
- Baumert, T.F.; Ito, S.; Wong, D.T.; Liang, T.J. Hepatitis C virus structural proteins assemble into viruslike particles in insect cells. J. Virol. 1998, 72, 3827–3836. [Google Scholar] [CrossRef] [PubMed]
- Baumert, T.F.; Vergalla, J.; Satoi, J.; Thomson, M.; Lechmann, M.; Herion, D.; Greenberg, H.B.; Ito, S.; Liang, T.J. Hepatitis C virus-like particles synthesized in insect cells as a potential vaccine candidate. Gastroenterology 1999, 117, 1397–1407. [Google Scholar] [CrossRef]
- Lechmann, M.; Murata, K.; Satoi, J.; Vergalla, J.; Baumert, T.F.; Liang, T.J. Hepatitis C virus-like particles induce virus-specific humoral and cellular immune responses in mice. Hepatology 2001, 34, 417–423. [Google Scholar] [CrossRef]
- Jeong, S.H.; Qiao, M.; Nascimbeni, M.; Hu, Z.; Rehermann, B.; Murthy, K.; Liang, T.J. Immunization with hepatitis C virus-like particles induces humoral and cellular immune responses in nonhuman primates. J. Virol. 2004, 78, 6995–7003. [Google Scholar] [CrossRef]
- Elmowalid, G.A.; Qiao, M.; Jeong, S.H.; Borg, B.B.; Baumert, T.F.; Sapp, R.K.; Hu, Z.; Murthy, K.; Liang, T.J. Immunization with hepatitis C virus-like particles results in control of hepatitis C virus infection in chimpanzees. Proc. Natl. Acad. Sci. USA 2007, 104, 8427–8432. [Google Scholar] [CrossRef]
- Chua, B.Y.; Johnson, D.; Tan, A.; Earnest-Silveira, L.; Sekiya, T.; Chin, R.; Torresi, J.; Jackson, D.C. Hepatitis C VLPs delivered to dendritic cells by a TLR2 targeting lipopeptide results in enhanced antibody and cell-mediated responses. PLoS ONE 2012, 7, e47492. [Google Scholar] [CrossRef]
- Christiansen, D.; Earnest-Silveira, L.; Chua, B.; Meuleman, P.; Boo, I.; Grubor-Bauk, B.; Jackson, D.C.; Keck, Z.Y.; Foung, S.K.H.; Drummer, H.E.; et al. Immunological responses following administration of a genotype 1a/1b/2/3a quadrivalent HCV VLP vaccine. Sci. Rep. 2018, 8, 6483. [Google Scholar] [CrossRef] [PubMed]
- Earnest-Silveira, L.; Chua, B.; Chin, R.; Christiansen, D.; Johnson, D.; Herrmann, S.; Ralph, S.A.; Vercauteren, K.; Mesalam, A.; Meuleman, P.; et al. Characterization of a hepatitis C virus-like particle vaccine produced in a human hepatocyte-derived cell line. J. Gen. Virol. 2016, 97, 1865–1876. [Google Scholar] [CrossRef] [PubMed]
- Collett, S.; Torresi, J.; Earnest-Silveira, L.; Christiansen, D.; Elbourne, A.; Ramsland, P.A. Probing and pressing surfaces of hepatitis C virus-like particles. J. Colloid Interface Sci. 2019, 545, 259–268. [Google Scholar] [CrossRef]
- Christiansen, D.; Earnest-Silveira, L.; Grubor-Bauk, B.; Wijesundara, D.K.; Boo, I.; Ramsland, P.A.; Vincan, E.; Drummer, H.E.; Gowans, E.J.; Torresi, J. Pre-clinical evaluation of a quadrivalent HCV VLP vaccine in pigs following microneedle delivery. Sci. Rep. 2019, 9, 9251. [Google Scholar] [CrossRef]
- Ura, T.; Okuda, K.; Shimada, M. Developments in Viral Vector-Based Vaccines. Vaccines 2014, 2, 624–641. [Google Scholar] [CrossRef] [PubMed]
- Barnes, E.; Folgori, A.; Capone, S.; Swadling, L.; Aston, S.; Kurioka, A.; Meyer, J.; Huddart, R.; Smith, K.; Townsend, R.; et al. Novel adenovirus-based vaccines induce broad and sustained T cell responses to HCV in man. Sci. Transl. Med. 2012, 4, 115ra1. [Google Scholar] [CrossRef] [PubMed]
- Capone, S.; Zampaglione, I.; Vitelli, A.; Pezzanera, M.; Kierstead, L.; Burns, J.; Ruggeri, L.; Arcuri, M.; Cappelletti, M.; Meola, A.; et al. Modulation of the immune response induced by gene electrotransfer of a hepatitis C virus DNA vaccine in nonhuman primates. J. Immunol. 2006, 177, 7462–7471. [Google Scholar] [CrossRef] [PubMed]
- Swadling, L.; Capone, S.; Antrobus, R.D.; Brown, A.; Richardson, R.; Newell, E.W.; Halliday, J.; Kelly, C.; Bowen, D.; Fergusson, J.; et al. A human vaccine strategy based on chimpanzee adenoviral and MVA vectors that primes, boosts, and sustains functional HCV-specific T cell memory. Sci. Transl. Med. 2014, 6, 261ra153. [Google Scholar] [CrossRef]
- Swadling, L.; Halliday, J.; Kelly, C.; Brown, A.; Capone, S.; Ansari, M.A.; Bonsall, D.; Richardson, R.; Hartnell, F.; Collier, J.; et al. Highly-Immunogenic Virally-Vectored T-cell Vaccines Cannot Overcome Subversion of the T-cell Response by HCV during Chronic Infection. Vaccines 2016, 4, 27. [Google Scholar] [CrossRef]
- Trial Evaluating Experimental Hepatitis C Vaccine Concludes|NIH: National Institute of Allergy and Infectious Diseases. Available online: https://www.niaid.nih.gov/news-events/trial-evaluating-experimental-hepatitis-c-vaccine-concludes (accessed on 29 December 2019).
- Beaumont, E.; Patient, R.; Hourioux, C.; Dimier-Poisson, I.; Roingeard, P. Chimeric hepatitis B virus/hepatitis C virus envelope proteins elicit broadly neutralizing antibodies and constitute a potential bivalent prophylactic vaccine. Hepatology 2013, 57, 1303–1313. [Google Scholar] [CrossRef]
- Wei, S.; Lei, Y.; Yang, J.; Wang, X.; Shu, F.; Wei, X.; Lin, F.; Li, B.; Cui, Y.; Zhang, H. Neutralization effects of antibody elicited by chimeric HBV S antigen viral-like particles presenting HCV neutralization epitopes. Vaccine 2018, 36, 2273–2281. [Google Scholar] [CrossRef] [PubMed]
- Garrone, P.; Fluckiger, A.C.; Mangeot, P.E.; Gauthier, E.; Dupeyrot-Lacas, P.; Mancip, J.; Cangialosi, A.; Du Chene, I.; LeGrand, R.; Mangeot, I.; et al. A prime-boost strategy using virus-like particles pseudotyped for HCV proteins triggers broadly neutralizing antibodies in macaques. Sci. Transl. Med. 2011, 3, 94ra71. [Google Scholar] [CrossRef] [PubMed]
| Vaccine Type | HCV Target | HCV Strain | Tested Species | Antibody Response * | CD4+ T Cell Response † | CD8+ T Cell Response † | Ref. |
|---|---|---|---|---|---|---|---|
| Subunit | |||||||
| HCV-1 rE1E2 | E1E2 | Gt1a HCV-1 | humans | homologous and heterologous | yes | N.D | [162,164] |
| H77 sE2Δ123 | E2core | Gt1a H77 | guinea pigs | homologous and heterologous | N.D | N.D | [170] |
| Virus-Like particles | |||||||
| core, E1, E2 from Gt1a, 1b, 2a and 3a | core, E1, E2 | Gt1a H77, Gt1b BK, Gt2a JFH1, Gt3a | mice, pigs | homologous neutralising antibodies | yes | yes | [181,184] |
| HBV/HCV-LPs | E1, E2 | Gt1a H77 | rabbit | homologous, heterologous activity towards Gt1a and 1b, reduced activity towards Gt2a and 3a isolates. | N.D | N.D | [191] |
| HBV/HCV-LPs | linear E1 and E2 epitopes | not stated | mice | heterologous towards Gt1a, 1b and 2a | N.D | N.D | [192] |
| murine leukaemia VLP-HCVE1E2 | E1, E2 | Gt1a H77 | mice, macaques | homologous, heterologous towards Gt1b, 2a, 2b and 4c | yes | N.D | [193] |
| Viral vector | |||||||
| ChAd3/MVA-Nsmut | NS3, NS4a, NS4b, NS5a, NS5b | Gt1a BK | humans | N/A | yes | yes | [188] |
| Peptide | |||||||
| p7 | p7 | Gt1b J4 | mice | N/A | yes | yes | [154] |
| HCVp6-MAP | E1, E2, NS4b, NS5a, NS5b | Gt4a ED43 | mice | homologous, heterologous towards JFH1 | yes | yes | [155] |
| DNA | |||||||
| pVax-sE1E2-IMX313P | E1, E2 | Gt1b | mice | homologous, heterologous towards Gt1a, 1b, 2a, 2b, 3a, 4a, 5, 6 | yes | N.D | [158] |
| DREP-HCV/MVA-HCV | core, E1, E1, p7, NS2, NS3 | Gt1a H77 | mice | non-neutralising IgG | yes | yes | [156] |
| pVax-N3-NS5b | NS3, NS4, NS5b | Gt1b, Gt3a | mice | N/A | yes | yes | [157] |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Duncan, J.D.; Urbanowicz, R.A.; Tarr, A.W.; Ball, J.K. Hepatitis C Virus Vaccine: Challenges and Prospects. Vaccines 2020, 8, 90. https://doi.org/10.3390/vaccines8010090
Duncan JD, Urbanowicz RA, Tarr AW, Ball JK. Hepatitis C Virus Vaccine: Challenges and Prospects. Vaccines. 2020; 8(1):90. https://doi.org/10.3390/vaccines8010090
Chicago/Turabian StyleDuncan, Joshua D., Richard A. Urbanowicz, Alexander W. Tarr, and Jonathan K. Ball. 2020. "Hepatitis C Virus Vaccine: Challenges and Prospects" Vaccines 8, no. 1: 90. https://doi.org/10.3390/vaccines8010090
APA StyleDuncan, J. D., Urbanowicz, R. A., Tarr, A. W., & Ball, J. K. (2020). Hepatitis C Virus Vaccine: Challenges and Prospects. Vaccines, 8(1), 90. https://doi.org/10.3390/vaccines8010090





