Considering Genomic and Immunological Correlates of Protection for a Dengue Intervention
Abstract
1. Introduction
2. Viral Lifecycle
3. Immune Correlates of Protection
4. Viral Dynamics
5. Current Vaccine Strategies
6. Conclusions
- ADE: The process where cross-reactive IgG antibodies from a previous infection with poor neutralising capability bind the heterologous serotype and enhances its uptake via FcRs into host cells.
- ADE Antibody: Antibody that binds viral particles but does not prevent viral entry into host cells.
- Antigenic-Sin: The domination of cross-reactive memory CD8 T-cells from the original serotype during a heterotypic infection which have low avidity.
- Heterotypic: A different DENV serotype.
- Homotypic: The same DENV serotype.
- Cross-reactive: Immune cells which react to multiple serotypes.
- Epitope Variants: Epitopes across serotypes with sequence differences between serotypes.
- Purifying Selection: Amino acid changes fixed by fitness advantage.
- Positive Selection: Amino acid changes fixed by chance.
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Bhatt, S.; Gething, P.W.; Brady, O.J.; Messina, J.P.; Farlow, A.W.; Moyes, C.L.; Drake, J.M.; Brownstein, J.S.; Hoen, A.G.; Sankoh, O.; et al. The global distribution and burden of dengue. Nature 2013, 496, 504–507. [Google Scholar] [CrossRef] [PubMed]
- Simmons, C.P.; Farrar, J.J.; Van Vinh Chau, N.; Wills, B. Dengue. N. Engl. J. Med. 2012, 366, 1423–1432. [Google Scholar] [CrossRef] [PubMed]
- Campion, E.W.; Weaver, S.C.; Lecuit, M. Chikungunya Virus and the Global Spread of a Mosquito-Borne Disease. N. Engl. J. Med. 2015, 372, 1231–1239. [Google Scholar]
- Guzman, M.G.; Harris, E. Dengue. Lancet 2015, 385, 453–465. [Google Scholar] [CrossRef]
- Bowman, L.R.; Donegan, S.; McCall, P.J. Is Dengue Vector Control Deficient in Effectiveness or Evidence? Systematic Review and Meta-analysis. PLoS Negl. Trop. Dis. 2016, 10, e0004551. [Google Scholar] [CrossRef]
- Hemingway, J.; Beaty, B.J.; Rowland, M.; Scott, T.W.; Sharp, B.L. The Innovative Vector Control Consortium: Improved control of mosquito-borne diseases. Trends Parasitol. 2006, 22, 308–312. [Google Scholar] [CrossRef]
- Lescar, J.; Luo, D.; Xu, T.; Sampath, A.; Lim, S.P.; Canard, B.; Vasudevan, S.G. Towards the design of antiviral inhibitors against flaviviruses: The case for the multifunctional NS3 protein from Dengue virus as a target. Antiviral Res. 2008, 80, 94–101. [Google Scholar] [CrossRef]
- Noble, C.G.; Chen, Y.-L.; Dong, H.; Gu, F.; Lim, S.P.; Schul, W.; Wang, Q.-Y.; Shi, P.-Y. Strategies for development of dengue virus inhibitors. Antiviral Res. 2010, 85, 450–462. [Google Scholar] [CrossRef]
- Cheung, Y.Y.; Chen, K.C.; Chen, H.; Seng, E.K.; Chu, J.J.H. Antiviral activity of lanatoside C against dengue virus infection. Antiviral Res. 2014, 111, 93–99. [Google Scholar] [CrossRef]
- Abdelnabi, R.; Neyts, J.; Delang, L. Towards antivirals against chikungunya virus. Antiviral Res. 2015, 121, 59–68. [Google Scholar] [CrossRef]
- Lani, R.; Hassandarvish, P.; Chiam, C.W.; Moghaddam, E.; Chu, J.J.H.; Rausalu, K.; Merits, A.; Higgs, S.; Vanlandingham, D.; Bakar, S.A.; et al. Antiviral activity of silymarin against chikungunya virus. Sci. Rep. 2015, 5, 11421. [Google Scholar] [CrossRef] [PubMed]
- Bin Masri, M.F.; Rathore, A.P.S.; John, A.L.S. Therapeutics for Dengue. Curr. Treat. Options Infect. Dis. 2019, 11, 199–214. [Google Scholar] [CrossRef]
- Vaughn, D.W.; Green, S.; Kalayanarooj, S.; Innis, B.L.; Nimmannitya, S.; Suntayakorn, S.; Rothman, A.L.; Ennis, F.A.; Nisalak, A. Dengue in the early febrile phase: Viremia and antibody responses. J. Infect. Dis. 1997, 176, 322–330. [Google Scholar] [CrossRef] [PubMed]
- Srikiatkhachorn, A.; Yoon, I.-K. Immune correlates for dengue vaccine development. Expert Rev. Vaccines 2016, 15, 455–465. [Google Scholar] [CrossRef]
- Diamond, M.S.; Pierson, T.C. Molecular Insight into Dengue Virus Pathogenesis and Its Implications for Disease Control. Cell 2015, 162, 488–492. [Google Scholar] [CrossRef] [PubMed]
- Halstead, S.B.; Russell, P.K. Protective and immunological behavior of chimeric yellow fever dengue vaccine. Vaccine 2016, 34, 1643–1647. [Google Scholar] [CrossRef]
- Capeding, M.R.; Tran, N.H.; Hadinegoro, S.R.S.; Ismail, H.I.H.M.; Chotpitayasunondh, T.; Chua, M.N.; Luong, C.Q.; Rusmil, K.; Wirawan, D.N.; Nallusamy, R.; et al. Clinical efficacy and safety of a novel tetravalent dengue vaccine in healthy children in Asia: A phase 3, randomised, observer-masked, placebo-controlled trial. Lancet 2014, 384, 1358–1365. [Google Scholar] [CrossRef]
- Villar, L.; Dayan, G.H.; Arredondo-García, J.L.; Rivera, D.M.; Cunha, R.; Deseda, C.; Reynales, H.; Costa, M.S.; Morales-Ramírez, J.O.; Carrasquilla, G.; et al. Efficacy of a Tetravalent Dengue Vaccine in Children in Latin America. N. Engl. J. Med. 2015, 372, 113–123. [Google Scholar] [CrossRef]
- Hadinegoro, S.R.; Arredondo-García, J.L.; Capeding, M.R.; Deseda, C.; Chotpitayasunondh, T.; Dietze, R.; Hj Muhammad Ismail, H.I.; Reynales, H.; Limkittikul, K.; Rivera-Medina, D.M.; et al. Efficacy and Long-Term Safety of a Dengue Vaccine in Regions of Endemic Disease. N. Engl. J. Med. 2015, 373, 1195–1206. [Google Scholar] [CrossRef]
- Sabchareon, A.; Wallace, D.; Sirivichayakul, C.; Limkittikul, K.; Chanthavanich, P.; Suvannadabba, S.; Jiwariyavej, V.; Dulyachai, W.; Pengsaa, K.; Wartel, T.A.; et al. Protective efficacy of the recombinant, live-attenuated, CYD tetravalent dengue vaccine in Thai schoolchildren: A randomised, controlled phase 2b trial. Lancet 2012, 380, 1559–1567. [Google Scholar] [CrossRef]
- Guzman, M.G.; Gubler, D.J.; Izquierdo, A.; Martínez, E.; Halstead, S.B. Dengue infection. Nat. Rev. Dis. Primers 2016, 2, 1–25. [Google Scholar] [CrossRef] [PubMed]
- Cardosa, J.; Ooi, M.H.; Tio, P.H.; Perera, D.; Holmes, E.C.; Bibi, K.; Manap, Z.A. Dengue Virus Serotype 2 from a Sylvatic Lineage Isolated from a Patient with Dengue Hemorrhagic Fever. PLoS Negl. Trop. Dis. 2009, 3, e423. [Google Scholar] [CrossRef] [PubMed]
- Wichit, S.; Ferraris, P.; Choumet, V.; Missé, D. The effects of mosquito saliva on dengue virus infectivity in humans. Curr. Opin. Virol. 2016, 21, 139–145. [Google Scholar] [CrossRef] [PubMed]
- Briant, L.; Desprès, P.; Choumet, V.; Missé, D. Role of skin immune cells on the host susceptibility to mosquito-borne viruses. Virology 2014, 464–465, 26–32. [Google Scholar] [CrossRef]
- Ferreira-de-Lima, V.H.; Lima-Camara, T.N. Natural vertical transmission of dengue virus in Aedes aegypti and Aedes albopictus: A systematic review. Parasites Vectors 2018, 11. [Google Scholar]
- Mukhopadhyay, S.; Kuhn, R.J.; Rossmann, M.G. A structural perspective of the flavivirus life cycle. Nat. Rev. Microbiol. 2005, 3, 13–22. [Google Scholar] [CrossRef]
- Cruz-Oliveira, C.; Freire, J.M.; Conceição, T.M.; Higa, L.M.; Castanho, M.A.; Da Poian, A.T. Receptors and routes of dengue virus entry into the host cells. FEMS Microbiol. Rev. 2015, 39, 155–170. [Google Scholar] [CrossRef]
- Screaton, G.; Mongkolsapaya, J.; Yacoub, S.; Roberts, C. New insights into the immunopathology and control of dengue virus infection. Nat. Rev. Immunol. 2015, 15, 745–759. [Google Scholar] [CrossRef]
- Unckless, R.L.; Jaenike, J. Structure of dengue virus: Implications for flavivirus organization, maturation, and fusion. Evolution 2011, 66, 678–689. [Google Scholar] [CrossRef]
- Allison, S.L.; Schalich, J.; Stiasny, K.; Mandl, C.W.; Heinz, F.X. Mutational Evidence for an Internal Fusion Peptide in Flavivirus Envelope Protein E. J. Virol. 2001, 75, 4268–4275. [Google Scholar] [CrossRef]
- Rodenhuis-Zybert, I.A.; Wilschut, J.; Smit, J.M. Dengue virus life cycle: Viral and host factors modulating infectivity. Cell. Mol. Life Sci. 2010, 67, 2773–2786. [Google Scholar] [CrossRef] [PubMed]
- Chanprapaph, S.; Saparpakorn, P.; Sangma, C.; Niyomrattanakit, P.; Hannongbua, S.; Angsuthanasombat, C.; Katzenmeier, G. Competitive inhibition of the dengue virus NS3 serine protease by synthetic peptides representing polyprotein cleavage sites. Biochem. Biophys. Res. Commun. 2005, 330, 1237–1246. [Google Scholar] [CrossRef] [PubMed]
- Chambers, T.J.; Hahn, C.S.; Galler, R.; Rice, C.M. Flavivirus Genome Organization, Expression, and Replication. Annu. Rev. Microbiol. 1990, 44, 649–688. [Google Scholar] [CrossRef] [PubMed]
- Falgout, B.; Pethel, M.; Zhang, Y.M.; Lai, C.J. Both nonstructural proteins NS2B and NS3 are required for the proteolytic processing of dengue virus nonstructural proteins. J. Virol. 1991, 65, 2467–2475. [Google Scholar]
- Yusof, R.; Clum, S.; Wetzel, M.; Murthy, H.M.; Padmanabhan, R. Purified NS2B/NS3 serine protease of dengue virus type 2 exhibits cofactor NS2B dependence for cleavage of substrates with dibasic amino acids in vitro. J. Biol. Chem. 2000, 275, 9963–9969. [Google Scholar] [CrossRef]
- Noble, C.G.; Seh, C.C.; Chao, A.T.; Shi, P.Y. Ligand-Bound Structures of the Dengue Virus Protease Reveal the Active Conformation. J. Virol. 2011, 86, 438–446. [Google Scholar] [CrossRef]
- Clyde, K.; Kyle, J.L.; Harris, E. Recent advances in deciphering viral and host determinants of dengue virus replication and pathogenesis. J. Virol. 2006, 80, 11418–11431. [Google Scholar] [CrossRef]
- Malet, H.; Massé, N.; Selisko, B.; Romette, J.-L.; Alvarez, K.; Guillemot, J.C.; Tolou, H.; Yap, T.L.; Vasudevan, S.G.; Lescar, J.; et al. The flavivirus polymerase as a target for drug discovery. Antiviral Res. 2008, 80, 23–35. [Google Scholar] [CrossRef]
- Selisko, B.; Wang, C.; Harris, E.; Canard, B. Regulation of Flavivirus RNA synthesis and replication. Curr. Opin. Virol. 2014, 9, 74–83. [Google Scholar] [CrossRef]
- Luo, D.; Xu, T.; Hunke, C.; Grüber, G.; Vasudevan, S.G.; Lescar, J. Crystal structure of the NS3 protease-helicase from dengue virus. J. Virol. 2008, 82, 173–183. [Google Scholar] [CrossRef]
- Li, L.; Lok, S.M.; Yu, I.-M.; Zhang, Y.; Kuhn, R.J.; Chen, J.; Rossmann, M.G. The Flavivirus Precursor Membrane-Envelope Protein Complex: Structure and Maturation. Science 2008, 319, 1830–1834. [Google Scholar] [CrossRef] [PubMed]
- Yu, I.-M.; Zhang, W.; Holdaway, H.A.; Li, L.; Kostyuchenko, V.A.; Chipman, P.R.; Kuhn, R.J.; Rossmann, M.G.; Chen, J. Structure of the immature dengue virus at low pH primes proteolytic maturation. Science 2008, 319, 1834–1837. [Google Scholar] [CrossRef] [PubMed]
- Junjhon, J.; Lausumpao, M.; Supasa, S.; Noisakran, S.; Songjaeng, A.; Saraithong, P.; Chaichoun, K.; Utaipat, U.; Keelapang, P.; Kanjanahaluethai, A.; et al. Differential Modulation of prM Cleavage, Extracellular Particle Distribution, and Virus Infectivity by Conserved Residues at Nonfurin Consensus Positions of the Dengue Virus pr-M Junction. J. Virol. 2008, 82, 10776–10791. [Google Scholar] [CrossRef] [PubMed]
- Miller, S.; Kastner, S.; Krijnse-Locker, J.; Bühler, S.; Bartenschlager, R. The non-structural protein 4A of dengue virus is an integral membrane protein inducing membrane alterations in a 2K-regulated manner. J. Biol. Chem. 2007, 282, 8873–8882. [Google Scholar] [CrossRef]
- Boon den, J.A.; Diaz, A.; Ahlquist, P. Cytoplasmic viral replication complexes. Cell Host Microbe 2010, 8, 77–85. [Google Scholar] [CrossRef]
- Scaturro, P.; Cortese, M.; Chatel-Chaix, L.; Fischl, W.; Bartenschlager, R. Dengue Virus Non-structural Protein 1 Modulates Infectious Particle Production via Interaction with the Structural Proteins. PLoS Pathog 2015, 11, e1005277. [Google Scholar]
- Xie, X.; Gayen, S.; Kang, C.; Yuan, Z.; Shi, P.-Y. Membrane topology and function of dengue virus NS2A protein. J. Virol. 2013, 87, 4609–4622. [Google Scholar] [CrossRef]
- Riedel, S. Edward Jenner and the history of smallpox and vaccination. Proceedings (Baylor University. Medical Center) 2005, 18, 21–689. [Google Scholar] [CrossRef]
- Dudas, R.A.; Karron, R.A. Respiratory syncytial virus vaccines. Clin. Microbiol. Rev. 1998, 11, 430–439. [Google Scholar] [CrossRef]
- Kim, H.W.; Canchola, J.G.; Brandt, C.D.; Pyles, G.; Chanock, R.M.; Jensen, K.; Parrott, R.H. Respiratory syncytial virus disease in infants despite prior administration of antigenic inactivated vaccine. Am. J. Epidemiol. 1969, 89, 422–434. [Google Scholar] [CrossRef]
- Halstead, S.B. Dengue Antibody-Dependent Enhancement: Knowns and Unknowns. Microbiol. Spectr. 2014, 2, 249–271. [Google Scholar]
- Gibbons, R.V.; Kalanarooj, S.; Jarman, R.G.; Nisalak, A.; Vaughn, D.W.; Endy, T.P.; Mammen, M.P.; Srikiatkhachorn, A. Analysis of repeat hospital admissions for dengue to estimate the frequency of third or fourth dengue infections resulting in admissions and dengue hemorrhagic fever, and serotype sequences. Am. J. Trop. Med. Hyg. 2007, 77, 910–913. [Google Scholar] [CrossRef] [PubMed]
- Burke, D.S.; Nisalak, A.; Johnson, D.E.; Scott, R.M. A prospective study of dengue infections in Bangkok. Am. J. Trop. Med. Hyg. 1988, 38, 172–180. [Google Scholar] [CrossRef] [PubMed]
- Halstead, S.B. Observations related to pathogensis of dengue hemorrhagic fever. VI. Hypotheses and discussion. Yale J. Biol. Med. 1970, 42, 350–362. [Google Scholar]
- Sabin, A.B. Research on dengue during World War II. Am. J. Trop. Med. Hyg. 1952, 1, 30–50. [Google Scholar] [CrossRef]
- Montoya, M.; Gresh, L.; Mercado, J.C.; Williams, K.L.; Vargas, M.J.; Gutierrez, G.; Kuan, G.; Gordon, A.; Balmaseda, A.; Harris, E. Symptomatic Versus Inapparent Outcome in Repeat Dengue Virus Infections Is Influenced by the Time Interval between Infections and Study Year. PLoS Negl. Trop. Dis. 2013, 7, e2357. [Google Scholar] [CrossRef]
- Anderson, K.B.; Gibbons, R.V.; Cummings, D.A.T.; Nisalak, A.; Green, S.; Libraty, D.H.; Jarman, R.G.; Srikiatkhachorn, A.; Mammen, M.P.; Darunee, B.; et al. A Shorter Time Interval Between First and Second Dengue Infections Is Associated With Protection From Clinical Illness in a School-based Cohort in Thailand. J. Infect Dis. 2014, 209, 360–368. [Google Scholar] [CrossRef]
- Salje, H.; Lessler, J.; Endy, T.P.; Curriero, F.C.; Gibbons, R.V.; Nisalak, A.; Nimmannitya, S.; Kalayanarooj, S.; Jarman, R.G.; Thomas, S.J.; et al. Revealing the microscale spatial signature of dengue transmission and immunity in an urban population. Proc. Natl. Acad. Sci. USA 2012, 109, 9535–9538. [Google Scholar] [CrossRef]
- Reich, N.G.; Shrestha, S.; King, A.A.; Rohani, P.; Lessler, J.; Kalayanarooj, S.; Yoon, I.-K.; Gibbons, R.V.; Burke, D.S.; Cummings, D.A.T. Interactions between serotypes of dengue highlight epidemiological impact of cross-immunity. J. R. Soc. Interface 2013, 10, 20130414. [Google Scholar] [CrossRef]
- Vaughn, D.W.; Green, S.; Kalayanarooj, S.; Innis, B.L.; Nimmannitya, S.; Suntayakorn, S.; Endy, T.P.; Raengsakulrach, B.; Rothman, A.L.; Ennis, F.A.; et al. Dengue viremia titer, antibody response pattern, and virus serotype correlate with disease severity. J. Infect Dis. 2000, 181, 2–9. [Google Scholar] [CrossRef]
- Libraty, D.H.; Endy, T.P.; Houng, H.S.H.; Green, S.; Kalayanarooj, S.; Suntayakorn, S.; Chansiriwongs, W.; Vaughn, D.W.; Nisalak, A.; Ennis, F.A.; et al. Differing influences of virus burden and immune activation on disease severity in secondary dengue-3 virus infections. J. Infect Dis. 2002, 185, 1213–1221. [Google Scholar] [CrossRef] [PubMed]
- Kliks, S.C.; Nimmanitya, S.; Nisalak, A.; Burke, D.S. Evidence that maternal dengue antibodies are important in the development of dengue hemorrhagic fever in infants. Am. J. Trop. Med. Hyg. 1988, 38, 411–419. [Google Scholar] [CrossRef] [PubMed]
- Halstead, S.B.; O’Rourke, E.J.; Allison, A.C. Dengue viruses and mononuclear phagocytes. II. Identity of blood and tissue leukocytes supporting in vitro infection. J. Expe. Med. 1977, 146, 218–229. [Google Scholar]
- Guzman, M.G.; Vazquez, S. The Complexity of Antibody-Dependent Enhancement of Dengue Virus Infection. Viruses 2010, 2, 2649–2662. [Google Scholar] [CrossRef] [PubMed]
- Dejnirattisai, W.; Jumnainsong, A.; Onsirisakul, N.; Fitton, P.; Vasanawathana, S.; Limpitikul, W.; Puttikhunt, C.; Edwards, C.; Duangchinda, T.; Supasa, S.; et al. Cross-reacting antibodies enhance dengue virus infection in humans. Science 2010, 328, 745–748. [Google Scholar] [CrossRef]
- Beltramello, M.; Williams, K.L.; Simmons, C.P.; Macagno, A.; Simonelli, L.; Quyen, N.T.H.; Sukupolvi-Petty, S.; Navarro-Sanchez, E.; Young, P.R.; de Silva, A.M.; et al. The Human Immune Response to Dengue Virus Is Dominated by Highly Cross-Reactive Antibodies Endowed with Neutralizing and Enhancing Activity. Cell Host Microbe 2010, 8, 271–283. [Google Scholar] [CrossRef] [PubMed]
- Halstead, S.B. In vivo enhancement of dengue virus infection in rhesus monkeys by passively transferred antibody. J. Infect Dis. 1979, 140, 527–533. [Google Scholar] [CrossRef]
- Goncalvez, A.P.; Engle, R.E.; St Claire, M.; Purcell, R.H.; Lai, C.-J. Monoclonal antibody-mediated enhancement of dengue virus infection in vitro and in vivo and strategies for prevention. Proc. Natl. Acad. Sci. USA 2007, 104, 9422–9427. [Google Scholar] [CrossRef]
- Dung, N.T.P.; Duyen, H.T.L.; Thuy, N.T.V.; Ngoc, T.V.; Chau, N.V.V.; Hien, T.T.; Rowland-Jones, S.L.; Dong, T.; Farrar, J.; Wills, B.; et al. Timing of CD8+ T cell responses in relation to commencement of capillary leakage in children with dengue. J. Immunol. 2010, 184, 7281–7287. [Google Scholar] [CrossRef]
- Flipse, J.; Diosa-Toro, M.A.; Hoornweg, T.E.; van de Pol, D.P.I.; Urcuqui-Inchima, S.; Smit, J.M. Antibody-Dependent Enhancement of Dengue Virus Infection in Primary Human Macrophages; Balancing Higher Fusion against Antiviral Responses. Sci. Rep. 2016, 6, 465. [Google Scholar] [CrossRef]
- Modhiran, N.; Kalayanarooj, S.; Ubol, S. Subversion of Innate Defenses by the Interplay between DENV and Pre-Existing Enhancing Antibodies: TLRs Signaling Collapse. PLoS Negl. Trop. Dis. 2010, 4, e924. [Google Scholar] [CrossRef] [PubMed]
- Chareonsirisuthigul, T.; Kalayanarooj, S.; Ubol, S. Dengue virus (DENV) antibody-dependent enhancement of infection upregulates the production of anti-inflammatory cytokines, but suppresses anti-DENV free radical and pro-inflammatory cytokine production, in THP-1 cells. J. Gen. Virol. 2007, 88, 365–375. [Google Scholar] [CrossRef] [PubMed]
- Ubol, S.; Phuklia, W.; Kalayanarooj, S.; Modhiran, N. Mechanisms of Immune Evasion Induced by a Complex of Dengue Virus and Preexisting Enhancing Antibodies. J. Infect. Dis. 2010, 201, 923–935. [Google Scholar] [CrossRef]
- Kou, Z.; Lim, J.Y.H.; Beltramello, M.; Quinn, M.; Chen, H.; Liu, S.-N.; Martnez-Sobrido, L.; Diamond, M.S.; Schlesinger, J.J.; de Silva, A.; et al. Human antibodies against dengue enhance dengue viral infectivity without suppressing type I interferon secretion in primary human monocytes. Virology 2011, 410, 240–247. [Google Scholar] [CrossRef] [PubMed]
- Boonnak, K.; Dambach, K.M.; Donofrio, G.C.; Tassaneetrithep, B.; Marovich, M.A. Cell Type Specificity and Host Genetic Polymorphisms Influence Antibody-Dependent Enhancement of Dengue Virus Infection. J. Virol. 2011, 85, 1671–1683. [Google Scholar] [CrossRef]
- Modhiran, N.; Watterson, D.; Muller, D.A.; Panetta, A.K.; Sester, D.P.; Liu, L.; Hume, D.A.; Stacey, K.J.; Young, P.R. Dengue virus NS1 protein activates cells via Toll-like receptor 4 and disrupts endothelial cell monolayer integrity. Sci. Transl. Med. 2015, 7, 304ra142. [Google Scholar] [CrossRef]
- Beatty, P.R.; Puerta-Guardo, H.; Killingbeck, S.S.; Glasner, D.R.; Hopkins, K.; Harris, E. Dengue virus NS1 triggers endothelial permeability and vascular leak that is prevented by NS1 vaccination. Sci. Transl. Med. 2015, 7, 304ra141. [Google Scholar] [CrossRef]
- Ng, J.K.W.; Zhang, S.L.; Tan, H.C.; Yan, B.; Gomez, J.M.M.; Tan, W.Y.; Lam, J.H.; Tan, G.K.X.; Ooi, E.E.; Alonso, S. First Experimental In Vivo Model of Enhanced Dengue Disease Severity through Maternally Acquired Heterotypic Dengue Antibodies. PLoS Pathog 2014, 10, e1004031. [Google Scholar] [CrossRef]
- Hober, D.; Poli, L.; Roblin, B.; Gestas, P.; Chungue, E.; Granic, G.; Imbert, P.; Pecarere, J.L.; Vergez-Pascal, R.; Wattre, P. Serum levels of tumor necrosis factor-alpha (TNF-alpha), interleukin-6 (IL-6), and interleukin-1 beta (IL-1 beta) in dengue-infected patients. Am. J. Trop. Med. Hyg. 1993, 48, 324–331. [Google Scholar] [CrossRef]
- Green, S.; Vaughn, D.W.; Kalayanarooj, S.; Nimmannitya, S.; Suntayakorn, S.; Nisalak, A.; Lew, R.; Innis, B.L.; Kurane, I.; Rothman, A.L.; et al. Early Immune Activation in Acute Dengue Illness Is Related to Development of Plasma Leakage and Disease Severity. J. Infect Dis. 1999, 179, 755–762. [Google Scholar] [CrossRef]
- Lok, S.-M.; Kostyuchenko, V.; Nybakken, G.E.; Holdaway, H.A.; Battisti, A.J.; Sukupolvi-Petty, S.; Sedlak, D.; Fremont, D.H.; Chipman, P.R.; Roehrig, J.T.; et al. Binding of a neutralizing antibody to dengue virus alters the arrangement of surface glycoproteins. Nat. Struct. Mol. Biol. 2008, 15, 312–317. [Google Scholar] [CrossRef] [PubMed]
- Pierson, T.C.; Diamond, M.S. Molecular mechanisms of antibody-mediated neutralisation of flavivirus infection. Expert Rev. Mol. Med. 2008, 10, e12. [Google Scholar] [CrossRef] [PubMed]
- Halstead, S.B. Neutralization and antibody-dependent enhancement of dengue viruses. Adv. Virus Res. 2003, 60, 421–467. [Google Scholar] [PubMed]
- Halstead, S.B.; Venkateshan, C.N.; Gentry, M.K.; Larsen, L.K. Heterogeneity of infection enhancement of dengue 2 strains by monoclonal antibodies. J. Immunol. 1984, 132, 1529–1532. [Google Scholar]
- Tsai, W.-Y.; Durbin, A.; Tsai, J.-J.; Hsieh, S.-C.; Whitehead, S.; Wang, W.-K. Complexity of Neutralizing Antibodies against Multiple Dengue Virus Serotypes after Heterotypic Immunization and Secondary Infection Revealed by In-Depth Analysis of Cross-Reactive Antibodies. J. Virol. 2015, 89, 7348–7362. [Google Scholar] [CrossRef]
- Kliks, S.C.; Nisalak, A.; Brandt, W.E.; Wahl, L.; Burke, D.S. Antibody-dependent enhancement of dengue virus growth in human monocytes as a risk factor for dengue hemorrhagic fever. Am. J. Trop. Med. Hyg. 1989, 40, 444–451. [Google Scholar] [CrossRef]
- Rodenhuis-Zybert, I.A.; van der Schaar, H.M.; da Silva Voorham, J.M.; van der Ende-Metselaar, H.; Lei, H.-Y.; Wilschut, J.; Smit, J.M. Immature Dengue Virus: A Veiled Pathogen? PLoS Pathog 2010, 6, e1000718. [Google Scholar] [CrossRef]
- Rodenhuis-Zybert, I.A.; Moesker, B.; da Silva Voorham, J.M.; van der Ende-Metselaar, H.; Diamond, M.S.; Wilschut, J.; Smit, J.M. A fusion-loop antibody enhances the infectious properties of immature flavivirus particles. J. Virol. 2011, 85, 11800–11808. [Google Scholar] [CrossRef]
- Lai, C.Y.; Tsai, W.Y.; Lin, S.R.; Kao, C.L.; Hu, H.P.; King, C.C.; Wu, H.C.; Chang, G.J.; Wang, W.K. Antibodies to Envelope Glycoprotein of Dengue Virus during the Natural Course of Infection Are Predominantly Cross-Reactive and Recognize Epitopes Containing Highly Conserved Residues at the Fusion Loop of Domain II. J. Virol. 2008, 82, 6631–6643. [Google Scholar] [CrossRef]
- De Alwis, R.; Smith, S.A.; Olivarez, N.P.; Messer, W.B.; Huynh, J.P.; Wahala, W.M.P.B.; White, L.J.; Diamond, M.S.; Baric, R.S.; Crowe, J.E.; et al. Identification of human neutralizing antibodies that bind to complex epitopes on dengue virions. Proc. Natl. Acad. Sci. USA 2012, 109, 7439–7444. [Google Scholar] [CrossRef]
- Teoh, E.P.; Kukkaro, P.; Teo, E.W.; Lim, A.P.C.; Tan, T.T.; Yip, A.; Schul, W.; Aung, M.; Kostyuchenko, V.A.; Leo, Y.S.; et al. The Structural Basis for Serotype-Specific Neutralization of Dengue Virus by a Human Antibody. Sci. Transl. Med. 2012, 4, 139ra83. [Google Scholar] [CrossRef] [PubMed]
- Fibriansah, G.; Tan, J.L.; Smith, S.A.; de Alwis, R.; Ng, T.-S.; Kostyuchenko, V.A.; Jadi, R.S.; Kukkaro, P.; de Silva, A.M.; Crowe, J.E.; et al. A highly potent human antibody neutralizes dengue virus serotype 3 by binding across three surface proteins. Nat. Commun. 2015, 6, 6341. [Google Scholar] [CrossRef] [PubMed]
- Rouvinski, A.; Guardado-Calvo, P.; Barba-Spaeth, G.; Duquerroy, S.; Vaney, M.-C.; Kikuti, C.M.; Navarro Sanchez, M.E.; Dejnirattisai, W.; Wongwiwat, W.; Haouz, A.; et al. Recognition determinants of broadly neutralizing human antibodies against dengue viruses. Nature 2015, 520, 109–113. [Google Scholar] [CrossRef] [PubMed]
- Dejnirattisai, W.; Wongwiwat, W.; Supasa, S.; Zhang, X.; Dai, X.; Rouvinsky, A.; Jumnainsong, A.; Edwards, C.; Quyen, N.T.H.; Duangchinda, T.; et al. A new class of highly potent, broadly neutralizing antibodies isolated from viremic patients infected with dengue virus. Nat. Immunol. 2014, 16, 170–177. [Google Scholar] [CrossRef]
- De Alwis, R.; Beltramello, M.; Messer, W.B.; Sukupolvi-Petty, S.; Wahala, W.M.P.B.; Kraus, A.; Olivarez, N.P.; Pham, Q.; Brian, J.; Tsai, W.-Y.; et al. In-Depth Analysis of the Antibody Response of Individuals Exposed to Primary Dengue Virus Infection. PLoS Negl. Trop. Dis. 2011, 5, e1188. [Google Scholar] [CrossRef]
- Avirutnan, P.; Punyadee, N.; Noisakran, S.; Komoltri, C.; Thiemmeca, S.; Auethavornanan, K.; Jairungsri, A.; Kanlaya, R.; Tangthawornchaikul, N.; Puttikhunt, C.; et al. Vascular Leakage in Severe Dengue Virus Infections: A Potential Role for the Nonstructural Viral Protein NS1 and Complement. J. Infect. Dis. 2006, 193, 1078–1088. [Google Scholar] [CrossRef]
- Sun, D.S.; King, C.C.; Huang, H.S.; Shih, Y.L.; Lee, C.C.; Tsai, W.J.; Yu, C.C.; Chang, H.H. Antiplatelet autoantibodies elicited by dengue virus non-structural protein 1 cause thrombocytopenia and mortality in mice. J. Throm. Haemostasis 2007, 5, 2291–2299. [Google Scholar] [CrossRef]
- Lin, C.F.; Lei, H.Y.; Liu, C.C.; Liu, H.S.; Yeh, T.M.; Wang, S.T.; Yang, T.I.; Sheu, F.C.; Kuo, C.F.; Lin, Y.S. Generation of IgM anti-platelet autoantibody in dengue patients. J. Med. Virol. 2001, 63, 143–149. [Google Scholar] [CrossRef]
- Falconar, A.K.I. The dengue virus nonstructural-1 protein (NS1) generatesantibodies to common epitopes on human blood clotting, integrin/adhesin proteins and binds to humanendothelial cells: Potential implications in haemorrhagic fever pathogenesis. Arch Virol. 2013, 142, 897–916. [Google Scholar] [CrossRef]
- Chuang, Y.-C.; Wang, S.-Y.; Lin, Y.-S.; Chen, H.-R.; Yeh, T.-M. Re-evaluation of the pathogenic roles of nonstructural protein 1 and its antibodies during dengue virus infection. J. Biomed. Sci. 2013, 20, 42. [Google Scholar] [CrossRef]
- Churdboonchart, V.; Bhamarapravati, N.; Peampramprecha, S.; Sirinavin, S. Antibodies against dengue viral proteins in primary and secondary dengue hemorrhagic fever. Am. J. Trop. Med. Hyg. 1991, 44, 481–493. [Google Scholar] [CrossRef] [PubMed]
- Valdés, K.; Alvarez, M.; Pupo, M.; Vázquez, S.; Rodríguez, R.; Guzmán, M.G. Human Dengue antibodies against structural and nonstructural proteins. Clin. Dia. Lab. Imm. 2000, 7, 856–857. [Google Scholar] [CrossRef] [PubMed]
- Vazeille, M.; Gaborit, P.; Mousson, L.; Girod, R.; Failloux, A.B. Competitive advantage of a dengue 4 virus when co-infecting the mosquito Aedes aegypti with a dengue 1 virus. BMC Infect. Dis. 2016 16:1 2016, 16, 318. [Google Scholar] [CrossRef] [PubMed]
- Sabchareon, A.; Lang, J.; Chanthavanich, P.; Yoksan, S.; Forrat, R.; Attanath, P.; Sirivichayakul, C.; Pengsaa, K.; Pojjaroen-Anant, C.; Chokejindachai, W.; et al. Safety and immunogenicity of tetravalent live-attenuated dengue vaccines in Thai adult volunteers: Role of serotype concentration, ratio, and multiple doses. Am. J. Trop. Med. Hyg. 2002, 66, 264–272. [Google Scholar] [CrossRef]
- Weiskopf, D.; Sette, A. T-Cell Immunity to Infection with Dengue Virus in Humans. Front. Immunol. 2014, 5, 571. [Google Scholar] [CrossRef]
- Weiskopf, D.; Angelo, M.A.; de Azeredo, E.L.; Sidney, J.; Greenbaum, J.A.; Fernando, A.N.; Broadwater, A.; Kolla, R.V.; De Silva, A.D.; de Silva, A.M.; et al. Comprehensive analysis of dengue virus-specific responses supports an HLA-linked protective role for CD8+ T cells. Proc. Natl. Acad. Sci. USA 2013, 110, E2046–E2053. [Google Scholar] [CrossRef]
- Simmons, C.P.; Dong, T.; Chau, N.V.; Dung, N.T.P.; Chau, T.N.B.; Thao, L.T.T.; Dung, N.T.; Hien, T.T.; Rowland-Jones, S.; Farrar, J. Early T-cell responses to dengue virus epitopes in Vietnamese adults with secondary dengue virus infections. J. Virol. 2005, 79, 5665–5675. [Google Scholar] [CrossRef]
- Duangchinda, T.; Dejnirattisai, W.; Vasanawathana, S.; Limpitikul, W.; Tangthawornchaikul, N.; Malasit, P.; Mongkolsapaya, J.; Screaton, G. Immunodominant T-cell responses to dengue virus NS3 are associated with DHF. Proc. Natl. Acad. Sci. USA 2010, 107, 16922–16927. [Google Scholar] [CrossRef]
- Weiskopf, D.; Yauch, L.E.; Angelo, M.A.; John, D.V.; Greenbaum, J.A.; Sidney, J.; Kolla, R.V.; De Silva, A.D.; de Silva, A.M.; Grey, H.; et al. Insights into HLA-Restricted T Cell Responses in a Novel Mouse Model of Dengue Virus Infection Point toward New Implications for Vaccine Design. J. Immunol. 2011, 187, 4268–4279. [Google Scholar] [CrossRef]
- Rivino, L.; Kumaran, E.A.P.; Jovanovic, V.; Nadua, K.; Teo, E.W.; Pang, S.W.; Teo, G.H.; Gan, V.C.H.; Lye, D.C.; Leo, Y.S.; et al. Differential Targeting of Viral Components by CD4+ versus CD8+ T Lymphocytes in Dengue Virus Infection. J. Virol. 2013, 87, 2693–2706. [Google Scholar] [CrossRef]
- Tian, Y.; Grifoni, A.; Sette, A.; Weiskopf, D. Human T Cell Response to Dengue Virus Infection. Front. Immunol. 2019, 10, 1043. [Google Scholar] [CrossRef] [PubMed]
- Cedillo-Barrón, L.; García-Cordero, J.; Bustos-Arriaga, J.; León-Juárez, M.; Gutiérrez-Castañeda, B. Antibody response to dengue virus. Microbes Infect. 2014, 16, 711–720. [Google Scholar] [CrossRef] [PubMed]
- Yu, L.; Nomaguchi, M.; Padmanabhan, R.; Markoff, L. Specific requirements for elements of the 5’ and 3’ terminal regions in flavivirus RNA synthesis and viral replication. Virology 2008, 374, 170–185. [Google Scholar] [CrossRef] [PubMed]
- Alvarez, D.E.; Lodeiro, M.F.; Luduena, S.J.; Pietrasanta, L.I.; Gamarnik, A.V. Long-Range RNA-RNA Interactions Circularize the Dengue Virus Genome. J. Virol. 2005, 79, 6631–6643. [Google Scholar] [CrossRef]
- Byk, L.A.; Gamarnik, A.V. Properties and Functions of the Dengue Virus Capsid Protein. Annu. Rev. 2016, 3, 263–281. [Google Scholar] [CrossRef]
- Wong, S.-S.; Haqshenas, G.; Gowans, E.J.; Mackenzie, J. The dengue virus M protein localises to the endoplasmic reticulum and forms oligomers. FEBS Lett. 2012, 586, 1032–1037. [Google Scholar] [CrossRef]
- Leung, D.; Schroder, K.; White, H.; Fang, N.X.; Stoermer, M.J.; Abbenante, G.; Martin, J.L.; Young, P.R.; Fairlie, D.P. Activity of recombinant dengue 2 virus NS3 protease in the presence of a truncated NS2B co-factor, small peptide substrates, and inhibitors. J. Biol. Chem. 2001, 276, 45762–45771. [Google Scholar] [CrossRef]
- Perera, R.; Kuhn, R.J. Structural proteomics of dengue virus. Curr. Opin. Microbiol. 2008, 11, 369–377. [Google Scholar] [CrossRef]
- Umareddy, I.; Chao, A.; Sampath, A.; Gu, F.; Vasudevan, S.G. Dengue virus NS4B interacts with NS3 and dissociates it from single-stranded RNA. J. Gen. Virol. 2006, 87, 2605–2614. [Google Scholar] [CrossRef]
- Egloff, M.-P.; Benarroch, D.; Selisko, B.; Romette, J.-L.; Canard, B. An RNA cap (nucleoside-2′-O-)-methyltransferase in the flavivirus RNA polymerase NS5: Crystal structure and functional characterization. EMBO J. 2002, 21, 2757–2768. [Google Scholar] [CrossRef]
- Mazzon, M.; Jones, M.; Davidson, A.; Chain, B.; Jacobs, M. Dengue virus NS5 inhibits interferon-alpha signaling by blocking signal transducer and activator of transcription 2 phosphorylation. J. Infect. Dis. 2009, 200, 1261–1270. [Google Scholar] [CrossRef] [PubMed]
- Hahn, C.S.; Hahn, Y.S.; Rice, C.M.; Lee, E.; Dalgarno, L.; Strauss, E.G.; Strauss, J.H. Conserved elements in the 3’ untranslated region of flavivirus RNAs and potential cyclization sequences. J. Mol. Biol. 1987, 198, 33–41. [Google Scholar] [CrossRef]
- Vita, R.; Overton, J.A.; Greenbaum, J.A.; Ponomarenko, J.; Clark, J.D.; Cantrell, J.R.; Wheeler, D.K.; Gabbard, J.L.; Hix, D.; Sette, A.; et al. The immune epitope database (IEDB) 3.0. Nucleic Acids Res. 2015, 43, D405–D412. [Google Scholar] [CrossRef] [PubMed]
- Murphy, K.; Travers, P.; Walport, M. Janeway’s Immunobiology, 7th ed.; Goatly, B., Ed.; Garland Science: New York, NY, USA, 2008. [Google Scholar]
- Remakus, S.; Sigal, L.J. Memory CD8+ T Cell Protection. In Crossroads Between Innate and Adaptive Immunity IV; Advances in Experimental Medicine and Biology; Springer: New York, NY, USA, 2013; Volume 785, pp. 77–86. [Google Scholar]
- Sant, A.J.; McMichael, A. Revealing the role of CD4(+) T cells in viral immunity. J. Exp. Med. 2012, 209, 1391–1395. [Google Scholar]
- Mongkolsapaya, J.; Dejnirattisai, W.; Xu, X.-N.; Vasanawathana, S.; Tangthawornchaikul, N.; Chairunsri, A.; Sawasdivorn, S.; Duangchinda, T.; Dong, T.; Rowland-Jones, S.; et al. Original antigenic sin and apoptosis in the pathogenesis of dengue hemorrhagic fever. Nat. Med. 2003, 9, 921–927. [Google Scholar] [CrossRef]
- Friberg, H.; Bashyam, H.; Toyosaki-Maeda, T.; Potts, J.A.; Greenough, T.; Kalayanarooj, S.; Gibbons, R.V.; Nisalak, A.; Srikiatkhachorn, A.; Green, S.; et al. Cross-Reactivity and Expansion of Dengue-Specific T cells During Acute Primary and Secondary Infections in Humans. Sci. Rep. 2011, 1, 2. [Google Scholar] [CrossRef]
- Zellweger, R.M.; Miller, R.; Eddy, W.E.; White, L.J.; Johnston, R.E.; Shresta, S. Role of Humoral versus Cellular Responses Induced by a Protective Dengue Vaccine Candidate. PLoS Pathog 2013, 9, e1003723. [Google Scholar] [CrossRef]
- Hatch, S.; Endy, T.P.; Thomas, S.; Mathew, A.; Potts, J.; Pazoles, P.; Libraty, D.H.; Gibbons, R.; Rothman, A.L. Intracellular Cytokine Production by Dengue Virus-specific T cells Correlates with Subclinical Secondary Infection. J. Infect. Dis. 2011, 203, 1282–1291. [Google Scholar] [CrossRef]
- Yauch, L.E.; Zellweger, R.M.; Kotturi, M.F.; Qutubuddin, A.; Sidney, J.; Peters, B.; Prestwood, T.R.; Sette, A.; Shresta, S. A Protective Role for Dengue Virus-Specific CD8+ T Cells. J. Immunol. 2009, 182, 4865–4873. [Google Scholar] [CrossRef]
- Mongkolsapaya, J.; Duangchinda, T.; Dejnirattisai, W.; Vasanawathana, S.; Avirutnan, P.; Jairungsri, A.; Khemnu, N.; Tangthawornchaikul, N.; Chotiyarnwong, P.; Sae-Jang, K.; et al. T Cell Responses in Dengue Hemorrhagic Fever: Are Cross-Reactive T Cells Suboptimal? J. Immunol. 2006, 176, 3821–3829. [Google Scholar] [CrossRef]
- Mathew, A.; Rothman, A.L. Understanding the contribution of cellular immunity to dengue disease pathogenesis. Immunol. Rev. 2008, 225, 300–313. [Google Scholar] [CrossRef] [PubMed]
- Kurane, I.; Rothman, A.L.; Livingston, P.G.; Green, S.; Gagnon, S.J.; Janus, J.; Innis, B.L.; Nimmannitya, S.; Nisalak, A.; Ennis, F.A. Immunopathologic mechanisms of dengue hemorrhagic fever and dengue shock syndrome. Arch. Virol. Suppl. 1994, 9, 59–64. [Google Scholar] [PubMed]
- Mangada, M.M.; Endy, T.P.; Nisalak, A.; Chunsuttiwat, S.; Vaughn, D.W.; Libraty, D.H.; Green, S.; Ennis, F.A.; Rothman, A.L. Dengue-specific T cell responses in peripheral blood mononuclear cells obtained prior to secondary dengue virus infections in Thai schoolchildren. J. Infect. Dis 2002, 185, 1697–1703. [Google Scholar] [CrossRef] [PubMed]
- Green, S.; Pichyangkul, S.; Vaughn, D.W.; Kalayanarooj, S.; Nimmannitya, S.; Nisalak, A.; Kurane, I.; Rothman, A.L.; Ennis, F.A. Early CD69 Expression on Peripheral Blood Lymphocytes from Children with Dengue Hemorrhagic Fever. J. Infect. Dis. 1999, 180, 1429–1435. [Google Scholar] [CrossRef] [PubMed]
- Friberg, H.; Burns, L.; Woda, M.; Kalayanarooj, S.; Endy, T.P.; Stephens, H.A.F.; Green, S.; Rothman, A.L.; Mathew, A. Memory CD8+ T cells from naturally acquired primary dengue virus infection are highly cross-reactive. Immunol. Cell Biol. 2011, 89, 122–129. [Google Scholar] [CrossRef]
- Almeida, J.R.; Sauce, D.; Price, D.A.; Papagno, L.; Shin, S.Y.; Moris, A.; Larsen, M.; Pancino, G.; Douek, D.C.; Autran, B.; et al. Antigen sensitivity is a major determinant of CD8+ T-cell polyfunctionality and HIV-suppressive activity. Blood 2009, 113, 6351–6360. [Google Scholar] [CrossRef] [PubMed]
- Park, S.H.; Shin, E.C.; Capone, S.; Caggiari, L.; Re, V.D.; Nicosia, A.; Folgori, A.; Rehermann, B. Successful Vaccination Induces Multifunctional Memory T-Cell Precursors Associated With Early Control of Hepatitis C Virus. Gastroenterology 2012, 143, 1048–1060.e4. [Google Scholar] [CrossRef]
- Yachi, P.P.; Ampudia, J.; Zal, T.; Gascoigne, N.R.J. Altered Peptide Ligands Induce Delayed CD8-T Cell Receptor Interaction—a Role for CD8 in Distinguishing Antigen Quality. Immunity 2006, 25, 203–211. [Google Scholar] [CrossRef]
- Rothman, A.L. Immunity to dengue virus: A tale of original antigenic sin and tropical cytokine storms. Nat. Rev. Immunol 2011, 11, 532–543. [Google Scholar] [CrossRef]
- Weaver, S.C.; Vasilakis, N. Molecular evolution of dengue viruses: Contributions of phylogenetics to understanding the history and epidemiology of the preeminent arboviral disease. Infect. Genet. Evol. 2009, 9, 523–540. [Google Scholar] [CrossRef]
- Twiddy, S.S.; Farrar, J.J.; Vinh Chau, N.; Wills, B.; Gould, E.A.; Gritsun, T.; Lloyd, G.; Holmes, E.C. Phylogenetic relationships and differential selection pressures among genotypes of dengue-2 virus. Virology 2002, 298, 63–72. [Google Scholar] [CrossRef] [PubMed]
- Zhang, C.; Mammen, M.P.; Chinnawirotpisan, P.; Klungthong, C.; Rodpradit, P.; Monkongdee, P.; Nimmannitya, S.; Kalayanarooj, S.; Holmes, E.C. Clade Replacements in Dengue Virus Serotypes 1 and 3 Are Associated with Changing Serotype Prevalence. J. Virol. 2005, 79, 15123–15130. [Google Scholar] [CrossRef] [PubMed]
- Nuegoonpipat, A.; Berlioz-Arthaud, A.; Chow, V.; Endy, T.; Lowry, K.; Mai, L.Q.; Ninh, T.U.; Pyke, A.; Reid, M.; Reynes, J.-M.; et al. Sustained transmission of dengue virus type 1 in the Pacific due to repeated introductions of different Asian strains. Virology 2004, 329, 505–512. [Google Scholar] [CrossRef] [PubMed]
- OhAinle, M.; Balmaseda, A.; Macalalad, A.R.; Tellez, Y.; Zody, M.C.; Saborío, S.; Nuñez, A.; Lennon, N.J.; Birren, B.W.; Gordon, A.; et al. Dynamics of Dengue Disease Severity Determined by the Interplay Between Viral Genetics and Serotype-Specific Immunity. Sci. Transl. Med. 2011, 3, 114ra128. [Google Scholar] [CrossRef] [PubMed]
- Lambrechts, L.; Fansiri, T.; Pongsiri, A.; Thaisomboonsuk, B.; Klungthong, C.; Richardson, J.H.; Ponlawat, A.; Jarman, R.G.; Scott, T.W. Dengue-1 Virus Clade Replacement in Thailand Associated with Enhanced Mosquito Transmission. J. Virol. 2012, 86, 1853–1861. [Google Scholar] [CrossRef]
- Wittke, V.; Robb, T.E.; Thu, H.M.; Nisalak, A.; Nimmannitya, S.; Kalayanrooj, S.; Vaughn, D.W.; Endy, T.P.; Holmes, E.C.; Aaskov, J.G. Extinction and Rapid Emergence of Strains of Dengue 3 Virus during an Interepidemic Period. Virology 2002, 301, 148–156. [Google Scholar] [CrossRef]
- Rico-Hesse, R. Dengue Virus Virulence and Transmission Determinants. In Dengue Virus; Current Topics in Microbiology and Immunology; Springer: Berlin/Heidelberg, Germany, 2009; Volume 338, pp. 45–55. [Google Scholar]
- Hang, V.T.T.; Holmes, E.C.; Veasna, D.; Quy, N.T.; Hien, T.T.; Quail, M.; Churcher, C.; Parkhill, J.; Cardosa, J.; Farrar, J.; et al. Emergence of the Asian 1 Genotype of Dengue Virus Serotype 2 in Viet Nam: In Vivo Fitness Advantage and Lineage Replacement in South-East Asia. PLoS Negl. Trop. Dis. 2010, 4, e757. [Google Scholar] [CrossRef]
- Costa, R.L.; Voloch, C.M.; Schrago, C.G. Comparative evolutionary epidemiology of dengue virus serotypes. Infect. Genet. Evol. 2012, 12, 309–314. [Google Scholar] [CrossRef]
- King, C.-C.; Chao, D.-Y.; Chien, L.-J.; Chang, G.-J.; Ling, T.-H.; Wu, Y.-C.; Huang, J.-H. Comparative analysis of full genomic sequences among different genotypes of dengue virus type 3. Virol. J. 2008, 5, 63. [Google Scholar] [CrossRef]
- Adams, B.; Holmes, E.C.; Zhang, C.; Mammen, M.P.; Nimmannitya, S.; Kalayanarooj, S.; Boots, M. Cross-protective immunity can account for the alternating epidemic pattern of dengue virus serotypes circulating in Bangkok. Proc. Natl. Acad. Sci. USA 2006, 103, 14234–14239. [Google Scholar] [CrossRef]
- Holmes, E.C.; Twiddy, S.S.; Woelk, C.H. Phylogenetic evidence for adaptive evolution of dengue viruses in nature. J. Gen. Virol. 2002, 83, 1679–1689. [Google Scholar]
- Wearing, H.J.; Rohani, P. Ecological and immunological determinants of dengue epidemics. Proc. Natl. Acad. Sci. USA 2006, 103, 11802–11807. [Google Scholar] [CrossRef] [PubMed]
- Adams, B.; Boots, M. Modelling the relationship between antibody-dependent enhancement and immunological distance with application to dengue. J. Theor. Biol. 2006, 242, 337–346. [Google Scholar] [CrossRef] [PubMed]
- Recker, M.; Blyuss, K.B.; Simmons, C.P.; Hien, T.T.; Wills, B.; Farrar, J.; Gupta, S. Immunological serotype interactions and their effect on the epidemiological pattern of dengue. Proc. R. Soc. Lond. B Biol. Sci. 2009, 276, 2541–2548. [Google Scholar] [CrossRef] [PubMed]
- Cologna, R.; Armstrong, P.M.; Rico-Hesse, R. Selection for Virulent Dengue Viruses Occurs in Humans and Mosquitoes. J. Virol. 2004, 79, 853–859. [Google Scholar] [CrossRef]
- Armstrong, P.M.; Rico-Hesse, R. Efficiency of dengue serotype 2 virus strains to infect and disseminate in Aedes aegypti. Am. J. Trop. Med. Hyg. 2003, 68, 539–544. [Google Scholar] [CrossRef]
- Armstrong, P.M.; Rico-Hesse, R. Differential susceptibility of Aedes aegypti to infection by the American and Southeast Asian genotypes of dengue type 2 virus. Vector-Borne Zoonotic Dis. 2001, 1, 159–168. [Google Scholar] [CrossRef]
- Libraty, D.H.; Young, P.R.; Pickering, D.; Endy, T.P.; Kalayanarooj, S.; Green, S.; Vaughn, D.W.; Nisalak, A.; Ennis, F.A.; Rothman, A.L. High Circulating Levels of the Dengue Virus Nonstructural Protein NS1 Early in Dengue Illness Correlate with the Development of Dengue Hemorrhagic Fever. J. Infect. Dis. 2002, 186, 1165–1168. [Google Scholar] [CrossRef]
- Bennett, S.N. Selection-Driven Evolution of Emergent Dengue Virus. Mol. Biol. Evol. 2003, 20, 1650–1658. [Google Scholar] [CrossRef]
- Vasilakis, N.; Holmes, E.C.; Fokam, E.B.; Faye, O.; Diallo, M.; Sall, A.A.; Weaver, S.C. Evolutionary Processes among Sylvatic Dengue Type 2 Viruses. J. Virol. 2007, 81, 9591–9595. [Google Scholar] [CrossRef]
- Holmes, E.C. Patterns of intra- and interhost nonsynonymous variation reveal strong purifying selection in dengue virus. J. Virol. 2003, 77, 11296–11298. [Google Scholar] [CrossRef] [PubMed]
- Holmes, E.C.; Twiddy, S.S. The origin, emergence and evolutionary genetics of dengue virus. Infect. Genet. Evol. 2003, 3, 19–28. [Google Scholar] [CrossRef]
- Foster, J.E.; Bennett, S.N.; Carrington, C.V.F.; Vaughan, H.; McMillan, W.O. Phylogeography and molecular evolution of dengue 2 in the Caribbean basin, 1981–2000. Virology 2004, 324, 48–59. [Google Scholar] [CrossRef] [PubMed]
- Thu, H.M.; Lowry, K.S.; Myint, T.T.; Shwe, T.N.; Han, A.M.; Khin, K.K.; Thant, K.Z.; Thein, S.; Aaskov, J.G. Myanmar dengue outbreak associated with displacement of serotypes 2, 3, and 4 by dengue 1. Facul. Scien. Tec. 2004, 10, 593–597. [Google Scholar] [CrossRef]
- Sittisombut, N.; Sistayanarain, A.; Cardosa, M.J. Possible occurrence of a genetic bottleneck in dengue serotype 2 viruses between the 1980 and 1987 epidemic seasons in Bangkok Thailand. Am. J. Tropical. 1997, 57, 100–108. [Google Scholar] [CrossRef]
- Halstead, S.B. Dengue virus-mosquito interactions. Annu. Rev. Entomol. 2008, 53, 273–291. [Google Scholar] [CrossRef]
- Morrison, A.C.; Minnick, S.L.; Rocha, C.; Forshey, B.M.; Stoddard, S.T.; Getis, A.; Focks, D.A.; Russell, K.L.; Olson, J.G.; Blair, P.J.; et al. Epidemiology of Dengue Virus in Iquitos, Peru 1999 to 2005: Interepidemic and Epidemic Patterns of Transmission. PLoS Negl. Trop. Dis. 2010, 4, e670. [Google Scholar] [CrossRef]
- Cummings, D.A.T.; Schwartz, I.B.; Billings, L.; Shaw, L.B.; Burke, D.S. Dynamic effects of antibody-dependent enhancement on the fitness of viruses. Proc. Natl. Acad. Sci. USA 2005, 102, 15259–15264. [Google Scholar] [CrossRef]
- Nagao, Y.; Koelle, K. Decreases in dengue transmission may act to increase the incidence of dengue hemorrhagic fever. Proc. Natl. Aca. Sci. USA 2008, 105, 2238–2243. [Google Scholar] [CrossRef]
- Pang, T.; Mak, T.K.; Gubler, D.J. Prevention and control of dengue—the light at the end of the tunnel. Lancet Infect. Dis. 2017, 17, e79–e87. [Google Scholar] [CrossRef]
- Halstead, S.B. Dengvaxia sensitizes seronegatives to vaccine enhanced disease regardless of age. Vaccine 2017, 35, 6355–6358. [Google Scholar] [CrossRef]
- Muller, D.A.; Young, P.R. The flavivirus NS1 protein: Molecular and structural biology, immunology, role in pathogenesis and application as a diagnostic biomarker. Antiviral Res. 2013, 98, 192–208. [Google Scholar] [CrossRef]
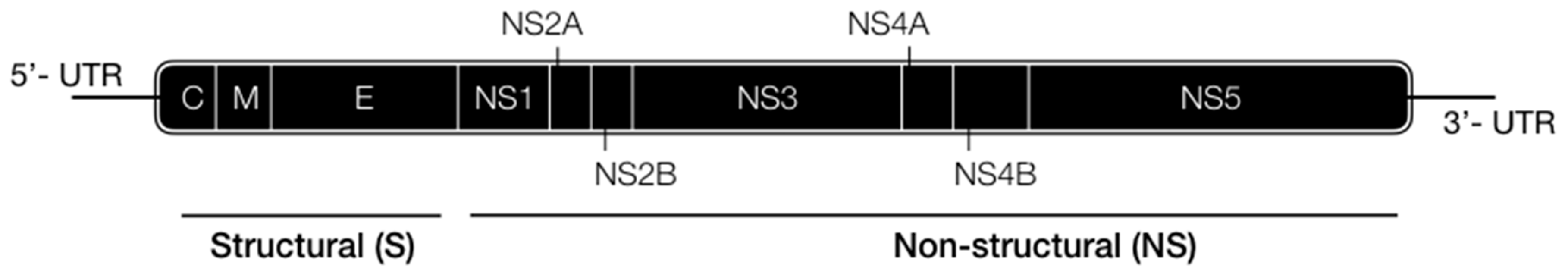
| Region | Function | T-cell Epitope [111] | B-cell Epitope [112] |
|---|---|---|---|
| 5′ UTR [113,114] 5′ untranslated regions | Assists viral RNA replication by direct binding with NS5 and stabilizes long range RNA–RNA interactions | --- | 1° Response: none 2° Response: none |
| C [115] Capsid Protein | Assists nucleic acid rearrangements, crucial for nucleocapsid formation | CD4, CD8 | 1° Response: none 2° Response: + |
| M [116] Membrane | Structural role in forming parts of the prM-E complex | --- | 1° Response: + 2° Response: +++ |
| E Envelope | Key role in interaction of DENV with host cells | CD4 | 1° Response: +++ 2° Response: +++ |
| NS1 [46] Non-structural 1 | Acts on immune evasion by binding into host complement proteins, modulates early events in RNA virus replication by interacting with virus dsRNA and NS4B protein. | --- | 1° Response: +++ 2° Response: +++ |
| NS2A [47] Non-structural 2A | Antagonises the host immune response, component of the viral replication complex and plays a role in virus assembly | CD4 | 1° Response: none 2° Response: + |
| NS2B [117] Non-structural 2B | Regulates viral protease activity (co-factor of the NS3 protease) | CD4 | 1° Response: none 2° Response: + |
| NS3 [118] Non-structural 3 | Acts as protease, RNA helicase and RTPase/NTPase | CD4, CD8 | 1° Response: + 2° Response: ++ |
| NS4A [44] Non-structural 4A | Induces host membrane alterations important for virus replication | CD4, CD8 | 1° Response: none 2° Response: + |
| NS4B [119] Non-structural 4B | Assists viral RNA replication through direct interaction with NS3 | CD4, CD8 | 1° Response: none 2° Response: + |
| NS5 [120,121] Non-structural 5 | Fundamental role in viral genome amplification, counteracts antiviral response through STAT2 degradation | CD4, CD8 | 1° Response: none 2° Response: + |
| 3′ UTR [122] 3′ untranslated regions | Assists long range RNA–RNA interaction between the ends of the viral genome | --- | 1° Response: none 2° Response: none |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Blight, J.; Alves, E.; Reyes-Sandoval, A. Considering Genomic and Immunological Correlates of Protection for a Dengue Intervention. Vaccines 2019, 7, 203. https://doi.org/10.3390/vaccines7040203
Blight J, Alves E, Reyes-Sandoval A. Considering Genomic and Immunological Correlates of Protection for a Dengue Intervention. Vaccines. 2019; 7(4):203. https://doi.org/10.3390/vaccines7040203
Chicago/Turabian StyleBlight, Joshua, Eduardo Alves, and Arturo Reyes-Sandoval. 2019. "Considering Genomic and Immunological Correlates of Protection for a Dengue Intervention" Vaccines 7, no. 4: 203. https://doi.org/10.3390/vaccines7040203
APA StyleBlight, J., Alves, E., & Reyes-Sandoval, A. (2019). Considering Genomic and Immunological Correlates of Protection for a Dengue Intervention. Vaccines, 7(4), 203. https://doi.org/10.3390/vaccines7040203





