1. Introduction
Innate immunity defines the first line of host defence, and interferons (IFNs) play central roles in mediating host resistance against viruses, bacteria, fungi and parasites, by directly and indirectly regulating multiple immune cells [
1]. IFNs are part of the class II cytokine family and are divided into three types (I, II and III) based on their receptor specificity, sequence homology, evolutionary relatedness, functional activity of these cytokines and nature of interferon-stimulated genes (ISGs) induction [
2,
3]. The ISGs display a spectrum of cellular functions and the cumulative actions of these ISGs define the antiviral state of the hosts.
Amongst all IFNs, type I IFNs were first identified, and represent a diverse and potent antiviral class of cytokines. Compared to multiple type I IFNs in human, only two well-characterized type I IFNs (IFN-α and IFN-β) are reported in chicken. We have recently identified a third novel chicken type I IFN (i.e., IFN-κ) that carried antiviral activities both in vitro and in ovo [
4]. While all type I IFNs are structurally diverse, they bind to the same heterodimeric receptor complex composed of one chain of the IFN-α receptor 1 (IFNAR1) and one chain of the IFN-α receptor 2 (IFNAR2). In contrast, IFN-γ (type II IFN) interacts with the IFNγR1 and IFNγR2 receptor complex and play pleiotropic roles in maturation and differentiation processes of several immune cells [
5]. A more recently identified class of IFNs is type III IFNs, or IL-28/29, which are encoded by a single gene of IFN-λ in chicken compared to four genes in humans [
4]. The type III IFN interacts with the heterodimeric receptor complex of IL-28Rα and IL-10Rβ, which are predominantly expressed on the epithelial cells of the host.
Upon binding to their cognate receptors, all types of IFNs initiate a cascade of intracellular events that culminate in the induction of several hundred genes; however, each type of IFN carries a certain level of distinction [
4]. In spite of several common properties of type I and III IFNs, recent discoveries have revealed context-specific functional differences [
6]. It has been shown that type I and III IFNs in mammals initiate the same signalling pathway through phosphorylation of signal transducer and activator of transcription 1 (STAT1) and STAT2 heterodimers possibly by tyrosine kinase 2 (TYK2) and Janus kinase 1 (JAK1). However, type II IFN triggers this pathway via the activation of STAT1 homodimers mediated by JAK1 and JAK2 kinases [
4].
Mammalian type I and type III IFNs signal through the JAK–STAT pathway to activate the heterotrimeric interferon-stimulated gene factor 3 (ISGF3) complex, comprised of phosphorylated STAT1 and STAT2, and interferon regulatory factor 9 (IRF9) [
7,
8]. (IRF9; yet to be identified in chickens.) In contrast, type II IFNs initiate the formation of a STAT1–STAT1 homodimer to assemble gamma-interferon activation factor (GAF), without the need of IRF9. Upon nuclear translocation, ISGF3 and GAF bind to IFN-stimulated response elements (ISREs) [
6] or gamma-IFN activation sequence (GAS) motifs, respectively [
7], leading to the transcriptional activation of hundreds of ISGs, which encode proteins that act via a variety of mechanisms to restrict viral infection [
9,
10,
11].
It has been demonstrated that the JAK–STAT pathway initiates transcriptional regulation of a string of cytokines, chemokine, antimicrobial products and regulators of apoptotic pathways [
2,
7]. In mammals, a range of genome scale transcriptomics screening approaches has been applied to map the diversity, kinetics and expression patterns of these antiviral factors. Based on these data, recent studies have applied large-scale screening platforms to elucidate functional importance of these ISGs regulated by IFN-α, IFN-λ and IFN-γ [
10,
11]. However, a primary focus of these studies was restricted to type I and type II IFNs whereas growing evidences are highlighting the crucial roles of IFN-λ (type III IFN) in establishing an antiviral state. Recently, it has been demonstrated that paracrine singling of IFN-λ is fundamental in early innate immune responses and to inhibit influenza virus spread [
12], which pose enormous zoonotic (e.g., H7N9, H5N1, H9N2 and H5N2) threats to public health and remains the most economically devastating disease in poultry [
13].
Most of our understandings on ISG-mediated antiviral functions in chicken are driven from human and other mammalian studies. Given the fact that chicken differs fundamentally from other mammals in innate immunity, the majority of current concepts are mainly based on assumptions. Some out of many examples include absence of RIG-I, IRF9 and IRF3 homologues, a lack of anti-viral roles of chicken Mx, an absence of many essential molecules in the innate immune signalling pathways, such as IFIT1/2/3, and shorter versions of ZC3HAV1 and many of the DNA sensors [
2,
3]. Intriguingly, despite the absence of these key components, chickens respond to highly pathogenic avian influenza (HPAIVs) and other viruses, and mount potent type I IFN responses [
14,
15,
16], highlighting the presence of yet unknown and evolutionary compensatory mechanisms in chicken.
Given the lack of comprehensive data on the nature of type I, II and type III IFN-mediated antiviral profiling, the uniqueness of the chicken as model system and being an important poultry species, studies have started to catalogue IFN-induced and IFN-regulated genes in chicken [
17,
18]. However, these studies have exclusively identified expression dynamics of ISGs induced by type I IFNs and the nature and regulation of type II- and type III-mediated ISGs are required to be explored in chicken at the genome-wide scale. Additionally, a side-by-side comparison of the type I, II, and III IFN-driven host cell transcriptomes has not been reported so far in chicken. Therefore, in order to lay the foundation for future research on functional annotation of ISGs, we performed intensive gene expression analysis in pan-IFN responsive chicken fibroblasts and provide a comprehensive and comparative dataset on chicken ISGs stimulated with chicken type I, II and III IFNs. This data will help to facilitate underpinning the comparative studies with human ISGs and to stimulate large-scale functional screens as has been performed in mammals [
10].
2. Materials and Methods
2.1. Cells Culture and Media
Immortalized chicken fibroblasts (DF-1) and human embryonic kidney cells 293T (HEK-293T) were maintained in Dulbecco’s modified eagle medium (DMEM) (Gibco, Carlsbad, CA, USA) supplemented with 10% foetal bovine serum (FBS), 1% penicillin and streptomycin (P/S) at 37 °C in a 5% CO2 incubator.
2.2. Production of Interferons
The open reading frame (ORF) sequences for chIFN-α, chIFN-γ and chIFN-λ were synthesized (GeneArt, Fisher Scientific, Loughborough, UK) and cloned into the pcDNA3.1+ vector. The verified pcDNA3.1+ encoding chIFN-α, chIFN-γ and chIFN-λ plasmids were transfected into HEK-293T cells using the Lipofectamine 2000 Transfection Reagent (Invitrogen, Carlsbad, CA, USA). Supernatants were collected at 48 h and 72 h post-transfection, and then were cleared by centrifugation (1500 rpm for 5 min) and pooled and quantified using an IFN-bioassay (see below).
2.3. IFN Bioassay
IFN-induced protection against VSV-GFP was used to identify IFN-producing stable clones (chIL28RA) and to quantify IFN preparations, as described before [
19] and we reported earlier [
3]. Briefly, DF-1 cells were seeded in 96-well plates until they are 90% confluent and treated with serial dilutions of supernatants containing interferons for 24 h. These interferon-stimulated cells were infected with VSV-GFP (MOI of 0.1). After 24 h post-infection, VSV-GFP replication was correlated with the change in GFP fluorescence signal intensities using a Luminometer (Promega, Madison, WI, USA). The percentage antiviral activities of IFNs were determined by comparing the percentage reduction of corrected GFP signal intensity (GFP signal intensity of IFN-treated and virus-infected wells minus the background fluorescence signal intensity of the uninfected control) with the mock-treated and VSV-GFP-infected control wells. One unit (U) of IFN in the tested IFN preparations was defined as the volume containing 50% inhibitory activity against VSV-GFP. A total of 1000 Us of IFNs were used for stimulation of chIL28RA DF-1 cells.
2.4. Library Preparation for mRNA Sequencing
Total RNA was extracted from IFN-primed or mock-treated chIL28RA DF-1 cells in duplicate for RNA-sequencing using the TRIzol reagent (Invitrogen, Carlsbad, CA, USA). Approximately 2.5 μg of RNA per sample was used as input material while the mRNA was enriched by Oligo (dT) beads and then split into short fragments using fragmentation buffer (Thermo) that were then reverse transcribed into cDNA using random primers. Second-strand cDNA was synthesized by DNA polymerase I, RNase H and dNTP. The cDNA fragments were purified using the QiaQuick PCR Extraction Kit, poly(A) tails were added and ends were repaired and ligated to Illumina sequencing adapters. The ligation products were size selected by agarose gel electrophoresis, PCR amplified and were then sequenced using the Illumina HiSeq in 2 × 150 bp configuration by Genewiz Co. (New York, NY, USA).
2.5. RNA-Seq Data Analysis
Sequence reads were trimmed to remove possible adapter sequences and nucleotides with poor quality using Trimmomatic v.0.36 [
20]. The reads were then mapped to the chicken reference genome available on ENSEMBL using the STAR aligner [
21]. The RNA-seq aligner was executed using a splice aligner, which detects splice junctions and incorporating them to help align the entire read sequences, which generate BAM files. Unique exon hit counts were calculated using feature counts from the Subread package. After mapping and total gene hit counts calculation, the total gene hit counts table was used for downstream differential expression analysis using EdgeR [
22]. Expression values from each sample were carried out to estimate the sample distances. The shorter the distance, the more closely related the samples are, which were used to identify if the two groups are closely related or not. Volcano plot analysis used to show the global transcriptional change across the groups [
23].
Gene ontology analysis was conducted using GeneSCF [
24]. Significant genes (absolute Log2 Fold Change, FC, >2) were first annotated with the Gene Ontology Biological Process information from the “goa_chicken” database. Then a Fisher’s exact test was conducted on each ontology. A list containing statistics of over- or under-representation of GO Biological Processes relative to the unfiltered gene list was generated for each comparison. After mapping and total transcript hit counts calculation, the total transcript hit counts table was used for downstream differential expression analysis using DEXSeq [
25]. To estimate the expression levels of alternatively spliced transcripts, the splice variant hit counts were extracted from the RNA-seq reads mapped to the genome. Differentially spliced genes were identified by testing for significant differences in read counts on exons (and junctions) of the genes using DEXSeq with an adjusted
p value of 0.05. Raw data is publically available (
Appendix A).
2.6. Quantitative Reverse Transcription-PCR and Cytokine Expression
Total RNA was extracted from chIFN-α, chIFN-β and chIFN-λ (1000 U) stimulated chIL28RA DF-1 cells using TRIzol reagents (Invitrogen, Carlsbad, CA, USA). The quantity and purity of extracted RNA was determined with a NanoDrop 1000 (PEQLAB Biotechnologie GMBH, Erlangen, Germany), while the RNA quality was analysed using a 2100 Bioanalyzer® (Agilent Technologies, Böblingen, Germany). A total of 200 ng of RNA was used in PCR reactions using SuperScript® III Platinum® SYBR® Green One-Step qRT-PCR Kit (Invitrogen, Carlsbad, CA, USA). The abundance of specific ISG mRNA was compared to the 28S rRNA in the Applied Biosystems Prism 7500 system. The reaction was carried out in an ABI 7500 light cycler using the following thermo profile: 50 °C held for 5 min and 95 °C held for 2 min, followed by 40 cycles of 95 °C for 3 s and 60 °C for 30 s. The melting curve was determined at 95 °C for 15 s, 60 °C for 1 min, 95 °C for 15 s and 60 °C for 15 s.
2.7. Prediction of ISREs and GAS Elements and Promotor Analysis
In order to investigate if the significantly induced ISGs by the chicken type I, II, and III IFNs carry ISRE and GAS elements in their promoter regions, we scanned a total of four kilo bases sequence upstream to the promoter regions according to the approach described before [
18,
26,
27]. Screening was performed based on the Ensembl ID of the most up-regulated genes for GAS and ISRE using a Shell Script (kindly provided by Thomas Lütteke, Germany) [
26].
2.8. Luciferase Reporter Assays
To determine responsiveness of chicken ISRE and GAS to chicken chIFN-α, chIFN-γ or chIFN-λ, chIL28RA DF-1 cells were grown in 96-well plate format at 2 × 104 to 4 × 104 cells/well and were co-transfected with 10 ng/well of a plasmid constitutively expressing Renilla luciferase (phRL-SV40; Promega, Madison, WI, USA) and 150 ng/well of full length pchMx or pchIFIT5 constructs. The pGL3.1 basic vector was used as a negative control. All transfections were performed using Lipofectamine 2000 and luciferase reporter assays were performed on the monolayers of chIL28RA DF-1 cells according to the manufacturer’s protocol. At 24 h post-transfection, the transfected DF1 cells were stimulated with chIFN-α, chIFN-γ, and chIFN-λ then lysed using 20 μL 1× passive lysis buffer (Promega, Madison, WI, USA) at 24 h post-IFN treatment, and samples were assayed for Firefly and Renilla luciferase activities using the Dual-Luciferase Reporter Assay System by following the supplier’s instructions.
2.9. Statistical Analyses
Pairwise comparisons of treated and control groups were performed using Student’s t-test. All statistical analysis and the majority of figures were performed in GraphPad Prism (GraphPad Software, La Jolla, CA, USA).
4. Discussion
Interferons were first described in “chicken” when Isaacs and Lindenmann noticed that supernatant from virus-infected chicken cells can “interfere” with the replication of influenza viruses [
29]. However, the vast majority of information on the nature of IFN-regulated genes and IFN pathways in chicken is derived from studies conducted in humans and mice. Moreover, substantial differences in several innate immune checkpoints have been observed in chickens, which highlight fundamental and evolutionary distinct mechanisms in birds, especially in chickens [
3]. Thus, understanding mechanisms and gaining insights into additional compensatory mechanisms in the IFN-mediated establishment of an antiviral state in chickens would provide foundations to underpin diversity and dynamics of innate immunity in animals.
Since ISGs play the most dynamic and versatile roles in inhibiting viruses at multiple scales, genetic annotation of these genes is essential to perform large-scale functional studies and to reveal functional plasticity of ISGs. While information in mammals, especially in human and mice, has started to expand in unravelling the roles of ISGs, studies have proposed the expression of ISGs both in vitro [
30] and in vivo [
18] in chicken; however, this information is only limited to type I IFN or to a limited set of type III IFN genes [
31]. At system and cellular levels, IFNs and their regulated genes function in a network and thus necessitate the exploration of these components at the genome-wide scale. Moreover, mapping and cataloguing genes that are specific to type I, II or III IFNs or identifying genes that span the responses against pan-IFN levels would provide foundation to functional investigations against viruses of birds.
Building upon existing information on chicken ISGs [
18,
30,
32] and our recent studies [
3,
33], we compared the expression of the genes that were induced by either chIFN-α, chIFN-λ or chIFN-γ, and catalogue the genes which are IFN-specific or genes that are shared between different types of IFNs in chicken. Since IFNs show cell and tissue-specific actions [
8], which are mainly attributed to their cognate expression patterns [
28], we attempted to establish a model system that could reliably respond to all type of IFNs. The chicken fibroblast (i.e., DF-1) is one of the most characterized cell lines in avian research and majority of virus-induced transcriptomics studies have been conducted on these cells. In order to stimulate future comparative studies and functional annotation, chicken fibroblasts were chosen. However, due to repressive expression of chIFN-λ receptors [
28], the treatment of DF1 with type III IFN showed defective genes inductions, IFN-promoter activation and cumulatively established an antiviral state against IFN-sensitive virus (e.g., VSV). We thus exploited this permanent DF1 cell line that exogenously expresses chIFN-λ receptors and thus revised the full responsiveness to type III as well as other IFNs. We propose this as an improved tool to study genome-scale ISGs induced by multiple stimuli and regulated by a complex cellular network.
Using genome-scale transcriptomes in chicken fibroblasts, we enlisted genes that were IFN-specific or were generic (induced by all IFNs). The overall numbers of genes that were either up- or down-regulated were lower compared to previous studies, which were performed on primary [
17] or established cell lines or in vivo [
18]. We reasoned that since we applied stringent-selection and fold-induction criteria in a cell line that could induce simultaneous responses, the overall gene transcription was weaker. Moreover, the in vitro experimental setting would differ from a naturally occurring infection where more potent local type I IFNs are activated soon after viral infections (or other stimuli) and which then expand to the system with the release of several additional cytokines and other affecters simultaneously [
34]. However, significantly up- or down-regulated genes in our study were known IFN-regulated genes (e.g., ISGs). Moreover, chIFIT5, which we have previously identified and characterized [
3], showed higher expression in type I and type III-treated chicken fibroblasts. Additionally, owing to the fact that different cells are known to respond IFNs differently [
4] and that ISGs express temporally depending on the nature of stimuli [
18], gauging the expression dynamics at different time points with both viral and IFNs stimulated cell can provide complete picture of cellular events. Based on our experiences (unpublished data), expression of receptors on different cell types is the primary factor to IFN responsiveness. Whether the nature of the ISGs is induced by a specific IFN in a particular cell population, vitalized by the paracrine establishment of antiviral responses, or selective induction of genes suffice the local antiviral states, warrant future investigations.
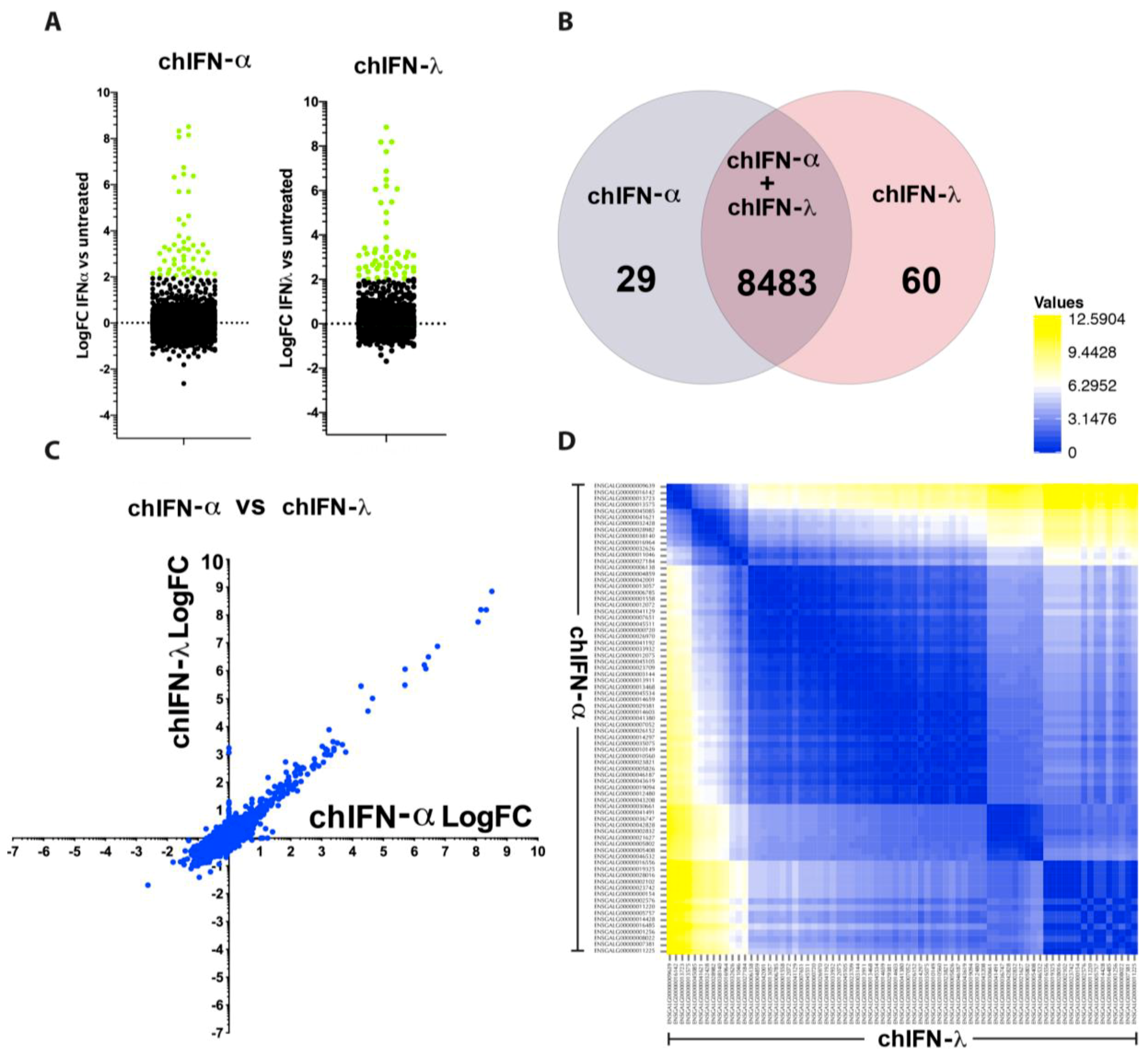
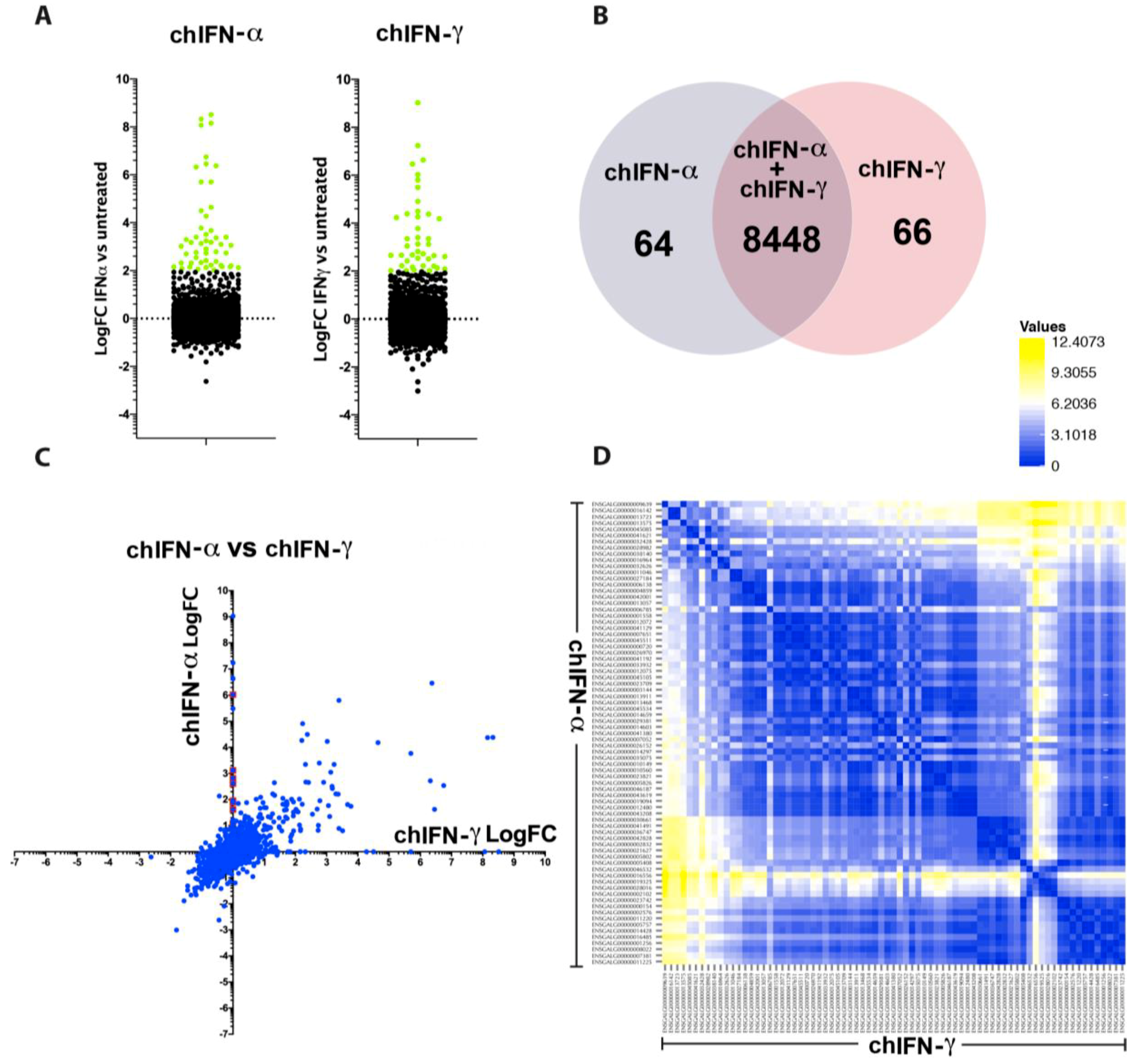
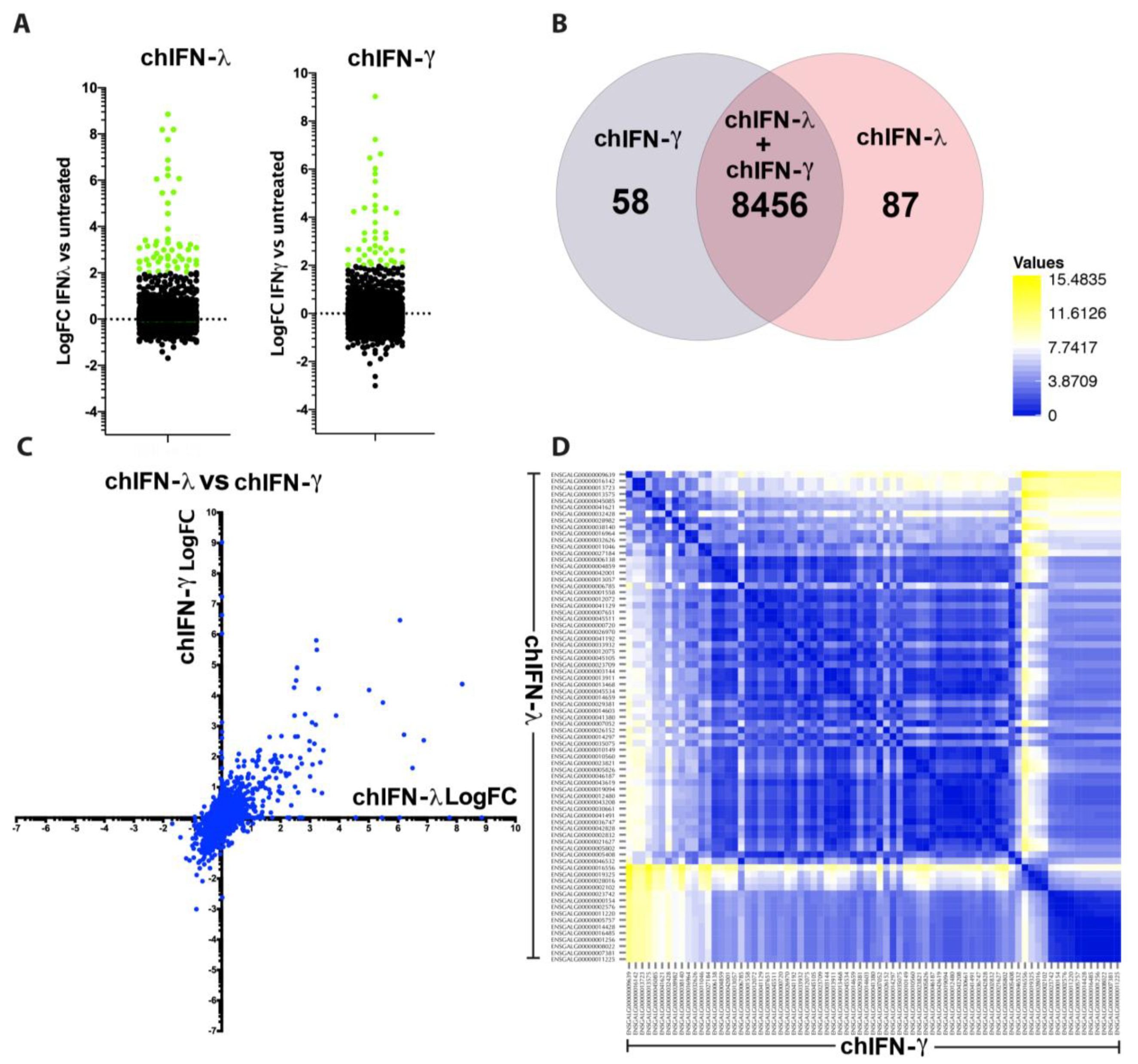
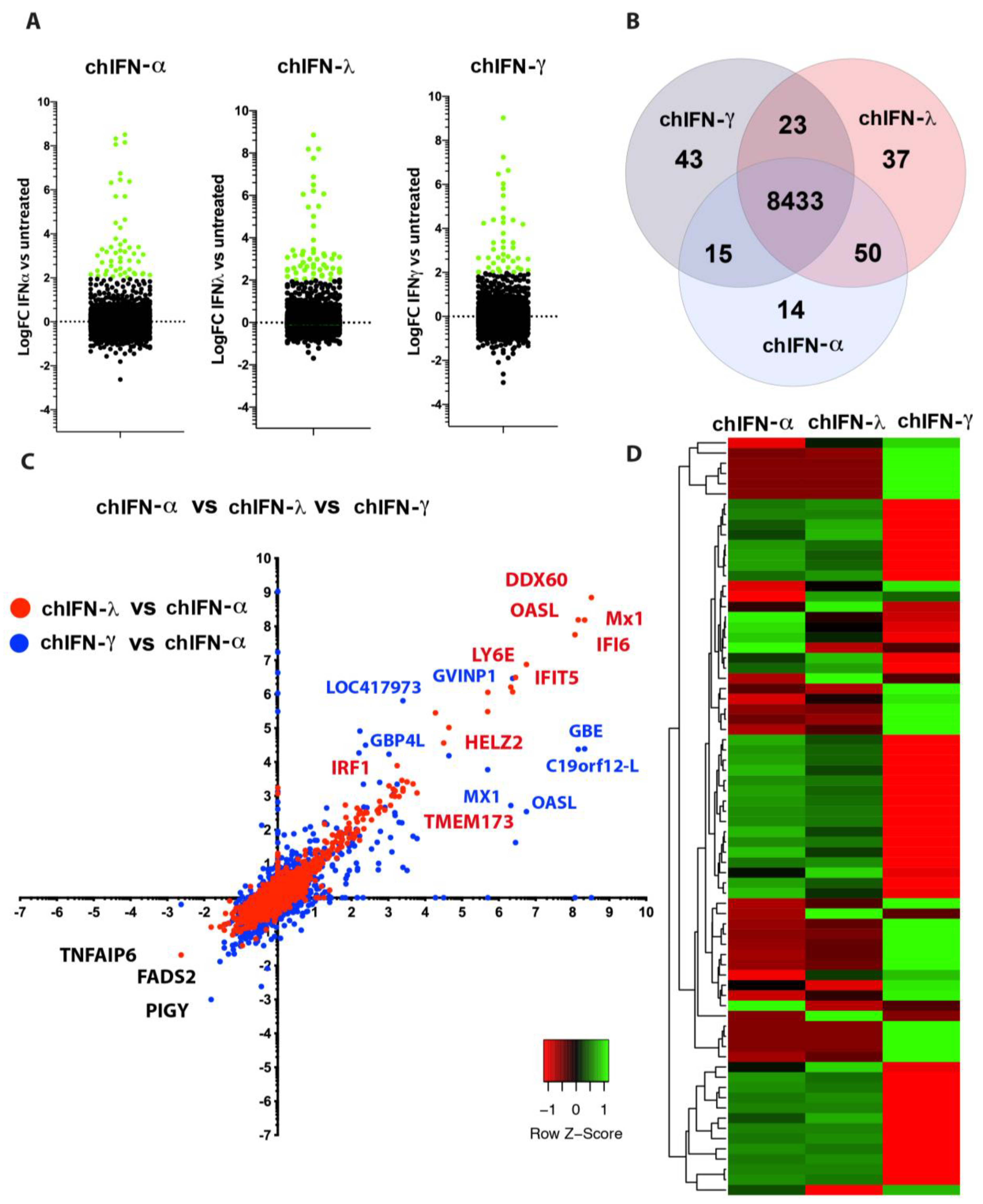
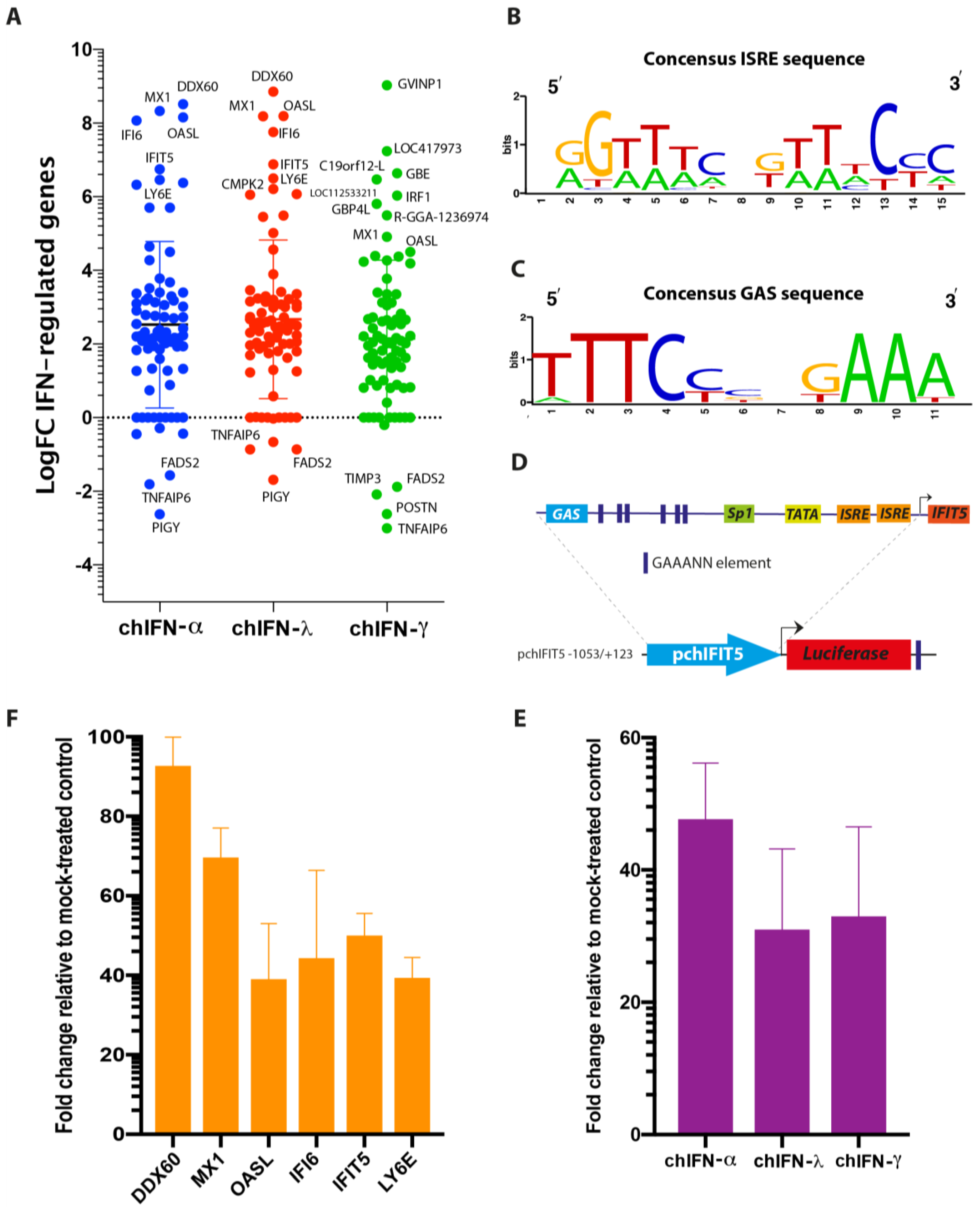
The direct comparison of all ISGs regulated by type I, II and III indicated that the effects of chIFN-α and chIFN-λ on the expression of ISGs were distinctive from the genes expressed in chIFN-γ-stimulated cells. As anticipated, a higher resolution analysis indicated that the differentially expressed genes included a list of known genes, such as
IFIT5,
Mx1 and
OASL, further supporting the reliability of the experimental approach and the suitability of the model. However, as Roll et al. [
18] have highlighted, poor annotation of the chicken genome may lead to either false gene annotation or a lack of gene assembly results in missed genes in the list of otherwise IFN-stimulated genes.
During the analysis of IFN-mediated gene expression, it was revealed that up-regulated genes were predominantly presented over the down-regulated genes in all tested type I, II and III IFNs. It was only PIGY (phosphatidylinositol glycan anchor biosynthesis) that was significantly down-regulated by the type I IFN and three genes named
TNFAIP6 (tumour necrosis factor-inducible gene 6 protein),
POSTN (periostin) and
TIMP3 (TIMP metallopeptidase inhibitor 3) were down-regulated by the type II IFNs. One plausible rationale for higher number of upregulated genes is the trait of all IFNs and is supported by the presence of ISRE and GAS elements in majority of analysed ISGs. Analysis of highly expressed ISGs in different IFN-treated lists revealed presence of a number of ISRE and GAS motifs. Intriguingly and in contrast to human promoter sequences [
27], a large number of GAS sequences were identified compared to ISRE motifs in chicken ISGs. Comparing these motifs with the nature of treated IFNs, no correlation was observed; all types of IFN-regulated genes carried both ISRE and GAS elements. While we demonstrated that chicken IFNs transcriptionally induced a reporter gene through GAS and ISRE in an IFN-independent manner, it requires future experimental studies to identify the biological relevance of in silico predicted motifs with functional relevance. The transcriptomes and individual reports on ISGs could not clearly distinguish between primary (direct IFN-dependents) and secondary (positive loop feedback of IFNs) responses, thus impeding a clear differentiation of genes expressions that are specific to each of these stimuli. Moreover, the majority of functional studies have focused on the up-regulated genes, the potential anti- or pro-viral roles of under-expressed genes would shed lights on the dynamics and plasticity of IFN regulated genes.
Owing to the fact that ISGs play fundamental roles in a wide range of cellular activities, including transcriptional and translational regulation of immune responses [
35,
36] and establishing a host antiviral state against viruses, the provided data would lay foundations to investigate functions of these important and complex domains of host–pathogen interactions using large-scale screening platforms. This comprehensive information is first of its kind to compare genes that are differentially regulated by type I, II and type III IFNs in chicken.