The Potential Role of Fc-Receptor Functions in the Development of a Universal Influenza Vaccine
Abstract
1. The Problem—The Elusive Influenza Virus
2. Antibodies to Influenza Virus—A Potential Solution
2.1. Antibody-Dependent Complement Mediated-Lysis (ADCL)
2.2. Antibody-Dependent Phagocytosis (ADP)
2.3. Antibody-Dependent Cellular Cytotoxicity (ADCC)
3. Antibody Structure and the Influence of Glycosylation on Fc-Receptor Function
4. The Rationale for Fc-Receptor Function as an Important Correlate of Antibody-Mediate Protection
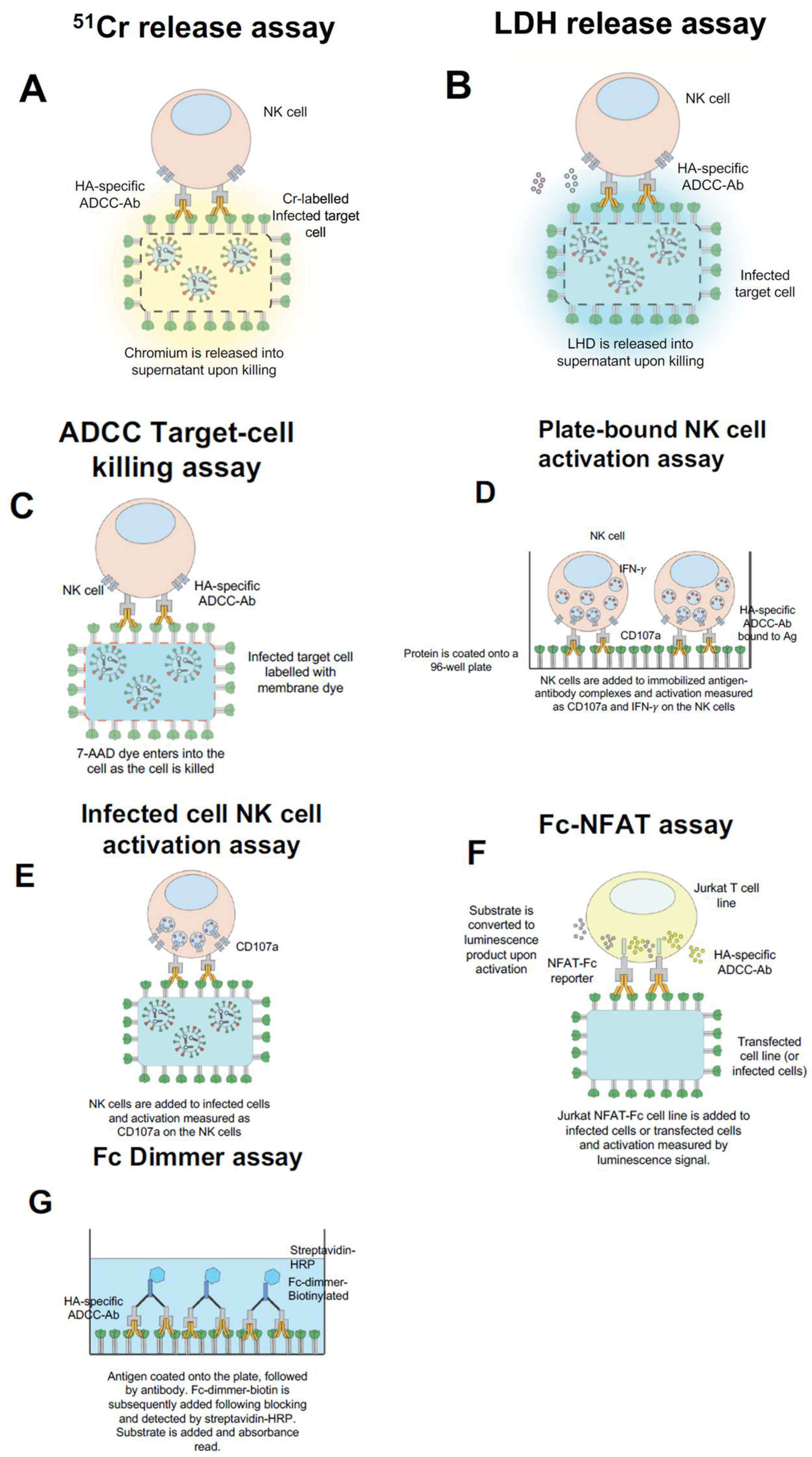
5. In Vitro Assays to Measure ADCC Function
6. Measuring ADCC Following Vaccination and Infection
7. Conclusions and Perspectives for the Future
Conflicts of Interest
References
- Organisation, W.H. Influenza (Seasonal) Fact Sheet. Available online: http://www.who.int/mediacentre/factsheets/fs211/en/ (accessed on 2 January 2018).
- Palache, A.; Oriol-Mathieu, V.; Fino, M.; Xydia-Charmanta, M. Seasonal influenza vaccine dose distribution in 195 countries (2004–2013): Little progress in estimated global vaccination coverage. Vaccine 2015, 33, 5598–5605. [Google Scholar] [CrossRef] [PubMed]
- Palache, A.; Abelin, A.; Hollingsworth, R.; Cracknell, W.; Jacobs, C.; Tsai, T.; Barbosa, P. Survey of distribution of seasonal influenza vaccine doses in 201 countries (2004–2015): The 2003 World Health Assembly resolution on seasonal influenza vaccination coverage and the 2009 influenza pandemic have had very little impact on improving influenza control and pandemic preparedness. Vaccine 2017, 35, 4681–4686. [Google Scholar] [PubMed]
- Huang, K.Y.; Rijal, P.; Schimanski, L.; Powell, T.J.; Lin, T.Y.; McCauley, J.W.; Daniels, R.S.; Townsend, A.R. Focused antibody response to influenza linked to antigenic drift. J. Clin. Investig. 2015, 125, 2631–2645. [Google Scholar] [CrossRef] [PubMed]
- Dinis, J.M.; Florek, N.W.; Fatola, O.O.; Moncla, L.H.; Mutschler, J.P.; Charlier, O.K.; Meece, J.K.; Belongia, E.A.; Friedrich, T.C. Deep Sequencing Reveals Potential Antigenic Variants at Low Frequencies in Influenza A Virus-Infected Humans. J. Virol. 2016, 90, 3355–3365. [Google Scholar] [CrossRef] [PubMed]
- DeDiego, M.L.; Anderson, C.S.; Yang, H.; Holden-Wiltse, J.; Fitzgerald, T.; Treanor, J.J.; Topham, D.J. Directed selection of influenza virus produces antigenic variants that match circulating human virus isolates and escape from vaccine-mediated immune protection. Immunology 2016, 148, 160–173. [Google Scholar] [CrossRef] [PubMed]
- Anderson, C.S.; Ortega, S.; Chaves, F.A.; Clark, A.M.; Yang, H.; Topham, D.J.; DeDiego, M.L. Natural and directed antigenic drift of the H1 influenza virus hemagglutinin stalk domain. Sci. Rep. 2017, 7, 14614. [Google Scholar] [CrossRef] [PubMed]
- Carrat, F.; Flahault, A. Influenza vaccine: The challenge of antigenic drift. Vaccine 2007, 25, 6852–6862. [Google Scholar] [CrossRef] [PubMed]
- Both, G.W.; Sleigh, M.J.; Cox, N.J.; Kendal, A.P. Antigenic drift in influenza virus H3 hemagglutinin from 1968 to 1980: Multiple evolutionary pathways and sequential amino acid changes at key antigenic sites. J. Virol. 1983, 48, 52–60. [Google Scholar] [PubMed]
- Gerhard, W.; Webster, R.G. Antigenic drift in influenza A viruses. I. Selection and characterization of antigenic variants of A/PR/8/34 (HON1) influenza virus with monoclonal antibodies. J. Exp. Med. 1978, 148, 383–392. [Google Scholar] [CrossRef] [PubMed]
- Webster, R.G.; Laver, W.G.; Air, G.M.; Ward, C.; Gerhard, W.; van Wyke, K.L. The mechanism of antigenic drift in influenza viruses: Analysis of Hong Kong (H3N2) variants with monoclonal antibodies to the hemagglutinin molecule. Ann. N. Y. Acad. Sci. 1980, 354, 142–161. [Google Scholar] [CrossRef] [PubMed]
- Goodwin, K.; Viboud, C.; Simonsen, L. Antibody response to influenza vaccination in the elderly: A quantitative review. Vaccine 2006, 24, 1159–1169. [Google Scholar] [CrossRef] [PubMed]
- Nichol, K.L. The efficacy, effectiveness and cost-effectiveness of inactivated influenza virus vaccines. Vaccine 2003, 21, 1769–1775. [Google Scholar] [CrossRef]
- Young, B.; Zhao, X.; Cook, A.R.; Parry, C.M.; Wilder-Smith, A.; I-Cheng, M.C. Do antibody responses to the influenza vaccine persist year-round in the elderly? A systematic review and meta-analysis. Vaccine 2017, 35, 212–221. [Google Scholar] [CrossRef] [PubMed]
- Petrie, J.G.; Ohmit, S.E.; Johnson, E.; Truscon, R.; Monto, A.S. Persistence of Antibodies to Influenza Hemagglutinin and Neuraminidase Following One or Two Years of Influenza Vaccination. J. Infect. Dis. 2015, 212, 1914–1922. [Google Scholar] [CrossRef] [PubMed]
- Gerhard, W.; Yewdell, J.; Frankel, M.E.; Webster, R. Antigenic structure of influenza virus haemagglutinin defined by hybridoma antibodies. Nature 1981, 290, 713–717. [Google Scholar] [CrossRef] [PubMed]
- Caton, A.J.; Brownlee, G.G.; Yewdell, J.W.; Gerhard, W. The antigenic structure of the influenza virus A/PR/8/34 hemagglutinin (H1 subtype). Cell 1982, 31, 417–427. [Google Scholar] [CrossRef]
- Wilson, I.A.; Skehel, J.J.; Wiley, D.C. Structure of the haemagglutinin membrane glycoprotein of influenza virus at 3 A resolution. Nature 1981, 289, 366–373. [Google Scholar] [CrossRef] [PubMed]
- Hobson, D.; Curry, R.L.; Beare, A.S.; Ward-Gardner, A. The role of serum haemagglutination-inhibiting antibody in protection against challenge infection with influenza A2 and B viruses. Epidemiol. Infect. 1972, 70, 767–777. [Google Scholar] [CrossRef]
- Ohmit, S.E.; Petrie, J.G.; Cross, R.T.; Johnson, E.; Monto, A.S. Influenza hemagglutination-inhibition antibody titer as a correlate of vaccine-induced protection. J. Infect. Dis. 2011, 204, 1879–1885. [Google Scholar] [CrossRef] [PubMed]
- Potter, C.W.; Oxford, J.S. Determinants of immunity to influenza infection in man. Br. Med. Bull. 1979, 35, 69–75. [Google Scholar] [CrossRef] [PubMed]
- Benoit, A.; Beran, J.; Devaster, J.M.; Esen, M.; Launay, O.; Leroux-Roels, G.; McElhaney, J.E.; Oostvogels, L.; van Essen, G.A.; Gaglani, M.; et al. Hemagglutination Inhibition Antibody Titers as a Correlate of Protection Against Seasonal A/H3N2 Influenza Disease. In Open Forum Infect Diseases; Oxford University Press: Oxford, UK, 2015. [Google Scholar]
- Fox, J.P.; Cooney, M.K.; Hall, C.E.; Foy, H.M. Influenzavirus infections in Seattle families, 1975–1979. II. Pattern of infection in invaded households and relation of age and prior antibody to occurrence of infection and related illness. Am. J. Epidemiol. 1982, 116, 228–242. [Google Scholar] [CrossRef] [PubMed]
- Coudeville, L.; Bailleux, F.; Riche, B.; Megas, F.; Andre, P.; Ecochard, R. Relationship between haemagglutination-inhibiting antibody titres and clinical protection against influenza: Development and application of a bayesian random-effects model. BMC Med. Res. Methodol. 2010, 10, 18. [Google Scholar] [CrossRef] [PubMed]
- Smith, D.J.; Lapedes, A.S.; de Jong, J.C.; Bestebroer, T.M.; Rimmelzwaan, G.F.; Osterhaus, A.D.; Fouchier, R.A. Mapping the antigenic and genetic evolution of influenza virus. Science 2004, 305, 371–376. [Google Scholar] [CrossRef] [PubMed]
- Anderson, C.S.; McCall, P.R.; Stern, H.A.; Yang, H.; Topham, D.J. Antigenic cartography of H1N1 influenza viruses using sequence-based antigenic distance calculation. BMC Bioinform. 2018, 19, 51. [Google Scholar] [CrossRef] [PubMed]
- Cobey, S.; Hensley, S.E. Immune history and influenza virus susceptibility. Curr. Opin. Virol. 2017, 22, 105–111. [Google Scholar] [CrossRef] [PubMed]
- Rattan, A.; Pawar, S.D.; Nawadkar, R.; Kulkarni, N.; Lal, G.; Mullick, J.; Sahu, A. Synergy between the classical and alternative pathways of complement is essential for conferring effective protection against the pandemic influenza A(H1N1) 2009 virus infection. PLoS Pathog. 2017, 13, e1006248. [Google Scholar] [CrossRef] [PubMed]
- O’Brien, K.B.; Morrison, T.E.; Dundore, D.Y.; Heise, M.T.; Schultz-Cherry, S. A protective role for complement C3 protein during pandemic 2009 H1N1 and H5N1 influenza A virus infection. PLoS ONE 2011, 6, e17377. [Google Scholar] [CrossRef] [PubMed]
- Kopf, M.; Abel, B.; Gallimore, A.; Carroll, M.; Bachmann, M.F. Complement component C3 promotes T-cell priming and lung migration to control acute influenza virus infection. Nat. Med. 2002, 8, 373–378. [Google Scholar] [CrossRef] [PubMed]
- Quinnan, G.V.; Ennis, F.A.; Tuazon, C.U.; Wells, M.A.; Butchko, G.M.; Armstrong, R.; McLaren, C.; Manischewitz, J.F.; Kiley, S. Cytotoxic lymphocytes and antibody-dependent complement-mediated cytotoxicity induced by administration of influenza vaccine. Infect. Immun. 1980, 30, 362–369. [Google Scholar] [PubMed]
- Co, M.D.; Cruz, J.; Takeda, A.; Ennis, F.A.; Terajima, M. Comparison of complement dependent lytic, hemagglutination inhibition and microneutralization antibody responses in influenza vaccinated individuals. Hum. Vaccine Immunother. 2012, 8, 1218–1222. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Verbonitz, M.W.; Ennis, F.A.; Hicks, J.T.; Albrecht, P. Hemagglutinin-specific complement-dependent cytolytic antibody response to influenza infection. J. Exp. Med. 1978, 147, 265–270. [Google Scholar] [CrossRef] [PubMed]
- Co, M.D.; Terajima, M.; Thomas, S.J.; Jarman, R.G.; Rungrojcharoenkit, K.; Fernandez, S.; Yoon, I.K.; Buddhari, D.; Cruz, J.; Ennis, F.A. Relationship of preexisting influenza hemagglutination inhibition, complement-dependent lytic, and antibody-dependent cellular cytotoxicity antibodies to the development of clinical illness in a prospective study of A(H1N1)pdm09 Influenza in children. Viral. Immunol. 2014, 27, 375–382. [Google Scholar] [CrossRef] [PubMed]
- Terajima, M.; Co, M.D.; Cruz, J.; Ennis, F.A. High Antibody-Dependent Cellular Cytotoxicity Antibody Titers to H5N1 and H7N9 Avian Influenza A Viruses in Healthy US Adults and Older Children. J. Infect. Dis. 2015, 212, 1052–1060. [Google Scholar] [CrossRef] [PubMed]
- Terajima, M.; Cruz, J.; Co, M.D.; Lee, J.H.; Kaur, K.; Wrammert, J.; Wilson, P.C.; Ennis, F.A. Complement-dependent lysis of influenza a virus-infected cells by broadly cross-reactive human monoclonal antibodies. J. Virol. 2011, 85, 13463–13467. [Google Scholar] [CrossRef] [PubMed]
- Ana-Sosa-Batiz, F.; Johnston, A.P.R.; Hogarth, P.M.; Wines, B.D.; Barr, I.; Wheatley, A.K.; Kent, S.J. Antibody-dependent phagocytosis (ADP) responses following trivalent inactivated influenza vaccination of younger and older adults. Vaccine 2017, 35, 6451–6458. [Google Scholar] [CrossRef] [PubMed]
- Vanderven, H.A.; Liu, L.; Ana-Sosa-Batiz, F.; Nguyen, T.H.; Wan, Y.; Wines, B.; Hogarth, P.M.; Tilmanis, D.; Reynaldi, A.; Parsons, M.S.; et al. Fc functional antibodies in humans with severe H7N9 and seasonal influenza. JCI Insight 2017, 6. [Google Scholar] [CrossRef] [PubMed]
- Ana-Sosa-Batiz, F.; Vanderven, H.; Jegaskanda, S.; Johnston, A.; Rockman, S.; Laurie, K.; Barr, I.; Reading, P.; Lichtfuss, M.; Kent, S.J. Influenza-Specific Antibody-Dependent Phagocytosis. PLoS ONE 2016, 11, e0154461. [Google Scholar] [CrossRef] [PubMed]
- Huber, V.C.; Lynch, J.M.; Bucher, D.J.; Le, J.; Metzger, D.W. Fc receptor-mediated phagocytosis makes a significant contribution to clearance of influenza virus infections. J. Immunol. 2001, 166, 7381–7388. [Google Scholar] [CrossRef] [PubMed]
- Quast, I.; Peschke, B.; Lunemann, J.D. Regulation of antibody effector functions through IgG Fc N-glycosylation. Cell. Mol. Life Sci. 2017, 74, 837–847. [Google Scholar] [CrossRef] [PubMed]
- Anthony, R.M.; Ravetch, J.V. A novel role for the IgG Fc glycan: The anti-inflammatory activity of sialylated IgG Fcs. J. Clin. Immunol. 2010, 30 (Suppl. 1), S9–S14. [Google Scholar] [CrossRef] [PubMed]
- Duncan, A.R.; Woof, J.M.; Partridge, L.J.; Burton, D.R.; Winter, G. Localization of the binding site for the human high-affinity Fc receptor on IgG. Nature 1988, 332, 563–564. [Google Scholar] [CrossRef] [PubMed]
- Nose, M.; Wigzell, H. Biological significance of carbohydrate chains on monoclonal antibodies. Proc. Natl. Acad. Sci. USA 1983, 80, 6632–6636. [Google Scholar] [CrossRef] [PubMed]
- Umana, P.; Jean-Mairet, J.; Moudry, R.; Amstutz, H.; Bailey, J.E. Engineered glycoforms of an antineuroblastoma IgG1 with optimized antibody-dependent cellular cytotoxic activity. Nat. Biotechnol. 1999, 17, 176–180. [Google Scholar] [CrossRef] [PubMed]
- Ferrara, C.; Grau, S.; Jager, C.; Sondermann, P.; Brunker, P.; Waldhauer, I.; Hennig, M.; Ruf, A.; Rufer, A.C.; Stihle, M.; et al. Unique carbohydrate-carbohydrate interactions are required for high affinity binding between FcgammaRIII and antibodies lacking core fucose. Proc. Natl. Acad. Sci. USA 2011, 108, 12669–12674. [Google Scholar] [CrossRef] [PubMed]
- Kaneko, Y.; Nimmerjahn, F.; Ravetch, J.V. Anti-inflammatory activity of immunoglobulin G resulting from Fc sialylation. Science 2006, 313, 670–673. [Google Scholar] [CrossRef] [PubMed]
- Anthony, R.M.; Nimmerjahn, F.; Ashline, D.J.; Reinhold, V.N.; Paulson, J.C.; Ravetch, J.V. Recapitulation of IVIG anti-inflammatory activity with a recombinant IgG Fc. Science 2008, 320, 373–376. [Google Scholar] [CrossRef] [PubMed]
- Kaveri, S.V.; Lacroix-Desmazes, S.; Bayry, J. The antiinflammatory IgG. N. Engl. J. Med. 2008, 359, 307–309. [Google Scholar] [CrossRef] [PubMed]
- Li, T.; DiLillo, D.J.; Bournazos, S.; Giddens, J.P.; Ravetch, J.V.; Wang, L.X. Modulating IgG effector function by Fc glycan engineering. Proc. Natl. Acad. Sci. USA 2017, 114, 3485–3490. [Google Scholar] [CrossRef] [PubMed]
- Corti, D.; Voss, J.; Gamblin, S.J.; Codoni, G.; Macagno, A.; Jarrossay, D.; Vachieri, S.G.; Pinna, D.; Minola, A.; Vanzetta, F.; et al. A neutralizing antibody selected from plasma cells that binds to group 1 and group 2 influenza A hemagglutinins. Science 2011, 333, 850–856. [Google Scholar] [CrossRef] [PubMed]
- He, W.; Chen, C.J.; Mullarkey, C.E.; Hamilton, J.R.; Wong, C.K.; Leon, P.E.; Uccellini, M.B.; Chromikova, V.; Henry, C.; Hoffman, K.W.; et al. Alveolar macrophages are critical for broadly-reactive antibody-mediated protection against influenza A virus in mice. Nat. Commun. 2017, 8, 846. [Google Scholar] [CrossRef] [PubMed]
- Mullarkey, C.E.; Bailey, M.J.; Golubeva, D.A.; Tan, G.S.; Nachbagauer, R.; He, W.; Novakowski, K.E.; Bowdish, D.M.; Miller, M.S.; Palese, P. Broadly Neutralizing Hemagglutinin Stalk-Specific Antibodies Induce Potent Phagocytosis of Immune Complexes by Neutrophils in an Fc-Dependent Manner. MBio 2016, 7, e01624-16. [Google Scholar] [CrossRef] [PubMed]
- DiLillo, D.J.; Tan, G.S.; Palese, P.; Ravetch, J.V. Broadly neutralizing hemagglutinin stalk-specific antibodies require FcgammaR interactions for protection against influenza virus in vivo. Nat. Med. 2014, 20, 143–151. [Google Scholar] [CrossRef] [PubMed]
- DiLillo, D.J.; Palese, P.; Wilson, P.C.; Ravetch, J.V. Broadly neutralizing anti-influenza antibodies require fc receptor engagement for in vivo protection. J. Clin. Investig. 2016, 126, 605–610. [Google Scholar] [CrossRef] [PubMed]
- He, W.; Tan, G.S.; Mullarkey, C.E.; Lee, A.J.; Lam, M.M.; Krammer, F.; Henry, C.; Wilson, P.C.; Ashkar, A.A.; Palese, P.; et al. Epitope specificity plays a critical role in regulating antibody-dependent cell-mediated cytotoxicity against influenza A virus. Proc. Natl. Acad. Sci. USA 2016, 113, 11931–11936. [Google Scholar] [CrossRef] [PubMed]
- Simhadri, V.R.; Dimitrova, M.; Mariano, J.L.; Zenarruzabeitia, O.; Zhong, W.; Ozawa, T.; Muraguchi, A.; Kishi, H.; Eichelberger, M.C.; Borrego, F. A Human Anti-M2 Antibody Mediates Antibody-Dependent Cell-Mediated Cytotoxicity (ADCC) and Cytokine Secretion by Resting and Cytokine-Preactivated Natural Killer (NK) Cells. PLoS ONE 2015, 10, e0124677. [Google Scholar] [CrossRef] [PubMed]
- Wang, R.; Song, A.; Levin, J.; Dennis, D.; Zhang, N.J.; Yoshida, H.; Koriazova, L.; Madura, L.; Shapiro, L.; Matsumoto, A.; et al. Therapeutic potential of a fully human monoclonal antibody against influenza A virus M2 protein. Antivir. Res. 2008, 80, 168–177. [Google Scholar] [CrossRef] [PubMed]
- Van den Hoecke, S.; Ehrhardt, K.; Kolpe, A.; El Bakkouri, K.; Deng, L.; Grootaert, H.; Schoonooghe, S.; Smet, A.; Bentahir, M.; Roose, K.; et al. Hierarchical and Redundant Roles of Activating FcgammaRs in Protection against Influenza Disease by M2e-Specific IgG1 and IgG2a Antibodies. J. Virol. 2017, 91, e02500-16. [Google Scholar] [CrossRef] [PubMed]
- Fujimoto, Y.; Tomioka, Y.; Takakuwa, H.; Uechi, G.; Yabuta, T.; Ozaki, K.; Suyama, H.; Yamamoto, S.; Morimatsu, M.; Mai le, Q.; et al. Cross-protective potential of anti-nucleoprotein human monoclonal antibodies against lethal influenza A virus infection. J. Gen. Virol. 2016, 97, 2104–2116. [Google Scholar] [CrossRef] [PubMed]
- Bullido, R.; Gomez-Puertas, P.; Albo, C.; Portela, A. Several protein regions contribute to determine the nuclear and cytoplasmic localization of the influenza A virus nucleoprotein. J. Gen. Virol. 2000, 81, 135–142. [Google Scholar] [CrossRef] [PubMed]
- Yewdell, J.W.; Frank, E.; Gerhard, W. Expression of influenza A virus internal antigens on the surface of infected P815 cells. J. Immunol. 1981, 126, 1814–1819. [Google Scholar] [PubMed]
- Bodewes, R.; Geelhoed-Mieras, M.M.; Wrammert, J.; Ahmed, R.; Wilson, P.C.; Fouchier, R.A.; Osterhaus, A.D.; Rimmelzwaan, G.F. In vitro assessment of the immunological significance of a human monoclonal antibody directed to the influenza a virus nucleoprotein. Clin. Vaccine Immunol. 2013, 20, 1333–1337. [Google Scholar] [CrossRef] [PubMed]
- Yassine, H.M.; Boyington, J.C.; McTamney, P.M.; Wei, C.J.; Kanekiyo, M.; Kong, W.P.; Gallagher, J.R.; Wang, L.; Zhang, Y.; Joyce, M.G.; et al. Hemagglutinin-stem nanoparticles generate heterosubtypic influenza protection. Nat. Med. 2015, 21, 1065–1070. [Google Scholar] [CrossRef] [PubMed]
- Impagliazzo, A.; Milder, F.; Kuipers, H.; Wagner, M.V.; Zhu, X.; Hoffman, R.M.; van Meersbergen, R.; Huizingh, J.; Wanningen, P.; Verspuij, J.; et al. A stable trimeric influenza hemagglutinin stem as a broadly protective immunogen. Science 2015, 349, 1301–1306. [Google Scholar] [CrossRef] [PubMed]
- Florek, N.W.; Weinfurter, J.T.; Jegaskanda, S.; Brewoo, J.N.; Powell, T.D.; Young, G.R.; Das, S.C.; Hatta, M.; Broman, K.W.; Hungnes, O.; et al. Modified vaccinia virus Ankara encoding influenza virus hemagglutinin induces heterosubtypic immunity in macaques. J. Virol. 2014, 88, 13418–13428. [Google Scholar] [CrossRef] [PubMed]
- El Bakkouri, K.; Descamps, F.; De Filette, M.; Smet, A.; Festjens, E.; Birkett, A.; Van Rooijen, N.; Verbeek, S.; Fiers, W.; Saelens, X. Universal vaccine based on ectodomain of matrix protein 2 of influenza A: Fc receptors and alveolar macrophages mediate protection. J. Immunol. 2011, 186, 1022–1031. [Google Scholar] [CrossRef] [PubMed]
- Jegerlehner, A.; Schmitz, N.; Storni, T.; Bachmann, M.F. Influenza A vaccine based on the extracellular domain of M2: Weak protection mediated via antibody-dependent NK cell activity. J. Immunol. 2004, 172, 5598–5605. [Google Scholar] [CrossRef] [PubMed]
- Kim, M.C.; Lee, Y.N.; Hwang, H.S.; Lee, Y.T.; Ko, E.J.; Jung, Y.J.; Cho, M.K.; Kim, Y.J.; Lee, J.S.; Ha, S.H.; et al. Influenza M2 virus-like particles confer a broader range of cross protection to the strain-specific pre-existing immunity. Vaccine 2014, 32, 5824–5831. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Lee, Y.N.; Kim, M.C.; Lee, Y.T.; Hwang, H.S.; Lee, J.; Kim, C.; Kang, S.M. Cross Protection against Influenza A Virus by Yeast-Expressed Heterologous Tandem Repeat M2 Extracellular Proteins. PLoS ONE 2015, 10, e0137822. [Google Scholar] [CrossRef] [PubMed]
- Lee, Y.N.; Lee, Y.T.; Kim, M.C.; Hwang, H.S.; Lee, J.S.; Kim, K.H.; Kang, S.M. Fc receptor is not required for inducing antibodies but plays a critical role in conferring protection after influenza M2 vaccination. Immunology 2014, 143, 300–309. [Google Scholar] [CrossRef] [PubMed]
- Sobhanie, M.; Matsuoka, Y.; Jegaskanda, S.; Fitzgerald, T.; Mallory, R.; Chen, Z.; Luke, C.; Treanor, J.; Subbarao, K. Evaluation of the Safety and Immunogenicity of a Candidate Pandemic Live Attenuated Influenza Vaccine (pLAIV) Against Influenza A(H7N9). J. Infect. Dis. 2015, 213, 922–929. [Google Scholar] [CrossRef] [PubMed]
- Zhong, W.; Liu, F.; Wilson, J.R.; Holiday, C.; Li, Z.N.; Bai, Y.; Tzeng, W.P.; Stevens, J.; York, I.A.; Levine, M.Z. Antibody-Dependent Cell-Mediated Cytotoxicity to Hemagglutinin of Influenza A Viruses After Influenza Vaccination in Humans. In Open Forum Infect Diseases; Oxford University Press: Oxford, UK, 2016. [Google Scholar]
- Jacobsen, H.; Rajendran, M.; Choi, A.; Sjursen, H.; Brokstad, K.A.; Cox, R.J.; Palese, P.; Krammer, F.; Nachbagauer, R. Influenza Virus Hemagglutinin Stalk-Specific Antibodies in Human Serum are a Surrogate Marker for In Vivo Protection in a Serum Transfer Mouse Challenge Model. MBio 2017, 8, e01463-17. [Google Scholar] [CrossRef] [PubMed]
- Jegaskanda, S.; Luke, C.; Hickman, H.D.; Sangster, M.Y.; Wieland-Alter, W.F.; McBride, J.M.; Yewdell, J.W.; Wright, P.F.; Treanor, J.; Rosenberger, C.M.; et al. Generation and Protective Ability of Influenza Virus-Specific Antibody-Dependent Cellular Cytotoxicity in Humans Elicited by Vaccination, Natural Infection, and Experimental Challenge. J. Infect. Dis. 2016, 214, 945–952. [Google Scholar] [CrossRef] [PubMed]
- Park, J.K.; Han, A.; Czajkowski, L.; Reed, S.; Athota, R.; Bristol, T.; Rosas, L.A.; Cervantes-Medina, A.; Taubenberger, J.K.; Memoli, M.J. Evaluation of Preexisting Anti-Hemagglutinin Stalk Antibody as a Correlate of Protection in a Healthy Volunteer Challenge with Influenza A/H1N1pdm Virus. MBio 2018, 9, e02284-17. [Google Scholar] [CrossRef] [PubMed]
- Brunner, K.T.; Mauel, J.; Cerottini, J.C.; Chapuis, B. Quantitative assay of the lytic action of immune lymphoid cells on 51-Cr-labelled allogeneic target cells in vitro; inhibition by isoantibody and by drugs. Immunology 1968, 14, 181–196. [Google Scholar] [PubMed]
- Hashimoto, G.; Wright, P.F.; Karzon, D.T. Antibody-dependent cell-mediated cytotoxicity against influenza virus-infected cells. J. Infect. Dis. 1983, 148, 785–794. [Google Scholar] [CrossRef] [PubMed]
- Korzeniewski, C.; Callewaert, D.M. An enzyme-release assay for natural cytotoxicity. J. Immunol. Methods 1983, 64, 313–320. [Google Scholar] [CrossRef]
- Decker, T.; Lohmann-Matthes, M.L. A quick and simple method for the quantitation of lactate dehydrogenase release in measurements of cellular cytotoxicity and tumor necrosis factor (TNF) activity. J. Immunol. Methods 1988, 115, 61–69. [Google Scholar] [CrossRef]
- Vanderven, H.A.; Jegaskanda, S.; Wines, B.D.; Hogarth, P.M.; Carmuglia, S.; Rockman, S.; Chung, A.W.; Kent, S.J. Antibody-Dependent Cellular Cytotoxicity Responses to Seasonal Influenza Vaccination in Older Adults. J. Infect. Dis. 2017, 217, 12–23. [Google Scholar] [CrossRef] [PubMed]
- Srivastava, V.; Yang, Z.; Hung, I.F.; Xu, J.; Zheng, B.; Zhang, M.Y. Identification of dominant antibody-dependent cell-mediated cytotoxicity epitopes on the hemagglutinin antigen of pandemic H1N1 influenza virus. J. Virol. 2013, 87, 5831–5840. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Ye, Z.W.; Yuan, S.; Poon, K.M.; Wen, L.; Yang, D.; Sun, Z.; Li, C.; Hu, M.; Shuai, H.; Zhou, J.; et al. Antibody-Dependent Cell-Mediated Cytotoxicity Epitopes on the Hemagglutinin Head Region of Pandemic H1N1 Influenza Virus Play Detrimental Roles in H1N1-Infected Mice. Front. Immunol. 2017, 8, 317. [Google Scholar] [CrossRef] [PubMed]
- Kim, G.G.; Donnenberg, V.S.; Donnenberg, A.D.; Gooding, W.; Whiteside, T.L. A novel multiparametric flow cytometry-based cytotoxicity assay simultaneously immunophenotypes effector cells: Comparisons to a 4 h 51Cr-release assay. J. Immunol. Methods 2007, 325, 51–66. [Google Scholar] [CrossRef] [PubMed]
- Perussia, B. The Cytokine Profile of Resting and Activated NK Cells. Methods 1996, 9, 370–378. [Google Scholar] [CrossRef] [PubMed]
- Wagstaffe, H.R.; Nielsen, C.M.; Riley, E.M.; Goodier, M.R. IL-15 Promotes Polyfunctional NK Cell Responses to Influenza by Boosting IL-12 Production. J. Immunol. 2018, 200, 2738–2747. [Google Scholar] [CrossRef] [PubMed]
- Alter, G.; Malenfant, J.M.; Altfeld, M. CD107a as a functional marker for the identification of natural killer cell activity. J. Immunol. Methods 2004, 294, 15–22. [Google Scholar] [CrossRef] [PubMed]
- Krzewski, K.; Gil-Krzewska, A.; Nguyen, V.; Peruzzi, G.; Coligan, J.E. LAMP1/CD107a is required for efficient perforin delivery to lytic granules and NK-cell cytotoxicity. Blood 2013, 121, 4672–4683. [Google Scholar] [CrossRef] [PubMed]
- Cohnen, A.; Chiang, S.C.; Stojanovic, A.; Schmidt, H.; Claus, M.; Saftig, P.; Janssen, O.; Cerwenka, A.; Bryceson, Y.T.; Watzl, C. Surface CD107a/LAMP-1 protects natural killer cells from degranulation-associated damage. Blood 2013, 122, 1411–1418. [Google Scholar] [CrossRef] [PubMed]
- Stratov, I.; Chung, A.; Kent, S.J. Robust NK cell-mediated human immunodeficiency virus (HIV)-specific antibody-dependent responses in HIV-infected subjects. J. Virol. 2008, 82, 5450–5459. [Google Scholar] [CrossRef] [PubMed]
- Chung, A.W.; Rollman, E.; Center, R.J.; Kent, S.J.; Stratov, I. Rapid degranulation of NK cells following activation by HIV-specific antibodies. J. Immunol. 2009, 182, 1202–1210. [Google Scholar] [CrossRef] [PubMed]
- Chung, A.W.; Isitman, G.; Navis, M.; Kramski, M.; Center, R.J.; Kent, S.J.; Stratov, I. Immune escape from HIV-specific antibody-dependent cellular cytotoxicity (ADCC) pressure. Proc. Natl. Acad. Sci. USA 2011, 108, 7505–7510. [Google Scholar] [CrossRef] [PubMed]
- Jegaskanda, S.; Job, E.R.; Kramski, M.; Laurie, K.; Isitman, G.; de Rose, R.; Winnall, W.R.; Stratov, I.; Brooks, A.G.; Reading, P.C.; et al. Cross-reactive influenza-specific antibody-dependent cellular cytotoxicity antibodies in the absence of neutralizing antibodies. J. Immunol. 2013, 190, 1837–1848. [Google Scholar] [CrossRef] [PubMed]
- Jegaskanda, S.; Weinfurter, J.T.; Friedrich, T.C.; Kent, S.J. Antibody-dependent cellular cytotoxicity is associated with control of pandemic H1N1 influenza virus infection of macaques. J. Virol. 2013, 87, 5512–5522. [Google Scholar] [CrossRef] [PubMed]
- Valkenburg, S.A.; Zhang, Y.; Chan, K.Y.; Leung, K.; Wu, J.T.; Poon, L.L. Preexisting Antibody-Dependent Cellular Cytotoxicity-Activating Antibody Responses Are Stable Longitudinally and Cross-reactive Responses Are Not Boosted by Recent Influenza Exposure. J. Infect. Dis. 2016, 214, 1159–1163. [Google Scholar] [CrossRef] [PubMed]
- De Vries, R.D.; Nieuwkoop, N.J.; Pronk, M.; de Bruin, E.; Leroux-Roels, G.; Huijskens, E.G.W.; van Binnendijk, R.S.; Krammer, F.; Koopmans, M.P.G.; Rimmelzwaan, G.F. Influenza virus-specific antibody dependent cellular cytoxicity induced by vaccination or natural infection. Vaccine 2017, 35, 238–247. [Google Scholar] [CrossRef] [PubMed]
- De Vries, R.D.; Nieuwkoop, N.J.; van der Klis, F.R.M.; Koopmans, M.P.G.; Krammer, F.; Rimmelzwaan, G.F. Primary Human Influenza B Virus Infection Induces Cross-Lineage Hemagglutinin Stalk-Specific Antibodies Mediating Antibody-Dependent Cellular Cytoxicity. J. Infect. Dis. 2017, 217, 3–11. [Google Scholar] [CrossRef] [PubMed]
- Jegaskanda, S.; Amarasena, T.H.; Laurie, K.L.; Tan, H.X.; Butler, J.; Parsons, M.S.; Alcantara, S.; Petravic, J.; Davenport, M.P.; Hurt, A.C.; et al. Standard trivalent influenza virus protein vaccination does not prime antibody-dependent cellular cytotoxicity in macaques. J. Virol. 2013, 87, 13706–13718. [Google Scholar] [CrossRef] [PubMed]
- Madhavi, V.; Ana-Sosa-Batiz, F.E.; Jegaskanda, S.; Center, R.J.; Winnall, W.R.; Parsons, M.S.; Ananworanich, J.; Cooper, D.A.; Kelleher, A.D.; Hsu, D.; et al. Antibody-dependent effector functions against HIV decline in subjects receiving antiretroviral therapy. J. Infect. Dis. 2015, 211, 529–538. [Google Scholar] [CrossRef] [PubMed]
- Wang, K.; Tomaras, G.D.; Jegaskanda, S.; Moody, M.A.; Liao, H.X.; Goodman, K.N.; Berman, P.W.; Rerks-Ngarm, S.; Pitisuttithum, P.; Nitayapan, S.; et al. Monoclonal Antibodies, Derived from Humans Vaccinated with the RV144 HIV Vaccine Containing the HVEM Binding Domain of Herpes Simplex Virus (HSV) Glycoprotein D, Neutralize HSV Infection, Mediate Antibody-Dependent Cellular Cytotoxicity, and Protect Mice from Ocular Challenge with HSV-1. J. Virol. 2017, 91, e00411-17. [Google Scholar] [PubMed]
- Henry Dunand, C.J.; Leon, P.E.; Huang, M.; Choi, A.; Chromikova, V.; Ho, I.Y.; Tan, G.S.; Cruz, J.; Hirsh, A.; Zheng, N.Y.; et al. Both Neutralizing and Non-Neutralizing Human H7N9 Influenza Vaccine-Induced Monoclonal Antibodies Confer Protection. Cell Host Microbe 2016, 19, 800–813. [Google Scholar] [CrossRef] [PubMed]
- Sutton, T.C.; Lamirande, E.W.; Bock, K.W.; Moore, I.N.; Koudstaal, W.; Rehman, M.; Weverling, G.J.; Goudsmit, J.; Subbarao, K. In Vitro Neutralization Is Not Predictive of Prophylactic Efficacy of Broadly Neutralizing Monoclonal Antibodies CR6261 and CR9114 against Lethal H2 Influenza Virus Challenge in Mice. J. Virol. 2017, 91, e01603-17. [Google Scholar] [CrossRef] [PubMed]
- Wines, B.D.; Billings, H.; McLean, M.R.; Kent, S.J.; Hogarth, P.M. Antibody Functional Assays as Measures of Fc Receptor-Mediated Immunity to HIV-New Technologies and their Impact on the HIV Vaccine Field. Curr. HIV Res. 2017, 15, 202–215. [Google Scholar] [PubMed]
- Kristensen, A.B.; Lay, W.N.; Ana-Sosa-Batiz, F.; Vanderven, H.A.; Madhavi, V.; Laurie, K.L.; Carolan, L.; Wines, B.D.; Hogarth, M.; Wheatley, A.K.; et al. Antibody Responses with Fc-Mediated Functions after Vaccination of HIV-Infected Subjects with Trivalent Influenza Vaccine. J. Virol. 2016, 90, 5724–5734. [Google Scholar] [CrossRef] [PubMed]
- Wines, B.D.; Vanderven, H.A.; Esparon, S.E.; Kristensen, A.B.; Kent, S.J.; Hogarth, P.M. Dimeric FcgammaR Ectodomains as Probes of the Fc Receptor Function of Anti-Influenza Virus IgG. J. Immunol. 2016, 197, 1507–1516. [Google Scholar] [CrossRef] [PubMed]
- Greenberg, S.B.; Criswell, B.S.; Six, H.R.; Couch, R.B. Lymphocyte cytotoxicity to influenza virus-infected cells: Response to vaccination and virus infection. Infect. Immun. 1978, 20, 640–645. [Google Scholar] [PubMed]
- Greenberg, S.B.; Criswell, B.S.; Six, H.R.; Couch, R.B. Lymphocyte cytotoxicity to influenza virus-infected cells. II. Requirement for antibody and non-T lymphocytes. J. Immunol. 1977, 119, 2100–2106. [Google Scholar] [PubMed]
- Jegaskanda, S.; Laurie, K.L.; Amarasena, T.H.; Winnall, W.R.; Kramski, M.; De Rose, R.; Barr, I.G.; Brooks, A.G.; Reading, P.C.; Kent, S.J. Age-associated cross-reactive antibody-dependent cellular cytotoxicity toward 2009 pandemic influenza A virus subtype H1N1. J. Infect. Dis. 2013, 208, 1051–1061. [Google Scholar] [CrossRef] [PubMed]
- Jegaskanda, S.; Vandenberg, K.; Laurie, K.L.; Loh, L.; Kramski, M.; Winnall, W.R.; Kedzierska, K.; Rockman, S.; Kent, S.J. Cross-reactive influenza-specific antibody-dependent cellular cytotoxicity in intravenous immunoglobulin as a potential therapeutic against emerging influenza viruses. J. Infect. Dis. 2014, 210, 1811–1822. [Google Scholar] [CrossRef] [PubMed]
- Hashimoto, G.; Wright, P.F.; Karzon, D.T. Ability of human cord blood lymphocytes to mediate antibody-dependent cellular cytotoxicity against influenza virus-infected cells. Infect. Immun. 1983, 42, 214–218. [Google Scholar] [PubMed]
- Kim, J.H.; Reber, A.J.; Kumar, A.; Ramos, P.; Sica, G.; Music, N.; Guo, Z.; Mishina, M.; Stevens, J.; York, I.A.; et al. Non-neutralizing antibodies induced by seasonal influenza vaccine prevent, not exacerbate A(H1N1)pdm09 disease. Sci. Rep. 2016, 6, 37341. [Google Scholar] [CrossRef] [PubMed]
- Khurana, S.; Loving, C.L.; Manischewitz, J.; King, L.R.; Gauger, P.C.; Henningson, J.; Vincent, A.L.; Golding, H. Vaccine-induced anti-HA2 antibodies promote virus fusion and enhance influenza virus respiratory disease. Sci. Transl. Med. 2013, 5, 200ra114. [Google Scholar] [CrossRef] [PubMed]
- Sridhar, S.; Begom, S.; Bermingham, A.; Hoschler, K.; Adamson, W.; Carman, W.; Bean, T.; Barclay, W.; Deeks, J.J.; Lalvani, A. Cellular immune correlates of protection against symptomatic pandemic influenza. Nat. Med. 2013, 19, 1305–1312. [Google Scholar] [CrossRef] [PubMed]
- Leon, P.E.; He, W.; Mullarkey, C.E.; Bailey, M.J.; Miller, M.S.; Krammer, F.; Palese, P.; Tan, G.S. Optimal activation of Fc-mediated effector functions by influenza virus hemagglutinin antibodies requires two points of contact. Proc. Natl. Acad. Sci. USA 2016, 113, E5944–E5951. [Google Scholar] [CrossRef] [PubMed]
- Cox, F.; Kwaks, T.; Brandenburg, B.; Koldijk, M.H.; Klaren, V.; Smal, B.; Korse, H.J.; Geelen, E.; Tettero, L.; Zuijdgeest, D.; et al. Antibody-Mediated FcgammaRIIIa Activity Is Both Dependent on FcR Engagement and Interactions between HA and Sialic Acids. Front. Immunol. 2016, 7, 399. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.; Boutz, D.R.; Chromikova, V.; Joyce, M.G.; Vollmers, C.; Leung, K.; Horton, A.P.; DeKosky, B.J.; Lee, C.H.; Lavinder, J.J.; et al. Molecular-level analysis of the serum antibody repertoire in young adults before and after seasonal influenza vaccination. Nat. Med. 2016, 22, 1456–1464. [Google Scholar] [CrossRef] [PubMed]
- Ekiert, D.C.; Kashyap, A.K.; Steel, J.; Rubrum, A.; Bhabha, G.; Khayat, R.; Lee, J.H.; Dillon, M.A.; O’Neil, R.E.; Faynboym, A.M.; et al. Cross-neutralization of influenza A viruses mediated by a single antibody loop. Nature 2012, 489, 526–532. [Google Scholar] [CrossRef] [PubMed]
- Zarnitsyna, V.I.; Lavine, J.; Ellebedy, A.; Ahmed, R.; Antia, R. Multi-epitope Models Explain How Pre-existing Antibodies Affect the Generation of Broadly Protective Responses to Influenza. PLoS Pathog. 2016, 12, e1005692. [Google Scholar] [CrossRef] [PubMed]
- Wheatley, A.K.; Kent, S.J. Prospects for antibody-based universal influenza vaccines in the context of widespread pre-existing immunity. Expert Rev. Vaccines 2015, 14, 1227–1239. [Google Scholar] [CrossRef] [PubMed]
- Goodier, M.R.; Lusa, C.; Sherratt, S.; Rodriguez-Galan, A.; Behrens, R.; Riley, E.M. Sustained Immune Complex-Mediated Reduction in CD16 Expression after Vaccination Regulates NK Cell Function. Front. Immunol. 2016, 7, 384. [Google Scholar] [CrossRef] [PubMed]
© 2018 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Jegaskanda, S. The Potential Role of Fc-Receptor Functions in the Development of a Universal Influenza Vaccine. Vaccines 2018, 6, 27. https://doi.org/10.3390/vaccines6020027
Jegaskanda S. The Potential Role of Fc-Receptor Functions in the Development of a Universal Influenza Vaccine. Vaccines. 2018; 6(2):27. https://doi.org/10.3390/vaccines6020027
Chicago/Turabian StyleJegaskanda, Sinthujan. 2018. "The Potential Role of Fc-Receptor Functions in the Development of a Universal Influenza Vaccine" Vaccines 6, no. 2: 27. https://doi.org/10.3390/vaccines6020027
APA StyleJegaskanda, S. (2018). The Potential Role of Fc-Receptor Functions in the Development of a Universal Influenza Vaccine. Vaccines, 6(2), 27. https://doi.org/10.3390/vaccines6020027



