Immunogenicity of a Recombinant Avian Influenza H2 Protein Using an Abdominal Inoculation Model in Chickens
Abstract
1. Introduction
2. Materials and Methods
2.1. Synthesis of H2 Gene
2.2. Cloning of H2 Gene into a Baculovirus Vector pAcGP67B
2.3. Production of Recombinant Baculovirus
2.4. Characterization of Recombinant H2 Protein
2.5. Chickens
2.6. Intra-Abdominal Inoculation
2.7. Isolation of Abdominal Leukocytes
2.8. Cell Counts and Viability
2.9. Leukocyte Populations
2.10. Cellular Activation
2.11. Flow Cytometry
2.12. Gene Expression
2.13. Antibody Detection
2.14. Statistical Analysis
3. Results
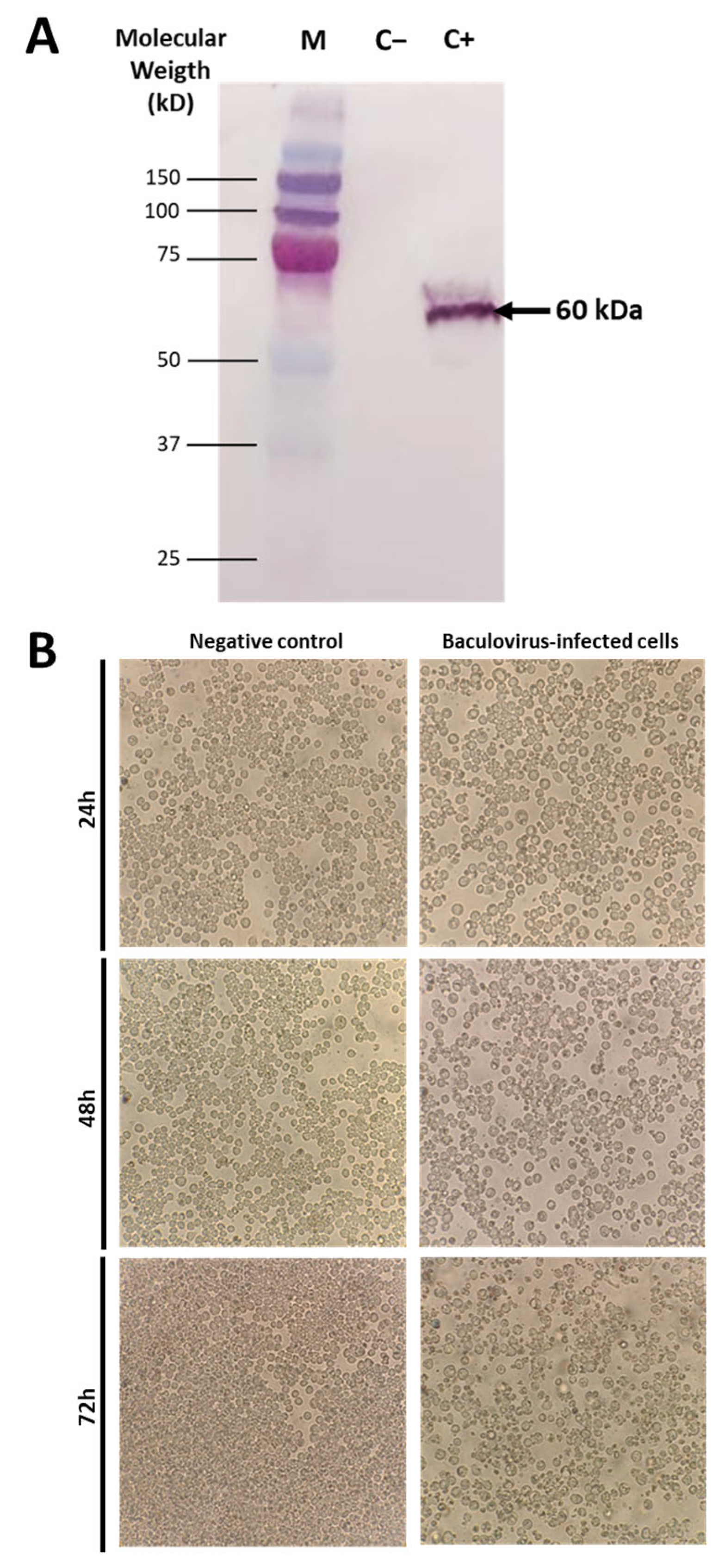
3.1. Modified H2 Gene Segment Inserted into Linearized Baculovirus Genome
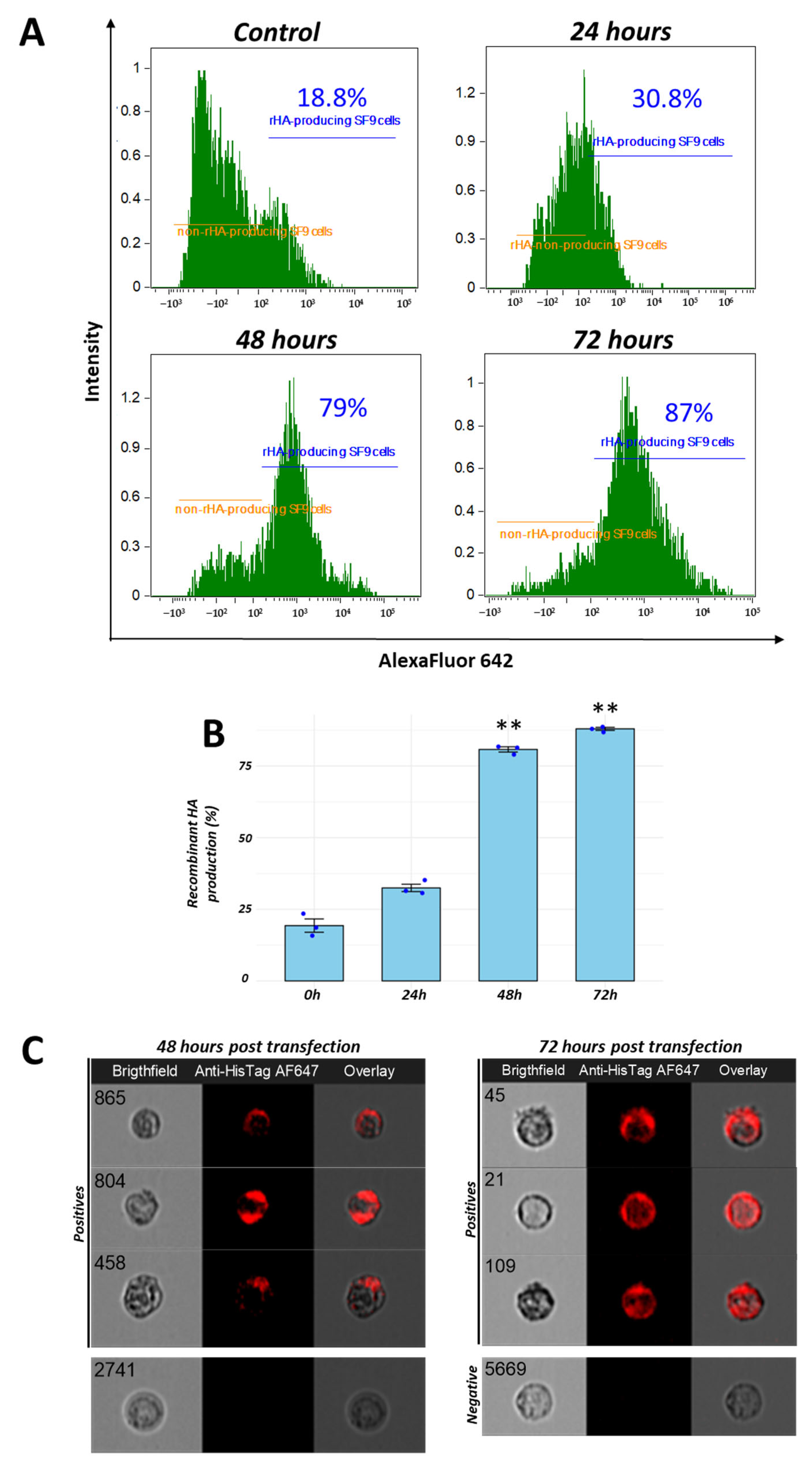
3.2. Maximum Recombinant H2 Protein Production Was Detected Following 48 h of Inoculation into Sf9 Cells
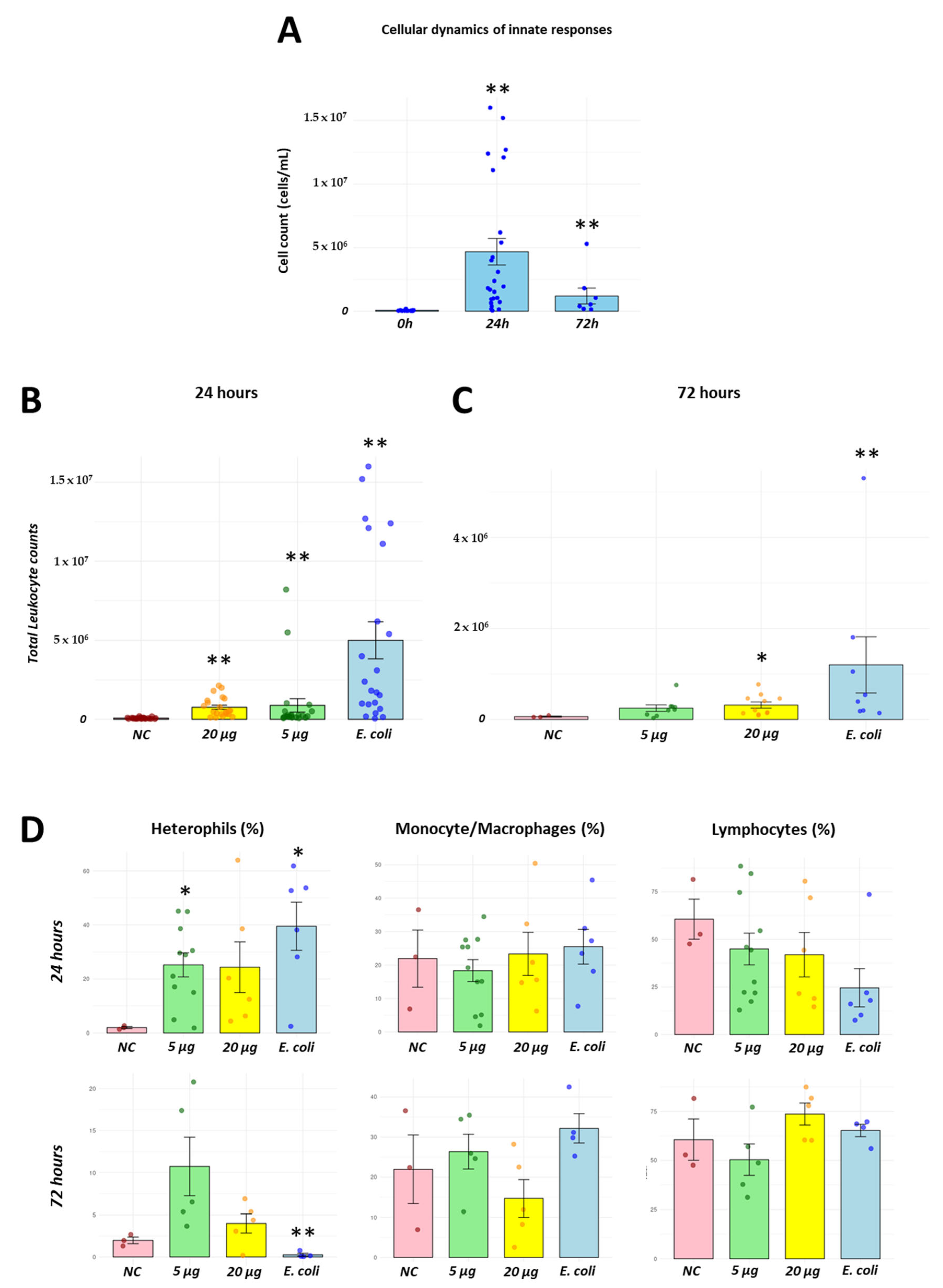
3.3. Recombinant H2 Protein Promotes a Leukocyte Infiltration in Chicken In Vivo Infection Model
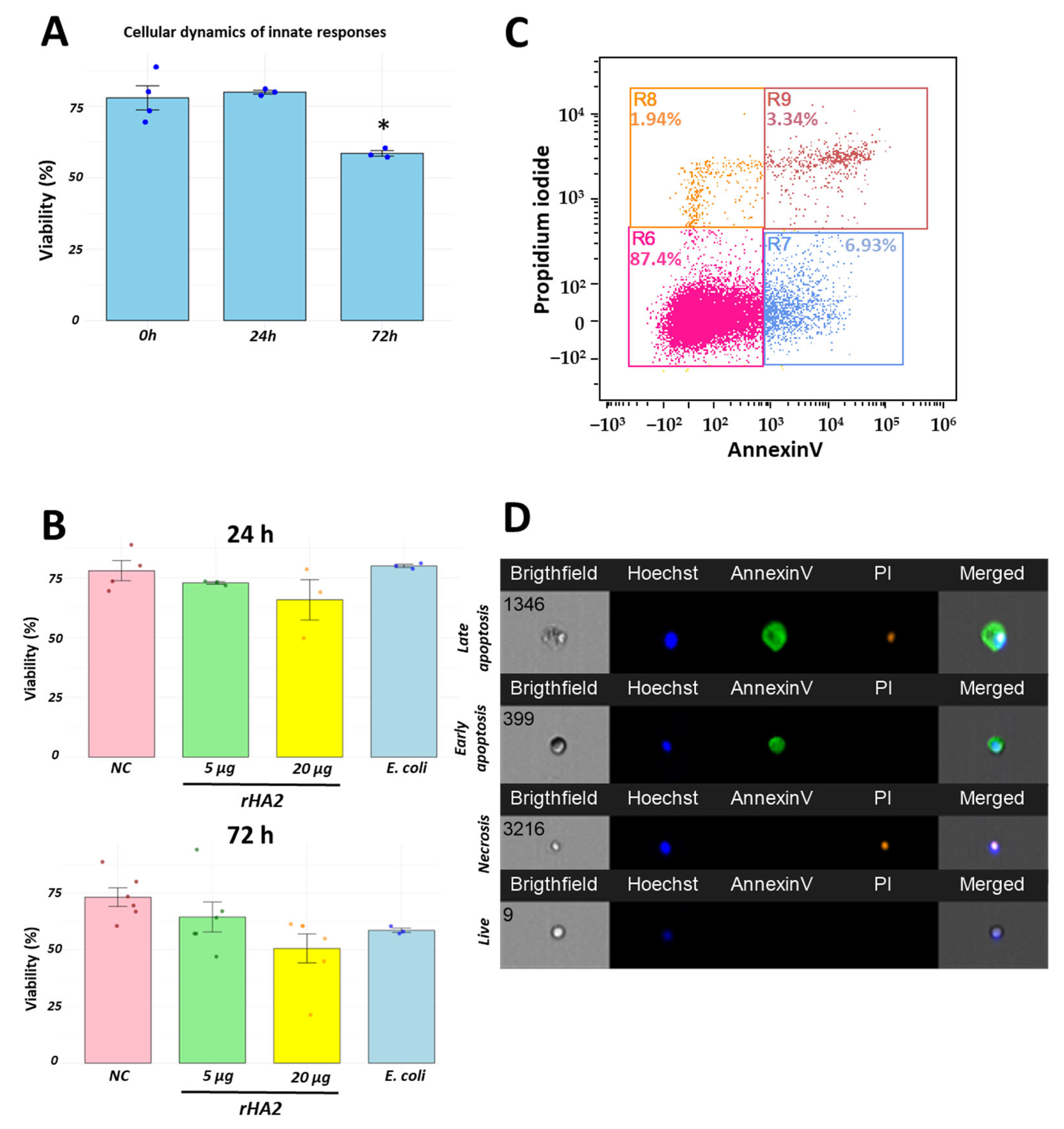
3.4. Recombinant H2 Protein Promotes Cellular Recruitment with High Cell Viability
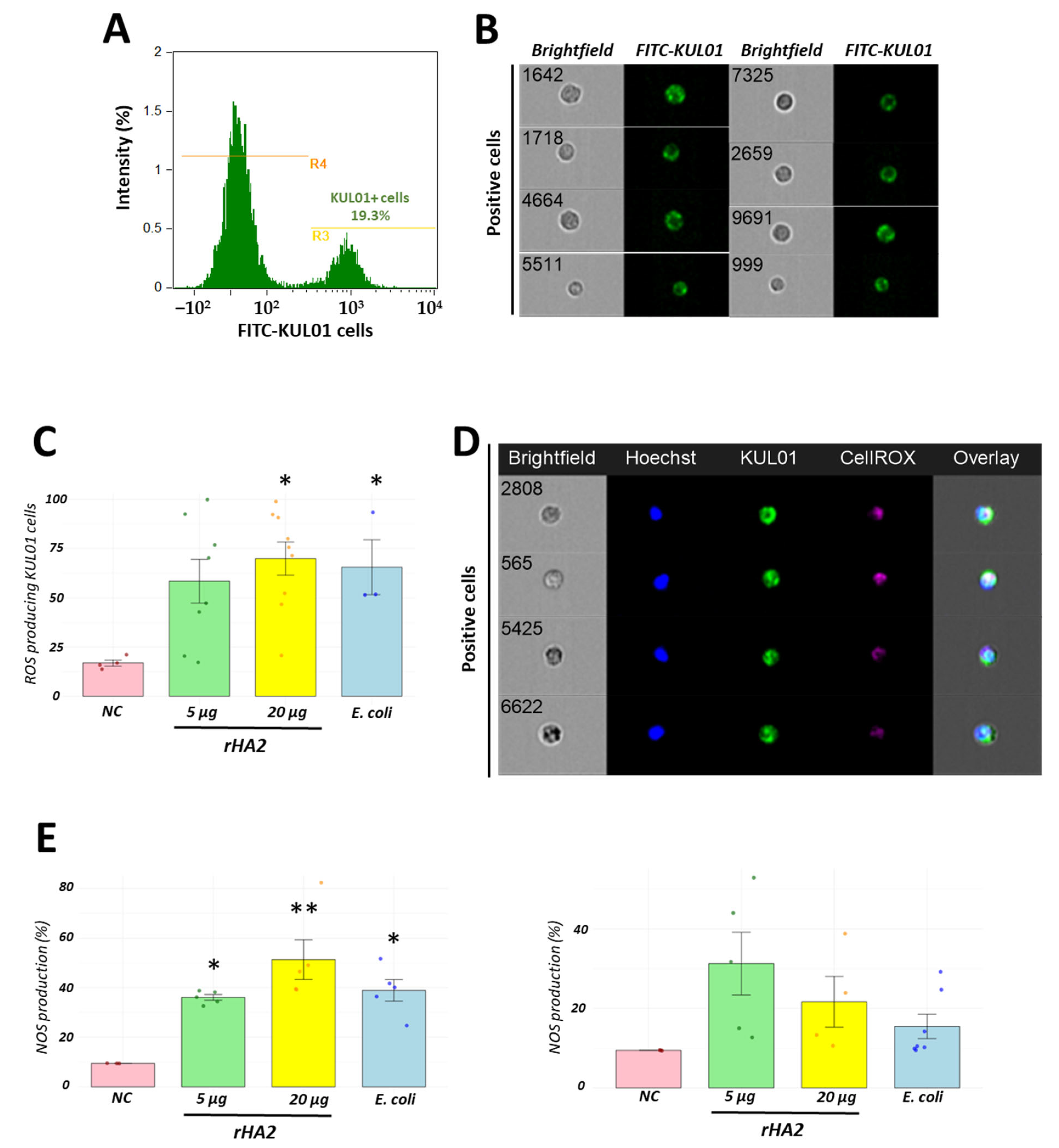
3.5. Recombinant H2 Protein Promotes In Vivo Macrophage Activation
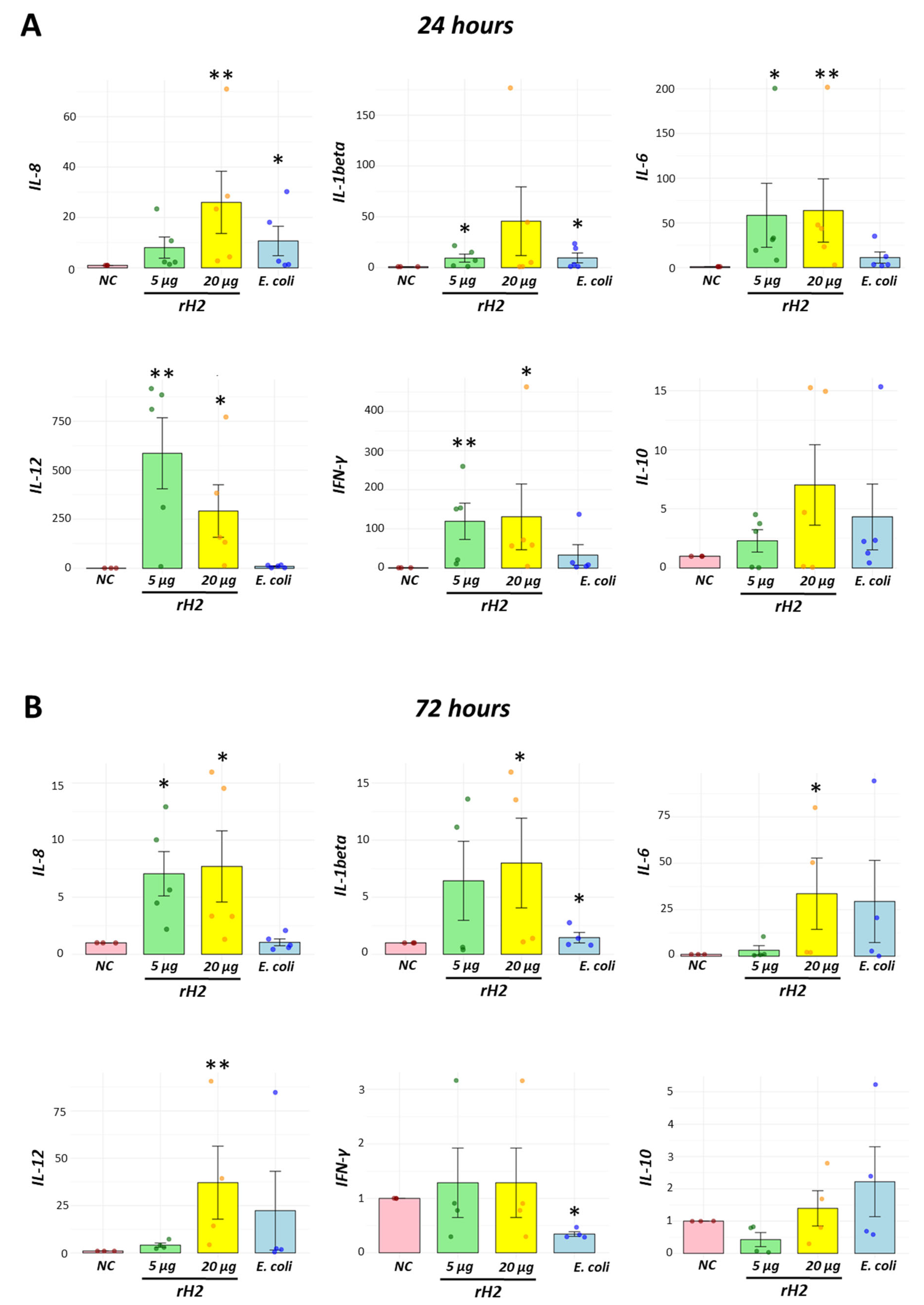
3.6. A Dynamic Immune Response Was Detected by Cytokine Expression in a Concentration-Dependent Manner
3.7. The Recombinant H2 Protein Promotes the Production of Inhibitory Antibodies
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
| HA | Hemagglutinin |
| H2 | subtype-2 hemagglutinin |
| µg | Micrograms |
| h.p.i. | Hours post inoculation |
| LPAIV | Low-pathogenic avian influenza virus |
| ROS | Reactive oxygen species |
| DNA | Desoxyribonucleic acid |
| SDS-PAGE | Sodium dodecyl sulfate polyacrylamide gel electrophoresis |
| PBS | Phosphate-buffered saline |
| FITC | Fluorescein isothiocyanate |
| PI | Propidium iodide |
| NOS | Nitric oxide species |
References
- Medina, R.A.; García-Sastre, A. Influenza A viruses: New research developments. Nat. Rev. Microbiol. 2011, 9, 590–603. [Google Scholar] [CrossRef]
- Bouvier, N.M.; Palese, P. The Biology of Influenza viruses. Vaccine 2008, 12, D49–D53. [Google Scholar] [CrossRef]
- Gamblin, S.J.; Skehel, J.J. Influenza hemagglutinin and neuraminidase membrane glycoproteins. J. Biol. Chem. 2010, 285, 28403–28409. [Google Scholar] [CrossRef] [PubMed]
- Cline, T.D.; Beck, D.; Bianchini, E. Influenza virus replication in macrophages: Balancing protection and pathogenesis. J. Gen. Virol. 2017, 98, 2401–2412. [Google Scholar] [CrossRef] [PubMed]
- Krammer, F.; Smith, G.J.D.; Fouchier, R.A.M.; Peiris, M.; Kedzierska, K.; Doherty, P.C.; Palese, P.; Shaw, M.L.; Treanor, J.; Webster, R.G.; et al. Influenza. Nat. Rev. Dis. Prim. 2018, 4, 3. [Google Scholar] [CrossRef]
- Hutchinson, E.C.; Yamauchi, Y. Understanding Influenza. In Influenza Virus; Yamauchi, Y., Ed.; Methods in Molecular Biology; Humana Press Inc.: New York, NY, USA, 2018; Volume 1836, pp. 1–21. [Google Scholar] [CrossRef]
- Lowen, A.C. It’s in the mix: Reassortment of segmented viral genomes. PLoS Pathog. 2018, 14, e1007200. [Google Scholar] [CrossRef]
- McDonald, S.M.; Nelson, M.I.; Turner, P.E.; Patton, J.T. Reassortment in segmented RNA viruses: Mechanisms and outcomes. Nat. Rev. Microbiol. 2016, 14, 448–460. [Google Scholar] [CrossRef]
- Ferhadian, D.; Contrant, M.; Printz-Schweigert, A.; Smyth, R.P.; Paillart, J.C.; Marquet, R. Structural and functional motifs in influenza virus RNAs. Front. Microbiol. 2018, 9, 559. [Google Scholar] [CrossRef]
- Webster, R.G.; Bean, W.J.; Gorman, O.T.; Chambers, T.M.; Kawaoka, Y. Evolution and ecology of influenza A viruses. Microbiol. Rev. 1992, 56, 152–179. [Google Scholar] [CrossRef] [PubMed]
- Machalaba, C.; Elwood, S.; Forcella, S.; Smith, K.; Hamilton, K.; Jebara, K.; Swayne, D.E.; Webby, R.J.; Mumford, E.; Mazet, J.A.; et al. Global avian influenza surveillance in wild birds: A strategy to capture viral diversity. Emerg. Infect. Dis. 2015, 21, e141415. [Google Scholar] [CrossRef]
- Venkatesh, D.; Poen, M.J.; Bestebroer, T.M.; Scheuer, R.D.; Vuong, O.; Chkhaidze, M.; Machablishvili, A.; Mamuchadze, J.; Ninua, L.; Fedorova, N.B.; et al. Avian influenza viruses in wild birds: Virus evolution in a multihost ecosystem. J. Virol. 2018, 92, 1–20. [Google Scholar] [CrossRef]
- Olsen, B.; Munster, V.J.; Wallensten, A.; Waldenström, J.; Osterhaus, A.; Fouchier, R. Global patterns of influenza A virus in wild birds. Science 2006, 384, 384–388. [Google Scholar] [CrossRef]
- The Global Consortium for H5N8 and Related Influenza Viruses. Role for migratory wild birds in the global spread of avian influenza H5N8. Science 2016, 354, 213–218. [Google Scholar] [CrossRef]
- Nelson, M.I.; Pollett, S.; Ghersi, B.; Silva, M.; Simons, M.P.; Icochea, E.; Gonzalez, A.E.; Segovia, K.; Kasper, M.R.; Montgomery, J.M.; et al. The genetic diversity of influenza A viruses in wild birds in Peru. PLoS ONE 2016, 11, e0146059. [Google Scholar] [CrossRef] [PubMed]
- Castro-Sanguinetti, G.R.; Simas, P.V.M.; Apaza-Chiara, A.P.; Callupe-Leyva, J.A.; Rondon-Espinoza, J.A.; Gavidia, C.M.; More-Bayona, J.A.; Veliz, R.I.G.; Vakharia, V.N.; Icochea, M.E.; et al. Genetic subtyping and phylogenetic analysis of HA and NA from avian influenza virus in wild birds from Peru reveals unique features among circulating strains in America. PLoS ONE 2022, 17, e0268957. [Google Scholar] [CrossRef] [PubMed]
- Castro-Sanguinetti, G.R.; González-Veliz, R.; Callupe-Leyva, A.; Apaza-Chiara, A.P.; Jara, J.; Silva, W.; Icochea, E.; More-Bayona, J.A. Highly pathogenic avian influenza virus H5N1 clade 2.3.4.4b from Peru forms a monophyletic group with Chilean isolates in South America. Sci. Rep. 2024, 14, 3635. [Google Scholar] [CrossRef]
- Andrews, S.F.; Raab, J.E.; Gorman, J.; Gillespie, R.A.; Cheung, C.S.F.; Rawi, R.; Cominsky, L.Y.; Boyington, J.C.; Creanga, A.; Shen, C.-H.; et al. A single residue in influenza virus H2 hemagglutinin enhances the breadth of the B cell response elicited by H2 vaccination. Nat. Med. 2022, 28, 373–382. [Google Scholar] [CrossRef]
- Reneer, Z.B.; Skarlupka, A.L.; Jamieson, P.J.; Ross, T.M. Broadly Reactive H2 Hemagglutinin Vaccines Elicit Cross-Reactive Antibodies in Ferrets Preimmune to Seasonal Influenza A Viruses. MSphere 2021, 6, 1–20. [Google Scholar] [CrossRef]
- Beau Reneer, Z.; Ross, T.M. H2 influenza viruses: Designing vaccines against future H2 pandemics. Biochem. Soc. Trans. 2019, 47, 251–264. [Google Scholar] [CrossRef]
- Beau Reneer, Z.; Abreu, R.B.; Jamal, U.S.; Corn, M.R.; Paugh, J.L.; Ross, T.M. Seasonal influenza vaccination does not effectively expand H2 cross-reactive antibodies in humans. Vaccine 2021, 39, 4173–4183. [Google Scholar] [CrossRef] [PubMed]
- Suarez, D.L.; Pantin-Jackwood, M.J. Recombinant viral-vectored vaccines for the control of avian influenza in poultry. Vet. Microbiol. 2017, 206, 144–151. [Google Scholar] [CrossRef] [PubMed]
- Casais, R.; Dove, B.; Cavanagh, D.; Britton, P. Recombinant avian infectious bronchitis virus expressing a heterologous spike gene demonstrates that the spike protein is a determinant of cell tropism. J. Virol. 2003, 77, 9084–9089. [Google Scholar] [CrossRef]
- Schultz-Cherry, S.; Jones, J.C. Influenza Vaccines: The Good, the Bad, and the Eggs, 1st ed.; Elsevier Inc.: Amsterdam, The Netherlands, 2010; Volume 77. [Google Scholar] [CrossRef]
- Rimmelzwaan, G.F.; Osterhaus, A.D.M.E. Influenza vaccines: New developments. Curr. Opin. Pharmacol. 2001, 1, 491–496. [Google Scholar] [CrossRef]
- Gouma, S.; Anderson, E.M.; Hensley, S.E. Challenges of making effective influenza vaccines. Annu. Rev. Virol. 2020, 7, 495–512. [Google Scholar] [CrossRef]
- More-Bayona, J.A.; Kumar, A.; Barreda, D.R. Contribution of leukocytes to the induction and resolution of the acute in fl ammatory response in chickens. Dev. Comp. Immunol. 2017, 74, 167–177. [Google Scholar] [CrossRef]
- O’Donnell Ea Ernst, D.N.; Hingorani, R. Multiparameter flow cytometry: Advances in high resolution analysis. Immune Netw. 2013, 13, 43–54. [Google Scholar] [CrossRef]
- George, T.C.; Basiji Da Hall, B.E.; Lynch, D.H.; Ortyn, W.E.; Perry, D.J.; Seo, M.J.; Zimmerman, C.A.; Morrissey, P.J. Distinguishing modes of cell death using the ImageStream multispectral imaging flow cytometer. Cytom. A 2004, 59, 237–245. [Google Scholar] [CrossRef]
- Havixbeck, J.J.; Wong, M.E.; More Bayona, J.A.; Barreda, D.R. Multi-parametric analysis of phagocyte antimicrobial responses using imaging flow cytometry. J. Immunol. Methods 2015, 423, 85–92. [Google Scholar] [CrossRef] [PubMed]
- More Bayona, J.A.; Karuppannan, A.K.; Trites, M.J.; Barreda, D.R. Application of imaging flow cytometry for characterization of acute inflammation in non-classical animal model systems. Methods 2016, 112, 167–174. [Google Scholar] [CrossRef] [PubMed]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real- time quantitative PCR and the 2^delta delta CT method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef]
- R Core Team. A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2021; Available online: https://www.R-project.org/ (accessed on 27 August 2025).
- Alexander, D.J.; Brown, I.H. History of highly pathogenic avian influenza. Rev. Sci. Et Tech. 2009, 28, 19–38. [Google Scholar] [CrossRef]
- Li, Y.T.; Linster, M.; Mendenhall, I.H.; Su, Y.C.F.; Smith, G.J.D. Avian influenza viruses in humans: Lessons from past outbreaks. Br. Med. Bull. 2019, 132, 81–95. [Google Scholar] [CrossRef]
- Scheibner, D.; Breithaupt, A.; Luttermann, C.; Blaurock, C.; Mettenleiter, T.C.; Abdelwhab, E.M. Genetic determinants for virulence and transmission of the panzootic avian influenza virus H5N8 clade 2.3.4.4 in Pekin ducks. J. Virol. 2022, 96, e0014922. [Google Scholar] [CrossRef]
- Horimoto, T.; Kawaoka, Y. Influenza: Lessons from past pandemics, warnings from current incidents. Nat. Rev. Microbiol. 2005, 3, 591–600. [Google Scholar] [CrossRef]
- Schaffer, J.; Kawaoka, Y.; Bean, W.; Suss, J.; Senne, D.; Webster, R. Origin of the pandemic 1957 H2 influenza A virus and the persistence of its possible progenitors in the avian reservoir. Virology 1993, 194, 781–788. [Google Scholar] [CrossRef] [PubMed]
- Sun, J.; Zheng, T.; Jia, M.; Wang, Y.; Yang, J.; Liu, Y.; Yang, P.; Xie, Y.; Sun, H.; Tong, Q.; et al. Dual receptor-binding, infectivity, and transmissibility of an emerging H2N2 low pathogenicity avian influenza virus. Nat. Commun. 2024, 15, 10012. [Google Scholar] [CrossRef] [PubMed]
- Thomazelli, L.M.; Pinho, J.R.R.; Dorlass, E.G.; Ometto, T.; Meneguin, C.; Paludo, D.; Frias, R.T.; Mancini, P.L.; Monteiro, C.; Aicher, S.M.; et al. Evidence of reassortment of avian influenza A (H2) viruses in Brazilian shorebirds. PLoS ONE 2024, 19, e0300862. [Google Scholar] [CrossRef] [PubMed]
- Havixbeck, J.J.; Rieger, A.M.; Wong, M.E.; Wilkie, M.P.; Barreda, D.R. Evolutionary conservation of divergent pro- inflammatory and homeostatic responses in lamprey phagocytes. PLoS ONE 2014, 9, e86255. [Google Scholar] [CrossRef]
| Monoclonal Antibody/Dyes | Fluorochrome | Manufacturer | Working Volume |
|---|---|---|---|
| Mouse anti-chicken KUL01 | FITC | Southern Biotech | 0.1 µL |
| CellRox Deep Red reagent | AF642 Deep red | Life Technologies | 0.4 µL |
| NucBlue live Reagent | Hoechst 33342 | Life Technologies | 2.5 µL |
| DAF-FM diacetate | FITC | Molecular Probes | 0.1 µL |
| Annexin V | FITC | BD Biosciences | 2.5 µL |
| Propidium iodide | PI | Thermo Fisher Scientific | 0.1 µL |
| Gene | Primer Name | Sequence (5′-3′) |
|---|---|---|
| IFN-γ | IFNG-F | TGTAGCTGACGGTGGACCTA |
| IFNG-R | GCGGCTTTGACTTGTCAGTG | |
| IL-1beta | IL-1beta-F | GCATCAAGGGCTACAAGCTC |
| IL-1beta-R | CAGGCGGTAGAAGATGAAGC | |
| IL-6 | IL6-F | CTCCTCGCCAATCTGAAGTC |
| IL6-R | CCCTCACGGTCTTCTCCATA | |
| IL-8 | IL8-F | CCTCCTCCTGGTTTCAGCTG |
| IL8-R | TGGCGTCAGCTTCACATCTT | |
| IL-10 | IL10-F | CGCTGTCACCGCTTCTTCA |
| IL10-R | TCCCGTTCTCATCCATCTTCTC | |
| IL-12 | IL12-F | TATCCCAAGACCTGGAGCAC |
| IL12-R | GCCCAGTCTTTGGAATCTGA | |
| TGF-beta | TGFbeta- | GATGGACCCGATGAGTATTGGGC |
| TGFbeta-R | GGGACACGTTGAACACGAAGAAG | |
| Β-actin | Bactin-F | CACCACAGCCGAGAGAGAAAT |
| Bactin-R | TGACCATCAGGGAGTTCATAGC |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Rondón-Espinoza, J.; Castro-Sanguinetti, G.; Apaza-Chiara, A.; Gonzalez-Veliz, R.; Callupe-Leyva, A.; Vakharia, V.N.; Icochea, E.; More-Bayona, J. Immunogenicity of a Recombinant Avian Influenza H2 Protein Using an Abdominal Inoculation Model in Chickens. Vaccines 2025, 13, 926. https://doi.org/10.3390/vaccines13090926
Rondón-Espinoza J, Castro-Sanguinetti G, Apaza-Chiara A, Gonzalez-Veliz R, Callupe-Leyva A, Vakharia VN, Icochea E, More-Bayona J. Immunogenicity of a Recombinant Avian Influenza H2 Protein Using an Abdominal Inoculation Model in Chickens. Vaccines. 2025; 13(9):926. https://doi.org/10.3390/vaccines13090926
Chicago/Turabian StyleRondón-Espinoza, Juan, Gina Castro-Sanguinetti, Ana Apaza-Chiara, Rosa Gonzalez-Veliz, Alonso Callupe-Leyva, Vikram N. Vakharia, Eliana Icochea, and Juan More-Bayona. 2025. "Immunogenicity of a Recombinant Avian Influenza H2 Protein Using an Abdominal Inoculation Model in Chickens" Vaccines 13, no. 9: 926. https://doi.org/10.3390/vaccines13090926
APA StyleRondón-Espinoza, J., Castro-Sanguinetti, G., Apaza-Chiara, A., Gonzalez-Veliz, R., Callupe-Leyva, A., Vakharia, V. N., Icochea, E., & More-Bayona, J. (2025). Immunogenicity of a Recombinant Avian Influenza H2 Protein Using an Abdominal Inoculation Model in Chickens. Vaccines, 13(9), 926. https://doi.org/10.3390/vaccines13090926





