ImmuniT Platform for Improved Neoantigen Prediction in Lung Cancer
Abstract
1. Introduction
2. Materials and Methods
2.1. Tissue and Blood Collection
2.2. Tissue Processing and Dissociation
2.3. Cell Culture and Co-Culture Assays
2.4. Whole-Exome and Bulk mRNA Sequencing
2.5. Neoantigen Prediction Pipeline
2.6. Neoantigen Expression Assays
2.7. Tetramer Staining and Flow Cytometry
2.8. In Silico Neoantigen Benchmarking
2.9. Statistical Analysis
3. Results
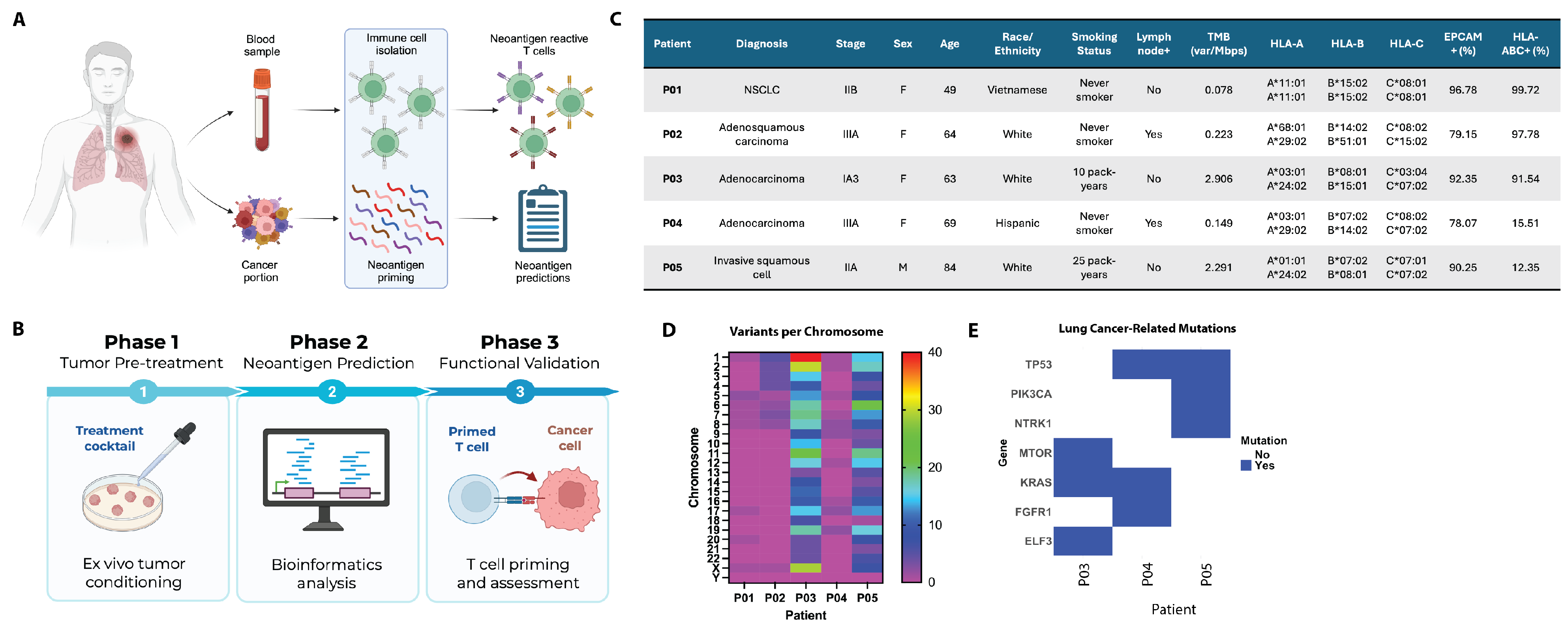
3.1. Patient Characteristics and Tumor Stage
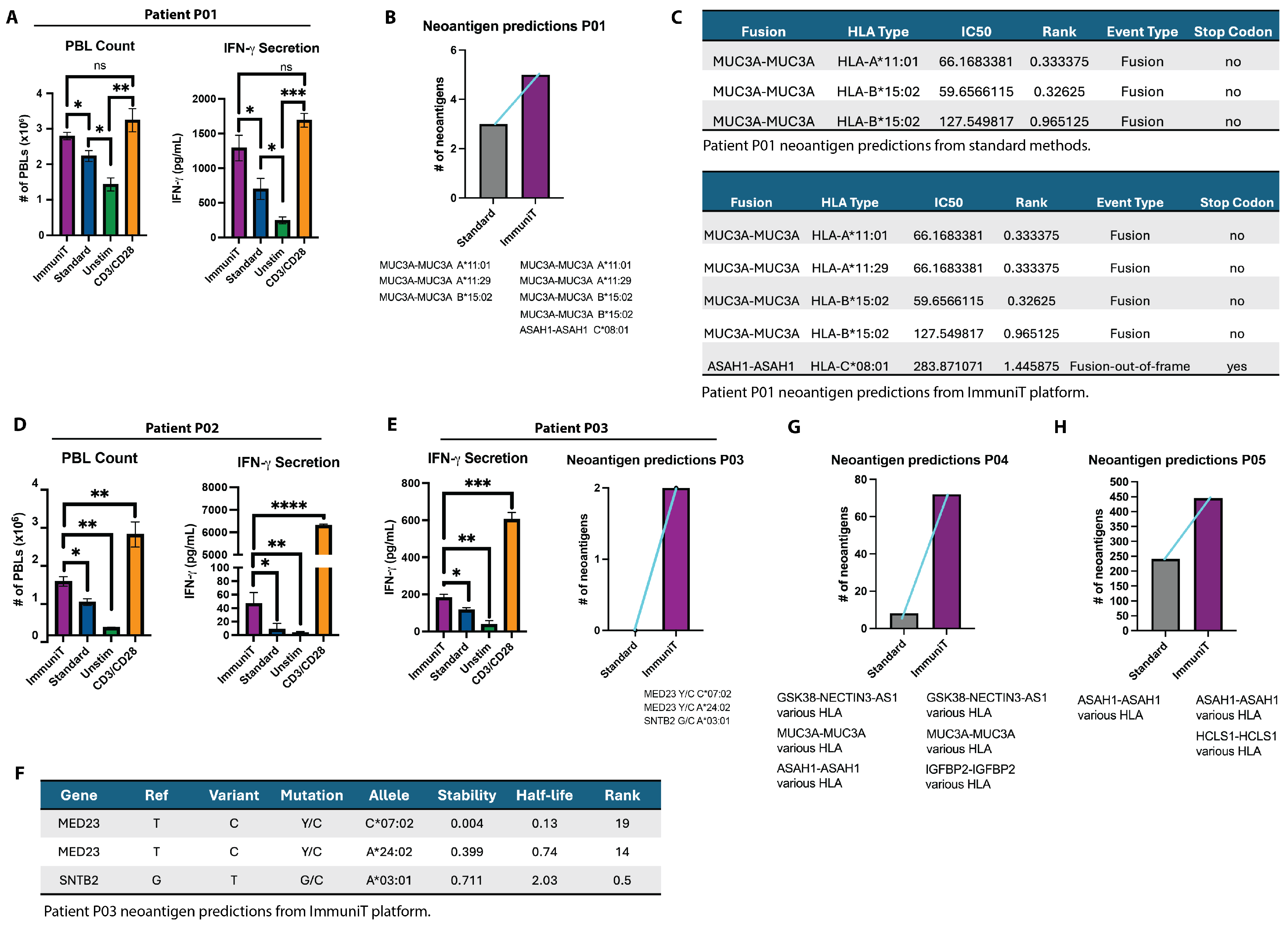
3.2. T Cell Activation and Neoantigen Prediction with the ImmuniT Platform
3.3. Functional Validation of Novel Neoantigens
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Siegel, R.L.; Miller, K.D.; Wagle, N.S.; Jemal, A. Cancer statistics, 2023. CA Cancer J. Clin. 2023, 73, 17–48. [Google Scholar] [CrossRef] [PubMed]
- Reck, M.; Rabe, K.F. Precision Diagnosis and Treatment for Advanced Non–Small-Cell Lung Cancer. N. Engl. J. Med. 2017, 377, 849–861. [Google Scholar] [CrossRef]
- Riley, R.S.; June, C.H.; Langer, R.; Mitchell, M.J. Delivery technologies for cancer immunotherapy. Nat. Rev. Drug Discov. 2019, 18, 175–196. [Google Scholar] [CrossRef]
- Chalmers, Z.R.; Connelly, C.F.; Fabrizio, D.; Gay, L.; Ali, S.M.; Ennis, R.; Schrock, A.; Campbell, B.; Shlien, A.; Chmielecki, J.; et al. Analysis of 100,000 human cancer genomes reveals the landscape of tumor mutational burden. Genome Med. 2017, 9, 34. [Google Scholar] [CrossRef]
- Spranger, S.; Gajewski, T.F. Impact of oncogenic pathways on evasion of antitumour immune responses. Nat. Rev. Cancer 2018, 18, 139–147. [Google Scholar] [CrossRef] [PubMed]
- Hu, Z.; Ott, P.A.; Wu, C.J. Towards personalized, tumour-specific, therapeutic vaccines for cancer. Nat. Rev. Immunol. 2018, 18, 168–182. [Google Scholar] [CrossRef]
- Peng, S.; Zaretsky, J.M.; Ng, A.H.; Chour, W.; Bethune, M.T.; Choi, J.; Hsu, A.; Holman, E.; Ding, X.; Guo, K.; et al. Sensitive Detection and Analysis of Neoantigen-Specific T Cell Populations from Tumors and Blood. Cell Rep. 2019, 28, 2728–2738.e7. [Google Scholar] [CrossRef]
- Ott, P.A.; Hu-Lieskovan, S.; Chmielowski, B.; Govindan, R.; Naing, A.; Bhardwaj, N.; Margolin, K.; Awad, M.M.; Hellmann, M.D.; Lin, J.J.; et al. A Phase Ib Trial of Personalized Neoantigen Therapy Plus Anti-PD-1 in Patients with Advanced Melanoma, Non-small Cell Lung Cancer, or Bladder Cancer. Cell 2020, 183, 347–362.e24. [Google Scholar] [CrossRef] [PubMed]
- Ingels, J.; De Cock, L.; Stevens, D.; Mayer, R.L.; Théry, F.; Sanchez, G.S.; Vermijlen, D.; Weening, K.; De Smet, S.; Lootens, N.; et al. Neoantigen-targeted dendritic cell vaccination in lung cancer patients induces long-lived T cells exhibiting the full differentiation spectrum. Cell Rep. Med. 2024, 5, 101516. [Google Scholar] [CrossRef] [PubMed]
- Yarchoan, M.; Johnson, B.A.; Lutz, E.R.; Laheru, D.A.; Jaffee, E.M. Targeting neoantigens to augment antitumour immunity. Nat. Rev. Cancer 2017, 17, 209–222. [Google Scholar] [CrossRef]
- Rosenthal, R.; Cadieux, E.L.; Salgado, R.; Bakir, M.A.; Moore, D.A.; Hiley, C.T.; Lund, T.; Tanić, M.; Reading, J.L.; Joshi, K.; et al. Neoantigen-directed immune escape in lung cancer evolution. Nature 2019, 567, 479–485. [Google Scholar] [CrossRef] [PubMed]
- Müller, M.; Huber, F.; Arnaud, M.; Kraemer, A.I.; Altimiras, E.R.; Michaux, J.; Taillandier-Coindard, M.; Chiffelle, J.; Murgues, B.; Gehret, T.; et al. Machine learning methods and harmonized datasets improve immunogenic neoantigen prediction. Immunity 2023, 56, 2650–2663.e6. [Google Scholar] [CrossRef] [PubMed]
- Wells, D.K.; van Buuren, M.M.; Dang, K.K.; Hubbard-Lucey, V.M.; Sheehan, K.C.; Campbell, K.M.; Lamb, A.; Ward, J.P.; Sidney, J.; Blazquez, A.B.; et al. Key Parameters of Tumor Epitope Immunogenicity Revealed Through a Consortium Approach Improve Neoantigen Prediction. Cell 2020, 183, 818–834.e13. [Google Scholar] [CrossRef]
- Rieder, D.; Fotakis, G.; Ausserhofer, M.; René, G.; Paster, W.; Trajanoski, Z.; Finotello, F. nextNEOpi: A comprehensive pipeline for computational neoantigen prediction. Bioinformatics 2022, 38, 1131–1132. [Google Scholar] [CrossRef]
- Chen, H.J.; Sun, J.; Huang, Z.; Hou, H.; Arcilla, M.; Rakhilin, N.; Joe, D.J.; Choi, J.; Gadamsetty, P.; Milsom, J.; et al. Comprehensive models of human primary and metastatic colorectal tumors in immunodeficient and immunocompetent mice by chemokine targeting. Nat. Biotechnol. 2015, 33, 656–660. [Google Scholar] [CrossRef]
- Kim, M.; Mun, H.; Sung, C.O.; Cho, E.J.; Jeon, H.J.; Chun, S.M.; Jung, D.J.; Shin, T.H.; Jeong, G.S.; Kim, D.K.; et al. Patient-derived lung cancer organoids as in vitro cancer models for therapeutic screening. Nat. Commun. 2019, 10, 3991. [Google Scholar] [CrossRef]
- Perea, F.; Bernal, M.; Sánchez-Palencia, A.; Carretero, J.; Torres, C.; Bayarri, C.; Gómez-Morales, M.; Garrido, F.; Ruiz-Cabello, F. The absence of HLA class I expression in non-small cell lung cancer correlates with the tumor tissue structure and the pattern of T cell infiltration. Int. J. Cancer 2017, 140, 888–899. [Google Scholar] [CrossRef]
- Zhang, P.; Chen, P.L.; Li, Z.H.; Zhang, A.; Zhang, X.R.; Zhang, Y.J.; Liu, D.; Mao, C. Association of smoking and polygenic risk with the incidence of lung cancer: A prospective cohort study. Br. J. Cancer 2022, 126, 1637–1646. [Google Scholar] [CrossRef]
- Ye, L.; Creaney, J.; Redwood, A.; Robinson, B. The Current Lung Cancer Neoantigen Landscape and Implications for Therapy. J. Thorac. Oncol. 2021, 16, 922–932. [Google Scholar] [CrossRef]
- Pao, W.; Girard, N. New driver mutations in non-small-cell lung cancer. Lancet Oncol. 2011, 12, 175–180. [Google Scholar] [CrossRef] [PubMed]
- Frankell, A.M.; Dietzen, M.; Al Bakir, M.; Lim, E.L.; Karasaki, T.; Ward, S.; Veeriah, S.; Colliver, E.; Huebner, A.; Bunkum, A.; et al. The evolution of lung cancer and impact of subclonal selection in TRACERx. Nature 2023, 616, 525–533. [Google Scholar] [CrossRef]
- Fu, X.; Liu, S.; Cao, D.; Li, C.; Ji, H.; Wang, G. Med23 deficiency reprograms the tumor microenvironment to promote lung tumorigenesis. Br. J. Cancer 2024, 130, 716–727. [Google Scholar] [CrossRef]
- Im, C.N.; Kim, B.M.; Moon, E.Y.; Hong, D.W.; Park, J.W.; Hong, S.H. Characterization of H460R, a Radioresistant Human Lung Cancer Cell Line, and Involvement of Syntrophin Beta 2 (SNTB2) in Radioresistance. Genom. Inform. 2013, 11, 245. [Google Scholar] [CrossRef] [PubMed]
- Guo, C.; Lu, H.; Gao, W.; Wang, L.; Lu, K.; Wu, S.; Pataer, A.; Huang, M.; El-Zein, R.; Lin, T.; et al. Insulin-Like Growth Factor Binding Protein-2 Level Is Increased in Blood of Lung Cancer Patients and Associated with Poor Survival. PLoS ONE 2013, 8, e74973. [Google Scholar] [CrossRef]
- Lai, M.; Realini, N.; La Ferla, M.; Passalacqua, I.; Matteoli, G.; Ganesan, A.; Pistello, M.; Mazzanti, C.M.; Piomelli, D. Complete Acid Ceramidase ablation prevents cancer-initiating cell formation in melanoma cells. Sci. Rep. 2017, 7, 7411. [Google Scholar] [CrossRef]
- Wermke, M.; Holderried, T.A.W.; Luke, J.J.; Morris, V.K.; Alsdorf, W.H.; Wetzko, K.; Andersson, B.S.; Wistuba, I.I.; Parra, E.R.; Hossain, M.B.; et al. First-in-human dose escalation trial to evaluate the clinical safety and efficacy of an anti-MAGEA1 autologous TCR-transgenic T cell therapy in relapsed and refractory solid tumors. J. Immunother. Cancer 2024, 12, e008668. [Google Scholar] [CrossRef]
- Neek, M.; Tucker, J.A.; Kim, T.I.; Molino, N.M.; Nelson, E.L.; Wang, S.W. Co-delivery of human cancer-testis antigens with adjuvant in protein nanoparticles induces higher cell-mediated immune responses. Biomaterials 2018, 156, 194–203. [Google Scholar] [CrossRef] [PubMed]
- Vansteenkiste, J.F.; Cho, B.C.; Vanakesa, T.; De Pas, T.; Zielinski, M.; Kim, M.S.; Jassem, J.; Yoshimura, M.; Dahabreh, J.; Nakayama, H.; et al. Efficacy of the MAGE-A3 cancer immunotherapeutic as adjuvant therapy in patients with resected MAGE-A3-positive non-small-cell lung cancer (MAGRIT): A randomised, double-blind, placebo-controlled, phase 3 trial. Lancet Oncol. 2016, 17, 822–835. [Google Scholar] [CrossRef] [PubMed]
- Altan, M.; Lopes, G.; Hiltermann, T.J.N.; Govindan, R.; Villaruz, L.C.; Calvo, E.; Edelman, M.J.; Furqan, M.; Neal, J.; Felip, E.; et al. Safety and Tolerability of Letetresgene Autoleucel (GSK3377794): Pilot Studies in Patients with Advanced Non–Small Cell Lung Cancer. Clin. Cancer Res. 2025, 31, 529–542. [Google Scholar] [CrossRef]
- Alves, P.M.S.; Viatte, S.; Fagerberg, T.; Michielin, O.; Bricard, G.; Bouzourene, H.; Vuilleumier, H.; Kruger, T.; Givel, J.C.; Lévy, F.; et al. Immunogenicity of the carcinoembryonic antigen derived peptide 694 in HLA-A2 healthy donors and colorectal carcinoma patients. Cancer Immunol. Immunother. 2007, 56, 1795–1805. [Google Scholar] [CrossRef] [PubMed]
- Mehrotra, S.; Britten, C.D.; Chin, S.; Garrett-Mayer, E.; Cloud, C.A.; Li, M.; Scurti, G.; Salem, M.L.; Nelson, M.H.; Thomas, M.B.; et al. Vaccination with poly(IC:LC) and peptide-pulsed autologous dendritic cells in patients with pancreatic cancer. J. Hematol. Oncol. 2017, 10, 82. [Google Scholar] [CrossRef]
- Ramlau, R.; Quoix, E.; Rolski, J.; Pless, M.; Lena, H.; Lévy, E.; Krzakowski, M.; Hess, D.; Tartour, E.; Chenard, M.P.; et al. A Phase II Study of Tg4010 (Mva-Muc1-Il2) in Association with Chemotherapy in Patients with Stage III/IV Non-small Cell Lung Cancer. J. Thorac. Oncol. 2008, 3, 735–744. [Google Scholar] [CrossRef]
- Roulois, D.; Grégoire, M.; Fonteneau, J.F. MUC1-Specific Cytotoxic T Lymphocytes in Cancer Therapy: Induction and Challenge. BioMed Res. Int. 2013, 2013, 871936. [Google Scholar] [CrossRef] [PubMed]
- Rappaport, A.R.; Kyi, C.; Lane, M.; Hart, M.G.; Johnson, M.L.; Henick, B.S.; Liao, C.Y.; Mahipal, A.; Shergill, A.; Spira, A.I.; et al. A shared neoantigen vaccine combined with immune checkpoint blockade for advanced metastatic solid tumors: Phase 1 trial interim results. Nat. Med. 2024, 30, 1013–1022. [Google Scholar] [CrossRef]
- Lu, D.; Chen, Y.; Jiang, M.; Wang, J.; Li, Y.; Ma, K.; Sun, W.; Zheng, X.; Qi, J.; Jin, W.; et al. KRAS G12V neoantigen specific T cell receptor for adoptive T cell therapy against tumors. Nat. Commun. 2023, 14, 6389. [Google Scholar] [CrossRef]
- Ghorani, E.; Reading, J.L.; Henry, J.Y.; de Massy, M.R.; Rosenthal, R.; Turati, V.; Joshi, K.; Furness, A.J.S.; Ben Aissa, A.; Saini, S.K.; et al. The T cell differentiation landscape is shaped by tumour mutations in lung cancer. Nat. Cancer 2020, 1, 546–561. [Google Scholar] [CrossRef]
- Caushi, J.X.; Zhang, J.; Ji, Z.; Vaghasia, A.; Zhang, B.; Hsiue, E.H.C.; Mog, B.J.; Hou, W.; Justesen, S.; Blosser, R.; et al. Transcriptional programs of neoantigen-specific TIL in anti-PD-1-treated lung cancers. Nature 2021, 596, 126–132. [Google Scholar] [CrossRef]
- Klebanoff, C.A.; Gattinoni, L.; Restifo, N.P. Sorting Through Subsets: Which T-Cell Populations Mediate Highly Effective Adoptive Immunotherapy? J. Immunother. 2012, 35, 651–660. [Google Scholar] [CrossRef]
- Tran, E.; Robbins, P.F.; Lu, Y.C.; Prickett, T.D.; Gartner, J.J.; Jia, L.; Pasetto, A.; Zheng, Z.; Ray, S.; Groh, E.M.; et al. T-Cell Transfer Therapy Targeting Mutant KRAS in Cancer. N. Engl. J. Med. 2016, 375, 2255–2262. [Google Scholar] [CrossRef] [PubMed]
- Ding, Z.; Li, Q.; Zhang, R.; Xie, L.; Shu, Y.; Gao, S.; Wang, P.; Su, X.; Qin, Y.; Wang, Y.; et al. Personalized neoantigen pulsed dendritic cell vaccine for advanced lung cancer. Signal Transduct. Target. Ther. 2021, 6, 26. [Google Scholar] [CrossRef] [PubMed]

| Antigen Name | Peptide Sequence | IEDB ID | HLA Allele | Affinity Rank | Affinity Score | Immunogenicity | Affinity (nM) | Binding Level |
|---|---|---|---|---|---|---|---|---|
| CEA | YLSGANLNL | 74915 | A*02:01 | 0.105 | 0.7986 | −0.00073 | 8.84 | Strong |
| KRAS (G12C) | VVGACGVGK | – | A*11:01 | 0.427 | 0.6035 | 0.08441 | 72.99 | Weak |
| KRAS (G12D) | VVGADGVGK | – | A*03:01 | 0.286 | 0.6045 | 0.24736 | 72.17 | Strong |
| KRAS (G12V) | VVGAVGVGK | – | A*11:01 | 0.384 | 0.6140 | 0.17711 | 65.10 | Strong |
| MAGE-A1 | KVLEYVIKV | 34095 | A*02:01 | 0.054 | 0.8371 | 0.11873 | 5.83 | Strong |
| MAGE-A3 | FLWGPRALV | 16970 | A*02:01 | 0.083 | 0.8120 | 0.17046 | 7.65 | Strong |
| MED23 (Y573C) | TYSRLLVCM | – | A*24:02 | 0.426 | 0.0002 | −0.01952 | 332.17 | Strong |
| MUC1 | LLLLTVLTV | 37528 | A*02:01 | 0.289 | 0.7231 | 0.07520 | 20.01 | Strong |
| NY-ESO-1 | SLLMWITQC | 59278 | A*02:01 | 2.553 | 0.4413 | 0.12576 | 422.15 | Weak |
| SNTB2 (G322C) | ATSTAGCSK | – | A*03:01 | 0.599 | 0.5183 | −0.08680 | 183.51 | Strong |
| Survivin | LTLGEFLKL | 237188 | A*02:01 | 4.719 | 4.7190 | 0.10284 | 1344.98 | Weak |
| hTERT | YLFFYRKSV | 2255432 | A*02:01 | 0.180 | 0.7606 | −0.07774 | 13.34 | Strong |
| TP53 (K132N) | TYSPALNNM | – | C*07:02 | 0.066 | 0.5441 | −0.04644 | 138.82 | Strong |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hachey, S.J.; Forsythe, A.G.; Keshava, H.B.; Hughes, C.C.W. ImmuniT Platform for Improved Neoantigen Prediction in Lung Cancer. Vaccines 2025, 13, 921. https://doi.org/10.3390/vaccines13090921
Hachey SJ, Forsythe AG, Keshava HB, Hughes CCW. ImmuniT Platform for Improved Neoantigen Prediction in Lung Cancer. Vaccines. 2025; 13(9):921. https://doi.org/10.3390/vaccines13090921
Chicago/Turabian StyleHachey, Stephanie J., Alexander G. Forsythe, Hari B. Keshava, and Christopher C. W. Hughes. 2025. "ImmuniT Platform for Improved Neoantigen Prediction in Lung Cancer" Vaccines 13, no. 9: 921. https://doi.org/10.3390/vaccines13090921
APA StyleHachey, S. J., Forsythe, A. G., Keshava, H. B., & Hughes, C. C. W. (2025). ImmuniT Platform for Improved Neoantigen Prediction in Lung Cancer. Vaccines, 13(9), 921. https://doi.org/10.3390/vaccines13090921






