Integrating Cellular Immune Biomarkers with Machine Learning to Identify Potential Correlates of Protection for a Trypanosoma cruzi Vaccine
Abstract
1. Introduction
2. Materials and Methods
2.1. Mice and Parasites
2.2. Treatments and Immunization Schedules
2.3. Infection Protocol, Parasitemia, and Survival
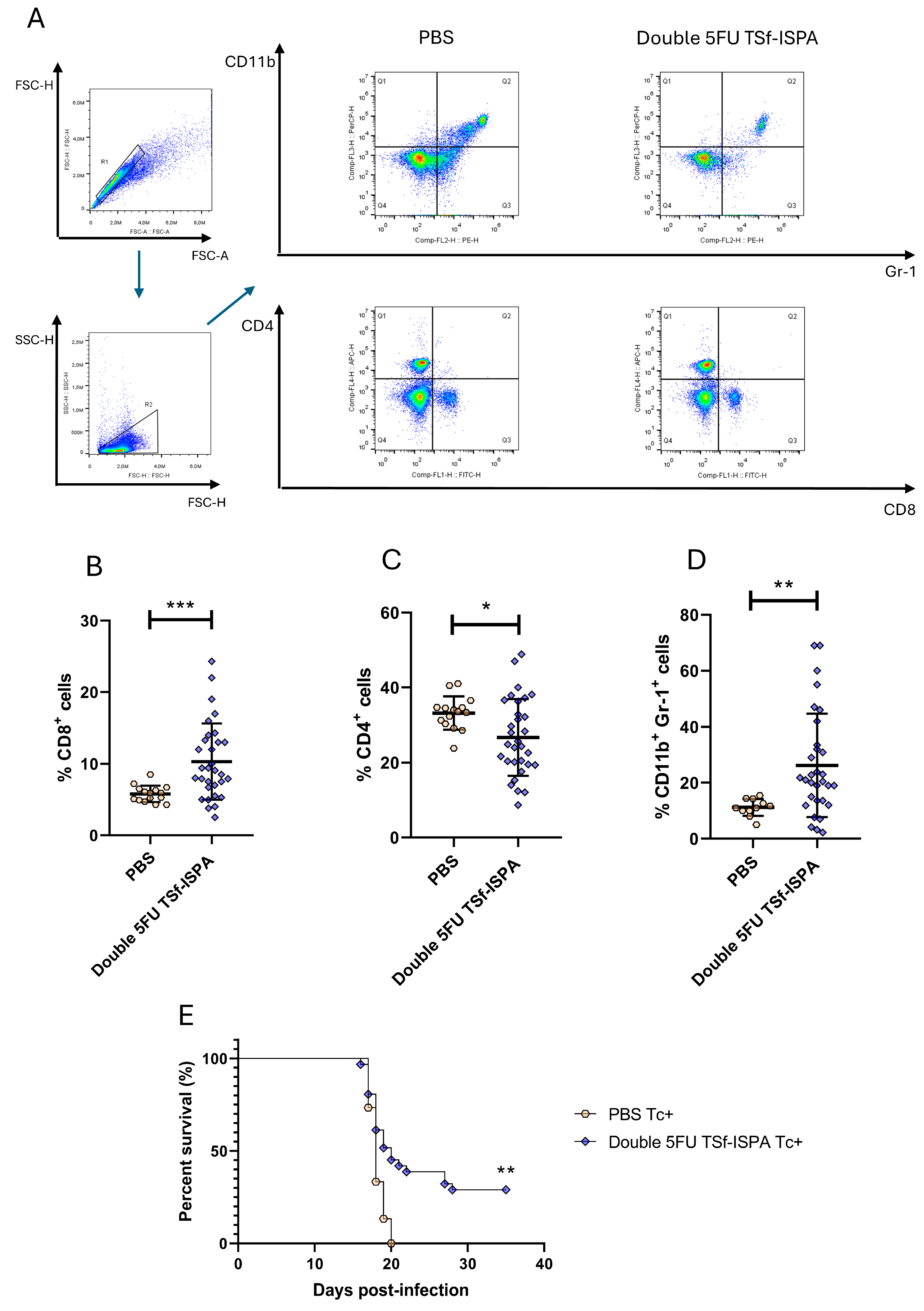
2.4. Flow Cytometry
2.5. Leukocyte Trafficking Assay: Adoptive Transfer and Flow Cytometry Analysis
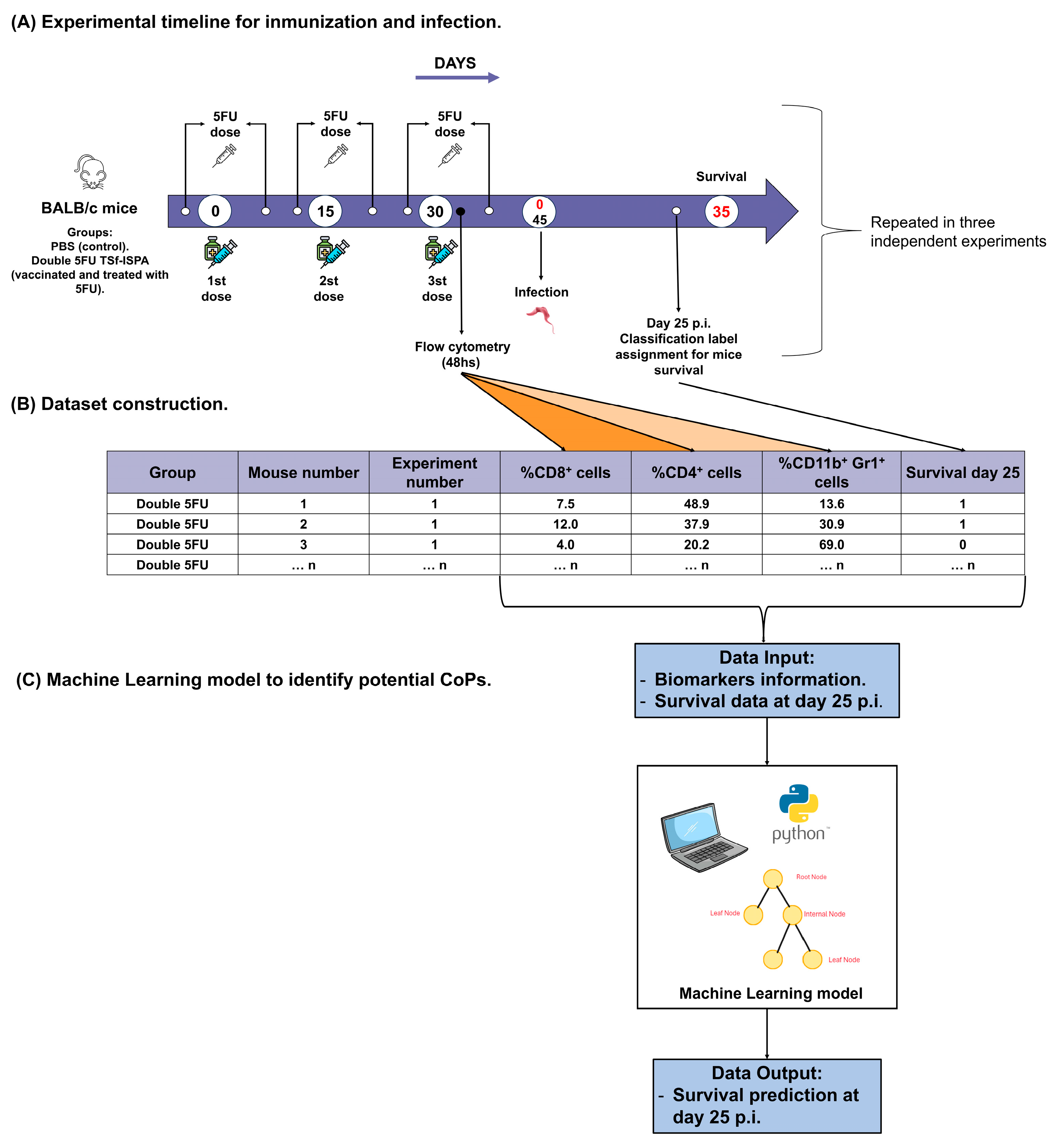
2.6. Dataset Construction
2.7. Machine-Learning Model
2.8. Performance Evaluation Metrics
2.9. Simulation of Input Variability to Evaluate the Potential CoP Decision Logic
2.10. Statistical Analysis
3. Results
3.1. Experimental Assays for Dataset Construction
Criteria Definition for Mice Survival
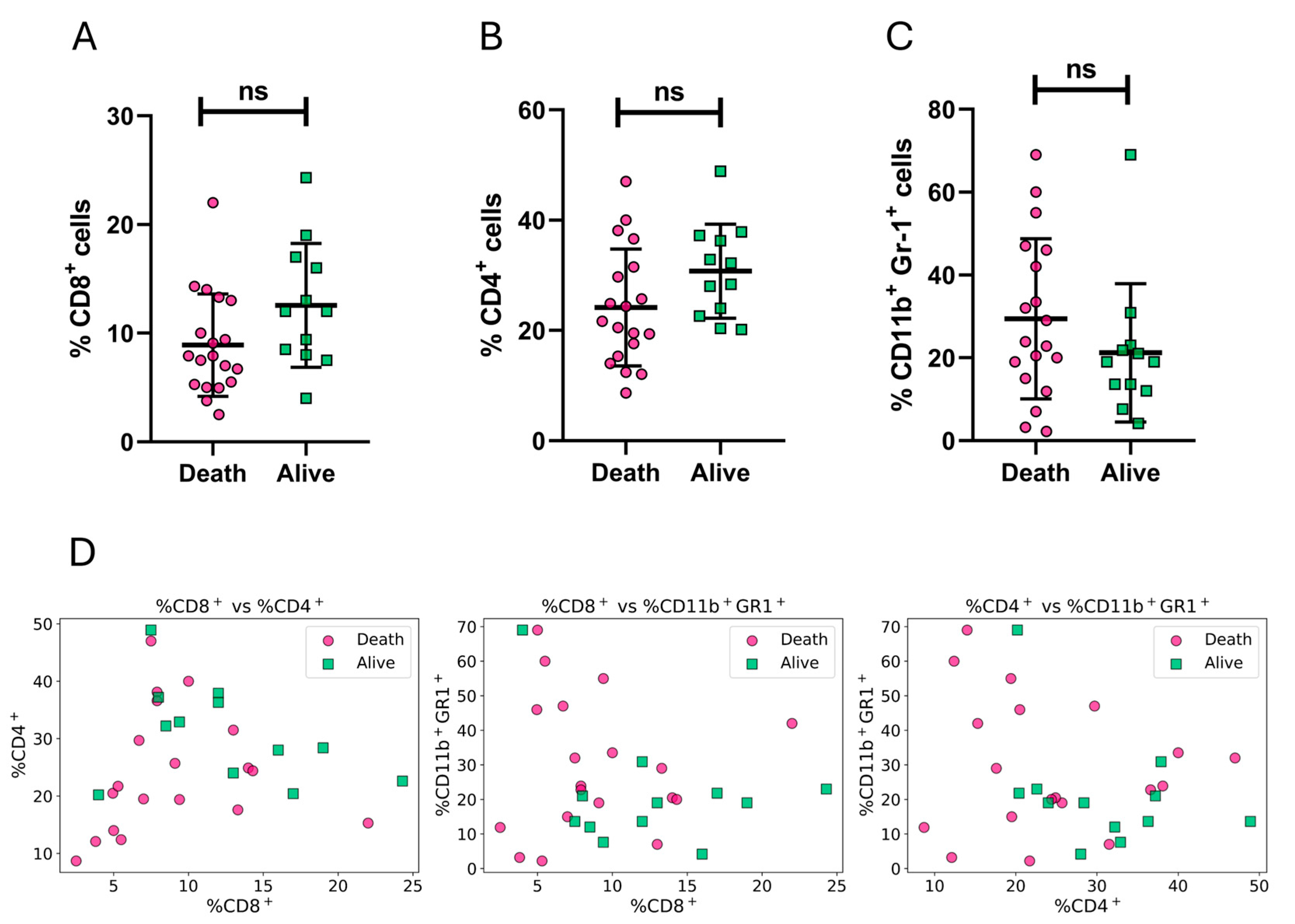
3.2. Exploratory Analysis
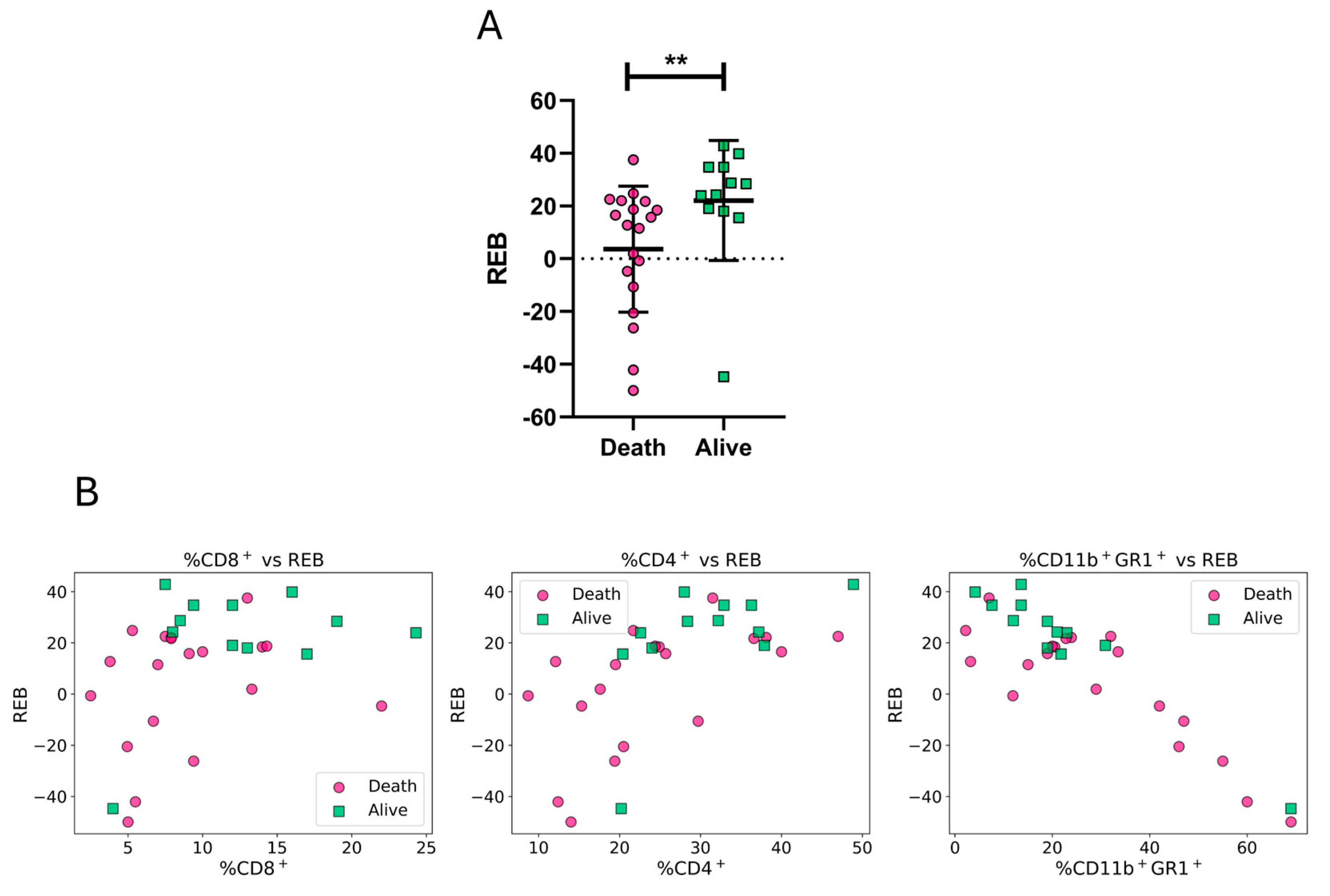
3.3. Rational Biomarker Engineering
3.4. Assessment of individual biomarkers and the rationally engineered biomarker (REB) as potential CoPs
3.5. Systematic Search of Weighted Combinations for the Engineered Biomarker
3.6. Evaluation of the pICoP Decision Logic Under Input Variability
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| T. cruzi | Trypanosoma cruzi |
| CoP | Correlate of Protection |
| mCoP | Mechanistic Correlate of Protection |
| nCoP | Non-Mechanistic Correlate of Protection |
| ML | Machine Learning |
| TSf | Trans-Sialidase Fragment |
| 5FU | 5-Fluorouracil |
| MDSC | Myeloid-Derived Suppressor Cell |
| CD4+ | Cluster of Differentiation 4 Positive |
| CD8+ | Cluster of Differentiation 8 Positive |
| CD11b+ | Cluster of Differentiation 11b Positive |
| Gr-1+ | Granulocyte marker 1 Positive |
| ISPA | Cage-Like Particle Adjuvant |
| PBS | Phosphate-Buffered Saline |
| CFSE | 5,6-Carboxyfluorescein Diacetate Succinimidyl Ester |
| REB | Rational Engineered Biomarker |
| pICoP | Potential Integrative Correlate of Protection |
| AUC-ROC | Area Under the Receiver Operating Characteristic Curve |
| Treg | Regulatory T Cell |
| DC | Dendritic Cell |
References
- Lidani, K.C.F.; Andrade, F.A.; Bavia, L.; Damasceno, F.S.; Beltrame, M.H.; Messias-Reason, I.J.; Sandri, T.L. Chagas Disease: From Discovery to a Worldwide Health Problem. Front. Public Health 2019, 7, 166. [Google Scholar] [CrossRef]
- Teixeira, S.M.; Burle-Caldas, G.D.A.; Castro, J.T.; Gazzinelli, R.T. The Time Has Come for a Vaccine against Chagas Disease. Lancet Reg. Health Am. 2025, 45, 101059. [Google Scholar] [CrossRef]
- Pérez-Molina, J.A.; Molina, I. Chagas Disease. Lancet Lond. Engl. 2018, 391, 82–94. [Google Scholar] [CrossRef]
- Morillo, C.A.; Marin-Neto, J.A.; Avezum, A.; Sosa-Estani, S.; Rassi, A.; Rosas, F.; Villena, E.; Quiroz, R.; Bonilla, R.; Britto, C.; et al. Randomized Trial of Benznidazole for Chronic Chagas’ Cardiomyopathy. N. Engl. J. Med. 2015, 373, 1295–1306. [Google Scholar] [CrossRef]
- Koff, W.C.; Burton, D.R.; Johnson, P.R.; Walker, B.D.; King, C.R.; Nabel, G.J.; Ahmed, R.; Bhan, M.K.; Plotkin, S.A. Accelerating Next-Generation Vaccine Development for Global Disease Prevention. Science 2013, 340, 1232910. [Google Scholar] [CrossRef]
- Amanna, I.J.; Slifka, M.K. Successful Vaccines. Curr. Top. Microbiol. Immunol. 2020, 428, 1–30. [Google Scholar] [CrossRef] [PubMed]
- Dumonteil, E.; Herrera, C. The Case for the Development of a Chagas Disease Vaccine: Why? How? When? Trop. Med. Infect. Dis. 2021, 6, 16. [Google Scholar] [CrossRef]
- Arnal, A.; Villanueva-Lizama, L.; Teh-Poot, C.; Herrera, C.; Dumonteil, E. Extent of Polymorphism and Selection Pressure on the Trypanosoma cruzi Vaccine Candidate Antigen Tc24. Evol. Appl. 2020, 13, 2663–2672. [Google Scholar] [CrossRef] [PubMed]
- Farani, P.S.G.; Jones, K.M.; Poveda, C. Treatments and the Perspectives of Developing a Vaccine for Chagas Disease. Vaccines 2024, 12, 870. [Google Scholar] [CrossRef] [PubMed]
- Goñi, O.; Alcaide, P.; Fresno, M. Immunosuppression during Acute Trypanosoma cruzi Infection: Involvement of Ly6G (Gr1+)CD11b+ Immature Myeloid Suppressor Cells. Int. Immunol. 2002, 14, 1125–1134. [Google Scholar] [CrossRef]
- Nardy, A.F.F.R.; Freire-de-Lima, C.G.; Pérez, A.R.; Morrot, A. Role of Trypanosoma cruzi Trans-Sialidase on the Escape from Host Immune Surveillance. Front. Microbiol. 2016, 7, 348. [Google Scholar] [CrossRef]
- Cardillo, F.; de Pinho, R.T.; Antas, P.R.Z.; Mengel, J. Immunity and Immune Modulation in Trypanosoma cruzi Infection. Pathog. Dis. 2015, 73, ftv082. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Cabrera, G.; Marcipar, I. Vaccines and the Regulatory Arm of the Immune System. An Overview from the Trypanosoma cruzi Infection Model. Vaccine 2019, 37, 3628–3637. [Google Scholar] [CrossRef]
- Cardoso, M.S.; Reis-Cunha, J.L.; Bartholomeu, D.C. Evasion of the Immune Response by Trypanosoma cruzi during Acute Infection. Front. Immunol. 2015, 6, 659. [Google Scholar] [CrossRef]
- Michelin, M.A.; Silva, J.S.; Cunha, F.Q.C. Inducible Cyclooxygenase Released Prostaglandin Mediates Immunosuppression in Acute Phase of Experimental Trypanosoma cruzi Infection. Exp. Parasitol. 2005, 111, 71–79. [Google Scholar] [CrossRef]
- Plotkin, S.A. Vaccines We Need But Don’t Have. Viral Immunol. 2018, 31, 114–116. [Google Scholar] [CrossRef]
- Pinazo, M.J.; Malchiodi, E.; Ioset, J.-R.; Bivona, A.; Gollob, K.J.; Dutra, W.O. Challenges and Advancements in the Development of Vaccines and Therapies against Chagas Disease. Lancet Microbe 2024, 5, 100972. [Google Scholar] [CrossRef]
- Plotkin, S.A.; Gilbert, P.B. Nomenclature for Immune Correlates of Protection after Vaccination. Clin. Infect. Dis. Off. Publ. Infect. Dis. Soc. Am. 2012, 54, 1615–1617. [Google Scholar] [CrossRef]
- Angel, J.; Steele, A.D.; Franco, M.A. Correlates of Protection for Rotavirus Vaccines: Possible Alternative Trial Endpoints, Opportunities, and Challenges. Hum. Vaccines Immunother. 2014, 10, 3659–3671. [Google Scholar] [CrossRef] [PubMed]
- Plotkin, S.A. Updates on Immunologic Correlates of Vaccine-Induced Protection. Vaccine 2020, 38, 2250–2257. [Google Scholar] [CrossRef] [PubMed]
- Samuel, A.L. Some Studies in Machine Learning Using the Game of Checkers. IBM J. Res. Dev. 1959, 3, 210–229. [Google Scholar] [CrossRef]
- Boulesteix, A.-L.; Schmid, M. Machine Learning versus Statistical Modeling. Biom. J. Biom. Z. 2014, 56, 588–593. [Google Scholar] [CrossRef]
- Srivastava, P.; Shukla, R.K.; Sharma, S.; Khanduja, D.; Gupta, R.; Alrasheedi, M.; Singh, G. Fuzzy Methodology Approach for Prioritizing Maintenance 4.0 Attributes. In Proceedings of the 2020 International Conference on Computation, Automation and Knowledge Management (ICCAKM), Dubai, United Arab Emirates, 9–10 January 2020; pp. 308–311. [Google Scholar]
- Bravi, B. Development and Use of Machine Learning Algorithms in Vaccine Target Selection. NPJ Vaccines 2024, 9, 15. [Google Scholar] [CrossRef]
- Javaid, M.; Haleem, A.; Pratap Singh, R.; Suman, R.; Rab, S. Significance of Machine Learning in Healthcare: Features, Pillars and Applications. Int. J. Intell. Netw. 2022, 3, 58–73. [Google Scholar] [CrossRef]
- Breiman, L.; Friedman, J.; Olshen, R.A.; Stone, C.J. Classification and Regression Trees; Chapman and Hall/CRC: New York, NY, USA, 2017; ISBN 978-1-315-13947-0. [Google Scholar]
- Mienye, I.D.; Jere, N. A Survey of Decision Trees: Concepts, Algorithms, and Applications. IEEE Access 2024, 12, 86716–86727. [Google Scholar] [CrossRef]
- Prochetto, E.; Roldán, C.; Bontempi, I.A.; Bertona, D.; Peverengo, L.; Vicco, M.H.; Rodeles, L.M.; Pérez, A.R.; Marcipar, I.S.; Cabrera, G. Trans-Sialidase-Based Vaccine Candidate Protects against Trypanosoma cruzi Infection, Not Only Inducing an Effector Immune Response but Also Affecting Cells with Regulatory/Suppressor Phenotype. Oncotarget 2017, 8, 58003–58020. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Bertona, D.; Pujato, N.; Bontempi, I.; Gonzalez, V.; Cabrera, G.; Gugliotta, L.; Hozbor, D.; Nicastro, A.; Calvinho, L.; Marcipar, I.S. Development and Assessment of a New Cage-like Particle Adjuvant. J. Pharm. Pharmacol. 2017, 69, 1293–1303. [Google Scholar] [CrossRef]
- Gamba, J.C.; Roldán, C.; Prochetto, E.; Lupi, G.; Bontempi, I.; Poncini, C.V.; Vermeulen, M.; Pérez, A.R.; Marcipar, I.; Cabrera, G. Targeting Myeloid-Derived Suppressor Cells to Enhance a Trans-Sialidase-Based Vaccine Against Trypanosoma cruzi. Front. Cell. Infect. Microbiol. 2021, 11, 671104. [Google Scholar] [CrossRef]
- Borgna, E.; Prochetto, E.; Gamba, J.C.; Vermeulen, E.M.; Poncini, C.V.; Cribb, P.; Pérez, A.R.; Marcipar, I.; González, F.B.; Cabrera, G. Control of Myeloid-Derived Suppressor Cell Dynamics Potentiates Vaccine Protection in Multiple Mouse Models of Trypanosoma cruzi Infection. Front. Immunol. 2024, 15, 1484290. [Google Scholar] [CrossRef]
- Vincent, J.; Mignot, G.; Chalmin, F.; Ladoire, S.; Bruchard, M.; Chevriaux, A.; Martin, F.; Apetoh, L.; Rébé, C.; Ghiringhelli, F. 5-Fluorouracil Selectively Kills Tumor-Associated Myeloid-Derived Suppressor Cells Resulting in Enhanced T Cell-Dependent Antitumor Immunity. Cancer Res. 2010, 70, 3052–3061. [Google Scholar] [CrossRef]
- Mathew, A.A.; Zakkariya, Z.T.; Ashokan, A.; Manohar, M.; Keechilat, P.; Nair, S.V.; Koyakutty, M. 5-FU Mediated Depletion of Myeloid Suppressor Cells Enhances T-Cell Infiltration and Anti-Tumor Response in Immunotherapy-Resistant Lung Tumor. Int. Immunopharmacol. 2023, 120, 110129. [Google Scholar] [CrossRef] [PubMed]
- Arocena, A.R.; Onofrio, L.I.; Pellegrini, A.V.; Carrera Silva, A.E.; Paroli, A.; Cano, R.C.; Aoki, M.P.; Gea, S. Myeloid-Derived Suppressor Cells Are Key Players in the Resolution of Inflammation during a Model of Acute Infection. Eur. J. Immunol. 2014, 44, 184–194. [Google Scholar] [CrossRef]
- Poncini, C.V.; González-Cappa, S.M. Dual Role of Monocyte-Derived Dendritic Cells in Trypanosoma cruzi Infection. Eur. J. Immunol. 2017, 47, 1936–1948. [Google Scholar] [CrossRef]
- Borgna, E.; Gamba, J.C.; Prochetto, E.; Marcipar, I.; Cabrera, G. Simple Protocol for Measuring CD11b+ GR-1+ (Ly6C+/Ly6G+) Myeloid Cells from a Minimum Volume of Mouse Peripheral Blood. Methods Cell Biol. 2024, 184, 59–68. [Google Scholar] [CrossRef] [PubMed]
- Powers, D.M.W. Evaluation: From Precision, Recall and F-Measure to ROC, Informedness, Markedness and Correlation. arXiv 2020, arXiv:2010.16061. [Google Scholar] [CrossRef]
- Mandrekar, J.N. Receiver Operating Characteristic Curve in Diagnostic Test Assessment. J. Thorac. Oncol. Off. Publ. Int. Assoc. Study Lung Cancer 2010, 5, 1315–1316. [Google Scholar] [CrossRef]
- Sceneay, J.; Griessinger, C.M.; Hoffmann, S.H.L.; Wen, S.W.; Wong, C.S.F.; Krumeich, S.; Kneilling, M.; Pichler, B.J.; Möller, A. Tracking the Fate of Adoptively Transferred Myeloid-Derived Suppressor Cells in the Primary Breast Tumor Microenvironment. PLoS ONE 2018, 13, e0196040. [Google Scholar] [CrossRef]
- Diao, W.; Jin, F.; Wang, B.; Zhang, C.-Y.; Chen, J.; Zen, K.; Li, L. The Protective Role of Myeloid-Derived Suppressor Cells in Concanavalin A-Induced Hepatic Injury. Protein Cell 2014, 5, 714–724. [Google Scholar] [CrossRef]
- Acosta Rodríguez, E.V.; Araujo Furlan, C.L.; Fiocca Vernengo, F.; Montes, C.L.; Gruppi, A. Understanding CD8+ T Cell Immunity to Trypanosoma cruzi and How to Improve It. Trends Parasitol. 2019, 35, 899–917. [Google Scholar] [CrossRef]
- Tarleton, R.L. CD8+ T Cells in Trypanosoma cruzi Infection. Semin. Immunopathol. 2015, 37, 233–238. [Google Scholar] [CrossRef]
- Arce-Fonseca, M.; Rios-Castro, M.; Carrillo-Sánchez, S.d.C.; Martínez-Cruz, M.; Rodríguez-Morales, O. Prophylactic and Therapeutic DNA Vaccines against Chagas Disease. Parasit. Vectors 2015, 8, 121. [Google Scholar] [CrossRef]
- da Costa, K.M.; Marques da Fonseca, L.; Dos Reis, J.S.; Santos, M.A.R.D.C.; Previato, J.O.; Mendonça-Previato, L.; Freire-de-Lima, L. Trypanosoma cruzi Trans-Sialidase as a Potential Vaccine Target Against Chagas Disease. Front. Cell. Infect. Microbiol. 2021, 11, 768450. [Google Scholar] [CrossRef]
- Borgna, E.; Prochetto, E.; Gamba, J.C.; Marcipar, I.; Cabrera, G. Role of Myeloid-Derived Suppressor Cells during Trypanosoma cruzi Infection. Int. Rev. Cell Mol. Biol. 2023, 375, 117–163. [Google Scholar] [CrossRef] [PubMed]
- Flippe, L.; Bézie, S.; Anegon, I.; Guillonneau, C. Future Prospects for CD8+ Regulatory T Cells in Immune Tolerance. Immunol. Rev. 2019, 292, 209–224. [Google Scholar] [CrossRef] [PubMed]
- Mercado, N.B.; Zahn, R.; Wegmann, F.; Loos, C.; Chandrashekar, A.; Yu, J.; Liu, J.; Peter, L.; McMahan, K.; Tostanoski, L.H.; et al. Single-Shot Ad26 Vaccine Protects against SARS-CoV-2 in Rhesus Macaques. Nature 2020, 586, 583–588. [Google Scholar] [CrossRef] [PubMed]
- Slyke, G.V.; Ehrbar, D.J.; Doering, J.; Yates, J.L.; Vitetta, E.S.; Donini, O.; Mantis, N.J. Endpoint and Epitope-Specific Antibody Responses as Correlates of Vaccine-Mediated Protection of Mice against Ricin Toxin. Vaccine 2020, 38, 6721–6729. [Google Scholar] [CrossRef]
- McIlwain, D.R.; Chen, H.; Rahil, Z.; Bidoki, N.H.; Jiang, S.; Bjornson, Z.; Kolhatkar, N.S.; Martinez, C.J.; Gaudillière, B.; Hedou, J.; et al. Human Influenza Virus Challenge Identifies Cellular Correlates of Protection for Oral Vaccination. Cell Host Microbe 2021, 29, 1828–1837.e5. [Google Scholar] [CrossRef]
- Sanchez Alberti, A.; Bivona, A.E.; Matos, M.N.; Cerny, N.; Schulze, K.; Weißmann, S.; Ebensen, T.; González, G.; Morales, C.; Cardoso, A.C.; et al. Mucosal Heterologous Prime/Boost Vaccination Induces Polyfunctional Systemic Immunity, Improving Protection Against Trypanosoma cruzi. Front. Immunol. 2020, 11, 128. [Google Scholar] [CrossRef]
- Panchanathan, V.; Chaudhri, G.; Karupiah, G. Correlates of Protective Immunity in Poxvirus Infection: Where Does Antibody Stand? Immunol. Cell Biol. 2008, 86, 80–86. [Google Scholar] [CrossRef]
- Plotkin, S.A. Correlates of Protection Induced by Vaccination. Clin. Vaccine Immunol. CVI 2010, 17, 1055–1065. [Google Scholar] [CrossRef]
- Darrah, P.A.; Patel, D.T.; De Luca, P.M.; Lindsay, R.W.B.; Davey, D.F.; Flynn, B.J.; Hoff, S.T.; Andersen, P.; Reed, S.G.; Morris, S.L.; et al. Multifunctional TH1 Cells Define a Correlate of Vaccine-Mediated Protection against Leishmania Major. Nat. Med. 2007, 13, 843–850. [Google Scholar] [CrossRef]
- Koch, T.; Mellinghoff, S.C.; Shamsrizi, P.; Addo, M.M.; Dahlke, C. Correlates of Vaccine-Induced Protection against SARS-CoV-2. Vaccines 2021, 9, 238. [Google Scholar] [CrossRef]
- Tomaras, G.D.; Plotkin, S.A. Complex Immune Correlates of Protection in HIV-1 Vaccine Efficacy Trials. Immunol. Rev. 2017, 275, 245–261. [Google Scholar] [CrossRef]
- Prochetto, E.; Borgna, E.; Jiménez-Cortegana, C.; Sánchez-Margalet, V.; Cabrera, G. Myeloid-Derived Suppressor Cells and Vaccination against Pathogens. Front. Cell. Infect. Microbiol. 2022, 12, 1003781. [Google Scholar] [CrossRef]
- Ndure, J.; Flanagan, K.L. Targeting Regulatory T Cells to Improve Vaccine Immunogenicity in Early Life. Front. Microbiol. 2014, 5, 477. [Google Scholar] [CrossRef]
- Batista-Duharte, A.; Téllez-Martínez, D.; Fuentes, D.L.P.; Carlos, I.Z. Molecular Adjuvants That Modulate Regulatory T Cell Function in Vaccination: A Critical Appraisal. Pharmacol. Res. 2018, 129, 237–250. [Google Scholar] [CrossRef] [PubMed]
- Stepkowski, S.; Bekbolsynov, D.; Oenick, J.; Brar, S.; Mierzejewska, B.; Rees, M.A.; Ekwenna, O. The Major Role of T Regulatory Cells in the Efficiency of Vaccination in General and Immunocompromised Populations: A Review. Vaccines 2024, 12, 992. [Google Scholar] [CrossRef]
- Bayry, J. Regulatory T Cells as Adjuvant Target for Enhancing the Viral Disease Vaccine Efficacy. Virusdisease 2014, 25, 18–25. [Google Scholar] [CrossRef]
- Lin, P.-H.; Wong, W.-I.; Wang, Y.-L.; Hsieh, M.-P.; Lu, C.-W.; Liang, C.-Y.; Jui, S.-H.; Wu, F.-Y.; Chen, P.-J.; Yang, H.-C. Vaccine-Induced Antigen-Specific Regulatory T Cells Attenuate the Antiviral Immunity against Acute Influenza Virus Infection. Mucosal Immunol. 2018, 11, 1239–1253. [Google Scholar] [CrossRef] [PubMed]
- Jaron, B.; Maranghi, E.; Leclerc, C.; Majlessi, L. Effect of Attenuation of Treg during BCG Immunization on Anti-Mycobacterial Th1 Responses and Protection against Mycobacterium Tuberculosis. PLoS ONE 2008, 3, e2833. [Google Scholar] [CrossRef] [PubMed]
- Felonato, M.; Pina, A.; de Araujo, E.F.; Loures, F.V.; Bazan, S.B.; Feriotti, C.; Calich, V.L.G. Anti-CD25 Treatment Depletes Treg Cells and Decreases Disease Severity in Susceptible and Resistant Mice Infected with Paracoccidioides Brasiliensis. PLoS ONE 2012, 7, e51071. [Google Scholar] [CrossRef]
- Galdino, N.A.L.; Loures, F.V.; de Araújo, E.F.; da Costa, T.A.; Preite, N.W.; Calich, V.L.G. Depletion of Regulatory T Cells in Ongoing Paracoccidioidomycosis Rescues Protective Th1/Th17 Immunity and Prevents Fatal Disease Outcome. Sci. Rep. 2018, 8, 16544. [Google Scholar] [CrossRef]
- Batista-Duharte, A.; Sendra, L.; Herrero, M.J.; Portuondo, D.L.; Téllez-Martínez, D.; Olivera, G.; Fernández-Delgado, M.; Javega, B.; Herrera, G.; Martínez, A.; et al. Foxp3 Silencing with Antisense Oligonucleotide Improves Immunogenicity of an Adjuvanted Recombinant Vaccine against Sporothrix Schenckii. Int. J. Mol. Sci. 2021, 22, 3470. [Google Scholar] [CrossRef]
- Batista-Duharte, A.; Téllez-Martínez, D.; Portuondo, D.L.; Carlos, I.Z. Selective Depletion of Regulatory T Cells Enhances the Immunogenicity of a Recombinant-Based Vaccine against Sporothrix Spp. Front. Cell. Infect. Microbiol. 2022, 12, 1084526. [Google Scholar] [CrossRef]
- Espinoza Mora, M.D.R.; Steeg, C.; Tartz, S.; Heussler, V.; Sparwasser, T.; Link, A.; Fleischer, B.; Jacobs, T. Depletion of Regulatory T Cells Augments a Vaccine-Induced T Effector Cell Response against the Liver-Stage of Malaria but Fails to Increase Memory. PLoS ONE 2014, 9, e104627. [Google Scholar] [CrossRef] [PubMed]
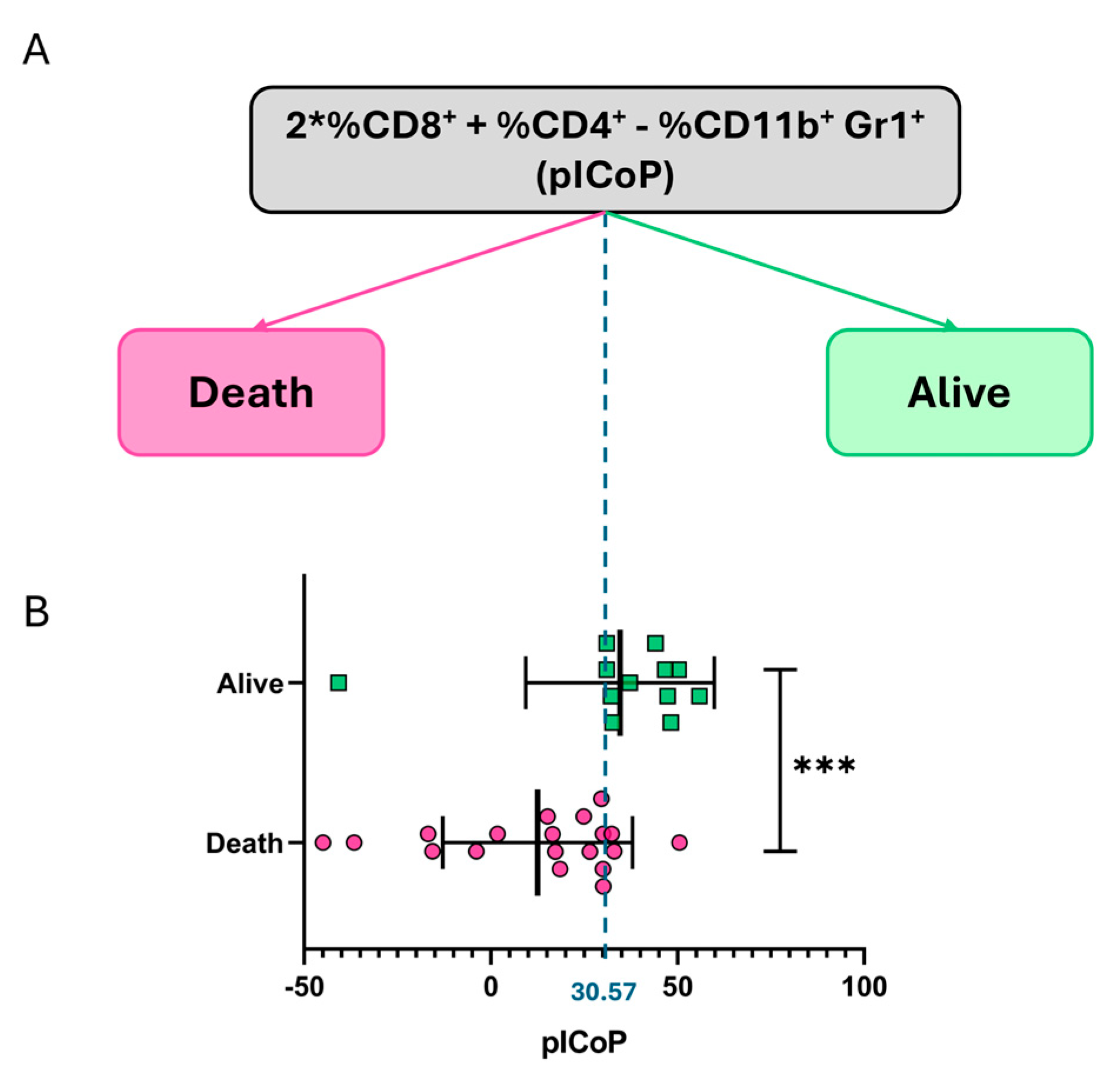
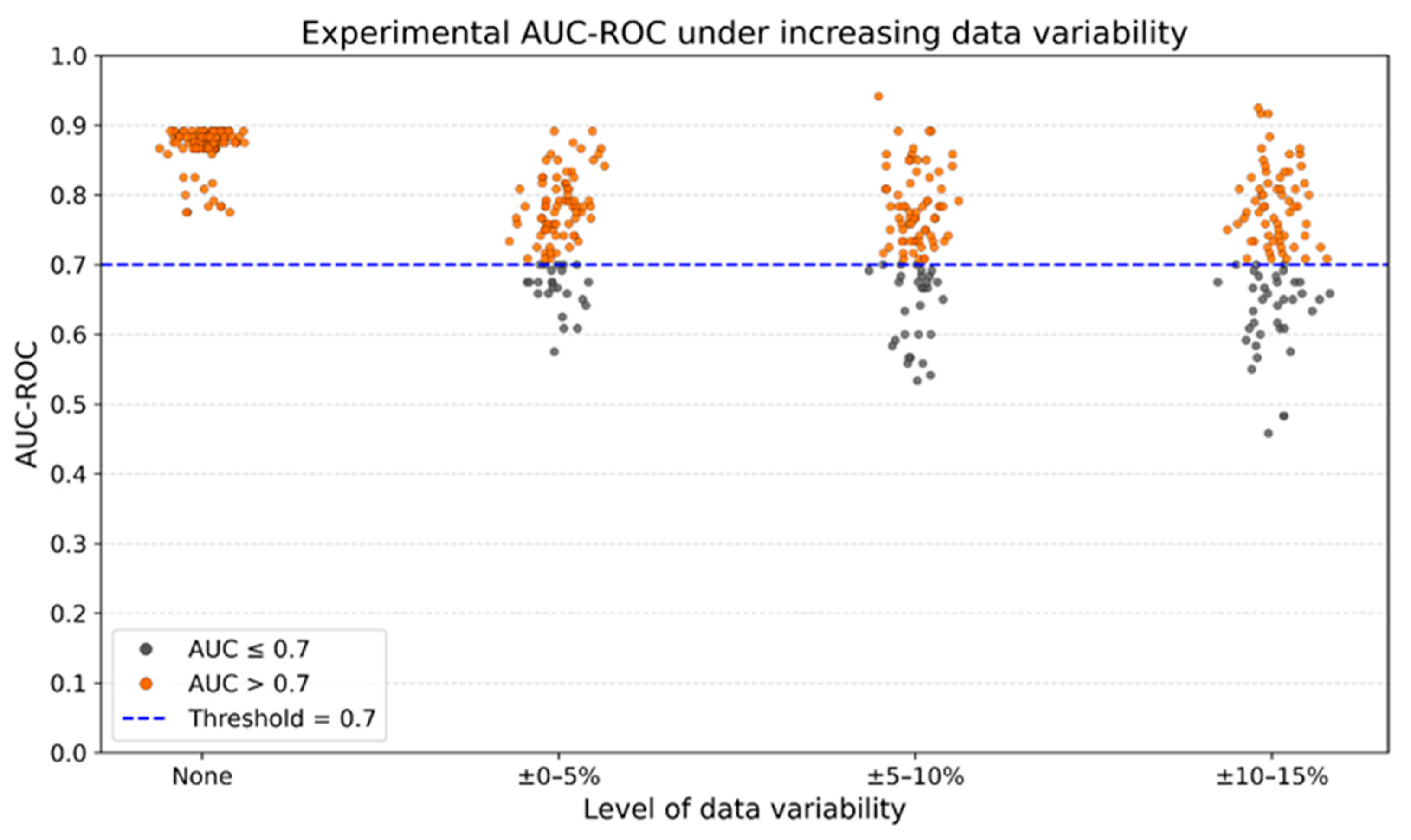
| Potential CoP | F1-Score of Alive Mice | Accuracy | AUC-ROC |
|---|---|---|---|
| 0.47 [95% CI: 0.46–0.49] | 0.47 [95% CI: 0.46–0.49] | 0.58 [95% CI: 0.57–0.59] | |
| 0.65 [95% CI: 0.64–0.66] | 0.61 [95% CI: 0.61–0.62] | 0.68 [95% CI: 0.67–0.69] | |
| 0.50 [95% CI: 0.49–0.51] | 0.50 [95% CI: 0.48–0.51] | 0.53 [95% CI: 0.52–0.55] | |
(REB) | 0.61 [95% CI: 0.59–0.63] | 0.72 [95% CI: 0.71–0.73] | 0.70 [95% CI: 0.69–0.71] |
| Potential CoP | F1-Score of Alive Mice | Accuracy | AUC-ROC |
|---|---|---|---|
(REB) | 0.61 [95% CI: 0.59–0.63] | 0.72 [95% CI: 0.71–0.73] | 0.70 [95% CI: 0.69–0.71] |
(pICoP) | 0.83 [95% CI: 0.82–0.84] | 0.86 [95% CI: 0.86–0.87] | 0.87 [95% CI: 0.86–0.87] |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gamba, J.C.; Borgna, E.; Prochetto, E.; Pérez, A.R.; Batista-Duharte, A.; Marcipar, I.; Gerard, M.; Cabrera, G. Integrating Cellular Immune Biomarkers with Machine Learning to Identify Potential Correlates of Protection for a Trypanosoma cruzi Vaccine. Vaccines 2025, 13, 915. https://doi.org/10.3390/vaccines13090915
Gamba JC, Borgna E, Prochetto E, Pérez AR, Batista-Duharte A, Marcipar I, Gerard M, Cabrera G. Integrating Cellular Immune Biomarkers with Machine Learning to Identify Potential Correlates of Protection for a Trypanosoma cruzi Vaccine. Vaccines. 2025; 13(9):915. https://doi.org/10.3390/vaccines13090915
Chicago/Turabian StyleGamba, Juan Cruz, Eliana Borgna, Estefanía Prochetto, Ana Rosa Pérez, Alexander Batista-Duharte, Iván Marcipar, Matías Gerard, and Gabriel Cabrera. 2025. "Integrating Cellular Immune Biomarkers with Machine Learning to Identify Potential Correlates of Protection for a Trypanosoma cruzi Vaccine" Vaccines 13, no. 9: 915. https://doi.org/10.3390/vaccines13090915
APA StyleGamba, J. C., Borgna, E., Prochetto, E., Pérez, A. R., Batista-Duharte, A., Marcipar, I., Gerard, M., & Cabrera, G. (2025). Integrating Cellular Immune Biomarkers with Machine Learning to Identify Potential Correlates of Protection for a Trypanosoma cruzi Vaccine. Vaccines, 13(9), 915. https://doi.org/10.3390/vaccines13090915








