Anti-HERV-K Drugs and Vaccines, Possible Therapies against Tumors
Abstract
1. Introduction
2. Evidence of Immunogenicity against HERV-K Genes and Viral Particles in Patients with Tumors
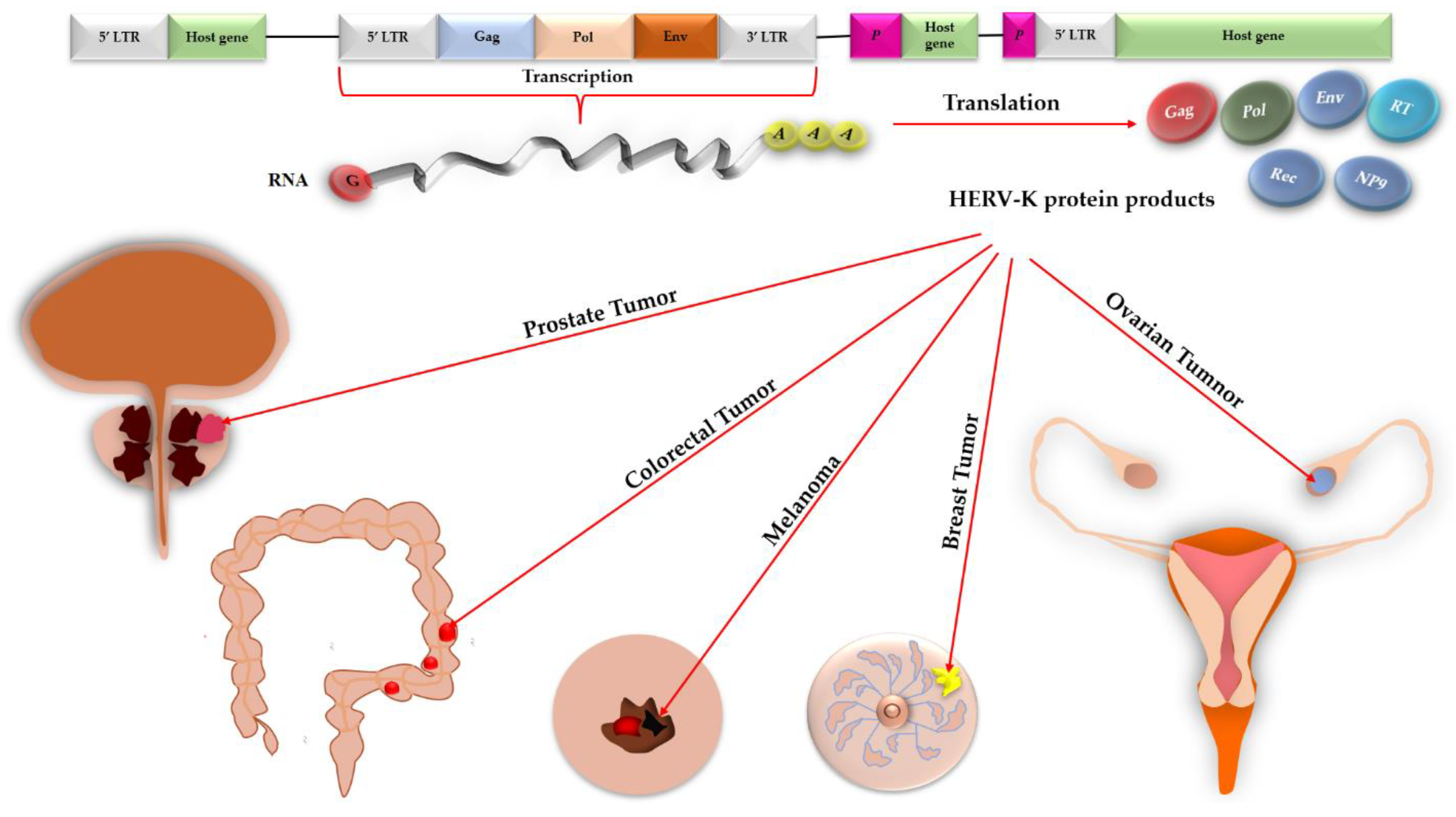
2.1. HERV-K and Melanoma
2.2. HERV-K and Teratocarcinoma
2.3. HERV-K and Osteosarcoma
2.4. HERV-K and Colorectal Tumors
2.5. HERV-K and Breast Tumors
2.6. HERV-K and Ovarian Cancer
2.7. HERV-K and Prostate Cancer
2.8. HERV-K and Atypical Teratoid Rhabdoid Tumors (AT/RT)
2.9. HERV-K and non-Hodgkin Lymphoma (NHL)
3. Therapeutic Strategies against Possible HERV-K-Induced Tumors
3.1. HERV-K Gag-Based Vaccines
3.2. Antibody-Based Therapies
3.2.1. Active Immunization against HERV-K as a Therapy against Solid Tumors
3.2.2. Anti-HERV-K Env Protein Antibodies
3.2.3. Anti-HERV-K Antibodies as The Future Breast Cancer Treatment?
3.3. HERV-K Inhibitors
3.3.1. HERV-K Env Inhibitors
3.3.2. HERV-K Reverse Transcriptase Inhibitors
3.4. Clustered Regulatory Interspaced Short Palindromic Repeats (CRISPR)/Cas9 Technology
3.5. Anti-HERV-K Expression
3.5.1. Anti-HIV-1 Drugs
3.5.2. Anti-Coronavirus Drugs
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Venkatesan, A.; Johnson, R.T. Infections and multiple sclerosis. In Multiple Sclerosis and Related Disorders; Goodin, D.S., Ed.; Elsevier: Amsterdam, The Netherlands, 2014; Volume 122, pp. 151–171. ISBN 0072-9752. [Google Scholar]
- Luzuriaga, K. Introduction to retroviridae. In Principles and Practice of Pediatric Infectious Diseases; Long, S.S., Ed.; Elsevier: London, UK, 2012; p. 1164. ISBN 978-1-4377-2702-9. [Google Scholar]
- Grandi, N.; Tramontano, E. Human endogenous retroviruses are ancient acquired elements still shaping innate immune responses. Front. Immunol. 2018, 9, 2039. [Google Scholar] [CrossRef] [PubMed]
- Nelson, P.N.; Carnegie, P.R.; Martin, J.; Davari Ejtehadi, H.; Hooley, P.; Roden, D.; Rowland-Jones, S.; Warren, P.; Astley, J.; Murray, P.G. Demystified. Human endogenous retroviruses. Mol. Pathol. 2003, 56, 11–18. [Google Scholar] [CrossRef] [PubMed]
- Bannert, N.; Kurth, R. The evolutionary dynamics of human endogenous retroviral families. Annu. Rev. Genomics Hum. Genet. 2006, 7, 149–173. [Google Scholar] [CrossRef]
- Pisano, M.P.; Grandi, N.; Cadeddu, M.; Blomberg, J.; Tramontano, E. Comprehensive characterization of the human endogenous retrovirus HERV-K(HML-6) Group: Overview of structure, phylogeny, and contribution to the human genome. J. Virol. 2019, 93, e00110-19. [Google Scholar] [CrossRef]
- Alcazer, V.; Bonaventura, P.; Depil, S. Human endogenous retroviruses (HERVs): Shaping the innate immune response in cancers. Cancers 2020, 12, 610. [Google Scholar] [CrossRef]
- Smit, A.F.A. Interspersed repeats and other mementos of transposable elements in mammalian genomes. Curr. Opin. Genet. Dev. 1999, 9, 657–663. [Google Scholar] [CrossRef]
- Larsson; Andersson beneficial role of human endogenous retroviruses: Facts and hypotheses. Scand. J. Immunol. 1998, 48, 329–338. [CrossRef]
- Nelson, P.N.; Hooley, P.; Roden, D.; Davari Ejtehadi, H.; Rylance, P.; Warren, P.; Martin, J.; Murray, P.G. Human endogenous retroviruses: Transposable elements with potential? Clin. Exp. Immunol. 2004, 138, 1–9. [Google Scholar] [CrossRef]
- Blomberg, J.; Ushameckis, D.; Jern, P. Evolutionary aspects of human endogenous retroviral sequences (HERVs) and disease. In Madame Curie Bioscience Database; Landes Bioscience: Austin, TX, USA, 2008. [Google Scholar]
- Gröger, V.; Cynis, H. Human endogenous retroviruses and their putative role in the development of autoimmune disorders such as multiple sclerosis. Front. Microbiol. 2018, 9, 265. [Google Scholar] [CrossRef]
- Boller, K.; König, H.; Sauter, M.; Mueller-Lantzsch, N.; Löwer, R.; Löwer, J.; Kurth, R. Evidence that HERV-K is the endogenous retrovirus sequence that codes for the human teratocarcinoma-derived retrovirus HTDV. Virology 1993, 196, 349–353. [Google Scholar] [CrossRef] [PubMed]
- Löwer, R.; Boller, K.; Hasenmaier, B.; Korbmacher, C.; Müller-Lantzsch, N.; Löwer, J.; Kurth, R. Identification of human endogenous retroviruses with complex MRNA expression and particle formation. Proc. Natl. Acad. Sci. USA 1993, 90, 4480–4484. [Google Scholar] [CrossRef] [PubMed]
- Nelson, P.N. Retroviruses in rheumatic diseases. Ann. Rheum. Dis. 1995, 54, 441–442. [Google Scholar] [CrossRef]
- Nelson, P.N.; Lever, A.M.L.; Smith, S.; Pitman, R.; Murray, P.; Perera, S.A.; Westwood, O.M.R.; Hay, F.C.; Ejtehadi, H.D.; Booth, J.C. Molecular investigations implicate human endogenous retroviruses as mediators of anti-retroviral antibodies in autoimmune rheumatic disease. Immunol. Investig. 1999, 28, 277–289. [Google Scholar] [CrossRef] [PubMed]
- Wilkinson, D.; Mager, D.; Leong, J. Endogenous human retroviruses. In The Retroviridae; Levy, J.A., Ed.; Plenum Press: New York, NY, USA, 1994; pp. 465–535. [Google Scholar]
- Subramanian, R.P.; Wildschutte, J.H.; Russo, C.; Coffin, J.M. Identification, characterization, and comparative genomic distribution of the HERV-K (HML-2) group of human endogenous retroviruses. Retrovirology 2011, 8, 90. [Google Scholar] [CrossRef] [PubMed]
- Perl, A.; Rosenblatt, J.D.; Chen, I.S.Y.; DiVincenzo, J.P.; Bever, R.; Poiesz, J.; Abraham, G.N. Detection and cloning of new HTLV-related endogenous sequences in man. Nucleic Acids Res. 1989, 17, 6841–6854. [Google Scholar] [CrossRef]
- Löwer, R.; Löwer, J.; Kurth, R. The viruses in all of us: Characteristics and biological significance of human endogenous retrovirus sequences. Proc. Natl. Acad. Sci. USA 1996, 93, 5177–5184. [Google Scholar] [CrossRef]
- Downey, R.F.; Sullivan, F.J.; Wang-Johanning, F.; Ambs, S.; Giles, F.J.; Glynn, S.A. Human endogenous retrovirus K and cancer: Innocent bystander or tumorigenic accomplice? Int. J. Cancer 2015, 137, 1249–1257. [Google Scholar] [CrossRef]
- Chan, S.M.; Sapir, T.; Park, S.-S.; Rual, J.-F.; Contreras-Galindo, R.; Reiner, O.; Markovitz, D.M. The HERV-K accessory protein Np9 controls viability and migration of teratocarcinoma cells. PLoS ONE 2019, 14, e0212970. [Google Scholar] [CrossRef]
- Ellinghaus, D.; Kurtz, S.; Willhoeft, U. LTRharvest, an efficient and flexible software for de novo detection of LTR retrotransposons. BMC Bioinform. 2008, 9, 18. [Google Scholar] [CrossRef]
- Guffanti, G.; Bartlett, A.; DeCrescenzo, P.; Macciardi, F.; Hunter, R. Transposable elements. In Behavioral Neurogenomics; Binder, E.B., Klengel, T., Eds.; Springer International Publishing: Cham, Switzerland, 2019; pp. 221–246. ISBN 978-3-030-31265-7. [Google Scholar]
- Sarah, P.; Guy, O.; François, M. A retroviral promoter and a cellular enhancer define a bipartite element which controls Env ERVWE1 placental expression. J. Virol. 2004, 78, 12157–12168. [Google Scholar] [CrossRef]
- Seifarth, W.; Frank, O.; Zeilfelder, U.; Spiess, B.; Greenwood, A.D.; Hehlmann, R.; Leib-Mösch, C. Comprehensive analysis of human endogenous retrovirus transcriptional activity in human tissues with a retrovirus-specific microarray. J. Virol. 2005, 79, 341–352. [Google Scholar] [CrossRef]
- van de Lagemaat, L.N.; Landry, J.-R.; Mager, D.L.; Medstrand, P. Transposable elements in mammals promote regulatory variation and diversification of genes with specialized functions. Trends Genet. 2003, 19, 530–536. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.; Xie, G.; Singh, M.; Ghanbarian, A.T.; Raskó, T.; Szvetnik, A.; Cai, H.; Besser, D.; Prigione, A.; Fuchs, N.V.; et al. Primate-specific endogenous retrovirus-driven transcription defines naive-like stem cells. Nature 2014, 516, 405–409. [Google Scholar] [CrossRef]
- Andersson, A.C.; Svensson, A.C.; Rolny, C.; Andersson, G.; Larsson, E. Expression of human endogenous retrovirus ERV3 (HERV-R) mRNA in normal and neoplastic tissues. Int. J. Oncol. 1998, 12, 309–322. [Google Scholar] [CrossRef]
- Sauter, M.; Schommer, S.; Kremmer, E.; Remberger, K.; Dölken, G.; Lemm, I.; Buck, M.; Best, B.; Neumann-Haefelin, D.; Mueller-Lantzsch, N. Human endogenous retrovirus K10: Expression of Gag protein and detection of antibodies in patients with seminomas. J. Virol. 1995, 69, 414–421. [Google Scholar] [CrossRef]
- Löwer, R.; Löwer, J.; Tondera-Koch, C.; Kurth, R. A general method for the identification of transcribed retrovirus sequences (R-U5 PCR) reveals the expression of the human endogenous retrovirus loci HERV-H and HERV-K in teratocarcinoma cells. Virology 1993, 192, 501–511. [Google Scholar] [CrossRef]
- Schulte, A.M.; Lai, S.; Kurtz, A.; Czubayko, F.; Riegel, A.T.; Wellstein, A. Human trophoblast and choriocarcinoma expression of the growth factor pleiotrophin attributable to germ-line insertion of an endogenous retrovirus. Proc. Natl. Acad. Sci. USA 1996, 93, 14759–14764. [Google Scholar] [CrossRef]
- Bera, T.K.; Tsukamoto, T.; Panda, D.K.; Huang, T.; Guzman, R.C.; Hwang, S.-I.; Nandi, S. Defective retrovirus insertion activates C-Ha-Ras proto-oncogene in an MNU-induced rat mammary carcinoma. Biochem. Biophys. Res. Commun. 1998, 248, 835–840. [Google Scholar] [CrossRef] [PubMed]
- Ahn, K.; Kim, H.S. Structural and quantitative expression analyses of HERV gene family in human tissues. Mol. Cells 2009, 28, 99–103. [Google Scholar] [CrossRef] [PubMed]
- Yang, C.; Guo, X.; Li, J.; Han, J.; Jia, L.; Wen, H.L.; Sun, C.; Wang, X.; Zhang, B.; Li, J.; et al. Significant upregulation of HERV-K (HML-2) transcription levels in human lung cancer and cancer cells. Front. Microbiol. 2022, 13, 850444. [Google Scholar] [CrossRef]
- Tönjes, R.R.; Löwer, R.; Boller, K.; Denner, J.; Hasenmaier, B.; Kirsch, H.; König, H.; Korbmacher, C.; Limbach, C.; Lugert, R.; et al. HERV-K: The biologically most active human endogenous retrovirus family. JAIDS J. Acquir. Immune Defic. Syndr. 1996, 13, S261–S267. [Google Scholar] [CrossRef]
- Barbulescu, M.; Turner, G.; Seaman, M.I.; Deinard, A.S.; Kidd, K.K.; Lenz, J. Many human endogenous retrovirus K (HERV-K) proviruses are unique to humans. Curr. Biol. 1999, 9, 861–868. [Google Scholar] [CrossRef] [PubMed]
- Etkind, P.R.; Lumb, K.; Du, J.; Racevskis, J. Type 1 HERV-K genome is spliced into subgenomic transcripts in the human breast tumor cell line T47D. Virology 1997, 234, 304–308. [Google Scholar] [CrossRef] [PubMed]
- Srinivasachar Badarinarayan, S.; Sauter, D. Not all viruses cause disease: HERV-K(HML-2) in healthy human tissues. PLoS Biol. 2022, 20, e3001884. [Google Scholar] [CrossRef] [PubMed]
- Dervan, E.; Bhattacharyya, D.D.; McAuliffe, J.D.; Khan, F.H.; Glynn, S.A. Ancient adversary—HERV-K (HML-2) in cancer. Front. Oncol. 2021, 11, 658489. [Google Scholar] [CrossRef] [PubMed]
- Dupressoir, A.; Lavialle, C.; Heidmann, T. From ancestral infectious retroviruses to bona fide cellular genes: Role of the captured syncytins in placentation. Placenta 2012, 33, 663–671. [Google Scholar] [CrossRef] [PubMed]
- Jean-Luc, B.; Dimitri, L.; Valérie, C.; Olivier, B.; Guy, O.; Sylvie, C.-F.; Bernard, M.; François, M.; François-Loïc, C. An envelope glycoprotein of the human endogenous retrovirus HERV-W is expressed in the human placenta and fuses cells expressing the type D mammalian retrovirus receptor. J. Virol. 2000, 74, 3321–3329. [Google Scholar] [CrossRef]
- Mi, S.; Lee, X.; Li, X.; Veldman, G.M.; Finnerty, H.; Racie, L.; LaVallie, E.; Tang, X.-Y.; Edouard, P.; Howes, S.; et al. Syncytin is a captive retroviral envelope protein involved in human placental morphogenesis. Nature 2000, 403, 785–789. [Google Scholar] [CrossRef]
- Blaise, S.; de Parseval, N.; Bénit, L.; Heidmann, T. Genomewide screening for fusogenic human endogenous retrovirus envelopes identifies syncytin 2, a gene conserved on primate evolution. Proc. Natl. Acad. Sci. USA 2003, 100, 13013–13018. [Google Scholar] [CrossRef]
- Best, S.; Le Tissier, P.; Towers, G.; Stoye, J.P. Positional cloning of the mouse retrovirus restriction gene Fvl. Nature 1996, 382, 826–829. [Google Scholar] [CrossRef]
- Morozov, V.A.; Dao Thi, V.L.; Denner, J. The transmembrane protein of the human endogenous retrovirus—K (HERV-K) modulates cytokine release and gene expression. PLoS ONE 2013, 8, e70399. [Google Scholar] [CrossRef] [PubMed]
- Bieda, K.; Hoffmann, A.; Boller, K. Phenotypic heterogeneity of human endogenous retrovirus particles produced by teratocarcinoma cell lines. J. Gen. Virol. 2001, 82, 591–596. [Google Scholar] [CrossRef] [PubMed]
- Gotzinger, N.; Sauter, M.; Roemer, K.; Mueller-Lantzsch, N. Regulation of human endogenous retrovirus-K Gag expression in teratocarcinoma cell lines and human tumours. J. Gen. Virol. 1996, 77, 2983–2990. [Google Scholar] [CrossRef]
- Depil, S.; Roche, C.; Dussart, P.; Prin, L. Expression of a human endogenous retrovirus, HERV-K, in the blood cells of leukemia patients. Leukemia 2002, 16, 254–259. [Google Scholar] [CrossRef] [PubMed]
- Brodsky, I.; Foley, B.; Gillespie, D. Expression of human endogenous retrovirus (HERV-K) in chronic myeloid leukemia. Leuk. Lymphoma 1993, 11, 119–123. [Google Scholar] [CrossRef]
- Brodsky, I.; Foley, B.; Haines, D.; Johnston, J.; Cuddy, K.; Gillespie, D. Expression of HERV-K proviruses in human leukocytes. Blood 1993, 81, 2369–2374. [Google Scholar] [CrossRef] [PubMed]
- Cowan, A.J.; Green, D.J.; Kwok, M.; Lee, S.; Coffey, D.G.; Holmberg, L.A.; Tuazon, S.; Gopal, A.K.; Libby, E.N. Diagnosis and management of multiple myeloma: A review. JAMA 2022, 327, 464–477. [Google Scholar] [CrossRef]
- Oganesyan, A.; Gregory, A.; Malard, F.; Ghahramanyan, N.; Mohty, M.; Kazandjian, D.; Mekinian, A.; Hakobyan, Y. Monoclonal gammopathies of clinical significance (MGCS): In pursuit of optimal treatment. Front. Immunol. 2022, 13, 1045002. [Google Scholar] [CrossRef]
- Masuda, Y.; Ishihara, R.; Murakami, Y.; Watanabe, S.; Asao, Y.; Gotoh, N.; Kasamatsu, T.; Takei, H.; Kobayashi, N.; Saitoh, T.; et al. Clinical significance of human endogenous retrovirus K (HERV-K) in multiple myeloma progression. Int. J. Hematol. 2022. [Google Scholar] [CrossRef]
- Kleiman, A.; Senyuta, N.; Trjakin, A.; Vinogradova, T.; Karseladze, A.; Gurtsrvitch, V.; Tjulandin, S. Expression of human endogenous retroviruses HERV-K/HTDV in germ cell tumours: Possible biological role and clinical application. In Germ Cell Tumours V.; Harnden, P., Joffe, J.K., Jones, W.G., Eds.; Springer London: London, UK, 2002; pp. 43–44. ISBN 978-1-4471-3281-3. [Google Scholar]
- Xue, B.; Zeng, T.; Jia, L.; Yang, D.; Lin, S.L.; Sechi, L.A.; Kelvin, D.J. Identification of the distribution of human endogenous retroviruses K (HML-2) by PCR-based target enrichment sequencing. Retrovirology 2020, 17, 10. [Google Scholar] [CrossRef]
- Lu, X.; Sachs, F.; Ramsay, L.; Jacques, P.-É.; Göke, J.; Bourque, G.; Ng, H.-H. The retrovirus HERVH Is a long noncoding RNA required for human embryonic stem cell identity. Nat. Struct. Mol. Biol. 2014, 21, 423–425. [Google Scholar] [CrossRef]
- Durruthy-Durruthy, J.; Sebastiano, V.; Wossidlo, M.; Cepeda, D.; Cui, J.; Grow, E.J.; Davila, J.; Mall, M.; Wong, W.H.; Wysocka, J.; et al. The primate-specific noncoding RNA HPAT5 regulates pluripotency during human preimplantation development and nuclear reprogramming. Nat. Genet. 2016, 48, 44–52. [Google Scholar] [CrossRef] [PubMed]
- Fuentes, D.R.; Swigut, T.; Wysocka, J. Systematic perturbation of retroviral LTRs reveals widespread long-range effects on human gene regulation. eLife 2018, 7, e35989. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Li, T.; Preissl, S.; Amaral, M.L.; Grinstein, J.D.; Farah, E.N.; Destici, E.; Qiu, Y.; Hu, R.; Lee, A.Y.; et al. Transcriptionally active HERV-H retrotransposons demarcate topologically associating domains in human pluripotent stem cells. Nat. Genet. 2019, 51, 1380–1388. [Google Scholar] [CrossRef] [PubMed]
- Jansz, N.; Faulkner, G.J. Endogenous retroviruses in the origins and treatment of cancer. Genome Biol. 2021, 22, 147. [Google Scholar] [CrossRef] [PubMed]
- Kaufmann, S.; Sauter, M.; Schmitt, M.; Baumert, B.; Best, B.; Boese, A.; Roemer, K.; Mueller-Lantzsch, N. Human endogenous retrovirus protein Rec interacts with the testicular Zinc-finger protein and androgen receptor. J. Gen. Virol. 2010, 91, 1494–1502. [Google Scholar] [CrossRef]
- Grow, E.J.; Flynn, R.A.; Chavez, S.L.; Bayless, N.L.; Wossidlo, M.; Wesche, D.J.; Martin, L.; Ware, C.B.; Blish, C.A.; Chang, H.Y.; et al. Intrinsic retroviral reactivation in human preimplantation embryos and pluripotent cells. Nature 2015, 522, 221–225. [Google Scholar] [CrossRef]
- Kassiotis, G. Endogenous retroviruses and the development of cancer. J. Immunol. 2014, 192, 1343–1349. [Google Scholar] [CrossRef]
- Hanke, K.; Hohn, O.; Bannert, N. HERV-K(HML-2), a seemingly silent subtenant—but still waters run deep. APMIS 2016, 124, 67–87. [Google Scholar] [CrossRef]
- Turner, G.; Barbulescu, M.; Su, M.; Jensen-Seaman, M.I.; Kidd, K.K.; Lenz, J. Insertional polymorphisms of full-length endogenous retroviruses in humans. Curr. Biol. 2001, 11, 1531–1535. [Google Scholar] [CrossRef]
- Schulz, W.A. Does HERV-K represent a potential therapeutic target for prostate cancer? Expert. Opin. Ther. Targets 2017, 21, 921–924. [Google Scholar] [CrossRef]
- Wright, C.J.; Smith, C.W.J.; Jiggins, C.D. Alternative splicing as a source of phenotypic diversity. Nat. Rev. Genet. 2022, 23, 697–710. [Google Scholar] [CrossRef] [PubMed]
- Nooraei, S.; Bahrulolum, H.; Hoseini, Z.S.; Katalani, C.; Hajizade, A.; Easton, A.J.; Ahmadian, G. Virus-like particles: Preparation, immunogenicity and their roles as nanovaccines and drug nanocarriers. J. Nanobiotechnology 2021, 19, 59. [Google Scholar] [CrossRef] [PubMed]
- Davis, L.E.; Shalin, S.C.; Tackett, A.J. Current state of melanoma diagnosis and treatment. Cancer Biol. Ther. 2019, 20, 1366–1379. [Google Scholar] [CrossRef] [PubMed]
- Hahn, S.; Ugurel, S.; Hanschmann, K.M.; Strobel, H.; Tondera, C.; Schadendorf, D.; Löwer, J.; Löwer, R. Serological response to human endogenous retrovirus K in melanoma patients correlates with survival probability. AIDS Res. Hum. Retrovir. 2008, 24, 717–723. [Google Scholar] [CrossRef]
- Schiavetti, F.; Thonnard, J.; Colau, D.; Boon, T.; Coulie, P. A human endogenous retroviral sequence encoding an antigen recognized on melanoma by cytolytic T lymphocytes. Cancer Res. 2002, 62, 5510–5516. [Google Scholar]
- Qi, X.; Sandmeyer, S.; Zaher, H. DNA repair | nonhomologous recombination: Retrotransposons. In Encyclopedia of Biological Chemistry III; Jez, J., Ed.; Elsevier: Oxford, UK, 2021; pp. 313–322. ISBN 978-0-12-822040-5. [Google Scholar]
- Briggs, E.M.; Ha, S.; Mita, P.; Brittingham, G.; Sciamanna, I.; Spadafora, C.; Logan, S.K. Long interspersed nuclear element-1 expression and retrotransposition in prostate cancer cells. Mob. DNA 2018, 9, 1. [Google Scholar] [CrossRef]
- Cardelli, M.; van Doorn, R.; Larcher, L.; Donato, M.D.; Piacenza, F.; Pierpaoli, E.; Giacconi, R.; Malavolta, M.; Rachakonda, S.; Gruis, N.A.; et al. Association of HERV-K and LINE-1 hypomethylation with reduced disease-free survival in melanoma patients. Epigenomics 2020, 12, 1689–1706. [Google Scholar] [CrossRef]
- Muster, T.; Waltenberger, A.; Grassauer, A.; Hirschl, S.; Caucig, P.; Romirer, I.; Födinger, D.; Seppele, H.; Schanab, O.; Magin-Lachmann, C.; et al. An endogenous retrovirus derived from human melanoma cells. Cancer Res. 2003, 63, 8735–8741. [Google Scholar]
- Büscher, K.; Trefzer, U.; Hofmann, M.; Sterry, W.; Kurth, R.; Denner, J. Expression of human endogenous retrovirus K in melanomas and melanoma cell lines. Cancer Res. 2005, 65, 4172–4180. [Google Scholar] [CrossRef]
- Schmitt, K.; Reichrath, J.; Roesch, A.; Meese, E.; Mayer, J. Transcriptional profiling of human endogenous retrovirus group HERV-K(HML-2) loci in melanoma. Genome Biol. Evol. 2013, 5, 307–328. [Google Scholar] [CrossRef] [PubMed]
- Singh, M.; Cai, H.; Bunse, M.; Feschotte, C.; Izsvák, Z. Human endogenous retrovirus K Rec forms a regulatory loop with MITF that opposes the progression of melanoma to an invasive stage. Viruses 2020, 12, 1303. [Google Scholar] [CrossRef]
- Miriam, D.; Marlies, S.; Vivienne, A.; Licht, J.D.; Klaus, R.; Nikolaus, M.-L. Physical and functional interactions of human endogenous retrovirus proteins Np9 and Rec with the promyelocytic leukemia zinc finger protein. J. Virol. 2007, 81, 5607–5616. [Google Scholar] [CrossRef]
- Kirsten, H.; Oliver, H.; Linda, L.; Katharina, F.; Jula, W.; Reinhard, K.; Norbert, B. Staufen-1 interacts with the human endogenous retrovirus family HERV-K(HML-2) Rec and Gag proteins and increases virion production. J. Virol. 2013, 87, 11019–11030. [Google Scholar] [CrossRef]
- Damjanov, I. Teratocarcinoma stem cells. Cancer Surv. 1990, 9, 303–319. [Google Scholar]
- Thowfeequ, S.; Srinivas, S. Embryonic and extraembryonic tissues during mammalian development: Shifting boundaries in time and space. Philos. Trans. R. Soc. B Biol. Sci. 2022, 377, 20210255. [Google Scholar] [CrossRef] [PubMed]
- Morozov, V.A.; Morozov, A.V. A comprehensive analysis of human endogenous retroviruses HERV-K (HML.2) from teratocarcinoma cell lines and detection of viral cargo in microvesicles. Int. J. Mol. Sci. 2021, 22, 12398. [Google Scholar] [CrossRef]
- Wang, C.; Liang, C. The insertion and dysregulation of transposable elements in osteosarcoma and their association with patient event-free survival. Sci. Rep. 2022, 12, 377. [Google Scholar] [CrossRef] [PubMed]
- Garcia-Montero, A.; Vasseur, S.; Mallo, G.V.; Soubeyran, P.; Dagorn, J.C.; Iovanna, J.L. Expression of the stress-induced P8 MRNA is transiently activated after culture medium change. Eur. J. Cell Biol. 2001, 80, 720–725. [Google Scholar] [CrossRef] [PubMed]
- Ferrari, L.; Cafora, M.; Rota, F.; Hoxha, M.; Iodice, S.; Tarantini, L.; Dolci, M.; Delbue, S.; Pistocchi, A.; Bollati, V. Extracellular vesicles released by colorectal cancer cell lines modulate innate immune response in zebrafish model: The possible role of human endogenous retroviruses. Int. J. Mol. Sci. 2019, 20, 3669. [Google Scholar] [CrossRef]
- Willer, A.; Saußele, S.; Gimbel, W.; Zeifarth, W.; Kister, P.; Leib-Mo¨sch, C.; Hehlmann, R. Two groups of endogenous MMTV related retroviral Env transcripts expressed in human tissues. Virus Genes 1997, 15, 123–133. [Google Scholar] [CrossRef] [PubMed]
- Alkabban, F.; Ferguson, T. Breast cancer. In StatPearls; StatPearls Publishing: Treasure Island, FL, USA, 2022. [Google Scholar]
- Lemaître, C.; Tsang, J.; Bireau, C.; Heidmann, T.; Dewannieux, M. A Human endogenous retrovirus-derived gene that can contribute to oncogenesis by activating the ERK pathway and inducing migration and invasion. PLoS Pathog. 2017, 13, e1006451. [Google Scholar] [CrossRef] [PubMed]
- Contreras-Galindo, R.; Kaplan, M.; Leissner, P.; Verjat, T.; Ferlenghi, I.; Bagnoli, F.; Giusti, F.; Dosik, M.; Hayes, D.; Gitlin, S.; et al. Human endogenous retrovirus K (HML-2) elements in the plasma of people with lymphoma and breast cancer. J. Virol. 2008, 82, 9329–9336. [Google Scholar] [CrossRef] [PubMed]
- Roett MA, E.P. Ovarian Cancer: An Overview. Am. Fam. Physician 2009, 80, 609. [Google Scholar] [PubMed]
- Rycaj, K.; Plummer, J.B.; Yin, B.; Li, M.; Garza, J.; Radvanyi, L.; Ramondetta, L.M.; Lin, K.; Johanning, G.L.; Tang, D.G.; et al. Cytotoxicity of human endogenous retrovirus K–specific T cells toward autologous ovarian cancer cells. Clin. Cancer Res. 2015, 21, 471–483. [Google Scholar] [CrossRef] [PubMed]
- Stephen, W.L.; Taylor, L.; Soon-Sutton Anu, R.I.; Sajjad, H.; Siref, L.E. Prostate cancer. In StatPearls; StatPearls Publishing: Treasure Island, FL, USA, 2022. [Google Scholar]
- Ishida, T.; Obata, Y.; Ohara, N.; Matsushita, H.; Sato, S.; Uenaka, A.; Saika, T.; Miyamura, T.; Chayama, K.; Nakamura, Y.; et al. Identification of the HERV-K Gag antigen in prostate cancer by SEREX using autologous patient serum and its immunogenicity. Cancer Immun. 2008, 8, 15. [Google Scholar]
- Johanning, G.L.; Malouf, G.G.; Zheng, X.; Esteva, F.J.; Weinstein, J.N.; Wang-Johanning, F.; Su, X. Expression of human endogenous retrovirus-K is strongly associated with the basal-like breast cancer phenotype. Sci. Rep. 2017, 7, 41960. [Google Scholar] [CrossRef] [PubMed]
- Goering, W.; Ribarska, T.; Schulz, W.A. Selective changes of retroelement expression in human prostate cancer. Carcinogenesis 2011, 32, 1484–1492. [Google Scholar] [CrossRef]
- Wallace, T.A.; Downey, R.F.; Seufert, C.J.; Schetter, A.; Dorsey, T.H.; Johnson, C.A.; Goldman, R.; Loffredo, C.A.; Yan, P.; Sullivan, F.J.; et al. Elevated HERV-K mRNA expression in PBMC is associated with a prostate cancer diagnosis particularly in older men and smokers. Carcinogenesis 2014, 35, 2074–2083. [Google Scholar] [CrossRef]
- Helgeson, B.E.; Tomlins, S.A.; Shah, N.; Laxman, B.; Cao, Q.; Prensner, J.R.; Cao, X.; Singla, N.; Montie, J.E.; Varambally, S.; et al. Characterization of TMPRSS2:ETV5 and SLC45A3:ETV5 gene fusions in prostate cancer. Cancer Res. 2008, 68, 73–80. [Google Scholar] [CrossRef]
- Mengying, L.; Lei, J.; Hanping, L.; Yongjian, L.; Jingwan, H.; Xiaolin, W.; Tianyi, L.; Jingyun, L.; Bohan, Z.; Xiuli, Z.; et al. P53 binding sites in long terminal repeat 5Hs (LTR5Hs) of human endogenous retrovirus K family (HML-2 subgroup) play important roles in the regulation of LTR5Hs transcriptional activity. Microbiol. Spectr. 2022, 10, e00485-22. [Google Scholar] [CrossRef]
- Rezaei, S.D.; Hayward, J.A.; Norden, S.; Pedersen, J.; Mills, J.; Hearps, A.C.; Tachedjian, G. HERV-K Gag RNA and protein levels are elevated in malignant regions of the prostate in males with prostate cancer. Viruses 2021, 13, 449. [Google Scholar] [CrossRef] [PubMed]
- Hua, T.; Zeng, Z.; Chen, J.; Xue, Y.; Li, Y.; Sang, Q. Human malignant rhabdoid tumor antigens as biomarkers and potential therapeutic targets. Cancers 2022, 14, 3685. [Google Scholar] [CrossRef] [PubMed]
- Doucet-O’Hare, T.T.; DiSanza, B.L.; DeMarino, C.; Atkinson, A.L.; Rosenblum, J.S.; Henderson, L.J.; Johnson, K.R.; Kowalak, J.; Garcia-Montojo, M.; Allen, S.J.; et al. SMARCB1 deletion in atypical teratoid rhabdoid tumors results in human endogenous retrovirus K (HML-2) expression. Sci. Rep. 2021, 11, 12893. [Google Scholar] [CrossRef]
- Sapkota, S.S.H. Non-hodgkin lymphoma. In StatPearls; StatPearls Publishing: Treasure Island, FL, USA, 2022. [Google Scholar]
- GlaxoSmithKline; GileadSciences. A Phase I/II Study of Safety and Efficacy of Lamivudine (EPIVIR®) and Tenofovir Disoproxil Fumarate (VIREAD®) Used to Lower the Plasma Level of Viral RNA of HERV-K(HML2) in Patients with Lymphoma. Identifier NCT01528865. University of Michigan Rogel Cancer Center. 2016. Available online: https://clinicaltrials.gov/ct2/show/NCT01528865 (accessed on 1 December 2022).
- Löwer, J.; Löwer, R.; Stegmann, J.; Frank, H.; Kurth, R. Retrovirus particle production in three of four human teratocarcinoma cell lines. In Modern Trends in Human; Leukemia, I.V., Neth, R., Gallo, R.C., Graf, T., Mannweiler, K., Winkler, K., Eds.; Springer: Berlin/Heidelberg, Germany, 1981; pp. 541–544. ISBN 978-3-642-67984-1. [Google Scholar]
- Li, M.D.; Bronson, D.L.; Lemke, T.D.; Faras, A.J. Restricted expression of New HERV-K members in human teratocarcinoma cells. Virology 1995, 208, 733–741. [Google Scholar] [CrossRef]
- Kassiotis, G.; Stoye, J.P. Immune responses to endogenous retroelements: Taking the bad with the good. Nat. Rev. Immunol. 2016, 16, 207–219. [Google Scholar] [CrossRef] [PubMed]
- Messerschmitt, P.J.; Garcia, R.M.; Abdul-Karim, F.W.; Greenfield, E.M.; Getty, P.J. Osteosarcoma. JAAOS J. Am. Acad. Orthop. Surg. 2009, 17, 515–527. [Google Scholar] [CrossRef] [PubMed]
- Moore, D.D.; Luu, H.H. Osteosarcoma. In Cancer Trearment and Research. Orthopaedic Oncology; Primary and Metastatic Tumors of the Skeletal, System; Peabody, T.D., Attar, S., Eds.; Springer International Publishing: Cham, Switzerland, 2014; Volume 162, pp. 65–92. ISBN 978-3-319-07323-1. [Google Scholar]
- Rodić, N.; Burns, K.H. Long Interspersed Element–1 (LINE-1): Passenger or driver in human neoplasms? PLoS Genet. 2013, 9, e1003402. [Google Scholar] [CrossRef]
- Levine, A.J.; Ting, D.T.; Greenbaum, B.D. P53 and the defenses against genome instability caused by transposons and repetitive elements. BioEssays 2016, 38, 508–513. [Google Scholar] [CrossRef]
- Wylie, A.; Jones, A.; D’Brot, A.; Lu, W.-J.; Kurtz, P.; Moran, J.; Rakheja, D.; Chen, K.; Hammer, R.; Comerford, S.; et al. P53 genes function to restrain mobile elements. Genes. Dev. 2015, 30, 64–77. [Google Scholar] [CrossRef]
- Clayton, E.A.; Wang, L.; Rishishwar, L.; Wang, J.; McDonald, J.F.; Jordan, I.K. Patterns of transposable element expression and insertion in cancer. Front. Mol. Biosci. 2016, 3, 76. [Google Scholar] [CrossRef] [PubMed]
- Dolci, M.; Favero, C.; Toumi, W.; Favi, E.; Tarantini, L.; Signorini, L.; Basile, G.; Bollati, V.; D’Alessandro, S.; Bagnoli, P.; et al. Human endogenous retroviruses long terminal repeat methylation, transcription, and protein expression in human colon cancer. Front. Oncol. 2020, 10, 569015. [Google Scholar] [CrossRef] [PubMed]
- Mármol, I.; Sánchez-de-Diego, C.; Pradilla Dieste, A.; Cerrada, E.; Rodriguez Yoldi, M.J. Colorectal carcinoma: A general overview and future perspectives in colorectal cancer. Int. J. Mol. Sci. 2017, 18, 197. [Google Scholar] [CrossRef]
- Lin, O.S. Acquired risk factors for colorectal cancer. In Cancer Epidemiology: Modifiable Factors; Verma, M., Ed.; Humana Press: Totowa, NJ, USA, 2009; pp. 361–372. ISBN 978-1-60327-492-0. [Google Scholar]
- Mallo, G.V.; Fiedler, F.; Calvo, E.L.; Ortiz, E.M.; Vasseur, S.; Keim, V.; Morisset, J.; Iovanna, J.L. Cloning and expression of the rat P8 CDNA, a new gene activated in pancreas during the acute phase of pancreatitis, pancreatic development, and regeneration, and which promotes cellular growth. J. Biol. Chem. 1997, 272, 32360–32369. [Google Scholar] [CrossRef] [PubMed]
- Malicet, C.; Dagorn, J.C.; Neira, J.L.; Iovanna, J.L. P8 and Prothymosin Alpha: Unity Is Strength. Cell Cycle 2006, 5, 829–830. [Google Scholar] [CrossRef]
- Malicet, C.; Giroux, V.; Vasseur, S.; Dagorn, J.C.; Neira, J.L.; Iovanna, J.L. Regulation of apoptosis by the P8/prothymosin α complex. Proc. Natl. Acad. Sci. USA 2006, 103, 2671–2676. [Google Scholar] [CrossRef]
- Martin, A.T.; Li Xinyu, A.; Sanders, J.A.; Ye, L.; Frewer, K.; Hargest, R.; Jiang, G.W. NUPR1 and its potential role in cancer and pathological conditions (review). Int. J. Oncol. 2021, 58, 21. [Google Scholar] [CrossRef]
- Ko, E.-J.; Ock, M.-S.; Choi, Y.-H.; Iovanna, J.L.; Mun, S.; Han, K.; Kim, H.-S.; Cha, H.-J. Human endogenous retrovirus (HERV)-K Env gene knockout affects tumorigenic characteristics of NUPR1 gene in DLD-1 colorectal cancer cells. Int. J. Mol. Sci. 2021, 22, 3941. [Google Scholar] [CrossRef]
- Gironella, M.; Malicet, C.; Cano, C.; Sandi, M.J.; Hamidi, T.; Tauil, R.M.N.; Baston, M.; Valaco, P.; Moreno, S.; Lopez, F.; et al. P8/NUPR1 regulates DNA-repair activity after double-strand gamma irradiation-induced DNA damage. J. Cell. Physiol. 2009, 221, 594–602. [Google Scholar] [CrossRef] [PubMed]
- Hamidi, T.; Algül, H.; Cano, C.E.; Sandi, M.J.; Molejon, M.I.; Riemann, M.; Calvo, E.L.; Lomberk, G.; Dagorn, J.-C.; Weih, F.; et al. Nuclear protein 1 promotes pancreatic cancer development and protects cells from stress by inhibiting apoptosis. J. Clin. Investig. 2012, 122, 2092–2103. [Google Scholar] [CrossRef]
- Encinar, J.A.; Mallo, G.V.; Mizyrycki, C.; Giono, L.; González-Ros, J.M.; Rico, M.; Cánepa, E.; Moreno, S.; Neira, J.L.; Iovanna, J.L. Human P8 is a HMG-I/Y-like protein with DNA binding activity enhanced by phosphorylation. J. Biol. Chem. 2001, 276, 2742–2751. [Google Scholar] [CrossRef] [PubMed]
- Grasso, D.; Garcia, M.N.; Hamidi, T.; Cano, C.; Calvo, E.; Lomberk, G.; Urrutia, R.; Iovanna, J.L. Genetic inactivation of the pancreatitis-inducible gene Nupr1 Impairs PanIN formation by modulating KrasG12D-induced senescence. Cell Death Differ. 2014, 21, 1633–1641. [Google Scholar] [CrossRef] [PubMed]
- Ashktorab, H.; Daremipouran, M.; Goel, A.; Varma, S.; Leavitt, R.; Sun, X.; Brim, H. DNA methylome profiling identifies novel methylated genes in African American patients with colorectal neoplasia. Epigenetics 2014, 9, 503–512. [Google Scholar] [CrossRef]
- Gualtieri, A.; Andreola, F.; Sciamanna, I.; Sinibaldi Vallebona, P.; Serafino, A.; Spadafora, C. Increased expression and copy number amplification of LINE-1 and SINE B1 retrotransposable elements in murine mammary carcinoma progression. Oncotarget 2013, 4, 1882–1893. [Google Scholar] [CrossRef]
- Chappell, G.; Kutanzi, K.; Uehara, T.; Tryndyak, V.; Hong, H.-H.; Hoenerhoff, M.; Beland, F.A.; Rusyn, I.; Pogribny, I.P. Genetic and epigenetic changes in fibrosis-associated hepatocarcinogenesis in mice. Int. J. Cancer 2014, 134, 2778–2788. [Google Scholar] [CrossRef] [PubMed]
- DeRoo, L.A.; Bolick, S.C.E.; Xu, Z.; Umbach, D.M.; Shore, D.; Weinberg, C.R.; Sandler, D.P.; Taylor, J.A. Global DNA methylation and one-carbon metabolism gene polymorphisms and the risk of breast cancer in the sister study. Carcinogenesis 2014, 35, 333–338. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Ogino, S.; Nishihara, R.; Lochhead, P.; Imamura, Y.; Kuchiba, A.; Morikawa, T.; Yamauchi, M.; Liao, X.; Qian, Z.R.; Sun, R.; et al. Prospective study of family history and colorectal cancer risk by tumor LINE-1 methylation level. JNCI J. Natl. Cancer Inst. 2013, 105, 130–140. [Google Scholar] [CrossRef]
- Rhyu, D.-W.; Kang, Y.-J.; Ock, M.-S.; Eo, J.-W.; Choi, Y.-H.; Kim, W.-J.; Leem, S.-H.; Yi, J.-M.; Kim, H.-S.; Cha, H.-J. Expression of human endogenous retrovirus Env genes in the blood of breast cancer patients. Int. J. Mol. Sci. 2014, 15, 9173–9183. [Google Scholar] [CrossRef]
- Liang, Q.; Ding, J.; Xu, R.; Xu, Z.; Zheng, S. Identification of a novel human endogenous retrovirus and promoter activity of its 5′ U3. Biochem. Biophys. Res. Commun. 2009, 382, 468–472. [Google Scholar] [CrossRef]
- Zare, M.; Mostafaei, S.; Ahmadi, A.; Azimzadeh Jamalkandi, S.; Abedini, A.; Esfahani-Monfared, Z.; Dorostkar, R.; Saadati, M. Human endogenous retrovirus Env genes: Potential blood biomarkers in lung cancer. Microb. Pathog. 2018, 115, 189–193. [Google Scholar] [CrossRef]
- Zhang, M.; Liang, J.Q.; Zheng, S. Expressional activation and functional roles of human endogenous retroviruses in cancers. Rev. Med. Virol. 2019, 29, e2025. [Google Scholar] [CrossRef] [PubMed]
- Manghera, M.; Ferguson, J.; Douville, R. HERV-K polyprotein processing and reverse transcriptase expression in human cell line models of neurological disease. Viruses 2015, 7, 320–332. [Google Scholar] [CrossRef] [PubMed]
- Harbeck, N.; Penault-Llorca, F.; Cortes, J.; Gnant, M.; Houssami, N.; Poortmans, P.; Ruddy, K.; Tsang, J.; Cardoso, F. Breast cancer. Nat. Rev. Dis. Prim. 2019, 5, 66. [Google Scholar] [CrossRef] [PubMed]
- Golan, M.; Hizi, A.; Resau, J.H.; Yaal-Hahoshen, N.; Reichman, H.; Keydar, I.; Tsarfaty, I. Human endogenous retrovirus (HERV-K) reverse transcriptase as a breast cancer prognostic marker. Neoplasia 2008, 10, 521-IN2. [Google Scholar] [CrossRef] [PubMed]
- Ejthadi, H.D.; Martin, J.H.; Junying, J.; Roden, D.A.; Lahiri, M.; Warren, P.; Murray, P.G.; Nelson, P.N. A novel multiplex RT-PCR system detects human endogenous retrovirus-K in breast cancer. Arch. Virol. 2005, 150, 177–184. [Google Scholar] [CrossRef] [PubMed]
- Wang-Johanning, F.; Frost, A.; Johanning, G.; Khazaeli, M.; LoBuglio, A.; Shaw, D.; Strong, T. Expression of human endogenous retrovirus k envelope transcripts in human breast cancer. Clin. Cancer Res. 2001, 7, 1553–1560. [Google Scholar] [PubMed]
- Wang-Johanning, F.; Li, M.; Esteva, F.J.; Hess, K.R.; Yin, B.; Rycaj, K.; Plummer, J.B.; Garza, J.G.; Ambs, S.; Johanning, G.L. Human endogenous retrovirus type K antibodies and mRNA as serum biomarkers of early-stage breast cancer. Int. J. Cancer 2014, 134, 587–595. [Google Scholar] [CrossRef]
- Wang-Johanning, F.; Radvanyi, L.; Rycaj, K.; Plummer, J.B.; Yan, P.; Sastry, K.J.; Piyathilake, C.J.; Hunt, K.K.; Johanning, G.L. Human endogenous retrovirus K triggers an antigen-specific immune response in breast cancer patients. Cancer Res. 2008, 68, 5869–5877. [Google Scholar] [CrossRef]
- Zhao, J.; Rycaj, K.; Geng, S.; Li, M.; Plummer, J.B.; Yin, B.; Liu, H.; Xu, X.; Zhang, Y.; Yan, Y.; et al. Expression of human endogenous retrovirus type K envelope protein is a novel candidate prognostic marker for human breast cancer. Genes Cancer 2011, 2, 914–922. [Google Scholar] [CrossRef]
- Wang-Johanning, F.; Rycaj, K.; Plummer, J.B.; Li, M.; Yin, B.; Frerich, K.; Garza, J.G.; Shen, J.; Lin, K.; Yan, P.; et al. Immunotherapeutic potential of anti-human endogenous retrovirus-K envelope protein antibodies in targeting breast tumors. JNCI J. Natl. Cancer Inst. 2012, 104, 189–210. [Google Scholar] [CrossRef]
- Rosen, R.; Sapra, A. TNM classification. In StatPearls; StatPearls Publishing: Treasure Island, FL, USA, 2022. [Google Scholar]
- Rivlin, N.; Brosh, R.; Oren, M.; Rotter, V. Mutations in the P53 tumor suppressor gene: Important milestones at the various steps of tumorigenesis. Genes Cancer 2011, 2, 466–474. [Google Scholar] [CrossRef] [PubMed]
- Reis, B.; Jungbluth, A.; Frosina, D.; Holz, M.; Ritter, E.; Nakayama, E.; Ishida, T.; Obata, Y.; Carver, B.; Scher, H.; et al. Prostate cancer progression correlates with increased humoral immune response to a human rndogenous retrovirus Gag protein. Clin. Cancer Res. 2013, 19, 6112–6125. [Google Scholar] [CrossRef] [PubMed]
- Hu, J.; Han, J.; Li, H.; Zhang, X.; Liu, L.; Chen, F.; Zeng, B. Human embryonic kidney 293 cells: A vehicle for biopharmaceutical manufacturing, structural biology, and electrophysiology. Cells Tissues Organs 2018, 205, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Wegener, M.; Müller-McNicoll, M. Nuclear retention of mRNAs—quality control, gene regulation and human disease. Semin. Cell. Dev. Biol. 2018, 79, 131–142. [Google Scholar] [CrossRef]
- Löwer, R.; Tönjes, R.R.; Korbmacher, C.; Kurth, R.; Löwer, J. Identification of a Rev-related protein by analysis of spliced transcripts of the human endogenous retroviruses HTDV/HERV-K. J. Virol. 1995, 69, 141–149. [Google Scholar] [CrossRef]
- Yang, J.; Bogerd, H.P.; Peng, S.; Wiegand, H.; Truant, R.; Cullen, B.R. An ancient family of human endogenous retroviruses encodes a functional homolog of the HIV-1 Rev protein. Proc. Natl. Acad. Sci. USA 1999, 96, 13404–13408. [Google Scholar] [CrossRef]
- Zeng, T.; Fedeli, M.A.; Tanda, F.; Wang, Y.; Yang, D.; Xue, B.; Jia, L.; Palmieri, G.; Sechi, L.A.; Kelvin, D.J. Whole-exome sequencing of prostate cancer in sardinian identify recurrent UDP-glucuronosyltransferase amplifications. J. Cancer 2021, 12, 438–450. [Google Scholar] [CrossRef]
- Wang, Z.; Wang, Y.; Zhang, J.; Hu, Q.; Zhi, F.; Zhang, S.; Mao, D.; Zhang, Y.; Liang, H. Significance of the TMPRSS2:ERG gene fusion in prostate cancer. Mol. Med. Rep. 2017, 16, 5450–5458. [Google Scholar] [CrossRef]
- Jia, L.; Liu, M.; Yang, C.; Li, H.; Liu, Y.; Han, J.; Zhai, X.; Wang, X.; Li, T.; Li, J.; et al. Comprehensive identification and characterization of the HERV-K (HML-9) group in the human genome. Retrovirology 2022, 19, 11. [Google Scholar] [CrossRef]
- Sizemore, G.M.; Pitarresi, J.R.; Balakrishnan, S.; Ostrowski, M.C. The ETS family of oncogenic transcription factors in solid tumours. Nat. Rev. Cancer 2017, 17, 337–351. [Google Scholar] [CrossRef]
- Stine, Z.E.; Walton, Z.E.; Altman, B.J.; Hsieh, A.L.; Dang, C.V. MYC, metabolism, and cancer. Cancer Discov. 2015, 5, 1024–1039. [Google Scholar] [CrossRef]
- Kraus, B.; Fischer, K.; Sliva, K.; Schnierle, B.S. Vaccination directed against the human endogenous retrovirus-K (HERV-K) Gag protein slows HERV-K Gag expressing cell growth in a murine model system. Virol. J. 2014, 11, 58. [Google Scholar] [CrossRef] [PubMed]
- Ragonnaud, E.; Neukirch, L.; Pedersen, I.; Daradoumis, J.; Daradoumis, J.; Grunddal, K.; Duvnjak, L.; Bermejo, A.; Schroedel, S.; Thirion, C.; et al. P03.03 Active immunization against human endogenous retrovirus type K (HERV-K) as an immunotherapeutic strategy against solid tumors. J. Immunother. Cancer 2022, 10, A17.2-A18. [Google Scholar] [CrossRef]
- Wang-Johanning, F.; Rycaj, K.; Huang, M.; Plummer, J.; Marks, J.; Johanning, G.; Rosenblum, M. Anti-HERV-K antibody 6H5 and 6H5/RGel inhibit cell growth and induce apoptosis in breast and ovarian cancer cells. Cancer Res. 2007, 67, 906. [Google Scholar]
- Perron, H.; Medina, J. Anti-HERV-K Envelope Antibody and Uses Thereof. EP3351265A1; European Patent Application. Assignee: Geneuro SA. Filed 20 January 2017, 25 July 2018. Available online: https://patents.google.com/patent/EP3351265A1/en (accessed on 1 November 2022).
- Karimi, A.; Sheervalilou, R.; Kahroba, H. A new insight on activation of human endogenous retroviruses (HERVs) in malignant melanoma upon exposure to CuSO4. Biol. Trace Elem. Res. 2019, 191, 70–74. [Google Scholar] [CrossRef]
- Carlini, F.; Ridolfi, B.; Molinari, A.; Parisi, C.; Bozzuto, G.; Toccacieli, L.; Formisano, G.; De Orsi, D.; Paradisi, S.; Grober, O.M.V.; et al. The reverse transcription inhibitor abacavir shows anticancer activity in prostate cancer cell lines. PLoS ONE 2010, 5, e14221. [Google Scholar] [CrossRef]
- Ko, E.J.; Kim, E.T.; Kim, H.; Lee, C.M.; Koh, S.B.; Eo, W.K.; Kim, H.; Oh, Y.L.; Ock, M.S.; Kim, K.H.; et al. Effect of human endogenous retrovirus-K Env gene knockout on proliferation of ovarian cancer cells. Genes. Genom. 2022, 44, 1091–1097. [Google Scholar] [CrossRef]
- Temerozo, J.R.; Fintelman-Rodrigues, N.; dos Santos, M.C.; Hottz, E.D.; Sacramento, C.Q.; de Paula Dias da Silva, A.; Mandacaru, S.C.; dos Santos Moraes, E.C.; Trugilho, M.R.O.; Gesto, J.S.M.; et al. Human endogenous retrovirus K in the respiratory tract is associated with COVID-19 physiopathology. Microbiome 2022, 10, 65. [Google Scholar] [CrossRef]
- Hashemian, S.M.; Farhadi, T.; Velayati, A.A. A review on remdesivir: A possible promising agent for the treatment of COVID-19. Drug. Des. Devel. Ther. 2020, 14, 3215–3222. [Google Scholar] [CrossRef]
- Sacramento, C.Q.; Fintelman-Rodrigues, N.; Temerozo, J.R.; de Paula Dias Da Silva, A.; de Paula Dias Da Silva, A.; da Silva, C.D.S.; Ferreira, A.C.; Mattos, M.; Pão, C.R.R.; de Freitas, C.S.; et al. In vitro antiviral activity of the anti-HCV drugs daclatasvir and sofosbuvir against SARS-CoV-2, the aetiological agent of COVID-19. J. Antimicrob. Chemother. 2021, 76, 1874–1885. [Google Scholar] [CrossRef]
- Li, P.; Zheng, Y.; Chen, X. Drugs for autoimmune inflammatory diseases: From small molecule compounds to anti-TNF biologics. Front. Pharmacol. 2017, 8, 460. [Google Scholar] [CrossRef] [PubMed]
- Moru, X.; Kun, Q.; Hongxia, S.; Yongxiu, Y.; Venugopal, N.; Jianqiang, Y.; Aijian, Q. Glycosylation of ALV-J envelope protein at sites 17 and 193 is pivotal in the virus infection. J. Virol. 2022, 96, e01549-21. [Google Scholar] [CrossRef]
- Bermejo, A.; Daradoumis, J.; Azcoaga, P.; Ragonnaud, E.; Neukrich, L.; Nielsen, K.; Andersoon, A.; Scroedel, S.; Thirion, C.; Ørskov, C.; et al. P03.05 vaccine immunotherapy against human endogenous retrovirus: A focus on anti-HERV-K antibodies. J. Immunother. Cancer 2022, 10, A19.1-A19. [Google Scholar] [CrossRef]
- Harris, E.D. Copper Homeostasis: The role of cellular transporters. Nutr. Rev. 2001, 59, 281–285. [Google Scholar] [CrossRef] [PubMed]
- Tapiero, H.; Townsend, D.M.; Tew, K.D. Trace elements in human physiology and pathology. Copper. Biomed. Pharmacother. 2003, 57, 386–398. [Google Scholar] [CrossRef]
- Osredkar, J.; Sustar, N. Copper and zinc, biological role and significance of copper/zinc imbalance. J. Clin. Toxicol. 2011, 3, 2161-0494. [Google Scholar] [CrossRef]
- D’Mello, S.A.N.; Finlay, G.J.; Baguley, B.C.; Askarian-Amiri, M.E. Signaling pathways in melanogenesis. Int. J. Mol. Sci. 2016, 17, 1144. [Google Scholar] [CrossRef]
- Grubman, A.; White, A.R. Copper as a key regulator of cell signalling pathways. Expert. Rev. Mol. Med. 2014, 16, e11. [Google Scholar] [CrossRef]
- Arigony, A.L.V.; de Oliveira, I.M.; Machado, M.; Bordin, D.L.; Bergter, L.; Prá, D.; Pêgas Henriques, J.A. The influence of micronutrients in cell culture: A reflection on viability and genomic stability. Biomed. Res. Int. 2013, 2013, 597282. [Google Scholar] [CrossRef]
- Ibba, G.; Piu, C.; Uleri, E.; Serra, C.; Dolei, A. Disruption by SaCas9 endonuclease of HERV-K Env, a retroviral gene with oncogenic and neuropathogenic potential, inhibits molecules involved in cancer and amyotrophic lateral sclerosis. Viruses 2018, 10, 412. [Google Scholar] [CrossRef]
- Brezgin, S.; Kostyusheva, A.; Kostyushev, D.; Chulanov, V. Dead Cas systems: Types, principles, and applications. Int. J. Mol. Sci. 2019, 20, 6041. [Google Scholar] [CrossRef] [PubMed]
- Nath, A. HERV-K Suppression Using Antiretroviral Therapy in Volunteers with Amyotrophic Lateral Sclerosis (ALS). Identifier NCT02437110. National Institute of Neurological Disorders and Stroke. 2022. Available online: https://clinicaltrials.gov/ct2/show/NCT02437110 (accessed on 1 December 2022).
- Olasupo, O.; Lowe, M.; Krishan, A.; Collins, P.; Iorio, A.; Matino, D. Clotting factor concentrates for preventing bleeding and bleeding-related complications in previously treated individuals with haemophilia A or B. Cochrane Database Syst. Rev. 2021, 18, CD014201. [Google Scholar] [CrossRef]
- Gabay, C.; Lamacchia, C.; Palmer, G. IL-1 Pathways in inflammation and human diseases. Nat. Rev. Rheumatol. 2010, 6, 232–241. [Google Scholar] [CrossRef] [PubMed]
- Zhao, R.; Chinai, J.M.; Buhl, S.; Scandiuzzi, L.; Ray, A.; Jeon, H.; Ohaegbulam, K.C.; Ghosh, K.; Zhao, A.; Scharff, M.D.; et al. HHLA2 is a member of the B7 family and inhibits human CD4 and CD8 T-cell function. Proc. Natl. Acad. Sci. USA 2013, 110, 9879–9884. [Google Scholar] [CrossRef] [PubMed]
- Amatya, N.; Garg, A.V.; Gaffen, S.L. IL-17 signaling: The yin and the yang. Trends Immunol. 2017, 38, 310–322. [Google Scholar] [CrossRef]
- Wang, X.; Zhao, C.; Zhang, C.; Mei, X.; Song, J.; Sun, Y.; Wu, Z.; Shi, W. Increased HERV-E Clone 4–1 expression contributes to DNA hypomethylation and IL-17 release from CD4+ T Cells via MiR-302d/MBD2 in systemic lupus erythematosus. Cell Commun. Signal. 2019, 17, 94. [Google Scholar] [CrossRef]
- Dorward, D.A.; Russell, C.D.; Um, I.H.; Elshani, M.; Armstrong, S.D.; Penrice-Randal, R.; Millar, T.; Lerpiniere, C.E.B.; Tagliavini, G.; Hartley, C.S.; et al. Tissue-specific immunopathology in fatal COVID-19. Am. J. Respir. Crit. Care Med. 2020, 203, 192–201. [Google Scholar] [CrossRef]
- Merad, M.; Martin, J.C. Pathological inflammation in patients with COVID-19: A key role for monocytes and macrophages. Nat. Rev. Immunol. 2020, 20, 355–362. [Google Scholar] [CrossRef]
- Ferreira, A.C.; Soares, V.C.; de Azevedo-Quintanilha, I.G.; da Silva Gomes Dias, S.; Fintelman-Rodrigues, N.; Sacramento, C.Q.; Mattos, M.; de Freitas, C.S.; Temerozo, J.R.; Teixeira, L.; et al. SARS-CoV-2 engages inflammasome and pyroptosis in human primary monocytes. Cell Death Discov. 2021, 7, 43. [Google Scholar] [CrossRef]
- Rodrigues, T.S.; de Sá, K.S.G.; Ishimoto, A.Y.; Becerra, A.; Oliveira, S.; Almeida, L.; Gonçalves, A.V.; Perucello, D.B.; Andrade, W.A.; Castro, R.; et al. Inflammasomes are activated in response to SARS-CoV-2 infection and are associated with COVID-19 severity in patients. J. Exp. Med. 2020, 218, e20201707. [Google Scholar] [CrossRef]
- Tay, M.Z.; Poh, C.M.; Rénia, L.; MacAry, P.A.; Ng, L.F.P. The trinity of COVID-19: Immunity, inflammation and intervention. Nat. Rev. Immunol. 2020, 20, 363–374. [Google Scholar] [CrossRef] [PubMed]
| Tumor Type | Involved Organs or Tissues | Tumor Risk Factors | Evidence of HERV-K Particles |
|---|---|---|---|
| Melanoma | Skin [70] | Expression/methylation of LINE-1, methylation of HERV-K [75] | HERV-K Gag, Pol, Env, and Rec proteins [76] |
| Teratocarcinoma | mixed germ-line tumors: female sexual gonads, placenta, extraembryonic membranes, and umbilical cord [82,83] | Changing the expression pattern of cytokines and cellular genes due to teratoma-liberated viral proteins [46], deglycosylation of HML-2 GH and Tera-1 [84] | TM 1, HERV-K Env [46] |
| Osteosarcoma | Bones | Dysfunctionality of TE 2 and DNA damages, overexpression of LINE-1/Alu/SVA/HERV-K [85] | Not yet defined |
| Colorectal cancer (CRC) | Colon and rectum | Conversion of NUPR1 into a chromatin protein due to stress condition [86], lower methylation rate of LINE-1 or HERV, EVs 3 [87] | HERV-K Env transcripts class II [88] |
| Breast tumors | Breast [89] | Activation of Ras-dependent ERK 4 1/2 mitogen-activated protein (MAP) kinase pathway and its transformation [90] | HML-2 Env mRNA [38,91], transcripts of Gag and Env genes [91] |
| Ovarian cancer | Ovary [92] | Induction of immunity by HERV-K Env protein and production of T cells, INFγ, and HERV-K-specific CTL 5 [93]. | HERV-K transcripts, Env protein, and active reverse transcriptase [93] |
| Prostate cancer | Prostate [94] | Expression of the NGO-Pr-54 region (a Gag-related antigen) [95], encoding chromosome 22q11.23 by HERV-K Gag proteins [96], overexpression of HERV-K_22q11.23 and accessory Np9 [97], elevated level of interferon-γ and expression of HERV-K Gag [98], chromosomal translocation and amplification, abnormal up-regulation of ETS factors [99], transcriptional activity of HERV-K LTRs [100] | HERV-K Gag and Gag RNA [101], HERV-K Env [98], |
| Atypical teratoid rhabdoid tumors (AT/RT) | Central nervous system (brain) [102] | Deletion of the SMARCB1 gene [103] | HERV-K Env [103] |
| Non-Hodgkin lymphoma (NHL) | Lymphoid tissue [104] | HERV-K expression and its impact in severity or recurrence of NHL [105] | HERV-K DNA, RNA, and proteins [105] |
| Types of Therapy | HERV-K Target | Immune Response | Impact on Tumor |
|---|---|---|---|
| MVA-HKcon vaccine [157] | Gag [157] | Inducing the T cell immune responses, knocking out the cells positive to HERV-K [157] | Suppressing growth of the tumor and reducing pulmonary metastases [157] |
| Active immunization via engineered adenovirus type 5 and 19a/64 carrying ISD-mutant HERV-K Gag and Env [158] | Gag and Env [158] | Rapid antibody T cell responses in mice, immune tolerance breakdown in non-human primates [158] | Suppressing growth of tumors [158] |
| Anti-HERV-K Env monoclonal antibodies [144] | Env [144] | mAbs 1 Env and T cell immune response in breast cancer, high number of cell surface molecules in human breast cancer that target the B cell response [144] | mAbs: generally restrained development of cancerous cells and promoted apoptosis of them in in vitro conditions [144]. 6H5 mAb: diminished expansion of xenograft tumors due to its cytotoxic profile [159]. GN_mAb_EnvK-01: treatment of HERV-K-associated diseases, such as ALS [160] |
| HERV-K Env inhibitors, such as CuSo4 [161] | Env [161] | Not yet defined | Higher concentration of CuSo4 and restriction of tumor growth [161] |
| HERV-K reverse transcriptase inhibitors, such as ABC 2 [162] | RT 3 [162] | Accelerating aging and cell death, affecting the genes responsible for remodeling of chromatin [162] | Declining cell growth/spread/ invasiveness of tumoral cells in prostate cancer [162] |
| CRISPR 4/Cas9 technology | Env | HERV-K HML-2 knock-out, regression of Ras protein [103], regulating RB and Cyclin B1 proteins, declining the cell multiplication, and inducing apoptosis in vitro [163] | Suppressing growth, invasiveness, and dissemination of ovarian tumors [163] |
| Anti-HIV-1 RT drugs [105] | Expression of HERV-K in general [105] | Not yet defined | Preventing HERV-K expression (such as lamivudine, zidovudine, etc.), Inhibiting HERV-K expression (Atazanavir) [105] |
Anti-coronavirus drugs
| Replication of SARS-CoV-2 RNA [165] and expression of HERV-K | Inhibiting inflammatory responses [166] | Not defined yet |
| HERV-K expression [164] | Inhibiting inflammatory responses [164] | Not defined yet |
| HERV-K expression [164] | Targeting TNF and ceasing the inflammatory responses [167] | Not defined yet |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Hosseiniporgham, S.; Sechi, L.A. Anti-HERV-K Drugs and Vaccines, Possible Therapies against Tumors. Vaccines 2023, 11, 751. https://doi.org/10.3390/vaccines11040751
Hosseiniporgham S, Sechi LA. Anti-HERV-K Drugs and Vaccines, Possible Therapies against Tumors. Vaccines. 2023; 11(4):751. https://doi.org/10.3390/vaccines11040751
Chicago/Turabian StyleHosseiniporgham, Sepideh, and Leonardo Antonio Sechi. 2023. "Anti-HERV-K Drugs and Vaccines, Possible Therapies against Tumors" Vaccines 11, no. 4: 751. https://doi.org/10.3390/vaccines11040751
APA StyleHosseiniporgham, S., & Sechi, L. A. (2023). Anti-HERV-K Drugs and Vaccines, Possible Therapies against Tumors. Vaccines, 11(4), 751. https://doi.org/10.3390/vaccines11040751








