Potential for a Plant-Made SARS-CoV-2 Neutralizing Monoclonal Antibody as a Synergetic Cocktail Component
Abstract
:1. Introduction
2. Methods and Materials
2.1. Plant Expression of CA1 and CB6
2.2. mAb Extraction and Purification
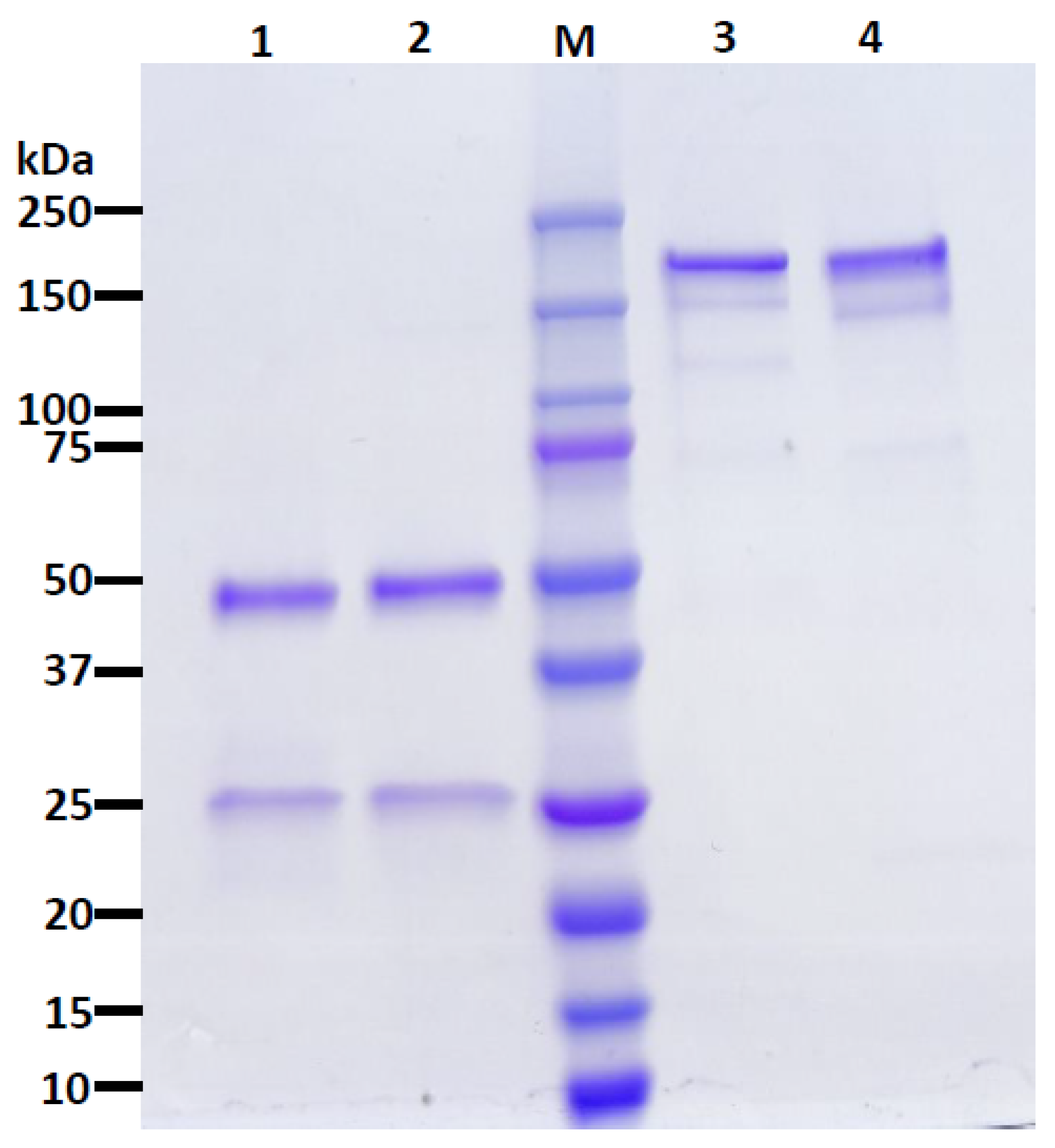
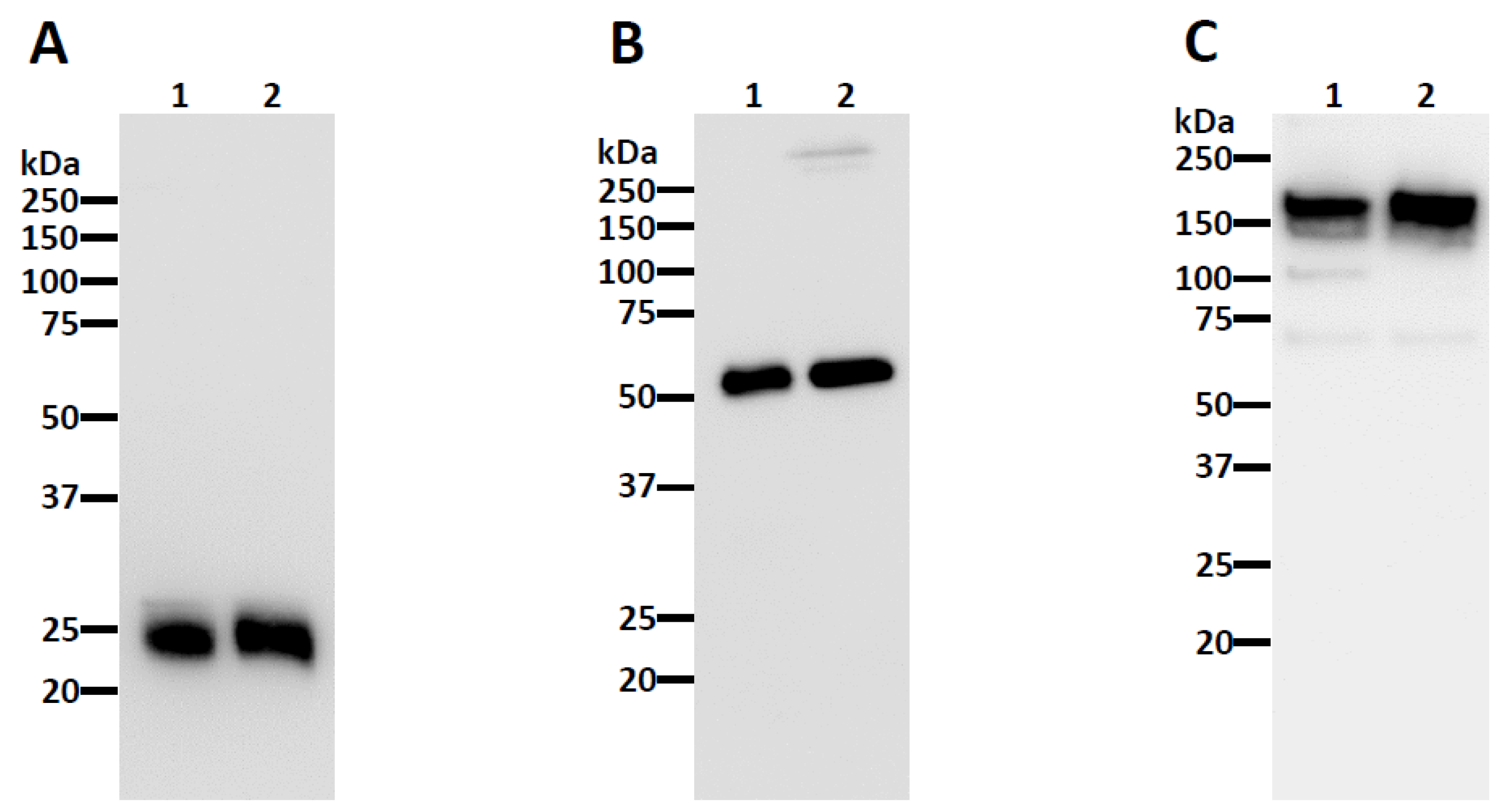
2.3. SDS-PAGE and Western Blotting
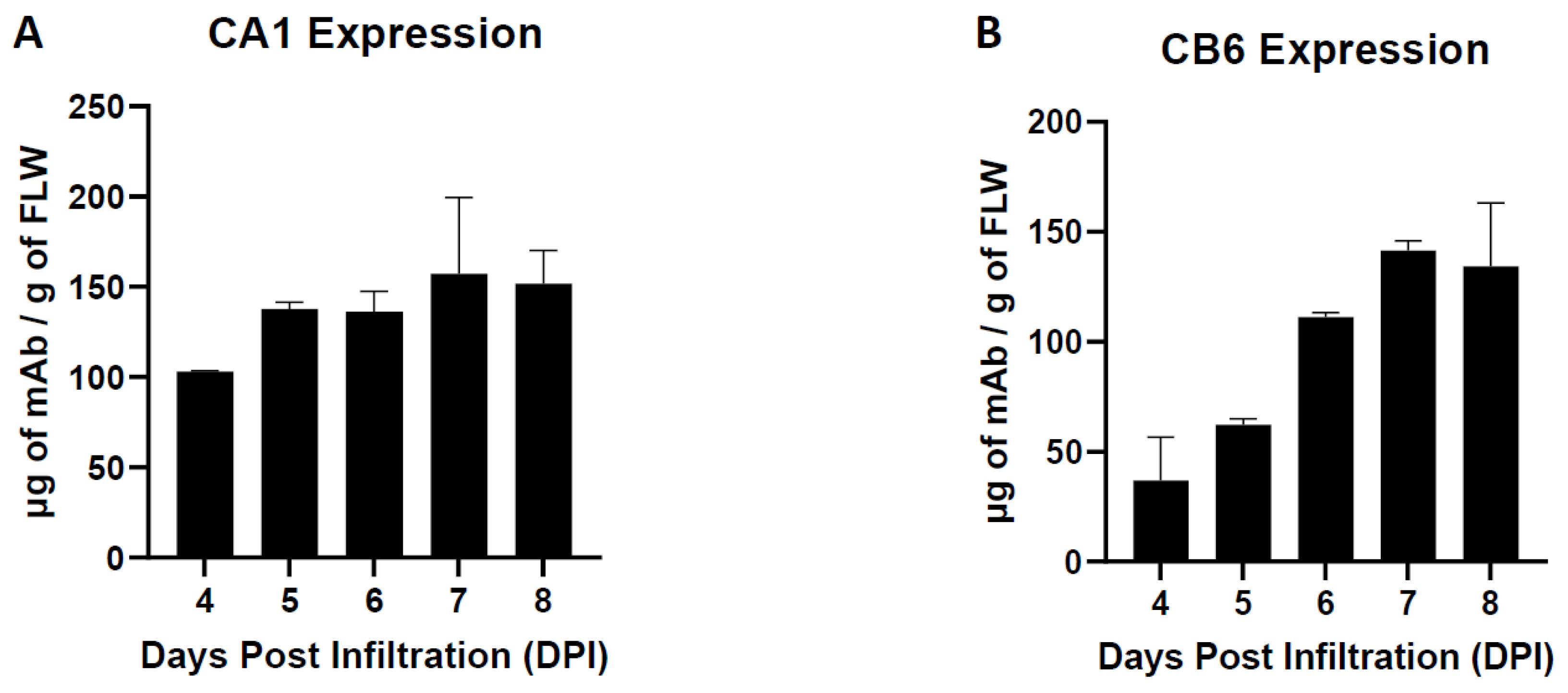
2.4. Temporal Expression ELISA
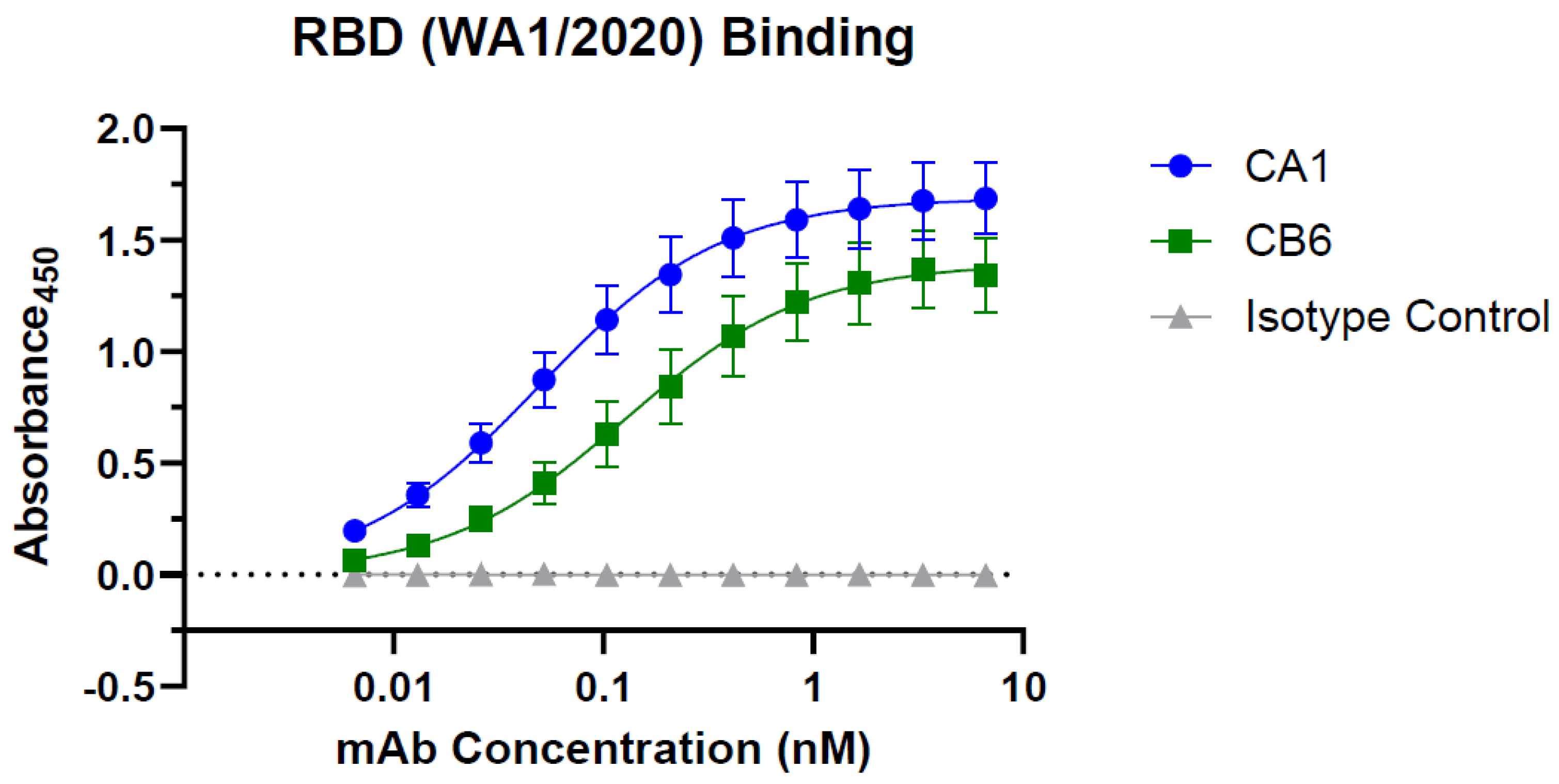
2.5. RBD-Binding ELISA
2.6. Competitive Sandwich ELISAs
2.7. Viruses and Cells
2.8. Focus-Forming Assay
2.9. Neutralization Synergy Analysis
2.10. Generation of Monoclonal Antibodies against RBD
2.11. Indirect ELISA for Hybridoma Screening
3. Results
3.1. Neutralizing Monoclonal Antibody (nAb) Expression in Nicotiana benthamiana
3.2. Plant-Made CA1 and CB6 Recognize SARS-CoV-2 RBD and Neutralize Multiple Variants
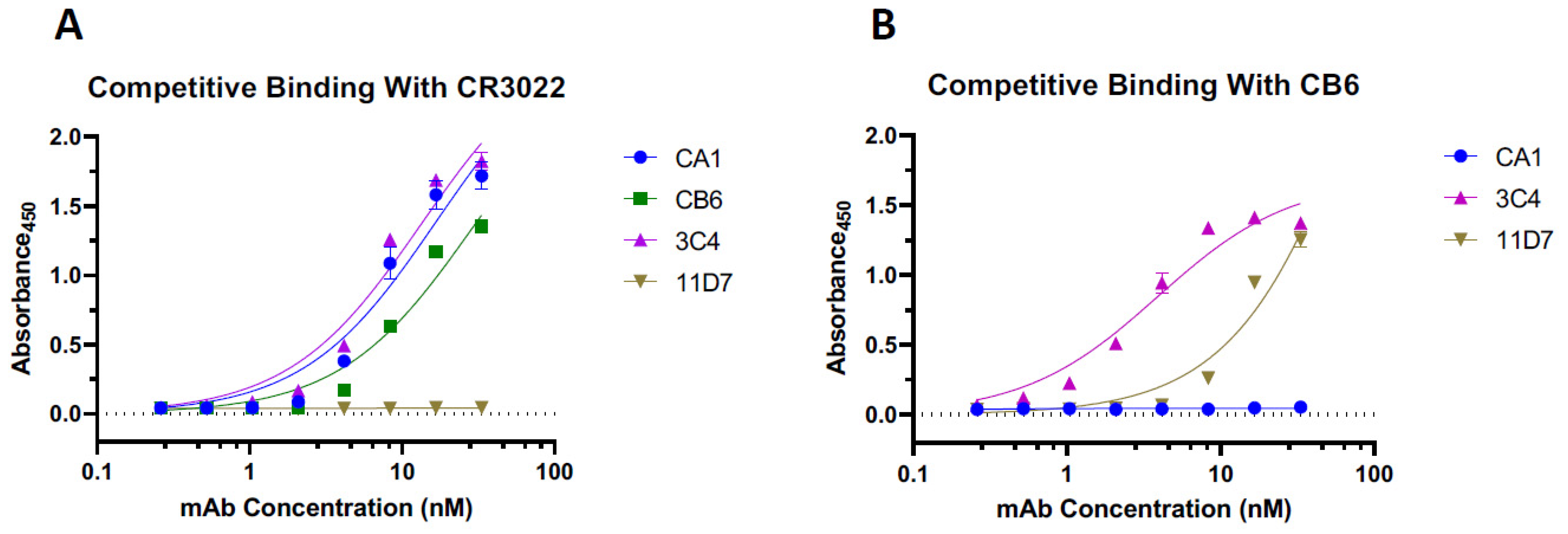
3.3. Plant-Made CB6 Has Synergy with Other Anti-RBD mAbs in Neutralizing SARS-CoV-2
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Lu, R.; Zhao, X.; Li, J.; Niu, P.; Yang, B.; Wu, H.; Wang, W.; Song, H.; Huang, B.; Zhu, N.; et al. Genomic characterisation and epidemiology of 2019 novel coronavirus: Implications for virus origins and receptor binding. Lancet 2020, 395, 565–574. [Google Scholar] [CrossRef] [Green Version]
- Singh, R.; Kang, A.; Luo, X.; Jeyanathan, M.; Gillgrass, A.; Afkhami, S.; Xing, Z. COVID-19: Current knowledge in clinical features, immunological responses, and vaccine development. FASEB J. 2021, 35, e21409. [Google Scholar] [CrossRef] [PubMed]
- Sahly, H.M.; Baden, L.R.; Essink, B.; Doblecki-Lewis, S.; Martin, J.M.; Anderson, E.J.; Campbell, T.B.; Clark, J.; Jackson, L.A.; Fichtenbaum, C.J.; et al. Efficacy of the mRNA-1273 SARS-CoV-2 Vaccine at Completion of Blinded Phase. N. Engl. J. Med. 2021, 385, 1774–1785. [Google Scholar] [CrossRef]
- Thomas, S.J.; Moreira, E.D., Jr.; Kitchin, N.; Absalon, J.; Gurtman, A.; Lockhart, S.; Perez, J.L.; Pérez Marc, G.; Polack, F.P.; Zerbini, C.; et al. Safety and Efficacy of the BNT162b2 mRNA Covid-19 Vaccine through 6 Months. N. Engl. J. Med. 2021, 385, 1761–1773. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.; Zhang, Y.; Wu, L.; Niu, S.; Song, C.; Zhang, Z.; Lu, G.; Qiao, C.; Hu, Y.; Yuen, K.; et al. Structural and functional basis of SARS-CoV-2 entry by using human ACE2. Cell 2020, 181, 894–904.e9. [Google Scholar] [CrossRef]
- Hoffmann, M.; Kleine-Weber, H.; Schroeder, S.; Krüger, N.; Herrler, T.; Erichsen, S.; Schiergens, T.S.; Herrler, G.; Wu, N.; Nitsche, A.; et al. SARS-CoV-2 Cell Entry Depends on ACE2 and TMPRSS2 and Is Blocked by a Clinically Proven Protease Inhibitor. Cell 2020, 181, 271–280.e8. [Google Scholar] [CrossRef]
- Walls, A.C.; Park, Y.; Tortorici, M.A.; Wall, A.; McGuire, A.T.; Veesler, D. Structure, Function, and Antigenicity of the SARS-CoV-2 Spike Glycoprotein. Cell 2020, 181, 281–292.e6. [Google Scholar] [CrossRef]
- Ke, Z.; Oton, J.; Qu, K.; Cortese, M.; Zila, V.; McKeane, L.; Nakane, T.; Zivanov, J.; Neufeldt, C.J.; Cerikan, B.; et al. Structures and distributions of SARS-CoV-2 spike proteins on intact virions. Nature 2020, 588, 498–502. [Google Scholar] [CrossRef]
- Pramanick, I.; Sengupta, N.; Mishra, S.; Pandey, S.; Girish, N.; Das, A.; Dutta, S. Conformational flexibility and structural variability of SARS-CoV2 S protein. Structure 2021, 29, 834–845. [Google Scholar] [CrossRef]
- Cai, Y.; Zhang, J.; Xiao, T.; Peng, H.; Sterling, S.M.; Walsh, R.M., Jr.; Rawson, S.; Rits-Volloch, S.; Chen, B. Distinct conformational states of SARS-CoV-2 spike protein. Science 2020, 369, 1586–1592. [Google Scholar] [CrossRef]
- Castelli, M.S.; McGonigle, P.; Hornby, P.J. The pharmacology and therapeutic applications of monoclonal antibodies. Pharmacol. Res. Perspect. 2019, 7, e00535. [Google Scholar] [CrossRef] [PubMed]
- Hansen, J.; Baum, A.; Pascal, K.E.; Russo, V.; Giordano, S.; Wloga, E.; Fulton, B.O.; Yan, Y.; Koon, K.; Patel, K.; et al. Studies in humanized mice and convalescent humans yield a SARS-CoV-2 antibody cocktail. Science 2020, 369, 1010–1014. [Google Scholar] [CrossRef] [PubMed]
- Rogers, T.F.; Zhao, F.; Huang, D.; Beutler, N.; Burns, A.; He, W.; Limbo, O.; Smith, C.; Song, G.; Woehl, J.; et al. Isolation of potent SARS-CoV-2 neutralizing antibodies and protection from disease in a small animal model. Science 2020, 369, 956–963. [Google Scholar] [CrossRef] [PubMed]
- Wu, Y.; Wang, F.; Shen, C.; Peng, W.; Li, D.; Zhao, C.; Li, Z.; Li, S.; Bi, Y.; Yang, Y.; et al. A noncompeting pair of human neutralizing antibodies block COVID-19 virus binding to its receptor ACE2. Science 2020, 368, 1274–1278. [Google Scholar] [CrossRef]
- Shi, R.; Shan, C.; Duan, X.; Chen, Z.; Lie, P.; Song, J.; Song, T.; Bi, X.; Han, C.; Wu, L.; et al. A human neutralizing antibody targets the receptor-binding site of SARS-CoV-2. Nature 2020, 584, 120–124. [Google Scholar] [CrossRef]
- Dussupt, V.; Sankhala, R.S.; Mendez-Rivera, L.; Townsley, S.M.; Schmidt, F.; Wieczorek, L.; Lal, K.G.; Donofrio, G.C.; Tran, U.; Jackson, N.D.; et al. Low-dose in vivo protection and neutralization across SARS-CoV-2 variants by monoclonal antibody combinations. Nat. Immunol. 2021, 22, 1503–1514. [Google Scholar] [CrossRef]
- Tuccori, M.; Ferraro, S.; Convertino, I.; Cappello, E.; Valdiserra, G.; Blandizzi, C.; Maggi, F.; Focosi, D. Anti-SARS-CoV-2 neutralizing monoclonal antibodies: Clinical pipeline. MAbs 2020, 12, 1854149. [Google Scholar] [CrossRef]
- Deshpande, A.; Harris, B.D.; Martinez-Sobrido, L.; Kobie, J.J.; Walter, M.R. Epitope Classification and RBD Binding Properties of Neutralizing Antibodies Against SARS-CoV-2 Variants of Concern. Front. Immunol. 2021, 12, 2185. [Google Scholar] [CrossRef]
- Barnes, C.O.; Jette, C.A.; Abernathy, M.E.; Dam, K.A.; Esswein, S.R.; Gristick, H.B.; Malyutin, A.G.; Sharaf, N.G.; Huey-Tubman, K.E.; Lee, Y.E.; et al. SARS-CoV-2 neutralizing antibody structures inform therapeutic strategies. Nature 2020, 588, 682–687. [Google Scholar] [CrossRef]
- Greaney, A.J.; Starr, T.N.; Barnes, C.O.; Weisblum, Y.; Schmidt, F.; Caskey, M.; Gaebler, C.; Cho, A.; Agudel, M.; Finkin, S.; et al. Mapping mutations to the SARS-CoV-2 RBD that escape binding by different classes of antibodies. Nat. Commun. 2021, 12, 1–14. [Google Scholar] [CrossRef]
- Starr, T.N.; Greaney, A.J.; Hilton, S.K.; Ellis, D.; Crawford, K.H.D.; Dingens, A.S.; Navarro, M.J.; Bowen, J.E.; Tortorici, M.A.; Walls, A.C.; et al. Deep Mutational Scanning of SARS-CoV-2 Receptor Binding Domain Reveals Constraints on Folding and ACE2 Binding. Cell 2020, 182, 1295–1310.e20. [Google Scholar] [CrossRef] [PubMed]
- Kovacech, B.; Fialova, L.; Filipcik, P.; Skrabana, R.; Zilkova, M.; Paulenka-Ivanovova, N.; Kovac, A.; Palova, D.; Rolkova, G.P.; Tomkova, K.; et al. Monoclonal antibodies targeting two immunodominant epitopes on the Spike protein neutralize emerging SARS-CoV-2 variants of concern. eBioMedicine 2022, 76, 103818. [Google Scholar] [CrossRef] [PubMed]
- Hwang, Y.C.; Lu, R.M.; Su, S.C.; Chiang, P.Y.; Ko, S.H.; Ke, F.Y.; Liang, K.H.; Hsieh, T.Y.; Wu, H.C. Monoclonal antibodies for COVID-19 therapy and SARS-CoV-2 detection. J. Biomed. Sci. 2022, 29, 1–50. [Google Scholar] [CrossRef] [PubMed]
- FDA Limits Use of Certain Monoclonal Antibodies to Treat COVID-19 Due to the Omicron Variant. Available online: https://www.fda.gov/news-events/press-announcements/coronavirus-covid-19-update-fda-limits-use-certain-monoclonal-antibodies-treat-covid-19-due-omicron (accessed on 6 April 2022).
- FDA Revokes Emergency Use Authorization for Monoclonal Antibody Bamlanivimab. Available online: https://www.fda.gov/news-events/press-announcements/coronavirus-covid-19-update-fda-revokes-emergency-use-authorization-monoclonal-antibody-bamlanivimab (accessed on 6 April 2022).
- Chen, Q. Development of plant-made monoclonal antibodies against viral infections. Curr. Opin. Virol. 2022, 52, 148–160. [Google Scholar] [CrossRef]
- Loos, A.; Steinkellner, H. Plant glyco-biotechnology on the way to synthetic biology. Front. Plant Sci. 2014, 5, 523. [Google Scholar] [CrossRef] [Green Version]
- Chen, Q. Glycoengineering of plants yields glycoproteins with polysialylation and other defined N-glycoforms. Proc. Natl. Acad. Sci. USA 2016, 113, 9404–9406. [Google Scholar] [CrossRef] [Green Version]
- Mastrangeli, R.; Palinsky, W.; Bierau, H. Glycoengineered antibodies: Towards the next-generation of immunotherapeutics. Glycobiology 2018, 29, 199–210. [Google Scholar] [CrossRef] [Green Version]
- Dent, M.; Hurtado, J.; Paul, A.M.; Sun, H.; Lai, H.; Yang, M.; Esqueda, A.; Bai, F.; Steinkellner, H.; Chen, Q. Plant-produced anti-dengue virus monoclonal antibodies exhibit reduced antibody-dependent enhancement of infection activity. J. Gen. Virol. 2016, 97, 3280–3290. [Google Scholar] [CrossRef] [Green Version]
- Chen, Q.; Davis, K.R. The potential of plants as a system for the development and production of human biologics [version 1; referees: 3 approved]. F1000Research 2016, 5, 912. [Google Scholar] [CrossRef] [Green Version]
- Sun, L.; Kallolimath, S.; Palt, R.; Stiasny, K.; Mayrhofer, P.; Maresch, D.; Eidenberger, L.; Steinkellner, H. Increased in vitro neutralizing activity of SARS-CoV-2 IgA1 dimers compared to monomers and IgG. Proc. Natl. Acad. Sci. USA 2021, 118, e2107148118. [Google Scholar] [CrossRef]
- Rattanapisit, K.; Bulaon, C.J.I.; Khorattanakulchai, N.; Shanmugaraj, B.; Wangkanont, K.; Phoolcharoen, W. Plant-produced SARS-CoV-2 receptor binding domain (RBD) variants showed differential binding efficiency with anti-spike specific monoclonal antibodies. PLoS ONE 2021, 16, e0253574. [Google Scholar] [CrossRef] [PubMed]
- Rattanapisit, K.; Shanmugaraj, B.; Manopwisedjaroen, S.; Purwono, P.B.; Siriwattananon, K.; Khorattanakulchai, N.; Hanittinan, O.; Boonyayothin, W.; Thitithanyanont, A.; Smith, D.R.; et al. Rapid production of SARS-CoV-2 receptor binding domain (RBD) and spike specific monoclonal antibody CR3022 in Nicotiana benthamiana. Sci. Rep. 2020, 10, 17698. [Google Scholar] [CrossRef] [PubMed]
- Siriwattananon, K.; Manowisedjaroen, S.; Shanmugaraj, B.; Rattanapisit, K.; Phumiamorn, S.; Sapsutthipas, S.; Trisiriwanich, S.; Prompetchara, E.; Ketloy, C.; Buranapraditkun, S.; et al. Plant-Produced Receptor-Binding Domain of SARS-CoV-2 Elicits Potent Neutralizing Responses in Mice and Non-human Primates. Front. Plant Sci. 2021, 12, 847. [Google Scholar] [CrossRef] [PubMed]
- Maharjan, P.M.; Choe, S. Plant-Based COVID-19 Vaccines: Current Status, Design, and Development Strategies of Candidate Vaccines. Vaccines 2021, 9, 992. [Google Scholar] [CrossRef] [PubMed]
- Gobeil, P.; Pillet, S.; Boulay, I.; Séguin, A.; Makarkov, A.; Heizer, G.; Bhutada, K.; Mahmood, A.; Charland, N.; Trépanier, S.; et al. Phase 2 Randomized Trial of an AS03 Adjuvanted Plant-Based Virus-Like Particle Vaccine for Covid-19 in Healthy Adults, Older Adults and Adults with Comorbidities. medRxiv 2021. [Google Scholar] [CrossRef]
- Kallolimath, S.; Sun, S.; Palt, R.; Stiasny, K.; Mayrhofer, P.; Gruber, C.; Kogelmann, B.; Chen, Q.; Steinkellner, H. Highly active engineered IgG3 antibodies against SARS-CoV-2. Proc. Natl. Acad. Sci. USA 2021, 118, e2107249118. [Google Scholar] [CrossRef] [PubMed]
- Diego-Martin, B.; González, B.; Vazquez-Vilar, M.; Selma, S.; Mateos-Fernández, R.; Gianoglio, S.; Fernández-del-Carmen, A.; Orzáez, D. Pilot Production of SARS-CoV-2 Related Proteins in Plants: A Proof of Concept for Rapid Repurposing of Indoor Farms into Biomanufacturing Facilities. Front. Plant Sci. 2020, 11, 2101. [Google Scholar] [CrossRef]
- Health Canada. Health Canada Authorizes Medicago COVID-19 Vaccine for Adults 18 to 64 Years of Age. Available online: https://www.canada.ca/en/health-canada/news/2022/02/health-canada-authorizes-medicago-covid-19-vaccine-for-adults-18-to-64-years-of-age.html (accessed on 6 April 2022).
- Jugler, C.; Sun, H.; Chen, Q. SARS-CoV-2 Spike Protein-Induced Interleukin 6 Signaling Is Blocked by a Plant-Produced Anti-Interleukin 6 Receptor Monoclonal Antibody. Vaccines 2021, 9, 1365. [Google Scholar] [CrossRef]
- He, J.; Lai, H.; Engle, M.; Gorlatov, S.; Gruber, C.; Steinkellner, H.; Diamond, M.S.; Chen, Q. Generation and Analysis of Novel Plant-Derived Antibody-Based Therapeutic Molecules against West Nile Virus. PLoS ONE 2014, 9, e93541. [Google Scholar] [CrossRef]
- Lai, H.; Engle, M.; Fuchs, A.; Keller, T.; Johnson, S.; Gorlatov, S.; Diamond, M.S.; Chen, Q. Monoclonal antibody produced in plants efficiently treats West Nile virus infection in mice. Proc. Natl. Acad. Sci. USA 2010, 107, 2419–2424. [Google Scholar] [CrossRef] [Green Version]
- Hurtado, J.; Acharya, D.; Lai, H.; Sun, H.; Kallolimath, S.; Steinkellner, H.; Bai, F.; Chen, Q. In vitro and in vivo efficacy of anti-chikungunya virus monoclonal antibodies produced in wild-type and glycoengineered Nicotiana benthamiana plants. Plant Biotechnol. J. 2020, 18, 266–273. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Leist, S.R.; Dinnon, K.H., III; Schäfer, A.; Tse, L.V.; Okuda, K.; Hou, Y.J.; West, A.; Edwards, C.E.; Sanders, W.; Fritch, E.J.; et al. A Mouse-Adapted SARS-CoV-2 Induces Acute Lung Injury and Mortality in Standard Laboratory Mice. Cell 2020, 183, 1070–1085.e12. [Google Scholar] [CrossRef] [PubMed]
- Case, J.B.; Bailey, A.L.; Kim, A.S.; Chen, R.E.; Diamond, M.S. Growth, detection, quantification, and inactivation of SARS-CoV-2. Virology 2020, 548, 39–48. [Google Scholar] [CrossRef] [PubMed]
- Ianevski, A.; Giri, A.K.; Aittokallio, T. SynergyFinder 2.0: Visual analytics of multi-drug combination synergies. Nucleic Acids Res. 2020, 48, W488–W493. [Google Scholar] [CrossRef]
- Berenbaum, M.C. What is synergy? Pharmacol. Rev. 1989, 41, 93–141. [Google Scholar]
- Loewe, S. The problem of synergism and antagonism of combined drugs. Arzneimittelforschung 1953, 3, 285–290. [Google Scholar]
- Köhler, G.; Milstein, C. Continuous cultures of fused cells secreting antibody of predefined specificity. Nature 1975, 256, 495–497. [Google Scholar] [CrossRef]
- Chen, Q.; Lai, H.; Hurtado, J.; Stahnke, J.; Leuzinger, K.; Dent, M. Agroinfiltration as an Effective and Scalable Strategy of Gene Delivery for Production of Pharmaceutical Proteins. Adv. Tech. Biol. Med. 2013, 1, 103. [Google Scholar] [CrossRef] [Green Version]
- Leuzinger, K.; Dent, M.; Hurtado, J.; Stahnke, J.; Lai, H.; Zhou, X.; Chen, Q. Efficient agroinfiltration of plants for high-level transient expression of recombinant proteins. J. Vis. Exp. 2013, 77, e50521. [Google Scholar] [CrossRef]
- Shanmugaraj, B.; Rattanapisit, K.; Manopwisedjaroen, S.; Thitithanyanont, A.; Phoolcharoen, W. Monoclonal Antibodies B38 and H4 Produced in Nicotiana benthamiana Neutralize SARS-CoV-2 in vitro. Front. Plant Sci. 2020, 11, 1811. [Google Scholar] [CrossRef]
- Jugler, C.; Joensuu, J.; Chen, Q. Hydrophobin-Protein A Fusion Protein Produced in Plants Efficiently Purified an Anti-West Nile Virus Monoclonal Antibody from Plant Extracts via Aqueous Two-Phase Separation. Int. J. Mol. Sci. 2020, 21, 2140. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- ter Meulen, J.; van den Brink, E.N.; Poon, L.L.M.; Marissen, W.E.; Leung, C.S.W.; Cox, F.; Cheung, C.Y.; Bakker, A.Q.; Bogaards, J.A.; van Deventer, E.; et al. Human Monoclonal Antibody Combination against SARS Coronavirus: Synergy and Coverage of Escape Mutants. PLoS Med. 2006, 3, e237. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tian, X.; Li, C.; Huang, A.; Xia, S.; Lu, S.; Shi, Z.; Lu, L.; Jiang, S.; Yang, Z.; Wu, Y.; et al. Potent binding of 2019 novel coronavirus spike protein by a SARS coronavirus-specific human monoclonal antibody. Emerg. Microbes Infect. 2020, 9, 382–385. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Loo, Y.M.; Mctamney, P.M.; Arends, R.H.; Abram, M.E.; Aksyuk, A.A.; Diallo, S.; Flores, D.J.; Kelly, E.J.; Ren, K.; Roque, R.; et al. The SARS-CoV-2 monoclonal antibody combination, AZD7442, is protective in non-human primates and has an extended half-life in humans. Sci. Transl. Med. 2022, 14, 635. [Google Scholar] [CrossRef] [PubMed]
- Roodink, I.; van Erp, M.; Li, A.; Potter, S.; van Duijnhoven, S.M.J.; Kuipers, A.J.; Kazemier, B.; van Geffen, E.; Hemrika, W.; Berkeveld, B.; et al. Cornering an Ever-Evolving Coronavirus: TATX-03, a fully human synergistic multi-antibody cocktail targeting the SARS-CoV-2 Spike Protein with in vivo efficacy. bioRxiv 2021. [Google Scholar] [CrossRef]
- VanBlargan, L.A.; Errico, J.M.; Halfmann, P.J.; Zost, S.J.; Crowe, J.E., Jr.; Purcell, L.A.; Kawaoka, Y.; Corti, D.; Fremont, D.H.; Diamond, M.S. An infectious SARS-CoV-2 B.1.1.529 Omicron virus escapes neutralization by therapeutic monoclonal antibodies. Nat. Med. 2022, 28, 490–495. [Google Scholar] [CrossRef]
- Planas, D.; Saunders, N.; Maes, P.; Guivel-Benhassine, F.; Planchais, C.; Buchrieser, J.; Bolland, W.H.; Porrot, F.; Staropoli, I.; Lemoine, F.; et al. Considerable escape of SARS-CoV-2 Omicron to antibody neutralization. Nature 2021, 602, 671–675. [Google Scholar] [CrossRef]
- Cao, Y.; Wang, J.; Fian, F.; Xiao, T.; Song, W.; Yisimayi, A.; Huang, W.; Li, Q.; Wang, P.; An, R.; et al. Omicron escapes the majority of existing SARS-CoV-2 neutralizing antibodies. Nature 2021, 602, 657–663. [Google Scholar] [CrossRef]
- Schähs, M.; Strasser, R.; Stadlmann, J.; Kunert, R.; Rademacher, T.; Steinkellner, H. Production of a monoclonal antibody in plants with a humanized N-glycosylation pattern. Plant Biotechnol. J. 2007, 5, 657–663. [Google Scholar] [CrossRef]
- Strasser, R.; Stadlmann, J.; Schähs, M.; Stiegler, G.; Quendler, H.; Mach, L.; Gloössl, J.; Weterings, K.; Pabst, M.; Steinkellner, H. Generation of glyco-engineered Nicotiana benthamiana for the production of monoclonal antibodies with a homogeneous human-like N-glycan structure. Plant Biotechnol. J. 2008, 6, 392–402. [Google Scholar] [CrossRef]
- Lai, H.; He, J.; Hurtado, J.; Stahnke, J.; Fuchs, A.; Mehlhop, E.; Gorlatov, S.; Loos, A.; Diamond, M.S.; Chen, Q. Structural and functional characterization of an anti-West Nile virus monoclonal antibody and its single-chain variant produced in glycoengineered plants. Plant Biotechnol. J. 2014, 12, 1098–1107. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zeitlin, L.; Pettitt, J.; Scully, C.; Bohorova, N.; Kim, D.; Pauly, M.; Hiatt, A.; Ngo, L.; Steinkellner, H.; Whaley, K.J.; et al. Enhanced potency of a fucose-free monoclonal antibody being developed as an Ebola virus immunoprotectant. Proc. Natl. Acad. Sci. USA 2011, 108, 20690–20694. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ma, J.; Drossard, J.; Lewis, D.; Altmann, F.; Boyle, J.; Christou, P.; Cole, T.; Dale, P.; van Dolleweerd, C.J.; Isitt, V.; et al. Regulatory approval and a first-in-human phase I clinical trial of a monoclonal antibody produced in transgenic tobacco plants. Plant Biotechnol. J. 2015, 13, 1106–1120. [Google Scholar] [CrossRef] [PubMed]
- Traynor, K. Taliglucerase alfa approved for Gaucher disease. Am. J. Health-Syst. Pharm. 2012, 69, 1009. [Google Scholar] [CrossRef]
- Chen, Q.; Lai, H. Plant-derived virus-like particles as vaccines. Hum. Vaccines Immunother. 2013, 9, 26–49. [Google Scholar] [CrossRef]
- Paul, M.J.; Thangaraj, H.; Ma, J. Commercialization of new biotechnology: A systematic review of 16 commercial case studies in a novel manufacturing sector. Plant Biotechnol. J. 2015, 13, 1209–1320. [Google Scholar] [CrossRef] [Green Version]
- Hager, K.J.; Pérez Marc, G.; Gobeil, P.; Diaz, R.S.; Heizer, G.; Llapur, C.; Makarkov, A.I.; Vasconcellos, E.; Pillet, S.; Riera, F.; et al. Efficacy and Safety of a Recombinant Plant-Based Adjuvanted Covid-19 Vaccine. N. Engl. J. Med. 2022. [Google Scholar] [CrossRef]
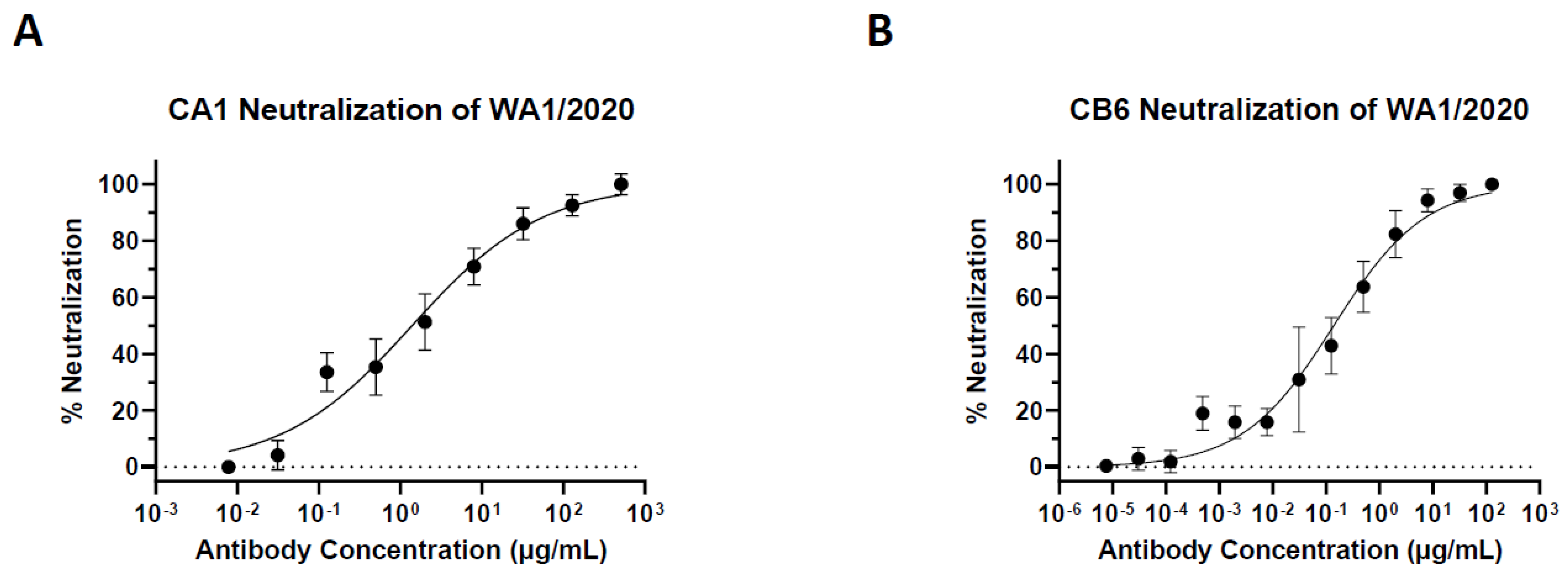
| Strain (Variant) | CA1 (IC50) | CB6 (IC50) |
|---|---|---|
| WA1/2020 | 9.29 nM | 0.93 nM |
| B.1.617.2 (Delta) | 89.87 nM | 0.75 nM |
| Mouse-Adapted (MA10) | 5.15 nM | 7.29 nM |
| Cocktail Combination & Concentration | Observed Percent Neutralization | HSA Predicted Neutralization | Loewe Predicted Neutralization |
|---|---|---|---|
| CB6 + 3C4 (IC20) | 51.66% | 33.97% | 33.94% |
| CB6 + 11D7 (IC20) | 63.18% | 33.97% | 36.06% |
| CB6 + 3C4 +11D7 (IC20) | 60.84% | 33.7% | 36.08% |
| CB6 + 3C4 (IC25) | 58.65% | 33.15% | 44.54% |
| CB6 + 11D7 (IC25) | 78.2% | 33.15% | 42.42% |
| CB6 + 3C4 + 11D7 (IC25) | 77.87% | 34.1% | 41.31% |
| CB6 + 3C4 (IC50) | 67.09% | 57.21% | 61.5% |
| CB6 + 11D7 (IC50) | 87.04% | 57.21% | 57.26% |
| CB6 + 3C4 + 11D7 (IC50) | 88.49% | 47.48% | 65.59% |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Jugler, C.; Sun, H.; Grill, F.; Kibler, K.; Esqueda, A.; Lai, H.; Li, Y.; Lake, D.; Chen, Q. Potential for a Plant-Made SARS-CoV-2 Neutralizing Monoclonal Antibody as a Synergetic Cocktail Component. Vaccines 2022, 10, 772. https://doi.org/10.3390/vaccines10050772
Jugler C, Sun H, Grill F, Kibler K, Esqueda A, Lai H, Li Y, Lake D, Chen Q. Potential for a Plant-Made SARS-CoV-2 Neutralizing Monoclonal Antibody as a Synergetic Cocktail Component. Vaccines. 2022; 10(5):772. https://doi.org/10.3390/vaccines10050772
Chicago/Turabian StyleJugler, Collin, Haiyan Sun, Francisca Grill, Karen Kibler, Adrian Esqueda, Huafang Lai, Yize Li, Douglas Lake, and Qiang Chen. 2022. "Potential for a Plant-Made SARS-CoV-2 Neutralizing Monoclonal Antibody as a Synergetic Cocktail Component" Vaccines 10, no. 5: 772. https://doi.org/10.3390/vaccines10050772
APA StyleJugler, C., Sun, H., Grill, F., Kibler, K., Esqueda, A., Lai, H., Li, Y., Lake, D., & Chen, Q. (2022). Potential for a Plant-Made SARS-CoV-2 Neutralizing Monoclonal Antibody as a Synergetic Cocktail Component. Vaccines, 10(5), 772. https://doi.org/10.3390/vaccines10050772







