A Vaccine Construction against COVID-19-Associated Mucormycosis Contrived with Immunoinformatics-Based Scavenging of Potential Mucoralean Epitopes
Abstract
:1. Introduction
2. Results
2.1. Target Proteins’ Selection
2.2. B-Cell Epitope Prediction
2.3. Protein Modeling, Refinement, and Validation
2.4. Discontinuous B-Cell Epitope Analysis
2.5. Selection of B-Cell Epitopes
2.6. T-Cell Epitope Analysis
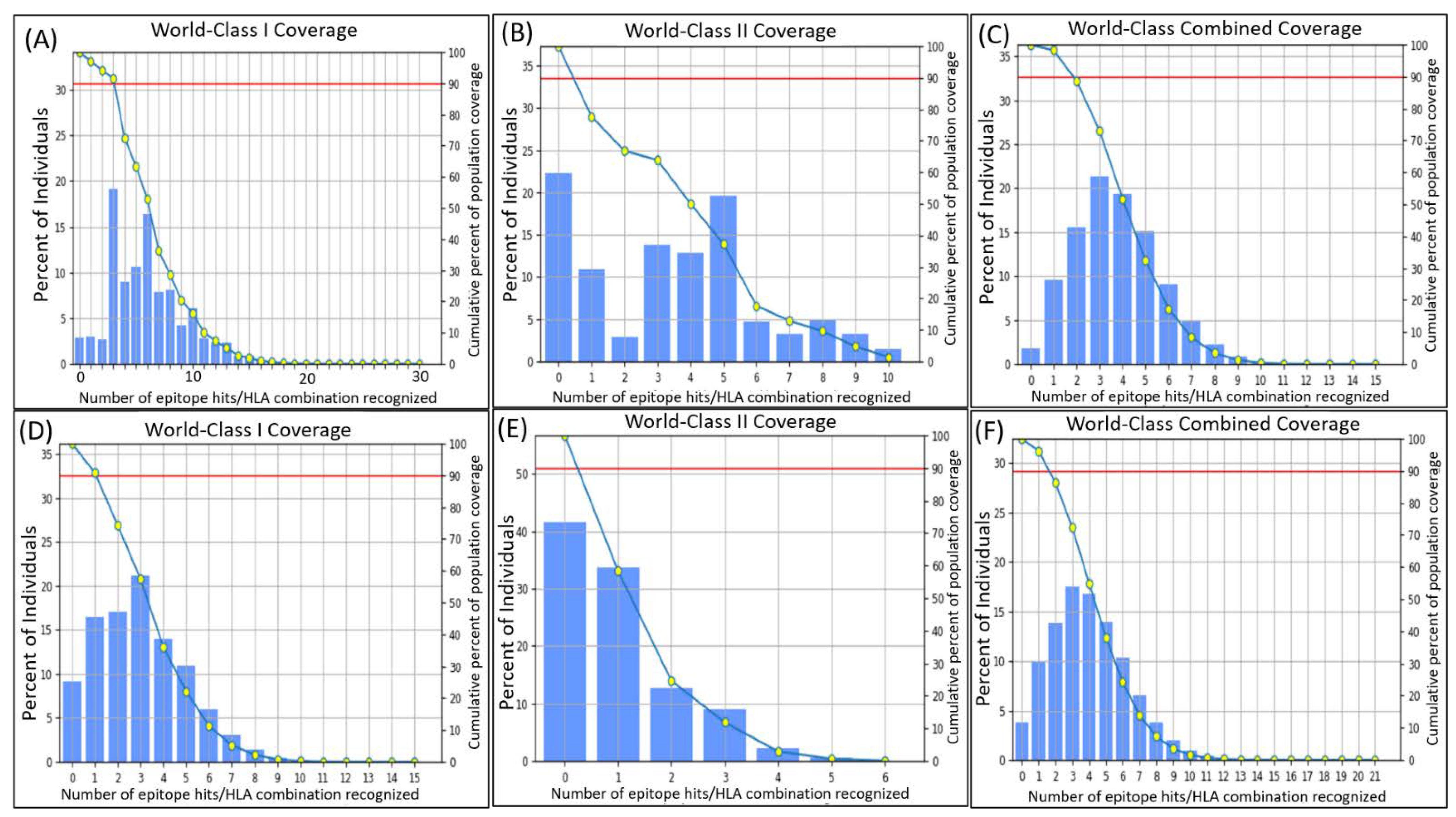
2.7. Population Coverage
2.8. Selection of T-Cell Epitopes
2.9. Final Vaccine Construct
2.10. Vaccine Antigenicity and Allergenicity
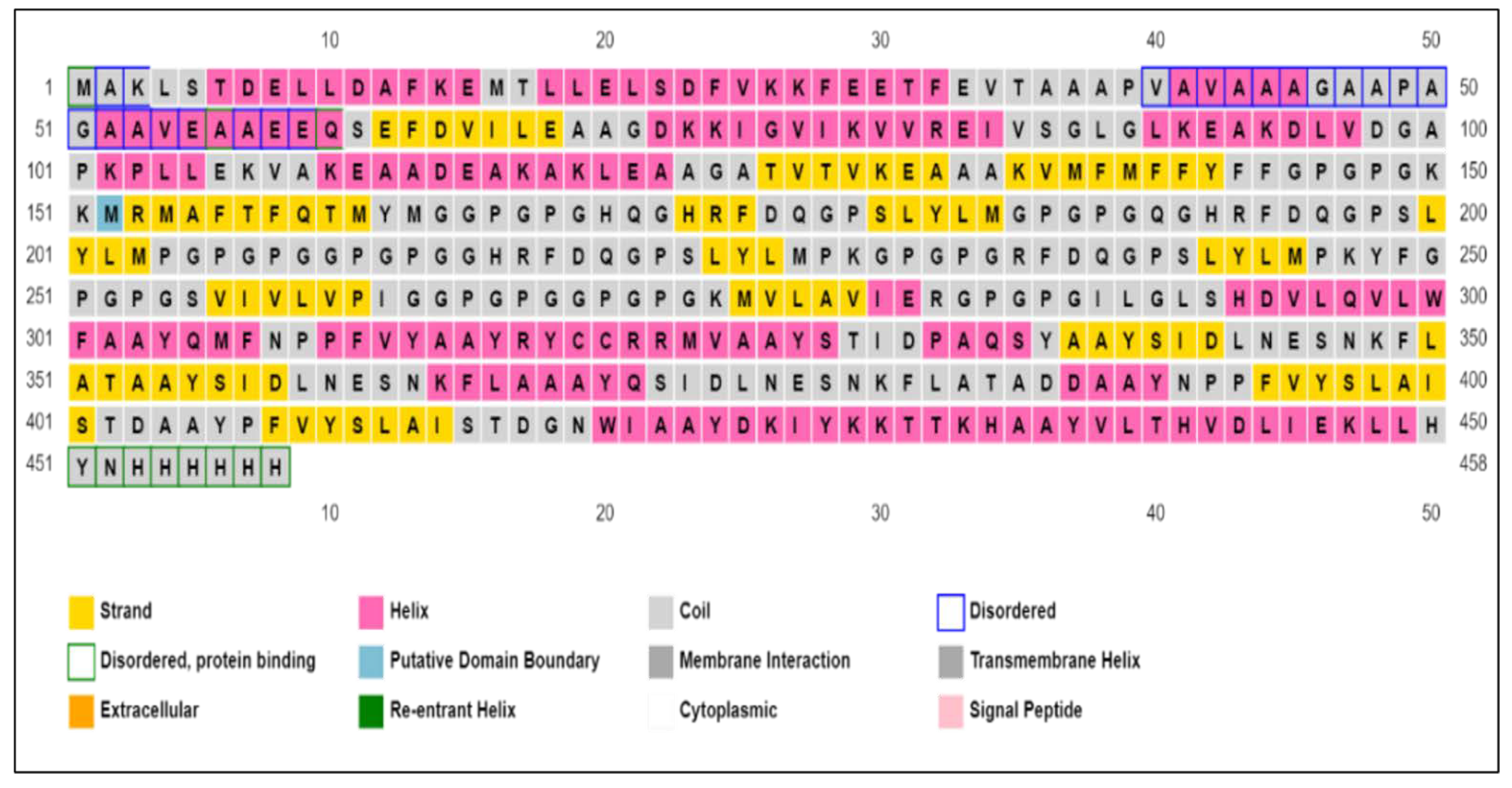
2.11. Prediction of Secondary Structure
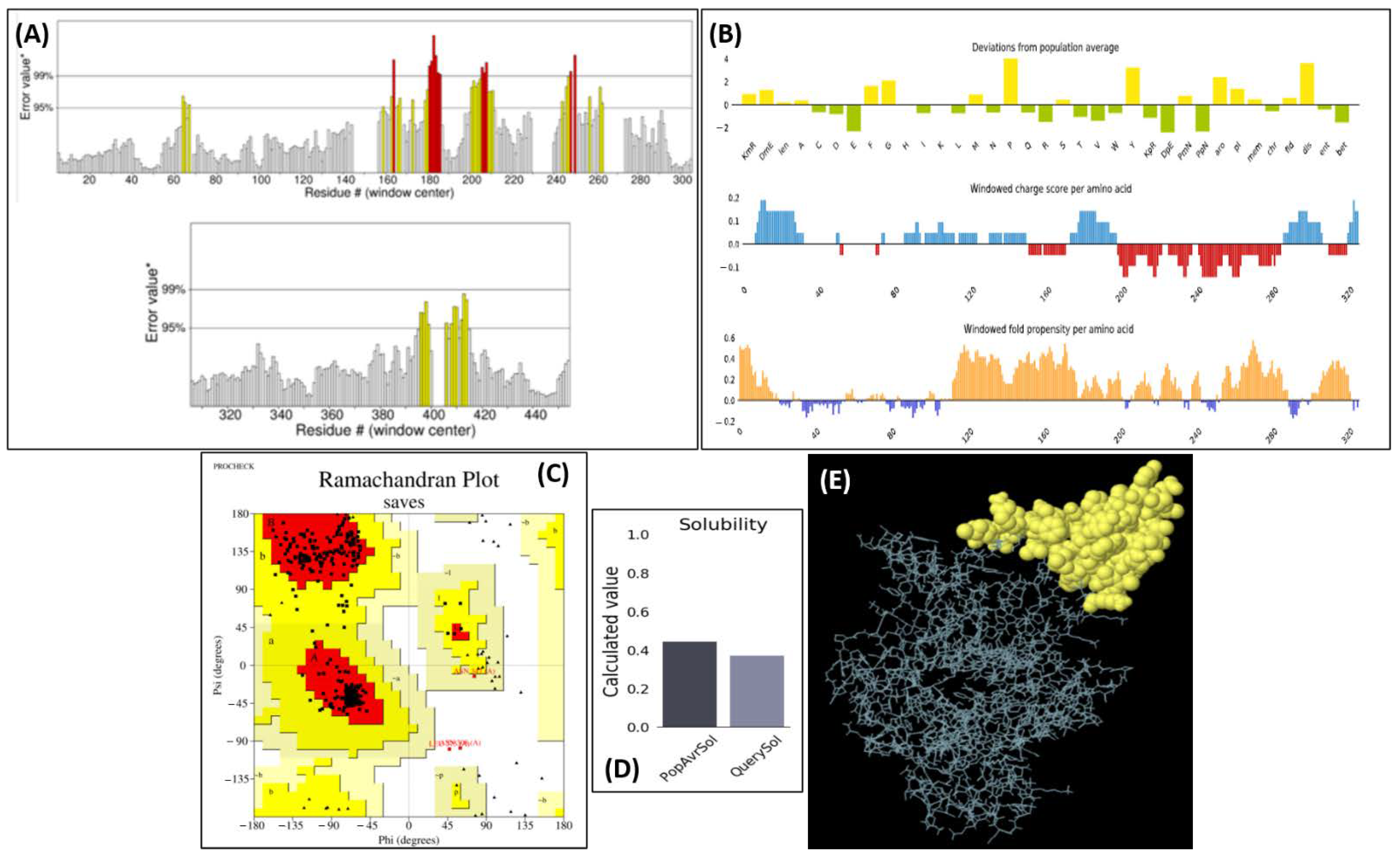
2.12. Tertiary Structure Prediction, Refinement, and Verification
2.13. Physicochemical Properties
2.14. ElliPro and Cleavage Analysis
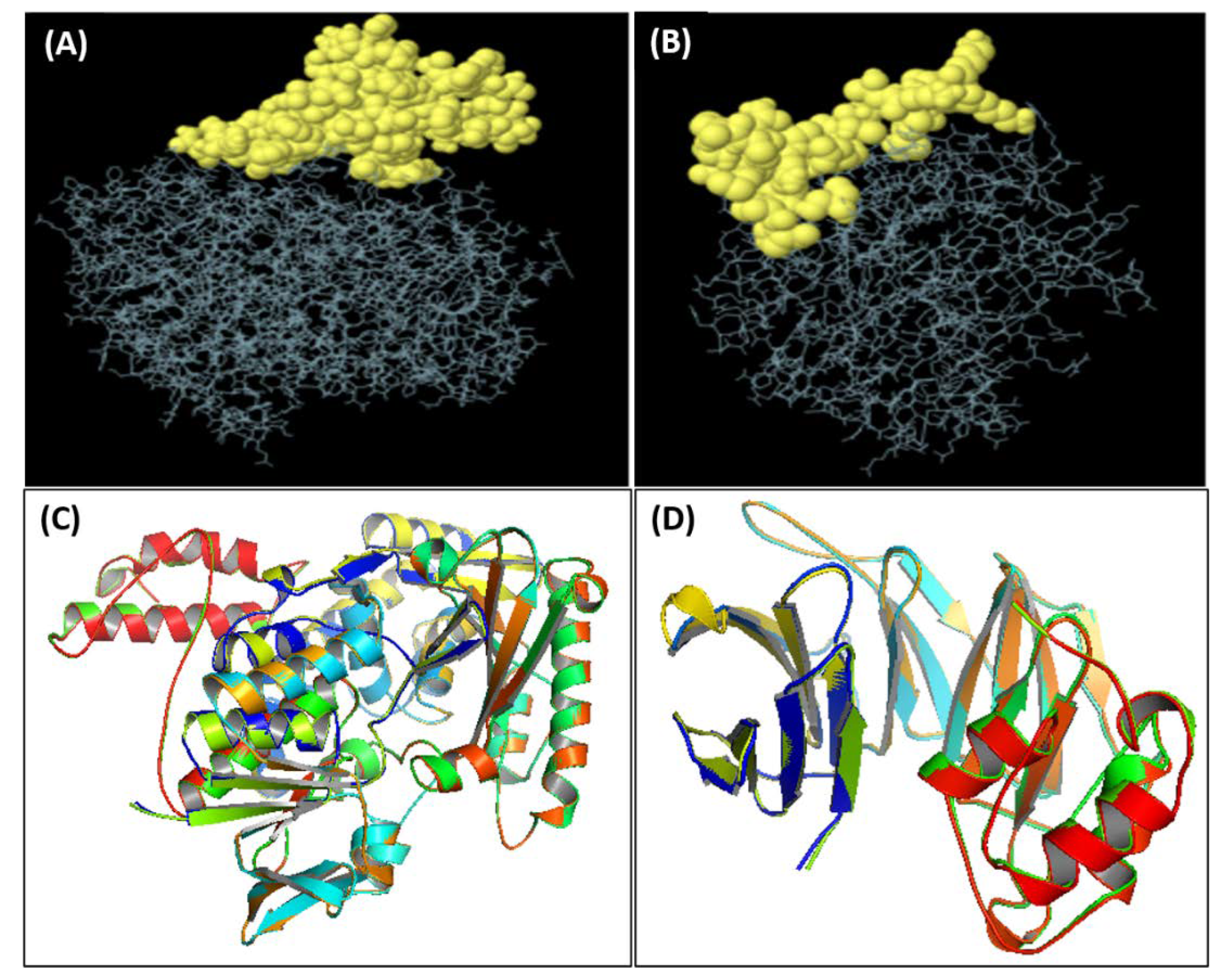
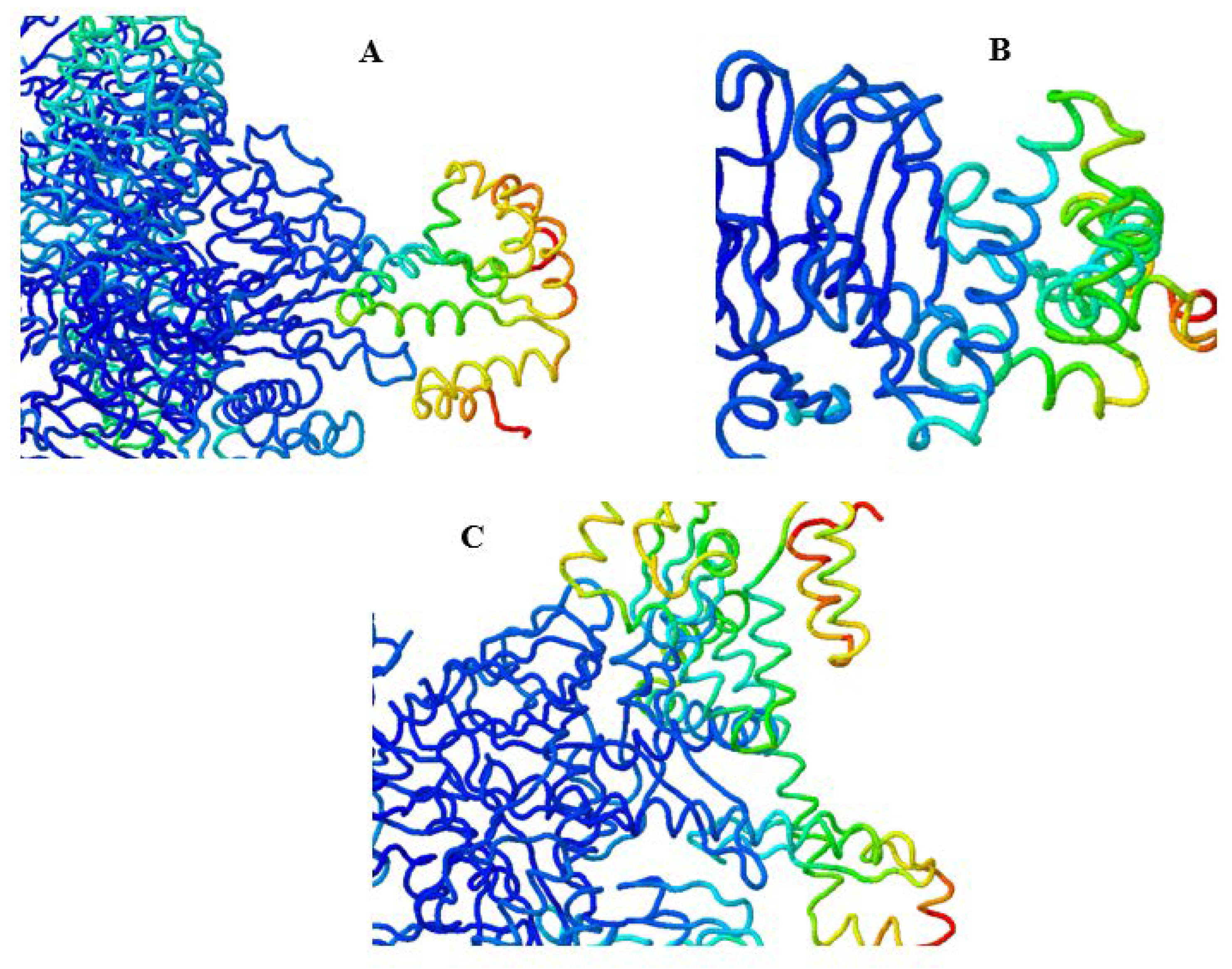
2.15. Molecular Docking
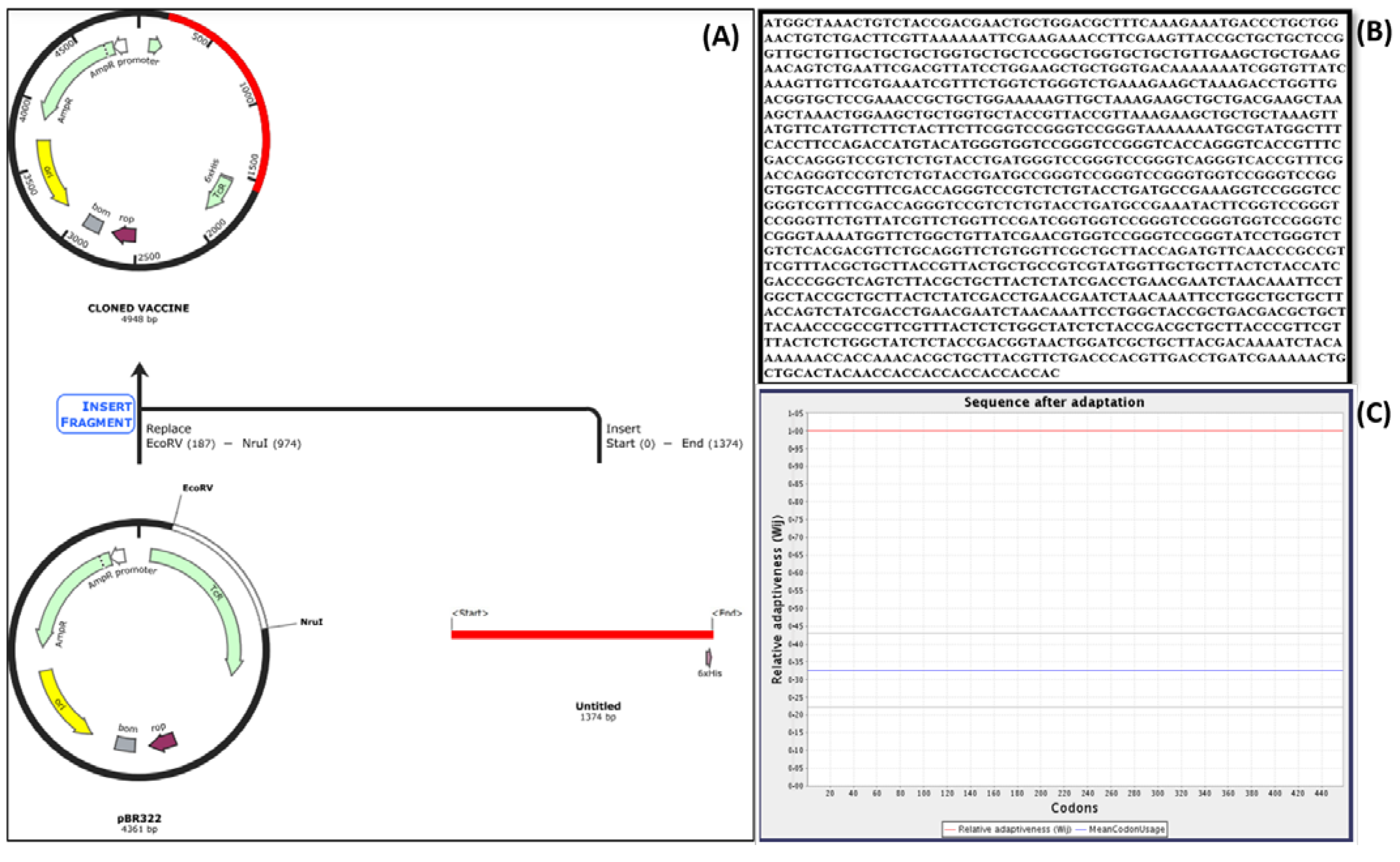
2.16. Codon Optimization
2.17. Cloning and Expression
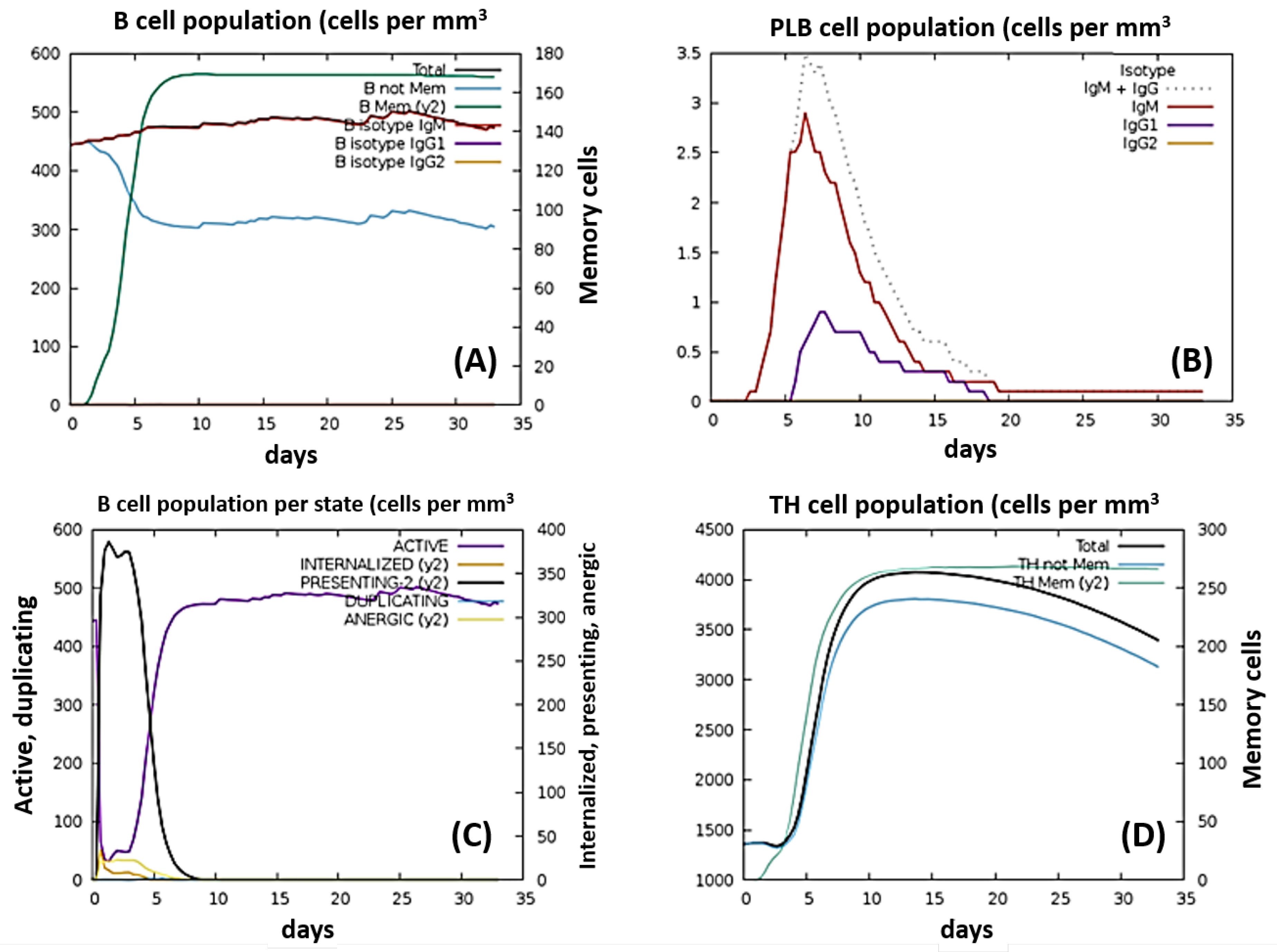
2.18. Immune Response Simulation
3. Discussion
4. Materials and Methods
4.1. Target Proteins’ Selection
4.2. B-Cell Epitope Prediction
4.3. Protein Modeling, Refinement, and Verification
4.4. Discontinuous B-Cell Epitope Analysis
4.5. Selection of B-Cell Epitopes
4.6. T-Cell Epitope Analyses
4.7. Population Coverage
4.8. Selection of T-Cell Epitopes
4.9. In Silico Construction of Vaccine
4.10. Antigenicity and Allergenicity Prediction of the Vaccine
4.11. Vaccine Secondary Structure Prediction
4.12. Vaccine Tertiary Structure Prediction, Refinement, and Verification
4.13. Physicochemical Properties
4.14. ElliPro and Cleavage Analysis of the Vaccine
4.15. Molecular Docking of Vaccine with Immune and General Receptors
4.16. Codon Optimization
4.17. Cloning and Expression Analysis
4.18. Immune Simulations
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Challa, S. Mucormycosis: Pathogenesis and pathology. Curr. Fungal Infect. Rep. 2019, 13, 11–20. [Google Scholar] [CrossRef]
- Spellberg, B.; Maertens, J. Mucormycosis. In Principles and Practice of Transplant Infectious Diseases; Springer: Berlin/Heidelberg, Germany, 2019; pp. 577–589. [Google Scholar]
- Baldin, C.; Ibrahim, A.S. Molecular mechanisms of mucormycosis—The bitter and the sweet. PLoS Pathog. 2017, 13, e1006408. [Google Scholar] [CrossRef]
- Beiglboeck, F.M.; Theofilou, N.E.; Fuchs, M.D.; Wiesli, M.G.; Leiggener, C.; Igelbrink, S.; Augello, M. Managing mucormycosis in diabetic patients: A case report with critical review of the literature. Oral Dis. 2021, 28, 568–576. [Google Scholar] [CrossRef]
- Hasham, K.; Ahmed, N.; Zeshan, B. Circulating microRNAs in oncogenic viral infections: Potential diagnostic biomarkers. SN Appl. Sci. 2020, 2, 1–13. [Google Scholar] [CrossRef]
- Singh, A.K.; Singh, R.; Joshi, S.R.; Misra, A. Mucormycosis in COVID-19: A systematic review of cases reported worldwide and in India. Diabetes Metab. Syndr. Clin. Res. Rev. 2021, 15, 102146. [Google Scholar] [CrossRef]
- Corzo-León, D.E.; Chora-Hernández, L.D.; Rodríguez-Zulueta, A.P.; Walsh, T.J.J.M.M. Diabetes mellitus as the major risk factor for mucormycosis in Mexico: Epidemiology, diagnosis, and outcomes of reported cases. Med. Mycol. 2017, 56, 29–43. [Google Scholar] [CrossRef]
- Jeong, W.; Keighley, C.; Wolfe, R.; Lee, W.L.; Slavin, M.; Kong, D.C.M.; Chen, S.-A.J. The epidemiology and clinical manifestations of mucormycosis: A systematic review and meta-analysis of case reports. Clin. Microbiol. Infect. 2019, 25, 26–34. [Google Scholar] [CrossRef]
- Serris, A.; Danion, F.; Lanternier, F.J.J.O.F. Disease entities in mucormycosis. J. Fungi 2019, 5, 23. [Google Scholar] [CrossRef]
- Ahmadikia, K.; Hashemi, S.J.; Khodavaisy, S.; Getso, M.I.; Alijani, N.; Badali, H.; Mirhendi, H.; Salehi, M.; Tabari, A.; Mohammadi Ardehali, M.J.M. The double-edged sword of systemic corticosteroid therapy in viral pneumonia: A case report and comparative review of influenza-associated mucormycosis versus COVID-19 associated mucormycosis. Mycoses 2021, 64, 798–808. [Google Scholar] [CrossRef]
- Ahmed, N.; Rizvi, A.; Naeem, A.; Saleem, W.; Ahmed, A.; Parveen, S.; Ilyas, M. COVID-19 and public awareness. Prof. Med. J. 2020, 27, 1710–1716. [Google Scholar] [CrossRef]
- Zurl, C.; Hoenigl, M.; Schulz, E.; Hatzl, S.; Gorkiewicz, G.; Krause, R.; Eller, P.; Prattes, J. Autopsy proven pulmonary mucormycosis due to Rhizopus microsporus in a critically ill COVID-19 patient with underlying hematological malignancy. J. Fungi 2021, 7, 88. [Google Scholar] [CrossRef]
- Asano-Mori, Y.J.M.m.j. Diagnosis and Treatment of Mucormycosis in Patients withHematological Malignancies [Translated Article]. Med. Mycol. J. 2017, 58, E97–E105. [Google Scholar] [CrossRef]
- Johnson, A.K.; Ghazarian, Z.; Cendrowski, K.D.; Persichino, J.G. Pulmonary aspergillosis and mucormycosis in a patient with COVID-19. Med. Mycol. Case Rep. 2021, 32, 64–67. [Google Scholar] [CrossRef]
- Sipsas, N.V.; Gamaletsou, M.N.; Anastasopoulou, A.; Kontoyiannis, D.P. Therapy of mucormycosis. J. Fungi 2018, 4, 90. [Google Scholar] [CrossRef]
- Bhadauria, D.; Etta, P.; Chelappan, A.; Gurjar, M.; Kaul, A.; Sharma, R.K.; Gupta, A.; Prasad, N.; Marak, R.S.; Jain, M.; et al. Isolated bilateral renal mucormycosis in apparently immunocompetent patients—A case series from India and review of the literature. Clin. Kidney J. 2018, 11, 769–776. [Google Scholar] [CrossRef]
- Ibrahim, A.S.; Gebremariam, T.; Lin, L.; Luo, G.; Husseiny, M.I.; Skory, C.D.; Fu, Y.; French, S.W.; Edwards, J.E., Jr. The high affinity iron permease is a key virulence factor required for Rhizopus oryzae pathogenesis. Mol. Microbiol. 2010, 77, 587–604. [Google Scholar] [CrossRef]
- Imran, M.; AS, A.; Tauseef, M.; Khan, S.A.; Hudu, S.A.; Abida. Mucormycosis medications: A patent review. Expert Opin. Ther. Pat. 2021, 31, 1059–1074. [Google Scholar] [CrossRef]
- Gupta, N.; Kumar, A. Designing an efficient multi-epitope vaccine against Campylobacter jejuni using immunoinformatics and reverse vaccinology approach. Microb. Pathog. 2020, 147, 104398. [Google Scholar] [CrossRef]
- Elamin Elhasan, L.M.; Hassan, M.B.; Elhassan, R.M.; Abdelrhman, F.A.; Salih, E.A.; Ibrahim, H.A.; Mohamed, A.A.; Osman, H.S.; Khalil, M.S.M.; Alsafi, A.A.; et al. Epitope-based peptide vaccine design against fructose bisphosphate aldolase of Candida glabrata: An immunoinformatics approach. J. Immunol. Res. 2021. [Google Scholar] [CrossRef]
- Chaudhuri, R.; Ramachandran, S. Immunoinformatics as a tool for new antifungal vaccines. In Vaccines for Invasive Fungal Infections; Springer: Berlin/Heidelberg, Germany, 2017; pp. 31–43. [Google Scholar]
- Dhanda, S.K.; Mahajan, S.; Paul, S.; Yan, Z.; Kim, H.; Jespersen, M.C.; Jurtz, V.; Andreatta, M.; Greenbaum, J.A.; Marcatili, P.; et al. IEDB-AR: Immune epitope database—Analysis resource in 2019. Nucleic Acids Res. 2019, 47, W502–W506. [Google Scholar] [CrossRef]
- Hoenigl, M.; Seidel, D.; Carvalho, A.; Rudramurthy, S.M.; Arastehfar, A.; Gangneux, J.-P.; Nasir, N.; Bonifaz, A.; Araiza, J.; Klimko, N.; et al. The emergence of COVID-19 associated mucormycosis: A review of cases from 18 countries. Lancet Microbe 2022. [Google Scholar] [CrossRef]
- Roden, M.M.; Zaoutis, T.E.; Buchanan, W.L.; Knudsen, T.A.; Sarkisova, T.A.; Schaufele, R.L.; Sein, M.; Sein, T.; Chiou, C.C.; Chu, J.H. Epidemiology and outcome of zygomycosis: A review of 929 reported cases. Clin. Infect. Dis. 2005, 41, 634–653. [Google Scholar] [CrossRef]
- Naveed, M.; Tehreem, S.; Arshad, S.; Bukhari, S.A.; Shabbir, M.A.; Essa, R.; Ali, N.; Zaib, S.; Khan, A.; Al-Harrasi, A. Design of a novel multiple epitope-based vaccine: An immunoinformatics approach to combat SARS-CoV-2 strains. J. Infect. Public Health 2021, 14, 938–946. [Google Scholar] [CrossRef]
- Tarang, S.; Kesherwani, V.; LaTendresse, B.; Lindgren, L.; Rocha-Sanchez, S.M.; Weston, M.D. In silico design of a multivalent vaccine against Candida albicans. Sci. Rep. 2020, 10, 1–7. [Google Scholar] [CrossRef]
- Abdelmoneim, A.H.; Mustafa, M.I.; Abdelmageed, M.I.; Murshed, N.S.; Dawoud, E.d.; Ahmed, E.M.; Kamal Eldein, R.M.; Elfadol, N.M.; Sati, A.O.M.; Makhawi, A.M. Immunoinformatics design of multiepitopes peptide-based universal cancer vaccine using matrix metalloproteinase-9 protein as a target. Immunol. Med. 2021, 44, 35–52. [Google Scholar] [CrossRef]
- Abraham Peele, K.; Srihansa, T.; Krupanidhi, S.; Ayyagari, V.S.; Venkateswarulu, T. Design of multi-epitope vaccine candidate against SARS-CoV-2: A in-silico study. J. Biomol. Struct. Dyn. 2021, 39, 3793–3801. [Google Scholar] [CrossRef]
- Yu, C.-S.; Cheng, C.-W.; Su, W.-C.; Chang, K.-C.; Huang, S.-W.; Hwang, J.-K.; Lu, C.-H. CELLO2GO: A web server for protein subCELlular LOcalization prediction with functional gene ontology annotation. PLoS ONE 2014, 9, e99368. [Google Scholar] [CrossRef]
- Zhang, J.; Madden, T.L. PowerBLAST: A new network BLAST application for interactive or automated sequence analysis and annotation. Genome Res. 1997, 7, 649–656. [Google Scholar] [CrossRef]
- Doytchinova, I.A.; Flower, D.R. VaxiJen: A server for prediction of protective antigens, tumour antigens and subunit vaccines. BMC Bioinform. 2007, 8, 1–7. [Google Scholar] [CrossRef]
- Dimitrov, I.; Flower, D.R.; Doytchinova, I. AllerTOP-a server for in silico prediction of allergens. BMC Bioinform. 2013, 14, S4. [Google Scholar] [CrossRef]
- Parker, J.; Guo, D.; Hodges, R. New hydrophilicity scale derived from high-performance liquid chromatography peptide retention data: Correlation of predicted surface residues with antigenicity and X-ray-derived accessible sites. Biochemistry 1986, 25, 5425–5432. [Google Scholar] [CrossRef] [PubMed]
- Kolaskar, A.S.; Tongaonkar, P.C. A semi-empirical method for prediction of antigenic determinants on protein antigens. FEBS Lett. 1990, 276, 172–174. [Google Scholar] [CrossRef]
- Yang, J.; Anishchenko, I.; Park, H.; Peng, Z.; Ovchinnikov, S.; Baker, D. Improved protein structure prediction using predicted interresidue orientations. Proc. Natl. Acad. Sci. USA 2020, 117, 1496–1503. [Google Scholar] [CrossRef] [PubMed]
- Lee, G.R.; Heo, L.; Seok, C. Effective protein model structure refinement by loop modeling and overall relaxation. Proteins: Struct. Funct. Bioinform. 2016, 84, 293–301. [Google Scholar] [CrossRef]
- Laskowski, R.A.; Rullmann, J.A.C.; MacArthur, M.W.; Kaptein, R.; Thornton, J.M. AQUA and PROCHECK-NMR: Programs for checking the quality of protein structures solved by NMR. J. Biomol. NMR 1996, 8, 477–486. [Google Scholar] [CrossRef]
- Colovos, C.; Yeates, T.O. Verification of protein structures: Patterns of nonbonded atomic interactions. Protein Sci. 1993, 2, 1511–1519. [Google Scholar] [CrossRef]
- Luthy, R.; Bowie, J.U.; Eisenberg, D. Assessment of protein models with three-dimensional profiles. Nature 1992, 356, 83–85. [Google Scholar] [CrossRef]
- Ponomarenko, J.; Bui, H.-H.; Li, W.; Fusseder, N.; Bourne, P.E.; Sette, A.; Peters, B. ElliPro: A new structure-based tool for the prediction of antibody epitopes. BMC Bioinform. 2008, 9, 1–8. [Google Scholar] [CrossRef]
- Tenzer, S.; Peters, B.; Bulik, S.; Schoor, O.; Lemmel, C.; Schatz, M.; Kloetzel, P.-M.; Rammensee, H.-G.; Schild, H.; Holzhütter, H.-G. Modeling the MHC class I pathway by combining predictions of proteasomal cleavage, TAP transport and MHC class I binding. Cell. Mol. Life Sci. CMLS 2005, 62, 1025–1037. [Google Scholar] [CrossRef]
- Bui, H.-H.; Sidney, J.; Peters, B.; Sathiamurthy, M.; Sinichi, A.; Purton, K.-A.; Mothé, B.R.; Chisari, F.V.; Watkins, D.I.; Sette, A. Automated generation and evaluation of specific MHC binding predictive tools: ARB matrix applications. Immunogenetics 2005, 57, 304–314. [Google Scholar] [CrossRef]
- Nielsen, M.; Lundegaard, C.; Lund, O.; Keşmir, C. The role of the proteasome in generating cytotoxic T-cell epitopes: Insights obtained from improved predictions of proteasomal cleavage. Immunogenetics 2005, 57, 33–41. [Google Scholar] [CrossRef] [PubMed]
- Bui, H.-H.; Sidney, J.; Dinh, K.; Southwood, S.; Newman, M.J.; Sette, A. Predicting population coverage of T-cell epitope-based diagnostics and vaccines. BMC Bioinform. 2006, 7, 1–5. [Google Scholar] [CrossRef] [PubMed]
- Cheng, J.; Randall, A.Z.; Sweredoski, M.J.; Baldi, P. SCRATCH: A protein structure and structural feature prediction server. Nucleic Acids Res. 2005, 33, W72–W76. [Google Scholar] [CrossRef]
- Gupta, S.; Kapoor, P.; Chaudhary, K.; Gautam, A.; Kumar, R.; Consortium, O.S.D.D.; Raghava, G.P. In silico approach for predicting toxicity of peptides and proteins. PLoS ONE 2013, 8, e73957. [Google Scholar] [CrossRef]
- Buchan, D.W.; Jones, D.T. The PSIPRED protein analysis workbench: 20 years on. Nucleic Acids Res. 2019, 47, W402–W407. [Google Scholar] [CrossRef] [PubMed]
- Jones, D.T.; Cozzetto, D. DISOPRED3: Precise disordered region predictions with annotated protein-binding activity. Bioinformatics 2015, 31, 857–863. [Google Scholar] [CrossRef]
- Nugent, T.; Jones, D.T. Transmembrane protein topology prediction using support vector machines. BMC Bioinform. 2009, 10, 1–11. [Google Scholar] [CrossRef]
- Bryson, K.; Cozzetto, D.; Jones, D.T. Computer-assisted protein domain boundary prediction using the Dom-Pred server. Curr. Protein Pept. Sci. 2007, 8, 181–188. [Google Scholar] [CrossRef]
- Greener, J.G.; Kandathil, S.M.; Jones, D.T. Deep learning extends de novo protein modelling coverage of genomes using iteratively predicted structural constraints. Nat. Commun. 2019, 10, 1–13. [Google Scholar] [CrossRef]
- Kandathil, S.M.; Greener, J.G.; Jones, D.T. Prediction of interresidue contacts with DeepMetaPSICOV in CASP13. Proteins: Struct. Funct. Bioinform. 2019, 87, 1092–1099. [Google Scholar] [CrossRef]
- Anderson, R.J.; Weng, Z.; Campbell, R.K.; Jiang, X. Main-chain conformational tendencies of amino acids. Proteins: Struct. Funct. Bioinform. 2005, 60, 679–689. [Google Scholar] [CrossRef] [PubMed]
- Gasteiger, E.; Hoogland, C.; Gattiker, A.; Wilkins, M.R.; Appel, R.D.; Bairoch, A. Protein identification and analysis tools on the ExPASy server. In The Proteomics Protocols Handbook; Walker, J.M., Ed.; Humana Press: Totowa, NJ, USA, 2005; pp. 571–607. [Google Scholar]
- Hebditch, M.; Carballo-Amador, M.A.; Charonis, S.; Curtis, R.; Warwicker, J. Protein–Sol: A web tool for predicting protein solubility from sequence. Bioinformatics 2017, 33, 3098–3100. [Google Scholar] [CrossRef] [PubMed]
- Burley, S.K.; Bhikadiya, C.; Bi, C.; Bittrich, S.; Chen, L.; Crichlow, G.V.; Christie, C.H.; Dalenberg, K.; Di Costanzo, L.; Duarte, J.M.; et al. RCSB Protein Data Bank: Powerful new tools for exploring 3D structures of biological macromolecules for basic and applied research and education in fundamental biology, biomedicine, biotechnology, bioengineering and energy sciences. Nucleic Acids Res. 2021, 49, D437–D451. [Google Scholar] [CrossRef] [PubMed]
- Desta, I.T.; Porter, K.A.; Xia, B.; Kozakov, D.; Vajda, S. Performance and its limits in rigid body protein-protein docking. Structure 2020, 28, 1071–1081.e1073. [Google Scholar] [CrossRef]
- Janson, G.; Zhang, C.; Prado, M.G.; Paiardini, A. PyMod 2.0: Improvements in protein sequence-structure analysis and homology modeling within PyMOL. Bioinformatics 2017, 33, 444–446. [Google Scholar] [CrossRef]
- Hospital, A.; Andrio, P.; Fenollosa, C.; Cicin-Sain, D.; Orozco, M.; Gelpí, J.L. MDWeb and MDMoby: An integrated web-based platform for molecular dynamics simulations. Bioinformatics 2012, 28, 1278–1279. [Google Scholar] [CrossRef]
- Grote, A.; Hiller, K.; Scheer, M.; Münch, R.; Nörtemann, B.; Hempel, D.C.; Jahn, D. JCat: A novel tool to adapt codon usage of a target gene to its potential expression host. Nucleic Acids Res. 2005, 33, W526–W531. [Google Scholar] [CrossRef]
- Rapin, N.; Lund, O.; Castiglione, F. Immune system simulation online. Bioinformatics 2011, 27, 2013–2014. [Google Scholar] [CrossRef]



| Protein | Antigenicity Score |
|---|---|
| Phytoene dehydrogenase | 0.5642 |
| Hypothetical protein | 0.9140 |
| No. | Start | End | Peptide | Length | Antigenicity | Surface Accessibility | Allergenicity |
|---|---|---|---|---|---|---|---|
| Phytoene dehydrogenase | |||||||
| 1 | 374 | 382 | SVIVLVPIG | 9 | 0.9144 | 1 | Nonallergen |
| 2 | 405 | 413 | KMVLAVIER | 9 | 1.1837 | 1 | Nonallergen |
| 3 | 446 | 459 | ILGLSHDVLQVLWF | 14 | 0.9727 | 1 | Nonallergen |
| Hypothetical protein | |||||||
| 1 | 31 | 41 | DKIYKKTTKH | 10 | 1.055 | 6.293 | Nonallergen |
| 2 | 260 | 274 | VLTHVDLIEKLLHYN | 15 | 1.164 | 2.059 | Nonallergen |
| 3 | 120 | 135 | IQLISPPSKKSKTT | 14 | 1.2688 | 4.217 | Nonallergen |
| Phytoene Dehydrogenase | |
|---|---|
| Epitope | Coverage |
| MHC-I coverage | |
| AAFWVMFMF | 27.24% |
| KIYDRASKY | 41.99% |
| SSISFYWSM | 27.64% |
| MAFTFQTMY | 62.52% |
| STFPVWFWL | 53.56% |
| VMFMFFYFF | 68.08% |
| LTSSSISFY | 64.02% |
| LVYAYHNILL | 59.37% |
| RMAFTFQTM | 37.16% |
| WVMFMFFYF | 50.11% |
| MHC-II coverage | |
| FDQGPSLYL | 44.03% |
| FIYNAPVAK | 16.02% |
| FKTKKMRMA | 22.39% |
| YMGMSPYDA | 18.55% |
| YFKTKKMRM | 36.85% |
| MRMAFTFQT | 60.71% |
| LRCDNNYKV | 6.69% |
| LAVIERRLG | 31.26% |
| FYWSMSTKV | 33.61% |
| FYVNVPSRI | 43.45% |
| Hypothetical Protein | |
| MHC-I coverage | |
| QMFNPPFVY | 66.02% |
| KVYEWDFSR | 45.05% |
| RYCCRRMVL | 49.95% |
| STIDPAQSY | 45.26% |
| YLSLLQAEY | 35.72% |
| MHC-II coverage | |
| AQMFNPPFV | 8.91% |
| FVYSLAIST | 26.79% |
| IDLNESNKF | 33.99% |
| IDPAQSYQL | 17.55% |
| IEKLLHYNP | 7.01% |
| Epitope | Antigenicity Score |
|---|---|
| Phytoene dehydrogenase | |
| VMFMFFYFF | 0.6814 |
| KKMRMAFTFQTMYMG | 0.5753 |
| HQGHRFDQGPSLYLM | 0.9144 |
| QGHRFDQGPSLYLMP | 1.3727 |
| GHRFDQGPSLYLMPK | 1.2474 |
| RFDQGPSLYLMPKYF | 1.8275 |
| Hypothetical protein | |
| QMFNPPFVY | 0.582 |
| RYCCRRMVL | 2.4632 |
| STIDPAQSY | 0.8302 |
| SIDLNESNKFLAT | 1.6163 |
| SIDLNESNKFLA | 1.5324 |
| QSIDLNESNKFLATADD | 1.3854 |
| NPPFVYSLAISTD | 2.0289 |
| PFVYSLAISTDGNWI | 1.8247 |
| Tool Used | VaxiJen v. 2.0 | Scratch | AllerTOP v.2.0 | AllergenFP v. 1.0 |
|---|---|---|---|---|
| Score | 0.7572 (antigenic) | 0.5 (antigenic) | Non-allergen | Nonallergen |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Naveed, M.; Ali, U.; Karobari, M.I.; Ahmed, N.; Mohamed, R.N.; Abullais, S.S.; Kader, M.A.; Marya, A.; Messina, P.; Scardina, G.A. A Vaccine Construction against COVID-19-Associated Mucormycosis Contrived with Immunoinformatics-Based Scavenging of Potential Mucoralean Epitopes. Vaccines 2022, 10, 664. https://doi.org/10.3390/vaccines10050664
Naveed M, Ali U, Karobari MI, Ahmed N, Mohamed RN, Abullais SS, Kader MA, Marya A, Messina P, Scardina GA. A Vaccine Construction against COVID-19-Associated Mucormycosis Contrived with Immunoinformatics-Based Scavenging of Potential Mucoralean Epitopes. Vaccines. 2022; 10(5):664. https://doi.org/10.3390/vaccines10050664
Chicago/Turabian StyleNaveed, Muhammad, Urooj Ali, Mohmed Isaqali Karobari, Naveed Ahmed, Roshan Noor Mohamed, Shahabe Saquib Abullais, Mohammed Abdul Kader, Anand Marya, Pietro Messina, and Giuseppe Alessandro Scardina. 2022. "A Vaccine Construction against COVID-19-Associated Mucormycosis Contrived with Immunoinformatics-Based Scavenging of Potential Mucoralean Epitopes" Vaccines 10, no. 5: 664. https://doi.org/10.3390/vaccines10050664
APA StyleNaveed, M., Ali, U., Karobari, M. I., Ahmed, N., Mohamed, R. N., Abullais, S. S., Kader, M. A., Marya, A., Messina, P., & Scardina, G. A. (2022). A Vaccine Construction against COVID-19-Associated Mucormycosis Contrived with Immunoinformatics-Based Scavenging of Potential Mucoralean Epitopes. Vaccines, 10(5), 664. https://doi.org/10.3390/vaccines10050664











