Comparative Immunogenicity of the Recombinant Receptor-Binding Domain of Protein S SARS-CoV-2 Obtained in Prokaryotic and Mammalian Expression Systems
Abstract
:1. Introduction
2. Materials and Methods
2.1. Creating the pET21-RBD, pVEAL2-RBD, pVEAL2-S, Expression Plasmids
2.2. Production of Recombinant Proteins
2.3. Protein Purification
2.4. Bio-Layer Interferometry
2.5. ELISA Assays
2.6. Animal Immunization
2.7. Neutralization Assay for SARS-CoV-2
2.8. Building of the RBD Model
3. Results
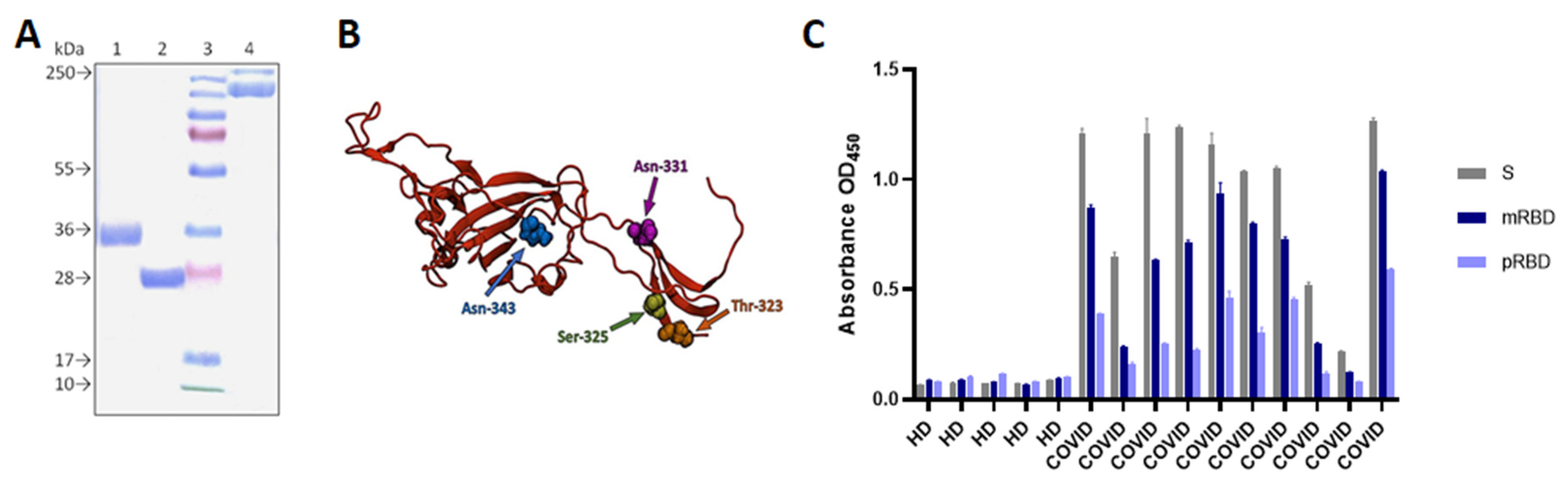
3.1. Construct Design, Expression, and Purification of pRBD, mRBD and S-Trimer
3.2. P. RBD and mRBD Characterization
3.3. PRBD and mRBD Immunogenicity in Mice
4. Discussion
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Wang, N.; Shang, J.; Jiang, S.; Du, L. Subunit Vaccines Against Emerging Pathogenic Human Coronaviruses. Front. Microbiol. 2020, 11, 298. [Google Scholar] [CrossRef]
- Dagotto, G.; Yu, J.; Barouch, D.H. Approaches and Challenges in SARS-CoV-2 Vaccine Development. Cell Host. Microbe 2020, 28, 364–370. [Google Scholar] [CrossRef]
- Du, L.; He, Y.; Jiang, S.; Zheng, B.J. Development of subunit vaccines against severe acute respiratory syndrome. Drugs Today 2008, 44, 63–73. [Google Scholar] [CrossRef]
- Zhang, N.; Jiang, S.; Du, L. Current advancements and potential strategies in the development of MERS-CoV vaccines. Expert Rev. Vaccines 2014, 13, 761–774. [Google Scholar] [CrossRef] [Green Version]
- Baxter, D. Active and passive immunity, vaccine types, excipients and licensing. Occup. Med. 2007, 57, 552–556. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Venters, C.L.; Graham, W.J.; Cassidy, W.M.; Denham, S. Recombivax-HB: Perspectives past, present and future. Expert Rev. Vaccines 2004, 3, 119–129. [Google Scholar] [CrossRef]
- Malik, H.; Khan, F.H.; Ahsan, H. Human papillomavirus: Current status and issues of vaccination. Arch. Virol. 2014, 159, 199–205. [Google Scholar] [CrossRef] [PubMed]
- Vartak, A.; Sucheck, S.J.; Harper, D.M.; Wall, K.A. Recent Advances in Subunit Vaccine Carriers. Vaccines 2016, 4, 12. [Google Scholar] [CrossRef] [Green Version]
- Mekonnen, D.; Mengist, H.M.; Jin, T. SARS-CoV-2 subunit vaccine adjuvants and their signaling pathways. Expert Rev. Vaccines 2021, 1, 1–13. [Google Scholar] [CrossRef]
- Li, T.; Zheng, Q.; Yu, H.; Wu, D.; Xue, W.; Xiong, H.; Huang, X.; Nie, M.; Yue, M.; Rong, R.; et al. SARS-CoV-2 spike produced in insect cells elicits high neutralization titres in non-human primates. Emerg. Microbes Infect. 2020, 9, 1–44. [Google Scholar] [CrossRef] [PubMed]
- Sambrook, J.; Fritsch, E.F.; Maniatis, T. Molecular Cloning: A Laboratory Manual, 2nd ed.; Cold Spring Harbor Laboratory Press, Cold Spring Harbor: Long Island, NY, USA, 1989. [Google Scholar]
- Reed, L.J.; Muench, H. A simple method of estimating fifty per cent endpoints12. Am. J. Epidemiol. 1938, 27, 493–497. [Google Scholar] [CrossRef]
- Shivakumar, D.; Harder, E.; Damm, W.; Friesner, R.A.; Sherman, W. Improving the prediction of absolute solvation free energies using the next generation opls force field. J. Chem. Theory Comput. 2012, 8, 2553–2558. [Google Scholar] [CrossRef] [PubMed]
- Humphrey, W.; Dalke, A.; Schulten, K. VMD: Visual molecular dynamics. J. Mol. Graph. 1996, 14, 33–38. [Google Scholar] [CrossRef]
- Gralinski, L.E.; Menachery, V.D. Return of the coronavirus: 2019-nCoV. Viruses 2020, 12, 135. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Shang, J.; Wan, Y.; Luo, C.; Ye, G.; Geng, Q.; Auerbach, A.; Li, F. Cell entry mechanisms of SARS-CoV-2. Proc. Natl. Acad. Sci. USA 2020, 117, 11727–11734. [Google Scholar] [CrossRef]
- Pallesen, J.; Wang, N.; Corbett, K.S.; Wrapp, D.; Kirchdoerfer, R.N.; Turner, H.L.; Cottrell, C.A.; Becker, M.M.; Wang, L.; Shi, W.; et al. Immunogenicity and structures of a rationally designed prefusion MERS-CoV spike antigen. Proc. Natl. Acad. Sci. USA 2017, 114, E7348–E7357. [Google Scholar] [CrossRef] [Green Version]
- Song, W.; Gui, M.; Wang, X.; Xiang, Y. Cryo-EM structure of the SARS coronavirus spike glycoprotein in complex with its host cell receptor ACE2. PLoS Pathog. 2018, 14, 1–19. [Google Scholar] [CrossRef]
- Wrapp, D.; Wang, N.; Corbett, K.S.; Goldsmith, J.A.; Hsieh, C.L.; Abiona, O.; Graham, B.S.; McLellan, J.S. Cryo-EM structure of the 2019-nCoV spike in the prefusion conformation. Science 2020, 367, 1260–1263. [Google Scholar] [CrossRef] [Green Version]
- Zhou, H.; Chen, Y.; Zhang, S.; Niu, P.; Qin, K.; Jia, W.; Huang, B.; Zhang, S.; Lan, J.; Zhang, L.; et al. Structural definition of a neutralization epitope on the N-terminal domain of MERS-CoV spike glycoprotein. Nat. Commun. 2019, 10, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Almofti, Y.A.; Ali Abd-elrahman, K.; Abd Elgadir Gassmallah, S.; Ahmed Salih, M. Multi Epitopes Vaccine Prediction against Severe Acute Respiratory Syndrome (SARS) Coronavirus Using Immunoinformatics Approaches. Am. J. Microbiol. Res. 2018, 6, 94–114. [Google Scholar] [CrossRef] [Green Version]
- Robbiani, D.F.; Gaebler, C.; Muecksch, F.; Lorenzi, J.C.C.; Wang, Z.; Cho, A.; Agudelo, M.; Barnes, C.O.; Gazumyan, A.; Finkin, S.; et al. Convergent antibody responses to SARS-CoV-2 in convalescent individuals. Nature 2020, 584, 437–442. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.; Zhang, Y.; Wu, L.; Niu, S.; Song, C.; Zhang, Z.; Lu, G.; Qiao, C.; Hu, Y.; Yuen, K.Y.; et al. Structural and Functional Basis of SARS-CoV-2 Entry by Using Human ACE2. Cell 2020, 181, 894–904.e9. [Google Scholar] [CrossRef]
- Huo, J.; Zhao, Y.; Ren, J.; Zhou, D.; Duyvesteyn, H.M.E.; Ginn, H.M.; Carrique, L.; Malinauskas, T.; Ruza, R.R.; Shah, P.N.M.; et al. Neutralization of SARS-CoV-2 by Destruction of the Prefusion Spike. Cell Host. Microbe 2020, 28, 445–454.e6. [Google Scholar] [CrossRef] [PubMed]
- Pinto, D.; Park, Y.J.; Beltramello, M.; Walls, A.C.; Tortorici, M.A.; Bianchi, S.; Jaconi, S.; Culap, K.; Zatta, F.; De Marco, A.; et al. Cross-neutralization of SARS-CoV-2 by a human monoclonal SARS-CoV antibody. Nature 2020, 583, 290–295. [Google Scholar] [CrossRef]
- Rogers, T.F.; Zhao, F.; Huang, D.; Beutler, N.; Burns, A.; He, W.; Limbo, O.; Smith, C.; Song, G.; Woehl, J.; et al. Isolation of potent SARS-CoV-2 neutralizing antibodies and protection from disease in a small animal model. Science 2020, 963, eabc7520. [Google Scholar] [CrossRef]
- Shi, R.; Shan, C.; Duan, X.; Chen, Z.; Liu, P.; Song, J.; Song, T.; Bi, X.; Han, C.; Wu, L.; et al. A human neutralizing antibody targets the receptor-binding site of SARS-CoV-2. Nature 2020, 584, 120–124. [Google Scholar] [CrossRef]
- Wu, Y.; Wang, F.; Shen, C.; Peng, W.; Li, D.; Zhao, C.; Li, Z.; Li, S.; Bi, Y.; Yang, Y.; et al. A noncompeting pair of human neutralizing antibodies block COVID-19 virus binding to its receptor ACE2. Science 2020, 368, 1274–1278. [Google Scholar] [CrossRef] [PubMed]
- Tahir Ul Qamar, M.; Saleem, S.; Ashfaq, U.A.; Bari, A.; Anwar, F.; Alqahtani, S. Epitope-based peptide vaccine design and target site depiction against Middle East Respiratory Syndrome Coronavirus: An immune-informatics study. J. Transl. Med. 2019, 17, 1–14. [Google Scholar] [CrossRef]
- Tai, W.; He, L.; Zhang, X.; Pu, J.; Voronin, D.; Jiang, S.; Zhou, Y.; Du, L. Characterization of the receptor-binding domain (RBD) of 2019 novel coronavirus: Implication for development of RBD protein as a viral attachment inhibitor and vaccine. Cell. Mol. Immunol. 2020, 17, 613–620. [Google Scholar] [CrossRef] [Green Version]
- Zang, J.; Gu, C.; Zhou, B.; Zhang, C.; Yang, Y.; Xu, S.; Bai, L.; Zhang, R.; Deng, Q.; Yuan, Z.; et al. Immunization with the receptor-binding domain of SARS-CoV-2 elicits antibodies cross-neutralizing SARS-CoV-2 and SARS-CoV without antibody-dependent enhancement. Cell Discov. 2020, 6, 4–7. [Google Scholar] [CrossRef]
- Liu, Z.; Xu, W.; Xia, S.; Gu, C.; Wang, X.; Wang, Q.; Zhou, J.; Wu, Y.; Cai, X.; Qu, D.; et al. RBD-Fc-based COVID-19 vaccine candidate induces highly potent SARS-CoV-2 neutralizing antibody response. Signal Transduct. Target. Ther. 2020, 5. [Google Scholar] [CrossRef]
- Sun, C.; Chen, L.; Yang, J.; Luo, C.; Zhang, Y.; Li, J.; Yang, J.; Zhang, J.; Xie, L. SARS-CoV-2 and SARS-CoV Spike-RBD Structure and Receptor Binding Comparison and Potential Implications on Neutralizing Antibody and Vaccine Development. Biorxiv 2020, 1–18. [Google Scholar] [CrossRef] [Green Version]
- Yang, J.; Wang, W.; Chen, Z.; Lu, S.; Yang, F.; Bi, Z.; Bao, L.; Mo, F.; Li, X.; Huang, Y.; et al. A vaccine targeting the RBD of the S protein of SARS-CoV-2 induces protective immunity. Nature 2020, 586, 572–577. [Google Scholar] [CrossRef]
- Malladi, S.K.; Singh, R.; Pandey, S.; Gayathri, S.; Kanjo, K.; Ahmed, S.; Khan, M.S.; Kalita, P.; Girish, N.; Upadhyaya, A.; et al. Design of a highly thermotolerant, immunogenic SARS-CoV-2 spike fragment. J. Biol. Chem. 2021, 296, 100025. [Google Scholar] [CrossRef]
- Tan, T.K.; Rijal, P.; Rahikainen, R.; Keeble, A.H.; Schimanski, L.; Hussain, S.; Harvey, R.; Hayes, J.W.P.; Edwards, J.C.; McLean, R.K.; et al. A COVID-19 vaccine candidate using SpyCatcher multimerization of the SARS-CoV-2 spike protein receptor-binding domain induces potent neutralising antibody responses. Nat. Commun. 2021, 12, 1–16. [Google Scholar] [CrossRef]
- Chen, W.H.; Hotez, P.J.; Bottazzi, M.E. Potential for developing a SARS-CoV receptor-binding domain (RBD) recombinant protein as a heterologous human vaccine against coronavirus infectious disease (COVID)-19. Hum. Vaccines Immunother. 2020, 16, 1239–1242. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bobbala, S.; Hook, S. Is There an Optimal Formulation and Delivery Strategy for Subunit Vaccines? Pharm. Res. 2016, 33, 2078–2097. [Google Scholar] [CrossRef] [PubMed]
- Shajahan, A.; Supekar, N.T.; Gleinich, A.S.; Azadi, P. Deducing the N- And O-glycosylation profile of the spike protein of novel coronavirus SARS-CoV-2. Glycobiology 2020, 30, 981–988. [Google Scholar] [CrossRef] [PubMed]
- Bernardi, A.; Huang, Y.; Harris, B.; Xiong, Y.; Nandi, S.; McDonald, K.A.; Faller, R. Development and simulation of fully glycosylated molecular models of ace2-fc fusion proteins and their interaction with the sars-cov-2 spike protein binding domain. PLoS ONE 2020, 15, e0237295. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Lai, D.; Zhang, H.; Jiang, H.; Tian, X.; Ma, M.; Qi, H.; Meng, Q.; Guo, S.; Wu, Y.; et al. Linear epitopes of SARS-CoV-2 spike protein elicit neutralizing antibodies in COVID-19 patients. Cell. Mol. Immunol. 2020, 17, 1095–1097. [Google Scholar] [CrossRef] [PubMed]
- Yi, Z.; Ling, Y.; Zhang, X.; Chen, J.; Hu, K.; Wang, Y.; Song, W.; Ying, T.; Zhang, R.; Lu, H.Z.; et al. Functional mapping of B-cell linear epitopes of SARS-CoV-2 in COVID-19 convalescent population. Emerg. Microbes Infect. 2020, 9, 1988–1996. [Google Scholar] [CrossRef] [PubMed]

Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Merkuleva, I.A.; Shcherbakov, D.N.; Borgoyakova, M.B.; Shanshin, D.V.; Rudometov, A.P.; Karpenko, L.I.; Belenkaya, S.V.; Isaeva, A.A.; Nesmeyanova, V.S.; Kazachinskaia, E.I.; et al. Comparative Immunogenicity of the Recombinant Receptor-Binding Domain of Protein S SARS-CoV-2 Obtained in Prokaryotic and Mammalian Expression Systems. Vaccines 2022, 10, 96. https://doi.org/10.3390/vaccines10010096
Merkuleva IA, Shcherbakov DN, Borgoyakova MB, Shanshin DV, Rudometov AP, Karpenko LI, Belenkaya SV, Isaeva AA, Nesmeyanova VS, Kazachinskaia EI, et al. Comparative Immunogenicity of the Recombinant Receptor-Binding Domain of Protein S SARS-CoV-2 Obtained in Prokaryotic and Mammalian Expression Systems. Vaccines. 2022; 10(1):96. https://doi.org/10.3390/vaccines10010096
Chicago/Turabian StyleMerkuleva, Iuliia A., Dmitry N. Shcherbakov, Mariya B. Borgoyakova, Daniil V. Shanshin, Andrey P. Rudometov, Larisa I. Karpenko, Svetlana V. Belenkaya, Anastasiya A. Isaeva, Valentina S. Nesmeyanova, Elena I. Kazachinskaia, and et al. 2022. "Comparative Immunogenicity of the Recombinant Receptor-Binding Domain of Protein S SARS-CoV-2 Obtained in Prokaryotic and Mammalian Expression Systems" Vaccines 10, no. 1: 96. https://doi.org/10.3390/vaccines10010096
APA StyleMerkuleva, I. A., Shcherbakov, D. N., Borgoyakova, M. B., Shanshin, D. V., Rudometov, A. P., Karpenko, L. I., Belenkaya, S. V., Isaeva, A. A., Nesmeyanova, V. S., Kazachinskaia, E. I., Volosnikova, E. A., Esina, T. I., Zaykovskaya, A. V., Pyankov, O. V., Borisevich, S. S., Shelemba, A. A., Chikaev, A. N., & Ilyichev, A. A. (2022). Comparative Immunogenicity of the Recombinant Receptor-Binding Domain of Protein S SARS-CoV-2 Obtained in Prokaryotic and Mammalian Expression Systems. Vaccines, 10(1), 96. https://doi.org/10.3390/vaccines10010096






