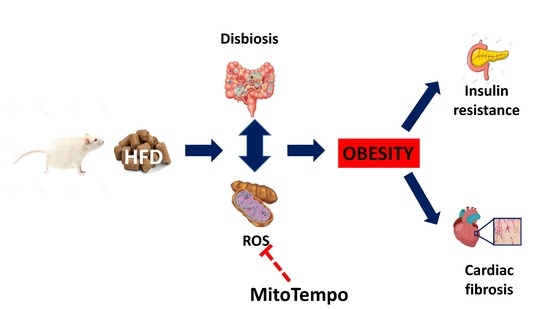
The Interaction between Mitochondrial Oxidative Stress and Gut Microbiota in the Cardiometabolic Consequences in Diet-Induced Obese Rats
Abstract
1. Introduction
2. Methods
2.1. MitoTempo Administration
2.2. Animals and Experimental Groups
2.3. Blood Biochemistry
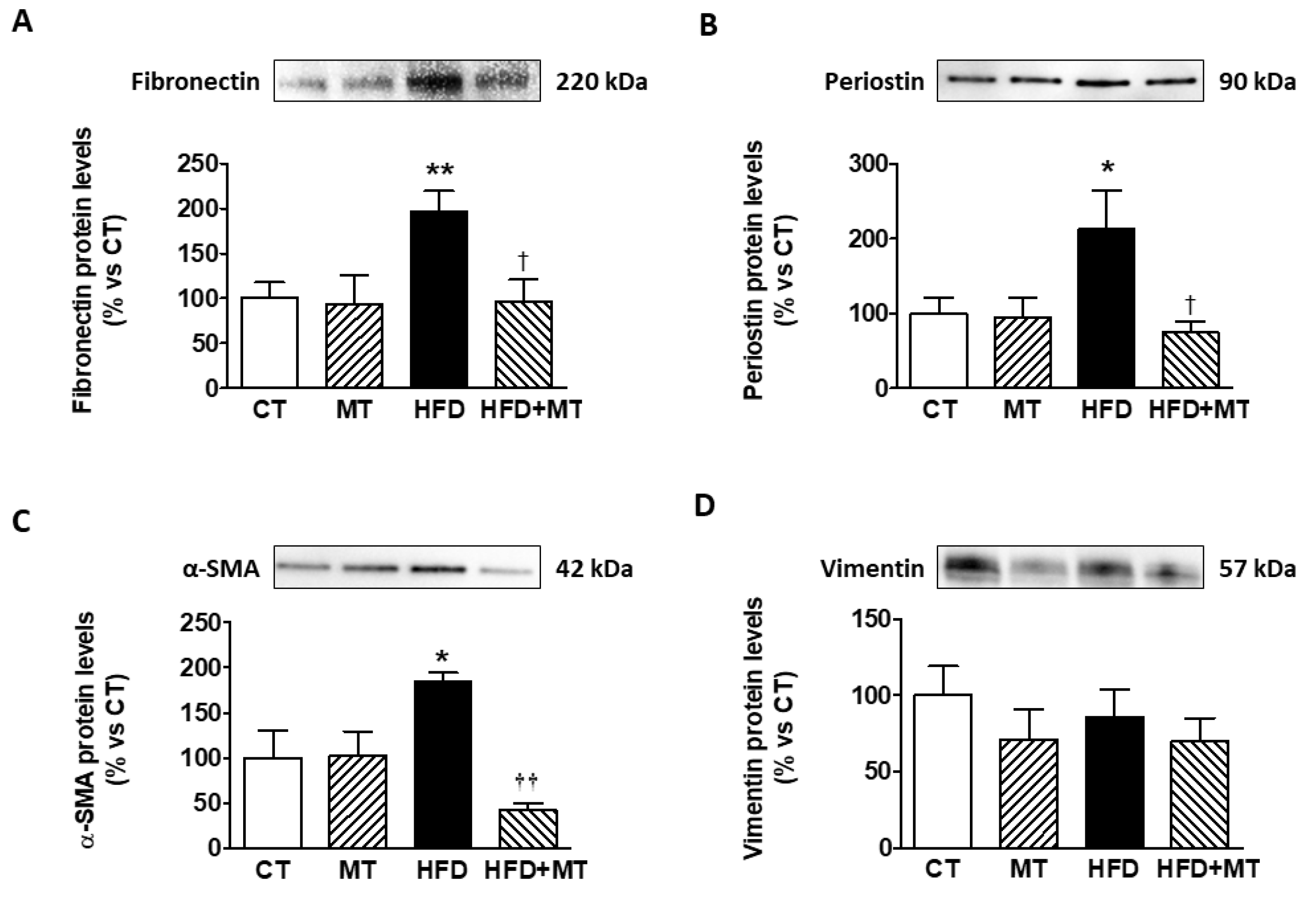
2.4. Western Blotting
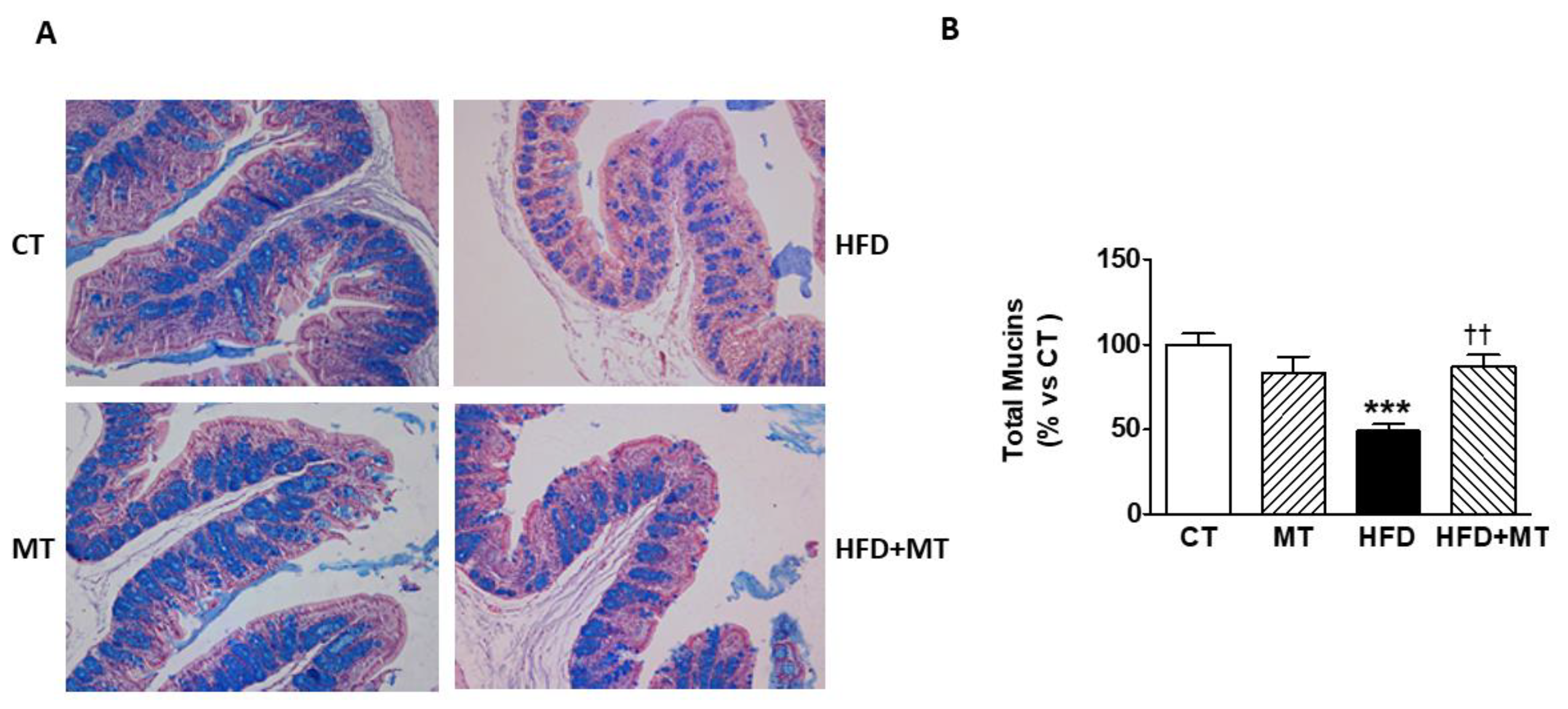
2.5. Mucin Level Analysis in Goblet Cells
2.6. Isolation of Microbial DNA
2.7. Amplification and Sequencing of the 16S rDNA Gene
2.8. Bioinformatic Analysis
2.9. Statistical Analysis
3. Results
3.1. Effects of MitoTempo in Metabolic and Cardiac Parameters of Diet-Induced Obese Rats
3.2. Effects of MitoTempo in the Mucin Levels in the Colon of Diet-Induced Obese Rats
3.3. Effects of MitoTempo in Gut Microbiota Diversity of Diet-Induced Obese Rats
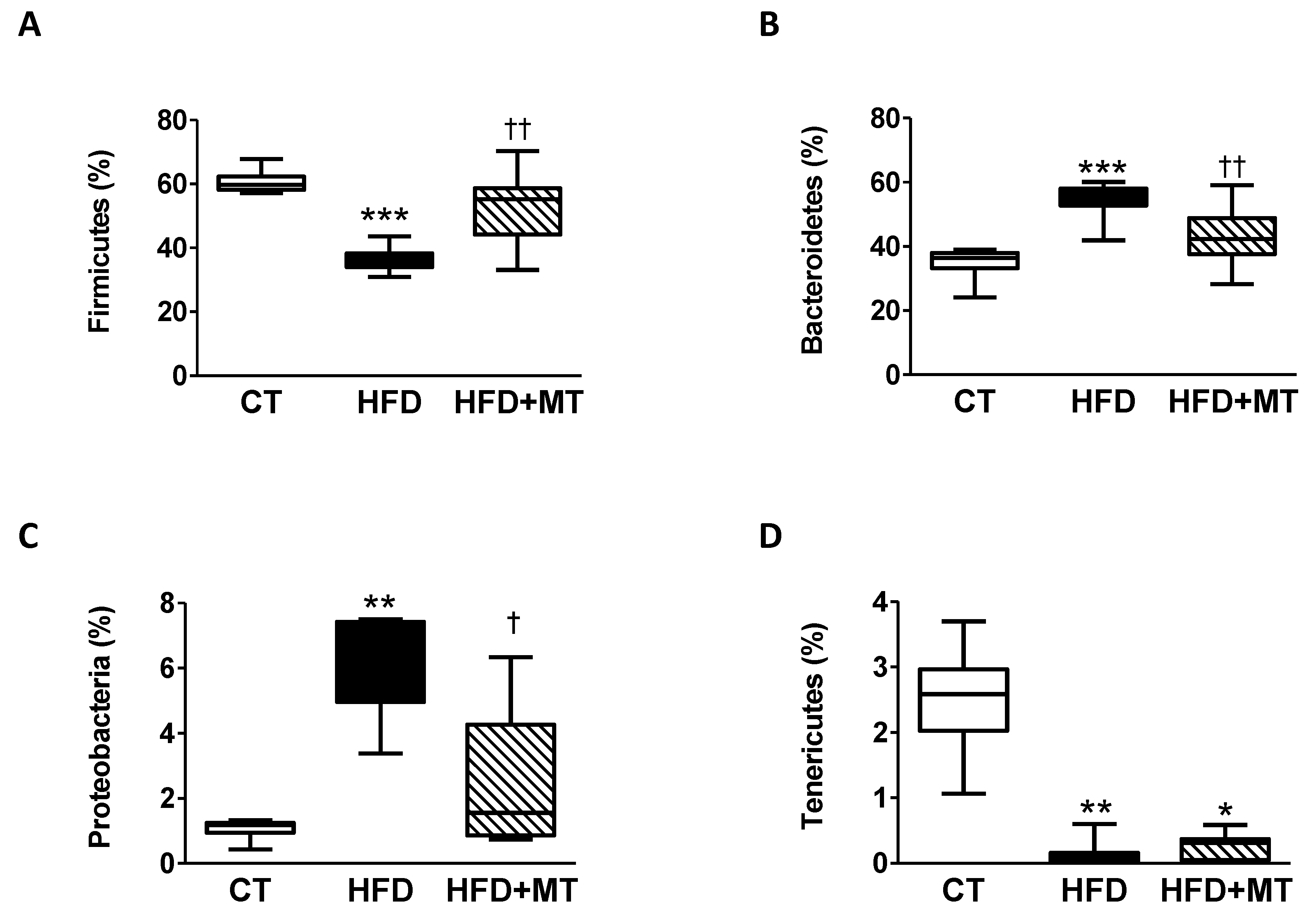
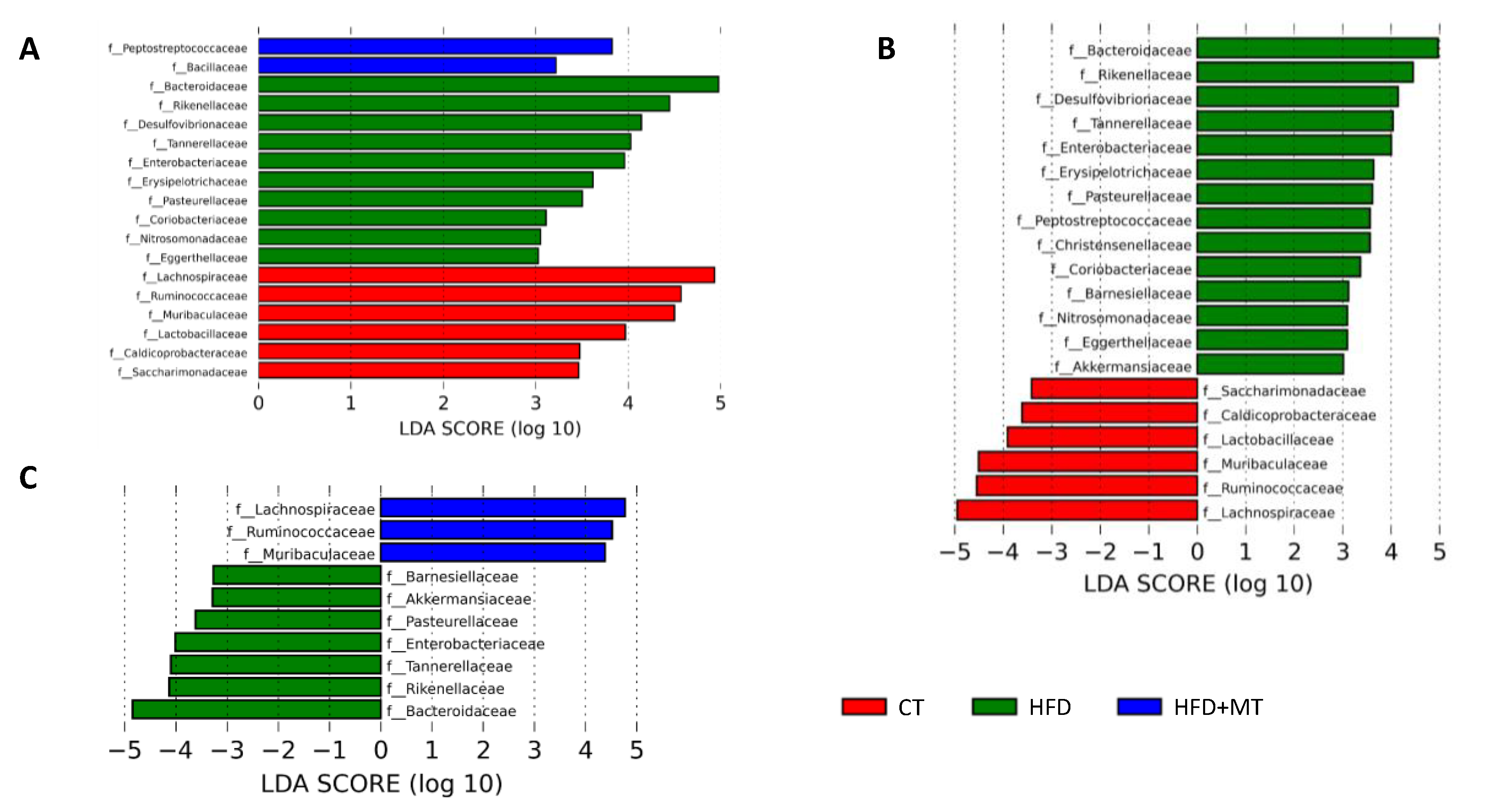
3.4. Effects of MitoTempo in Gut Microbiota Composition of HFD-Induced Obese Rats
3.5. Relationship between Genera Abundance and Metabolic and Cardiac Parameters
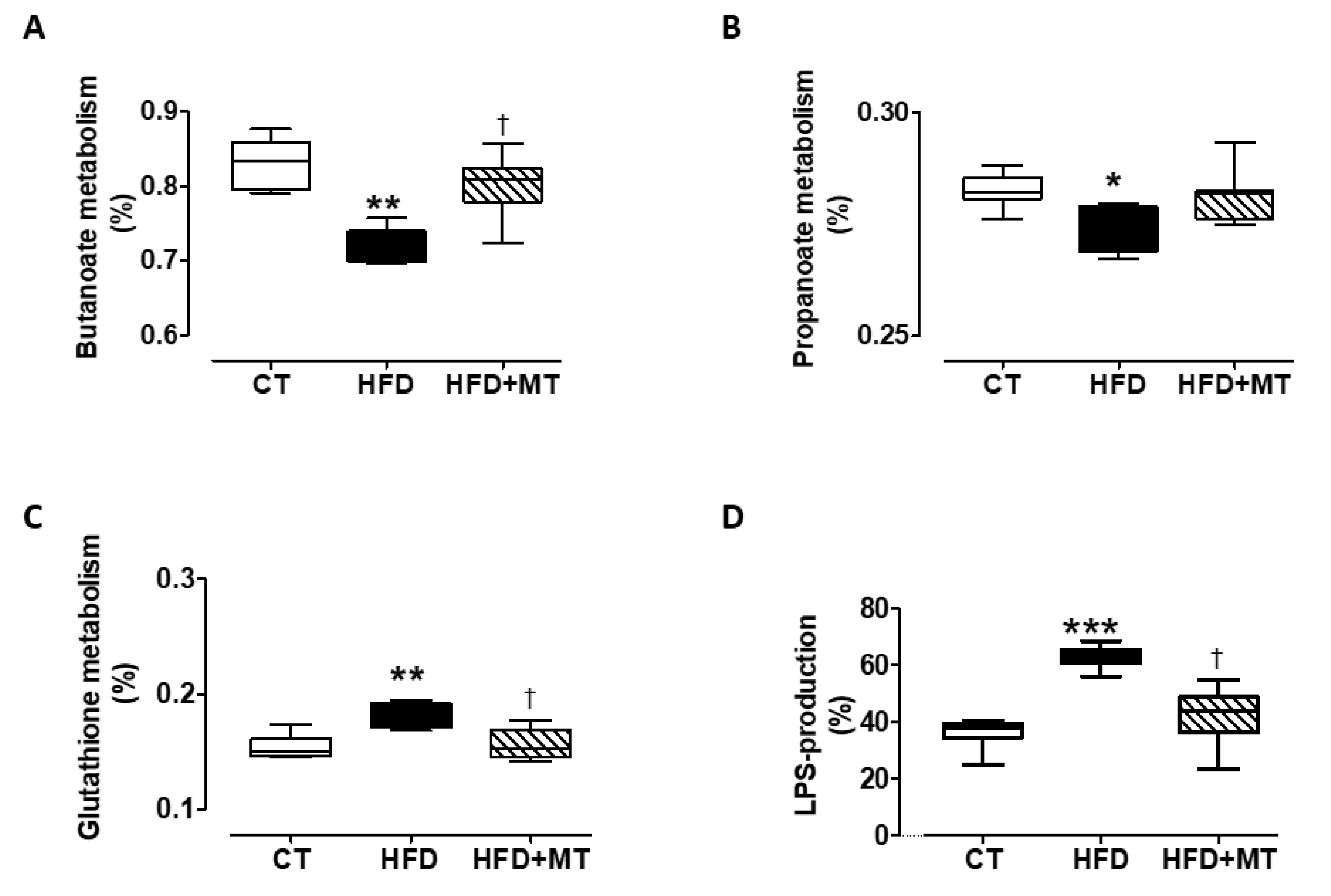
3.6. Effects of MitoTempo in Gut microbiota Metabolism of Diet-Induced Obese Rats
4. Discussion
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Diamanti-Kandarakis, E.; Papalou, O.; Kandaraki, E.A.; Kassi, G. Mechanisms in Endocrinology: Nutrition as a mediator of oxidative stress in metabolic and reproductive disorders in women. Eur. J. Endocrinol. 2017, 176, R79–R99. [Google Scholar] [CrossRef] [PubMed]
- Green, D.R.; Galluzzi, L.; Kroemer, G. Mitochondria and the autophagy-inflammation-cell death axis in organismal aging. Science 2011, 333, 1109–1112. [Google Scholar] [CrossRef]
- Redza-Dutordoir, M.; Averill-Bates, D.A. Activation of apoptosis signalling pathways by reactive oxygen species. Biochim. Biophys. Acta 2016, 1863, 2977–2992. [Google Scholar] [CrossRef] [PubMed]
- Wei, H.; Cong, X. The effect of reactive oxygen species on cardiomyocyte differentiation of pluripotent stem cells. Free Radic. Res. 2018, 52, 150–158. [Google Scholar] [CrossRef]
- de Mello, A.H.; Costa, A.B.; Engel, J.D.G.; Rezin, G.T. Mitochondrial dysfunction in obesity. Life Sci. 2018, 192, 26–32. [Google Scholar] [CrossRef]
- Jimenez-Gonzalez, S.; Marin-Royo, G.; Jurado-Lopez, R.; Bartolome, M.V.; Romero-Miranda, A.; Luaces, M.; Islas, F.; Nieto, M.L.; Martinez-Martinez, E.; Cachofeiro, V. The Crosstalk between Cardiac Lipotoxicity and Mitochondrial Oxidative Stress in the Cardiac Alterations in Diet-Induced Obesity in Rats. Cells 2020, 9, 451. [Google Scholar] [CrossRef]
- Marin-Royo, G.; Rodriguez, C.; Le Pape, A.; Jurado-Lopez, R.; Luaces, M.; Antequera, A.; Martinez-Gonzalez, J.; Souza-Neto, F.V.; Nieto, M.L.; Martinez-Martinez, E.; et al. The role of mitochondrial oxidative stress in the metabolic alterations in diet-induced obesity in rats. FASEB J. Off. Publ. Fed. Am. Soc. Exp. Biol. 2019, 33, 12060–12072. [Google Scholar] [CrossRef]
- Gutierrez-Tenorio, J.; Marin-Royo, G.; Martinez-Martinez, E.; Martin, R.; Miana, M.; Lopez-Andres, N.; Jurado-Lopez, R.; Gallardo, I.; Luaces, M.; San Roman, J.A.; et al. The role of oxidative stress in the crosstalk between leptin and mineralocorticoid receptor in the cardiac fibrosis associated with obesity. Sci. Rep. 2017, 7, 16802. [Google Scholar] [CrossRef] [PubMed]
- Ortega, F.B.; Lavie, C.J.; Blair, S.N. Obesity and Cardiovascular Disease. Circ. Res. 2016, 118, 1752–1770. [Google Scholar] [CrossRef]
- Isomaa, B.; Almgren, P.; Tuomi, T.; Forsen, B.; Lahti, K.; Nissen, M.; Taskinen, M.R.; Groop, L. Cardiovascular morbidity and mortality associated with the metabolic syndrome. Diabetes Care 2001, 24, 683–689. [Google Scholar] [CrossRef] [PubMed]
- Miyamoto, J.; Igarashi, M.; Watanabe, K.; Karaki, S.I.; Mukouyama, H.; Kishino, S.; Li, X.; Ichimura, A.; Irie, J.; Sugimoto, Y.; et al. Gut microbiota confers host resistance to obesity by metabolizing dietary polyunsaturated fatty acids. Nat. Commun. 2019, 10, 4007. [Google Scholar] [CrossRef]
- Maslowski, K.M.; Mackay, C.R. Diet, gut microbiota and immune responses. Nat. Immunol. 2011, 12, 5–9. [Google Scholar] [CrossRef]
- Flint, H.J.; Duncan, S.H.; Scott, K.P.; Louis, P. Links between diet, gut microbiota composition and gut metabolism. Proc. Nutr. Soc. 2015, 74, 13–22. [Google Scholar] [CrossRef]
- Wang, S.; Huang, M.; You, X.; Zhao, J.; Chen, L.; Wang, L.; Luo, Y.; Chen, Y. Gut microbiota mediates the anti-obesity effect of calorie restriction in mice. Sci. Rep. 2018, 8, 13037. [Google Scholar] [CrossRef]
- Khan, M.J.; Gerasimidis, K.; Edwards, C.A.; Shaikh, M.G. Role of Gut Microbiota in the Aetiology of Obesity: Proposed Mechanisms and Review of the Literature. J. Obes. 2016, 2016, 7353642. [Google Scholar] [CrossRef]
- Angelakis, E.; Armougom, F.; Million, M.; Raoult, D. The relationship between gut microbiota and weight gain in humans. Future Microbiol. 2012, 7, 91–109. [Google Scholar] [CrossRef]
- Duncan, S.H.; Lobley, G.E.; Holtrop, G.; Ince, J.; Johnstone, A.M.; Louis, P.; Flint, H.J. Human colonic microbiota associated with diet, obesity and weight loss. Int. J. Obes. 2008, 32, 1720–1724. [Google Scholar] [CrossRef]
- Kabouridis, P.S.; Lasrado, R.; McCallum, S.; Chng, S.H.; Snippert, H.J.; Clevers, H.; Pettersson, S.; Pachnis, V. The gut microbiota keeps enteric glial cells on the move; prospective roles of the gut epithelium and immune system. Gut Microbes 2015, 6, 398–403. [Google Scholar] [CrossRef]
- Natividad, J.M.; Verdu, E.F. Modulation of intestinal barrier by intestinal microbiota: Pathological and therapeutic implications. Pharmacol. Res. 2013, 69, 42–51. [Google Scholar] [CrossRef]
- Saint-Georges-Chaumet, Y.; Edeas, M. Microbiota-mitochondria inter-talk: Consequence for microbiota-host interaction. Pathog. Dis. 2016, 74, ftv096. [Google Scholar] [CrossRef]
- Clark, A.; Mach, N. The Crosstalk between the Gut Microbiota and Mitochondria during Exercise. Front. Physiol. 2017, 8, 319. [Google Scholar] [CrossRef] [PubMed]
- Donohoe, D.R.; Garge, N.; Zhang, X.; Sun, W.; O′Connell, T.M.; Bunger, M.K.; Bultman, S.J. The microbiome and butyrate regulate energy metabolism and autophagy in the mammalian colon. Cell Metab. 2011, 13, 517–526. [Google Scholar] [CrossRef] [PubMed]
- den Besten, G.; van Eunen, K.; Groen, A.K.; Venema, K.; Reijngoud, D.J.; Bakker, B.M. The role of short-chain fatty acids in the interplay between diet, gut microbiota, and host energy metabolism. J. Lipid Res. 2013, 54, 2325–2340. [Google Scholar] [CrossRef] [PubMed]
- Kozak, L.P.; Newman, S.; Chao, P.M.; Mendoza, T.; Koza, R.A. The early nutritional environment of mice determines the capacity for adipose tissue expansion by modulating genes of caveolae structure. PLoS ONE 2010, 5, e11015. [Google Scholar] [CrossRef]
- Gutch, M.; Kumar, S.; Razi, S.M.; Gupta, K.K.; Gupta, A. Assessment of insulin sensitivity/resistance. Indian J. Endocrinol. Metab. 2015, 19, 160–164. [Google Scholar] [CrossRef]
- Rastelli, M.; Knauf, C.; Cani, P.D. Gut Microbes and Health: A Focus on the Mechanisms Linking Microbes, Obesity, and Related Disorders. Obesity 2018, 26, 792–800. [Google Scholar] [CrossRef]
- Vallianou, N.; Stratigou, T.; Christodoulatos, G.S.; Dalamaga, M. Understanding the Role of the Gut Microbiome and Microbial Metabolites in Obesity and Obesity-Associated Metabolic Disorders: Current Evidence and Perspectives. Curr. Obes. Rep. 2019, 8, 317–332. [Google Scholar] [CrossRef]
- Sze, M.A.; Schloss, P.D. Looking for a Signal in the Noise: Revisiting Obesity and the Microbiome. mBio 2016, 7. [Google Scholar] [CrossRef]
- Lee, C.J.; Sears, C.L.; Maruthur, N. Gut microbiome and its role in obesity and insulin resistance. Ann. N. Y. Acad. Sci. 2020, 1461, 37–52. [Google Scholar] [CrossRef]
- Truax, A.D.; Chen, L.; Tam, J.W.; Cheng, N.; Guo, H.; Koblansky, A.A.; Chou, W.C.; Wilson, J.E.; Brickey, W.J.; Petrucelli, A.; et al. The Inhibitory Innate Immune Sensor NLRP12 Maintains a Threshold against Obesity by Regulating Gut Microbiota Homeostasis. Cell Host Microbe 2018, 24, 364–378.e366. [Google Scholar] [CrossRef]
- Vujicic, M.; Saksida, T.; Despotovic, S.; Bajic, S.S.; Lalic, I.; Koprivica, I.; Gajic, D.; Golic, N.; Tolinacki, M.; Stojanovic, I. The Role of Macrophage Migration Inhibitory Factor in the Function of Intestinal Barrier. Sci. Rep. 2018, 8, 6337. [Google Scholar] [CrossRef]
- Vital, M.; Karch, A.; Pieper, D.H. Colonic Butyrate-Producing Communities in Humans: An Overview Using Omics Data. mSystems 2017, 2. [Google Scholar] [CrossRef]
- Do, M.H.; Lee, E.; Oh, M.J.; Kim, Y.; Park, H.Y. High-Glucose or -Fructose Diet Cause Changes of the Gut Microbiota and Metabolic Disorders in Mice without Body Weight Change. Nutrients 2018, 10, 761. [Google Scholar] [CrossRef]
- Lippert, K.; Kedenko, L.; Antonielli, L.; Kedenko, I.; Gemeier, C.; Leitner, M.; Kautzky-Willer, A.; Paulweber, B.; Hackl, E. Gut microbiota dysbiosis associated with glucose metabolism disorders and the metabolic syndrome in older adults. Benef. Microbes 2017, 8, 545–556. [Google Scholar] [CrossRef] [PubMed]
- Han, L.; Zhao, L.H.; Zhang, M.L.; Li, H.T.; Gao, Z.Z.; Zheng, X.J.; Wang, X.M.; Wu, H.R.; Zheng, Y.J.; Jiang, X.T.; et al. A Novel Antidiabetic Monomers Combination Alleviates Insulin Resistance Through Bacteria-Cometabolism-Inflammation Responses. Front. Microbiol. 2020, 11, 173. [Google Scholar] [CrossRef] [PubMed]
- Walker, J.M.; Eckardt, P.; Aleman, J.O.; da Rosa, J.C.; Liang, Y.; Iizumi, T.; Etheve, S.; Blaser, M.J.; Breslow, J.L.; Holt, P.R. The effects of trans-resveratrol on insulin resistance, inflammation, and microbiota in men with the metabolic syndrome: A pilot randomized, placebo-controlled clinical trial. J. Clin. Transl. Res. 2019, 4, 122–135. [Google Scholar] [PubMed]
- Walker, J.T.; McLeod, K.; Kim, S.; Conway, S.J.; Hamilton, D.W. Periostin as a multifunctional modulator of the wound healing response. Cell Tissue Res. 2016, 365, 453–465. [Google Scholar] [CrossRef]
- Wu, H.; Li, G.N.; Xie, J.; Li, R.; Chen, Q.H.; Chen, J.Z.; Wei, Z.H.; Kang, L.N.; Xu, B. Resveratrol ameliorates myocardial fibrosis by inhibiting ROS/ERK/TGF-beta/periostin pathway in STZ-induced diabetic mice. BMC Cardiovasc. Disord. 2016, 16, 5. [Google Scholar] [CrossRef]
- Valiente-Alandi, I.; Potter, S.J.; Salvador, A.M.; Schafer, A.E.; Schips, T.; Carrillo-Salinas, F.; Gibson, A.M.; Nieman, M.L.; Perkins, C.; Sargent, M.A.; et al. Inhibiting Fibronectin Attenuates Fibrosis and Improves Cardiac Function in a Model of Heart Failure. Circulation 2018, 138, 1236–1252. [Google Scholar] [CrossRef]
- Plummer, J.L.; Ossowicz, C.J.; Whibley, C.; Ilsley, A.H.; Hall, P.D. Influence of intestinal flora on the development of fibrosis and cirrhosis in a rat model. J. Gastroenterol. Hepatol. 2000, 15, 1307–1311. [Google Scholar] [CrossRef]
- Knippenberg, S.; Ueberberg, B.; Maus, R.; Bohling, J.; Ding, N.; Tort Tarres, M.; Hoymann, H.G.; Jonigk, D.; Izykowski, N.; Paton, J.C.; et al. Streptococcus pneumoniae triggers progression of pulmonary fibrosis through pneumolysin. Thorax 2015, 70, 636–646. [Google Scholar] [CrossRef] [PubMed]
- Singh, H.; Torralba, M.G.; Moncera, K.J.; DiLello, L.; Petrini, J.; Nelson, K.E.; Pieper, R. Gastro-intestinal and oral microbiome signatures associated with healthy aging. GeroScience 2019, 41, 907–921. [Google Scholar] [CrossRef] [PubMed]
- Cui, X.; Ye, L.; Li, J.; Jin, L.; Wang, W.; Li, S.; Bao, M.; Wu, S.; Li, L.; Geng, B.; et al. Metagenomic and metabolomic analyses unveil dysbiosis of gut microbiota in chronic heart failure patients. Sci. Rep. 2018, 8, 635. [Google Scholar] [CrossRef] [PubMed]
- Kimura, I.; Ozawa, K.; Inoue, D.; Imamura, T.; Kimura, K.; Maeda, T.; Terasawa, K.; Kashihara, D.; Hirano, K.; Tani, T.; et al. The gut microbiota suppresses insulin-mediated fat accumulation via the short-chain fatty acid receptor GPR43. Nat. Commun. 2013, 4, 1829. [Google Scholar] [CrossRef]
- Ghosh, S.S.; Wang, J.; Yannie, P.J.; Ghosh, S. Intestinal Barrier Dysfunction, LPS Translocation, and Disease Development. J. Endocr. Soc. 2020, 4, bvz039. [Google Scholar] [CrossRef]
- Ellulu, M.S.; Patimah, I.; Khaza′ai, H.; Rahmat, A.; Abed, Y. Obesity and inflammation: The linking mechanism and the complications. Arch. Med Sci. AMS 2017, 13, 851–863. [Google Scholar] [CrossRef]
- Gregor, M.F.; Hotamisligil, G.S. Inflammatory mechanisms in obesity. Annu. Rev. Immunol. 2011, 29, 415–445. [Google Scholar] [CrossRef]
- Huang, W.; Zhou, L.; Guo, H.; Xu, Y.; Xu, Y. The role of short-chain fatty acids in kidney injury induced by gut-derived inflammatory response. Metab. Clin. Exp. 2017, 68, 20–30. [Google Scholar] [CrossRef]
- Chang, P.V.; Hao, L.; Offermanns, S.; Medzhitov, R. The microbial metabolite butyrate regulates intestinal macrophage function via histone deacetylase inhibition. Proc. Natl. Acad. Sci. USA 2014, 111, 2247–2252. [Google Scholar] [CrossRef]
- Ghosh, S.; Sulistyoningrum, D.C.; Glier, M.B.; Verchere, C.B.; Devlin, A.M. Altered glutathione homeostasis in heart augments cardiac lipotoxicity associated with diet-induced obesity in mice. J. Biol. Chem. 2011, 286, 42483–42493. [Google Scholar] [CrossRef]

| Variable | CT | MT | HFD | HFD + MT |
|---|---|---|---|---|
| Adiposity index (%) | 3.49 ± 0.26 | 3.52 ± 0.22 | 8.39 ± 0.41 *** | 6.9 ± 0.55 *,††† |
| Glucose (mg/dL) | 98.8 ± 4.6 | 95.2 ± 2.8 | 114.6 ± 2.1 ** | 104.3 ± 2.5 † |
| Insulin (pg/mL) | 72.2 ± 7.7 | 60.9 ± 5.9 | 241.6 ± 30.8 *** | 132.7 ± 16.5 *,††† |
| HOMA index | 2.50 ± 0.33 | 1.98 ± 0.19 | 9.28 ± 1.1 *** | 4.8 ± 0.7 *,††† |
| Variable | CT | HFD | HFD + MT |
|---|---|---|---|
| Observed OTUs | 3267 ± 121 | 3495 ± 117 | 3458 ± 98 |
| Chao1 richness index | 3945 ± 100 | 4168 ± 1128 | 4095 ± 61 |
| Shannon index | 7.93 ± 0.14 | 7.51 ± 0.10 * | 7.95 ± 0.14 † |
| Pielou’s evenness index | 0.68 ± 0.011 | 0.64 ± 0.01 * | 0.68 ± 0.013 † |
| Bray–Curtis dissimilarity | 0.549 ± 0.017 | 0.423 ± 0.02 ** | 0.62 ± 0.041 *,† |
| JSD distance | 0.483 ± 0.013 | 0.388 ± 0.015 ** | 0.521 ± 0.033 *,† |
| Genus | Collagen Content | HOMA Index | ||
|---|---|---|---|---|
| - | Β (95% CI) | p | Β (95% CI) | p |
| Eubacterium oxidoreducens group | −0.73 (−1.17 to −0.28) | 0.003 | −2.36 (−3.73 to −0.98) | 0.002 |
| Streptococcus | 6.26 (0.30 to 12.22) | 0.041 | ||
| Lachnospiraceae ND3007 group | −3.42 (−6.17 to −0.67) | 0.018 | ||
| Erysipelotrichaceae UCG-003 | −63.64 (−109.45 to −17.08) | 0.01 | ||
| Ruminococcus 1 | −0.13 (−0.25 to −0.11) | 0.034 | ||
| Morganella | 10.71 (6.91 to 14.51) | <0.0001 | ||
| Muribaculaceae (uncultured bacterium) | −0.457 (−0.667 to −0.246) | <0.0001 | ||
| Candidatus Soleaferrea | 253.77 (125.49 to 382.05) | 0.001 | ||
| Holdmania | −107.70 (−177.80 to −37.70) | 0.005 | ||
| Peptoclostridium | 82.47 (5.56 to 159.38) | 0.037 | ||
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ortega-Hernández, A.; Martínez-Martínez, E.; Gómez-Gordo, R.; López-Andrés, N.; Fernández-Celis, A.; Gutiérrrez-Miranda, B.; Nieto, M.L.; Alarcón, T.; Alba, C.; Gómez-Garre, D.; et al. The Interaction between Mitochondrial Oxidative Stress and Gut Microbiota in the Cardiometabolic Consequences in Diet-Induced Obese Rats. Antioxidants 2020, 9, 640. https://doi.org/10.3390/antiox9070640
Ortega-Hernández A, Martínez-Martínez E, Gómez-Gordo R, López-Andrés N, Fernández-Celis A, Gutiérrrez-Miranda B, Nieto ML, Alarcón T, Alba C, Gómez-Garre D, et al. The Interaction between Mitochondrial Oxidative Stress and Gut Microbiota in the Cardiometabolic Consequences in Diet-Induced Obese Rats. Antioxidants. 2020; 9(7):640. https://doi.org/10.3390/antiox9070640
Chicago/Turabian StyleOrtega-Hernández, Adriana, Ernesto Martínez-Martínez, Ruben Gómez-Gordo, Natalia López-Andrés, Amaya Fernández-Celis, Beatriz Gutiérrrez-Miranda, María Luisa Nieto, Teresa Alarcón, Claudio Alba, Dulcenombre Gómez-Garre, and et al. 2020. "The Interaction between Mitochondrial Oxidative Stress and Gut Microbiota in the Cardiometabolic Consequences in Diet-Induced Obese Rats" Antioxidants 9, no. 7: 640. https://doi.org/10.3390/antiox9070640
APA StyleOrtega-Hernández, A., Martínez-Martínez, E., Gómez-Gordo, R., López-Andrés, N., Fernández-Celis, A., Gutiérrrez-Miranda, B., Nieto, M. L., Alarcón, T., Alba, C., Gómez-Garre, D., & Cachofeiro, V. (2020). The Interaction between Mitochondrial Oxidative Stress and Gut Microbiota in the Cardiometabolic Consequences in Diet-Induced Obese Rats. Antioxidants, 9(7), 640. https://doi.org/10.3390/antiox9070640