Physiological Roles of Plant Methionine Sulfoxide Reductases in Redox Homeostasis and Signaling
Abstract
1. Introduction
2. Subcellular Localization and Organ Distribution of Plant MSRs
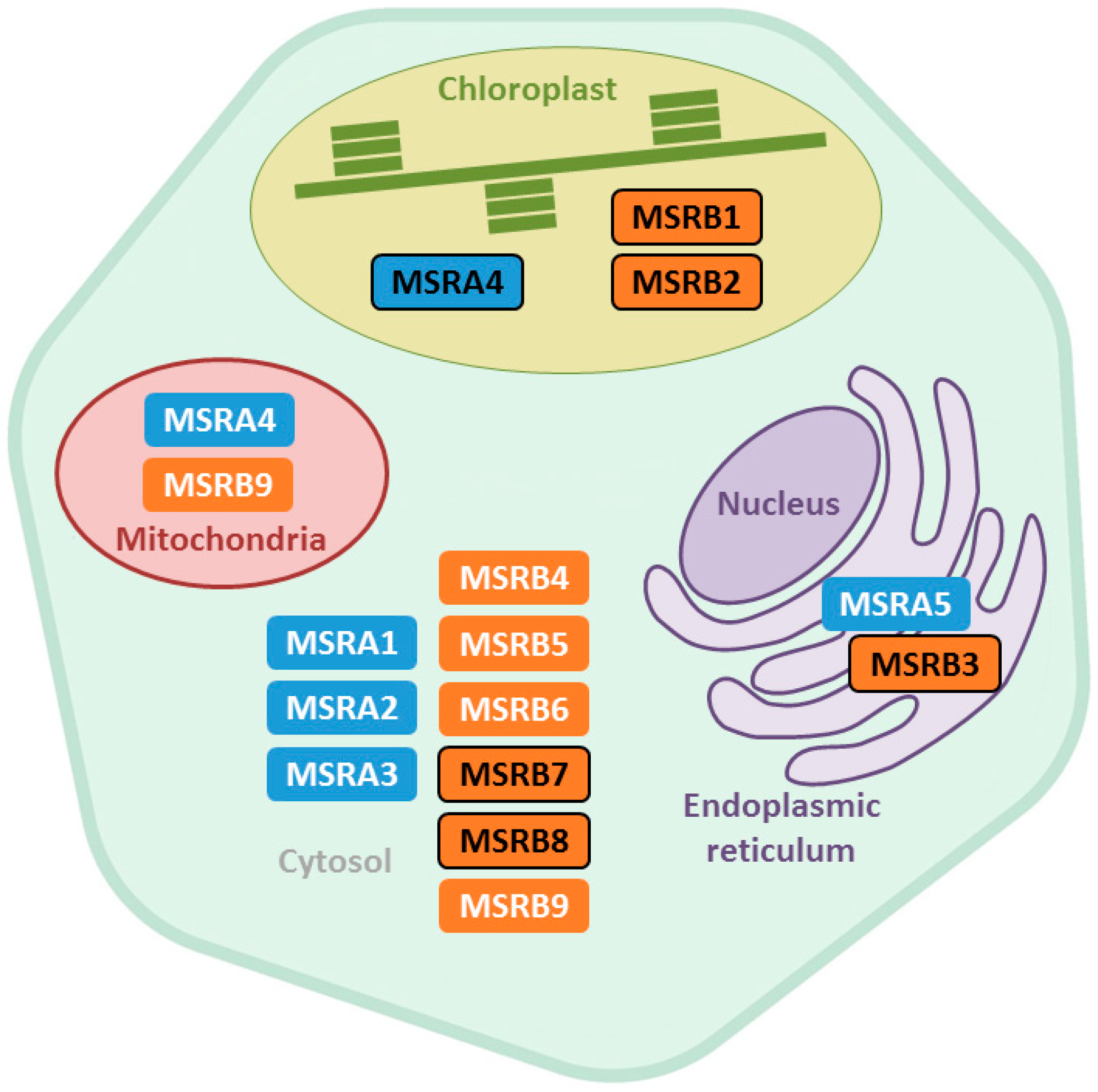
2.1. Subcellular Localization
2.2. Organ Distribution
3. Regulation of the Expression of MSR Genes in Photosynthetic Organisms
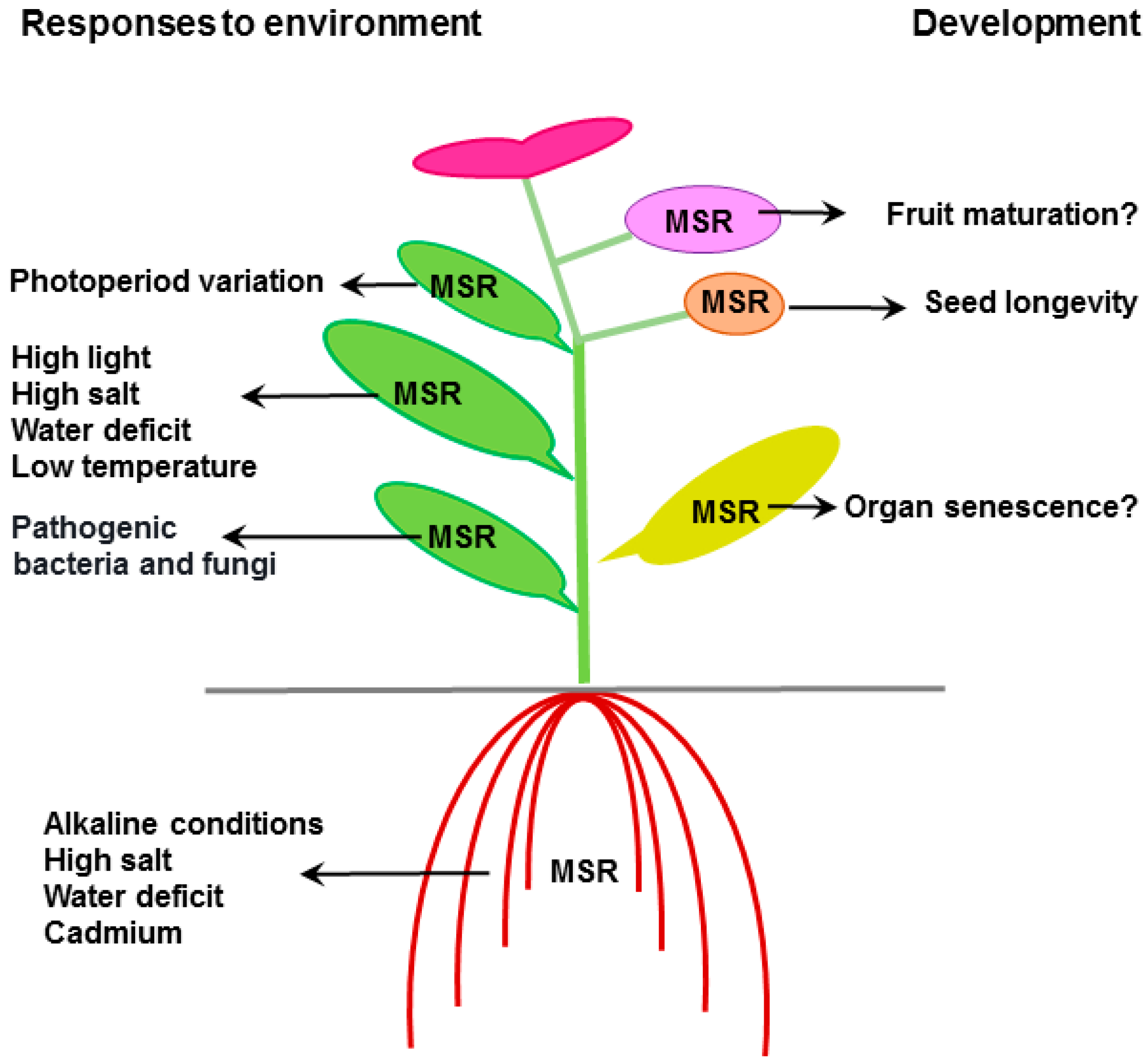
3.1. Effect of Environmental Conditions
3.2. Signaling Actors Involved in the Control of MSR Gene Expression
3.2.1. Involvement of ROS and Reactive Nitrogen Species (RNS) in MSR Gene Expression
3.2.2. Involvement of Phytohormones in MSR Gene Expression
3.2.3. Conclusions
4. MSR Activity and MetO Content in Higher Plants
4.1. MSR Activity in Plant Extracts
4.2. MetO Content in Plants
4.3. Signals Involved in the Control of MSR Activity and MetO Level
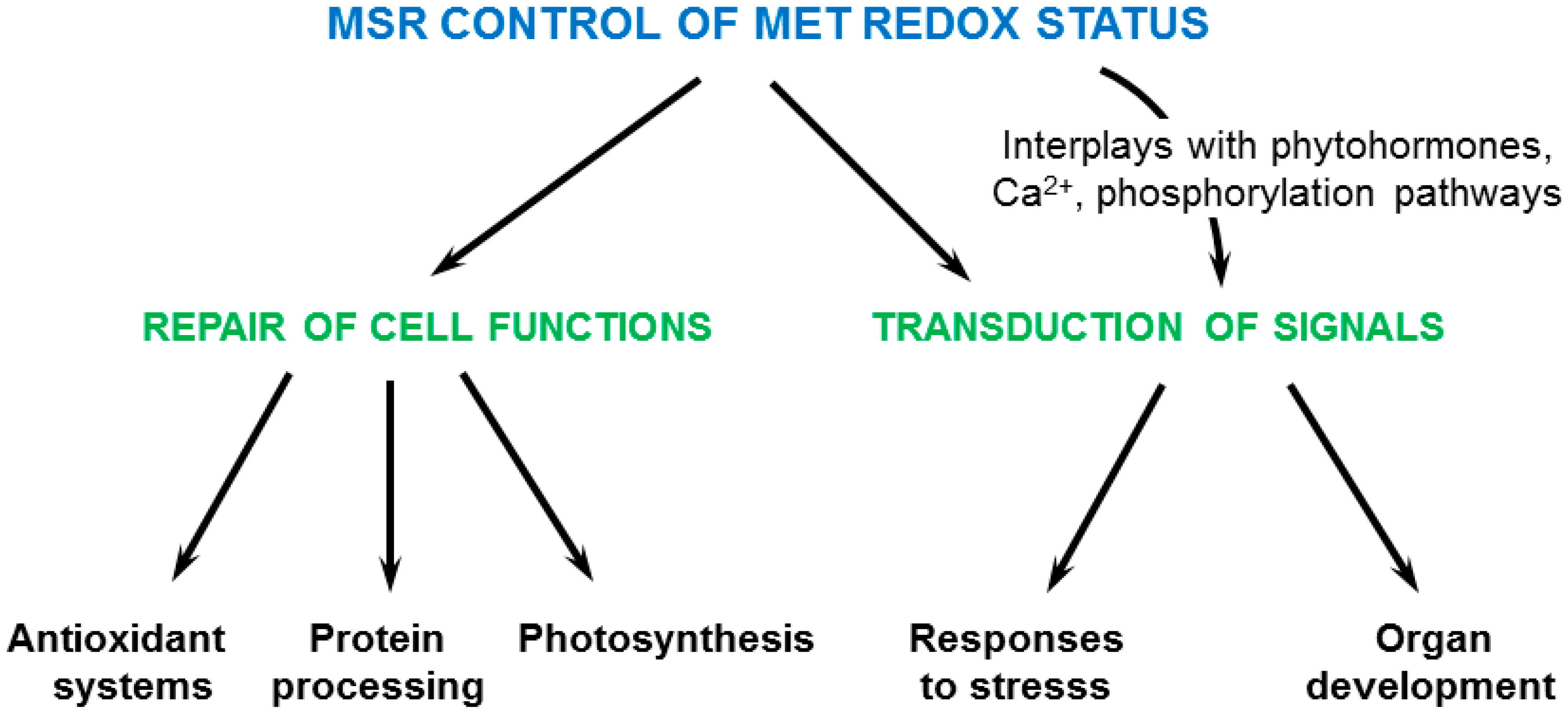
5. Physiological Functions of Plant MSRs
5.1. Oxidative Treatments and Photooxidative Conditions
5.2. Abiotic Constraints
5.3. Biotic Constraints
5.4. Involvement in Ageing Process
6. Mode of Action and Substrates of Plant MSRs
6.1. Strategies for Searching Plant MSR Targets
6.1.1. Proteins Displaying High Met Content
6.1.2. Proteins Exhibiting Modified MetO Content in Response to Oxidative Treatments or Signaling Molecules
6.1.3. Proteins Interacting with MSRs
6.1.4. Genes Displaying Modified Expression in Lines Up- or Down-Regulated for MSR Expression
6.2. Identity and Functions of MSR Partners or Possible Targets
6.2.1. Translation and Folding of Proteins
6.2.2. Chlorophyll Metabolism and Photosynthetic Activity
6.2.3. Antixoxidant Mechanisms
6.2.4. Signaling in Relation with Calmodulin
6.2.5. Proteins Responsive to Stress
7. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Davies, M.J. The oxidative environment and protein damage. Biochim. Biophys. Acta 2005, 1703, 93–109. [Google Scholar] [CrossRef] [PubMed]
- Møller, I.M.; Jensen, P.E.; Hansson, A. Oxidative modifications to cellular components in plants. Annu. Rev. Plant Biol. 2007, 58, 459–481. [Google Scholar] [CrossRef] [PubMed]
- Couturier, J.; Chibani, K.; Jacquot, J.P.; Rouhier, N. Cysteine-based redox regulation and signaling in plants. Front. Plant Sci. 2013, 4, 105. [Google Scholar] [CrossRef] [PubMed]
- Meyer, Y.; Reichheld, J.P.; Vignols, F. Thioredoxins in Arabidopsis and other plants. Photosynth. Res. 2005, 86, 419–433. [Google Scholar] [CrossRef] [PubMed]
- Rouhier, N.; Couturier, J.; Jacquot, J.P. Genome-wide analysis of plant glutaredoxin systems. J. Exp. Bot. 2006, 57, 1685–1696. [Google Scholar] [CrossRef] [PubMed]
- Rouhier, N.; Lemaire, S.D.; Jacquot, J.P. The role of glutathione in photosynthetic organisms: Emerging functions for glutaredoxins and glutathionylation. Annu. Rev. Plant Biol. 2008, 59, 143–166. [Google Scholar] [CrossRef] [PubMed]
- Couturier, J.; Jacquot, J.P.; Rouhier, N. Evolution and diversity of glutaredoxins in photosynthetic organisms. Cell. Mol. Life Sci. 2009, 66, 2539–2557. [Google Scholar] [CrossRef] [PubMed]
- Vieira Dos Santos, C.; Rey, P. Plant thioredoxins are key actors in the oxidative stress response. Trends Plant Sci. 2006, 11, 329–334. [Google Scholar] [CrossRef] [PubMed]
- Meyer, Y.; Belin, C.; Delorme-Hinoux, V.; Reichheld, J.P.; Riondet, C. Thioredoxin and glutaredoxin systems in plants: Molecular mechanisms, crosstalks, and functional significance. Antioxid. Redox Signal. 2012, 17, 1124–1160. [Google Scholar] [CrossRef] [PubMed]
- Rouhier, N.; Cerveau, D.; Couturier, J.; Reichheld, J.P.; Rey, P. Involvement of thiol-based mechanisms in plant development Biochim. Biophys. Acta 2015, 1850, 1479–1496. [Google Scholar] [CrossRef] [PubMed]
- Rouhier, N.; Villarejo, A.; Srivastava, M.; Gelhaye, E.; Keech, O.; Droux, M.; Finkemeier, I.; Samuelsson, G.; Dietz, K.J.; Jacquot, J.P.; et al. Identification of plant glutaredoxin targets. Antioxid. Redox Signal. 2005, 7, 919–929. [Google Scholar] [CrossRef] [PubMed]
- Montrichard, F.; Alkhalfioui, F.; Yano, H.; Vensel, W.H.; Hurkman, W.J.; Buchanan, B.B. Thioredoxin targets in plants: The first 30 years. J. Proteom. 2009, 72, 452–474. [Google Scholar] [CrossRef] [PubMed]
- Pérez-Pérez, M.E.; Mauriès, A.; Maes, A.; Tourasse, N.J.; Hamon, M.; Lemaire, S.D.; Marchand, C.H. The deep thioredoxome in Chlamydomonas reinhardtii: New insights into redox regulation. Mol. Plant 2017, 10, 1107–1125. [Google Scholar] [CrossRef] [PubMed]
- Tarrago, L.; Péterfi, Z.; Lee, B.C.; Michel, T.; Gladyshev, V.N. Monitoring methionine sulfoxide with stereospecific mechanism-based fluorescent sensors. Nat. Chem. Biol. 2015, 11, 332–338. [Google Scholar] [CrossRef]
- Luo, S.; Levine, R.L. Methionine in proteins defends against oxidative stress. FASEB J. 2009, 23, 464–472. [Google Scholar] [CrossRef] [PubMed]
- Brot, N.; Weissbach, L.; Werth, J.; Weissbach, H. Enzymatic reduction of protein-bound methionine sulfoxide. Proc. Natl. Acad. Sci. USA 1981, 78, 2155–2158. [Google Scholar] [CrossRef] [PubMed]
- Grimaud, R.; Ezraty, B.; Mitchell, J.K.; Lafitte, D.; Briand, C.; Derrick, P.J.; Barras, F. Repair of oxidized proteins. Identification of a new methionine sulfoxide reductase. J. Biol. Chem. 2001, 276, 48915–48920. [Google Scholar] [CrossRef] [PubMed]
- Lowther, W.T.; Brot, N.; Weissbach, H.; Honek, J.F.; Matthews, B.W. Thiol-disulfide exchange is involved in the catalytic mechanism of peptide methionine sulfoxide reductase. Proc. Natl. Acad. Sci. USA 2000, 97, 6463–6468. [Google Scholar] [CrossRef] [PubMed]
- Weissbach, H.; Etienne, F.; Hoshi, T.; Heinemann, S.H.; Lowther, W.T.; Matthews, B.; St John, G.; Nathan, C.; Brot, N. Peptide methionine sulfoxide reductase: Structure, mechanism of action, and biological function. Arch. Biochem. Biophys. 2002, 397, 172–178. [Google Scholar] [CrossRef] [PubMed]
- Tarrago, L.; Kaya, A.; Weerapana, E.; Marino, S.M.; Gladyshev, V.N. Methionine sulfoxide reductases preferentially reduce unfolded oxidized proteins and protect cells from oxidative protein unfolding. J. Biol. Chem. 2012, 287, 2448–2459. [Google Scholar] [CrossRef] [PubMed]
- Le, D.T.; Tarrago, L.; Watanabe, Y.; Kaya, A.; Lee, B.C.; Tran, U.; Nishiyama, R.; Fomenko, D.E.; Gladyshev, V.N.; Tran, L.S. Diversity of plant methionine sulfoxide reductases B and evolution of a form specific for free methionine sulfoxide. PLoS ONE 2013, 8, e65637. [Google Scholar] [CrossRef] [PubMed]
- Boschi-Muller, S.; Gand, A.; Branlant, G. The methionine sulfoxide reductases: Catalysis and substrate specificities. Arch. Biochem. Biophys. 2008, 474, 266–273. [Google Scholar] [CrossRef] [PubMed]
- Rouhier, N.; Kauffmann, B.; Tete-Favier, F.; Palladino, P.; Gans, P.; Branlant, G.; Jacquot, J.P.; Boschi-Muller, S. Functional and structural aspects of poplar cytosolic and plastidial type a methionine sulfoxide reductases. J. Biol. Chem. 2007, 282, 3367–3378. [Google Scholar] [CrossRef] [PubMed]
- Tarrago, L.; Laugier, E.; Zaffagnini, M.; Marchand, C.; Le Maréchal, P.; Rouhier, N.; Lemaire, S.D.; Rey, P. Regeneration mechanisms of Arabidopsis thaliana methionine sulfoxide reductases B by glutaredoxins and thioredoxins. J. Biol. Chem. 2009, 284, 18963–18971. [Google Scholar] [CrossRef] [PubMed]
- Rey, P.; Cuine, S.; Eymery, F.; Garin, J.; Court, M.; Jacquot, J.P.; Rouhier, N.; Broin, M. Analysis of the proteins targeted by CDSP32, a plastidic thioredoxin participating in oxidative stress responses. Plant J. 2005, 41, 31–42. [Google Scholar] [CrossRef] [PubMed]
- Sagher, D.; Brunell, D.; Hejtmancik, J.F.; Kantorow, M.; Brot, N.; Weissbach, H. Thionein can serve as a reducing agent for the methionine sulfoxide reductases. Proc. Natl. Acad. Sci. USA 2006, 103, 8656–8661. [Google Scholar] [CrossRef] [PubMed]
- Vieira Dos Santos, C.; Laugier, E.; Tarrago, L.; Massot, V.; Issakidis-Bourguet, E.; Rouhier, N.; Rey, P. Specificity of thioredoxins and glutaredoxins as electron donors to two distinct classes of Arabidopsis plastidial methionine sulfoxide reductases B. FEBS Lett. 2007, 581, 4371–4376. [Google Scholar] [CrossRef] [PubMed]
- Ding, D.; Sagher, D.; Laugier, E.; Rey, P.; Weissbach, H.; Zhang, X.H. Studies on the reducing systems for plant and animal methionine sulfoxide reductases B lacking the resolving cysteine. Biochem. Biophys. Res. Commun. 2007, 361, 629–633. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.Y.; Kim, J.R. Thioredoxin as a reducing agent for mammalian methionine sulfoxide reductases B lacking resolving cysteine. Biochem. Biophys. Res. Commun. 2008, 371, 490–494. [Google Scholar] [CrossRef] [PubMed]
- Tarrago, L.; Laugier, E.; Zaffagnini, M.; Marchand, C.; Le Maréchal, P.; Lemaire, S.D.; Rey, P. Plant thioredoxin CDSP32 regenerates 1-Cys methionine sulfoxide reductase B activity through the direct reduction of sulfenic acid. J. Biol. Chem. 2010, 285, 14964–14972. [Google Scholar] [CrossRef] [PubMed]
- Couturier, J.; Vignols, F.; Jacquot, J.P.; Rouhier, N. Glutathione- and glutaredoxin-dependent reduction of methionine sulfoxide reductase A. FEBS Lett. 2012, 586, 3894–3899. [Google Scholar] [CrossRef] [PubMed]
- Le, D.T.; Nguyen, K.; Chu, H.D.; Vu, N.T.; Pham, T.T.L.; Tran, L.P. Function of the evolutionarily conserved plant methionine-S-sulfoxide reductase without the catalytic residue. Protoplasma 2018, 1–10. [Google Scholar] [CrossRef] [PubMed]
- Koc, A.; Gasch, A.P.; Rutherford, J.C.; Kim, H.Y.; Gladyshev, V.N. Methionine sulfoxide reductase regulation of yeast lifespan reveals reactive oxygen species-dependent and -independent components of aging. Proc. Natl. Acad. Sci. USA 2004, 101, 7999–8004. [Google Scholar] [CrossRef] [PubMed]
- Gabbita, S.P.; Aksenov, M.Y.; Lovell, M.A.; Markesbery, W.R. Decrease in peptide methionine sulfoxide reductase in Alzheimer’s disease brain. J. Neurochem. 1999, 73, 1660–1666. [Google Scholar] [CrossRef] [PubMed]
- Petropoulos, I.; Mary, J.; Perichon, M.; Friguet, B. Rat peptide methionine sulphoxide reductase: Cloning of the cDNA, and down-regulation of gene expression and enzyme activity during aging. Biochem. J. 2001, 355 Pt 3, 819–825. [Google Scholar] [CrossRef]
- Moskovitz, J.; Flescher, E.; Berlett, B.S.; Azare, J.; Poston, J.M.; Stadtman, E.R. Overexpression of peptide-methionine sulfoxide reductase in Saccharomyces cerevisiae and human T cells provides them with high resistance to oxidative stress. Proc. Natl. Acad. Sci. USA 1998, 95, 14071–14075. [Google Scholar] [CrossRef] [PubMed]
- Ruan, H.; Tang, X.D.; Chen, M.L.; Joiner, M.L.; Sun, G.; Brot, N.; Weissbach, H.; Heinemann, S.H.; Iverson, L.; Wu, C.F.; et al. High-quality life extension by the enzyme peptide methionine sulfoxide reductase. Proc. Natl. Acad. Sci. USA 2002, 99, 2748–2753. [Google Scholar] [CrossRef] [PubMed]
- Stadtman, E.R.; Moskovitz, J.; Berlett, B.S.; Levine, R.L. Cyclic oxidation and reduction of protein methionine residues is an important antioxidant mechanism. Mol. Cell Biochem. 2002, 234–235, 3–9. [Google Scholar] [CrossRef]
- Drazic, A.; Miura, H.; Peschek, J.; Le, Y.; Bach, N.C.; Kriehuber, T.; Winter, J. Methionine oxidation activates a transcription factor in response to oxidative stress. Proc. Natl. Acad. Sci. USA 2013, 110, 9493–9498. [Google Scholar] [CrossRef] [PubMed]
- Erickson, J.R.; Joiner, M.L.; Guan, X.; Kutschke, W.; Yang, J.; Oddis, C.V.; Bartlett, R.K.; Lowe, J.S.; O’Donnell, S.E.; Aykin-Burns, N.; et al. A dynamic pathway for calcium-independent activation of CaMKII by methionine oxidation. Cell 2008, 133, 462–474. [Google Scholar] [CrossRef] [PubMed]
- Lee, B.C.; Péterfi, Z.; Hoffmann, F.W.; Moore, R.E.; Kaya, A.; Avanesov, A.; Tarrago, L.; Zhou, Y.; Weerapana, E.; Fomenko, D.E.; et al. MsrB1 and MICALs regulate actin assembly and macrophage function via reversible stereoselective methionine oxidation. Mol. Cell 2013, 51, 397–404. [Google Scholar] [CrossRef] [PubMed]
- Hung, R.J.; Spaeth, C.S.; Yesilyurt, H.G.; Terman, J.R. SelR reverses Mical-mediated oxidation of actin to regulate F-actin dynamics. Nat. Cell Biol. 2013, 15, 1445–1454. [Google Scholar] [CrossRef] [PubMed]
- Sanchez, J.; Nikolau, B.J.; Stumpf, P.K. Reduction of N-Acetyl methionine sulfoxide in plants. Plant Physiol. 1983, 73, 619–623. [Google Scholar] [CrossRef] [PubMed]
- Sadanandom, A.; Piffanelli, P.; Knott, T.; Robinson, C.; Sharpe, A.; Lydiate, D.; Murphy, D.; Fairbairn, D.J. Identification of a peptide methionine sulphoxide reductase gene in an oleosin promoter from Brassica napus. Plant J. 1996, 10, 235–242. [Google Scholar] [CrossRef] [PubMed]
- Rodrigo, M.J.; Moskovitz, J.; Salamini, F.; Bartels, D. Reverse genetic approaches in plants and yeast suggest a role for novel, evolutionarily conserved, selenoprotein-related genes in oxidative stress defense. Mol. Genet. Genom. 2002, 267, 613–621. [Google Scholar] [CrossRef] [PubMed]
- Rouhier, N.; Vieira Dos Santos, C.; Tarrago, L.; Rey, P. Plant methionine sulfoxide reductase A and B multigenic families. Photosynth. Res. 2006, 89, 247–262. [Google Scholar] [CrossRef] [PubMed]
- Tarrago, L.; Laugier, E.; Rey, P. Protein-repairing methionine sulfoxide reductases in photosynthetic organisms, gene organization, reduction mechanisms, and physiological roles. Mol. Plant 2009, 2, 202–217. [Google Scholar] [CrossRef] [PubMed]
- Zhu, J.; Ding, P.; Li, Q.; Gao, Y.; Chen, F.; Xia, G. Molecular characterization and expression profile of methionine sulfoxide reductase gene family in maize (Zea mays) under abiotic stresses. Gene 2015, 562, 159–168. [Google Scholar] [CrossRef] [PubMed]
- Sadanandom, A.; Poghosyan, Z.; Fairbairn, D.J.; Murphy, D.J. Differential regulation of plastidial and cytosolic isoforms of peptide methionine sulfoxide reductase in Arabidopsis. Plant Physiol. 2000, 123, 255–264. [Google Scholar] [CrossRef] [PubMed]
- Vieira Dos Santos, C.; Cuine, S.; Rouhier, N.; Rey, P. The Arabidopsis plastidic methionine sulfoxide reductase B proteins. Sequence and activity characteristics, comparison of the expression with plastidic methionine sulfoxide reductase A, and induction by photooxidative stress. Plant Physiol. 2005, 138, 909–922. [Google Scholar] [CrossRef] [PubMed]
- Li, C.W.; Lee, S.H.; Chieh, P.S.; Lin, C.S.; Wang, Y.C.; Chan, M.T. Arabidopsis root-abundant cytosolic methionine sulfoxide reductase B genes MsrB7 and MsrB8 are involved in tolerance to oxidative stress. Plant Cell Physiol. 2012, 53, 1707–1719. [Google Scholar] [CrossRef] [PubMed]
- Kwon, S.J.; Kwon, S.I.; Bae, M.S.; Cho, E.J.; Park, O.K. Role of the methionine sulfoxide reductase MsrB3 in cold acclimation in Arabidopsis. Plant Cell Physiol. 2007, 48, 1713–1723. [Google Scholar] [CrossRef] [PubMed]
- Dai, C.; Liu, L.; Wang, M.H. Characterization of a methionine sulfoxide reductase B from tomato (Solanum lycopersicum), and its protecting role in Saccharomyces cerevisiae. Protein J. 2013, 32, 39–47. [Google Scholar] [CrossRef] [PubMed]
- Gao, L.; Shen, W.; Yan, P.; Tuo, D.; Li, X.; Zhou, P. NIa-pro of Papaya ringspot virus interacts with papaya methionine sulfoxide reductase B1. Virology 2012, 434, 78–87. [Google Scholar] [CrossRef] [PubMed]
- Guo, X.; Wu, Y.; Wang, Y.; Chen, Y.; Chu, C. OsMSRA4.1 and OsMSRB1.1, two rice plastidial methionine sulfoxide isoforms, are involved in abiotic stress responses. Planta 2009, 230, 227–238. [Google Scholar] [CrossRef] [PubMed]
- Oh, S.K.; Baek, K.H.; Seong, E.S.; Joung, Y.H.; Choi, G.J.; Park, J.M.; Cho, H.S.; Kim, E.A.; Lee, S.; Choi, D. MsrB2, pepper methionine sulfoxide reductase B2, is a novel defense regulator against oxidative stress and pathogen attack. Plant Physiol. 2010, 154, 245–261. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.S.; Park, H.M.; Chae, S.; Lee, T.H.; Hwang, D.J.; Oh, S.D.; Park, J.S.; Song, D.G.; Pan, C.H.; Choi, D.; et al. A pepper MSRB2 gene confers drought tolerance in rice through the protection of chloroplast-targeted genes. PLoS ONE 2014, 9, e90588. [Google Scholar] [CrossRef] [PubMed]
- Liu, L.; Wang, M.H. Cloning, expression, and characterization of a methionine sulfoxide reductase B gene from Nicotiana tabacum. Protein J. 2013, 32, 543–550. [Google Scholar] [CrossRef] [PubMed]
- Liu, L.; Wang, M.H. Expression and biological properties of a novel methionine sulfoxide reductase A in tobacco (Nicotiana tabacum). Protein J. 2013, 32, 266–274. [Google Scholar] [CrossRef] [PubMed]
- Sun, X.; Sun, M.; Jia, B.; Qin, Z.; Yang, K.; Chen, C.; Yu, Q.; Zhu, Y. A Glycine soja methionine sulfoxide reductase B5a interacts with the Ca(2+)/CAM-binding kinase GsCBRLK and activates ROS signaling under carbonate alkaline stress. Plant J. 2016, 86, 514–529. [Google Scholar] [CrossRef] [PubMed]
- Jiang, G.; Xiao, L.; Yan, H.; Zhang, D.; Wu, F.; Liu, X.; Su, X.; Dong, X.; Wang, J.; Duan, X.; et al. Redox regulation of methionine in calmodulin affects the activity levels of senescence-related transcription factors in litchi. Biochim. Biophys. Acta 2017, 1861, 1140–1151. [Google Scholar] [CrossRef] [PubMed]
- Jiang, G.; Wu, F.; Li, Z.; Li, T.; Gupta, V.K.; Duan, X.; Jiang, Y. Sulfoxidation regulation of Musa acuminata calmodulin (MaCaM) influences the functions of MaCaM-binding proteins. Plant Cell Physiol. 2018, 59, 1214–1224. [Google Scholar] [CrossRef] [PubMed]
- Klodmann, J.; Senkler, M.; Rode, C.; Braun, H.P. Defining the protein complex proteome of plant mitochondria. Plant Physiol. 2011, 157, 587–598. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.; Lei, Z.; Watson, B.S.; Sumner, L.W. Sub-cellular proteomics of Medicago truncatula. Front. Plant Sci. 2013, 4, 112. [Google Scholar] [CrossRef] [PubMed]
- Senkler, J.; Senkler, M.; Eubel, H.; Hildebrandt, T.; Lengwenus, C.; Schertl, P.; Schwarzländer, M.; Wagner, S.; Wittig, I.; Braun, H.P. The mitochondrial complexome of Arabidopsis thaliana. Plant J. 2017, 89, 1079–1092. [Google Scholar] [CrossRef] [PubMed]
- Noctor, G.; Reichheld, J.P.; Foyer, C.H. ROS-related redox regulation and signaling in plants. Semin. Cell Dev. Biol. 2017, 80, 3–12. [Google Scholar] [CrossRef] [PubMed]
- In, O.; Berberich, T.; Romdhane, S.; Feierabend, J. Changes in gene expression during dehardening of cold-hardened winter rye (Secale cereale L.) leaves and potential role of a peptide methionine sulfoxide reductase in cold-acclimation. Planta 2005, 220, 941–950. [Google Scholar] [CrossRef] [PubMed]
- Châtelain, E.; Satour, P.; Laugier, E.; Ly Vu, B.; Payet, N.; Rey, P.; Montrichard, F. Evidence for the participation of the methionine sulfoxide reductase repair system in plant seed longevity. Proc. Natl. Acad. Sci. USA 2013, 110, 3633–3638. [Google Scholar] [CrossRef] [PubMed]
- Chu, H.D.; Nguyen, K.L.; Watanabe, Y.; Le, D.T.; Tran, L.P. Expression analyses of soybean genes encoding methionine-R-sulfoxide reductase under various conditions suggest a possible role in the adaptation to stress. Appl. Biol. Chem. 2016, 59, 681–687. [Google Scholar] [CrossRef]
- Xiao, T.; Memgmeng, M.; Wang, G.; Quian, M.; Chen, Y.; Zheng, L.; Zhang, H.; Hu, Z.; Shen, Z.; Xia, Y. A methionine-R-sulfoxide reductase, OsMSRB5, is required for rice defense against copper toxicity. Environ. Exp. Bot. 2018, 153, 45–53. [Google Scholar] [CrossRef]
- Lopez, A.P.; Portales, R.; López-Ráez, J.; Medina-Escobar, N.; Blanco, J.; Franco, A. Characterization of a strawberry late-expressed and fruit-specific peptide methionine sulphoxide reductase. Physiol. Plant 2006, 126, 129–139. [Google Scholar] [CrossRef]
- Allen, M.D.; Kropat, J.; Tottey, S.; Del Campo, J.A.; Merchant, S.S. Manganese deficiency in Chlamydomonas results in loss of photosystem II and MnSOD function, sensitivity to peroxides, and secondary phosphorus and iron deficiency. Plant Physiol. 2007, 143, 263–277. [Google Scholar] [CrossRef] [PubMed]
- Wu, T.M.; Hsu, Y.T.; Sung, M.S.; Hsu, Y.T.; Lee, T.M. Expression of genes involved in redox homeostasis and antioxidant defense in a marine macroalga Ulva fasciata by excess copper. Aquat. Toxicol. 2009, 94, 275–285. [Google Scholar] [CrossRef] [PubMed]
- Zhao, L.; Chen, M.; Cheng, D.; Yang, H.; Sun, Y.; Zhou, H.; Huang, F. Different B-type methionine sulfoxide reductases in Chlamydomonas may protect the alga against high-light, sulfur-depletion, or oxidative stress. J. Integr. Plant Biol. 2013, 55, 1054–1068. [Google Scholar] [CrossRef] [PubMed]
- Hsu, Y.T.; Lee, T.M. Photosynthetic electron transport mediates the light-ciontrolled up-regulation of expression of methionine sulfoxide reductase A and B from marine macroalga Ulva fasciata. J. Phycol. 2010, 46, 112–122. [Google Scholar] [CrossRef]
- Romero, H.M.; Berlett, B.S.; Jensen, P.J.; Pell, E.J.; Tien, M. Investigations into the role of the plastidial peptide methionine sulfoxide reductase in response to oxidative stress in Arabidopsis. Plant Physiol. 2004, 136, 3784–3794. [Google Scholar] [CrossRef] [PubMed]
- Dai, C.; Singh, N.K.; Park, M. Characterization of a novel methionine sulfoxide reductase A from tomato (Solanum lycopersicum), and its protecting role in Escherichia coli. BMB Rep. 2011, 44, 805–810. [Google Scholar] [CrossRef] [PubMed]
- Méndez, A.A.; Pena, L.B.; Benavides, M.P.; Gallego, S.M. Priming with NO controls redox state and prevents cadmium-induced general up-regulation of methionine sulfoxide reductase gene family in Arabidopsis. Biochimie 2016, 131, 128–136. [Google Scholar] [CrossRef] [PubMed]
- Collin, V.C.; Eymery, F.; Genty, B.; Rey, P.; Havaux, M. Vitamin E is essential for the tolerance of Arabidopsis thaliana to metal-induced oxidative stress. Plant Cell Environ. 2008, 31, 244–257. [Google Scholar] [CrossRef] [PubMed]
- Alvarez, S.; Berla, B.M.; Sheffield, J.; Cahoon, R.E.; Jez, J.M.; Hicks, L.M. Comprehensive analysis of the Brassica juncea root proteome in response to cadmium exposure by complementary proteomic approaches. Proteomics 2009, 9, 2419–2431. [Google Scholar] [CrossRef] [PubMed]
- Simonović, A.D.; Anderson, M.D. Analysis of methionine oxides and nitrogen-transporting amino acids in chilled and acclimated maize seedlings. Amino Acids 2007, 33, 607–613. [Google Scholar] [CrossRef] [PubMed]
- Dai, C.; Wang, M.H. Characterization and functional analysis of methionine sulfoxide reductase A gene family in tomato. Mol. Biol. Rep. 2012, 39, 6297–6308. [Google Scholar] [CrossRef] [PubMed]
- Bouchenak, F.; Henri, P.; Benrebiha, F.Z.; Rey, P. Differential responses to salinity of two Atriplex halimus populations in relation to organic solutes and antioxidant systems involving thiol reductases. J. Plant Physiol. 2012, 169, 1445–1553. [Google Scholar] [CrossRef] [PubMed]
- Fatehi, F.; Hosseinzadeh, A.; Alizadeh, H.; Brimavandi, T.; Struik, P.C. The proteome response of salt-resistant and salt-sensitive barley genotypes to long-term salinity stress. Mol. Biol. Rep. 2012, 39, 6387–6397. [Google Scholar] [CrossRef] [PubMed]
- Marok, M.A.; Tarrago, L.; Ksas, B.; Henri, P.; Abrous-Belbachir, O.; Havaux, M.; Rey, P. A drought-sensitive barley variety displays oxidative stress and strongly increased contents in low-molecular weight antioxidant compounds during water deficit compared to a tolerant variety. J. Plant Physiol. 2013, 170, 633–645. [Google Scholar] [CrossRef] [PubMed]
- Roy, S.; Nandi, A.K. Arabidopsis thaliana methionine sulfoxide reductase B8 influences stress-induced cell death and effector-triggered immunity. Plant Mol. Biol. 2017, 93, 109–120. [Google Scholar] [CrossRef] [PubMed]
- Vieira Dos Santos, C.; Delavault, P.; Letousey, P.; Thalouarn, P. Identification by suppression subtractive hybridization and expression analysis of Arabidopsis thaliana putative defence genes during Orobanche ramosa infection. Physiol. Mol. Plant Pathol. 2003, 62, 297–303. [Google Scholar] [CrossRef]
- Choudhury, F.K.; Rivero, R.M.; Blumwald, E.; Mittler, R. Reactive oxygen species, abiotic stress and stress combination. Plant J. 2017, 90, 856–867. [Google Scholar] [CrossRef] [PubMed]
- Yu, M.; Lamattina, L.; Spoel, S.H.; Loake, G.J. Nitric oxide function in plant biology: A redox cue in deconvolution. New Phytol. 2014, 202, 1142–1156. [Google Scholar] [CrossRef] [PubMed]
- Foyer, C.; Noctor, G. Stress-triggered redox signalling: What’s in pROSpect? Plant Cell Environ. 2016, 39, 951–964. [Google Scholar] [CrossRef] [PubMed]
- Umbreen, S.; Lubega, J.; Cui, B.; Pan, Q.; Jiang, J.; Loake, G.J. Specificity in nitric oxide signaling. J. Exp. Bot. 2018, 69, 3439–3448. [Google Scholar] [CrossRef] [PubMed]
- D’Autréaux, B.; Toledano, M.B. ROS as signalling molecules: Mechanisms that generate specificity in ROS homeostasis. Nat. Rev. Mol. Cell Biol. 2007, 10, 813–824. [Google Scholar] [CrossRef] [PubMed]
- Frederickson Matika, D.E.; Loake, G.J. Redox regulation in plant immune function. Antioxid. Redox Signal. 2014, 21, 1373–1388. [Google Scholar] [CrossRef] [PubMed]
- Chang, H.L.; Tseng, Y.L.; Ho, K.L.; Shie, S.C.; Wu, P.S.; Hsu, Y.T.; Lee, T.M. Reactive oxygen species modulate the differential expression of methionine sulfoxide reductasegenes in Chlamydomonas reinhardtii under high light illumination. Physiol. Plant 2014, 150, 550–564. [Google Scholar] [CrossRef] [PubMed]
- Hsu, Y.T.; Lee, T.M. Nitric oxide up-regulates the expression of methionine sulfoxide reductase genes in the intertidal macroalga Ulva fasciata for high light acclimation. Plant Cell Physiol. 2012, 53, 445–456. [Google Scholar] [CrossRef] [PubMed]
- Oh, J.E.; Hong, S.W.; Lee, Y.; Koh, E.J.; Kim, K.; Seo, Y.W.; Chung, N.; Jeong, M.; Jang, C.S.; Lee, B.; et al. Modulation of gene expressions and enzyme activities of methionine sulfoxide reductases by cold, ABA or high salt treatments in Arabidopsis. Plant Sci. 2005, 169, 1030–1036. [Google Scholar] [CrossRef]
- Danon, A.; Coll, N.S.; Apel, K. Cryptochrome-1-dependent execution of programmed cell death induced by singlet oxygen in Arabidopsis thaliana. Proc. Natl. Acad. Sci. USA 2006, 103, 17036–17041. [Google Scholar] [CrossRef] [PubMed]
- Begara-Morales, J.C.; Sánchez-Calvo, B.; Luque, F.; Leyva-Pérez, M.O.; Leterrier, M.; Corpas, F.J.; Barroso, J.B. Differential transcriptomic analysis by RNA-Seq of GSNO-responsive genes between Arabidopsis roots and leaves. Plant Cell Physiol. 2014, 55, 1080–1095. [Google Scholar] [CrossRef] [PubMed]
- Xia, X.J.; Zhou, Y.H.; Shi, K.; Zhou, J.; Foyer, C.H.; Yu, J.Q. Interplay between reactive oxygen species and hormones in the control of plant development and stress tolerance. J. Exp. Bot. 2015, 66, 2839–2856. [Google Scholar] [CrossRef] [PubMed]
- Mata-Pérez, C.; Sánchez-Calvo, B.; Begara-Morales, J.C.; Luque, F.; Jiménez-Ruiz, J.; Padilla, M.N.; Fierro-Risco, J.; Valderrama, R.; Fernández-Ocaña, A.; Corpas, F.J.; et al. Yanscriptomic profiling of linolenic acid-responsive genes in ROS signaling from RNA-seq data in Arabidopsis. Front. Plant Sci. 2015, 6, 122. [Google Scholar] [CrossRef] [PubMed]
- Wasternack, C.; Hause, B. Jasmonates: Biosynthesis, perception, signal transduction and action in plant stress response, growth and development. An update to the 2007 review in Annals of Botany. Ann. Bot. 2013, 111, 1021–1058. [Google Scholar] [CrossRef] [PubMed]
- Yan, S.; Dong, X. Perception of the plant immune signal salicylic acid. Curr. Opin. Plant Biol. 2014, 20, 64–68. [Google Scholar] [CrossRef] [PubMed]
- Zhu, J.K. Salt and drought stress signal transduction in plants. Annu. Rev. Plant Biol. 2002, 53, 247–273. [Google Scholar] [CrossRef] [PubMed]
- Ferguson, D.L.; Burke, J.J. A new method of measuring protein-methionine-s-oxide reductase activity. Plant Physiol. 1992, 100, 529–532. [Google Scholar] [CrossRef] [PubMed]
- Laugier, E.; Tarrago, L.; Vieira Dos Santos, C.; Eymery, F.; Havaux, M.; Rey, P. Arabidopsis thaliana plastidial methionine sulfoxide reductases B, MSRBs, account for most leaf peptide MSR activity and are essential for growth under environmental constraints through a role in the preservation of photosystem antennae. Plant J. 2010, 61, 271–282. [Google Scholar] [CrossRef] [PubMed]
- Laugier, E.; Tarrago, L.; Courteille, A.; Innocenti, G.; Eymery, F.; Rumeau, D.; Issakidis-Bourguet, E.; Rey, P. Involvement of thioredoxin y2 in the preservation of leaf methionine sulfoxide reductase capacity and growth under high light. Plant Cell Envirion. 2013, 36, 670–682. [Google Scholar] [CrossRef] [PubMed]
- Bechtold, U.; Murphy, D.J.; Mullineaux, P.M. Arabidopsis peptide methionine sulfoxide reductase2 prevents cellular oxidative damage in long nights. Plant Cell 2004, 16, 908–919. [Google Scholar] [CrossRef] [PubMed]
- Ferguson, D.L.; Burke, J.J. Methionyl sulfoxide content and protein-methionine-S-oxide reductase activity in response to water deficits or high temperature. Physiol. Plant 1994, 90, 253–258. [Google Scholar] [CrossRef]
- Bechtold, U.; Rabbani, N.; Mullineaux, P.M.; Thornalley, P.J. Quantitative measurement of specific biomarkers for protein oxidation, nitration and glycation in Arabidopsis leaves. Plant J. 2009, 59, 661–671. [Google Scholar] [CrossRef] [PubMed]
- Jacques, S.; Ghesquière, B.; De Bock, P.J.; Demol, H.; Wahni, K.; Willems, P.; Messens, J.; Van Breusegem, F.; Gevaert, K. Protein methionine sulfoxide dynamics in Arabidopsis thaliana under oxidative stress. Mol. Cell Proteom. 2015, 14, 1217–1229. [Google Scholar] [CrossRef] [PubMed]
- Marondedze, C.; Turek, I.; Parrott, B.; Thomas, L.; Jankovic, B.; Lilley, K.S.; Gehring, C. Structural and functional characteristics of cGMP-dependent methionine oxidation in Arabidopsis thaliana proteins. Cell Commun. Signal. 2013, 11, 1. [Google Scholar] [CrossRef] [PubMed]
- Oh, J.E.; Hossain Md, A.; Kim, J.H.; Chu, S.H.; Lee, E.H.; Hwang, K.Y.; Noh, H.; Hong, S.W.; Lee, H. The ectopic expression of methionine sulfoxide reductase 4 results in enhanced growth performance in arabidopsis. Plant Sci. 2010, 178, 265–270. [Google Scholar] [CrossRef]
- Han, Y.; Du, Y.; Wang, J.; Wu, T. Overexpression of Chinese flowering cabbage BpPMSR3 enhances the tolerance of Arabidopsis thaliana to cadmium. J. Plant Nutr. Soil Sci. 2018. [Google Scholar] [CrossRef]
- Siddiqui, Z.S.; Cho, J.; Kwon, T.R.; Ahn, B.O.; Lee, S.B.; Jeong, M.J.; Ryu, T.H.; Lee, S.K.; Park, S.C.; Park, S.H. Physiological mechanism of drought tolerance in transgenic rice plants expressing Capsicum annuum methionine sulfoxide reductase B2 (CaMsrB2) gene. Acta Physiol. Plant 2014, 36, 1143–1153. [Google Scholar] [CrossRef]
- El Hassouni, M.E.; Chambost, J.P.; Expert, D.; Van Gijsegem, F.; Barras, F. The minimal gene set member msrA, encoding peptide methionine sulfoxide reductase, is a virulence determinant of the plant pathogen Erwinia chrysanthemi. Proc. Natl. Acad. Sci. USA 1999, 96, 887–892. [Google Scholar] [CrossRef]
- Gustavsson, N.; Kokke, B.; Härndahl, U.; Silow, M.; Bechtold, U.; Poghosyan, Z.; Murphy, D.; Boelens, W.; Sundby, C. A peptide methionine sulfoxide reductase highly expressed in photosynthetic tissue in Arabidopsis thaliana can protect the chaperone-like activity of a chloroplast-localized small heat shock protein. Plant J. 2002, 29, 545–553. [Google Scholar] [CrossRef] [PubMed]
- Ezraty, B.; Grimaud, R.; El Hassouni, M.; Moinier, D.; Barras, F. Methionine sulfoxide reductases protect Ffh from oxidative damages in Escherichia coli. EMBO J. 2004, 23, 1868–1877. [Google Scholar] [CrossRef] [PubMed]
- Chu, H.D.; Le, Q.N.; Nguyen, H.Q.; Le, D.T. Genome-wide analysis of genes encoding methionine-rich proteins in Arabidopsis and soybean suggesting their roles in the adaptation of plants to abiotic stress. Int. J. Genom. 2016, 2016, 5427062. [Google Scholar] [CrossRef]
- Wehr, N.B.; Levine, R.L. Wanted and wanting: Antibody against methionine sulfoxide. Free Radic. Biol. Med. 2012, 53, 1222–1225. [Google Scholar] [CrossRef] [PubMed]
- Ghesquière, B.; Gevaert, K. Proteomics methods to study methionine oxidation. Mass Spectrom. Rev. 2014, 33, 147–156. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.H.; Li, C.W.; Koh, K.W.; Chuang, H.Y.; Chen, Y.R.; Lin, C.S.; Chan, M.T. MSRB7 reverses oxidation of GSTF2/3 to confer tolerance of Arabidopsis thaliana to oxidative stress. J. Exp. Bot. 2014, 65, 5049–5062. [Google Scholar] [CrossRef] [PubMed]
- Tarrago, L.; Kieffer-Jaquinod, S.; Lamant, T.; Marcellin, M.; Garin, J.; Rouhier, N.; Rey, P. Affinity chromatography: A valuable strategy to isolate substrates of methionine sulfoxide reductases? Antioxid. Redox Signal. 2012, 16, 79–84. [Google Scholar] [CrossRef] [PubMed]
- Sundby, C.; Harndahl, U.; Gustavsson, N.; Ahrman, E.; Murphy, D.J. Conserved methionines in chloroplasts. Biochim. Biophys. Acta 2005, 1703, 191–202. [Google Scholar] [CrossRef] [PubMed]
- Lallement, P.A.; Brouwer, B.; Keech, O.; Hecker, A.; Rouhier, N. The still mysterious roles of cysteine-containing glutathione transferases in plants. Front. Pharmacol. 2014, 5, 192. [Google Scholar] [CrossRef] [PubMed]
- Martínez-Márquez, A.; Martínez-Esteso, M.J.; Vilella-Antón, M.T.; Sellés-Marchart, S.; Morante-Carriel, J.A.; Hurtado, E.; Palazon, J.; Bru-Martínez, R.A. Tau Class Glutathione-S-Transferase is Involved in Trans-Resveratrol Transport Out of Grapevine Cells. Front. Plant Sci. 2017, 8, 1457. [Google Scholar] [CrossRef] [PubMed]
- Pégeot, H.; Mathiot, S.; Perrot, T.; Gense, F.; Hecker, A.; Didierjean, C.; Rouhier, N. Structural plasticity among glutathione transferase Phi members: Natural combination of catalytic residues confers dual biochemical activities. FEBS J. 2017, 284, 2442–2463. [Google Scholar] [CrossRef] [PubMed]
- Tossounian, M.A.; Van Molle, I.; Wahni, K.; Jacques, S.; Gevaert, K.; Van Breusegem, F.; Vertommen, D.; Young, D.; Rosado, L.A.; Messens, J. Disulfide bond formation protects Arabidopsis thaliana glutathione transferase tau 23 from oxidative damage. Biochim. Biophys. Acta 2018, 1862, 775–789. [Google Scholar] [CrossRef] [PubMed]
- Tossounian, M.A.; Wahni, K.; Van Molle, I.; Vertommen, D.; Astolfi Rosado, L.; Messens, J. Redox regulated methionine oxidation of Arabidopsis thaliana glutathione transferase Phi9 induces H-site flexibility. Protein Sci. 2018. [Google Scholar] [CrossRef] [PubMed]
- Benoit, S.L.; Bayyareddy, K.; Mahawar, M.; Sharp, J.S.; Maier, R.J. Alkyl hydroperoxide reductase repair by Helicobacter pylori methionine sulfoxide reductase. J. Bacteriol. 2013, 195, 5396–5401. [Google Scholar] [CrossRef] [PubMed]
- O’Neil, K.T.; DeGrado, W.F. How calmodulin binds its targets: Sequence independent recognition of amphiphilic alpha-helices. Trends Biochem. Sci. 1990, 15, 59–64. [Google Scholar] [CrossRef]
- Bernstein, H.D.; Poritz, M.A.; Strub, K.; Hoben, P.J.; Brenner, S.; Walter, P. Model for signal sequence recognition from amino-acid sequence of 54K subunit of signal recognition particle. Nature 1989, 340, 482–486. [Google Scholar] [CrossRef] [PubMed]
- Xiong, Y.; Chen, B.; Smallwood, H.S.; Urbauer, R.J.; Markille, L.M.; Galeva, N.; Williams, T.D.; Squier, T.C. High-affinity and cooperative binding of oxidized calmodulin by methionine sulfoxide reductase. Biochemistry 2006, 45, 14642–14654. [Google Scholar] [CrossRef] [PubMed]
- Bigelow, D.J.; Squier, T.C. Redox modulation of cellular signaling and metabolism through reversible oxidation of methionine sensors in calcium regulatory proteins. Biochim. Biophys. Acta 2005, 1703, 121–134. [Google Scholar] [CrossRef] [PubMed]
- Carruthers, N.J.; Stemmer, P.M. Methionine oxidation in the calmodulin-binding domain of calcineurin disrupts calmodulin binding and calcineurin activation. Biochemistry 2008, 47, 3085–3095. [Google Scholar] [CrossRef] [PubMed]
- Bigelow, D.J.; Squier, T.C. Thioredoxin-dependent redox regulation of cellular signaling and stress response through reversible oxidation of methionines. Mol. Biosyst. 2011, 7, 2101–2109. [Google Scholar] [CrossRef] [PubMed]
- Snijder, J.; Rose, R.J.; Raijmakers, R.; Heck, A.J. Site-specific methionine oxidation in calmodulin affects structural integrity and interaction with Ca2+/calmodulin-dependent protein kinase II. J. Struct. Biol. 2011, 174, 187–195. [Google Scholar] [CrossRef] [PubMed]
- Bouché, N.; Yellin, A.; Snedden, W.A.; Fromm, H. Plant-specific calmodulin-binding proteins. Annu. Rev. Plant Biol. 2005, 56, 435–466. [Google Scholar] [CrossRef] [PubMed]
- Yang, T.; Poovaiah, B.W. Calcium/calmodulin-mediated signal network in plants. Trends Plant Sci. 2003, 8, 505–512. [Google Scholar] [CrossRef] [PubMed]
- Hardin, S.C.; Larue, C.T.; Oh, M.H.; Jain, V.; Huber, S.C. Coupling oxidative signals to protein phosphorylation via methionine oxidation in Arabidopsis. Biochem. J. 2009, 422, 305–312. [Google Scholar] [CrossRef] [PubMed]
- Veredas, F.J.; Cantón, F.R.; Aledo, J.C. Methionine residues around phosphorylation sites are preferentially oxidized in vivo under stress conditions. Sci. Rep. 2017, 7, 40403. [Google Scholar] [CrossRef] [PubMed]
- Lim, J.C.; You, Z.; Kim, G.; Levine, R.L. Methionine sulfoxide reductase A is a stereospecific methionine oxidase. Proc. Natl. Acad. Sci. USA 2011, 108, 10472–10477. [Google Scholar] [CrossRef] [PubMed]
| Condition | Variation in MSR Expression. Species | References |
|---|---|---|
| Abiotic constraints | ||
| High light | ↗ A. thaliana | [76] |
| ↗ S. cereale | [67] | |
| High light/low temperature | ↗ A. thaliana | [50] |
| Low temperature | ↗ N. tabacum, S. lycopersicum | [59,82] |
| ↗ O. sativa, S. cereale, Z. mays | [55,67,81] | |
| Water deficit | ↗ G. max, N. tabacum | [58,69] |
| High salt (NaCl) | ↗ A. halimus, A. thaliana, N. tabacum, S. lycopersicum | [58,59,82,83,96] |
| ↗ H. vulgare, O. sativa, Z. mays | [48,55,84] | |
| ↗ ↘ G. max | [69] | |
| High carbonate | ↗ G. soja | [60] |
| Cadmium | ↗ ↘ A. thaliana, B. juncea | [78,79,80] |
| Biotic constraints | ||
| Virus | ↗ A. thaliana, C. papaya | [49,54] |
| Bacteria | ↗ A. thaliana | [86] |
| ↘ C. annuum | [56] | |
| Fungi | ↗ ↘ Populus × interamericana | [50] |
| Parasite plants | ↗ A. thaliana | [87] |
| Oxidative treatments | ||
| Methyl viologen | ↗ A. thaliana, N. tabacum, S. lycopersicum | [51,59,76,82] |
| ↗ O. sativa, S. cereale | [55,67] | |
| Hydrogen peroxide | ↗ ↘ S. lycopersicum | [82] |
| ↗ A. thaliana | [96] | |
| Singlet oxygen | ↗ A. thaliana | [97] |
| S-nitrosoglutathione | ↗ A. thaliana | [98] |
| Copper excess | ↗ O. sativa | [70] |
| Hormone treatments | ||
| Abscisic acid | ↗ ↘ A. thaliana, G. max, N. tabacum, S. lycopersicum | [58,59,69,82,96] |
| Jasmonic acid | ↗ ↘ A. thaliana, S. lycopersicum | [82,96,100] |
| ↘ C. annuum | [56] | |
| Salicylic acid | ↗ ↘ S. lycopersicum | [82] |
| ↘ C. annuum | [56] | |
| Ethylene | ↗ S. lycopersicum | [82] |
| ↗ M. acuminata | [62] |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Rey, P.; Tarrago, L. Physiological Roles of Plant Methionine Sulfoxide Reductases in Redox Homeostasis and Signaling. Antioxidants 2018, 7, 114. https://doi.org/10.3390/antiox7090114
Rey P, Tarrago L. Physiological Roles of Plant Methionine Sulfoxide Reductases in Redox Homeostasis and Signaling. Antioxidants. 2018; 7(9):114. https://doi.org/10.3390/antiox7090114
Chicago/Turabian StyleRey, Pascal, and Lionel Tarrago. 2018. "Physiological Roles of Plant Methionine Sulfoxide Reductases in Redox Homeostasis and Signaling" Antioxidants 7, no. 9: 114. https://doi.org/10.3390/antiox7090114
APA StyleRey, P., & Tarrago, L. (2018). Physiological Roles of Plant Methionine Sulfoxide Reductases in Redox Homeostasis and Signaling. Antioxidants, 7(9), 114. https://doi.org/10.3390/antiox7090114





