Using Integrated Bioinformatics Analysis to Identify Saponin Formosanin C as a Ferroptosis Inducer in Colorectal Cancer with p53 and Oncogenic KRAS
Abstract
1. Introduction
2. Materials and Methods
2.1. Genomic Data Commons—The Cancer Genome Atlas (GDC-TCGA)
2.2. Kaplan–Meier Plotter
2.3. University of California at Santa Cruz (UCSC) Xena Platform
2.4. Connectivity Map and Library of Integrated Network-Based Cellular Signatures Unified Environment (CLUE) and ConsensusPathDB (CPDB)
2.5. Gene Set Enrichment Analysis (GSEA)
2.6. The cBio Cancer Genomics Portal (Cbioportal)
2.7. Dependency Map (DepMap)
2.8. Reagents and Cell Culture
2.9. Cell Population Growth Assay
2.10. Compound Interaction Analysis
2.11. Flow Cytometric Analysis
2.12. Intracellular Iron Measurement
2.13. Label-Free Live Cell Imaging and Analysis
2.14. Western Blot Analysis
2.15. Statistical Analysis
3. Results
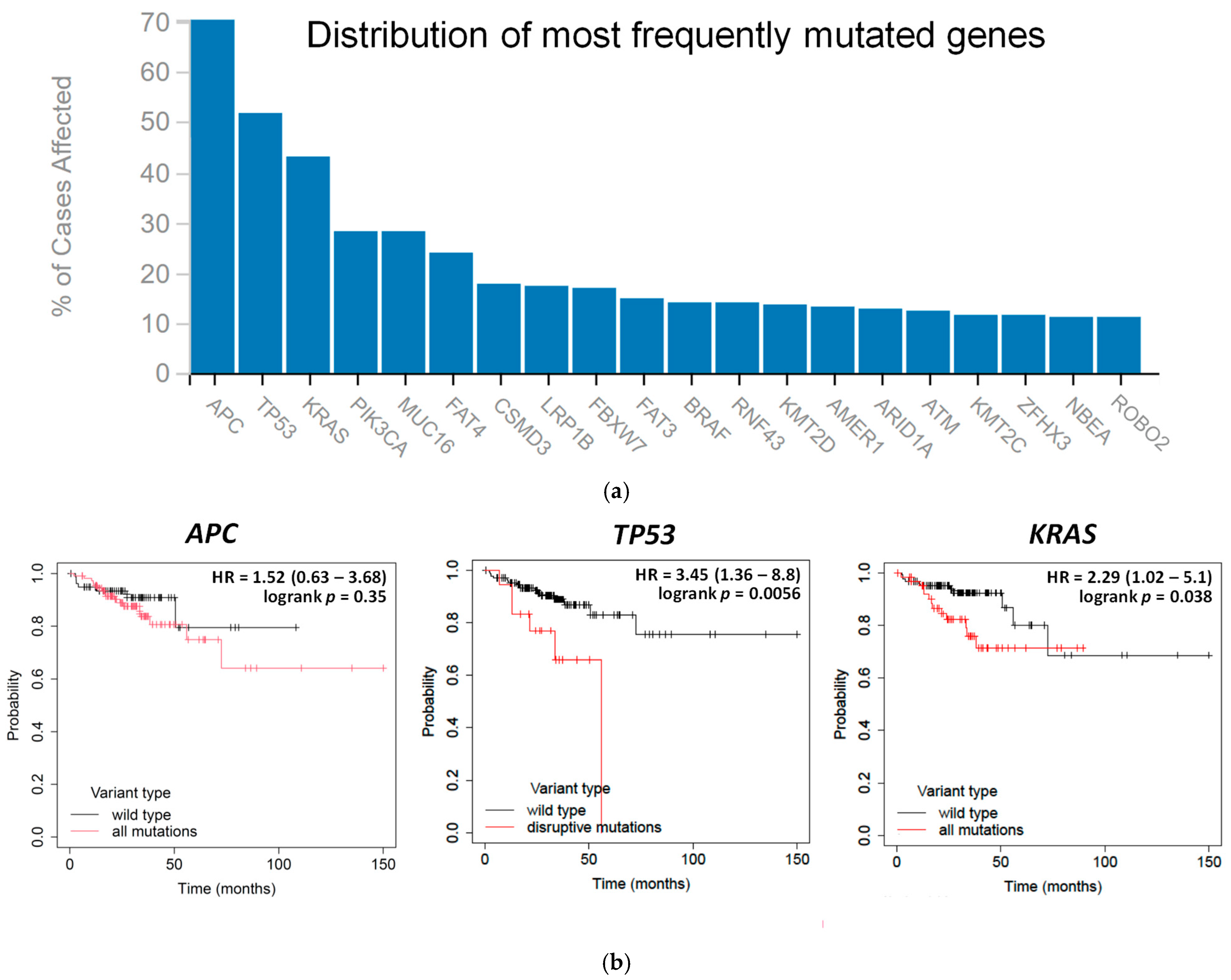
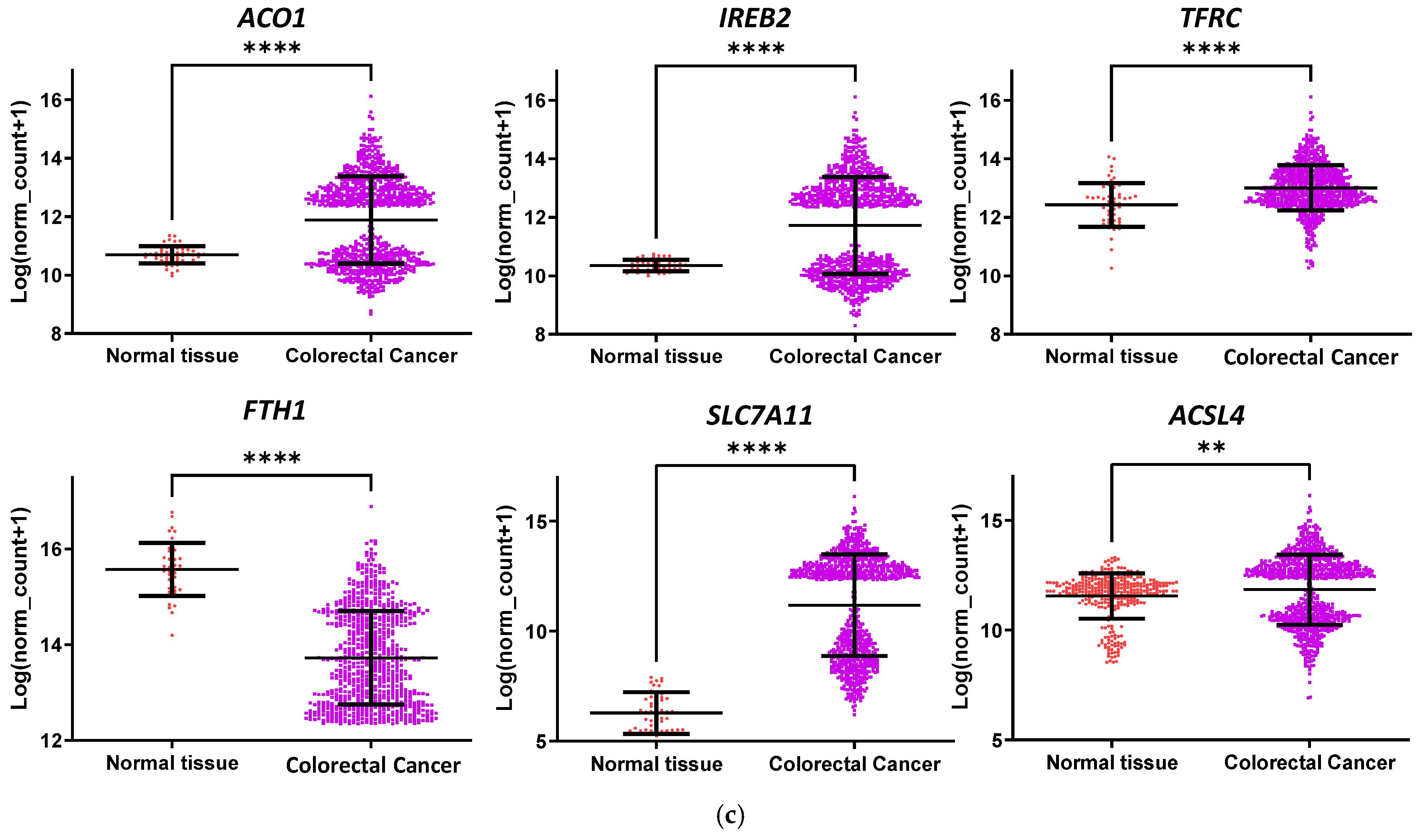
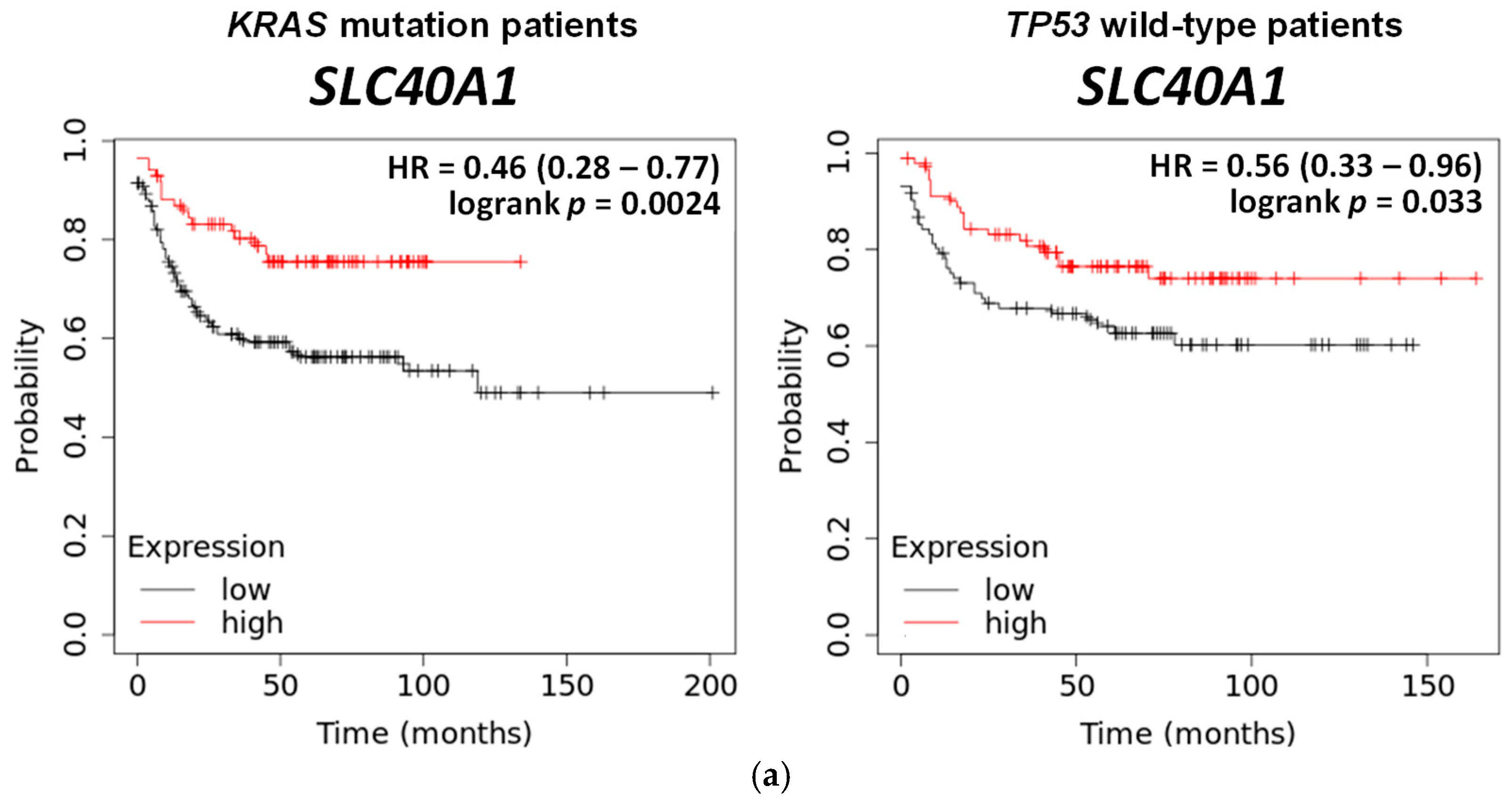
3.1. CRC Is Susceptible to Ferroptosis, and Poor Prognosis Is Linked to TP53 and KRAS Mutations
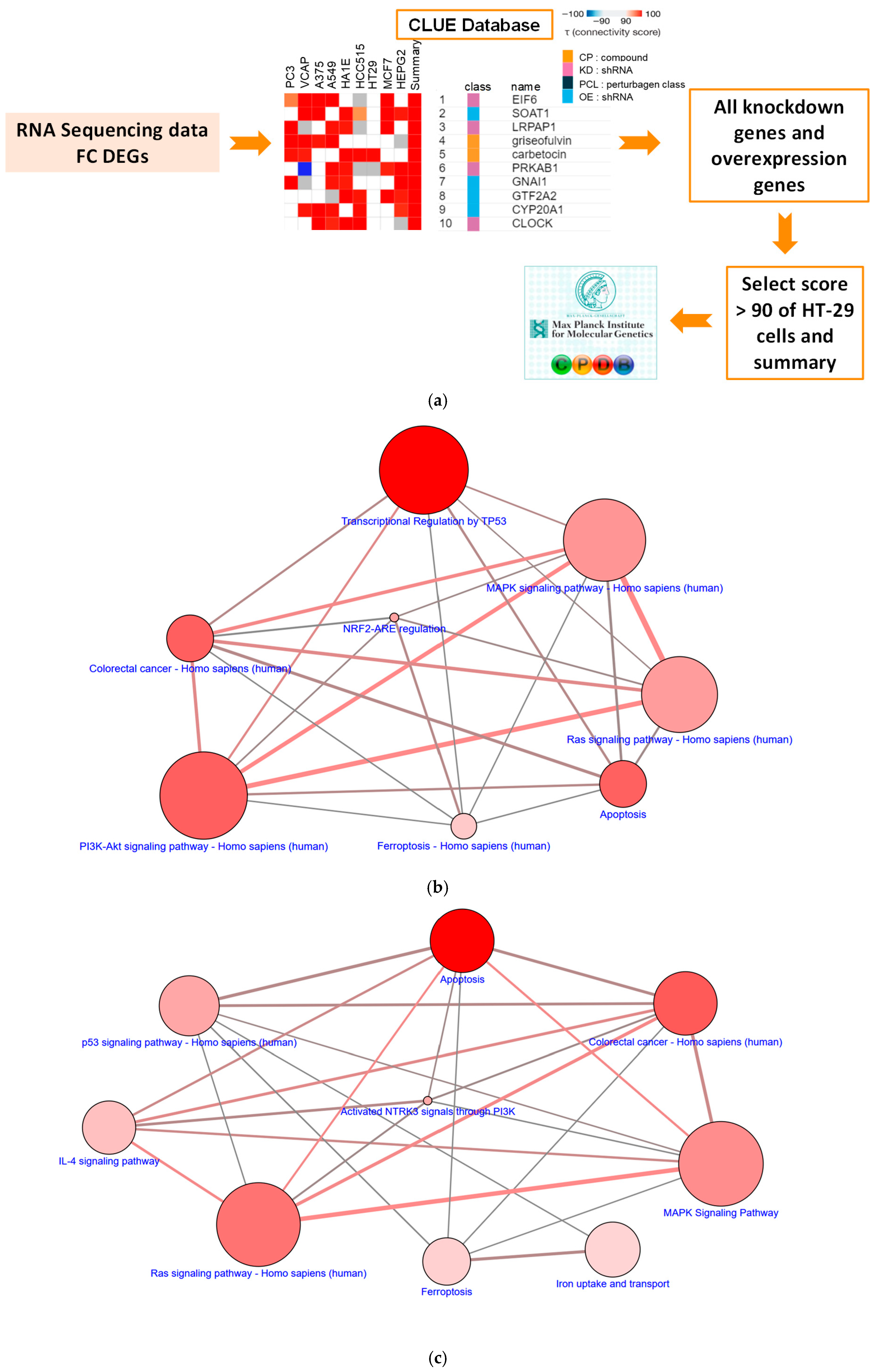
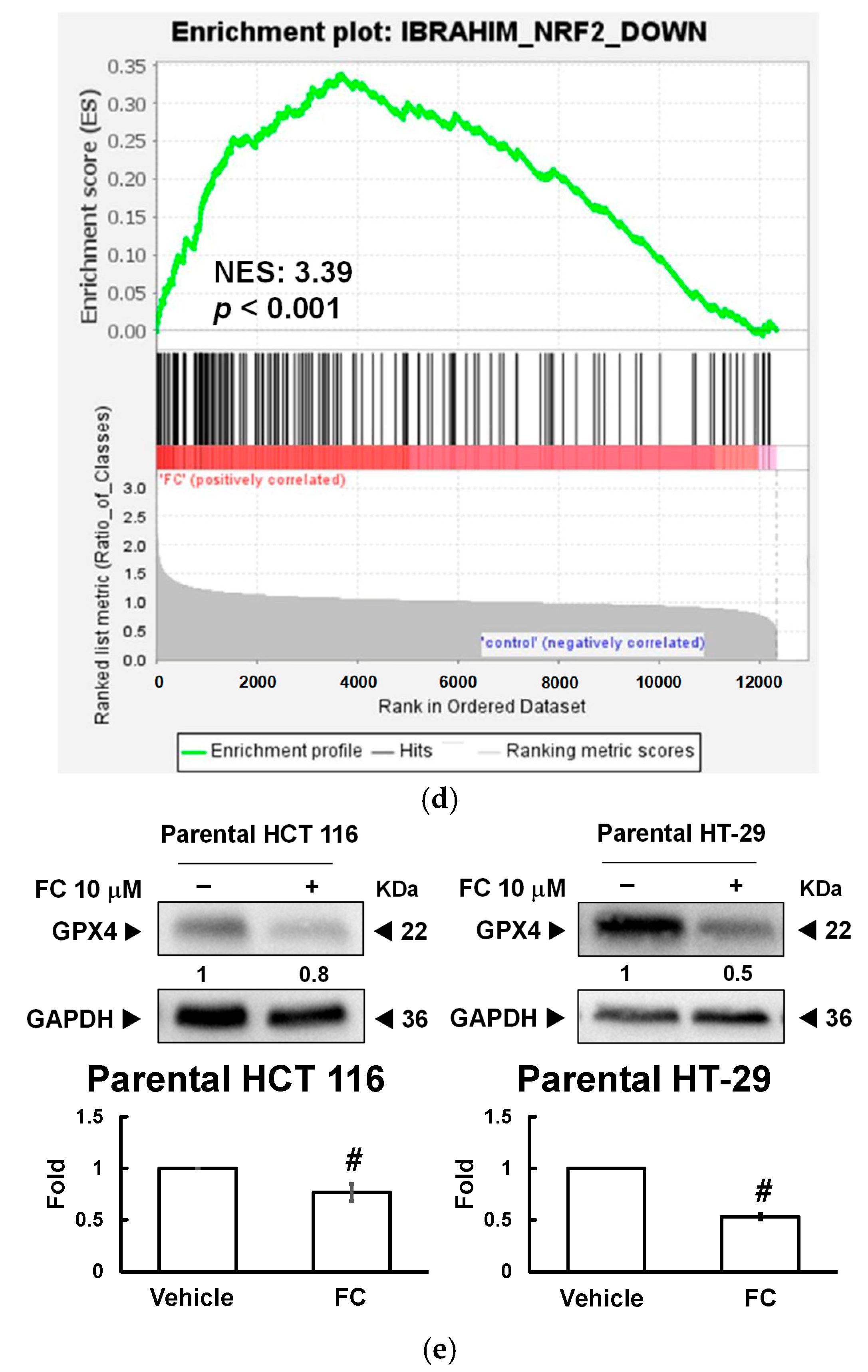
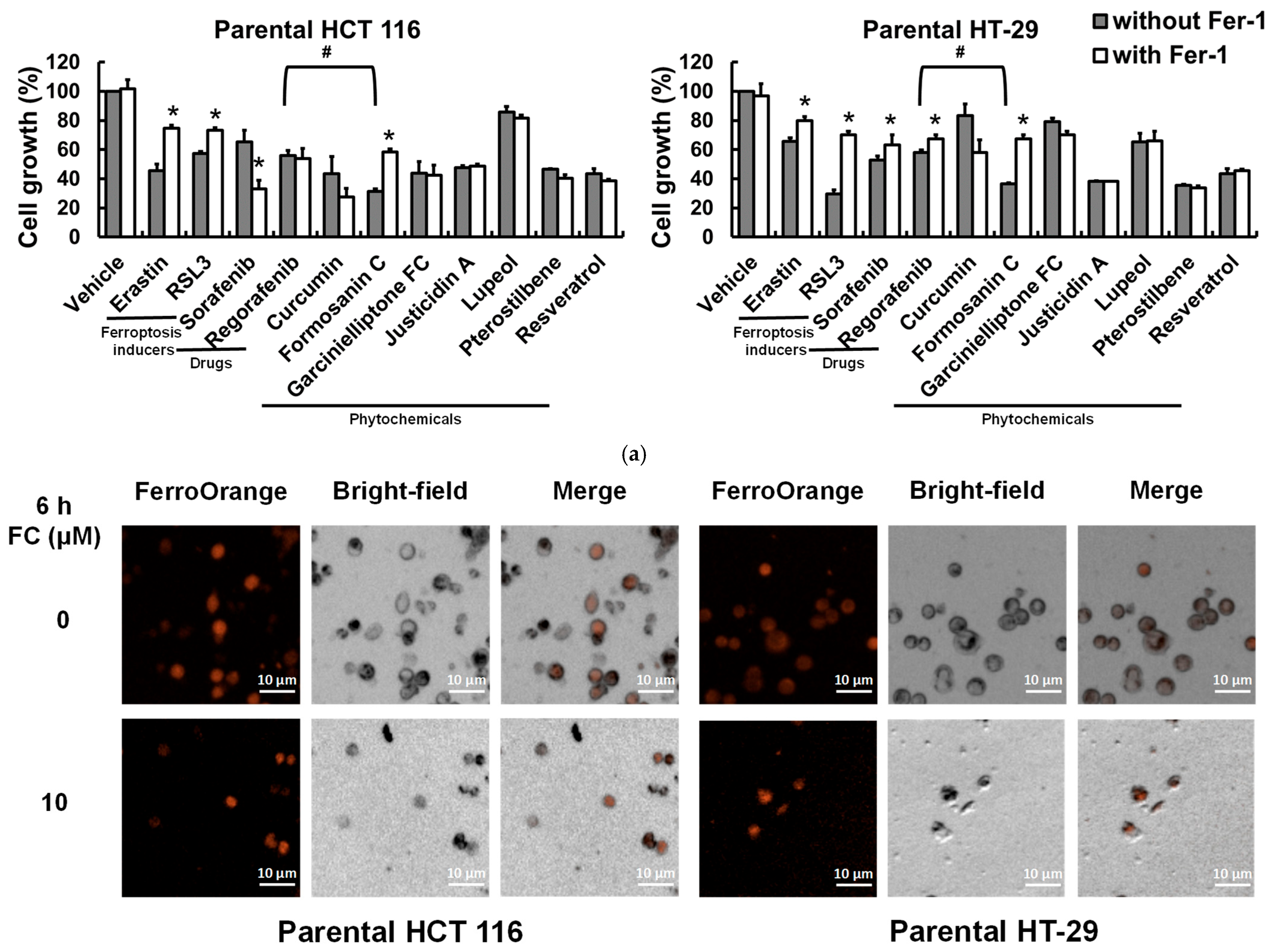
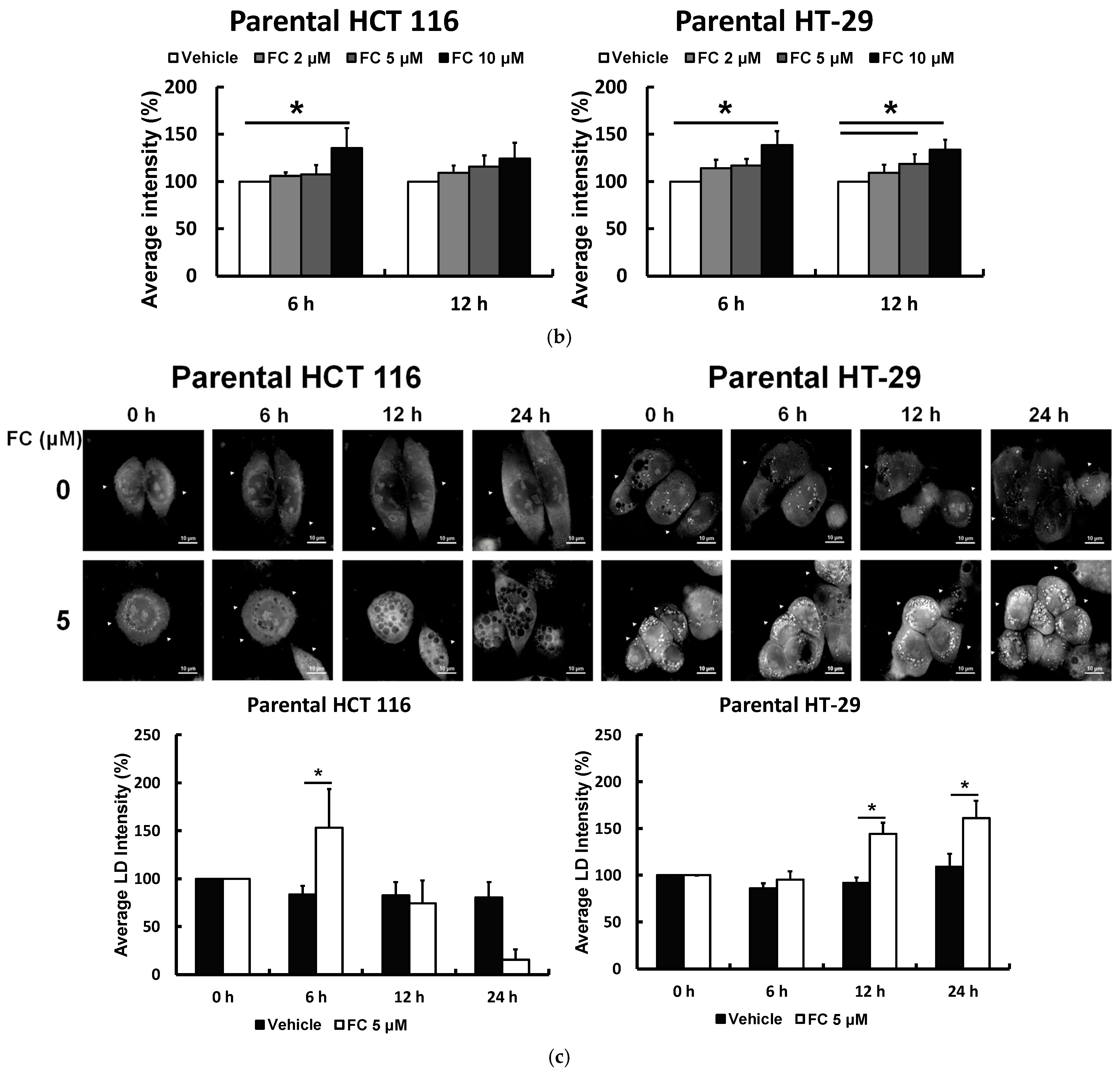
3.2. Saponin Formosanin C Is Predicted to Trigger Ferroptosis in CRC Cells via Diminished Antioxidation Capacity
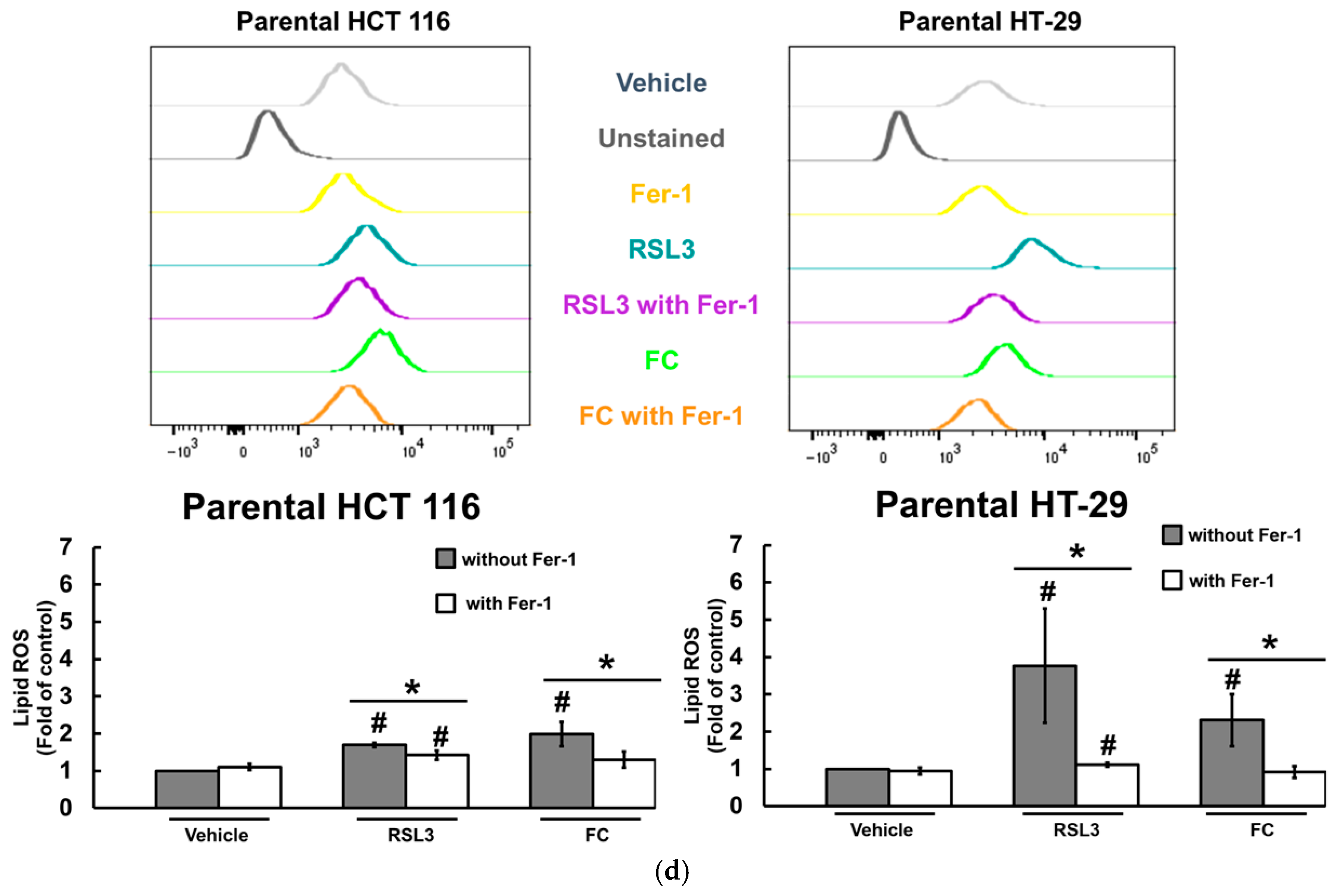
3.3. Saponin Formosanin C-Induced Ferroptosis Is Indeed Associated with Lipid ROS Formation
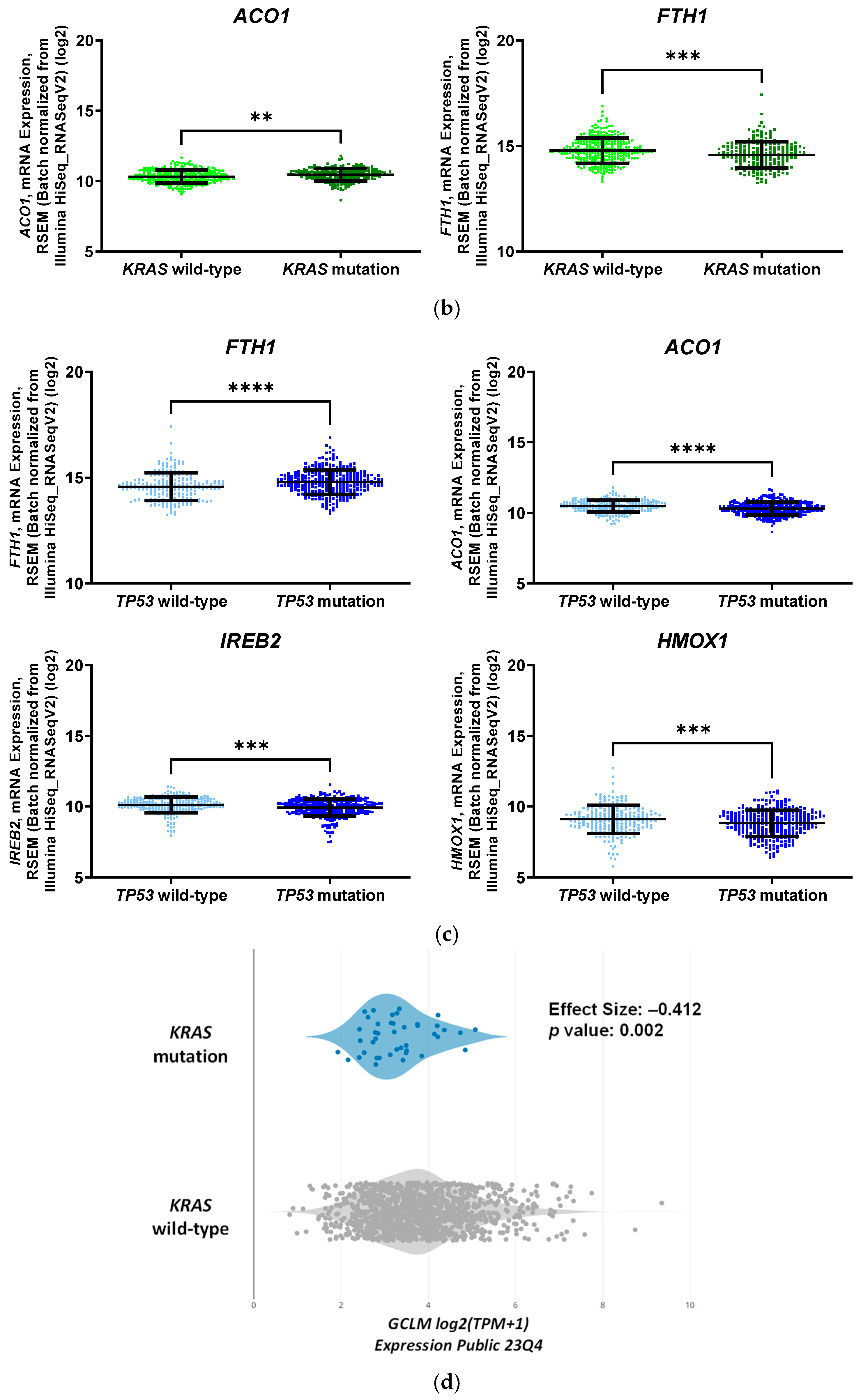
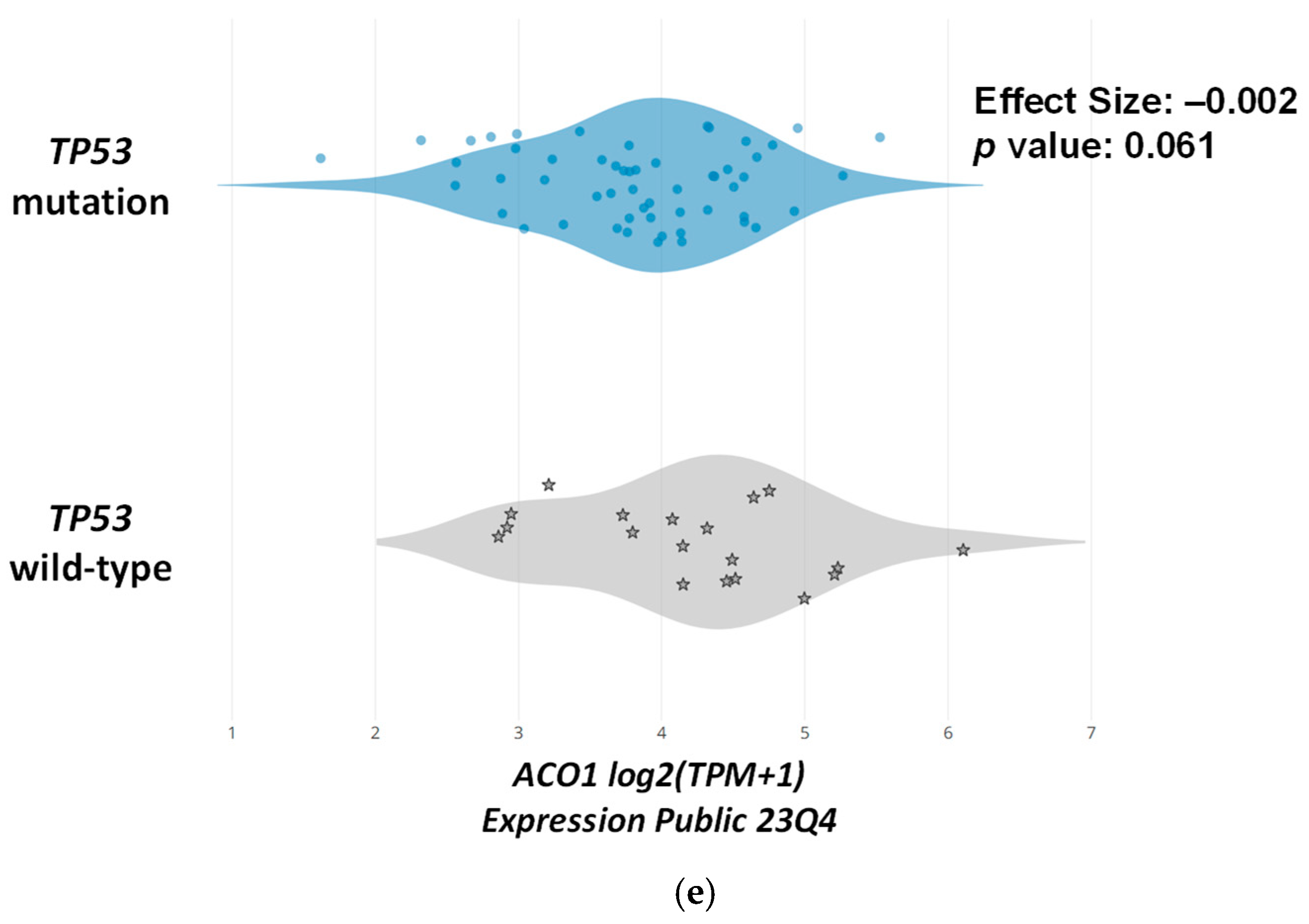
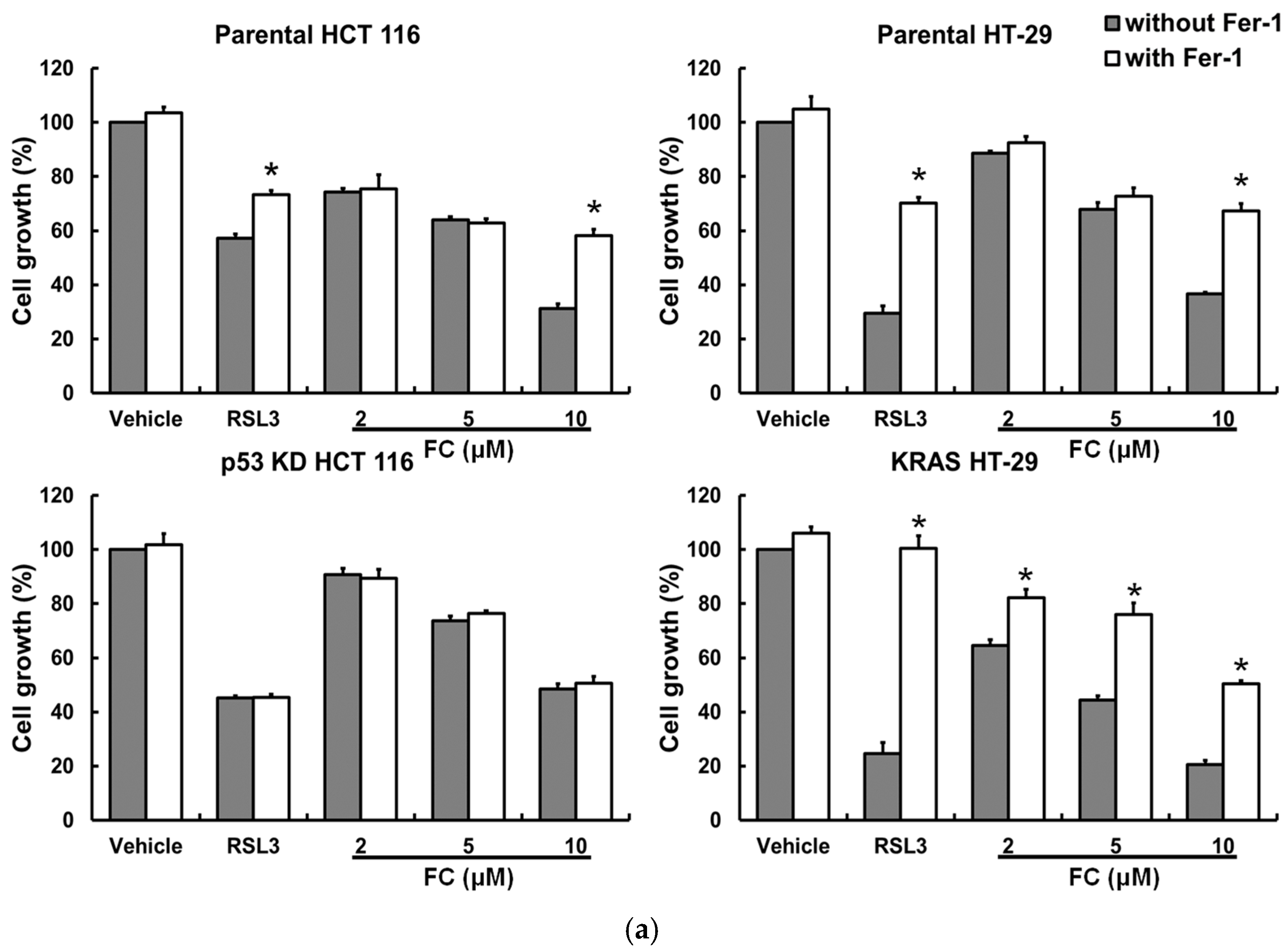
3.4. Wild-Type TP53 and Mutant KRAS Separately Favors Ferroptosis, and p53 and Oncogenic KRAS Sensitize CRC Cells to Formosanin C
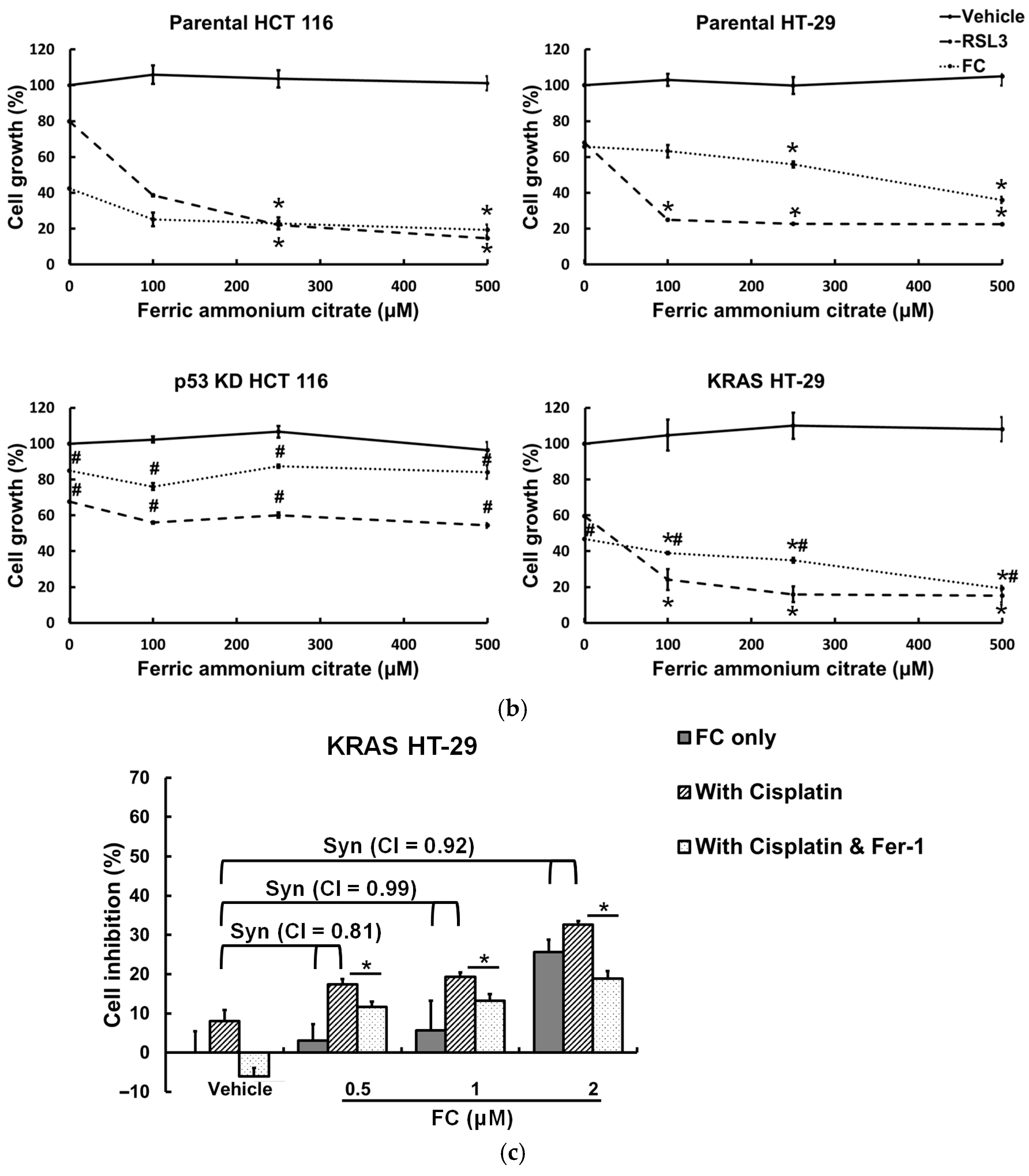
3.5. Formosanin C Treatment Sensitizes KRAS HT-29 Cells to Cisplatin
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
Abbreviations
References
- Lu, B.; Chen, X.B.; Ying, M.D.; He, Q.J.; Cao, J.; Yang, B. The Role of Ferroptosis in Cancer Development and Treatment Response. Front. Pharmacol. 2017, 8, 992. [Google Scholar] [CrossRef]
- Dixon, S.J.; Lemberg, K.M.; Lamprecht, M.R.; Skouta, R.; Zaitsev, E.M.; Gleason, C.E.; Patel, D.N.; Bauer, A.J.; Cantley, A.M.; Yang, W.S.; et al. Ferroptosis: An iron-dependent form of nonapoptotic cell death. Cell 2012, 149, 1060–1072. [Google Scholar] [CrossRef]
- Hangauer, M.J.; Viswanathan, V.S.; Ryan, M.J.; Bole, D.; Eaton, J.K.; Matov, A.; Galeas, J.; Dhruv, H.D.; Berens, M.E.; Schreiber, S.L.; et al. Drug-tolerant persister cancer cells are vulnerable to GPX4 inhibition. Nature 2017, 551, 247–250. [Google Scholar] [CrossRef]
- Co, H.K.C.; Wu, C.C.; Lee, Y.C.; Chen, S.H. Emergence of large-scale cell death through ferroptotic trigger waves. Nature 2024, 631, 654–662. [Google Scholar] [CrossRef] [PubMed]
- Favoriti, P.; Carbone, G.; Greco, M.; Pirozzi, F.; Pirozzi, R.E.; Corcione, F. Worldwide burden of colorectal cancer: A review. Updates Surg. 2016, 68, 7–11. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Ma, X.; Chakravarti, D.; Shalapour, S.; DePinho, R.A. Genetic and biological hallmarks of colorectal cancer. Genes Dev. 2021, 35, 787–820. [Google Scholar] [CrossRef] [PubMed]
- Foltran, L.; De Maglio, G.; Pella, N.; Ermacora, P.; Aprile, G.; Masiero, E.; Giovannoni, M.; Iaiza, E.; Cardellino, G.G.; Lutrino, S.E.; et al. Prognostic role of KRAS, NRAS, BRAF and PIK3CA mutations in advanced colorectal cancer. Future Oncol. 2015, 11, 629–640. [Google Scholar] [CrossRef]
- Dekker, E.; Tanis, P.J.; Vleugels, J.L.A.; Kasi, P.M.; Wallace, M.B. Colorectal cancer. Lancet 2019, 394, 1467–1480. [Google Scholar] [CrossRef]
- Leowattana, W.; Leowattana, P.; Leowattana, T. Systemic treatment for metastatic colorectal cancer. World J. Gastroenterol. 2023, 29, 1569–1588. [Google Scholar] [CrossRef]
- Dang, Q.; Sun, Z.; Wang, Y.; Wang, L.; Liu, Z.; Han, X. Ferroptosis: A double-edged sword mediating immune tolerance of cancer. Cell Death Dis. 2022, 13, 925. [Google Scholar] [CrossRef]
- Liu, X.; Tuerxun, H.; Li, Y.; Li, Y.; He, Y.; Zhao, Y. Ferroptosis: Reviewing CRC with the Third Eye. J. Inflamm. Res. 2022, 15, 6801–6812. [Google Scholar] [CrossRef] [PubMed]
- Lanczky, A.; Gyorffy, B. Web-based survival analysis tool tailored for medical research (KMplot): Development and implementation. J. Med. Internet Res. 2021, 23, e27633. [Google Scholar] [CrossRef] [PubMed]
- Goldman, M.J.; Craft, B.; Hastie, M.; Repecka, K.; McDade, F.; Kamath, A.; Banerjee, A.; Luo, Y.; Rogers, D.; Brooks, A.N.; et al. Visualizing and interpreting cancer genomics data via the Xena platform. Nat. Biotechnol. 2020, 38, 675–678. [Google Scholar] [CrossRef] [PubMed]
- Kamburov, A.; Stelzl, U.; Lehrach, H.; Herwig, R. The ConsensusPathDB interaction database: 2013 update. Nucleic Acids Res. 2013, 41, D793–D800. [Google Scholar] [CrossRef]
- Herwig, R.; Hardt, C.; Lienhard, M.; Kamburov, A. Analyzing and interpreting genome data at the network level with ConsensusPathDB. Nat. Protoc. 2016, 11, 1889–1907. [Google Scholar] [CrossRef]
- Chen, H.C.; Tang, H.H.; Hsu, W.H.; Wu, S.Y.; Cheng, W.H.; Wang, B.Y.; Su, C.L. Vulnerability of triple-negative breast cancer to saponin formosanin C-induced ferroptosis. Antioxidants 2022, 11, 298. [Google Scholar] [CrossRef]
- Subramanian, A.; Tamayo, P.; Mootha, V.K.; Mukherjee, S.; Ebert, B.L.; Gillette, M.A.; Paulovich, A.; Pomeroy, S.L.; Golub, T.R.; Lander, E.S.; et al. Gene set enrichment analysis: A knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. USA 2005, 102, 15545–15550. [Google Scholar] [CrossRef]
- Martens, M.; Ammar, A.; Riutta, A.; Waagmeester, A.; Slenter, D.N.; Hanspers, K.; Miller, R.A.; Digles, D.; Lopes, E.N.; Ehrhart, F.; et al. WikiPathways: Connecting communities. Nucleic Acids Res. 2021, 49, D613–D621. [Google Scholar] [CrossRef]
- Cerami, E.; Gao, J.; Dogrusoz, U.; Gross, B.E.; Sumer, S.O.; Aksoy, B.A.; Jacobsen, A.; Byrne, C.J.; Heuer, M.L.; Larsson, E.; et al. The cBio cancer genomics portal: An open platform for exploring multidimensional cancer genomics data. Cancer Discov. 2012, 2, 401–404. [Google Scholar] [CrossRef]
- Corsello, S.M.; Nagari, R.T.; Spangler, R.D.; Rossen, J.; Kocak, M.; Bryan, J.G.; Humeidi, R.; Peck, D.; Wu, X.; Tang, A.A.; et al. Discovering the anti-cancer potential of non-oncology drugs by systematic viability profiling. Nat. Cancer 2020, 1, 235–248. [Google Scholar] [CrossRef]
- Yu, C.; Mannan, A.M.; Yvone, G.M.; Ross, K.N.; Zhang, Y.L.; Marton, M.A.; Taylor, B.R.; Crenshaw, A.; Gould, J.Z.; Tamayo, P.; et al. High-throughput identification of genotype-specific cancer vulnerabilities in mixtures of barcoded tumor cell lines. Nat. Biotechnol. 2016, 34, 419–423. [Google Scholar] [CrossRef] [PubMed]
- Bos, J.L. The ras gene family and human carcinogenesis. Mutat. Res. 1988, 195, 255–271. [Google Scholar] [CrossRef] [PubMed]
- Wu, C.S.; Wu, S.Y.; Chen, H.C.; Chu, C.A.; Tang, H.H.; Liu, H.S.; Hong, Y.R.; Huang, C.F.; Huang, G.C.; Su, C.L. Curcumin functions as a MEK inhibitor to induce a synthetic lethal effect on KRAS mutant colorectal cancer cells receiving targeted drug regorafenib. J. Nutr. Biochem. 2019, 74, 108227. [Google Scholar] [CrossRef]
- Chen, X.; Kang, R.; Kroemer, G.; Tang, D. Broadening horizons: The role of ferroptosis in cancer. Nat. Rev. Clin. Oncol. 2021, 18, 280–296. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.C.; Su, C.L.; Chen, L.L.; Won, S.J. Formosanin C-induced apoptosis requires activation of caspase-2 and change of mitochondrial membrane potential. Cancer Sci. 2009, 100, 503–513. [Google Scholar] [CrossRef]
- Wu, C.C.; Lu, Y.H.; Wei, B.L.; Yang, S.C.; Won, S.J.; Lin, C.N. Phloroglucinols with prooxidant activity from Garcinia subelliptica. J. Nat. Prod. 2008, 71, 246–250. [Google Scholar] [CrossRef]
- Day, S.H.; Lin, Y.C.; Tsai, M.L.; Tsao, L.T.; Ko, H.H.; Chung, M.I.; Lee, J.C.; Wang, J.P.; Won, S.J.; Lin, C.N. Potent cytotoxic lignans from Justicia procumbens and their effects on nitric oxide and tumor necrosis factor-alpha production in mouse macrophages. J. Nat. Prod. 2002, 65, 379–381. [Google Scholar] [CrossRef]
- Takahashi, N.; Li, W.; Banerjee, D.; Guan, Y.; Wada-Takahashi, Y.; Brennan, M.F.; Chou, T.C.; Scotto, K.W.; Bertino, J.R. Sequence-dependent synergistic cytotoxicity of ecteinascidin-743 and paclitaxel in human breast cancer cell lines in vitro and in vivo. Cancer Res. 2002, 62, 6909–6915. [Google Scholar]
- Lin, P.L.; Tang, H.H.; Wu, S.Y.; Shaw, N.S.; Su, C.L. Saponin Formosanin C-induced Ferritinophagy and Ferroptosis in Human Hepatocellular Carcinoma Cells. Antioxidants 2020, 9, 682. [Google Scholar] [CrossRef]
- Cao, J.Y.; Dixon, S.J. Mechanisms of ferroptosis. Cell Mol. Life Sci. 2016, 73, 2195–2209. [Google Scholar] [CrossRef]
- Liao, S.C.; Hsu, H.W.; Chuang, K.L.; Huang, Z.Y.; Lin, K.T.; Hsu, W.H.; Chang, K.H.; Huang, C.F.; Su, C.L. Using the Pleiotropic Characteristics of Curcumin to Validate the Potential Application of a Novel Gene Expression Screening Platform. Nutrients 2019, 11, 1397. [Google Scholar] [CrossRef] [PubMed]
- Liao, S.C.; Hsu, W.H.; Huang, Z.Y.; Chuang, K.L.; Lin, K.T.; Tseng, C.L.; Tsai, T.H.; Dao, A.H.; Su, C.L.; Huang, C.F. Bioactivity Evaluation of a Novel Formulated Curcumin. Nutrients 2019, 11, 2982. [Google Scholar] [CrossRef]
- Won, S.-J.; Yen, C.-H.; Hsieh, H.-W.; Chang, S.-W.; Lin, C.-N.; Huang, C.-Y.F.; Su, C.-L. Using connectivity map to identify natural lignan justicidin A as a NF-kappaB suppressor. J. Funct. Foods 2017, 34, 68–76. [Google Scholar] [CrossRef]
- Tang, D.; Chen, X.; Kang, R.; Kroemer, G. Ferroptosis: Molecular mechanisms and health implications. Cell Res. 2021, 31, 107–125. [Google Scholar] [CrossRef]
- Yang, W.S.; SriRamaratnam, R.; Welsch, M.E.; Shimada, K.; Skouta, R.; Viswanathan, V.S.; Cheah, J.H.; Clemons, P.A.; Shamji, A.F.; Clish, C.B.; et al. Regulation of ferroptotic cancer cell death by GPX4. Cell 2014, 156, 317–331. [Google Scholar] [CrossRef]
- Kim, S.E.; Zhang, L.; Ma, K.; Riegman, M.; Chen, F.; Ingold, I.; Conrad, M.; Turker, M.Z.; Gao, M.; Jiang, X.; et al. Ultrasmall nanoparticles induce ferroptosis in nutrient-deprived cancer cells and suppress tumour growth. Nat. Nanotechnol. 2016, 11, 977–985. [Google Scholar] [CrossRef]
- Lang, L. FDA approves sorafenib for patients with inoperable liver cancer. Gastroenterology 2008, 134, 379. [Google Scholar] [CrossRef]
- Yan, Y.; Grothey, A. Molecular profiling in the treatment of colorectal cancer: Focus on regorafenib. OncoTargets Ther. 2015, 8, 2949–2957. [Google Scholar] [CrossRef]
- Berndt, C.; Alborzinia, H.; Amen, V.S.; Ayton, S.; Barayeu, U.; Bartelt, A.; Bayir, H.; Bebber, C.M.; Birsoy, K.; Bottcher, J.P.; et al. Ferroptosis in health and disease. Redox Biol. 2024, 75, 103211. [Google Scholar] [CrossRef]
- Sastry, J.; Kellie, S.J. Severe neurotoxicity, ototoxicity and nephrotoxicity following high-dose cisplatin and amifostine. Pediatr. Hematol. Oncol. 2005, 22, 441–445. [Google Scholar] [CrossRef]
- Lamb, J.; Crawford, E.D.; Peck, D.; Modell, J.W.; Blat, I.C.; Wrobel, M.J.; Lerner, J.; Brunet, J.P.; Subramanian, A.; Ross, K.N.; et al. The Connectivity Map: Using gene-expression signatures to connect small molecules, genes, and disease. Science 2006, 313, 1929–1935. [Google Scholar] [CrossRef] [PubMed]
- Subramanian, A.; Narayan, R.; Corsello, S.M.; Peck, D.D.; Natoli, T.E.; Lu, X.; Gould, J.; Davis, J.F.; Tubelli, A.A.; Asiedu, J.K.; et al. A Next Generation Connectivity Map: L1000 Platform and the First 1,000,000 Profiles. Cell 2017, 171, 1437–1452.e1417. [Google Scholar] [CrossRef] [PubMed]
- Schott, C.; Graab, U.; Cuvelier, N.; Hahn, H.; Fulda, S. Oncogenic RAS Mutants Confer Resistance of RMS13 Rhabdomyosarcoma Cells to Oxidative Stress-Induced Ferroptotic Cell Death. Front. Oncol. 2015, 5, 131. [Google Scholar] [CrossRef] [PubMed]
- Park, S.; Oh, J.; Kim, M.; Jin, E.J. Bromelain effectively suppresses Kras-mutant colorectal cancer by stimulating ferroptosis. Anim. Cells Syst. 2018, 22, 334–340. [Google Scholar] [CrossRef]
- Yang, M.; Zou, J.; Zhu, H.; Liu, S.; Wang, H.; Bai, P.; Xiao, X. Paris saponin II inhibits human ovarian cancer cell-induced angiogenesis by modulating NF-kappaB signaling. Oncol. Rep. 2015, 33, 2190–2198. [Google Scholar] [CrossRef]
- Wu, R.T.; Chiang, H.C.; Fu, W.C.; Chien, K.Y.; Chung, Y.M.; Horng, L.Y. Formosanin-C, an immunomodulator with antitumor activity. Int. J. Immunopharmacol. 1990, 12, 777–786. [Google Scholar] [CrossRef]
- Fenwick, D.E.; Oakenfull, D. Saponin content of food plants and some prepared foods. J. Sci. Food Agric. 1983, 34, 186–191. [Google Scholar] [CrossRef]
- Xiao, X.; Zou, J.; Bui-Nguyen, T.M.; Bai, P.; Gao, L.; Liu, J.; Liu, S.; Xiao, J.; Chen, X.; Zhang, X.; et al. Paris saponin II of Rhizoma Paridis--a novel inducer of apoptosis in human ovarian cancer cells. Biosci. Trends 2012, 6, 201–211. [Google Scholar] [CrossRef]
- Pottier, C.; Montero-Ruiz, L.; Jehay, R.; Wery, C.; Baiwir, D.; Mazzucchelli, G.; Bekisz, S.; Thissen, R.; Josse, C.; Rorive, A.; et al. Targeting ferroptosis resistance resensitizes metastatic HR(+)HER2(−) breast cancer cells to palbociclib-hormone therapy. Cancer Commun. 2025, 45, 460–464. [Google Scholar] [CrossRef]
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chen, H.-C.; Chen, C.-Y.; Wang, P.-Y.; Su, P.-Y.; Tsai, S.-P.; Hsu, C.-P.; Liu, H.-S.; Huang, C.-Y.F.; Cheng, W.-H.; Lee, M.-F.; et al. Using Integrated Bioinformatics Analysis to Identify Saponin Formosanin C as a Ferroptosis Inducer in Colorectal Cancer with p53 and Oncogenic KRAS. Antioxidants 2025, 14, 1027. https://doi.org/10.3390/antiox14081027
Chen H-C, Chen C-Y, Wang P-Y, Su P-Y, Tsai S-P, Hsu C-P, Liu H-S, Huang C-YF, Cheng W-H, Lee M-F, et al. Using Integrated Bioinformatics Analysis to Identify Saponin Formosanin C as a Ferroptosis Inducer in Colorectal Cancer with p53 and Oncogenic KRAS. Antioxidants. 2025; 14(8):1027. https://doi.org/10.3390/antiox14081027
Chicago/Turabian StyleChen, Hsin-Chih, Ching-Ying Chen, Pao-Yuan Wang, Pin-Yu Su, Shu-Ping Tsai, Chi-Pei Hsu, Hsiao-Sheng Liu, Chi-Ying F. Huang, Wen-Hsing Cheng, Ming-Fen Lee, and et al. 2025. "Using Integrated Bioinformatics Analysis to Identify Saponin Formosanin C as a Ferroptosis Inducer in Colorectal Cancer with p53 and Oncogenic KRAS" Antioxidants 14, no. 8: 1027. https://doi.org/10.3390/antiox14081027
APA StyleChen, H.-C., Chen, C.-Y., Wang, P.-Y., Su, P.-Y., Tsai, S.-P., Hsu, C.-P., Liu, H.-S., Huang, C.-Y. F., Cheng, W.-H., Lee, M.-F., & Su, C.-L. (2025). Using Integrated Bioinformatics Analysis to Identify Saponin Formosanin C as a Ferroptosis Inducer in Colorectal Cancer with p53 and Oncogenic KRAS. Antioxidants, 14(8), 1027. https://doi.org/10.3390/antiox14081027





