Extracellular Vesicles and Cancer Therapy: Insights into the Role of Oxidative Stress
Abstract
1. Introduction
2. Extracellular Vesicles: Biogenesis and Characterization
2.1. Biogenesis of EVs
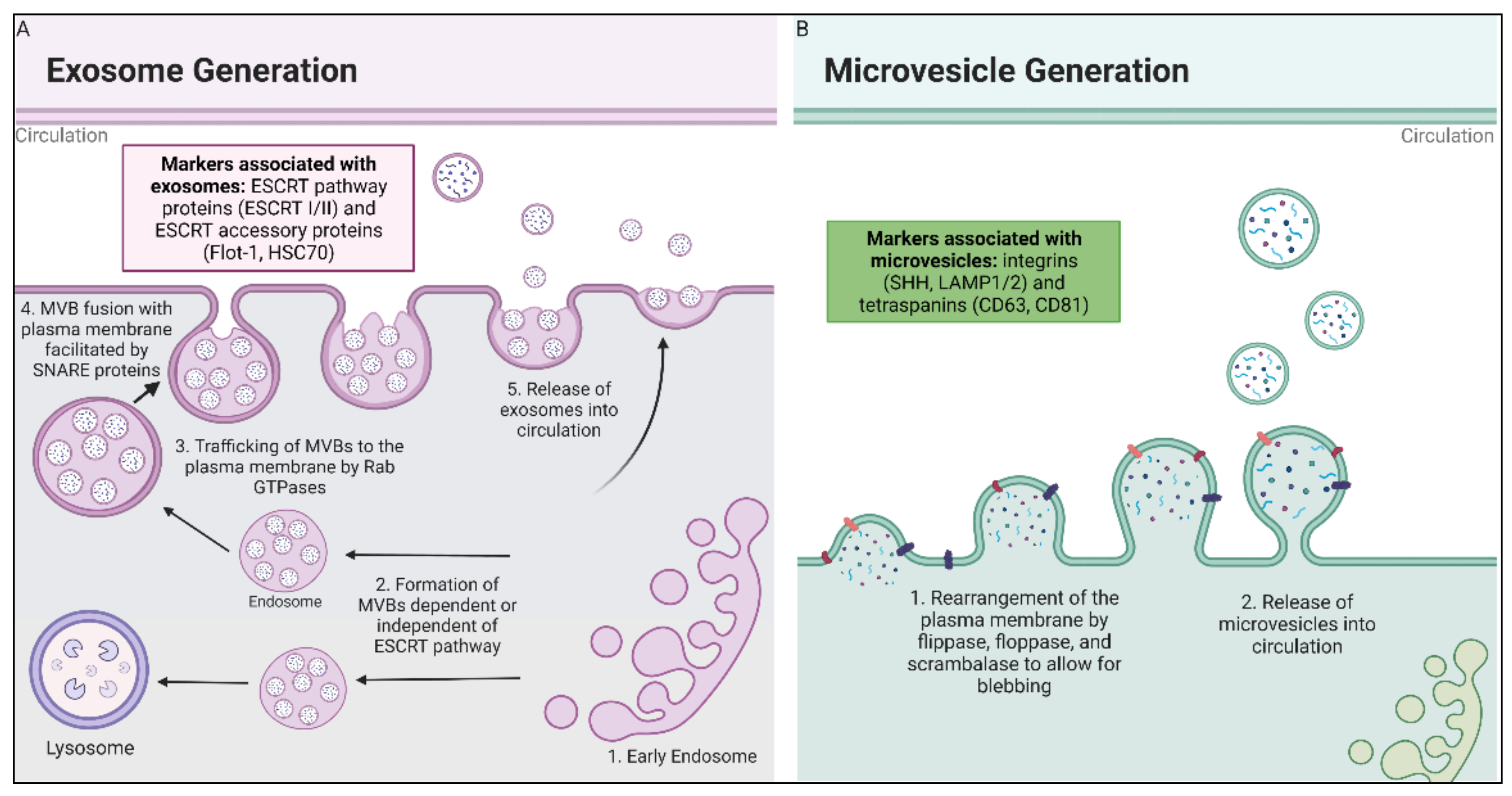
2.1.1. Exosome Biogenesis
2.1.2. Microvesicle Biogenesis
2.2. EVs Content
2.3. EVs Characterization
2.4. EVs Isolation
3. Oxidative Stress: Pro-Oxidants and Antioxidants
3.1. Oxidants and ROS
3.2. Antioxidants and Redox Couples
3.3. Redox Signaling
3.4. Oxidative Stress and Cancer Therapy
3.5. HNE and EVs
4. Role and Function of EVs: Current Understanding and Future Directions
4.1. EVs in Cancer
4.2. Translational Uses of EVs
4.3. EVs and Cancer Therapy
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
Abbreviations
References
- Van Niel, G.; D’Angelo, G.; Raposo, G. Shedding light on the cell biology of extracellular vesicles. Nat. Rev. Mol. Cell Biol. 2018, 19, 213–228. [Google Scholar] [CrossRef]
- Shah, R.; Patel, T.; Freedman, J.E. Circulating Extracellular Vesicles in Human Disease. N. Engl. J. Med. 2018, 379, 2180–2181. [Google Scholar] [CrossRef]
- Shao, H.; Chung, J.; Balaj, L.; Charest, A.; Bigner, D.D.; Carter, B.S.; Hochberg, F.H.; Breakefield, X.O.; Weissleder, R.; Lee, H. Protein typing of circulating microvesicles allows real-time monitoring of glioblastoma therapy. Nat. Med. 2012, 18, 1835–1840. [Google Scholar] [CrossRef]
- Vader, P.; Mol, E.A.; Pasterkamp, G.; Schiffelers, R.M. Extracellular vesicles for drug delivery. Adv. Drug Deliv. Rev. 2016, 106, 148–156. [Google Scholar] [CrossRef]
- Skog, J.; Wurdinger, T.; van Rijn, S.; Meijer, D.H.; Gainche, L.; Sena-Esteves, M.; Curry, W.T., Jr.; Carter, B.S.; Krichevsky, A.M.; Breakefield, X.O. Glioblastoma microvesicles transport RNA and proteins that promote tumour growth and provide diagnostic biomarkers. Nat. Cell Biol. 2008, 10, 1470–1476. [Google Scholar] [CrossRef]
- Peinado, H.; Alec kovic, M.; Lavotshkin, S.; Matei, I.; Costa-Silva, B.; Moreno-Bueno, G.; Hergueta-Redondo, M.; Williams, C.; Garcia-Santos, G.; Ghajar, C.M.; et al. Corrigendum: Melanoma exosomes educate bone marrow progenitor cells toward a pro-metastatic phenotype through MET. Nat. Med. 2016, 22, 1502. [Google Scholar] [CrossRef] [PubMed]
- Al-Nedawi, K.; Meehan, B.; Micallef, J.; Lhotak, V.; May, L.; Guha, A.; Rak, J. Intercellular transfer of the oncogenic receptor EGFRvIII by microvesicles derived from tumour cells. Nat. Cell Biol. 2008, 10, 619–624. [Google Scholar] [CrossRef]
- Lachenal, G.; Pernet-Gallay, K.; Chivet, M.; Hemming, F.J.; Belly, A.; Bodon, G.; Blot, B.; Haase, G.; Goldberg, Y.; Sadoul, R. Release of exosomes from differentiated neurons and its regulation by synaptic glutamatergic activity. Mol. Cell. Neurosci. 2011, 46, 409–418. [Google Scholar] [CrossRef]
- Thery, C.; Ostrowski, M.; Segura, E. Membrane vesicles as conveyors of immune responses. Nat. Rev. Immunol. 2009, 9, 581–593. [Google Scholar] [CrossRef]
- Harmati, M.; Gyukity-Sebestyen, E.; Dobra, G.; Janovak, L.; Dekany, I.; Saydam, O.; Hunyadi-Gulyas, E.; Nagy, I.; Farkas, A.; Pankotai, T.; et al. Small extracellular vesicles convey the stress-induced adaptive responses of melanoma cells. Sci. Rep. 2019, 9, 15329. [Google Scholar] [CrossRef]
- Siegel, R.L.; Miller, K.D.; Fuchs, H.E.; Jemal, A. Cancer statistics, 2022. CA Cancer J. Clin. 2022, 72, 7–33. [Google Scholar] [CrossRef]
- Vitali, M.; Ripamonti, C.I.; Roila, F.; Proto, C.; Signorelli, D.; Imbimbo, M.; Corrao, G.; Brissa, A.; Rosaria, G.; de Braud, F.; et al. Cognitive impairment and chemotherapy: A brief overview. Crit. Rev. Oncol. Hematol. 2017, 118, 7–14. [Google Scholar] [CrossRef]
- Halliwell, B.G.J.M.C. Free Radicals in Biology and Medicine; Oxford University Press: Oxford, UK, 2015. [Google Scholar]
- Dalleau, S.; Baradat, M.; Gueraud, F.; Huc, L. Cell death and diseases related to oxidative stress: 4-hydroxynonenal (HNE) in the balance. Cell Death Differ. 2013, 20, 1615–1630. [Google Scholar] [CrossRef]
- Lovell, M.A.; Markesbery, W.R. Oxidative DNA damage in mild cognitive impairment and late-stage Alzheimer’s disease. Nucleic Acids Res. 2007, 35, 7497–7504. [Google Scholar] [CrossRef]
- Sultana, R.; Butterfield, D.A. Role of oxidative stress in the progression of Alzheimer’s disease. J. Alzheimers Dis. 2010, 19, 341–353. [Google Scholar] [CrossRef] [PubMed]
- Di Domenico, F.; Tramutola, A.; Butterfield, D.A. Role of 4-hydroxy-2-nonenal (HNE) in the pathogenesis of alzheimer disease and other selected age-related neurodegenerative disorders. Free Radic. Biol. Med. 2017, 111, 253–261. [Google Scholar] [CrossRef] [PubMed]
- Perluigi, M.; Coccia, R.; Butterfield, D.A. 4-Hydroxy-2-nonenal, a reactive product of lipid peroxidation, and neurodegenerative diseases: A toxic combination illuminated by redox proteomics studies. Antioxid. Redox Signal. 2012, 17, 1590–1609. [Google Scholar] [CrossRef] [PubMed]
- Green, J.L.; Knight, S.J.; McCarthy, M.; De Luca, C.R. Motor functioning during and following treatment with chemotherapy for pediatric acute lymphoblastic leukemia. Pediatr. Blood Cancer 2013, 60, 1261–1266. [Google Scholar] [CrossRef] [PubMed]
- Anderson, F.S.; Kunin-Batson, A.S. Neurocognitive late effects of chemotherapy in children: The past 10 years of research on brain structure and function. Pediatr. Blood Cancer 2009, 52, 159–164. [Google Scholar] [CrossRef]
- Chen, Y.; Jungsuwadee, P.; Vore, M.; Butterfield, D.A.; St Clair, D.K. Collateral damage in cancer chemotherapy: Oxidative stress in nontargeted tissues. Mol. Interv. 2007, 7, 147–156. [Google Scholar] [CrossRef]
- Hall, E.J. Radiobiology for the Radiologist; Medical Dept., Harper & Row: Hagerstown, MD, USA, 1973. [Google Scholar]
- Andersen, J.K. Oxidative stress in neurodegeneration: Cause or consequence? Nat. Med. 2004, 10, S18–S25. [Google Scholar] [CrossRef] [PubMed]
- Salim, S. Oxidative Stress and the Central Nervous System. J. Pharmacol. Exp. Ther. 2017, 360, 201–205. [Google Scholar] [CrossRef] [PubMed]
- Tangpong, J.; Cole, M.P.; Sultana, R.; Joshi, G.; Estus, S.; Vore, M.; St Clair, W.; Ratanachaiyavong, S.; St Clair, D.K.; Butterfield, D.A. Adriamycin-induced, TNF-alpha-mediated central nervous system toxicity. Neurobiol. Dis. 2006, 23, 127–139. [Google Scholar] [CrossRef] [PubMed]
- Ren, X.; Keeney, J.T.R.; Miriyala, S.; Noel, T.; Powell, D.K.; Chaiswing, L.; Bondada, S.; St Clair, D.K.; Butterfield, D.A. The triangle of death of neurons: Oxidative damage, mitochondrial dysfunction, and loss of choline-containing biomolecules in brains of mice treated with doxorubicin. Advanced insights into mechanisms of chemotherapy induced cognitive impairment (“chemobrain”) involving TNF-alpha. Free Radic. Biol. Med. 2019, 134, 1–8. [Google Scholar] [CrossRef]
- Suematsu, N.; Tsutsui, H.; Wen, J.; Kang, D.; Ikeuchi, M.; Ide, T.; Hayashidani, S.; Shiomi, T.; Kubota, T.; Hamasaki, N.; et al. Oxidative stress mediates tumor necrosis factor-alpha-induced mitochondrial DNA damage and dysfunction in cardiac myocytes. Circulation 2003, 107, 1418–1423. [Google Scholar] [CrossRef]
- Cheung, Y.T.; Sabin, N.D.; Reddick, W.E.; Bhojwani, D.; Liu, W.; Brinkman, T.M.; Glass, J.O.; Hwang, S.N.; Srivastava, D.; Pui, C.H.; et al. Leukoencephalopathy and long-term neurobehavioural, neurocognitive, and brain imaging outcomes in survivors of childhood acute lymphoblastic leukaemia treated with chemotherapy: A longitudinal analysis. Lancet Haematol. 2016, 3, e456–e466. [Google Scholar] [CrossRef]
- Teng, F.; Fussenegger, M. Shedding Light on Extracellular Vesicle Biogenesis and Bioengineering. Adv. Sci. (Weinh) 2020, 8, 2003505. [Google Scholar] [CrossRef]
- Valadi, H.; Ekstrom, K.; Bossios, A.; Sjostrand, M.; Lee, J.J.; Lotvall, J.O. Exosome-mediated transfer of mRNAs and microRNAs is a novel mechanism of genetic exchange between cells. Nat. Cell Biol. 2007, 9, 654–659. [Google Scholar] [CrossRef]
- Thery, C.; Witwer, K.W.; Aikawa, E.; Alcaraz, M.J.; Anderson, J.D.; Andriantsitohaina, R.; Antoniou, A.; Arab, T.; Archer, F.; Atkin-Smith, G.K.; et al. Minimal information for studies of extracellular vesicles 2018 (MISEV2018): A position statement of the International Society for Extracellular Vesicles and update of the MISEV2014 guidelines. J. Extracell. Vesicles 2018, 7, 1535750. [Google Scholar] [CrossRef]
- Jahn, R.; Scheller, R.H. SNAREs—engines for membrane fusion. Nat. Rev. Mol. Cell Biol. 2006, 7, 631–643. [Google Scholar] [CrossRef]
- Bonifacino, J.S.; Glick, B.S. The Mechanisms of Vesicle Budding and Fusion. Cell 2004, 116, 153–166. [Google Scholar] [CrossRef]
- Pap, E.; Pallinger, E.; Pasztoi, M.; Falus, A. Highlights of a new type of intercellular communication: Microvesicle-based information transfer. Inflamm. Res. 2009, 58, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Christ, L.; Raiborg, C.; Wenzel, E.M.; Campsteijn, C.; Stenmark, H. Cellular Functions and Molecular Mechanisms of the ESCRT Membrane-Scission Machinery. Trends Biochem. Sci. 2017, 42, 42–56. [Google Scholar] [CrossRef] [PubMed]
- Kowal, J.; Tkach, M.; Thery, C. Biogenesis and secretion of exosomes. Curr. Opin. Cell Biol. 2014, 29, 116–125. [Google Scholar] [CrossRef] [PubMed]
- Stuffers, S.; Sem Wegner, C.; Stenmark, H.; Brech, A. Multivesicular endosome biogenesis in the absence of ESCRTs. Traffic 2009, 10, 925–937. [Google Scholar] [CrossRef]
- Strauss, K.; Goebel, C.; Runz, H.; Mobius, W.; Weiss, S.; Feussner, I.; Simons, M.; Schneider, A. Exosome secretion ameliorates lysosomal storage of cholesterol in Niemann-Pick type C disease. J. Biol. Chem. 2010, 285, 26279–26288. [Google Scholar] [CrossRef]
- Hoshino, D.; Kirkbride, K.C.; Costello, K.; Clark, E.S.; Sinha, S.; Grega-Larson, N.; Tyska, M.J.; Weaver, A.M. Exosome Secretion Is Enhanced by Invadopodia and Drives Invasive Behavior. Cell Rep. 2013, 5, 1159–1168. [Google Scholar] [CrossRef]
- Stenmark, H. Rab GTPases as coordinators of vesicle traffic. Nat. Rev. Mol. Cell Biol. 2009, 10, 513–525. [Google Scholar] [CrossRef]
- Hankins, H.M.; Baldridge, R.D.; Xu, P.; Graham, T.R. Role of flippases, scramblases and transfer proteins in phosphatidylserine subcellular distribution. Traffic 2015, 16, 35–47. [Google Scholar] [CrossRef]
- Laulagnier, K.; Grand, D.; Dujardin, A.; Hamdi, S.; Vincent-Schneider, H.; Lankar, D.; Salles, J.P.; Bonnerot, C.; Perret, B.; Record, M. PLD2 is enriched on exosomes and its activity is correlated to the release of exosomes. FEBS Lett. 2004, 572, 11–14. [Google Scholar] [CrossRef]
- Bicalho, B.; Holovati, J.L.; Acker, J.P. Phospholipidomics reveals differences in glycerophosphoserine profiles of hypothermically stored red blood cells and microvesicles. Biochim. Biophys. Acta 2013, 1828, 317–326. [Google Scholar] [CrossRef] [PubMed]
- Abels, E.R.; Breakefield, X.O. Introduction to Extracellular Vesicles: Biogenesis, RNA Cargo Selection, Content, Release, and Uptake. Cell. Mol. Neurobiol. 2016, 36, 301–312. [Google Scholar] [CrossRef] [PubMed]
- Huber, V.; Fais, S.; Iero, M.; Lugini, L.; Canese, P.; Squarcina, P.; Zaccheddu, A.; Colone, M.; Arancia, G.; Gentile, M.; et al. Human Colorectal Cancer Cells Induce T-Cell Death Through Release of Proapoptotic Microvesicles: Role in Immune Escape. Gastroenterology 2005, 128, 1796–1804. [Google Scholar] [CrossRef]
- Pigati, L.; Yaddanapudi, S.C.; Iyengar, R.; Kim, D.J.; Hearn, S.A.; Danforth, D.; Hastings, M.L.; Duelli, D.M. Selective release of microRNA species from normal and malignant mammary epithelial cells. PLoS ONE 2010, 5, e13515. [Google Scholar] [CrossRef]
- Sukati, S.; Ho, J.; Chaiswing, L.; Sompol, P.; Pandit, H.; Wei, W.; Izumi, T.; Chen, Q.; Weiss, H.; Noel, T.; et al. Extracellular vesicles released after cranial radiation: An insight into an early mechanism of brain injury. Brain Res. 2022, 1782, 147840. [Google Scholar] [CrossRef] [PubMed]
- Vinik, Y.; Ortega, F.G.; Mills, G.B.; Lu, Y.; Jurkowicz, M.; Halperin, S.; Aharoni, M.; Gutman, M.; Lev, S. Proteomic analysis of circulating extracellular vesicles identifies potential markers of breast cancer progression, recurrence, and response. Sci. Adv. 2020, 6. [Google Scholar] [CrossRef] [PubMed]
- Bandu, R.; Oh, J.W.; Kim, K.P. Mass spectrometry-based proteome profiling of extracellular vesicles and their roles in cancer biology. Exp. Mol. Med. 2019, 51, 1–10. [Google Scholar] [CrossRef]
- Skliar, M.; Chernyshev, V.S. Imaging of Extracellular Vesicles by Atomic Force Microscopy. J. Vis. Exp. 2019. [Google Scholar] [CrossRef]
- Nguyen, U.; Squaglia, N.; Boge, A.; Fung, P.A. The Simple Western™: A gel-free, blot-free, hands-free Western blotting reinvention. Nat. Methods 2011, 8, v. [Google Scholar] [CrossRef]
- Cvjetkovic, A.; Jang, S.C.; Konečná, B.; Höög, J.L.; Sihlbom, C.; Lässer, C.; Lötvall, J. Detailed Analysis of Protein Topology of Extracellular Vesicles-Evidence of Unconventional Membrane Protein Orientation. Sci. Rep. 2016, 6, 36338. [Google Scholar] [CrossRef]
- Gardiner, C.; Vizio, D.D.; Sahoo, S.; Théry, C.; Witwer, K.W.; Wauben, M.; Hill, A.F. Techniques used for the isolation and characterization of extracellular vesicles: Results of a worldwide survey. J. Extracell. Vesicles 2016, 5, 32945. [Google Scholar] [CrossRef] [PubMed]
- Carnino, J.M.; Lee, H.; Jin, Y. Isolation and characterization of extracellular vesicles from Broncho-alveolar lavage fluid: A review and comparison of different methods. Respir. Res. 2019, 20, 240. [Google Scholar] [CrossRef] [PubMed]
- Brennan, K.; Martin, K.; FitzGerald, S.P.; O’Sullivan, J.; Wu, Y.; Blanco, A.; Richardson, C.; Mc Gee, M.M. A comparison of methods for the isolation and separation of extracellular vesicles from protein and lipid particles in human serum. Sci. Rep. 2020, 10, 1039. [Google Scholar] [CrossRef] [PubMed]
- Cvjetkovic, A.; Lötvall, J.; Lässer, C. The influence of rotor type and centrifugation time on the yield and purity of extracellular vesicles. J. Extracell. Vesicles 2014, 3, 23111. [Google Scholar] [CrossRef]
- Reuter, S.; Gupta, S.C.; Chaturvedi, M.M.; Aggarwal, B.B. Oxidative stress, inflammation, and cancer: How are they linked? Free Radic. Biol. Med. 2010, 49, 1603–1616. [Google Scholar] [CrossRef]
- Xia, C.; Meng, Q.; Liu, L.Z.; Rojanasakul, Y.; Wang, X.R.; Jiang, B.H. Reactive oxygen species regulate angiogenesis and tumor growth through vascular endothelial growth factor. Cancer Res. 2007, 67, 10823–10830. [Google Scholar] [CrossRef]
- Kang, S.W.; Lee, S.; Lee, E.K. ROS and energy metabolism in cancer cells: Alliance for fast growth. Arch. Pharm. Res. 2015, 38, 338–345. [Google Scholar] [CrossRef]
- Brand, M.D. The sites and topology of mitochondrial superoxide production. Exp. Gerontol. 2010, 45, 466–472. [Google Scholar] [CrossRef]
- Chaiswing, L.; Zhong, W.; Cullen, J.J.; Oberley, L.W.; Oberley, T.D. Extracellular redox state regulates features associated with prostate cancer cell invasion. Cancer Res. 2008, 68, 5820–5826. [Google Scholar] [CrossRef]
- Harris, C.; Hansen, J.M. Oxidative stress, thiols, and redox profiles. Methods Mol. Biol. 2012, 889, 325–346. [Google Scholar] [CrossRef]
- Jones, D.P.; Go, Y.M.; Anderson, C.L.; Ziegler, T.R.; Kinkade, J.M., Jr.; Kirlin, W.G. Cysteine/cystine couple is a newly recognized node in the circuitry for biologic redox signaling and control. FASEB J. 2004, 18, 1246–1248. [Google Scholar] [CrossRef] [PubMed]
- Fukai, T.; Ushio-Fukai, M. Superoxide dismutases: Role in redox signaling, vascular function, and diseases. Antioxid. Redox Signal. 2011, 15, 1583–1606. [Google Scholar] [CrossRef] [PubMed]
- Yarana, C.; St Clair, D.K. Chemotherapy-Induced Tissue Injury: An Insight into the Role of Extracellular Vesicles-Mediated Oxidative Stress Responses. Antioxidants 2017, 6, 75. [Google Scholar] [CrossRef] [PubMed]
- Wardman, P.; Candeias, L.P. Fenton Chemistry: An Introduction. Radiat. Res. 1996, 145, 523–531. [Google Scholar] [CrossRef] [PubMed]
- Dhar, S.K.; St Clair, D.K. Manganese superoxide dismutase regulation and cancer. Free Radic. Biol. Med. 2012, 52, 2209–2222. [Google Scholar] [CrossRef] [PubMed]
- Miriyala, S.; Spasojevic, I.; Tovmasyan, A.; Salvemini, D.; Vujaskovic, Z.; St Clair, D.; Batinic-Haberle, I. Manganese superoxide dismutase, MnSOD and its mimics. Biochim. Biophys. Acta 2012, 1822, 794–814. [Google Scholar] [CrossRef]
- Nandi, A.; Yan, L.-J.; Jana, C.K.; Das, N. Role of Catalase in Oxidative Stress- and Age-Associated Degenerative Diseases. Oxidative Med. Cell. Longev. 2019, 2019, 9613090. [Google Scholar] [CrossRef]
- Lubos, E.; Loscalzo, J.; Handy, D.E. Glutathione peroxidase-1 in health and disease: From molecular mechanisms to therapeutic opportunities. Antioxid. Redox Signal. 2011, 15, 1957–1997. [Google Scholar] [CrossRef]
- Valko, M.; Leibfritz, D.; Moncol, J.; Cronin, M.T.; Mazur, M.; Telser, J. Free radicals and antioxidants in normal physiological functions and human disease. Int. J. Biochem. Cell Biol. 2007, 39, 44–84. [Google Scholar] [CrossRef]
- Shadel, G.S.; Horvath, T.L. Mitochondrial ROS signaling in organismal homeostasis. Cell 2015, 163, 560–569. [Google Scholar] [CrossRef]
- Wilson, C.; Gonzalez-Billault, C. Regulation of cytoskeletal dynamics by redox signaling and oxidative stress: Implications for neuronal development and trafficking. Front. Cell. Neurosci. 2015, 9, 381. [Google Scholar] [CrossRef] [PubMed]
- Li, J.; Stouffs, M.; Serrander, L.; Banfi, B.; Bettiol, E.; Charnay, Y.; Steger, K.; Krause, K.H.; Jaconi, M.E. The NADPH oxidase NOX4 drives cardiac differentiation: Role in regulating cardiac transcription factors and MAP kinase activation. Mol. Biol. Cell 2006, 17, 3978–3988. [Google Scholar] [CrossRef] [PubMed]
- Veal, E.A.; Day, A.M.; Morgan, B.A. Hydrogen peroxide sensing and signaling. Mol. Cell 2007, 26, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Florence, T.M. The production of hydroxyl radical from hydrogen peroxide. J. Inorg. Biochem. 1984, 22, 221–230. [Google Scholar] [CrossRef]
- Sonntag, C.V. The Chemical Basis of Radiation Biology; Taylor & Francis: London, UK; Philadelphia, PA, USA, 1989. [Google Scholar]
- Liou, G.Y.; Storz, P. Reactive oxygen species in cancer. Free Radic. Res. 2010, 44, 479–496. [Google Scholar] [CrossRef]
- Chan, D.W.; Liu, V.W.; Tsao, G.S.; Yao, K.M.; Furukawa, T.; Chan, K.K.; Ngan, H.Y. Loss of MKP3 mediated by oxidative stress enhances tumorigenicity and chemoresistance of ovarian cancer cells. Carcinogenesis 2008, 29, 1742–1750. [Google Scholar] [CrossRef]
- Chaiswing, L.; Bourdeau-Heller, J.M.; Zhong, W.; Oberley, T.D. Characterization of redox state of two human prostate carcinoma cell lines with different degrees of aggressiveness. Free Radic. Biol. Med. 2007, 43, 202–215. [Google Scholar] [CrossRef]
- Zhou, D.; Shao, L.; Spitz, D.R. Reactive oxygen species in normal and tumor stem cells. Adv. Cancer Res. 2014, 122, 1–67. [Google Scholar] [CrossRef]
- Ralph, S.J.; Rodríguez-Enríquez, S.; Neuzil, J.; Saavedra, E.; Moreno-Sánchez, R. The causes of cancer revisited: “mitochondrial malignancy” and ROS-induced oncogenic transformation - why mitochondria are targets for cancer therapy. Mol. Asp. Med. 2010, 31, 145–170. [Google Scholar] [CrossRef]
- Conklin, K.A. Chemotherapy-Associated Oxidative Stress: Impact on Chemotherapeutic Effectiveness. Integr. Cancer Ther. 2004, 3, 294–300. [Google Scholar] [CrossRef]
- Li, Y.; Xue, Z.; Dong, X.; Liu, Q.; Liu, Z.; Li, H.; Xing, H.; Xu, Y.; Tang, K.; Tian, Z.; et al. Mitochondrial dysfunction and oxidative stress in bone marrow stromal cells induced by daunorubicin leads to DNA damage in hematopoietic cells. Free Radic. Biol. Med. 2020, 146, 211–221. [Google Scholar] [CrossRef] [PubMed]
- Doroshow, J.H.; Davies, K.J. Redox cycling of anthracyclines by cardiac mitochondria. II. Formation of superoxide anion, hydrogen peroxide, and hydroxyl radical. J. Biol. Chem. 1986, 261, 3068–3074. [Google Scholar] [CrossRef]
- Ayala, A.; Muñoz, M.F.; Argüelles, S. Lipid Peroxidation: Production, Metabolism, and Signaling Mechanisms of Malondialdehyde and 4-Hydroxy-2-Nonenal. Oxidative Med. Cell. Longev. 2014, 2014, 360438. [Google Scholar] [CrossRef] [PubMed]
- Yen, H.C.; Oberley, T.D.; Vichitbandha, S.; Ho, Y.S.; St Clair, D.K. The protective role of manganese superoxide dismutase against adriamycin-induced acute cardiac toxicity in transgenic mice. J. Clin. Investig. 1996, 98, 1253–1260. [Google Scholar] [CrossRef] [PubMed]
- Chaiswing, L.; Cole, M.P.; St Clair, D.K.; Ittarat, W.; Szweda, L.I.; Oberley, T.D. Oxidative damage precedes nitrative damage in adriamycin-induced cardiac mitochondrial injury. Toxicol. Pathol. 2004, 32, 536–547. [Google Scholar] [CrossRef]
- Nithipongvanitch, R.; Ittarat, W.; Velez, J.M.; Zhao, R.; St Clair, D.K.; Oberley, T.D. Evidence for p53 as guardian of the cardiomyocyte mitochondrial genome following acute adriamycin treatment. J. Histochem. Cytochem. 2007, 55, 629–639. [Google Scholar] [CrossRef]
- Chen, Y.; Daosukho, C.; Opii, W.O.; Turner, D.M.; Pierce, W.M.; Klein, J.B.; Vore, M.; Butterfield, D.A.; St Clair, D.K. Redox proteomic identification of oxidized cardiac proteins in adriamycin-treated mice. Free Radic. Biol. Med. 2006, 41, 1470–1477. [Google Scholar] [CrossRef]
- Petersen, D.R.; Doorn, J.A. Reactions of 4-hydroxynonenal with proteins and cellular targets. Free Radic. Biol. Med. 2004, 37, 937–945. [Google Scholar] [CrossRef]
- Butterfield, D.A.; Halliwell, B. Oxidative stress, dysfunctional glucose metabolism and Alzheimer disease. Nat. Rev. Neurosci. 2019, 20, 148–160. [Google Scholar] [CrossRef]
- Grune, T.; Davies, K.J.A. The proteasomal system and HNE-modified proteins. Mol. Asp. Med. 2003, 24, 195–204. [Google Scholar] [CrossRef]
- Markesbery, W.R.; Lovell, M.A. Four-hydroxynonenal, a product of lipid peroxidation, is increased in the brain in Alzheimer’s disease. Neurobiol. Aging 1998, 19, 33–36. [Google Scholar] [CrossRef]
- Bruce-Keller, A.J.; Li, Y.J.; Lovell, M.A.; Kraemer, P.J.; Gary, D.S.; Brown, R.R.; Markesbery, W.R.; Mattson, M.P. 4-Hydroxynonenal, a product of lipid peroxidation, damages cholinergic neurons and impairs visuospatial memory in rats. J. Neuropathol. Exp. Neurol. 1998, 57, 257–267. [Google Scholar] [CrossRef] [PubMed]
- Perry, E.A.; Castellani, R.J.; Moreira, P.I.; Nunomura, A.; Lui, Q.; Harris, P.L.; Sayre, L.M.; Szweda, P.A.; Szweda, L.I.; Zhu, X.; et al. Neurofilaments are the major neuronal target of hydroxynonenal-mediated protein cross-links. Free Radic. Res. 2013, 47, 507–510. [Google Scholar] [CrossRef] [PubMed][Green Version]
- Shringarpure, R.; Grune, T.; Sitte, N.; Davies, K.J. 4-Hydroxynonenal-modified amyloid-beta peptide inhibits the proteasome: Possible importance in Alzheimer’s disease. Cell. Mol. Life Sci. 2000, 57, 1802–1809. [Google Scholar] [CrossRef] [PubMed]
- VanWinkle, W.B.; Snuggs, M.; Miller, J.C.; Buja, L.M. Cytoskeletal alterations in cultured cardiomyocytes following exposure to the lipid peroxidation product, 4-hydroxynonenal. Cell Motil. Cytoskelet. 1994, 28, 119–134. [Google Scholar] [CrossRef] [PubMed]
- Leonarduzzi, G.; Chiarpotto, E.; Biasi, F.; Poli, G. 4-Hydroxynonenal and cholesterol oxidation products in atherosclerosis. Mol. Nutr. Food Res. 2005, 49, 1044–1049. [Google Scholar] [CrossRef]
- Zhao, Y.; Miriyala, S.; Miao, L.; Mitov, M.; Schnell, D.; Dhar, S.K.; Cai, J.; Klein, J.B.; Sultana, R.; Butterfield, D.A.; et al. Redox proteomic identification of HNE-bound mitochondrial proteins in cardiac tissues reveals a systemic effect on energy metabolism after doxorubicin treatment. Free. Radic. Biol. Med. 2014, 72, 55–65. [Google Scholar] [CrossRef]
- Yarana, C.; Carroll, D.; Chen, J.; Chaiswing, L.; Zhao, Y.; Noel, T.; Alstott, M.; Bae, Y.; Dressler, E.V.; Moscow, J.A.; et al. Extracellular Vesicles Released by Cardiomyocytes in a Doxorubicin-Induced Cardiac Injury Mouse Model Contain Protein Biomarkers of Early Cardiac Injury. Clin. Cancer Res. 2018, 24, 1644–1653. [Google Scholar] [CrossRef]
- Chaiswing, L.; St Clair, W.H.; St Clair, D.K. Redox Paradox: A Novel Approach to Therapeutics-Resistant Cancer. Antioxid. Redox Signal. 2018, 29, 1237–1272. [Google Scholar] [CrossRef]
- El Andaloussi, S.; Mager, I.; Breakefield, X.O.; Wood, M.J. Extracellular vesicles: Biology and emerging therapeutic opportunities. Nat. Rev. Drug Discov. 2013, 12, 347–357. [Google Scholar] [CrossRef]
- Alexander, M.; Hu, R.; Runtsch, M.C.; Kagele, D.A.; Mosbruger, T.L.; Tolmachova, T.; Seabra, M.C.; Round, J.L.; Ward, D.M.; O’Connell, R.M. Exosome-delivered microRNAs modulate the inflammatory response to endotoxin. Nat. Commun. 2015, 6, 7321. [Google Scholar] [CrossRef] [PubMed]
- Ratajczak, M.Z.; Kucia, M.; Jadczyk, T.; Greco, N.J.; Wojakowski, W.; Tendera, M.; Ratajczak, J. Pivotal role of paracrine effects in stem cell therapies in regenerative medicine: Can we translate stem cell-secreted paracrine factors and microvesicles into better therapeutic strategies? Leukemia 2012, 26, 1166–1173. [Google Scholar] [CrossRef] [PubMed]
- Xu, R.; Rai, A.; Chen, M.; Suwakulsiri, W.; Greening, D.W.; Simpson, R.J. Extracellular vesicles in cancer - implications for future improvements in cancer care. Nat. Rev. Clin. Oncol. 2018, 15, 617–638. [Google Scholar] [CrossRef]
- Sung, B.H.; Ketova, T.; Hoshino, D.; Zijlstra, A.; Weaver, A.M. Directional cell movement through tissues is controlled by exosome secretion. Nat. Commun. 2015, 6, 7164. [Google Scholar] [CrossRef] [PubMed]
- Yang, M.; Chen, J.; Su, F.; Yu, B.; Su, F.; Lin, L.; Liu, Y.; Huang, J.D.; Song, E. Microvesicles secreted by macrophages shuttle invasion-potentiating microRNAs into breast cancer cells. Mol. Cancer 2011, 10, 117. [Google Scholar] [CrossRef] [PubMed]
- Buzas, E.I.; György, B.; Nagy, G.; Falus, A.; Gay, S. Emerging role of extracellular vesicles in inflammatory diseases. Nat. Rev. Rheumatol. 2014, 10, 356–364. [Google Scholar] [CrossRef]
- Thompson, A.G.; Gray, E.; Heman-Ackah, S.M.; Mager, I.; Talbot, K.; Andaloussi, S.E.; Wood, M.J.; Turner, M.R. Extracellular vesicles in neurodegenerative disease—pathogenesis to biomarkers. Nat. Rev. Neurol. 2016, 12, 346–357. [Google Scholar] [CrossRef]
- Aykin-Burns, N.; Ahmad, I.M.; Zhu, Y.; Oberley, L.W.; Spitz, D.R. Increased levels of superoxide and H2O2 mediate the differential susceptibility of cancer cells versus normal cells to glucose deprivation. Biochem. J. 2009, 418, 29–37. [Google Scholar] [CrossRef]
- Ohno, S.; Takanashi, M.; Sudo, K.; Ueda, S.; Ishikawa, A.; Matsuyama, N.; Fujita, K.; Mizutani, T.; Ohgi, T.; Ochiya, T.; et al. Systemically injected exosomes targeted to EGFR deliver antitumor microRNA to breast cancer cells. Mol. Ther. 2013, 21, 185–191. [Google Scholar] [CrossRef]
- Yao, J.; Wang, Z.; Cheng, Y.; Ma, C.; Zhong, Y.; Xiao, Y.; Gao, X.; Li, Z. M2 macrophage-derived exosomal microRNAs inhibit cell migration and invasion in gliomas through PI3K/AKT/mTOR signaling pathway. J. Transl. Med. 2021, 19, 99. [Google Scholar] [CrossRef]
- Walker, S.; Busatto, S.; Pham, A.; Tian, M.; Suh, A.; Carson, K.; Quintero, A.; Lafrence, M.; Malik, H.; Santana, M.X.; et al. Extracellular vesicle-based drug delivery systems for cancer treatment. Theranostics 2019, 9, 8001–8017. [Google Scholar] [CrossRef] [PubMed]
- Bunggulawa, E.J.; Wang, W.; Yin, T.; Wang, N.; Durkan, C.; Wang, Y.; Wang, G. Recent advancements in the use of exosomes as drug delivery systems. J. Nanobiotechnol. 2018, 16, 81. [Google Scholar] [CrossRef] [PubMed]
- Goetzl, E.J.; Abner, E.L.; Jicha, G.A.; Kapogiannis, D.; Schwartz, J.B. Declining levels of functionally specialized synaptic proteins in plasma neuronal exosomes with progression of Alzheimer’s disease. FASEB J. 2018, 32, 888–893. [Google Scholar] [CrossRef]
- Balaj, L.; Lessard, R.; Dai, L.; Cho, Y.-J.; Pomeroy, S.L.; Breakefield, X.O.; Skog, J. Tumour microvesicles contain retrotransposon elements and amplified oncogene sequences. Nat. Commun. 2011, 2, 180. [Google Scholar] [CrossRef] [PubMed]
- Kahlert, C.; Melo, S.A.; Protopopov, A.; Tang, J.; Seth, S.; Koch, M.; Zhang, J.; Weitz, J.; Chin, L.; Futreal, A.; et al. Identification of double-stranded genomic DNA spanning all chromosomes with mutated KRAS and p53 DNA in the serum exosomes of patients with pancreatic cancer. J. Biol. Chem. 2014, 289, 3869–3875. [Google Scholar] [CrossRef]
- Liu, C.; Zhao, J.; Tian, F.; Cai, L.; Zhang, W.; Feng, Q.; Chang, J.; Wan, F.; Yang, Y.; Dai, B.; et al. Low-cost thermophoretic profiling of extracellular-vesicle surface proteins for the early detection and classification of cancers. Nat. Biomed. Eng. 2019, 3, 183–193. [Google Scholar] [CrossRef]
- Baulch, J.E.; Acharya, M.M.; Allen, B.D.; Ru, N.; Chmielewski, N.N.; Martirosian, V.; Giedzinski, E.; Syage, A.; Park, A.L.; Benke, S.N.; et al. Cranial grafting of stem cell-derived microvesicles improves cognition and reduces neuropathology in the irradiated brain. Proc. Natl. Acad. Sci. USA 2016, 113, 4836–4841. [Google Scholar] [CrossRef]
- Tome, W.A.; Gokhan, S.; Brodin, N.P.; Gulinello, M.E.; Heard, J.; Mehler, M.F.; Guha, C. A mouse model replicating hippocampal sparing cranial irradiation in humans: A tool for identifying new strategies to limit neurocognitive decline. Sci. Rep. 2015, 5, 14384. [Google Scholar] [CrossRef]
- Pui, C.H.; Campana, D.; Pei, D.; Bowman, W.P.; Sandlund, J.T.; Kaste, S.C.; Ribeiro, R.C.; Rubnitz, J.E.; Raimondi, S.C.; Onciu, M.; et al. Treating childhood acute lymphoblastic leukemia without cranial irradiation. N. Engl. J. Med. 2009, 360, 2730–2741. [Google Scholar] [CrossRef]
- Hardy, S.J.; Krull, K.R.; Wefel, J.S.; Janelsins, M. Cognitive Changes in Cancer Survivors. Am. Soc. Clin. Oncol. Educ. Book 2018, 795–806. [Google Scholar] [CrossRef]
- Howlader, N.; Noone, A.M.; Krapcho, M.; Miller, D.; Brest, A.; Yu, M.; Ruhl, J.; Tatalovich, Z.; Mariotto, A.; Lewis, D.R.; et al. SEER Cacer Statistics Review, 1975–2017; National Cancer Institute: Bethesda, MD, USA, 2020. Available online: https://seer.cancer.gov/csr/1975_2017/ (accessed on 25 April 2022).
- Brinkman, T.M.; Ness, K.K.; Li, Z.; Huang, I.C.; Krull, K.R.; Gajjar, A.; Merchant, T.E.; Klosky, J.L.; Partin, R.E.; Tonning Olsson, I.; et al. Attainment of Functional and Social Independence in Adult Survivors of Pediatric CNS Tumors: A Report From the St Jude Lifetime Cohort Study. J. Clin. Oncol. 2018, 36, 2762–2769. [Google Scholar] [CrossRef] [PubMed]
- Cooper, S.L.; Brown, P.A. Treatment of pediatric acute lymphoblastic leukemia. Pediatr. Clin. N. Am. 2015, 62, 61–73. [Google Scholar] [CrossRef] [PubMed]
- Terwilliger, T.; Abdul-Hay, M. Acute lymphoblastic leukemia: A comprehensive review and 2017 update. Blood Cancer J. 2017, 7, e577. [Google Scholar] [CrossRef] [PubMed]
- Angiolillo, A.L.; Schore, R.J.; Kairalla, J.A.; Devidas, M.; Rabin, K.R.; Zweidler-McKay, P.; Borowitz, M.J.; Wood, B.; Carroll, A.J.; Heerema, N.A.; et al. Excellent Outcomes With Reduced Frequency of Vincristine and Dexamethasone Pulses in Standard-Risk B-Lymphoblastic Leukemia: Results From Children’s Oncology Group AALL0932. J. Clin. Oncol. 2021, 39, 1437–1447. [Google Scholar] [CrossRef] [PubMed]
- Iacobini, M.; Menichelli, A.; Palumbo, G.; Multari, G.; Werner, B.; Del Principe, D. Involvement of oxygen radicals in cytarabine-induced apoptosis in human polymorphonuclear cells. Biochem. Pharmacol. 2001, 61, 1033–1040. [Google Scholar] [CrossRef]
- Gibson, E.M.; Nagaraja, S.; Ocampo, A.; Tam, L.T.; Wood, L.S.; Pallegar, P.N.; Greene, J.J.; Geraghty, A.C.; Goldstein, A.K.; Ni, L.; et al. Methotrexate Chemotherapy Induces Persistent Tri-glial Dysregulation that Underlies Chemotherapy-Related Cognitive Impairment. Cell 2019, 176, 43–55. [Google Scholar] [CrossRef]
- Baker, W.J.; Royer, G.L., Jr.; Weiss, R.B. Cytarabine and neurologic toxicity. J. Clin. Oncol. 1991, 9, 679–693. [Google Scholar] [CrossRef]
- Zheng, D.J.; Lu, X.; Schore, R.J.; Balsamo, L.; Devidas, M.; Winick, N.J.; Raetz, E.A.; Loh, M.L.; Carroll, W.L.; Sung, L.; et al. Longitudinal analysis of quality-of-life outcomes in children during treatment for acute lymphoblastic leukemia: A report from the Children’s Oncology Group AALL0932 trial. Cancer 2018, 124, 571–579. [Google Scholar] [CrossRef]
- Johansson, A.C.; Appelqvist, H.; Nilsson, C.; Kågedal, K.; Roberg, K.; Ollinger, K. Regulation of apoptosis-associated lysosomal membrane permeabilization. Apoptosis 2010, 15, 527–540. [Google Scholar] [CrossRef]
- Hara, T.; Nakamura, K.; Matsui, M.; Yamamoto, A.; Nakahara, Y.; Suzuki-Migishima, R.; Yokoyama, M.; Mishima, K.; Saito, I.; Okano, H.; et al. Suppression of basal autophagy in neural cells causes neurodegenerative disease in mice. Nature 2006, 441, 885–889. [Google Scholar] [CrossRef]
- Prolo, L.M.; Vogel, H.; Reimer, R.J. The lysosomal sialic acid transporter sialin is required for normal CNS myelination. J. Neurosci. 2009, 29, 15355–15365. [Google Scholar] [CrossRef] [PubMed]
| Parameter | Purpose | Suggested Methods | Additional Notes |
|---|---|---|---|
| 1. Quantification of EVs | To best quantify the amount of EVs present in a sample. Starting volume that EVs are being isolated from also needs to be taken into account [31]. | 1. Protein concentration (e.g., BCA, Bradford, total protein on SDS-PAGE) 2. EVs’ particle number quantification (e.g., nanoparticle tracking analysis, standard or high-resolution flow cytometry) | Either total protein as measured by BCA or particle number is most commonly used. Quantification of total lipids, specific molecules, or total RNA may also be used. |
| 2. General characteristics of EVs by protein composition | To quantify the purity of EVs isolation with a minimum of three positive markers (at least one transmembrane or lipid-bound protein and at least one cytosolic protein). Additionally, a negative or contamination marker must be used for a minimum total of at least four protein markers [31]. | 1. Western blotting 2. Flow cytometry 3. Capillary-based automated Western blot (Jess) [47] 4. Reverse phase protein array (RPPA) [48] 5. Mass spectrometry [49] | Two other categories of protein markers are suggested in MISEV2018 but are not required [31]: 1. Proteins localized in/on intracellular compartments of eukaryotic cells to identify specificity of small EVs’ subtype(s) (e.g., LMNA and CYC1) 2. Secreted or luminal proteins that can bind to receptors on the EV surface for mode of association of EVs (e.g., EGF, VEGFA, and collagen) |
| 3. Characterization of single vesicles | Provide some parameters regarding the individual EVs present in the bulk population of EVs that are being used for study. Two methods must be used. The first should provide a high-resolution image of the EVs and the second should calculate biophysical parameters of single EVs that can be used to quantify a large number of EVs [31]. | 1. Electron microscopy (SEM, TEM, cryo-EM), SPM, Atomic Force Microscopy (AFM) [50], super-resolution microscopy 2. Nanoparticle tracking analysis, high-resolution flow cytometry, fluorescence correlation spectroscopy | The authors of MISEV2018 provide many potential methods that can be used for the characterization of single vesicles but emphasize the significance of proper documentation of the experimental conditions, such as documentation of the source of the EVs, the starting volume of the source, the conditions of isolation, etc. |
| 4. Characterization of topology of EV-associated components | To determine the location of some proteins between the lumen and the surface of EVs [31]. | 1. Mild digestions, permeabilizations, or antibody studies followed by SDS-PAGE, RT-PCR, etc. 2. Flow cytometry and fluorescence microscopy with antibodies 3. EM with immunolabeling | Topology may be a result of unknown mechanisms localizing cytosolic components to the surface and may be important for function. |
| Method | Advantages | Disadvantages |
|---|---|---|
| Ultracentrifugation |
|
|
| Density gradient centrifugation |
|
|
| Filtration |
|
|
| Size exclusion chromatography |
|
|
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ho, J.; Chaiswing, L.; St. Clair, D.K. Extracellular Vesicles and Cancer Therapy: Insights into the Role of Oxidative Stress. Antioxidants 2022, 11, 1194. https://doi.org/10.3390/antiox11061194
Ho J, Chaiswing L, St. Clair DK. Extracellular Vesicles and Cancer Therapy: Insights into the Role of Oxidative Stress. Antioxidants. 2022; 11(6):1194. https://doi.org/10.3390/antiox11061194
Chicago/Turabian StyleHo, Jenni, Luksana Chaiswing, and Daret K. St. Clair. 2022. "Extracellular Vesicles and Cancer Therapy: Insights into the Role of Oxidative Stress" Antioxidants 11, no. 6: 1194. https://doi.org/10.3390/antiox11061194
APA StyleHo, J., Chaiswing, L., & St. Clair, D. K. (2022). Extracellular Vesicles and Cancer Therapy: Insights into the Role of Oxidative Stress. Antioxidants, 11(6), 1194. https://doi.org/10.3390/antiox11061194






