In Silico Study of Superoxide Dismutase Gene Family in Potato and Effects of Elevated Temperature and Salicylic Acid on Gene Expression
Abstract
:1. Introduction
2. Materials and Methods
2.1. Potato Genomic Resources
2.2. StSOD Promoter Analysis
2.3. In Silico Characterization of StSOD Protein Features
2.4. Phylogenetic Analysis of StSOD Protein Sequences
2.5. Plant Material and Growth Conditions
2.6. SA and Temperature Treatments
2.7. RNA Extraction and cDNA Synthesis
2.8. qRT-PCR Analysis
2.9. Statistical Analysis
3. Results
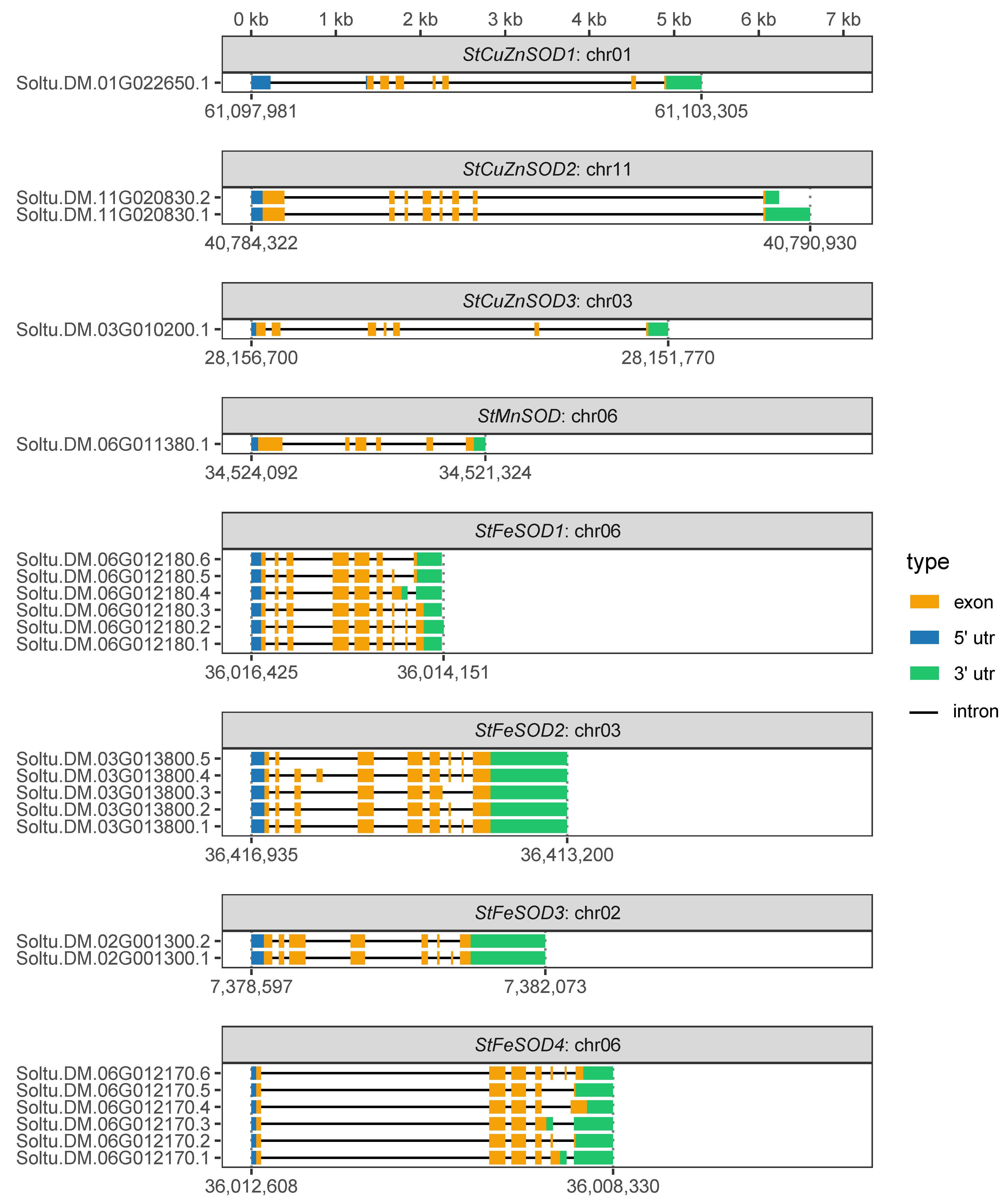
3.1. StSOD Genes: Structure and Chromosomal Distribution
3.2. Analysis of Promoter Regions of StSOD Genes
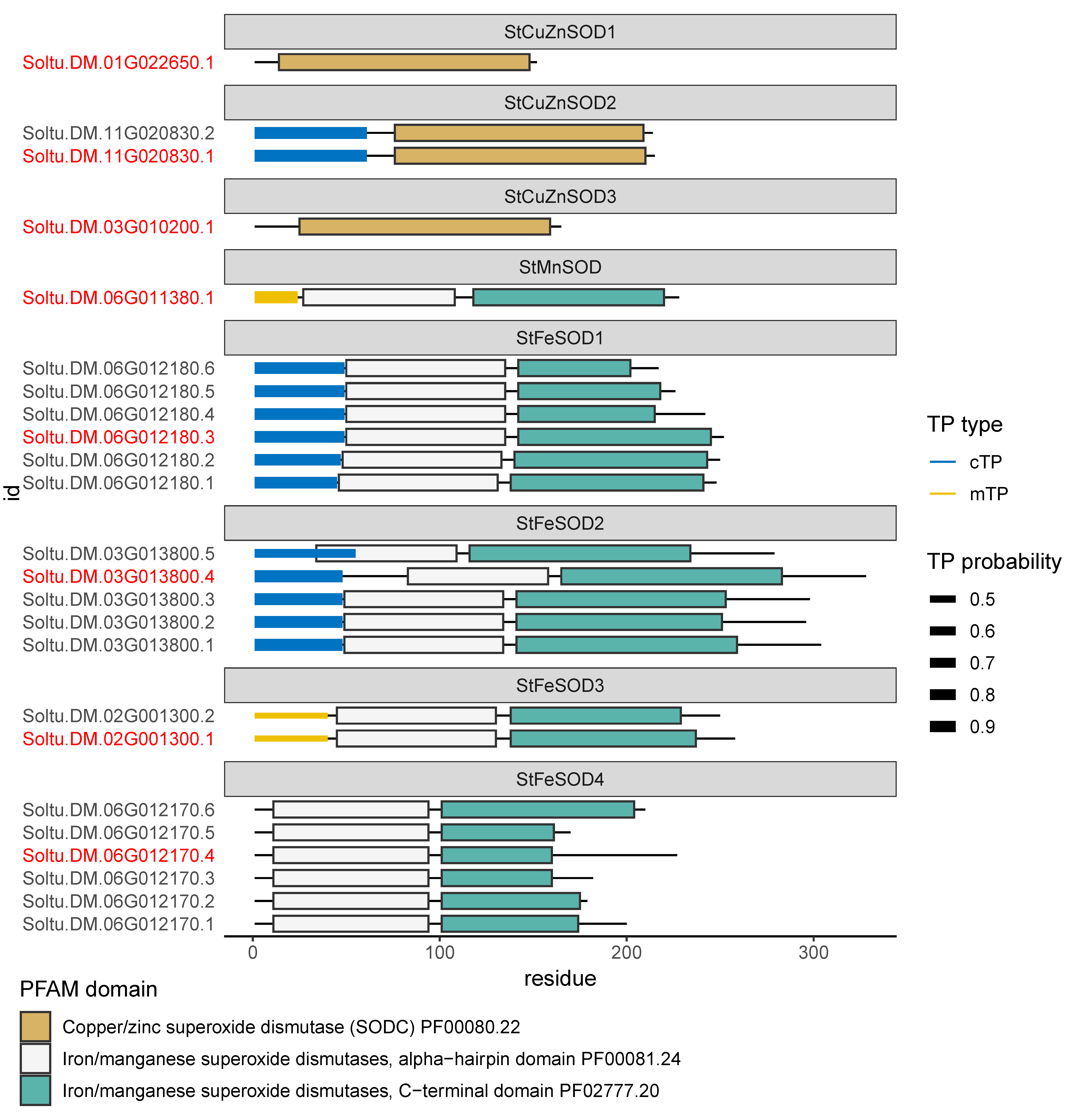
3.3. Characteristics of StSOD Proteins
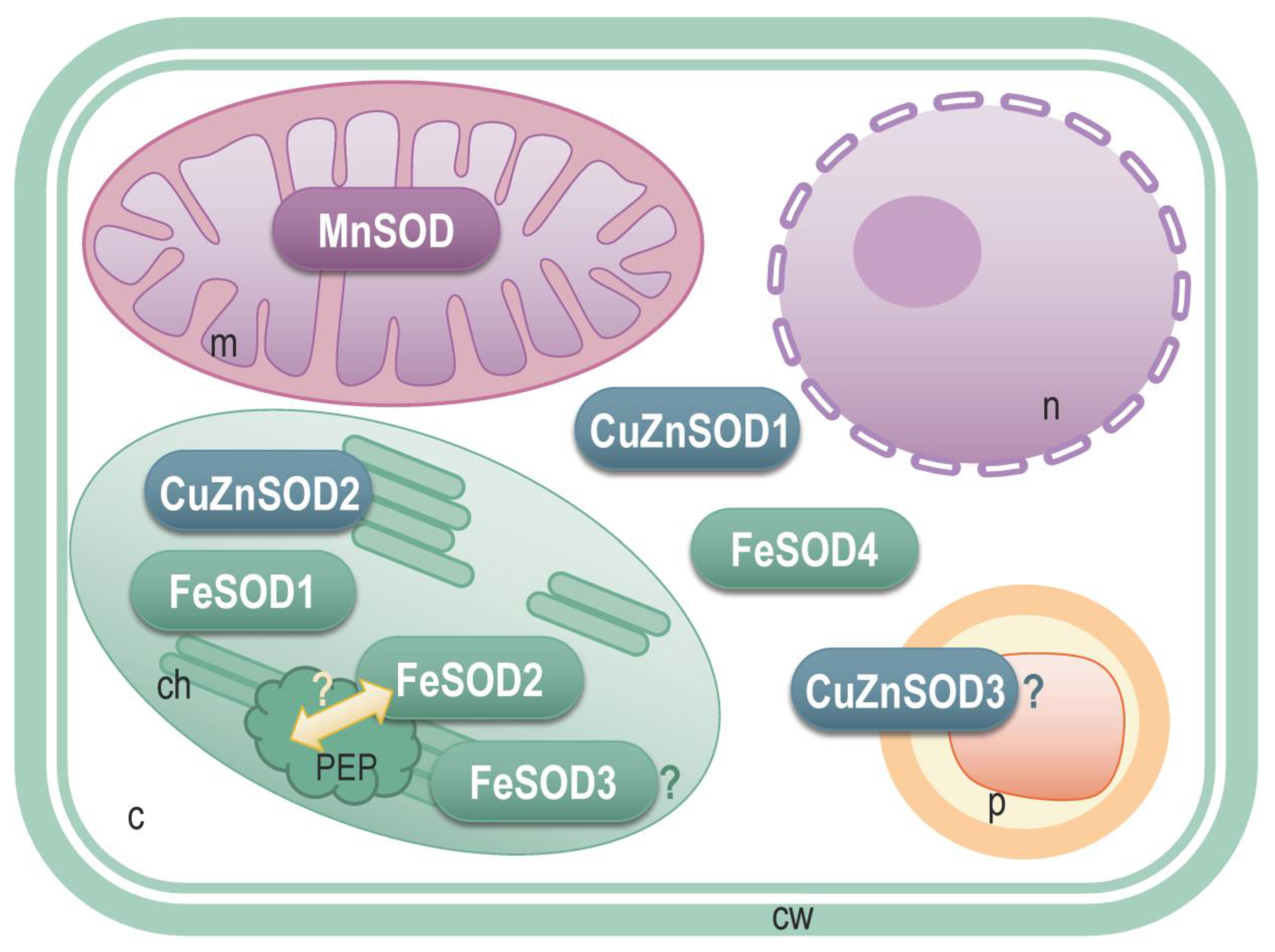
3.4. StSOD Protein Structure and Subcellular Localization
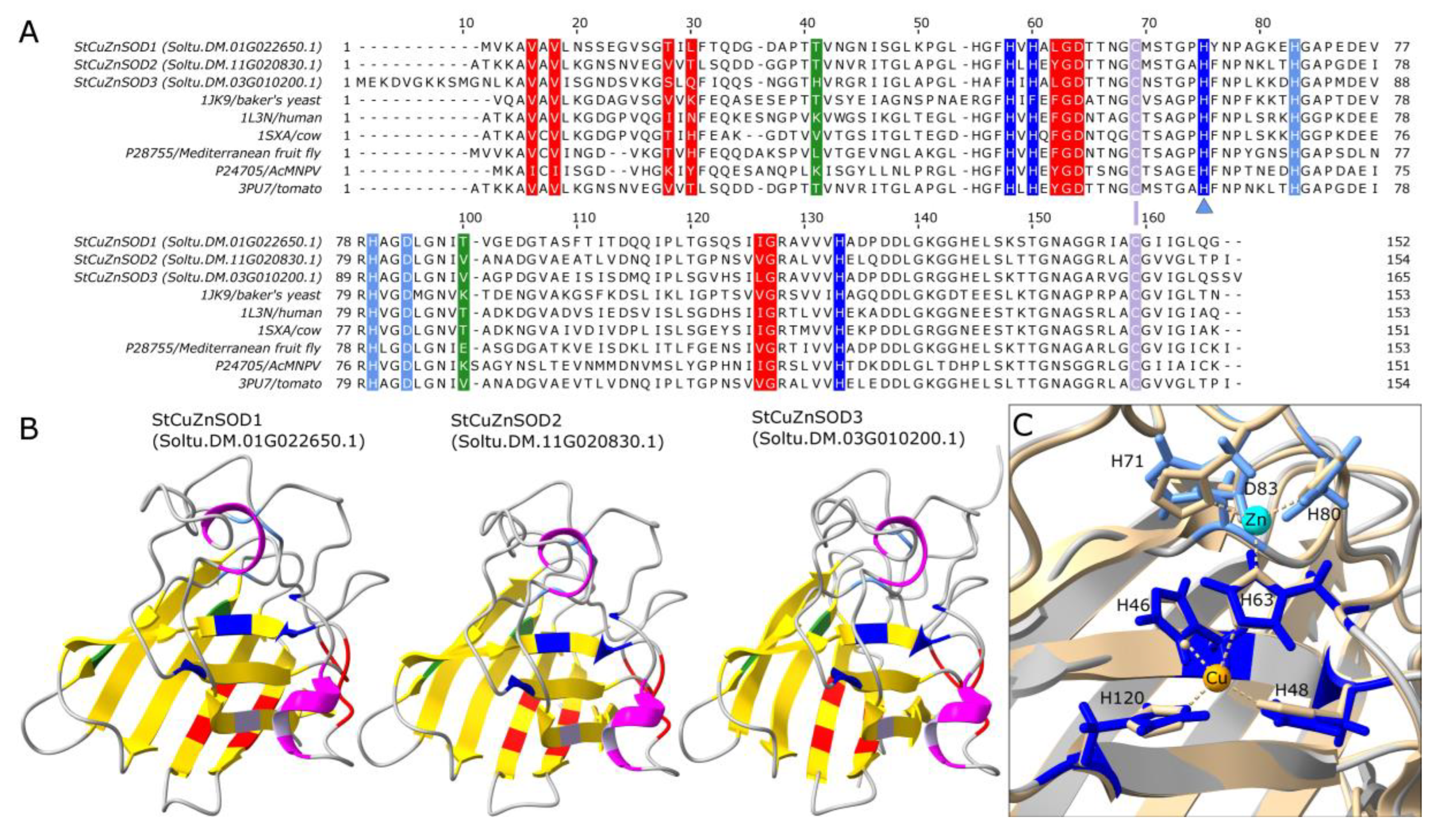
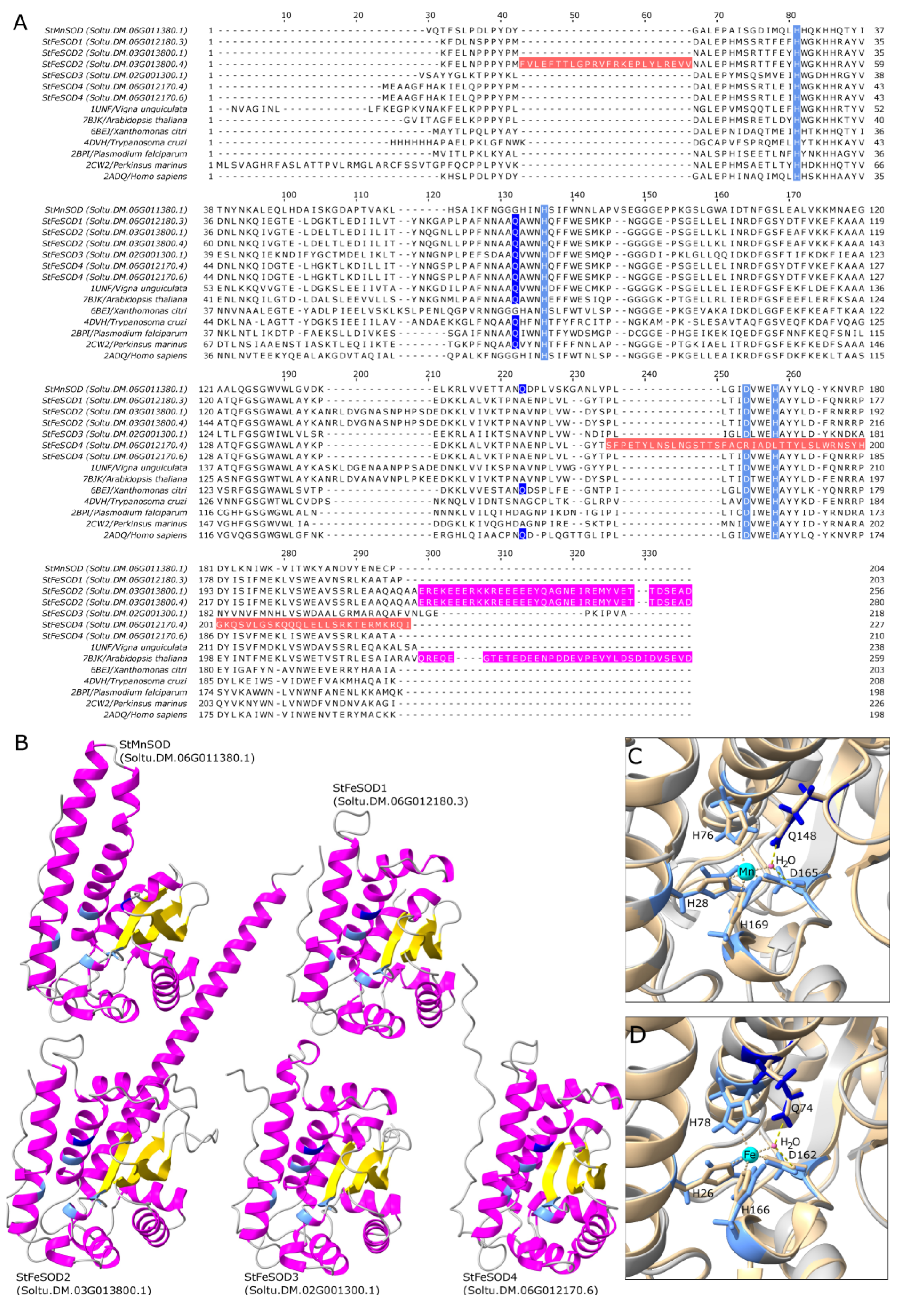
3.5. Predicted Tertiary Structure of Potato SODs
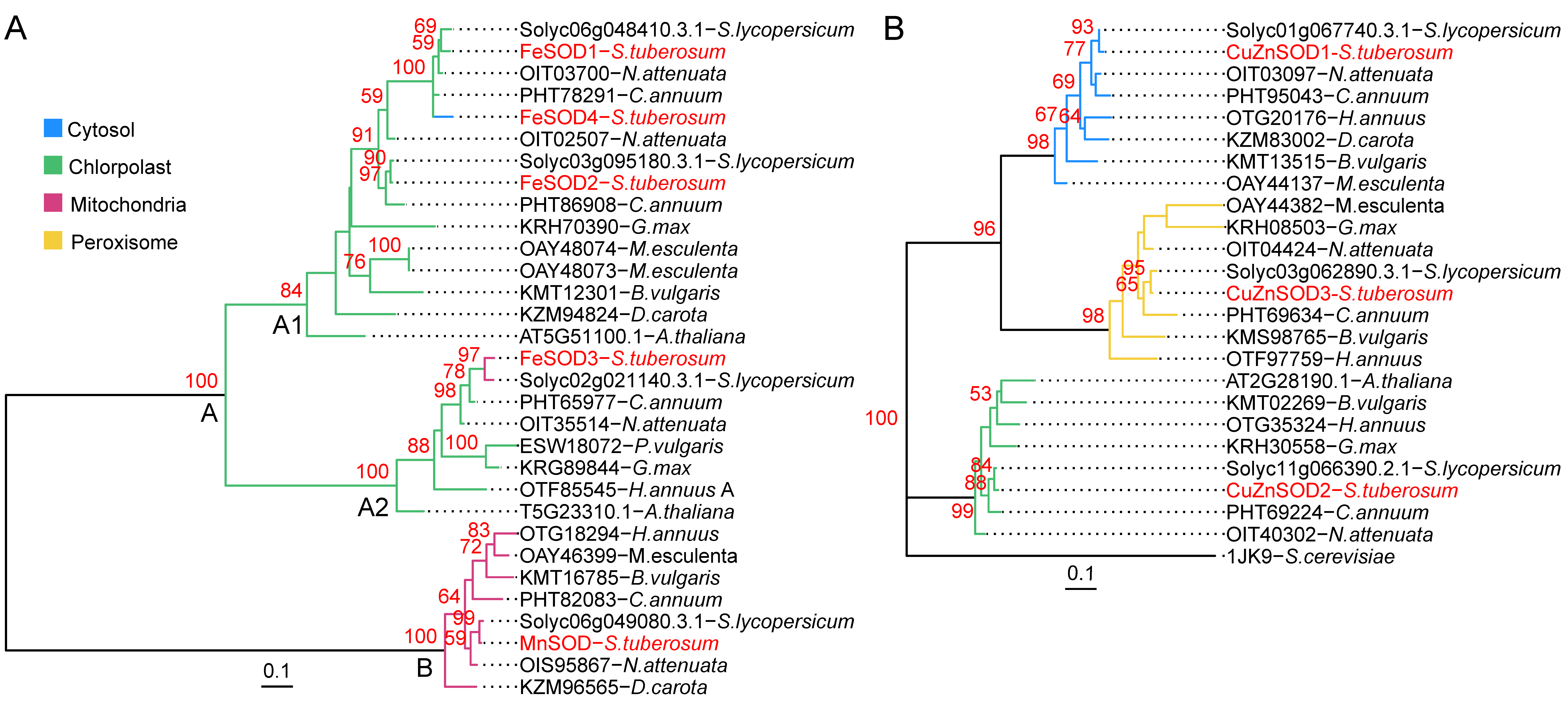
3.6. Phylogenetic Relations of StSOD Proteins
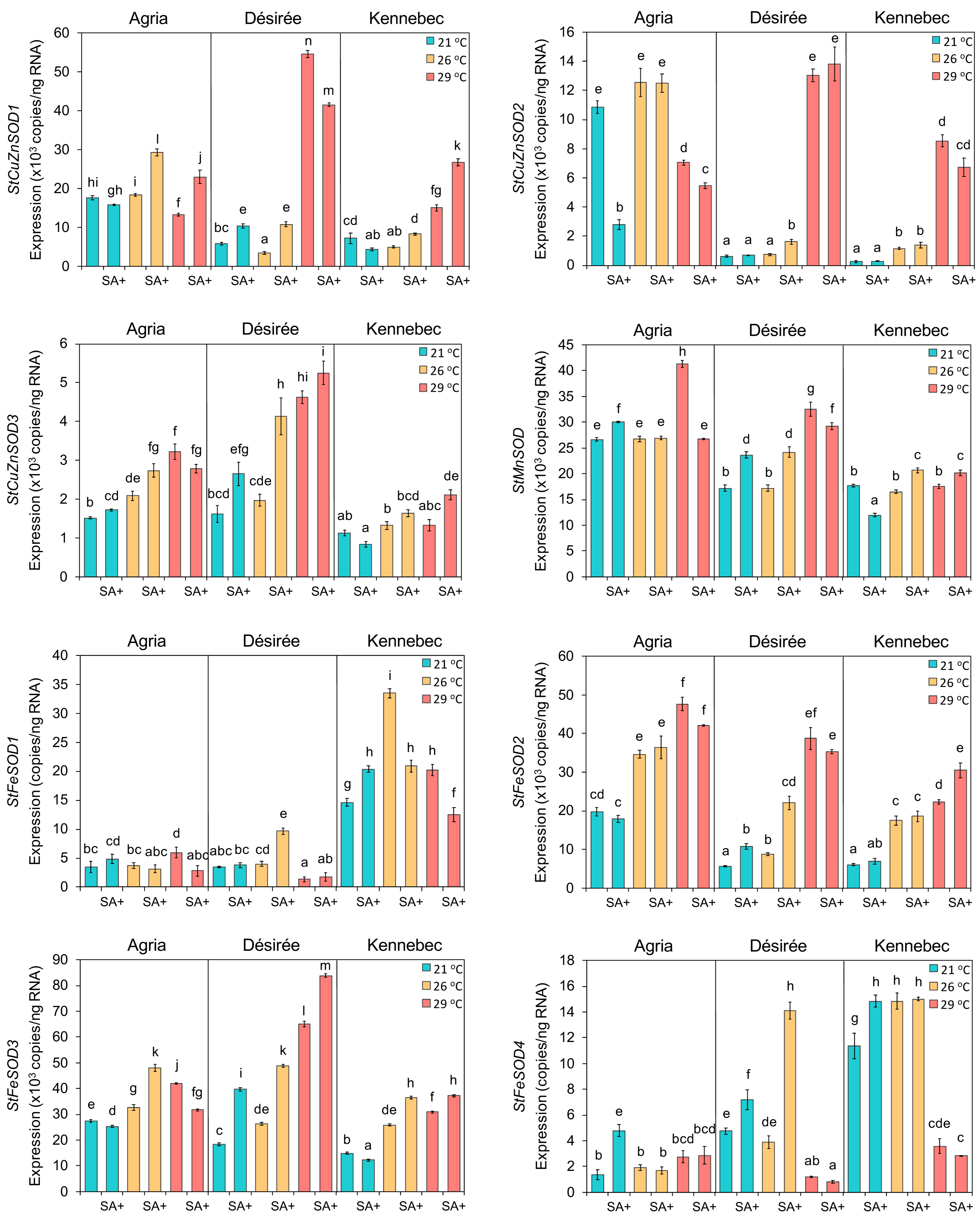
3.7. Expression Profiles of the StSOD Genes in Response to Elevated Temperatures and SA Application
4. Discussion
4.1. StSOD Gene Family
4.2. Tandem Duplication of the FeSOD Genes Is a Characteristic of Solanum Species
4.3. Gene Models of StSODs
4.4. Structural Features, Subcellular Localization and Phylogenetic Relations of StCuZnSODs
4.5. StMnSOD Is a Mitochondrial Enzyme with Distinguishable Structural Features
4.6. Not All StFeSOD Splice Variants Encode Functional Proteins
4.7. Expression of StSOD Genes in Response to Elevated Temperatures and Exogenous SA
4.8. StCuZnSOD1 Promoter Features Many More Cis-Acting Regulatory Elements Than Other StSOD Promoters and Is the Only One Predicted to Be Regulated by Heat or SA
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Struik, P.C. Responses of the potato plant to temperature. In Potato Biology and Biotechnology, 1st ed.; Vreugdenhil, D., Bradshaw, J., Gebhardt, C., Govers, F., Taylor, M., MacKerron, D., Ross, H., Eds.; Elsevier B.V.: Amsterdam, The Netherlands, 2007; pp. 367–393. [Google Scholar] [CrossRef]
- Hancock, R.D.; Morris, W.L.; Ducreux, L.J.; Morris, J.A.; Usman, M.; Verrall, S.R.; Fuller, J.; Simpson, C.G.; Zhang, R.; Hedley, P.E.; et al. Physiological, biochemical and molecular responses of the potato (Solanum tuberosum L.) plant to moderately elevated temperature. Plant Cell Environ. 2014, 37, 439–450. [Google Scholar] [CrossRef] [PubMed]
- Momčilović, I.; Fu, J.; Pantelić, D.; Rudić, J.; Broćić, Z. Impact of Heat Stress on Potato: Plant Responses and Approaches to Tolerance Improvement. In The Potato Crop: Management, Production, and Food Security; Villa, P.M., Ed.; Nova: New York, NY, USA, 2021; pp. 91–122. [Google Scholar] [CrossRef]
- Fu, J.; Momcilovic, I.; Prasad, P.V.V. Molecular bases and improvement of heat tolerance in crop plants. In Heat Stress: Causes, Prevention and Treatments; Josipovic, S., Ludwig, E., Eds.; Nova: New York, NY, USA, 2012; pp. 185–214. [Google Scholar]
- Momčilović, I.; Pantelić, D. Plant superoxide dismutases: Important players in abiotic stress tolerance. In Superoxide Dismutase: Structure, Synthesis and Applications; Magliozzi, S., Ed.; Nova: New York, NY, USA, 2018; pp. 45–100. [Google Scholar]
- Karuppanapandian, T.; Moon, J.C.; Kim, C.; Manoharan, K.; Kim, W. Reactive oxygen species in plants: Their generation, signal transduction, and scavenging mechanisms. Aust. J. Crop Sci. 2011, 5, 709–725. [Google Scholar]
- Devireddy, A.R.; Tschaplinski, T.J.; Tuskan, G.A.; Muchero, W.; Chen, J.G. Role of reactive oxygen species and hormones in plant responses to temperature changes. Int. J. Mol. Sci. 2021, 22, 8843. [Google Scholar] [CrossRef] [PubMed]
- Saleem, M.; Fariduddin, Q.; Castroverde, C.D.M. Salicylic acid: A key regulator of redox signalling and plant immunity. Plant Physiol. Biochem. 2021, 168, 381–397. [Google Scholar] [CrossRef]
- Gill, S.S.; Anjum, N.A.; Gill, R.; Yadav, S.; Hasanuzzaman, M.; Fujita, M.; Mishra, P.; Sabat, S.C.; Tuteja, N. Superoxide dismutase—Mentor of abiotic stress tolerance in crop plants. Environ. Sci. Pollut. Res. 2015, 22, 10375–10394. [Google Scholar] [CrossRef] [PubMed]
- Janků, M.; Luhová, L.; Petřivalský, M. On the origin and fate of reactive oxygen species in plant cell compartments. Antioxidants 2019, 8, 105. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kwon, S.Y.; Jeong, Y.J.; Lee, H.S.; Kim, J.S.; Cho, K.Y.; Allen, R.D.; Kwak, S.S. Enhanced tolerances of transgenic tobacco plants expressing both superoxide dismutase and ascorbate peroxidase in chloroplasts against methyl viologen-mediated oxidative stress. Plant Cell Environ. 2002, 25, 873–882. [Google Scholar] [CrossRef]
- Kaouthar, F.; Ameny, F.K.; Yosra, K.; Walid, S.; Ali, G.; Faiçal, B. Responses of transgenic Arabidopsis plants and recombinant yeast cells expressing a novel durum wheat manganese superoxide dismutase TdMnSOD to various abiotic stresses. J. Plant Physiol. 2016, 198, 56–68. [Google Scholar] [CrossRef] [PubMed]
- Waterer, D.; Benning, N.T.; Wu, G.; Luo, X.; Liu, X.; Gusta, M.; McHughen, A.; Gusta, L.V. Evaluation of abiotic stress tolerance of genetically modified potatoes (Solanum tuberosum cv. Desiree). Mol. Breed. 2010, 25, 527–540. [Google Scholar] [CrossRef]
- Mohamed, H.I.; El-Shazly, H.H.; Badr, A. Role of salicylic acid in biotic and abiotic stress tolerance in plants. In Plant Phenolics in Sustainable Agriculture; Lone, R., Shuab, R., Kamili, A., Eds.; Springer: Singapore, 2020; pp. 533–554. [Google Scholar] [CrossRef]
- Snyman, M.; Cronjé, M.J. Modulation of heat shock factors accompanies salicylic acid-mediated potentiation of Hsp70 in tomato seedlings. J. Exp. Bot. 2008, 59, 2125–2132. [Google Scholar] [CrossRef] [Green Version]
- Wang, L.J.; Fan, L.; Loescher, W.; Duan, W.; Liu, G.J.; Cheng, J.S.; Luo, H.B.; Li, S.H. Salicylic acid alleviates decreases in photosynthesis under heat stress and accelerates recovery in grapevine leaves. BMC Plant Biol. 2010, 10, 34. [Google Scholar] [CrossRef] [Green Version]
- Wang, L.J.; Li, S.H. Salicylic acid-induced heat or cold tolerance in relation to Ca2+ homeostasis and antioxidant systems in young grape plants. Plant Sci. 2006, 170, 685–694. [Google Scholar] [CrossRef]
- Jahan, M.S.; Wang, Y.; Shu, S.; Zhong, M.; Chen, Z.; Wu, J.; Sun, J.; Guo, S. Exogenous salicylic acid increases the heat tolerance in Tomato (Solanum lycopersicum L) by enhancing photosynthesis efficiency and improving antioxidant defense system through scavenging of reactive oxygen species. Sci. Hortic. 2019, 247, 421–429. [Google Scholar] [CrossRef]
- Khan, M.I.R.; Iqbal, N.; Masood, A.; Per, T.S.; Khan, N.A. Salicylic acid alleviates adverse effects of heat stress on photosynthesis through changes in proline production and ethylene formation. Plant Signal. Behav. 2013, 8, e26374. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cingoz, G.S.; Gurel, E. Effects of salicylic acid on thermotolerance and cardenolide accumulation under high temperature stress in Digitalis trojana Ivanina. Plant Physiol. Biochem. 2016, 105, 145–149. [Google Scholar] [CrossRef] [PubMed]
- Shen, H.; Zhao, B.; Xu, J.; Zheng, X.; Huang, W. Effects of salicylic acid and calcium chloride on heat tolerance in Rhododendron ‘Fen Zhen Zhu’. J. Am. Soc. Hortic. Sci. 2016, 141, 363–372. [Google Scholar] [CrossRef] [Green Version]
- Miura, K.; Tada, Y. Regulation of water, salinity, and cold stress responses by salicylic acid. Front. Plant Sci. 2014, 5, 4. [Google Scholar] [CrossRef] [Green Version]
- The Potato Genome Sequencing Consortium. Genome sequence and analysis of the tuber crop potato. Nature 2011, 475, 189–195. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pham, G.M.; Hamilton, J.P.; Wood, J.C.; Burke, J.T.; Zhao, H.; Vaillancourt, B.; Ou, S.; Jiang, J.; Buell, C.R. Construction of a chromosome-scale long-read reference genome assembly for potato. Gigascience 2020, 9, giaa100. [Google Scholar] [CrossRef] [PubMed]
- Hijmans, R.J. The effect of climate change on global potato production. Am. J. Potato Res. 2003, 80, 271–279. [Google Scholar] [CrossRef]
- Tian, F.; Yang, D.C.; Meng, Y.Q.; Jin, J.; Gao, G. PlantRegMap: Charting functional regulatory maps in plants. Nucleic Acids Res. 2020, 48, D1104–D1113. [Google Scholar] [CrossRef] [PubMed]
- Storey, J.D.; Tibshirani, R. Statistical significance for genomewide studies. Proc. Natl. Acad. Sci. USA 2003, 100, 9440–9445. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jin, J.; Tian, F.; Yang, D.C.; Meng, Y.Q.; Kong, L.; Luo, J.; Gao, G. PlantTFDB 4.0: Toward a central hub for transcription factors and regulatory interactions in plants. Nucleic Acids Res. 2017, 45, D1040–D1045. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lescot, M.; Déhais, P.; Thijs, G.; Marchal, K.; Moreau, Y.; Van de Peer, Y.; Rouzé, P.; Rombauts, S. PlantCARE, a database of plant cis-acting regulatory elements and a portal to tools for in silico analysis of promoter sequences. Nucleic Acids Res. 2002, 30, 325–327. [Google Scholar] [CrossRef] [PubMed]
- Gasteiger, E.; Hoogland, C.; Gattiker, A.; Duvaud, S.; Wilkins, M.R.; Appel, R.D.; Bairoch, A. Protein Identification and Analysis Tools on the ExPASy Server. In The Proteomics Protocols Handbook; Walker, J.M., Ed.; Humana Press: Totowa, NJ, USA, 2005; pp. 571–607. [Google Scholar] [CrossRef]
- Armenteros, J.J.A.; Sønderby, C.K.; Sønderby, S.K.; Nielsen, H.; Winther, O. DeepLoc: Prediction of protein subcellular localization using deep learning. Bioinformatics 2017, 33, 3387–3395. [Google Scholar] [CrossRef] [PubMed]
- Yu, C.S.; Chen, Y.C.; Lu, C.H.; Hwang, J.K. Prediction of protein subcellular localization. Proteins Struct. Funct. Bioinform. 2006, 64, 643–651. [Google Scholar] [CrossRef] [PubMed]
- Stäerk, H.; Dallago, C.; Heinzinger, M.; Rost, B. Light attention predicts protein location from the language of life. bioRxiv 2021. [Google Scholar] [CrossRef]
- Armenteros, J.J.A.; Salvatore, M.; Emanuelsson, O.; Winther, O.; Von Heijne, G.; Elofsson, A.; Nielsen, H. Detecting sequence signals in targeting peptides using deep learning. Life Sci. Alliance 2019, 2, e201900429. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Neuberger, G.; Maurer-Stroh, S.; Eisenhaber, B.; Hartig, A.; Eisenhaber, F. Motif refinement of the peroxisomal targeting signal 1 and evaluation of taxon-specific differences. J. Mol. Biol. 2003, 328, 567–579. [Google Scholar] [CrossRef]
- Neuberger, G.; Maurer-Stroh, S.; Eisenhaber, B.; Hartig, A.; Eisenhaber, F. Prediction of peroxisomal targeting signal 1 containing proteins from amino acid sequence. J. Mol. Biol. 2003, 328, 581–592. [Google Scholar] [CrossRef]
- Reumann, S.; Buchwald, D.; Lingner, T. PredPlantPTS1: A web server for the prediction of plant peroxisomal proteins. Front. Plant Sci. 2012, 3, 194. [Google Scholar] [CrossRef] [Green Version]
- Mistry, J.; Chuguransky, S.; Williams, L.; Qureshi, M.; Salazar, G.A.; Sonnhammer, E.L.; Tosatto, S.C.E.; Paladin, L.; Raj, S.; Richardson, L.J.; et al. Pfam: The protein families database in 2021. Nucleic Acids Res. 2021, 49, D412–D419. [Google Scholar] [CrossRef] [PubMed]
- Eddy, S.R. Accelerated profile HMM searches. PLoS Comput. Biol. 2011, 7, e1002195. [Google Scholar] [CrossRef] [Green Version]
- Lu, S.; Wang, J.; Chitsaz, F.; Derbyshire, M.K.; Geer, R.C.; Gonzales, N.R.; Gwadz, M.; Hurwitz, D.I.; Marchler, G.H.; Song, J.S.; et al. CDD/SPARCLE: The conserved domain database in 2020. Nucleic Acids Res. 2020, 48, D265–D268. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Marchler-Bauer, A.; Bryant, S.H. CD-Search: Protein domain annotations on the fly. Nucleic Acids Res. 2004, 32, W327–W331. [Google Scholar] [CrossRef]
- Wright, E.S. DECIPHER: Harnessing local sequence context to improve protein multiple sequence alignment. BMC Bioinform. 2015, 16, 322. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jumper, J.; Evans, R.; Pritzel, A.; Green, T.; Figurnov, M.; Ronneberger, O.; Tunyasuvunakool, K.; Bates, R.; Žídek, A.; Potapenko, A.; et al. Highly accurate protein structure prediction with AlphaFold. Nature 2021, 596, 583–589. [Google Scholar] [CrossRef]
- Pettersen, E.F.; Goddard, T.D.; Huang, C.C.; Meng, E.C.; Couch, G.S.; Croll, T.I.; Morris, J.H.; Ferrin, T.E. UCSF ChimeraX: Structure visualization for researchers, educators, and developers. Protein Sci. 2021, 30, 70–82. [Google Scholar] [CrossRef]
- Chen, V.B.; Arendall, W.B., III; Headd, J.J.; Keedy, D.A.; Immormino, R.M.; Kapral, G.J.; Murray, L.W.; Richardson, J.S.; Richardson, D.C. MolProbity: All-atom structure validation for macromolecular crystallography. Acta Cryst. 2010, D66, 1–21. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Waterhouse, A.; Bertoni, M.; Bienert, S.; Studer, G.; Tauriello, G.; Gumienny, R.; Heer, F.T.; de Beer, T.A.P.; Rempfer, C.; Bordoli, L.; et al. SWISS-MODEL: Homology modelling of protein structures and complexes. Nucleic Acids Res. 2018, 46, W296–W303. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dragićević, M. ggrama: Ramachandran Diagrams using ggplot2 Graphics. 2021. Available online: https://github.com/missuse/ggrama (accessed on 30 November 2021). [CrossRef]
- Yu, Y.K.; Capra, J.A.; Stojmirović, A.; Landsman, D.; Altschul, S.F. Log-odds sequence logos. Bioinformatics 2015, 31, 324–331. [Google Scholar] [CrossRef] [Green Version]
- Le, S.Q.; Gascuel, O. An improved general amino acid replacement matrix. Mol. Biol. Evol. 2008, 25, 1307–1320. [Google Scholar] [CrossRef] [Green Version]
- Schliep, K.P. phangorn: Phylogenetic analysis in R. Bioinformatics 2011, 27, 592–593. [Google Scholar] [CrossRef] [Green Version]
- Murashige, T.; Skoog, F. A revised medium for rapid growth and bio assays with tobacco tissue cultures. Physiol. Plant. 1962, 15, 473–497. [Google Scholar] [CrossRef]
- Linsmaier, E.M.; Skoog, F. Organic growth factor requirements of tobacco tissue cultures. Physiol. Plant. 1965, 18, 100–127. [Google Scholar] [CrossRef]
- López-Delgado, H.; Mora-Herrera, M.E.; Zavaleta-Mancera, H.A.; Cadena-Hinojosa, M.; Scott, I.M. Salicylic acid enhances heat tolerance and potato virus X (PVX) elimination during thermotherapy of potato microplants. Am. J. Potato Res. 2004, 81, 171–176. [Google Scholar] [CrossRef]
- Tiwari, J.K.; Buckseth, T.; Devi, S.; Varshney, S.; Sahu, S.; Patil, V.U.; Zinta, R.; Ali, N.; Moudgil, V.; Singh, R.K.; et al. Physiological and genome-wide RNA-sequencing analyses identify candidate genes in a nitrogen-use efficient potato cv. Kufri Gaurav. Plant Physiol. Biochem. 2020, 154, 171–183. [Google Scholar] [CrossRef]
- Klatte, M.; Bauer, P. Accurate real-time reverse transcription quantitative PCR. In Plant Signal Transduction; Pfannschmidt, T., Ed.; Human Press: Totowa, NJ, USA, 2009; Volume 479, pp. 61–77. [Google Scholar] [CrossRef]
- Cannon, S.B.; Mitra, A.; Baumgarten, A.; Young, N.D.; May, G. The roles of segmental and tandem gene duplication in the evolution of large gene families in Arabidopsis thaliana. BMC Plant Biol. 2004, 4, 10. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ikai, A. Thermostability and aliphatic index of globular proteins. J. Biochem. 1980, 88, 1895–1898. [Google Scholar] [CrossRef]
- Getzoff, E.D.; Tainer, J.A.; Stempien, M.M.; Bell, G.I.; Hallewell, R.A. Evolution of CuZn superoxide dismutase and the Greek key β-barrel structural motif. Proteins Struct. Funct. Genet. 1989, 5, 322–336. [Google Scholar] [CrossRef]
- Favier, A.; Gans, P.; Erba, E.B.; Signor, L.; Muthukumar, S.S.; Pfannschmidt, T.; Blanvillain, R.; Cobessi, D. The plastid-encoded RNA polymerase-associated protein PAP9 is a superoxide dismutase with unusual structural features. Front. Plant Sci. 2021, 12, 668897. [Google Scholar] [CrossRef]
- Quint, P.; Reutzel, R.; Mikulski, R.; McKenna, R.; Silverman, D.N. Crystal structure of nitrated human manganese superoxide dismutase: Mechanism of inactivation. Free Radic. Biol. Med. 2006, 40, 453–458. [Google Scholar] [CrossRef]
- Zhou, Y.; Hu, L.; Wu, H.; Jiang, L.; Liu, S. Genome-wide identification and transcriptional expression analysis of cucumber superoxide dismutase (SOD) family in response to various abiotic stresses. Int. J. Genom. 2017, 2017, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Furukawa, Y.; Torres, A.S.; O’Halloran, T.V. Oxygen-induced maturation of SOD1: A key role for disulfide formation by the copper chaperone CCS. EMBO J. 2004, 23, 2872–2881. [Google Scholar] [CrossRef] [PubMed]
- Feng, X.; Lai, Z.; Lin, Y.; Lai, G.; Lian, C. Genome-wide identification and characterization of the superoxide dismutase gene family in Musa acuminata cv. Tianbaojiao (AAA group). BMC Genom. 2015, 16, 823. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jiang, W.; Yang, L.; He, Y.; Zhang, H.; Li, W.; Chen, H.; Ma, D.; Yin, J. Genome-wide identification and transcriptional expression analysis of superoxide dismutase (SOD) family in wheat (Triticum aestivum). PeerJ 2019, 7, e8062. [Google Scholar] [CrossRef] [Green Version]
- Yadav, S.; Gill, S.S.; Passricha, N.; Gill, R.; Badhwar, P.; Anjum, N.A.; Francisco, J.B.J.; Tuteja, N. Genome-wide analysis and transcriptional expression pattern-assessment of superoxide dismutase (SOD) in rice and Arabidopsis under abiotic stresses. Plant Gene 2019, 17, 100165. [Google Scholar] [CrossRef]
- Filiz, E.; Tombuloğlu, H. Genome-wide distribution of superoxide dismutase (SOD) gene families in Sorghum bicolor. Turk. J. Biol. 2015, 39, 49–59. [Google Scholar] [CrossRef]
- Wang, W.; Xia, M.; Chen, J.; Deng, F.; Yuan, R.; Zhang, X.; Shen, F. Genome-wide analysis of superoxide dismutase gene family in Gossypium raimondii and G. arboreum. Plant Gene 2016, 6, 18–29. [Google Scholar] [CrossRef] [Green Version]
- Feng, K.; Yu, J.; Cheng, Y.; Ruan, M.; Wang, R.; Ye, Q.; Zhou, G.; Li, Z.; Yao, Z.; Yang, Y.; et al. The SOD gene family in tomato: Identification, phylogenetic relationships, and expression patterns. Front. Plant Sci. 2016, 7, 1279. [Google Scholar] [CrossRef] [Green Version]
- Xu, G.; Guo, C.; Shan, H.; Kong, H. Divergence of duplicate genes in exon–intron structure. Proc. Natl. Acad. Sci. USA 2012, 109, 1187–1192. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Srivastava, V.; Srivastava, M.K.; Chibani, K.; Nilsson, R.; Rouhier, N.; Melzer, M.; Wingsle, G. Alternative splicing studies of the reactive oxygen species gene network in Populus reveal two isoforms of high-isoelectric-point superoxide dismutase. Plant Physiol. 2009, 149, 1848–1859. [Google Scholar] [CrossRef] [Green Version]
- Feng, W.; Hongbin, W.; Bing, L.; Jinfa, W. Cloning and characterization of a novel splicing isoform of the iron-superoxide dismutase gene in rice (Oryza sativa L.). Plant Cell Rep. 2006, 24, 734–742. [Google Scholar] [CrossRef]
- Perry, J.J.P.; Shin, D.S.; Getzoff, E.D.; Tainer, J.A. The structural biochemistry of the superoxide dismutases. Biochim. Biophys. Acta-Proteins Proteom. 2010, 1804, 245–262. [Google Scholar] [CrossRef] [Green Version]
- Alscher, R.G.; Erturk, N.; Heath, L.S. Role of superoxide dismutases (SODs) in controlling oxidative stress in plants. J. Exp. Bot. 2002, 53, 1331–1341. [Google Scholar] [CrossRef] [PubMed]
- Momčilović, I.; Pantelić, D.; Hfidan, M.; Savić, J.; Vinterhalter, D. Improved procedure for detection of superoxide dismutase isoforms in potato, Solanum tuberosum L. Acta Physiol. Plant. 2014, 36, 2059–2066. [Google Scholar] [CrossRef]
- Yamakura, F.; Kawasaki, H. Post-translational modifications of superoxide dismutase. Biochim. Biophys. Acta Proteins Proteom. 2010, 1804, 318–325. [Google Scholar] [CrossRef] [PubMed]
- Reumann, S.; Chowdhary, G. Prediction of peroxisomal matrix proteins in plants. In Proteomics of Peroxisomes; del Río, L., Schrader, M., Eds.; Springer: Singapore, 2018; Volume 89, pp. 125–138. [Google Scholar] [CrossRef]
- Corpas, F.J.; González-Gordo, S.; Palma, J.M. Plant peroxisomes: A factory of reactive species. Front. Plant Sci. 2020, 11, 853. [Google Scholar] [CrossRef] [PubMed]
- Fink, R.C.; Scandalios, J.G. Molecular evolution and structure–function relationships of the superoxide dismutase gene families in angiosperms and their relationship to other eukaryotic and prokaryotic superoxide dismutases. Arch. Biochem. Biophys. 2002, 399, 19–36. [Google Scholar] [CrossRef] [PubMed]
- Sheng, Y.; Abreu, I.A.; Cabelli, D.E.; Maroney, M.J.; Miller, A.F.; Teixeira, M.; Valentine, J.S. Superoxide dismutases and superoxide reductases. Chem. Rev. 2014, 114, 3854–3918. [Google Scholar] [CrossRef] [PubMed]
- Retnoningrum, D.S.; Yoshida, H.; Razani, M.D.; Meidianto, V.F.; Hartanto, A.; Artarini, A.; Ismaya, W.T. Unprecedented Role of the N73-F124 Pair in the Staphylococcus equorum MnSOD Activity. Curr. Enzym. Inhib. 2021, 17, 2–8. [Google Scholar] [CrossRef]
- Cabrejos, D.A.L.; Alexandrino, A.V.; Pereira, C.M.; Mendonça, D.C.; Pereira, H.D.; Novo-Mansur, M.T.M.; Garratt, R.C.; Goto, L.S. Structural characterization of a pathogenicity-related superoxide dismutase codified by a probably essential gene in Xanthomonas citri subsp. citri. PLoS ONE 2019, 14, e0209988. [Google Scholar] [CrossRef]
- Sheng, Y.; Durazo, A.; Schumacher, M.; Gralla, E.B.; Cascio, D.; Cabelli, D.E.; Valentine, J.S. Tetramerization reinforces the dimer interface of MnSOD. PLoS ONE 2013, 8, e62446. [Google Scholar] [CrossRef] [Green Version]
- Su, Z.; Wang, J.; Yu, J.; Huang, X.; Gu, X. Evolution of alternative splicing after gene duplication. Genome Res. 2006, 16, 182–189. [Google Scholar] [CrossRef] [Green Version]
- Van Dam, J.; Kooman, P.L.; Struik, P.C. Effects of temperature and photoperiod on early growth and final number of tubers in potato (Solanum tuberosum L.). Potato Res. 1996, 39, 51–62. [Google Scholar] [CrossRef]
- Ahn, Y.J.; Claussen, K.; Zimmerman, J.L. Genotypic differences in the heat-shock response and thermotolerance in four potato cultivars. Plant Sci. 2004, 166, 901–911. [Google Scholar] [CrossRef]
- Tang, R.; Niu, S.; Zhang, G.; Chen, G.; Haroon, M.; Yang, Q.; Rajora, O.P.; Li, X.Q. Physiological and growth responses of potato cultivars to heat stress. Botany 2018, 96, 897–912. [Google Scholar] [CrossRef]
- Demirel, U.; Morris, W.L.; Ducreux, L.J.; Yavuz, C.; Asim, A.; Tindas, I.; Campbell, R.; Morris, J.A.; Verrall, S.R.; Hedlez, P.E.; et al. Physiological, biochemical, and transcriptional responses to single and combined abiotic stress in stress-tolerant and stress-sensitive potato genotypes. Front. Plant Sci. 2020, 11, 169. [Google Scholar] [CrossRef] [PubMed]
- Hayat, Q.; Hayat, S.; Irfan, M.; Ahmad, A. Effect of exogenous salicylic acid under changing environment: A review. Environ. Exp. Bot. 2010, 68, 14–25. [Google Scholar] [CrossRef]
- Herrera-Vásquez, A.; Salinas, P.; Holuigue, L. Salicylic acid and reactive oxygen species interplay in the transcriptional control of defense genes expression. Front. Plant Sci. 2015, 6, 171. [Google Scholar] [CrossRef] [Green Version]
- Li, Q.; Wang, G.; Wang, Y.; Yang, D.; Guan, C.; Ji, J. Foliar application of salicylic acid alleviate the cadmium toxicity by modulation the reactive oxygen species in potato. Ecotoxicol. Environ. Saf. 2019, 172, 317–325. [Google Scholar] [CrossRef] [PubMed]
- Antonić, D.; Milošević, S.; Cingel, A.; Lojić, M.; Trifunović-Momčilov, M.; Petrić, M.; Subotić, A.; Simonović, A. Effects of exogenous salicylic acid on Impatiens walleriana L. grown in vitro under polyethylene glycol-imposed drought. S. Afr. J. Bot. 2016, 105, 226–233. [Google Scholar] [CrossRef]
- Sharma, L.; Priya, M.; Kaushal, N.; Bhandhari, K.; Chaudhary, S.; Dhankher, O.P.; Prasad, P.V.V.; Siddique, K.H.M.; Nayyar, H. Plant growth-regulating molecules as thermoprotectants: Functional relevance and prospects for improving heat tolerance in food crops. J. Exp. Bot. 2020, 71, 569–594. [Google Scholar] [CrossRef] [PubMed]
- Guo, M.; Liu, J.H.; Ma, X.; Luo, D.X.; Gong, Z.H.; Lu, M.H. The plant heat stress transcription factors (HSFs): Structure, regulation, and function in response to abiotic stresses. Front. Plant Sci. 2016, 7, 114. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Zhao, Y.; Tian, X.; Wang, F.; Zhang, L.; Xin, M.; Hu, Z.; Yao, Y.; Ni, Z.; Sun, Q.; Peng, H. Characterization of wheat MYB genes responsive to high temperatures. BMC Plant Biol. 2017, 17, 208. [Google Scholar] [CrossRef] [PubMed]
- Islam, M.M.; Sandhu, J.; Walia, H.; Saha, R. Transcriptomic data-driven discovery of global regulatory features of rice seeds developing under heat stress. Comput. Struct. Biotechnol. J. 2020, 18, 2556–2567. [Google Scholar] [CrossRef] [PubMed]
- Waqas, M.; Shahid, L.; Shoukat, K.; Aslam, U.; Azeem, F.; Atif, R.M. Role of DNA-binding with one finger (Dof) transcription factors for abiotic stress tolerance in plants. In Transcription Factors for Abiotic Stress Tolerance in Plants; Wani, S.H., Ed.; Academic Press: Cambridge, MA, USA, 2020; pp. 1–14. [Google Scholar] [CrossRef]
- Charfeddine, M.; Saïdi, M.N.; Charfeddine, S.; Hammami, A.; Bouzid, R.G. Genome-wide analysis and expression profiling of the ERF transcription factor family in potato (Solanum tuberosum L.). Mol. Biotechnol. 2015, 57, 348–358. [Google Scholar] [CrossRef]
- Sunkar, R.; Kapoor, A.; Zhu, J.K. Posttranscriptional induction of two Cu/Zn superoxide dismutase genes in Arabidopsis is mediated by downregulation of miR398 and important for oxidative stress tolerance. Plant Cell 2006, 18, 2051–2065. [Google Scholar] [CrossRef] [Green Version]
- Cohu, C.M.; Pilon, M. Regulation of superoxide dismutase expression by copper availability. Physiol. Plant. 2007, 129, 747–755. [Google Scholar] [CrossRef]

| Proposed Name | Gene ID | Chromosomal Localization | Transcript ID | Transcript Length (bp) | Protein Length (aa) | Isoelectric Point (pI) | Instability Index | Aliphatic Index | Molecular Weight (kDa) |
|---|---|---|---|---|---|---|---|---|---|
| StCuZnSOD1 | Soltu.DM.01G022650 | Chr.01: 61,097,981-61,103,305 | Soltu.DM.01G022650.1 * | 459 | 152 | 5.28 | 32.24 | 80.20 | 15.3 |
| StCuZnSOD2 | Soltu.DM.11G020830 | Chr.11: 40,784,322-40,790,930 Chr.11: 40,784,322-40,790,564 | Soltu.DM.11G020830.1 * Soltu.DM.11G020830.2 | 648 645 | 215 214 | 6.34 6.34 | 24.76 24.25 | 87.95 88.36 | 22.1 22.1 |
| StCuZnSOD3 | Soltu.DM.03G010200 | Chr.03: 28,151,770-28,156,700 | Soltu.DM.03G010200.1 * | 498 | 165 | 6.78 | 18.64 | 88.61 | 16.8 |
| StMnSOD | Soltu.DM.06G011380 | Chr.06: 34,521,324-34,524,092 | Soltu.DM.06G011380.1 * | 687 | 228 | 7.13 | 35.60 | 91.14 | 25.3 |
| StFeSOD1 | Soltu.DM.06G012180 | Chr.06: 36,014,174-36,016,425 Chr.06: 36,014,151-36,016,425 Chr.06: 36,014,174-36,016,425 Chr.06: 36,014,174-36,016,425 Chr.06: 36,014,174-36,016,425 Chr.06: 36,014,174-36,016,425 | Soltu.DM.06G012180.1 Soltu.DM.06G012180.2 Soltu.DM.06G012180.3 * Soltu.DM.06G012180.4 Soltu.DM.06G012180.5 Soltu.DM.06G012180.6 | 747 753 759 729 681 654 | 248 250 252 242 226 217 | 6.31 6.60 6.60 7.77 5.96 6.52 | 33.40 33.86 34.62 35.05 35.28 35.44 | 70.56 70.00 70.99 75.08 74.38 70.74 | 27.8 28.1 28.3 26.9 25.1 24.1 |
| StFeSOD2 | Soltu.DM.03G013800 | Chr.03: 36,413,200-36,416,935 Chr.03: 36,413,200-36,416,935 Chr.03: 36,413,200-36,416,935 Chr.03: 36,413,200-36,416,935 Chr.03: 36,413,200-36,416,935 | Soltu.DM.03G013800.1 Soltu.DM.03G013800.2 Soltu.DM.03G013800.3 Soltu.DM.03G013800.4 * Soltu.DM.03G013800.5 | 915 891 897 987 840 | 304 296 298 328 279 | 5.56 5.49 5.83 5.69 5.20 | 41.47 41.99 40.95 40.11 42.01 | 73.26 73.58 72.11 76.19 73.87 | 34.7 33.7 33.8 37.6 31.9 |
| StFeSOD3 | Soltu.DM.02G001300 | Chr.02: 7,378,597-7,382,073 Chr.02: 7,378,597-7,382,073 | Soltu.DM.02G001300.1 * Soltu.DM.02G001300.2 | 777 753 | 258 250 | 6.07 5.96 | 42.61 43.46 | 85.04 85.80 | 29.6 28.5 |
| StFeSOD4 | Soltu.DM.06G012170 | Chr.06: 36,008,330-36,012,608 Chr.06: 36,008,330-36,012,608 Chr.06: 36,008,330-36,012,608 Chr.06: 36,008,330-36,012,608 Chr.06: 36,008,330-36,012,608 | Soltu.DM.06G012170.1 Soltu.DM.06G012170.2 Soltu.DM.06G012170.3 Soltu.DM.06G012170.4 * Soltu.DM.06G012170.5 | 603 540 549 684 513 | 200 179 182 227 170 | 5.58 5.81 6.29 7.85 6.12 | 31.53 33.93 30.48 39.93 35.64 | 82.00 78.60 76.26 73.17 74.18 | 22.5 20.2 20.4 25.6 19.1 |
| Chr.06: 36,008,330-36,012,608 | Soltu.DM.06G012170.6 | 633 | 210 | 5.89 | 31.93 | 77.71 | 23.8 |
| Gene | Protein ID | TargetP2 | DeepLoc | CELLO | LA | PTS1 Predictor | PredPlant PTS1 |
|---|---|---|---|---|---|---|---|
| StCuZnSOD1 | Soltu.DM.01G022650.1 | O | C | C | C | ||
| StCuZnSOD2 | Soltu.DM.11G020830.1 Soltu.DM.11G020830.2 | Ch Ch | Ch Ch | Ch Ch | Ch Ch | ||
| StCuZnSOD3 | Soltu.DM.03G010200.1 | O | P | C | P | Not-P | Not-P |
| StMnSOD | Soltu.DM.06G011380.1 | Mt | Mt | Mt | Mt | ||
| StFeSOD1 | Soltu.DM.06G012180.1 Soltu.DM.06G012180.2 Soltu.DM.06G012180.3 Soltu.DM.06G012180.4 Soltu.DM.06G012180.5 Soltu.DM.06G012180.6 | Ch Ch Ch Ch Ch Ch | Ch Ch Ch Ch Ch Ch | Ch Ch Ch Ch Ch Ch | Ch Ch Ch Ch Ch Ch | ||
| StFeSOD2 | Soltu.DM.03G013800.1 Soltu.DM.03G013800.2 Soltu.DM.03G013800.3 Soltu.DM.03G013800.4 Soltu.DM.03G013800.5 | Ch Ch Ch Ch Ch | Ch Ch Ch Ch Ch | Ch Ch Ch Ch, C Ch | Ch Ch Ch Ch Mt | ||
| StFeSOD3 | Soltu.DM.02G001300.1 Soltu.DM.02G001300.2 | Mt Mt | Ch Ch | Ch Ch | Mt Mt | ||
| StFeSOD4 | Soltu.DM.06G012170.1 Soltu.DM.06G012170.2 Soltu.DM.06G012170.3 Soltu.DM.06G012170.4 Soltu.DM.06G012170.5 | O O O O O | Mt Mt Mt Mt Mt | Ch, C Ch, C Ch, C, Mt N, Mt Ch, C, Mt | Mt C C Mt Mt | ||
| Soltu.DM.06G012170.6 | O | Mt | Ch | C |
| Protein ID | Ramachandran Favored | Ramachandran Outliers | Rotamer Outliers | Clashscore |
|---|---|---|---|---|
| Soltu.DM.01G022650.1 | 98.67% | 0.00% | 0.00% | 0.95 |
| Soltu.DM.11G020830.1 | 98.03% | 0.00% | 0.00% | 0.00 |
| Soltu.DM.03G010200.1 | 92.02% | 2.45% | 0.00% | 0.00 |
| Soltu.DM.06G011380.1 | 98.51% | 0.00% | 0.00% | 0.95 |
| Soltu.DM.06G012180.3 | 98.01% | 0.00% | 0.00% | 0.93 |
| Soltu.DM.03G013800.1 | 97.24% | 0.39% | 0.91% | 0.49 |
| Soltu.DM.02G001300.1 | 95.83% | 0.00% | 0.00% | 2.58 |
| Soltu.DM.06G012170.6 | 97.60% | 0.48% | 0.57% | 2.11 |
| Species | Genome Size (Mbp) | CuZnSOD | MnSOD | FeSOD | Total | Chr. num. | Introns num. | References |
|---|---|---|---|---|---|---|---|---|
| Solanum tuberosum | 844 | 3 | 1 | 4 | 8 | 5 | 4–9 | Current work |
| Arabidopsis thaliana | 125 | 3 | 2 | 3 | 8 | 5 | 5–8 | [9,65] |
| Oryza sativa | 389 | 4 | 1 | 2 | 7 | 6 | 5–9 | [9,65] |
| Sorghum bicolor | 730 | 5 * | 1 | 2 | 8 | 6 | 5–7 | [66] |
| Gossypium raimondii | 885 | 5 | 2 | 2 | 9 | 6+ | 4–8 | [67] |
| Gossypium arboreum | 1746 | 5 | 2 | 2 | 9 | 6 | 5–8 | [67] |
| Cucumis sativus | 367 | 5 | 1 | 3 | 9 | 5+ | 3–8 | [61] |
| Musa acuminate | 523 | 6 | 4 | 2 | 12 | 8 | 5–8 | [63] |
| Solanum lycopersicum | 828 | 4 * | 1 | 4 | 9 | 6 | 4–8 | [68] |
| Triticum aestivum | ~17,000 | 17 | 3 | 6 | 26 | 3 × 3 | 4–7 | [64] |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Rudić, J.; Dragićević, M.B.; Momčilović, I.; Simonović, A.D.; Pantelić, D. In Silico Study of Superoxide Dismutase Gene Family in Potato and Effects of Elevated Temperature and Salicylic Acid on Gene Expression. Antioxidants 2022, 11, 488. https://doi.org/10.3390/antiox11030488
Rudić J, Dragićević MB, Momčilović I, Simonović AD, Pantelić D. In Silico Study of Superoxide Dismutase Gene Family in Potato and Effects of Elevated Temperature and Salicylic Acid on Gene Expression. Antioxidants. 2022; 11(3):488. https://doi.org/10.3390/antiox11030488
Chicago/Turabian StyleRudić, Jelena, Milan B. Dragićević, Ivana Momčilović, Ana D. Simonović, and Danijel Pantelić. 2022. "In Silico Study of Superoxide Dismutase Gene Family in Potato and Effects of Elevated Temperature and Salicylic Acid on Gene Expression" Antioxidants 11, no. 3: 488. https://doi.org/10.3390/antiox11030488
APA StyleRudić, J., Dragićević, M. B., Momčilović, I., Simonović, A. D., & Pantelić, D. (2022). In Silico Study of Superoxide Dismutase Gene Family in Potato and Effects of Elevated Temperature and Salicylic Acid on Gene Expression. Antioxidants, 11(3), 488. https://doi.org/10.3390/antiox11030488










